Abstract
Coronary artery disease, myocardial infarction, and secondary damages of the myocardium in the form of ischemic heart disease remain major causes of death in Western countries. Beyond traditional risk factors such as smoking, hypertension, dyslipidemia, or diabetes, a positive family history is known to increase risk. The genetic factors underlying this observation remained unknown for decades until genetic studies were able to identify multiple genomic loci contributing to the heritability of the trait. Knowledge of the affected genes and the resulting molecular and cellular mechanisms leads to improved understanding of the pathophysiology leading to coronary atherosclerosis. Major goals are also to improve prevention and therapy of coronary artery disease and its sequelae via improved risk prediction tools and pharmacological targets. In this chapter, we recapitulate recent major findings. We focus on established novel targets and discuss possible further targets which are currently explored in translational studies.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Coronary artery disease (CAD) and myocardial infarction (MI) are the main causes of morbidity and mortality. The identification of risk factors is a prerequisite to improve prevention and therapy of the disease via gaining knowledge about the underlying pathophysiological processes as well as the identification of therapeutic targets. Hyperlipidemia, hypertension, and diabetes mellitus are examples for risk factors which can be treated via pharmacological intervention, whereas smoking and obesity can be addressed by lifestyle interventions. Age and male gender, which are also major risk factors, cannot be addressed therapeutically. In the past, a positive family history has also been regarded as a non-modifiable risk factor. However, the underlying risk factors were not known for decades. In this chapter, we summarize the developments in the past years which led to the identification of a plethora of genomic loci which are associated with CAD with high statistical certainty.
2 Methodological Aspects
Over the past decades, the methodological spectrum has been vastly expanded to identify novel genetic risk factors of CAD and MI. Details on these methods have been discussed elsewhere (Kessler et al. 2016). In this section, we aim to briefly mention some important points.
2.1 Techniques
The development of arrays with hundred thousands of common single nucleotide polymorphisms (SNPs) distributed all over the genome of an individual enabled researchers to deeply genotype large numbers of individuals. An important prerequisite was knowledge of the human genome which has been mainly gained via projects as the human genome project (Lander et al. 2001; Sachidanandam et al. 2001). Subsequent projects as the 1000 Genomes Project (1000 Genomes Project Consortium et al. 2012, 2015) raised further possibilities with the imputation of SNPs which had not been genotyped directly. As a consequence genome-wide association studies investigating millions of SNPs have been performed, and mostly common variants were found to be associated with the disease and lend support to the common disease-common variant hypothesis (Reich and Lander 2001). Thereby, since 2007, mostly genome-wide association studies and subsequent analyses led to the identification of genetic risk variants for various traits including CAD and traditional risk factors as blood pressure or lipids. Subsequently, large international consortia were formed that were responsible for the identification of most of the currently known CAD risk loci. Figure 1 illustrates the evolution of genome-wide association studies focusing on the CARDIoGRAMplusC4D consortium, a coalition of several individual studies to facilitate the identification of CAD risk factors (Schunkert et al. 2019) and further studies using data from UK Biobank (Littlejohns et al. 2019). The idea of exome-wide association studies is comparable to that of a genome-wide association study with the exception that the investigated SNPs are enriched for coding variants distributed over the genome (Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia Investigators et al. 2016). Most of the currently known genetic CAD risk factors have been identified using these methods (Kessler et al. 2016; Khera and Kathiresan 2017a; Erdmann et al. 2018).
Discovery of genomic variants associated with CAD/MI using genome-/exome-wide association studies (modified after Kessler et al. 2016). (a) Over the past years, ongoing research led to the identification of increasing numbers of SNPs which are associated with CAD/MI at the genome-wide level of significance. Results from the CARDIoGRAMplusC4D-Million Hearts Initiative are awaited for 2019/2020. (b) The number of investigated individuals correlates with the number of identified risk variants which highlights the importance of international collaborations
Apart from common variants, the association of rare variants and private mutations has also been extensively studied. To identify such genetic variation, mostly exome sequencing has been used in the past (Erdmann et al. 2013). As the name says, this next-generation sequencing method is able to determine the genomic sequence of an individual enriched for coding regions and thus renders the identification of missense variants co-segregating with a certain disease in, e.g., a family with high prevalence possible. Whole-genome sequencing is not restricted to coding sequences. However, due to costs and computational challenges of the different methods, whole-genome sequencing has not yet contributed to the current knowledge of genetic variation in CAD.
2.2 Selection of Individuals
Most large-scale genomic studies use a case-control design, i.e., a cohort of patients suffering from the trait of interest is compared to a group of healthy individuals free of the disease. Obviously, clear definitions of phenotypes are important to gain reliable results. Most GWAS on CAD/MI focused on cases suffering from the disease at an early age. This increased the possibility of identification of the genetic predisposition which, together with gender, still represents the strongest risk factor. For controls, population-based samples as well as individuals which have been diagnosed to not to suffer from CAD/MI have been included. An important resource that has increasingly been used in recent projects is the UK Biobank (https://www.ukbiobank.ac.uk/). It includes health information as well as genetic data for 500,000 individuals which can be accessed by researchers after approval of specific projects (Littlejohns et al. 2019). In the USA, the Million Veteran Project has formed and stated to provide likewise genome-wide data (Klarin et al. 2018).
2.3 Statistical Analysis
To reduce the possibility of a false-positive finding, rigorous statistical thresholds have been determined to identify a certain variant to be associated with a trait as CAD/MI. Currently, a p-value below 5 × 10−8 is commonly accepted as a genome-wide significant finding. The strict p-value is a result of the large number of statistical tests which are performed in parallel on one dataset in a genome-wide association study (Pe’er et al. 2008). However, it cannot be excluded that true positive signals are discarded because they do not reach the stringent p-value. Recent studies also reported loci below certain false discovery rate thresholds to address this issue.
2.4 Mendelian Randomization Studies
Mendelian randomization studies (for an overview see Jansen et al. 2014) enable to investigate whether a biomarker or a trait is causal in the development of a disease. In epidemiological research, associations between biomarkers and diseases are frequently observed. However, the association can be influenced by unknown factors. On the other hand, the investigated disease itself can influence a given biomarker. In Mendelian randomization studies, causality between a biomarker and development of a disease can be assumed if a genetic variant influences this biomarker in the same direction as the biomarker is associated with the disease. Important examples in CAD are biomarkers such as LDL-cholesterol or C-reactive protein as well as behavioral traits such as educational attainment (Zeng et al. 2019a) which are discussed in more detail below.
3 Genetic Risk Factors Associated with Coronary Artery Disease
3.1 Genome-Wide Association Studies
The discovery of genetic CAD/MI risk factors was launched in 2007 by three independent studies which reported the chromosome 9p21 locus as the first and, until today, strongest genetic risk locus (Helgadottir et al. 2007; McPherson et al. 2007; Samani et al. 2007). Since then, with the inclusion of more individuals and the possibility of analyzing a larger number of SNPs, more and more variants associated with CAD/MI have been reported (for an overview see Kessler et al. (2016), Khera and Kathiresan (2017b), and Erdmann et al. (2018)). The currently known loci are depicted in Table 1. Surprisingly, most of the SNPs tag genes which have not been studied in the pathophysiology of coronary atherosclerosis before. Another surprising finding was that only the minority of genes is also associated with traditional risk factors as hypertension or lipid metabolism rendering the involvement of other cellular mechanisms likely. Additionally, almost every lead SNP is located in a non-coding region of the genome. The most prominent example is again the chromosome 9p21 locus. Here, it is still unclear which gene might be responsible for the strong signal. Rather, there is much evidence that the effect might be mediated by the circular non-coding RNA ANRIL (Holdt et al. 2010, 2013, 2016). Overall, it has been estimated that the 163 so far known CAD risk loci explain between 30% and 40% of CAD heritability (Nelson et al. 2017). It has also been found that a large number of the reported loci also harbor multiple independent signals (van der Harst and Verweij 2018). Additionally, most of the loci show pleiotropy, i.e., a risk variant is associated with several phenotypes. About half of the currently known CAD risk loci have been reported to be associated with other traits (Webb et al. 2017).
