Abstract
Research on the innate immunity has accelerated over the last decades. The main reason for this has been the discovery of receptors recognizing danger molecules from pathogens. This has been facilitated through genome and transcriptome sequencing of different fish species. Also, endogenous host molecules from sterile physiological insults may also bind to certain receptors and induce immunological processes. The magnitude and quality of adaptive immunity are known to be dependent on the instructions the innate response gives. This chapter gives an overview of selected innate immune organs/tissues, factors, and processes that have been suggested to possess important roles during innate immune response in fish.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Innate immunity
- Fish
- TLR
- Complement
- Cytokines
- Acute-phase proteins
- Antimicrobial peptides
- Chemokines
- Epigenetics
2.1 Introduction
2.1.1 Innate Immunity: The Concept
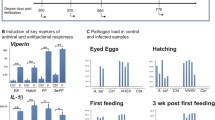
Innate immune defense is important for protecting the host from infection, not only in naïve fish but also in fish that have previously been infected. “Innate immunity has shed its older, disparaging title of ‘non-specific immunity’ and now stands as a proud partner with the adaptive immune system in protecting human hosts from infectious insults. For any who doubt the impressive protective capacity of the innate immune system, it is instructive to consider that only vertebrates boast the added benefits of an adaptive immune system, leaving most organisms on our planet to survive on innate immunity alone” (Turvey and Broide 2010). Indeed, this applies also to fish. The immune system of teleost fish is composed of two kinds of receptor types: The germline-encoded pattern recognition receptors (PRRs) and the antigen-specific receptors are made from gene arrangement after, e.g., pathogen infection. The latter consist of, e.g., antibodies, MHC I and MHC II, and T-cell receptors. In addition, numerous other receptors/molecules can take part in the innate immunity. The innate mechanisms can be divided into constitutive and inducible. The former represents rapid ongoing ligand binding to receptors and a quick response, while the inducible (e.g., many PPRs) acts slower—but with a higher magnitude (Paludan et al. 2020) (Figs. 2.1 and 2.2).
Simplified illustration shows how inducible innate immunity changes over time, whereas the constitutive is stable vs. time. To get complete sterilization and resolution, both inducible and constitutive innate immune responses plus antibodies are often needed. Epigenetic changes may contribute to better fitness/increased protection when fish are exposed to a second infection. Targeted gene expression surveys or transcriptomics has focused primarily on describing or identifying inducible genes (e.g., DEGs), while in contrast, factors contributing to the constitutive arm have been poorly described
Hypothetical time-course study of a gene (qPCR) expression. The orange line represents a gene with constitutive expression upon treatment with stimulant 1, and the black line demonstrates an early expression of the same gene after the fish were treated with stimulant 2, whereas the dotted red line represents a delayed expression of this gene after the fish were treated by stimulant 3. A specific gene may be induced by a certain stimulant and not by others, or there may be a stable, rapid, or delayed induction. The magnitude of induction may likely be dependent on number of specific receptors on cells or/and the number of cells that harbor specific receptors
2.1.2 Innate Receptors
Innate and adaptive immunity can cooperate to clear the infections. Central receptors in the early innate responses are so-called Toll-like receptors (TLRs) and are vital for the communication between the innate and adaptive branches (Rivera et al. 2016). The germline-encoded pattern recognition receptors (PRRs) are central in the recognition of microbial components and for the activation of innate immunity, which may induce inflammatory response to eliminate pathogens. The PRRs, expressed in innate immune cells, include receptors such as Toll-like receptors (TLRs), RIG-I-like receptors (RLRs), NOD-like receptors (NLRs), and C-type lectin receptors (CLRs). Upon recognition of microbial components known as pathogen-associated molecular patterns (PAMPs), PRRs induce intracellular signaling networks to activate transcription factors that regulate genes involved in inflammatory responses. Importantly, these innate immune signals also trigger dynamic chromatin changes. Such changes may in turn induce modulated gene-specific expression patterns resulting in even more pathogen elimination.
2.2 Cells in the Innate Immune Response
The traditional view that the adaptive and innate immune defense is divided into two compartments is now more or less history. The fish innate immune cells comprise not only the “traditional” innate cells such as macrophages (Kordon et al. 2018; Rieger and Barreda 2011; Grayfer et al. 2014, 2018) and granulocytes (Pijanowski et al. 2013; Schmidt 1905) but also red blood cells (Puente-Marin et al. 2018, 2019b; Dahle et al. 2015; Wessel et al. 2015), thrombocytes (Stosik et al. 2019), B cells (Wu et al. 2020), and subtypes of T cells (Scapigliati et al. 2018).
