Abstract
In the era of genomic medicine, targeted next generation sequencing strategies (NGS) are becoming increasingly adopted by clinical molecular diagnostic laboratories to identify genetic diagnostic and prognostic biomarkers in hemato-oncology. We describe the EuroClonality-NGS DNA Capture (EuroClonality-NDC) assay, which is designed to simultaneously detect B and T cell clonal rearrangements, translocations, copy number alterations, and sequence variants. The accompanying validated bioinformatics pipeline enables production of an integrated report. The combination of the laboratory protocol and bioinformatics pipeline in the EuroClonality-NDC minimizes the potential for human error, reduces economic costs compared to current molecular testing strategies, and should improve diagnostic outcomes.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- EuroClonality
- Next generation sequencing
- BIOMED-2
- Immunoglobulin
- T cell receptor
- Copy number alteration
- Translocation
- Lymphoma
1 Introduction
Lymphoproliferative disorders (LPD) can be classified based on multiple parameters, including morphology, immunophenotyping, and genetic analysis. While a large number of lymphoproliferative disorders can be classified solely by assessment of morphology and immunophenotyping, there is an increasing role for the evaluation of genetic features as evidenced by the publication of the updated WHO guidelines in 2016 [1]. The updated guidelines include a vast array of genomic alterations that can significantly improve the diagnostic criteria and the prognostic relevance of existing entities and has led to the introduction of new disease entities. This may result in a change in practice in clinical laboratories with the validation of multiple molecular tests covering the required genetic alterations.
As all cells within a tumor are presumed to arise from a common clonal progenitor, lymphoid malignancies should exhibit clonal rearrangements of the immunoglobulin (IG) and/or T cell receptor (TR) loci. Detection of a clonal IG /TR rearrangement, which can aid in differentiating between a clonal B/T cell proliferation and a reactive hyperplasia, is typically performed using a PCR-based method with primers designed, optimized, and validated during the EuroClonality BIOMED-2 study [2, 3]. PCR products are commonly analyzed using either capillary electrophoresis (i.e., GeneScan) or heteroduplex analysis on polyacrylamide gels. Analysis of clonality using these detection methods is prone to subjectivity, particularly in cases with low tumor infiltration, where it can be difficult to distinguish a clonal peak within a polyclonal background, or for targets with limited complementary determining region 3 (CDR3 ) variability (e.g., IGKV-J rearrangements). Interpretation of the results is also subject to confounding variables such as the impact of somatic hypermutation (SHM) on detection of IGH , IGK , and IGL rearrangements. The presence of mutations within the binding sites of the PCR primers can prevent annealing, leading to false-negative results which can be addressed in the majority of cases by performing PCR for alternate IG targets that are less prone to SHM such as IGH D-J and IGK Kde rearrangements [3].
Translocations are another genetic alteration tested for in molecular pathology laboratories as they are a hallmark of specific entities within non-Hodgkin lymphoma (NHL) as well as acute lymphoblastic leukemia (ALL) or plasma cell myeloma (PCM), among others. Translocations involving the IG /TR loci such as t(8;14)(q24;q32) in Burkitt lymphoma (BL), t(14;18)(q32;q21) in follicular lymphoma (FL), t(11;14)(q13;q32) in mantle cell lymphoma (MCL), and ALK translocations (to various partners such as NPM1, ATIC and RANBP2) in anaplastic large cell lymphoma (ALCL) are part of the diagnostic testing regime for those particular lymphomas. Currently, translocations are commonly assessed by FISH or PCR methods, although a number of tests are often required, particularly in B-NHL types to encompass different IG loci (IGH , IGK , and IGL ) and to accommodate the large region where translocation breakpoints can occur. Multiple tests are often required to accurately define double/triple hit lymphomas as the recent 2016 WHO guidelines established a new classification of high-grade B-NHL based on the presence of a MYC translocation along with BCL2 and/or BCL6 translocations.
Single-nucleotide DNA alterations and/or small insertions or deletions, traditionally detected using Sanger sequencing and more recently by amplicon and capture targeted NGS, can aid in diagnostic and prognostic classification of the disease [4,5,6]. Mutations can often show a higher frequency in particular LPD subtypes such as mutations of TCF3 or ID3 which have been reported in 70% of sporadic BL, mutations of MYD88 in >90% of Waldenström’s macroglobulinemia, or mutations of TET2, IDH2, and RHOA in a large percentage of angioimmunoblastic T cell lymphoma (AITL). From several sequencing studies, specific mutation profiles can define molecular subtypes such as in diffuse large B cell lymphoma (DLBCL) where specific mutations are associated with the germinal center B-cell (GCB) and activated B cell (ABC) subtypes which have pronounced survival differences with standard chemotherapy [7,8,9]. The presence of TP53 mutations in chronic lymphocytic leukemia (CLL) has been shown to be an independent prognostic factor and predictor of chemotherapy refractoriness [10]; similarly, NOTCH1 and SF3B1 mutations can be independent prognostic markers in CLL [11, 12]. Activating mutations of NOTCH1 are observed in approximately 60% of T-ALL cases and are reported to be associated with shorter survival in adults [13, 14].