The studies which led to the discovery of the currently known CAD/MI risk variants as well as studies which gave first insights into their involvement in CAD/MI are referenced in Table 1. Specific genes and pathways are further discussed below.
3.2 Exome-Wide Association Studies
As mentioned above, GWAS mainly led to the identification of non-coding variation associated with the disease. To specifically investigate the role of coding variants, dedicated arrays enriched for exonic variants were used. In an international effort, the Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia Investigators performed the largest exome-wide association study so far (Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia Investigators et al. 2016). The results are depicted in Table 2. In summary, only four variants were identified. First, it could be replicated that coding variation in the LPA and PCSK9 genes is associated with CAD/MI. Second, two novel coding variants in ANGPTL4 and SVEP1 were found. Whereas LPA, PCSK9, and ANGPTL4 are associated with lipid metabolism (see below), SVEP1 only displayed association with blood pressure (Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia Investigators et al. 2016). However, this effect cannot fully explain the association signal for CAD/MI.
3.3 Exome and Whole-Genome Sequencing
As discussed above, due to still comparatively high costs, whole-genome sequencing has not yet significantly contributed to the knowledge of genetic CAD risk factors. In contrast, several studies made use of exome sequencing to identify variation in the coding sequence which is associated with CAD/MI. One scope of application the investigation of members of families with a high disease prevalence. Using this approach and subsequent co-segregation analyses genetic factors underlying the development of CAD/MI or its risk factors was possible. An important example is a mutation in the GUCY1A3 gene, which has been – together with a coding mutation in the CCT7 gene – shown to be responsible for the phenotype of premature CAD/MI in a family (Erdmann et al. 2013). As shown in Table 1, the GUCY1A3 locus also harbors common, non-coding variants associated with CAD/MI. Thus, an allelic series has been shown at the locus with a mutation and common variants which lead to a strong or only moderate risk increase, respectively. Further loci have been identified by analyzing large cohorts of cases and controls comparable to the GWAS approach. Most of the genes that were thereby identified play a role in lipid metabolism. Some of the genes, e.g., LDLR or PCSK9, also demonstrate allelic series. A selection of genes which have been identified using exome sequencing is depicted in Table 3 and discussed below in more detail.
4 Genetic Overlap and Demarcation with Other Atherosclerotic Diseases
An increased prevalence of CAD risk alleles can be traced in a large number of cardiovascular conditions including heart failure, peripheral arterial disease, or atrial fibrillation (Ntalla et al. 2019). Large-scale genomic studies have also been published in particular for stroke (for a review see Dichgans et al. (2019)). Despite there is genetic overlap between CAD and atherosclerotic stroke (Dichgans et al. 2014; Kessler et al. 2015a), the genetic risk factors of CAD and stroke are not similar. The formation of large, international consortia has advanced the identification of risk genes in the fields of CAD and stroke genetics. In other atherosclerotic diseases as peripheral artery disease, genetic research will be facilitated by publicly available data from large-scale biobanks. Nevertheless, the identification of common genetic risk factors will be important to evaluate novel therapeutic strategies. Other genetic risk factors for CAD might also not play an important role in related diseases: as such, the strongest genetic risk factor reported so far, chromosome 9p21, is not associated with calcified aortic stenosis; in contrast, the LPA gene is associated with both diseases (Trenkwalder et al. 2018).
5 Pharmacological Targets Identified by Genomic Studies
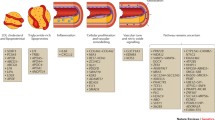
Genomic studies identified putative pharmacological targets at every stage of plaque formation, progression, and rupture (Fig. 2) which are discussed in this section.
5.1 Lipid Metabolism
While less than half of the identified variants tag genes which are associated with traditional risk factors, lipid metabolism represents a cluster of such variants.
LDL-Cholesterol Metabolism
High levels of LDL-cholesterol are an established risk factors for CAD. Not surprisingly, the most prominent genes which are directly involved in LDL-cholesterol metabolism and have been tagged by GWAS are LDLR, PCSK9, and SORT1. In line, mutations in the LDLR gene leading to reduced hepatic uptake of LDL-cholesterol have been shown to underlie familial hypercholesterolemia which itself increases CAD risk (Tolleshaug et al. 1983; Brown and Goldstein 1986), and common variants at the LDLR locus have been associated with both phenotypes (Myocardial Infarction Genetics Consortium et al. 2009; Teslovich et al. 2010; Schunkert et al. 2011; Do et al. 2015). PCSK9 has been identified as an interaction partner of the LDL-receptor. If PCSK9 binds to the LDL-receptor, it is internalized and degraded resulting in reduced hepatic uptake and high plasma levels of LDL-cholesterol (Cameron et al. 2006). Gain-of-function mutations in PCSK9 have been shown to increase LDL-cholesterol (Abifadel et al. 2003), whereas loss-of-function variants have been shown to reduce LDL-cholesterol and CAD risk (Cohen et al. 2006). PCSK9 has been targeted pharmacologically via different approaches. The inhibition of PCSK9 using, e.g., neutralizing antibodies resulted in a marked reduction of LDL-cholesterol and reduced risk of cardiovascular events in CAD patients (Sabatine et al. 2015, 2017). Sortilin 1 encoded by SORT1 has also been shown to affect LDL-cholesterol (Samani et al. 2007; Musunuru et al. 2010; Teslovich et al. 2010; Schunkert et al. 2011). However, as it seems to also play a role in other processes and as the mechanism involving sortilin 1 in LDL-cholesterol metabolism has still not been fully understood, it might not represent an ideal drug target. NPC1L1, a membrane transporter leading to the uptake of cholesterol from the intestine encoded by the NPC1L1 gene, also revealed a signal. Using exome sequencing, loss-of-function mutations were identified to be associated with reduced LDL-cholesterol and protection from CAD (Myocardial Infarction Genetics Consortium Investigators et al. 2014). In parallel, it has been shown that pharmacological targeting of NPC1L1 with ezetimibe also reduces the incidence of cardiovascular events in addition to a statin (Cannon et al. 2015). Figure 3 summarizes the multiple lines of evidence for an involvement of LDL-cholesterol metabolism in CAD/MI from a genetic point of view.
Multiple lines of genetic evidence for an involvement of LDL-cholesterol metabolism in CAD/MI. 1. Pharmacological treatments targeting genes which have been associated with LDL-cholesterol reduce risk of cardiovascular events. 2. Mendelian randomization studies have proven the causal influence of LDL-cholesterol metabolism in CAD/MI. 3. CAD/MI and LDL-cholesterol show shared genetic risk variants which are associated with both traits at the genome-wide level of statistical significance. 4. Genetic risk scores for LDL-cholesterol including genome-wide significant variants are also associated with CAD/MI risk, i.e., individuals with a high genetic LDL-cholesterol risk score are also at higher risk for CAD/MI and vice versa. Contains modified image material available at Servier Medical Art under a Creative Commons Attribution 3.0 Unported License
Triglyceride Metabolism
Lipoprotein lipase has been identified as a central enzyme regulating triglyceride levels. In line, genetic variants in LPL have been associated with both triglyceride levels and CAD risk (Khera et al. 2017). Additionally, the genes encoding several modulators of lipoprotein lipase activity were found to be associated with CAD/MI risk: (1) apolipoprotein C-III reduces lipoprotein lipase activity, and mutations in APOC3 are associated with both increased triglyceride levels and CAD risk (The TG and HDL Working Group of the Exome Sequencing Project, National Heart, Lung, and Blood Institute 2014; Klarin et al. 2018); (2) apolipoprotein A-V increases lipoprotein lipase activity, and mutations in APOA5 increase triglyceride levels and CAD risk (Do et al. 2015); and (3) angiopoietin-like 4 inhibits lipoprotein lipase activity, and mutations in ANGPTL4 reduce triglyceride levels and CAD risk (Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia Investigators et al. 2016). Angiopoietin-like 3 seems to have comparable effects as deficiency was also associated with reduced triglycerides and CAD risk (Stitziel et al. 2017). Mainly apolipoprotein C-III but also angiopoietin-like 3 are subjects of research for novel strategies targeting lipoprotein lipase metabolism (Graham et al. 2013; Gaudet et al. 2015; Ahmad et al. 2019).