2.2.1 Monocytes/Macrophages
Monocytes are large mononuclear circulating leukocytes, which become macrophages when they settle tissues and organs. The nucleus may display different oval-, kidney-, or bean-shaped conformations, while the cytoplasm is usually pale and agranular, with varying amounts of vesicles and lysosomes. Macrophages are one of the main immune cells performing phagocytosis, the other presumably being the neutrophilic granulocytes. Phagocytosis is a multistage process for removal and cellular ingestion and destruction by intracellular enzymes and other substances (Grayfer et al. 2014; Hodgkinson et al. 2015). In addition to being professional phagocytes, macrophages can also function as professional APCs by presenting antigen to T cells on MHC class II. Such functions have also been suggested from studies on fish monocytes/macrophages (Sugamata et al. 2009; Wittamer et al. 2011). It has been suggested that fish display at least three different phenotypes of macrophages, based on their activation processes: innately, classically, and alternatively activated macrophages (Wentzel et al. 2020; Hodgkinson et al. 2015). Innate activation occurs when a macrophage receives a stimulus from the recognition of a microbial substance (e.g., PAMP) through cell receptor(s) without any need for any co-stimulation. Classical activation, however, occurs with the combination of such a stimulus and the cytokine interferon-gamma (IFN-gamma). Both innate activation and classical activation typically lead to increased pro-inflammatory response as opposed to alternative activation. A presence of cytokines (interleukin 4 (IL4) and/or interleukin 13 (IL13)) induces a macrophage phenotype with a resolving function (wound healing and tissue repair). There is also a suggestion for a fourth type of macrophages, namely regulatory macrophage. Regulatory activation is associated with the cytokine interleukin 10 (IL10) and important for downregulation of the inflammatory process (Wiegertjes et al. 2016). However, macrophages are apparently able to change between different phenotypes, and there is still some uncertainty whether all these activation pathways perform the same way in fish, as in mammals (Forlenza et al. 2011). Please see Chap. 6 for a more thorough overview of macrophages in fish.
2.2.2 Dendritic Cells
Dendritic cells (DCs) are categorized as a professional APCs found within several different tissues and are very effective at initiating both innate and adaptive immune responses in mammals (Banchereau et al. 2000; Worbs et al. 2017). DCs are typically small cells, with several elongated, cytoplasmic processes (dendrites) that increase the cell surface area (Collin and Bigley 2018). Cell populations with DC-like morphology and functions have been reported from teleost fish (Shao et al. 2015; Bassity and Clark 2012; Haugland et al. 2012), but due to lack of specific markers it is currently unknown whether they are true homologs of the mammalian cell type.
2.2.3 Granulocytes
Granulocytes are leukocytes with cytoplasmic granules and often nucleus with varying shapes (lobes) (Flerova and Balabanova 2013). They are central pool of the innate immune cells (Lieschke and Trede 2009). The granulocytes have traditionally been grouped into neutrophilic granulocytes (neutrophils), eosinophilic granulocytes (eosinophils), and basophilic granulocytes (basophils), based on their staining characteristics with different dyes. However, this classification was originally developed for use in mammalian hematology and does not appear to always correlate well with characteristics of fish granulocytes (Kelenyi and Nemeth 1969; Drzewina 1909; Rombout et al. 2005).
2.2.3.1 Neutrophils
Neutrophils typically possess nucleus with varying degrees of lobulation and contain granules that usually do not display marked affinity for staining with basic or acid dyes (such as hematoxylin and eosin). Neutrophils are generally most abundant between the granulocytes. In mammals, neutrophils are very mobile and are usually among the first cells to infiltrate tissue during onset and early phases of inflammation (Rosales 2018). Similar cellular recruitment speed has also been reported from teleosts (Lamas and Ellis 1994; Katzenback and Belosevic 2009; Havixbeck and Barreda 2015). Neutrophils are armed with a diverse arsenal of cellular weapons, making them effective combatants against invading pathogens (Havixbeck and Barreda 2015). Like macrophages, they are able to degrade ingested microbes and particles through production and release of reactive oxygen species (ROS) and proteases (Katzenback and Belosevic 2009; Rieger and Barreda 2011). They can also release different granules, upon degranulation, containing antimicrobial proteins and enzymes such as myeloperoxidase (MPO) (Lieschke and Trede 2009). In addition, neutrophils are able to form extracellular traps, which contain antimicrobial factors (Palic et al. 2007; Pijanowski et al. 2013; Chi and Sun 2016; Zhao et al. 2017; Van et al. 2020).