Finally, copy number alterations (CNA ) are also prevalent in LPD and can be associated with the underlying biology, with 17p deletion in CLL and PCM being associated with a less favorable outcome. The European Research Initiative on Chronic Lymphocytic Leukemia (ERIC) recommends analysis of del(17p) and TP53 gene mutations as an integral part of routine diagnostics for CLL patients requiring treatment [15].
The overarching objective of the EuroClonality-NDC is to enable a single NGS test to integrate genomic analyses that are currently performed by a number of molecular testing strategies. As part of the EuroClonality-NGS working group, we have developed the EuroClonality-NGS DNA capture assay (EuroClonality-NDC) to detect clinically relevant genetic alterations in LPD using a capture-hybridization approach. To achieve this objective, EuroClonality-NDC was designed to capture all functional variable (V), diversity (D), and joining (J) genes of the IG and TR loci along with additional probes to identify structural variants (SV) in the form of chromosomal translocations and detect CNA and somatic mutations. The accompanying purpose-built bioinformatics pipeline, ARResT/Interrogate, which was originally developed for amplicon assays, was customized and validated for the EuroClonality-NDC [16]. An optimized standard operating procedure (SOP), which has undergone a multi-site validation, ensures robust assay performance [17]. The development and validation of both the EuroClonality-NDC capture panel and the bioinformatics platform provides an end-to-end workflow which minimizes subjective interpretation of results. The methods detailed in this chapter relate to an updated version of the SOP to reflect recent improvements in library preparation and target enrichment.
2 Materials
2.1 DNA Quantification
The following products and equipment from Thermo Fisher Scientific (Waltham, MA, USA) are required:
-
1.
Qubit dsDNA broad-range (BR) assay.
-
2.
Qubit dsDNA high-sensitivity (HS) assay.
-
3.
Qubit Assay Tubes.
-
4.
Qubit Fluorometer.
2.2 DNA Integrity Assessment
The following products and equipment from Agilent Technologies (Santa Clara, CA, USA) are required:
-
1.
Genomic DNA Reagents.
-
2.
Genomic DNA ScreenTape.
-
3.
D1000 Reagents.
-
4.
D1000 ScreenTape.
-
5.
High Sensitivity D1000 Reagents.
-
6.
High Sensitivity D1000 ScreenTape.
-
7.
4150/4200 TapeStation System.
2.3 DNA Library Preparation
The following products from Roche Sequencing Solutions (Pleasanton, CA, USA) are required:
-
1.
KAPA HyperPlus Kit.
-
2.
KAPA UDI Primer Mixes.
-
3.
KAPA Universal Adapter.
-
4.
KAPA HyperPure Beads.
The following items will be required for the multiple bead cleanup steps that are performed in both a pre- and post-PCR environment:
-
1.
96-well magnetic plate.
-
2.
Magnetic stands for 0.2 mL PCR strips.
-
3.
Magnetic stands for 1.5 mL microfuge tubes.
2.4 DNA Hybridization
The following products from Roche Sequencing Solutions (Pleasanton, CA, USA) are required:
-
1.
KAPA HyperCapture Reagent Kit.
-
2.
KAPA HyperCapture Bead.
The following product from Univ8 Genomics Ltd. (Belfast, UK) is required:
-
1.
EuroClonality-NDC.
2.5 Sequencing of Enriched DNA Library
The following products from Illumina, Inc. (San Diego, CA, USA) are required:
-
1.
PhiX Sequencing Control V3.
-
2.
NextSeq 500/550 Mid Output Kit v2.5 (150 cycles).
-
3.
NextSeq 500/550 Sequencing System.
3 Methods
3.1 Genomic DNA Evaluation and Preparation for DNA Library Generation
-
1.
If the extraction of genomic DNA leads to the DNA being eluted into a buffer containing EDTA (see Note 1), a column or bead-based purification should be performed prior to performing any additional steps as the fragmentation enzyme is sensitive to EDTA.
-
2.
The gDNA concentration is assessed using the Qubit broad range assay. Manufacturer guidelines are followed with two modifications: (1) the standard/sample is added to the Qubit assay tubes first followed by the Qubit working solution, and (2) the incubation time prior to reading the standard/sample is 20 min.
-
3.
The gDNA integrity assessment is performed using the Genomic DNA ScreenTape Assay. Manufacturer guidelines are followed without any modifications.
-
4.
For the EuroClonality-NDC protocol, a positive control, a no template control (NTC), and 22 samples are processed in each batch (see Note 2). In well A1 of a 96-well PCR plate, place 100 ng of the positive control in a total of 35 μL, and in well A2, place 35 μL of the NTC.
-
5.
For the EuroClonality-NDC assay, 100 ng of high-molecular-weight genomic DNA is required or for genomic DNA extracted from formalin-fixed DNA 100 ng (average fragment size >1000 bp) or 200 ng (average fragment size <1000 bp) is used in a total of 35 μL. Each sample to be prepared should be placed into a separate well of a 96-well PCR plate (see Note 3).
3.2 DNA Library Generation
-
1.