5.2 Inflammation
Vascular inflammation and the recruitment of leukocytes are the hallmarks of atherosclerosis (for reviews see Lusis (2000), Libby et al. (2011), and Swirski and Nahrendorf (2013)). Several clinical studies already in the past decades thus targeted inflammatory parameters. A prominent example is C-reactive protein (CRP), which has been regarded for a long time as a pathophysiological player. This was mainly due to the observation of elevated CRP levels in individuals suffering from CAD in epidemiological studies (Koenig et al. 1999; Danesh et al. 2004) making CRP a promising therapeutic target (Pepys et al. 2006). Indeed, clinical trials, e.g., the JUPITER trial which investigated the effect of rosuvastatin in individuals with elevated CRP levels, showed a benefit (Ridker et al. 2008). Genetic studies were, however, able to dissect this association and trial outcomes. As such, LDL-cholesterol could be clearly proven as a causal risk factor for CAD as a genetic risk score for LDL-cholesterol elevating variants was also associated with CAD risk (Kathiresan et al. 2008). In contrast, several variants found to increase CRP levels did not show an association with CAD risk (Lange et al. 2006; Zacho et al. 2008; Linsel-Nitschke et al. 2008; Schunkert and Samani 2008; C Reactive Protein Coronary Heart Disease Genetics Collaboration (CCGC) et al. 2011). While these studies on CRP remained disappointing, targeting inflammation has indeed lately been proven to be effective in CAD. In the CANTOS trial, the administration of an interleukin-1β neutralizing antibody was able to reduce the incidence of cardiovascular events ultimately proving the inflammation hypothesis (Ridker et al. 2017). This is in line with Mendelian randomization studies that revealed evidence for a causal role of interleukin-6/interleukin-6 receptor signaling, which is downstream of interleukin-1β, in CAD (Interleukin-6 Receptor Mendelian Randomisation Analysis Consortium et al. 2012).
GWAS led to the identification of a number of variants tagging genes which play a role in inflammatory processes to be associated with CAD/MI.
Autoimmune Processes
CAD/MI and autoimmune disease share genetic risk factors. As such, the SH2B3 locus has been identified to be associated with CAD/MI (Coronary Artery Disease C4D Genetics Consortium 2011b; Schunkert et al. 2011) but also, e.g., type 1 diabetes mellitus (Barrett et al. 2009) and celiac disease (Hunt et al. 2008). A role for the encoded protein, SH2B adaptor protein 3 (SH2B3), has been shown in post MI remodeling: lack of Sh2b3 in rats was associated with increased fibrosis, increased leukocyte infiltration, and decreased cardiac function (Flister et al. 2015). Additionally, it is involved in dendritic cell function leading to T-cell interferon signaling (Mori et al. 2014). Nevertheless, uncertainties remain as the locus was also found to be associated with blood pressure (Levy et al. 2009; Newton-Cheh et al. 2009) and hematologic parameters (Soranzo et al. 2009). The latter observation might be important as an involvement in megakaryopoiesis and stabilization of thrombi has also been shown experimentally (Tong and Lodish 2004; Takizawa et al. 2010). In a recent study, the SH2B3 risk allele was found to be associated with decreased SH2B3 expression which, at least for the murine counterpart Sh2b3/Lnk, together with cholesterol loading promoted platelet production and activation (Wang et al. 2016).
Innate Immunity
CXCL12 is a chemokine which is encoded by the CAD risk gene CXCL12 (Myocardial Infarction Genetics Consortium et al. 2009; Coronary Artery Disease C4D Genetics Consortium 2011b) and has a function in various cellular processes. In atherosclerosis the CXCL12/CXCR4 axis has initially been described to have a rather protective role (Döring et al. 2014). Disruption of this axis led to increased plaque formation and proinflammatory plaque phenotypes in vivo (Zernecke et al. 2008). In line, CXCL12/CXCR4 signaling was shown to be important in endothelial cells and smooth muscle cells via maintaining barrier function and contractile responses, respectively (Döring et al. 2017). However, the interaction of CXCL12 and CXCR4 in atherosclerosis seems to be more complex. Whereas the atheroprotective findings were mainly derived from studies in which the interaction was inhibited by small molecules or genetic deletion of CXCR4, endothelial cell-derived CXCL12 was found to promote atherosclerosis (Döring et al. 2019). CXCR4 in contrast seems to be atheroprotective: in addition to the mentioned in vivo studies, the rs2322864 C-allele, which was found to be associated with CAD risk in a candidate gene study, was also associated with CXCR4 expression in plaques (Döring et al. 2017). The double-edged effects of CXCL12/CXCR4 signaling need to be considered to successfully target this pathway.
5.3 Platelet Function and Nitric Oxide Signaling
Genetic studies led to the identification of several genes which play a role in nitric oxide (NO) signaling leading to the formation or degradation of the second messenger cyclic guanosine monophosphate (cGMP), an endogenous inhibitor of platelet aggregation (Moro et al. 1996). Whereas several genes (e.g., NOS3, PDE5A, MRVI1, PDE3A) fulfill important functions in this pathway, most is known about GUCY1A3 which encodes the α1-subunit of the soluble guanylyl cyclase (α1-sGC). As discussed above, a digenic mutation in GUCY1A3 and the CCT7 gene, which encodes the chaperone protein, was identified in a family by exome sequencing (Erdmann et al. 2013). At the same time, also a common non-coding variant (rs7678555) was identified in Europeans to be associated with CAD by GWAS (CARDIoGRAMplusC4D Consortium et al. 2013). Whereas the digenic mutation led to loss of α1-sGC due to premature stop of translation (Erdmann et al. 2013), the common variant has been shown to influence GUCY1A3 expression, i.e., the risk allele G was linked to reduced expression (Kessler et al. 2017). Both the digenic mutation and the common risk variant led to reduced cGMP formation in platelets (Erdmann et al. 2013; Kessler et al. 2017). As a consequence, platelets of carriers of the common risk variants showed impaired inhibition of platelet aggregation secondary to NO stimulation (Kessler et al. 2017). The sGC is a known pharmacological target. Specific stimulators and activators of the sGC are available (for an overview see Stasch et al. (2011)) and approved for other traits, e.g., pulmonary hypertension (Ghofrani et al. 2013a, b). First preclinical data also render a positive influence of sGC stimulators on atherosclerotic phenotypes possible (Tsou et al. 2014). Additionally, the impaired response to nitric oxide in carriers of the common risk allele G in platelets might also be targeted via unspecific inhibitors of platelet aggregation. Recently, we showed that homozygous risk allele carriers might benefit from aspirin treatment in the primary prevention of cardiovascular diseases, whereas homozygous or heterozygous carriers of the non-risk allele A seem to even display increased risk (Hall et al. 2019). Additionally, homozygous carriers of the risk allele G are at an increased risk of ischemic events after the implantation of coronary stents, at least in part via higher on-aspirin platelet reactivity (Kessler et al. 2019). While platelets, in addition to their well-known role in atherothrombosis, are also involved in atherosclerotic plaque formation (Gawaz et al. 2005) and despite the known influence of sGC function on inflammatory phenotypes (Ahluwalia et al. 2004), the exact mechanisms are still unknown. This is complicated by the fact that a complete knockout of the murine counterpart Gucy1a3 led to the unexpected finding of reduced atherosclerotic plaque formation (Segura-Puimedon et al. 2016), whereas genetically determined reduced but not lacking expression of Gucy1a3 was also associated with increased plaque formation (Kessler et al. 2017). The genetic findings at the further mentioned NO-cGMP-signaling loci associated with CAD/MI have been extensively discussed elsewhere (Wobst et al. 2018). A promising target in addition to sGC is phosphodiesterase 5A encoded by the PDE5A gene (Nelson et al. 2017) which leads to degradation of cGMP. However, there is currently no evidence for a beneficial effect of PDE5A inhibition in atherosclerosis.