Eosinophils, or acidophils, are described to contain cytoplasmic granules that stain bright red with the acidic dye (eosin). However, cellular identification based solely on cytochemical and/or histochemical staining characteristics may lead to misinterpretation as basophils, eosinophilic granule cells, mast cells, some neutrophils (also called heterophils), and rodlet cells also are capable to be dyed to various degrees. Consequently, it has been suggested that mammalian terminology should be used whenever possible for describing these cell types (Watanabe et al. 1997; Suljevic et al. 2017). In mammals, the eosinophils have immunological roles regarding both immune regulation, defense against parasitic infections, and allergic inflammatory reactions (Hogan et al. 2008). Teleost eosinophils have been reported to be phagocytic (Watson et al. 1963), and they increase in cell numbers and increase the degranulation activity as a response to infection (Balla et al. 2010).
2.2.3.2 Basophils
Basophils are large granulocytes with staining of their cytoplasmic granules with a basic dye (hematoxylin). Basophils are rarely observed in teleost species (Tavares-Dias 2006). Their granules contain histamine, an inflammatory mediator, and basophils are associated with anaphylaxis, allergy, and hypersensitivity reactions (Chirumbolo 2012). As such, they are similar to the mast cells. Although not fully established, these granulocytes might also have other functions within the fish immune system (Odaka et al. 2018).
2.2.3.3 Eosinophilic Granule Cells
Eosinophilic granule cells (EGCs) and rodlet cells have been observed in fish (Reite and Evensen 2006). Such cells resemble the classical mast cells (Reite 1998). Teleost EGCs have been identified in several species, as part of the host inflammatory response to injected vaccines, bacterial infection, parasite infestations, or other types of noxious stimuli (Rombout et al. 2011).
2.2.4 Thrombocytes
Thrombocytes are oval-shaped, nucleated, and agranular cells located in fish. In some fish species, thrombocytes have been shown to be phagocytic and it has been discussed whether thrombocytes can function as APCs and/or is coupled to the innate immunity (Stosik et al. 2019; Passantino et al. 2005).
2.2.5 Red Blood Cells
From a RNAseq study on trout red blood cells exposed to either poly I:C, it was found that the cells expressed numerous transcripts of immune molecules—such as ifna, tlr3, tlr9, mx, and ccl4 (Morera et al. 2011). Thus, the authors suggested that red blood cells indeed participate in innate immune response.
2.3 Epigenetic Control of Innate Immunity
2.3.1 Epigenetics: The Concept
Epigenetics involves heritable factors that regulate spatiotemporal genome expression, which may induce different phenotypes. Two of the molecular mechanisms, histone modifications and DNA methylation, regulate gene expression at the chromatin level. In contrast, microRNAs are molecules that affect gene expression at the posttranslational level. Epigenetic histone modification involves acetylation/deacetylation, methylation/demethylation, and phosphorylation/dephosphorylation of specific histone amino acids. Pathogens have evolved a variety of strategies to modify host epigenetics. For example, they can (1) directly modify host proteins and chromatin, (2) attenuate PRR binding and signaling pathways, and (3) modulate the expression of activators and repressors of innate immunity. Hosts can abrogate pathogen-induced epigenetic changes to maintain their innate defense characters (Zhang and Cao 2019). Analysis of posttranslational processes on immunity has not generally been well studied in fish. However, in one study, the impact of histone modification after infectious necrosis virus infection (IPNV) and temperature control has been shown (Boltana et al. 2018). In this study, IPNV-infected fish that preferred a given temperature showed histone modification, which could explain modulated expression of il1, il2, ifng, and ifnrg receptor. The pattern of histone modification was different from IPNV-infected fish kept at constant temperature. In another study, spring viremia of carp virus (SVCV) infection induced histone modification in zebrafish (Danio rerio). The authors indicated that the ifn, tlr, and C-reactive protein promoters were methylated postinfection; thus, these genes were upregulated compared to controls (Medina-Gali et al. 2018). Since epigenetic modification of the genome is a heritable trait, epigenetic programming of brood stockfish, by, e.g., immunostimulants, may be a viable approach to produce offspring with higher innate disease resistance (Zhang et al. 2019).