Remove the following products from the KAPA HyperPlus Kit and thaw on ice:
-
(a)
KAPA Frag Buffer (10×).
-
(b)
End Repair & A-Tailing Buffer.
-
(c)
Ligation Buffer.
-
(d)
KAPA HiFi HotStart ReadyMix (2×).
-
(e)
Library Amplification Primer Mix (10×).
-
(a)
-
2.
Prepare a thermocycler by selecting the fragmentation program (Table 1) and pausing prior to the commencement of the first step to ensure the block is pre-cooled to 4 °C (see Note 4).
-
3.
While keeping the reagents on ice, prepare a mastermix which contains 5 μL KAPA Frag Buffer (10×) and 10 μL KAPA Frag Enzyme for each reaction to be performed.
-
4.
While on ice, add 15 μL of the fragmentation mastermix to the well containing 35 μL double-stranded genomic DNA to achieve a total volume of 50 μL (see Note 5). Vortex gently before spinning down briefly.
-
5.
Place the reaction in the pre-cooled thermocycler and start the paused fragmentation program.
-
6.
While samples are undergoing fragmentation, prepare the End Repair and A-Tailing Buffer mastermix which contains 7 μL KAPA End Repair & A-tailing Buffer and 3 μL HyperPlus ERAT Enzyme Mix for each reaction to be performed (see Note 6).
-
7.
Following completion of the fragmentation reaction, place samples onto the plate cooler.
-
8.
Add 10 μL of the End Repair and A-Tailing Buffer mastermix to the well containing 50 μL of fragmented genomic DNA to achieve a total volume of 60 μL. Vortex gently before spinning down briefly.
-
9.
Incubate samples on a thermocycler using the selected End Repair and A-Tailing Buffer program (Table 2).
-
10.
While the End Repair and A-Tailing program is underway, prepare the Ligation mastermix which contains 30 μL Ligation buffer and 10 μL DNA Ligase for each reaction to be performed.
-
11.
Also, while the End Repair and A-Tailing program is underway, KAPA HyperPure Beads are removed from 4 °C to ensure they are equilibrated to room temperature in time for later paramagnetic bead clean up steps (see Note 7).
-
12.
Following completion of the End Repair and A-Tailing reaction, place samples onto the plate cooler.
-
13.
Add 10 μL of the universal adapter followed by 40 μL Ligation mastermix to each well to achieve a total volume of 110 μL (see Note 8). Vortex gently before spinning down briefly.
-
14.
Incubate samples on a thermocycler using the selected Adapter Ligation program (Table 3).
-
15.
While the Adapter Ligation program is running, remove the required number of UDI primer mixes from the freezer and thaw on ice.
-
16.
Following completion of the Adapter Ligation thermocycler program, remove samples from the thermocycler.
-
17.
Resuspend the room temperature KAPA HyperPure beads by vortexing vigorously.
-
18.
Perform a 0.8× bead cleanup by adding 88 μL of KAPA HyperPure beads to each well to achieve a total volume of 198 μL before pipette mixing ten times taking care not to generate bubbles.
-
19.
Incubate the bead/sample mixture for 15 min at room temperature to allow the DNA to bind to the beads (see Note 9).
-
20.
While the bead/sample mixture is incubating, prepare 20 mL of fresh 80% ethanol by adding 4 mL PCR grade water to 16 mL molecular grade ethanol. Vortex and leave at room temperature until required.
-
21.
While the bead/sample mixture is incubating, prepare 5 mL of fresh 10 mM Tris-HCl, pH 8.0, by adding 50 μL 1 M Tris-HCl, pH 8.0, to 4.95 mL PCR grade water. Vortex and leave at room temperature until required.
-
22.
Place samples onto a magnetic stand and wait approximately 5 min for the solution to clear (see Note 10).
-
23.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
24.
With the plate remaining on the magnetic stand, perform an ethanol wash by adding 200 μL of freshly prepared 80% ethanol (see Note 11).
-
25.
Incubate the sample in 80% ethanol for 30 s.
-
26.
Carefully remove the ethanol taking care not to disturb the pellet.
-
27.
Repeat steps 24–26 until a total of two ethanol washes have been performed.
-
28.
Remove residual ethanol without disturbing the beads.
-
29.
Air-dry the beads at room temperature to enable evaporation of any remaining ethanol (see Note 12).
-
30.
Remove the sample from the magnetic stand.
-
31.
Resuspend each bead pellet in 22 μL of 10 mM Tris-HCl, pH 8.0 (see Note 11).
-
32.
Incubate the sample for 2 min to enable DNA to elute from the beads.
-
33.
Place the sample on the magnetic stand to pellet the beads and for the solution to clear.
-
34.
With the plate remaining on the magnetic stand, transfer 20 μL of the eluate to a new 200 μL PCR plate.
-
35.
To the 20 μL of eluate, add 5 μL of KAPA UDI Primer mix to each individual sample library followed by 25 μL of KAPA HiFi HotStart ReadyMix. Vortex gently before spinning down briefly (see Note 3).
-
36.
Incubate samples on a thermocycler using the selected Pre-Capture PCR Amplification program (Table 4).
-
37.