5.4 Vascular Phenotypes
Several genes identified by GWAS have been linked to vascular phenotypes including the regulation of vascular tone and vascular remodeling.
Vascular Tone and Blood Pressure
Hypertension is a known risk factor for CAD (Yusuf et al. 2004). Some of the CAD risk genes also display genome-wide association with blood pressure. In particular, NOS3, encoding endothelial NO synthase (eNOS), and GUCY1A3 (see above) have an established role also in smooth muscle cells leading to vasodilatation after production of cGMP (Moro et al. 1996). Both have been identified as blood pressure genes (International Consortium for Blood Pressure Genome-Wide Association Studies et al. 2011; Salvi et al. 2012). In line, a genetic risk score for NOS3 and GUCY1A3 was associated with increased CAD risk. However, the effect could only in part be explained by effects on blood pressure (Emdin et al. 2018). Further genes associated with both CAD and blood pressure include SH2B3, CYP17A1, FURIN, AGT, and ARHGAP42. Profound knowledge about the underlying mechanisms is still lacking. However, a genetic risk score for hypertension was strongly associated with CAD/MI and vice versa highlighting the importance of this risk factor which also has a strong heritability (Ntalla et al. 2019).
Vascular Remodeling
A large number of genes have been linked to vascular remodeling (for an overview see Erdmann et al. (2018)). One example is the REST gene which has been identified as a CAD risk gene in 2017 (Nelson et al. 2017). REST encodes the RE-1 silencing transcription factor (REST) which has been mainly studied in neuronal diseases as Huntington disease (Zuccato et al. 2003) or seizures (McClelland et al. 2014). However, there is also evidence for a role in cardiac development where REST led to increased proliferation (Zhang et al. 2017). Additionally, REST has been described to inhibit microRNA-21 and to inhibit proliferation in vascular smooth muscle cells. As a consequence, REST might influence the formation of vulnerable atherosclerotic plaques (Jin et al. 2018). As it acts as a transcriptional regulator, several downstream transcripts might play important pathophysiological roles and represent novel targets. Another novel target is a disintegrin and metalloproteinase with thrombospondin motifs 7 (ADAMTS-7) which has been identified as a CAD risk gene (ADAMTS7) in 2011 (Reilly et al. 2011; Schunkert et al. 2011; Coronary Artery Disease C4D Genetics Consortium 2011a). It has been shown that Adamts7 deficiency leads to reduced neointima formation after vascular injury (Bauer et al. 2015; Kessler et al. 2015b) as well as reduced atherosclerotic plaque formation under proatherogenic conditions (Bauer et al. 2015). Whereas the influence on vascular remodeling is influenced by ADAMTS-7-dependent degradation of cartilage oligomeric matrix protein (COMP) (Wang et al. 2009) and thrombospondin-1 (TSP-1) (Kessler et al. 2015b) with effects on vascular smooth muscle cells and endothelial cells, respectively, the mechanism underlying reduced atherosclerotic plaque formation in mice lacking Adamts7 remains unknown so far. Of note, the ADAMTS7 locus is the only locus that has been identified to exert a gene-environment interaction with smoking. In smokers, the protective effect of the ADAMTS7 non-risk allele was outweighed in smokers, presumably via upregulation of ADAMTS7 expression secondary to exposure to tobacco ingredients (Saleheen et al. 2017).
5.5 Further Directions
Other CAD risk genes have been clustered into similarly interesting pathways which could in principle be targets of therapeutic interventions. Genes have, for example, also been annotated to transcriptional gene regulation, mitosis and proliferation, or neovascularization and angiogenesis (Lempiäinen et al. 2018; Zeng et al. 2019b). The majority of variants and genes has nevertheless not yet been classified to such pathways (Erdmann et al. 2018). Also, the role of the first and strongest risk locus chromosome 9p21 is still not fully understood. Whereas studies initially focused on the role of the cyclin-dependent kinase inhibitors 2A/2B (CDKN2A/B) (Harismendy et al. 2011), the circular form of the non-coding RNA ANRIL, which is also located at the locus (Pasmant et al. 2011), was found to have an atheroprotective role through balanced regulation of vascular cells. In contrast, the linear form which is increased in carriers of the risk allele increases atherosclerosis risk (Holdt et al. 2016).
6 Risk Scores and Risk Prediction
Apart from identifying novel pharmacological targets, GWAS raised hope to identify patients at risk early on to improve prevention of CAD. Whereas initial studies including some of the identified variants only led to modest success in this regard (Hughes et al. 2012; for an overview see Kessler and Schunkert (2012)), recent studies using polygenic risk scores changed the scenario. A polygenic risk score for CSD including more than six millions of variants was able to identify individuals with a substantially elevated CAD risk. Strikingly, the score identified 20 times more individuals than familial hypercholesterolemia patients, while the carriers of a high genetic risk score were at even higher CAD risk (Khera et al. 2018). Even in a scenario in which high polygenic risk score and the presence of familial hypercholesterolemia lead to a same increase in CAD risk, the prevalence of a high polygenic risk score was ten times higher than that of familial hypercholesterolemia (Khera et al. 2019) illustrating the potential of including such scores in prevention programs.
Another field in which polygenic risk scores can be used is to investigate associations with other phenotypes. In particular, as discussed above it has been shown that CAD/MI and other cardiovascular phenotypes, e.g., peripheral arterial disease, stroke, or abdominal aneurysm, share genetic predisposition. In contrast, risk of migraine was reduced with increased genetic CAD/MI risk (Ntalla et al. 2019). Risk scores were also able to dissect an interrelationship between educational attainment, which is regarded as a determinant of lifestyle factors, and CAD/MI risk. Here, a genetic risk score including variants which are known to be associated with educational attainment was also associated with CAD risk. As the signal was lost after adjusting for body mass index and smoking, it can be hypothesized that a genetic predisposition to educational attainment might influence a healthier lifestyle and, subsequently, reduce CAD/MI risk (Zeng et al. 2019a).
Furthermore, there is hope that knowledge of particular genetic risk factors can be used to design individualized treatment strategies. GUCY1A3 as an example has been discussed above. Here, knowledge of genotype could inform aspirin therapy in the setting of primary prevention (Hall et al. 2019) and ischemic risk in CAD patients after PCI (Kessler et al. 2019). Genetic risk scores have also been able to identify individuals who could have a larger benefit from statin treatment (Mega et al. 2015; Natarajan et al. 2017). In the future, polygenic risk scores might therefore also be used in the design of clinical trials. A recent post hoc study of the ODYSSEY trial which investigated the benefit from the PCSK9 inhibitor alirocumab in CAD (Schwartz et al. 2018) revealed that individuals with a higher polygenic risk score particularly benefited from treatment (Damask et al. 2020). Whereas this clearly indicated that polygenic risk scores might provide an excellent tool for risk stratification, a prospective benefit in the design of clinical trials needs to be demonstrated.