2.3.2 Micro RNA
MicroRNAs (miRNAs) are a family of small noncoding RNAs that play vital roles in modulating host immune response. Accumulating evidence demonstrates that host miRNAs are involved as mediators in regulating viral replication and host antiviral immunity in mammals. In a miiuy croaker macrophages, miR-3570 that was upregulated after rhabdovirus infection interfered and led to downregulation of type I interferon in the cells. In turn, this downregulation caused increased virus replication in cells (Xu et al. 2018a). Binding to Toll-like receptors (TLRs) and subsequent intracellular signaling may also bring about production of microRNA. This may result in a positive or negative feedback loop system regulating immune response. More on this complex issue is described in a review authored by Zhou et al. (2018).
2.3.3 Long Noncoding RNA
lncRNAs have been demonstrated to play pivotal roles in various biological processes, especially gene expression regulations, including transcriptional regulation, posttranscriptional control, and epigenetic processes. The functional significance of lncRNA lags far behind what is the status on mammals. However, a novel lncRNA (SETD3-OT) in turbot (Scophthalmus maximus) has been identified. From the annotation of neighboring adjacent genes, SETD3-OT might be involved in the regulation of cell apoptosis and cycle, the immune cell development, and the immune response against infection. The expression pattern of SETD3-OT was similar to the majority of the neighboring genes following Aeromonas salmonicida challenge. The SETD3-OT expression was high levels in mucosal surfaces in controls fish (intestine, gill, and skin), but was downregulated following Vibrio anguillarum infection (Yang et al. 2020). In another study, Nodavirus infected European sea bass (Dicentrarchus labrax) displayed many putative lncRNA, suggested to possibly be involved in immune responses (Pereiro et al. 2020). Other studies have also suggested lncRNA to be involved in the regulation of immune responses (Boltana et al. 2016; Valenzuela-Miranda and Gallardo-Escarate 2016; Paneru et al. 2016; Valenzuela-Munoz et al. 2018, 2019).
2.3.4 Small Interfering RNA and Circular RNA
In addition to microRNA and lncRNA, the methylation of mRNA, occurrence of small interfering RNAs, and circular RNAs may all contribute to epigenetic modulation of gene expression in vertebrates, including fish (Wang et al. 2018a). Olive flounder (Paralichthys olivaceus) experimentally infected with Edwardsiella tarda showed differentially expressed circRNA. The authors suggested that these belonged to the circRNA-miRNA-mRNA network, where KEGG analysis indicated that they were part of the Herpes simplex infection and intestinal immune network for IgA production (Xiu et al. 2019). Another study showed that circRNAs are involved in mammalian antiviral immunity (Wang et al. 2017). KEGG (Kyoto Encyclopedia of Genes and Genomes; www.genome.jp), a huge database integrating genomic, chemical, and systemic functional information, is often used to find what cellular networks/pathways the DEGs belong to. It refers to what is described in and annotated from human/mice systems.
2.4 Mucosal Innate Defense
2.4.1 Innate Immune Molecules of the Fish Skin
The skin of fishes protects fish from external pathogens. The outermost layer is mainly composed of epithelial cells, termed keratocytes. These cells cover scales and are highly phagocytic toward certain particles. They are also motile. The motility of fish keratocytes is studied in a number of fish species (Asbakk and Dalmo 1998; Tsuchida and Theriot 2013; Galbraith and Sheetz 1998; Jurado et al. 2005; Okimura et al. 2018; Ream et al. 2003). There has been limited research on the production of innate defense factors, but this topic deserves more attention. Whether the cells possess phagocytic receptors is not known, it seems that the cells are able to discriminate the uptake dependent on the kind of bacteria (Karlsen et al. 2012). The skin mucus contains an array of molecules enabling protection from pathogens. In a study on yellow catfish (Pelteobagrus fulvidraco), 133 differentially expressed proteins were found after bath infection with Edwardsiella ictaluri. A minority of these differentially expressed proteins were directly immune-related. Examples of the upregulated genes were complement component c3, MAP kinase 1, and interferon-induced 35 kDa protein (Xiong et al. 2020).