Following completion of the Pre-Capture PCR Amplification program, remove samples from the thermocycler.
-
38.
To the 50 μL PCR reaction, add 70 μL of KAPA HyperPure beads before pipette mixing ten times taking care not to generate bubbles.
-
39.
Incubate the bead/sample mixture for 15 min at room temperature to allow the DNA to bind to the beads (see Note 9).
-
40.
Place samples onto a magnetic stand and wait approximately 3 min for the solution to clear (see Note 10).
-
41.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
42.
With the plate remaining on the magnetic stand, perform an ethanol wash by adding 200 μL of freshly prepared 80% ethanol (see Note 11).
-
43.
Incubate the sample in 80% ethanol for 30 s.
-
44.
Carefully remove the ethanol taking care not to disturb the pellet.
-
45.
Repeat steps 42–44 until a total of two ethanol washes have been performed.
-
46.
Remove residual ethanol without disturbing the beads.
-
47.
Air-dry the beads at room temperature to enable evaporation of any remaining ethanol (see Note 12).
-
48.
Remove the sample from the magnetic stand.
-
49.
Resuspend each bead pellet in 32 μL of 10 mM Tris-HCl, pH 8.0 (see Note 11).
-
50.
Incubate the sample for 2 min to enable DNA to elute from the beads.
-
51.
Place the sample on the magnetic stand to pellet the beads and for the solution to clear.
-
52.
With the plate remaining on the magnetic stand, transfer 31 μL of the eluate to a new 200 μL PCR plate, labelled “Master Plate” which is to be retained for preparation of the hybridization.
-
53.
From the 31 μL of the transferred eluate, remove 4 μL of the eluate and transfer to a new plate, labelled “QC Plate” for the purposes of quality control assessment. The Master Plate can be stored at −20 °C until ready to perform the hybridization. Proceed to Subheading 3.3 with the QC Plate.
3.3 Quality Control of DNA Libraries
-
1.
The concentration of each individual library is assessed using the Qubit broad range assay (see Note 13). Manufacturer guidelines are followed with two modifications: (1) the standard/sample is added to the Qubit assay tubes first followed by the Qubit working solution, and (2) the incubation time prior to reading the standard/sample is 20 min.
-
2.
The average fragment size of each individual library is assessed using the TapeStation D1000 assay. Manufacturer guidelines are followed without any modifications.
3.4 DNA Hybridization
-
1.
Thaw reagents required for the DNA hybridization step which include COT Human DNA, Universal Enhancing Oligos, Hybridization Buffer, and Hybridization Component H.
-
2.
Remove the KAPA HyperPure beads from 4 °C and allow to equilibrate to room temperature for 30 min.
-
3.
For the EuroClonality-NDC protocol, 22 clinical samples are pooled, in equal amounts, into one hybridization reaction to achieve a total of 1.5 μg of DNA (i.e., 68.2 ng of each individual library). To achieve this, calculate the volume of each library to enable 68.2 ng of each library to be added to the hybridization reaction. For the NTC, which should not have a measurable DNA concentration, the average volume of library being added from the 22 samples is calculated to determine the amount of volume of the NTC library to add (see Note 14).
-
4.
Label a LoBind DNA 1.5 mL tube and add the required volume of each of the 22 libraries and the NTC to this tube. Calculate the total volume of the 22 pooled libraries plus the NTC library. If the total volume of libraries is <45 μL (i.e., libraries), then PCR grade water is added to adjust volume to 45 μL.
-
5.
To the pooled libraries, add 20 μL COT Human DNA. Vortex gently before spinning down briefly.
-
6.
Calculate the total volume of the 22 pooled libraries, the NTC library plus the 20 μL of COT DNA. The volume of beads required in the next step is 2× this total volume (i.e., if the total volume was calculated to be 75 μL, then 150 μL KAPA HyperPure beads will be required).
-
7.
Vortex the KAPA HyperPure beads until a homogenous solution is achieved.
-
8.
To the pooled libraries, add the volume of KAPA HyperPure beads calculated in the step 6. Seal the tube and vortex vigorously for 10 s.
-
9.
Incubate the bead/sample mixture for 10 min at room temperature to allow the pooled libraries and COT Human DNA to bind to the beads.
-
10.
Place samples onto a magnetic stand and wait approximately 3 min for the solution to clear (see Note 10).
-
11.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
12.
With the plate remaining on the magnetic stand, perform an ethanol wash by adding 200 μL of freshly prepared 80% ethanol.
-
13.
Incubate the sample in 80% ethanol for 30 s.
-
14.
Carefully remove the ethanol taking care not to disturb the pellet. Remove residual ethanol with an additional pipetting step without disturbing the beads.
-
15.
Air-dry the beads at room temperature for approximately 5 min to enable evaporation of any remaining ethanol (see Note 12).
-
16.
Remove the sample from the magnetic stand.
-
17.
Add 13.4 μL of Universal Enhancing Oligos (UEO) to the tube, before sealing the tube and vortexing vigorously for 10 s to ensure a homogeneous mixture is achieved.
-
18.