7 Summary
Genomic studies led to the identification of a large and still growing number of genes which play a role in the pathophysiology of CAD/MI. While only a few have been functionally investigated so far, novel therapeutic strategies have been developed in, e.g., LDL-cholesterol metabolism, and further promising targets might be identified. In addition, knowledge of genetic risk factors might facilitate prevention of the disease through early identification of individuals at risk and therapy via individualized treatment strategies.
References
1000 Genomes Project Consortium, Abecasis GR, Auton A et al (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65. https://doi.org/10.1038/nature11632
1000 Genomes Project Consortium, Auton A, Brooks LD et al (2015) A global reference for human genetic variation. Nature 526:68–74. https://doi.org/10.1038/nature15393
Abifadel M, Varret M, Rabès J-P et al (2003) Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat Genet 34:154–156. https://doi.org/10.1038/ng1161
Ahluwalia A, Foster P, Scotland RS et al (2004) Antiinflammatory activity of soluble guanylate cyclase: cGMP-dependent down-regulation of P-selectin expression and leukocyte recruitment. Proc Natl Acad Sci U S A 101:1386–1391. https://doi.org/10.1073/pnas.0304264101
Ahmad Z, Banerjee P, Hamon S et al (2019) Inhibition of Angiopoietin-Like Protein 3 with a Monoclonal antibody reduces triglycerides in hypertriglyceridemia. Circulation 140:470–486. https://doi.org/10.1161/CIRCULATIONAHA.118.039107
Barrett JC, Clayton DG, Concannon P et al (2009) Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet 41:703–707. https://doi.org/10.1038/ng.381
Bauer RC, Tohyama J, Cui J et al (2015) Knockout of Adamts7, a novel coronary artery disease locus in humans, reduces atherosclerosis in mice. Circulation 131:1202–1213. https://doi.org/10.1161/CIRCULATIONAHA.114.012669
Braenne I, Willenborg C, Tragante V et al (2017) A genomic exploration identifies mechanisms that may explain adverse cardiovascular effects of COX-2 inhibitors. Sci Rep 7:10252. https://doi.org/10.1038/s41598-017-10928-4
Brown MS, Goldstein JL (1986) A receptor-mediated pathway for cholesterol homeostasis. Science 232:34–47
C Reactive Protein Coronary Heart Disease Genetics Collaboration (CCGC), Wensley F, Gao P et al (2011) Association between C reactive protein and coronary heart disease: mendelian randomisation analysis based on individual participant data. BMJ 342:d548. https://doi.org/10.1136/bmj.d548
Cameron J, Holla ØL, Ranheim T et al (2006) Effect of mutations in the PCSK9 gene on the cell surface LDL receptors. Hum Mol Genet 15:1551–1558. https://doi.org/10.1093/hmg/ddl077
Cannon CP, Blazing MA, Giugliano RP et al (2015) Ezetimibe added to statin therapy after acute coronary syndromes. N Engl J Med 372:2387–2397. https://doi.org/10.1056/NEJMoa1410489
CARDIoGRAMplusC4D Consortium, Deloukas P, Kanoni S et al (2013) Large-scale association analysis identifies new risk loci for coronary artery disease. Nat Genet 45:25–33. https://doi.org/10.1038/ng.2480
Cohen JC, Boerwinkle E, Mosley TH, Hobbs HH (2006) Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N Engl J Med 354:1264–1272. https://doi.org/10.1056/NEJMoa054013
Coronary Artery Disease C4D Genetics Consortium (2011a) A genome-wide association study in Europeans and South Asians identifies five new loci for coronary artery disease. Nat Genet 43:339–344. https://doi.org/10.1038/ng.782
Coronary Artery Disease C4D Genetics Consortium (2011b) A genome-wide association study in Europeans and South Asians identifies five new loci for coronary artery disease. Nat Genet 43:339–344. https://doi.org/10.1038/ng.782
Damask A, Steg PG, Schwartz GG et al (2020) Patients with high genome-wide polygenic risk scores for coronary artery disease may receive greater clinical benefit from alirocumab treatment in the ODYSSEY OUTCOMES Trial. Circulation 141:624–636. https://doi.org/10.1161/CIRCULATIONAHA.119.044434
Danesh J, Wheeler JG, Hirschfield GM et al (2004) C-reactive protein and other circulating markers of inflammation in the prediction of coronary heart disease. N Engl J Med 350:1387–1397. https://doi.org/10.1056/NEJMoa032804
Dichgans M, Malik R, König IR et al (2014) Shared genetic susceptibility to ischemic stroke and coronary artery disease: a genome-wide analysis of common variants. Stroke 45:24–36. https://doi.org/10.1161/STROKEAHA.113.002707
Dichgans M, Pulit SL, Rosand J (2019) Stroke genetics: discovery, biology, and clinical applications. Lancet Neurol 18:587–599. https://doi.org/10.1016/S1474-4422(19)30043-2
Do R, Stitziel NO, Won H-H et al (2015) Exome sequencing identifies rare LDLR and APOA5 alleles conferring risk for myocardial infarction. Nature 518:102–106. https://doi.org/10.1038/nature13917
Döring Y, Pawig L, Weber C, Noels H (2014) The CXCL12/CXCR4 chemokine ligand/receptor axis in cardiovascular disease. Front Physiol 5:212. https://doi.org/10.3389/fphys.2014.00212
Döring Y, Noels H, van der Vorst EPC et al (2017) Vascular CXCR4 limits atherosclerosis by maintaining arterial integrity: evidence from mouse and human studies. Circulation 136:388–403. https://doi.org/10.1161/CIRCULATIONAHA.117.027646
Döring Y, van der Vorst EPC, Duchene J et al (2019) CXCL12 derived from endothelial cells promotes atherosclerosis to drive coronary artery disease. Circulation 139:1338–1340. https://doi.org/10.1161/CIRCULATIONAHA.118.037953
Emdin CA, Khera AV, Klarin D et al (2018) Phenotypic consequences of a genetic predisposition to enhanced nitric oxide signaling. Circulation 137:222–232. https://doi.org/10.1161/CIRCULATIONAHA.117.028021
Erdmann J, Grosshennig A, Braund PS et al (2009) New susceptibility locus for coronary artery disease on chromosome 3q22.3. Nat Genet 41:280–282. https://doi.org/10.1038/ng.307
Erdmann J, Willenborg C, Nahrstaedt J et al (2011) Genome-wide association study identifies a new locus for coronary artery disease on chromosome 10p11.23. Eur Heart J 32:158–168. https://doi.org/10.1093/eurheartj/ehq405
Erdmann J, Stark K, Esslinger UB et al (2013) Dysfunctional nitric oxide signalling increases risk of myocardial infarction. Nature 504:432–436. https://doi.org/10.1038/nature12722
Erdmann J, Kessler T, Munoz Venegas L, Schunkert H (2018) A decade of genome-wide association studies for coronary artery disease: the challenges ahead. Cardiovasc Res 114:1241–1257. https://doi.org/10.1093/cvr/cvy084
Flister MJ, Hoffman MJ, Lemke A et al (2015) SH2B3 is a genetic determinant of cardiac inflammation and fibrosis. Circ Cardiovasc Genet 8:294–304. https://doi.org/10.1161/CIRCGENETICS.114.000527
Gaudet D, Alexander VJ, Baker BF et al (2015) Antisense inhibition of apolipoprotein C-III in patients with hypertriglyceridemia. N Engl J Med 373:438–447. https://doi.org/10.1056/NEJMoa1400283