Among the antibacterial enzymes, the best studied in the fish skin is lysozyme. Lysozyme is a glycoside hydrolase that catalyzes the hydrolysis of 1,4-beta-linkages between N-acetylmuramic acid and N-acetyl-D-glucosamine residues in peptidoglycan, which is the major component of gram-positive bacterial cell wall. However, it seems that lysozyme-like enzymes have activity also against gram-negatives, parasites, and virus, as reviewed by Dash et al. (2018). This review also contains detailed description of other skin-related innate immune factors (Dash et al. 2018).
How the mucus is obtained, for concomitant analysis of factors, will inevitably decide which substances will be found during a screening process. As an example: If the mucus sample contains cells or scales, it is clear that the samples also contain cellular factors and most probably also immune factors normally localized in deeper layers (e.g., connective tissue and muscle). The most gentle and sensible protocol is to adsorb the mucus using a tissue paper. While a wiping method using tissue paper also gives a good protein yield, this method comes with some degradations. If the research requires a high mucus yield together with substances from the epithelial layer, the wiping method is preferable (Faeste et al. 2020). It is quite difficult to discriminate between substances normally found in mucus compared to what is intracellularly or extracellularly localized in epidermis, subdermis, and connective tissue. Thus, many reports describe the presence of substances not (only) found in the mucus itself but also found in the underlaying tissue. As an example of the latter, the transcriptomic analysis of a skin sample (3 × 1 cm) from large yellow croaker (Larimichthys crocea) followed by Cryptocaryon irritans challenge revealed up to 1055 DEGs (differentially expressed genes) (96 h postinfection). Since many of the DEGs were clearly innate immune-related, it would have been interesting to see how many and which transcripts were from epithelial cells, connective tissue cells, blood cells, and muscle cells, respectively. Probably, a similar sampling protocol was followed by Liu et al. (2020a, b) where zebrafish were challenged with spring viremia of carp virus (SVCV)—causing skin lesions. This study revealed 320 DEPs (differentially expressed proteins) (48 h postinfection) and 181 DEPs (96 h postinfection). Sixteen of these were confirmed by means of QPCR analysis (Bai et al. 2020). DEPs often found are complement factors, and chemokines, heat shock proteins, MHC, cell adhesion molecules, TNF-induced protein, and many more were regulated (Liu et al. 2020b). In conclusion, analysis of skin innate defense mechanisms should discriminate between mucus itself, epidermis, subdermis, and connective tissue.
The epidermis consists of keratocytes, which are highly mobile cells and also possess (mostly overlooked) phagocytic activity (Asbakk and Dalmo 1998; Sveen et al. 2020). The immunological significance of their phagocytic ability is not yet fully understood. One theory is that they engulf as many particles they can before going into a cell death pathway and are sloughed off from the epidermis (Asbakk and Dalmo 1998). It is speculated that these cells possess some innate defense mechanisms (e.g., receptors) (Lindell et al. 2012). The epithelial layer of the fisheye cornea consists of cells that highly resemble skin keratocytes. These cells are not studied with respect to their innate defense abilities. We have preliminary results showing that these cells also engulf foreign particles (Fig. 2.3). For more details on mucosal immunity in fish, see Chap. 12.
2.4.2 Nasopharynx-Associated Lymphoid Tissue (NALT)
NALT has been discovered to harbor lymphocytes, but also genes central in induction of innate immunity. These include mx1, tlr3, il1r, il8, tnfr, myD88, c3, c4, c7–1, cxc9, cxcl9, cathl1, ccl19, and il6 (Tacchi et al. 2014; Yu et al. 2018). The significance of NALT-mediated innate response, compared to, e.g., skin or intestine, is not clear. Another assemblance of lymphoid cells can be found in the buccal cavity of rainbow trout (Oncorhynchus mykiss) that have been infected by Flavobacterium columnare. After infection, this buccal cavity lymphoid tissue was found to express innate factors such as il8, il1b, chemokine like 19, cathl1 and cathl2, rig1, among other adaptive immune genes (Xu et al. 2020).