To the library pool and UEO mixture, add 43 μL of mastermix prepared using the following components: 28 μL Hybridization Buffer, 12 μL Hybridization Component H, and 3 μL PCR grade water.
-
19.
Vortex before spinning down briefly. Incubate for 2 min at room temperature.
-
20.
Place samples onto a magnetic stand and wait approximately 3 min for the solution to clear.
-
21.
Transfer 56.4 μL of the eluate into a new well containing 4 μL of the EuroClonality-NDC panel.
-
22.
Vortex vigorously before spinning down briefly.
-
23.
Incubate samples on a thermocycler using the selected “Hybridization” program (Table 5).
-
24.
Dilute wash buffers provided in the KAPA HyperCapture Reagent Kit using the volumes of stock buffer solution and PCR grade water detailed in Table 6.
-
25.
Split the 400 μL of 1× Stringent Wash Buffer into two aliquots of 200 μL in 0.2 mL PCR tubes, and incubate on the thermocycler at 55 °C for at least 15 min.
-
26.
Place the 100 μL aliquot of 1× Wash Buffer I into the thermocycler at 55 °C for at least 15 min.
-
27.
Vortex the Capture Beads from the KAPA HyperCapture Bead kit thoroughly to ensure a homogenous solution.
-
28.
Remove 50 μL of Capture Beads for each pool and place into a 1.5 mL tube, and equilibrate to room temperature for 30 min.
-
29.
Following the 30-min incubation, place the tube containing Capture Beads onto a magnetic stand, and wait approximately 3 min for the solution to clear.
-
30.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
31.
Add a volume of 1× Bead Wash Buffer which is twice the original volume of Capture Bead used in step 28, to the pelleted Capture Beads.
-
32.
Remove from the magnetic stand and vortex for 10 s before spinning down briefly.
-
33.
Place the tube back onto the magnetic stand and wait until the beads have pelleted and the solution is clear.
-
34.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
35.
Perform a second wash of the Capture Beads by performing steps 31–34 again.
-
36.
Add a volume of 1× Bead Wash Buffer which is the same volume of Capture Beads used in step 28 to the pelleted Capture Beads.
-
37.
Remove from the magnetic stand and vortex for 10 s before spinning down briefly.
-
38.
Aliquot 50 μL of the resuspended Capture Beads into a 0.2 mL tube for each capture to be performed.
-
39.
Place the tube onto the magnetic stand and wait until the beads have pelleted and the solution is clear (see Note 15).
-
40.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
41.
The Capture Beads are now ready to bind the hybridized DNA.
-
42.
Proceed to the next step immediately to prevent the Capture Beads from drying out.
-
43.
Transfer the hybridization sample (60.4 μL) to the tube containing the pelleted Capture Beads from the previous step (see Note 16).
-
44.
Vortex for 10 s before spinning down briefly (see Note 17).
-
45.
Incubate the tube now containing the hybridized DNA and the Capture Beads on the thermocycler at 55 °C for 15 min.
-
46.
After the 15-min incubation, remove the samples and the tube containing the 100 μL aliquot of 1× Wash Buffer from the thermocycler.
-
47.
Add the 100 μL of 55 °C 1× Wash Buffer I before vortexing for 10 s and spinning down briefly.
-
48.
Place the tube onto the magnetic stand and wait until the beads have pelleted and the solution is clear.
-
49.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
50.
Remove a tube containing a 200 μL aliquot of 1× Stringent Wash Buffer from the thermocycler, and add the 200 μL of 1× Stringent Wash Buffer to the sample.
-
51.
Remove the sample from the magnet before vortexing for 10 s.
-
52.
Incubate the sample on the thermocycler at 55 °C for 5 min.
-
53.
After the incubation, spin the sample briefly before placing the sample onto the magnetic stand. Wait until the beads have pelleted and the solution is clear.
-
54.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
55.
Repeat steps 50–54 with the only remaining 200 μL aliquot of pre-warmed 1× Stringent Wash Buffer.
-
56.
Add 200 μL of room temperature 1× Wash Buffer I to the sample followed by vortexing for 10 s.
-
57.
Incubate the sample for 1 min at room temperature.
-
58.
After the incubation, spin the sample briefly before placing the sample onto the magnetic stand. Wait until the beads have pelleted and the solution is clear.
-
59.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
60.
Add 200 μL of room temperature 1× Wash Buffer II to the sample followed by vortexing for 10 s.
-
61.
Incubate the sample for 1 min at room temperature.
-
62.
After the incubation, spin the sample briefly before placing the sample onto the magnetic stand. Wait until the beads have pelleted and the solution is clear.
-
63.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
64.
Add 200 μL of room temperature 1× Wash Buffer III to the sample followed by vortexing for 10 s.
-
65.
Incubate the sample for 1 min at room temperature.
-
66.
After the incubation, spin the sample briefly before placing the sample onto the magnetic stand. Wait until the beads have pelleted and the solution is clear.
-
67.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
68.
Add 20 μL of PCR grade water to the sample followed by vortexing for 10 s and subsequently spin the sample briefly.
-
69.
Remove KAPA HyperPure Beads (70 μL required for each capture ) for use in later steps within a post-PCR area, and allow to equilibrate to room temperature.