Gawaz M, Langer H, May AE (2005) Platelets in inflammation and atherogenesis. J Clin Invest 115:3378–3384. https://doi.org/10.1172/JCI27196
Ghofrani H-A, D’Armini AM, Grimminger F et al (2013a) Riociguat for the treatment of chronic thromboembolic pulmonary hypertension. N Engl J Med 369:319–329. https://doi.org/10.1056/NEJMoa1209657
Ghofrani H-A, Galiè N, Grimminger F et al (2013b) Riociguat for the treatment of pulmonary arterial hypertension. N Engl J Med 369:330–340. https://doi.org/10.1056/NEJMoa1209655
Graham MJ, Lee RG, Bell TA et al (2013) Antisense oligonucleotide inhibition of apolipoprotein C-III reduces plasma triglycerides in rodents, nonhuman primates, and humans. Circ Res 112:1479–1490. https://doi.org/10.1161/CIRCRESAHA.111.300367
Gudbjartsson DF, Bjornsdottir US, Halapi E et al (2009) Sequence variants affecting eosinophil numbers associate with asthma and myocardial infarction. Nat Genet 41:342–347. https://doi.org/10.1038/ng.323
Hall KT, Kessler T, Buring JE et al (2019) Genetic variation at the coronary artery disease risk locus GUCY1A3 modifies cardiovascular disease prevention effects of aspirin. Eur Heart J 40:3385–3392. https://doi.org/10.1093/eurheartj/ehz384
Harismendy O, Notani D, Song X et al (2011) 9p21 DNA variants associated with coronary artery disease impair interferon-γ signalling response. Nature 470:264–268. https://doi.org/10.1038/nature09753
Helgadottir A, Thorleifsson G, Manolescu A et al (2007) A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316:1491–1493. https://doi.org/10.1126/science.1142842
Holdt LM, Beutner F, Scholz M et al (2010) ANRIL expression is associated with atherosclerosis risk at chromosome 9p21. Arterioscler Thromb Vasc Biol 30:620–627. https://doi.org/10.1161/ATVBAHA.109.196832
Holdt LM, Hoffmann S, Sass K et al (2013) ALU elements in ANRIL non-coding RNA at chromosome 9p21 modulate atherogenic cell functions through trans-regulation of gene networks. PLoS Genet 9:e1003588. https://doi.org/10.1371/journal.pgen.1003588
Holdt LM, Stahringer A, Sass K et al (2016) Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat Commun 7:12429. https://doi.org/10.1038/ncomms12429
Howson JMM, Zhao W, Barnes DR et al (2017) Fifteen new risk loci for coronary artery disease highlight arterial-wall-specific mechanisms. Nat Genet 385:117–119. https://doi.org/10.1038/ng.3874
Hughes MF, Saarela O, Stritzke J et al (2012) Genetic markers enhance coronary risk prediction in men: the MORGAM prospective cohorts. PLoS One 7:e40922
Hunt KA, Zhernakova A, Turner G et al (2008) Newly identified genetic risk variants for celiac disease related to the immune response. Nat Genet 40:395–402. https://doi.org/10.1038/ng.102
IBC 50K CAD Consortium (2011) Large-scale gene-centric analysis identifies novel variants for coronary artery disease. PLoS Genet 7:e1002260. https://doi.org/10.1371/journal.pgen.1002260
Interleukin-6 Receptor Mendelian Randomisation Analysis (IL6R MR) Consortium, Swerdlow DI, Holmes MV et al (2012) The interleukin-6 receptor as a target for prevention of coronary heart disease: a mendelian randomisation analysis. 379:1214–1224. https://doi.org/10.1016/S0140-6736(12)60110-X
International Consortium for Blood Pressure Genome-Wide Association Studies, Ehret GB, Munroe PB et al (2011) Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 478:103–109. https://doi.org/10.1038/nature10405
Jansen H, Samani NJ, Schunkert H (2014) Mendelian randomization studies in coronary artery disease. Eur Heart J 35:1917–1924. https://doi.org/10.1093/eurheartj/ehu208
Jin H, Li DY, Chernogubova E et al (2018) Local delivery of miR-21 stabilizes fibrous caps in vulnerable atherosclerotic lesions. Mol Ther 26:1040–1055. https://doi.org/10.1016/j.ymthe.2018.01.011
Kathiresan S, Melander O, Anevski D et al (2008) Polymorphisms associated with cholesterol and risk of cardiovascular events. N Engl J Med 358:1240–1249. https://doi.org/10.1056/NEJMoa0706728
Kessler T, Schunkert H (2012) Clinical validation of genetic markers for improved risk estimation. Eur J Prev Cardiol 19:25–32. https://doi.org/10.1177/2047487312448993
Kessler T, Erdmann J, Dichgans M, Schunkert H (2015a) Shared genetic aetiology of coronary artery disease and atherosclerotic stroke-2015. Curr Atheroscler Rep 17:498. https://doi.org/10.1007/s11883-015-0498-5
Kessler T, Zhang L, Liu Z et al (2015b) ADAMTS-7 inhibits re-endothelialization of injured arteries and promotes vascular remodeling through cleavage of Thrombospondin-1. Circulation 131:1191–1201. https://doi.org/10.1161/CIRCULATIONAHA.114.014072
Kessler T, Vilne B, Schunkert H (2016) The impact of genome-wide association studies on the pathophysiology and therapy of cardiovascular disease. EMBO Mol Med 8:688–701. https://doi.org/10.15252/emmm.201506174
Kessler T, Wobst J, Wolf B et al (2017) Functional characterization of the GUCY1A3 coronary artery disease risk locus. Circulation 136:476–489. https://doi.org/10.1161/CIRCULATIONAHA.116.024152
Kessler T, Wolf B, Eriksson N et al (2019) Association of the coronary artery disease risk gene GUCY1A3 with ischaemic events after coronary intervention. Cardiovasc Res 115:1512–1518. https://doi.org/10.1093/cvr/cvz015
Khera AV, Kathiresan S (2017a) Genetics of coronary artery disease: discovery, biology and clinical translation. Nat Rev Genet 18:331–344. https://doi.org/10.1038/nrg.2016.160
Khera AV, Kathiresan S (2017b) Genetics of coronary artery disease: discovery, biology and clinical translation. Nat Rev Genet 18:1–14. https://doi.org/10.1038/nrg.2016.160
Khera AV, Won H-H, Peloso GM et al (2017) Association of rare and common variation in the lipoprotein lipase gene with coronary artery disease. JAMA 317:937–946. https://doi.org/10.1001/jama.2017.0972
Khera AV, Chaffin M, Aragam KG et al (2018) Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet 50:1219–1224. https://doi.org/10.1038/s41588-018-0183-z
Khera AV, Chaffin M, Zekavat SM et al (2019) Whole-genome sequencing to characterize monogenic and polygenic contributions in patients hospitalized with early-onset myocardial infarction. Circulation 139:1593–1602. https://doi.org/10.1161/CIRCULATIONAHA.118.035658
Klarin D, Zhu QM, Emdin CA et al (2017) Genetic analysis in UK Biobank links insulin resistance and transendothelial migration pathways to coronary artery disease. Nat Genet 49:1392–1397. https://doi.org/10.1038/ng.3914
Klarin D, Damrauer SM, Cho K et al (2018) Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat Genet 50:1514–1523
Koenig W, Sund M, Fröhlich M et al (1999) C-Reactive protein, a sensitive marker of inflammation, predicts future risk of coronary heart disease in initially healthy middle-aged men: results from the MONICA (Monitoring Trends and Determinants in Cardiovascular Disease) Augsburg Cohort Study, 1984 to 1992. Circulation 99:237–242. https://doi.org/10.1161/01.cir.99.2.237