2.4.3 Gills in Innate Immunity
Gill-associated lymphoid tissue (GIALT) has been characterized in Atlantic salmon and different fish species (Resseguier et al. 2020; Haugarvoll et al. 2008). This tissue was, in Atlantic salmon, identified to express upregulated genes such as complement component c3, il18, mx3, il20, ifn type II, viperin, rig1, and ifna after ISAV challenge (Austbo et al. 2014; Valenzuela-Miranda et al. 2015). Pro-inflammatory (il6, il17c1) and anti-inflammatory (il10, tgfb) genes have been found, in rainbow trout gills, after Ich (Ichthyophthirius multifiliis) infection (Syahputra et al. 2019). Another study aimed at doing a transcriptomic survey of Atlantic salmon gills suffering from multifactorial pathologies. Genes that were differentially expressed were depicted to be involved in pathways such as cellular immune response (IL-17 signaling, IL-6 signaling, granzyme A signaling, crosstalk between dendritic cells and natural killer cells, granulocyte adhesion and diapedesis, and HMGB1 signaling), cytokine signaling (IL-17 signaling, IL-6 signaling, acute-phase response signaling, role of JAK family kinases in IL-6 type cytokine signaling, TNFR2 signaling, and HMGB1 signaling), and tissue damage and repair (Krol et al. 2020). Some of these genes possess central functions in innate immunity. More details on the gills’ function in the immune response, please see Chap. 1.
2.4.4 Intestine in Innate Immunity
During the recent years, many excellent review articles describing the fish’s intestinal immunity have been published (Dawood 2020; Nadal et al. 2020; Sitja-Bobadilla et al. 2016; Brugman 2016; Dezfuli et al. 2016; Scapigliati et al. 2018; Brinchmann et al. 2018). Recently, there have been many innovative approaches to better understand intestinal immunity. In one of these studies, proteomic and transcriptomic examination of the intestinal mucus in Tilapia infected with Streptococcus agalactiae showed that innate factors such as c1r-like EGF domain, c1q-binding protein, hsp1, hsp90b, galectin, and membrane attack complex component/perforin domain, conserved site, complement factor D, C-type lectin fold, il1, il1r, and foxp3 (Wu et al. 2016). Another study, in grass carp, did a transciptomic and proteomic examination of the intestine after oral DNA vaccination (Li et al. 2020). The study revealed 250 and 50 immune-related DEGs and DEPs, respectively, after the oral vaccination. KEGG enrichment analysis showed genes and proteins participating in the Toll-like receptor signaling pathway, MAPK signaling pathway, NOD-like receptor signaling pathway, and the complement cascade were present both in the mucous and tissue homogenates. It is obvious that the intestinal innate mechanisms are quite diverse. More research using modern omic technologies will inevitably give us more information about the significance of the various intestinal innate factors that have on disease resistance.
Recently, a lymphoid structure in the cloacal region was discovered in Atlantic salmon (Loken et al. 2020). This may be the same as Inami et al. (2009) described in Atlantic cod—although the work on cod did not specify the anatomical localization properly (Inami et al. 2009). Whether this collection of lymphatic cells has any function in innate defense is not clear. However, through gene expression studies, genes encoding il1b, il8, il10, hepcidin, and ccl19 were found. These genes likely play roles in the innate immunity.
Future studies of intestinal mucus must be carefully planned and executed to avoid contaminant cells and blood. This may give false assumptions with regard to the actual presence of innate factors.
2.5 Innate Defense Mechanisms in Muscle Tissues
Skeletal muscles have been implicated in several atypical physiological processes including immune response, especially after pathogen challenge. When zebrafish were intramuscularly challenged by Salmonella enterica, pro-inflammatory il1b and tnfa were highly expressed in their skeletal muscle. Likewise, hep (hepcidin) and il10 were also expressed (Chatterjee et al. 2016). The authors did not examine any presence of leukocytes in muscle tissue samples after the fish were challenged with Salmonella. Thus, it is likely that inflammatory leukocytes did contribute to the expression of the pro-inflammatory cytokines. The contribution of incoming leukocytes to the inflammatory event is discussed by Valenzuela et al. (2017) and Kaitetzidou et al. (2012). Similar to the induction of innate immune genes in skeletal muscle, Atlantic salmon infected with salmonid alphavirus and piscine reovirus showed altered gene expression in the heart tissue (Johansen et al. 2015). In this study, several innate genes were expressed in the heart muscle. The authors did not elaborate whether this contribution was caused by inflammatory cells or not.