-
70.
For each hybridization, add to a fresh 0.2 mL tube 25 μL of KAPA HiFi HotStart ReadyMix and 5 μL of Post-Capture PCR Oligos.
-
71.
Add the 20 μL of bead-bound captured DNA from step 68 to the 0.2 mL tube containing the PCR reagents to achieve a total volume of 50 μL. Mix thoroughly by pipette mixing.
-
72.
Within a post-PCR designated area, place samples on a thermocycler and run the selected Post-Capture PCR Amplification program detailed in Table 7 (see Note 18).
-
73.
While the Post-Capture amplification is underway, prepare 80% ethanol by adding 200 μL PCR grade water to 800 μL molecular grade ethanol in a 1.5 mL microfuge tube. Vortex gently before spinning down briefly.
-
74.
Following completion of the Post-Capture amplification thermocycler program, vortex the KAPA HyperPure Beads which are now equilibrated to room temperature.
-
75.
Add 70 μL KAPA HyperPure Beads to each 50 μL PCR reaction which contains the amplified and enriched DNA library pool before vortexing for 10 s and spinning down briefly.
-
76.
Incubate the bead/sample mixture for 15 min at room temperature to allow the sample to bind to the beads.
-
77.
Place samples onto a magnetic stand and wait approximately 3 min for the solution to clear.
-
78.
Carefully remove and discard the supernatant taking care not to disturb the pellet.
-
79.
With the plate remaining on the magnetic stand, perform an ethanol wash by adding 200 μL of freshly prepared 80% ethanol.
-
80.
Incubate the sample in 80% ethanol for 30 s.
-
81.
Carefully remove the ethanol taking care not to disturb the pellet.
-
82.
Repeat steps 79–81 for a total of 2 ethanol washes.
-
83.
Carefully remove any residual ethanol with an additional pipetting step without disturbing the beads.
-
84.
Air-dry the beads at room temperature for approximately 5 min to enable evaporation of any remaining ethanol (see Note 12).
-
85.
Remove the sample from the magnetic stand.
-
86.
Resuspend the bead pellet in 22 μL of PCR grade water before vortexing for 10 s and spinning down briefly (see Note 17).
-
87.
Incubate the sample for 2 min to enable DNA to elute from the beads.
-
88.
Place the sample on the magnetic stand to pellet the beads and for the solution to clear.
-
89.
With the plate remaining on the magnetic stand, transfer 20 μL of the eluate to a new 200 μL PCR plate.
-
90.
The amplified and enriched library is now ready for the final quality control steps prior to sequencing.
3.5 Quality Control of Enriched DNA Library
-
1.
The concentration of the amplified and enriched library is assessed using the Qubit high sensitivity assay. Manufacturer guidelines are followed with two modifications: (1) the standard/sample is added to the Qubit assay tubes first followed by the Qubit working solution, and (2) the incubation time prior to reading the standard/sample is 20 min.
-
2.
The average fragment size of each individual library is assessed using the High Sensitivity D1000 TapeStation assay. Manufacturer guidelines are followed without any modifications.
3.6 Sequencing of Enriched DNA Library
-
1.
Prepare the amplified and enriched pooled library for sequencing on the NextSeq 500/550 by adhering to the Illumina “Denature and Dilute Guidelines” using the following parameters (see Note 19):
-
(a)
Protocol A (Standard Normalization Method).
-
(b)
Final dilution of library is to 1.5 pM for Mid Output kits.
-
(c)
Final PhiX (sequencing control) spike-in percentage is 1% of the final library and PhiX composition.
-
(a)
3.7 Bioinformatic Analysis of the Sequencing Data Using ARResT/Interrogate
-
1.
The bioinformatics pipeline, ARResT/Interrogate, can be accessed at arrest.tools/interrogate and requires an account which can be created by emailing contact@arrest.tools (see Note 20).
-
2.
Once logged in to the ARResT/Interrogate bioinformatics website, switch to the “Interrogate.EC NDC” user mode at the top left of the user interface.
-
3.
Switch to the “processing” tab, and follow the instructions to upload your samples in compressed FASTQ format; the extension should be “.fastq.gz” (see Note 21).
-
4.
Following upload of the FASTQ files, no further options require selection.
-
5.
Click on blue “test it” button and if the subsequent response is “OK,” click on the green “process” button.
-
6.
Progress of the bioinformatic pipeline can be followed using the user interface or an email can be sent to notify user of completion of the analysis.
-
7.
When the run is complete, follow the instructions on the user interface to retrieve the result files.
-
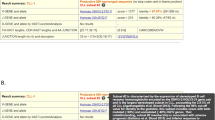
8.
The file with extension “.gathered.xlsx” contains results for samples contained within one analysis and is the main output. It contains panel-supported events, i.e., rearrangements, translocations, and somatic mutations (see Note 22).
-
9.
Open the file with extension “.gathered.xlsx” with Microsoft Excel or a suitable equivalent. Column descriptions are provided.
4 Notes
-
1.