Lander ES, Linton LM, Birren B et al (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Lange LA, Carlson CS, Hindorff LA et al (2006) Association of polymorphisms in the CRP gene with circulating C-reactive protein levels and cardiovascular events. JAMA 296:2703–2711. https://doi.org/10.1001/jama.296.22.2703
Lempiäinen H, Brænne I, Michoel T et al (2018) Network analysis of coronary artery disease risk genes elucidates disease mechanisms and druggable targets. Sci Rep 8:3434
Levy D, Ehret GB, Rice K et al (2009) Genome-wide association study of blood pressure and hypertension. Nat Genet 41:677–687. https://doi.org/10.1038/ng.384
Libby P, Ridker PM, Hansson GK (2011) Progress and challenges in translating the biology of atherosclerosis. Nature 473:317–325. https://doi.org/10.1038/nature10146
Linsel-Nitschke P, Götz A, Erdmann J et al (2008) Lifelong reduction of LDL-cholesterol related to a common variant in the LDL-receptor gene decreases the risk of coronary artery disease--a Mendelian Randomisation study. PLoS One 3:e2986. https://doi.org/10.1371/journal.pone.0002986
Littlejohns TJ, Sudlow C, Allen NE, Collins R (2019) UK Biobank: opportunities for cardiovascular research. Eur Heart J 40:1158–1166. https://doi.org/10.1093/eurheartj/ehx254
Lusis AJ (2000) Atherosclerosis. Nature 407:233–241. https://doi.org/10.1038/35025203
McClelland S, Brennan GP, Dubé C et al (2014) The transcription factor NRSF contributes to epileptogenesis by selective repression of a subset of target genes. Elife 3:e01267. https://doi.org/10.7554/eLife.01267
McPherson R, Tybjaerg-Hansen A (2016) Genetics of coronary artery disease. Circ Res 118:564–578. https://doi.org/10.1161/CIRCRESAHA.115.306566
McPherson R, Pertsemlidis A, Kavaslar N et al (2007) A common allele on chromosome 9 associated with coronary heart disease. Science 316:1488–1491. https://doi.org/10.1126/science.1142447
Mega JL, Stitziel NO, Smith JG et al (2015) Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: an analysis of primary and secondary prevention trials. Lancet 385:2264–2271. https://doi.org/10.1016/S0140-6736(14)61730-X
Mori T, Iwasaki Y, Seki Y et al (2014) Lnk/Sh2b3 controls the production and function of dendritic cells and regulates the induction of IFN-γ-producing T cells. J Immunol 193:1728–1736. https://doi.org/10.4049/jimmunol.1303243
Moro MA, Russel RJ, Cellek S et al (1996) cGMP mediates the vascular and platelet actions of nitric oxide: confirmation using an inhibitor of the soluble guanylyl cyclase. Proc Natl Acad Sci U S A 93:1480–1485
Musunuru K, Strong A, Frank-Kamenetsky M et al (2010) From noncoding variant to phenotype via SORT1 at the 1p13 cholesterol locus. Nature 466:714–719. https://doi.org/10.1038/nature09266
Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia Investigators, Stitziel NO, Stirrups KE et al (2016) Coding variation in ANGPTL4, LPL, and SVEP1 and the risk of coronary disease. N Engl J Med 374:1134–1144. https://doi.org/10.1056/NEJMoa1507652
Myocardial Infarction Genetics Consortium Investigators, Stitziel NO, Won H-H et al (2014) Inactivating mutations in NPC1L1 and protection from coronary heart disease. N Engl J Med 371:2072–2082. https://doi.org/10.1056/NEJMoa1405386
Myocardial Infarction Genetics Consortium, Voight BF, Purcell S et al (2009) Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat Genet 41:334–341. https://doi.org/10.1038/ng.327
Natarajan P, Young R, Stitziel NO et al (2017) Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation 135:2091–2101. https://doi.org/10.1161/CIRCULATIONAHA.116.024436
Nelson CP, Goel A, Butterworth AS et al (2017) Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat Genet 49:1385–1391. https://doi.org/10.1038/ng.3913
Newton-Cheh C, Johnson T, Gateva V et al (2009) Genome-wide association study identifies eight loci associated with blood pressure. Nat Genet 41:666–676. https://doi.org/10.1038/ng.361
Nikpay M, Goel A, Won H-H et al (2015) A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat Genet 47:1121–1130. https://doi.org/10.1038/ng.3396
Nomura A, Won H-H, Khera AV et al (2017) Protein-truncating variants at the Cholesteryl Ester transfer protein gene and risk for coronary heart disease. Circ Res 121:81–88. https://doi.org/10.1161/CIRCRESAHA.117.311145
Ntalla I, Kanoni S, Zeng L et al (2019) Genetic risk score for coronary disease identifies predispositions to cardiovascular and noncardiovascular diseases. J Am Coll Cardiol 73:2932–2942
Pasmant E, Sabbagh A, Vidaud M, Bièche I (2011) ANRIL, a long, noncoding RNA, is an unexpected major hotspot in GWAS. FASEB J 25:444–448. https://doi.org/10.1096/fj.10-172452
Pe’er I, Yelensky R, Altshuler D, Daly MJ (2008) Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet Epidemiol 32:381–385. https://doi.org/10.1002/gepi.20303
Peloso GM, Nomura A, Khera AV et al (2019) Rare protein-truncating variants in APOB, lower low-density lipoprotein cholesterol, and protection against coronary heart disease. Circ Genom Precis Med 12:e002376. https://doi.org/10.1161/CIRCGEN.118.002376
Pepys MB, Hirschfield GM, Tennent GA et al (2006) Targeting C-reactive protein for the treatment of cardiovascular disease. Nature 440:1217–1221. https://doi.org/10.1038/nature04672
Reich DE, Lander ES (2001) On the allelic spectrum of human disease. Trends Genet 17:502–510
Reilly MP, Li M, He J et al (2011) Identification of ADAMTS7 as a novel locus for coronary atherosclerosis and association of ABO with myocardial infarction in the presence of coronary atherosclerosis: two genome-wide association studies. Lancet 377:383–392. https://doi.org/10.1016/S0140-6736(10)61996-4
Ridker PM, Danielson E, Fonseca FAH et al (2008) Rosuvastatin to prevent vascular events in men and women with elevated C-reactive protein. N Engl J Med 359:2195–2207. https://doi.org/10.1056/NEJMoa0807646
Ridker PM, Everett BM, Thuren T et al (2017) Antiinflammatory therapy with Canakinumab for atherosclerotic disease. N Engl J Med 377:1119–1131. https://doi.org/10.1056/NEJMoa1707914
Sabatine MS, Giugliano RP, Wiviott SD et al (2015) Efficacy and safety of evolocumab in reducing lipids and cardiovascular events. N Engl J Med 372:1500–1509. https://doi.org/10.1056/NEJMoa1500858
Sabatine MS, Giugliano RP, Keech AC et al (2017) Evolocumab and clinical outcomes in patients with cardiovascular disease. N Engl J Med 376:1713–1722. https://doi.org/10.1056/NEJMoa1615664
Sachidanandam R, Weissman D, Schmidt SC et al (2001) A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409:928–933. https://doi.org/10.1038/35057149
Saleheen D, Zhao W, Young R et al (2017) Loss of cardioprotective effects at the ADAMTS7 locus as a result of gene-smoking interactions. Circulation 135:2336–2353. https://doi.org/10.1161/CIRCULATIONAHA.116.022069
Salvi E, Kutalik Z, Glorioso N et al (2012) Genomewide association study using a high-density single nucleotide polymorphism array and case-control design identifies a novel essential hypertension susceptibility locus in the promoter region of endothelial NO synthase. Hypertension 59:248–255. https://doi.org/10.1161/HYPERTENSIONAHA.111.181990