2.6 Innate Defense Mechanisms in Kidney and Spleen
The kidney and spleen are hematopoietic organs capable of inducing and exerting innate immune responses (Uribe et al. 2011; Svingerud et al. 2012; Kumar et al. 2018). The head kidney is the principal immune organ responsible for phagocytosis, antigen processing, and formation of Igm and immune memory (Page et al. 2013; Stosik et al. 2018; Rauta et al. 2012; Rombout et al. 2005; Kim et al. 2017). Kidneys in fish are paired and have a Y shape along the body axis. The immune relevant part, the head kidney, is located anteriorly. The posterior is mostly the renal system. The form of the head kidney varies between species. In some species, there are two separate extensions in the most anterior part of the organ, while in salmonid species the kidney is present as a single organ (Press and Evensen 1999). It is acknowledged that the head kidneys’ main function is hematopoiesis of lymphocytes, phagocytosis, antigen presentation, and maturation of lymphocytes. Its significance in innate immunity is not very well researched yet, although the head kidney leukocytes are armed with innate factors (Aballai et al. 2017; Gerdol et al. 2015; Cao et al. 2020; Hwang et al. 2017; Rozas-Serri et al. 2019; Zhou et al. 2019). It should be clear that cells in the posterior part of the kidney also have capability to express immune genes after pathogen challenge, as reported by Sudhagar et al. (2019).
2.7 Innate Defense Mechanisms in the Spleen
It is acknowledged that the main functions of the spleen are in hematopoiesis of lymphocytes, antigen trapping, and destruction of red blood cells (Press and Evensen 1999). However, from RNAseq analysis it is evident that the spleen cells contain and express numerous innate immune genes such as those involved in chemokine signaling, Toll-like receptor signaling, RIG-1, and NOD-mediated signaling and complement cascade (Ali et al. 2014).
2.8 Innate Defense Mechanisms in the Liver
The liver is acknowledged to produce acute-phase proteins, including complement components following infection or physiological insult. An array of innate defense factors has been found following a transcriptomic study of rainbow trout. This study revealed transcripts coding for genes important in acute-phase response, inflammatory response, genes coding for PAMP-binding receptors, and molecules central in chemotaxis (Martin et al. 2010). This finding suggests that the liver also has capacity to mount innate responses.
2.9 Receptors and Molecules of the Innate Immune Defense
Innate immunity is orchestrated by numerous molecules such as cytokines, complement factor, and receptors. Many molecules participate in both innate immunity and adaptive immunity. The following chapters describe the roles of selected innate molecules that have been ascribed to innate immunity—as central components.
2.9.1 Toll-Like Receptors (TLRs) as Pattern Recognition Receptors (PPRs)
The number of TLRs adds to other pattern recognition receptors (Tribouley et al. 1978) (including splice variants) such as different C-type lectin receptors, NOD-like (nucleotide-binding oligomerization domain-like) receptors (NLRs), RIG-1-like receptors, and scavenger receptors (Brubaker et al. 2015), and suggests that fish may very well be equipped with innate receptors that may likely be targets for innate immune training. TLRs are a family of pattern recognition receptors that bind pathogen-associated molecular patterns (PAMPs) (Pietretti & Wiegertjes, 2014). In addition, several TLRs are able to bind certain endogenous molecules called damage-associated molecular patterns (DAMPs) (following, e.g., trauma). TLRs are highly important since they represent a considerable diversity in their ligand-binding properties and thus facilitate responses against a wide array of pathogens. Genome duplication events in fishes during evolution have been attributed to the diversity of TLRs; therefore, differences with respect to the number of TLR loci exist between mammalian species and many fish species (Palti 2011). As an example: The genome of a mudskipper species (Periophthalmodon schlosseri) contains 11 copies of tlr13 (You et al. 2014). Most vertebrate genomes are recognized to have at least one gene representing each of the seven major tlr1, tlr2, tlr3, tlr4, tlr5, tlr7 and tlr11 families (Roach et al. 2005). Within Osteichthyes, the large tlr1 subfamily members include tlr1, tlr2, tlr14, tlr18, tlr25, tlr27, and tlr28 (Nie et al. 2018). The tlr3, tlr4, and tlr5 subfamilies recognize dsRNA, LPS, and bacterial flagellin. The tlr7 subfamily ligands