An alternative method to address EDTA-containing genomic DNA samples is to add 5 μL of a conditioning solution, provided in the KAPA HyperPlus Kit to genomic DNA in a total volume of 30 μL. The conditioning solution, provided in the KAPA HyperPlus Kit, is diluted to a concentration dependent on the EDTA concentration in the DNA sample.
-
2.
NTC is either PCR grade water or the same buffer used to elute the gDNA following DNA extraction. Users can use commercially sourced high-molecular-weight gDNA as a positive control to monitor the consistency in performance of library preparation across multiple batches.
-
3.
It is worthwhile generating a template to record the order the samples are added to the 96-well plate and for later stages of library preparation to record the UDI primer mix assigned to each sample. Plates can be prepared the day before to minimize set up times on the day of library preparation.
-
4.
Fragmentation is key to the size distribution of the final library and is impacted by fragmentation time and temperature. With the wide range of different thermocyclers on the market, optimization of the fragmentation time is advised to ensure the ideal size distribution profile following fragmentation is achieved.
-
5.
Maintaining the temperature of the reaction at 4 °C during the setup of the fragmentation reaction is critical, and it is advised using a PCR cooler to ensure fragmentation does not being prior to loading the plate on the thermocycler.
-
6.
After thawing, the End Repair and A-Tailing buffer may contain precipitates which may require incubation at 37 °C and thorough vortexing before use to ensure they have been completely resuspended.
-
7.
Aliquots of KAPA HyperPure Beads can be made to reduce the number of times the KAPA HyperPure Beads are removed from storage at 4 °C.
-
8.
Universal adapter stocks are aliquoted to avoid repeated freeze/thaw cycles.
-
9.
We use a 15-min incubation period with beads to ensure maximal recovery of library in samples with a poor gDNA integrity.
-
10.
Various plate magnets are available on the market with different locations of the magnets and variable magnetic strengths. This variation can impact on the time required for beads to pellet, and it is worthwhile determining optimal times for incubation of bead containing samples for specific magnetic plates.
-
11.
Steps such as ethanol washes of beads and elution of beads should be performed using a multichannel pipette to ensure consistency and prevent beads drying out.
-
12.
Air-drying of beads is dependent on room temperature. The bead pellet should still be dark brown and glossy but show little sign of excess liquid. Over-drying of beads can lead to poor elution of DNA from the beads and therefore lower yields.
-
13.
With this version of the library preparation method, the concentration of individual libraries tends to lie within the range of the Qubit broad range assay.
-
14.
Inclusion of the positive control for the hybridization steps is not required. While the NTC is included in the hybridization reaction, the NTC DNA concentration should be negligible and not factored into the calculations for the combined DNA mass of 1.5 μg.
-
15.
Different magnetic stands will be required for the workflow and some will be required in both pre- and post-PCR environments. From our experience, sourcing good magnetic stands is essential to the success of the workflow.
-
16.
Keep the “Hybridization” program on thermocycler running for incubations with the Capture Beads and subsequent wash steps.
-
17.
This step is to ensure all the sample-bead mix is at the bottom of the well for the incubation steps on the thermocycler. Do not spin long enough for the beads to pellet at the bottom of the tube.
-
18.
It is advised the Post-Capture PCR and all further steps are performed in an area designated for post-PCR activities to minimize contamination risks.
-
19.
The EuroClonality-NDC protocol was developed and validated to achieve an optimal mean target coverage depth for the detection of clonal rearrangements, translocation , copy number alterations, and single nucleotide variant /indels in 22 samples, using a single hybridization reaction and sequenced on an Illumina NextSeq 500/550 system with the Mid Output kit sequencing reagents. It is at the readers’ discretion if they want to adapt the protocol to:
-
(a)
Employ different Illumina sequencing platforms.
-
(b)
Utilize sequencing reagents with increased output.
-
(c)
Alter the number of samples being applied to the flow cell for sequencing.
-
(a)
-
20.
The method of accessing the bioinformatics pipeline may change. The latest version will always be available either through the user interface of ARResT/Interrogate or through the authors.
-
21.
Dependent on the number of samples and quality of internet connection, this can take time.
-
22.
Additional files are available for in-depth analysis such as a comprehensive QC metrics summary, BAM files, CNV files and VCF files.