Samani NJ, Erdmann J, Hall AS et al (2007) Genomewide association analysis of coronary artery disease. N Engl J Med 357:443–453. https://doi.org/10.1056/NEJMoa072366
Schunkert H, Samani NJ (2008) Elevated C-reactive protein in atherosclerosis--chicken or egg? N Engl J Med 359:1953–1955
Schunkert H, König IR, Kathiresan S et al (2011) Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat Genet 43:333–338. https://doi.org/10.1038/ng.784
Schunkert H, Erdmann J, Samani SNJ (2019) CARDIoGRAM celebrates its 10th Anniversary. Eur Heart J 40:1664–1666. https://doi.org/10.1093/eurheartj/ehz347
Schwartz GG, Steg PG, Szarek M et al (2018) Alirocumab and cardiovascular outcomes after acute coronary syndrome. N Engl J Med 379:2097–2107. https://doi.org/10.1056/NEJMoa1801174
Segura-Puimedon M, Mergia E, Al-Hasani J et al (2016) Proatherosclerotic effect of the α1-subunit of soluble Guanylyl Cyclase by promoting smooth muscle phenotypic switching. Am J Pathol 186:2220–2231. https://doi.org/10.1016/j.ajpath.2016.04.010
Soranzo N, Spector TD, Mangino M et al (2009) A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat Genet 41:1182–1190. https://doi.org/10.1038/ng.467
Stasch J-P, Pacher P, Evgenov OV (2011) Soluble guanylate cyclase as an emerging therapeutic target in cardiopulmonary disease. Circulation 123:2263–2273. https://doi.org/10.1161/CIRCULATIONAHA.110.981738
Stitziel NO, Khera AV, Wang X et al (2017) ANGPTL3 deficiency and protection against coronary artery disease. J Am Coll Cardiol 69:2054–2063. https://doi.org/10.1016/j.jacc.2017.02.030
Swirski FK, Nahrendorf M (2013) Leukocyte behavior in atherosclerosis, myocardial infarction, and heart failure. Science 339:161–166. https://doi.org/10.1126/science.1230719
Takizawa H, Nishimura S, Takayama N et al (2010) Lnk regulates integrin αIIbβ3 outside-in signaling in mouse platelets, leading to stabilization of thrombus development in vivo. J Clin Invest 120:179–190. https://doi.org/10.1172/JCI39503DS1
Teslovich TM, Musunuru K, Smith AV et al (2010) Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466:707–713. https://doi.org/10.1038/nature09270
The TG and HDL Working Group of the Exome Sequencing Project, National Heart, Lung, and Blood Institute (2014) Loss-of-function mutations in APOC3, triglycerides, and coronary disease. N Engl J Med 371:22–31. https://doi.org/10.1056/NEJMoa1307095
Tolleshaug H, Hobgood KK, Brown MS, Goldstein JL (1983) The LDL receptor locus in familial hypercholesterolemia: multiple mutations disrupt transport and processing of a membrane receptor. Cell 32:941–951
Tong W, Lodish HF (2004) Lnk inhibits Tpo-mpl signaling and Tpo-mediated megakaryocytopoiesis. J Exp Med 200:569–580. https://doi.org/10.1084/jem.20040762
Tregouet D-A, König IR, Erdmann J et al (2009) Genome-wide haplotype association study identifies the SLC22A3-LPAL2-LPA gene cluster as a risk locus for coronary artery disease. Nat Genet 41:283–285. https://doi.org/10.1038/ng.314
Trenkwalder T, Nelson CP, Musameh MD et al (2018) Effects of the coronary artery disease associated LPA and 9p21 loci on risk of aortic valve stenosis. Int J Cardiol 276:212–217. https://doi.org/10.1016/j.ijcard.2018.11.094
Tsou C-Y, Chen C-Y, Zhao J-F et al (2014) Activation of soluble guanylyl cyclase prevents foam cell formation and atherosclerosis. Acta Physiol (Oxf) 210:799–810. https://doi.org/10.1111/apha.12210
van der Harst P, Verweij N (2018) Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ Res 122:433–443. https://doi.org/10.1161/CIRCRESAHA.117.312086
Verweij N, Eppinga RN, Hagemeijer Y, van der Harst P (2017) Identification of 15 novel risk loci for coronary artery disease and genetic risk of recurrent events, atrial fibrillation and heart failure. Sci Rep 7:2761. https://doi.org/10.1038/s41598-017-03062-8
Wang L, Zheng J, Bai X et al (2009) ADAMTS-7 mediates vascular smooth muscle cell migration and neointima formation in balloon-injured rat arteries. Circ Res 104:688–698. https://doi.org/10.1161/CIRCRESAHA.108.188425
Wang W, Tang Y, Wang Y et al (2016) LNK/SH2B3 loss of function promotes atherosclerosis and thrombosis. Circ Res 119:e91–e103. https://doi.org/10.1161/CIRCRESAHA.116.308955
Webb TR, Erdmann J, Stirrups KE et al (2017) Systematic evaluation of pleiotropy identifies 6 further loci associated with coronary artery disease. J Am Coll Cardiol 69:823–836. https://doi.org/10.1016/j.jacc.2016.11.056
Wobst J, Ameln Von S, Wolf B et al (2016) Stimulators of the soluble guanylyl cyclase: promising functional insights from rare coding atherosclerosis-related GUCY1A3 variants. Basic Res Cardiol 111:51. https://doi.org/10.1007/s00395-016-0570-5
Wobst J, Schunkert H, Kessler T (2018) Genetic alterations in the NO-cGMP pathway and cardiovascular risk. Nitric Oxide 76:105–112. https://doi.org/10.1016/j.niox.2018.03.019
Yusuf S, Hawken S, Ounpuu S et al (2004) Effect of potentially modifiable risk factors associated with myocardial infarction in 52 countries (the INTERHEART study): case-control study. Lancet 364:937–952. https://doi.org/10.1016/S0140-6736(04)17018-9
Zacho J, Tybjaerg-Hansen A, Jensen JS et al (2008) Genetically elevated C-reactive protein and ischemic vascular disease. N Engl J Med 359:1897–1908. https://doi.org/10.1056/NEJMoa0707402
Zeng L, Ntalla I, Kessler T et al (2019a) Genetically modulated educational attainment and coronary disease risk. Eur Heart J 40:2413–2420
Zeng L, Talukdar HA, Koplev S et al (2019b) Contribution of gene regulatory networks to heritability of coronary artery disease. J Am Coll Cardiol 73:2946–2957
Zernecke A, Bot I, Djalali-Talab Y et al (2008) Protective role of CXC receptor 4/CXC ligand 12 unveils the importance of neutrophils in atherosclerosis. Circ Res 102:209–217. https://doi.org/10.1161/CIRCRESAHA.107.160697
Zhang D, Wang Y, Lu P et al (2017) REST regulates the cell cycle for cardiac development and regeneration. Nat Commun 8:1979. https://doi.org/10.1038/s41467-017-02210-y
Zuccato C, Tartari M, Crotti A et al (2003) Huntingtin interacts with REST/NRSF to modulate the transcription of NRSE-controlled neuronal genes. Nat Genet 35:76–83. https://doi.org/10.1038/ng1219
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2020 The Author(s)
About this chapter
Cite this chapter
Kessler, T., Schunkert, H. (2020). Genomic Strategies Toward Identification of Novel Therapeutic Targets. In: von Eckardstein, A., Binder, C.J. (eds) Prevention and Treatment of Atherosclerosis . Handbook of Experimental Pharmacology, vol 270. Springer, Cham. https://doi.org/10.1007/164_2020_360
Download citation
DOI: https://doi.org/10.1007/164_2020_360
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-86075-2
Online ISBN: 978-3-030-86076-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)