References
Swerdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R et al (2016) The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 127(20):2375–2390. https://doi.org/10.1182/blood-2016-01-643569
van Dongen JJ, Langerak AW, Bruggemann M, Evans PA, Hummel M, Lavender FL et al (2003) Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17(12):2257–2317. https://doi.org/10.1038/sj.leu.2403202
Langerak AW, Groenen PJ, Bruggemann M, Beldjord K, Bellan C, Bonello L et al (2012) EuroClonality/BIOMED-2 guidelines for interpretation and reporting of Ig/TCR clonality testing in suspected lymphoproliferations. Leukemia 26(10):2159–2171. https://doi.org/10.1038/leu.2012.246
Pestinger V, Smith M, Sillo T, Findlay JM, Laes JF, Martin G et al (2020) Use of an integrated pan-cancer oncology enrichment next-generation sequencing assay to measure tumour mutational burden and detect clinically actionable variants. Mol Diagn Ther 24(3):339–349. https://doi.org/10.1007/s40291-020-00462-x
Bratman SV, Newman AM, Alizadeh AA, Diehn M (2015) Potential clinical utility of ultrasensitive circulating tumor DNA detection with CAPP-Seq. Expert Rev Mol Diagn 15(6):715–719. https://doi.org/10.1586/14737159.2015.1019476
Cheng DT, Mitchell TN, Zehir A, Shah RH, Benayed R, Syed A et al (2015) Memorial sloan kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J Mol Diagn 17(3):251–264. https://doi.org/10.1016/j.jmoldx.2014.12.006
Richter J, Schlesner M, Hoffmann S, Kreuz M, Leich E, Burkhardt B et al (2012) Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nat Genet 44(12):1316–1320. https://doi.org/10.1038/ng.2469
Lopez C, Kleinheinz K, Aukema SM, Rohde M, Bernhart SH, Hubschmann D et al (2019) Genomic and transcriptomic changes complement each other in the pathogenesis of sporadic Burkitt lymphoma. Nat Commun 10(1):1459. https://doi.org/10.1038/s41467-019-08578-3
Karube K, Enjuanes A, Dlouhy I, Jares P, Martin-Garcia D, Nadeu F et al (2018) Integrating genomic alterations in diffuse large B-cell lymphoma identifies new relevant pathways and potential therapeutic targets. Leukemia 32(3):675–684. https://doi.org/10.1038/leu.2017.251
Gonzalez D, Martinez P, Wade R, Hockley S, Oscier D, Matutes E et al (2011) Mutational status of the TP53 gene as a predictor of response and survival in patients with chronic lymphocytic leukemia: results from the LRF CLL4 trial. J Clin Oncol 29(16):2223–2229. https://doi.org/10.1200/JCO.2010.32.0838
Rossi D, Rasi S, Spina V, Bruscaggin A, Monti S, Cresta S et al (2012) The genome of chemorefractory chronic lymphocytic leukemia reveals frequent mutations of NOTCH1 and SF3B1. Leukemia Suppl 1(Suppl 2):S26–S28. https://doi.org/10.1038/leusup.2012.16
Oscier DG, Rose-Zerilli MJ, Winkelmann N, Gonzalez de Castro D, Gomez B, Forster J et al (2013) The clinical significance of NOTCH1 and SF3B1 mutations in the UK LRF CLL4 trial. Blood 121(3):468–475. https://doi.org/10.1182/blood-2012-05-429282
Weng AP, Ferrando AA, Lee W, Morris JP, Silverman LB, Sanchez-Irizarry C et al (2004) Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306(5694):269–271. https://doi.org/10.1126/science.1102160
Zhu YM, Zhao WL, Fu JF, Shi JY, Pan Q, Hu J et al (2006) NOTCH1 mutations in T-cell acute lymphoblastic leukemia: prognostic significance and implication in multifactorial leukemogenesis. Clin Cancer Res 12(10):3043–3049. https://doi.org/10.1158/1078-0432.CCR-05-2832
Malcikova J, Tausch E, Rossi D, Sutton LA, Soussi T, Zenz T et al (2018) ERIC recommendations for TP53 mutation analysis in chronic lymphocytic leukemia-update on methodological approaches and results interpretation. Leukemia 32(5):1070–1080. https://doi.org/10.1038/s41375-017-0007-7
Bystry V, Reigl T, Krejci A, Demko M, Hanakova B, Grioni A et al (2017) ARResT/Interrogate: an interactive immunoprofiler for IG/TR NGS data. Bioinformatics 33(3):435–437. https://doi.org/10.1093/bioinformatics/btw634
Stewart JP, Gazdova J, Darzentas N, Wren D, Proszek P, Fazio G (2021) EuroClonality-NGS Working Group. Validation of the EuroClonality-NGS DNA capture panel as an integrated genomic tool for lymphoproliferative disorders. Blood Adv 5(16):3188–3198. https://doi.org/10.1182/bloodadvances.2020004056 PMID: 34424321; PMCID: PMC8405189
Acknowledgments
We would like to acknowledge the EuroClonality-NGS working group and, in particular, those members that directly or indirectly contributed to the validation of the EuroClonality-NDC assay. The EuroClonality-NGS Working Group is an independent scientific subdivision of EuroClonality that aims at innovation, standardization, and education in the field of diagnostic clonality analysis. The revenues of the previously obtained patent (PCT/NL2003/000690), which is collectively owned by the EuroClonality Foundation and licensed to InVivoScribe, are exclusively used for EuroClonality activities, such as for covering costs of the Working Group meetings, collective Work Packages, and the EuroClonality Educational Workshops. The EuroClonality consortium operates under an umbrella of ESLHO, which is an official EHA Scientific Working Group.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this protocol
Cite this protocol
Stewart, J.P. et al. (2022). Immunoglobulin/T Cell Receptor Capture Strategy for Comprehensive Immunogenetics. In: Langerak, A.W. (eds) Immunogenetics. Methods in Molecular Biology, vol 2453. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2115-8_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2115-8_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2114-1
Online ISBN: 978-1-0716-2115-8
eBook Packages: Springer Protocols