are nucleic acid motifs, whereas the tlr11 family TLRs recognize an array of different molecules—from proteins to nucleic acids—reviewed by Nie et al. (2018). The ligand specificities for each TLR have not been very well studied in fish, though flagellin, synthetic triacetylated lipopeptide (Pam3CSK4), lipopeptides from gram-positive bacteria, and short double-stranded RNA (dsRNA) have been shown to interact with/bind to tlr1/2, tlr5, and tlr22, respectively (Nie et al. 2018). This means that fish immunologists assume that tlr localization and ligand specificities of fish tlrs are similar to mammalian counterparts. This is reviewed by Pietretti et al. (2014) and Kanwal et al. (2014). Tlrs are, in mice and humans, localized in the cell membrane and in the endoplasmic reticulum (ER), endosomes, and lysosomes (Fink et al. 2016). Tlr receptors 1, 2, 6, and 10 have, in human or mice models, been found to recognize a broad range of peptidoglycans and lipoproteins from, e.g., bacteria and parasites. These are located on the cell surface and, following engagement, there is intracellular signaling ending in NF-kB-dependent gene expression. NF-kB promotes expression of pro-inflammatory cytokines. Viral recognition may be brought about by tlr3, tlr7, tlr8, and tlr9 where they potentially can bind dsRNA, single-stranded RNA (ssRNA), and CpG DNA. As found in zebrafish, the tlr22 may also bind dsRNA or poly I:C (a dsRNA mimic) (Li et al. 2017b) [Fitzgerald, 2020 #954]. These “antiviral receptors,” upon ligand binding, confer (via TRIF or/and MyD88) activation of interferon regulatory factors 3 and 7 (transcription factors), which in turn facilitates transcription of interferon type I expression.
Taken together, the common interpretation is that TLR activation results in the production of pro-inflammatory cytokines (e.g., tnfa and IL-1ß) and/or in the expression and synthesis of transcription factors involved in protection against viruses, bacteria, and parasites (Sahoo 2020; Kanwal et al. 2014; Rauta et al. 2014; Zhang and Gui 2012; Palti 2011).
Tables 2.1, 2.2, 2.3, 2.4, 2.5, 2.6, 2.7, 2.8, 2.9, 2.10, 2.11, 2.12, 2.13, 2.14, 2.15, 2.16, 2.17, 2.18, 2.19, 2.20, 2.21, and 2.22 give an updated overview of tlrs 1–5, 7–9, 12–14, and 18–28 found in different fish species. This list is continuously growing as genome, and transcriptome sequences from new species are completed and analyzed. This will be done during the “Fish10K” project where the aim is to genome sequence 10,000 fish species during a ten-year period (Fan et al. 2020), and through the ongoing “Fish1K” project (Sun et al. 2016b) and Earth Biogenome Project (Lewin et al. 2018).
2.9.2 Interferon Type I
Interferons (IFNs) are a group of cytokines with important roles in defense against viral pathogens (cf. Chaps. 13 and 14). They are divided into two families, type I and type II, based on structural properties and functions. Both the type I and II IFN systems are essential to antiviral defense in innate and adaptive immunity (Zou and Secombes 2011) (Tables 2.23 and 2.24). In contrast to type I IFNs, which are more important in innate immunity, IFN-γ (type II IFN) is exclusively produced in immune-related cells and is more important later in the immune response. In innate immune responses, IFN-γ is produced by natural killer cells (Jung et al. 2012). During adaptive cell-mediated immune responses, IFN-γ is produced by CD4-positive Th1 cells and CD8-positive cytotoxic T lymphocytes. IFNs induce the expression of a broad array of IFN-stimulated genes (isgs), which encode for proteins with direct antiviral activity, including inhibition of viral transcription, degradation of viral RNA, inhibition of translation, or modification of protein function. Several reviews of the interferon system of teleost fish have been presented over the years (Robertsen 2006; Workenhe et al. 2010; Zou and Secombes 2011; Secombes and Zou 2017). Chaves-Pozo and coworkers investigated the interferon response in the ovary of rainbow trout (O. mykiss). They found that the VHS virus strongly upregulated all the ifn genes studied, while the IPN virus either had no effect or strongly suppressed ifn gene expression (Chaves-Pozo et al. 2010). Valero and coworkers investigated ifns in the gonads of gilthead sea bream (Sparus aurata) and European sea bass (D. labrax). They evaluated the expression after infection with the disease viral nervous necrosis (VNN) in the brain (Valero et al. 2015). The orange-spotted grouper, Epinephelus coioides, is a commercially important fish that is widely farmed in tropical waters, e.g., in Taiwan, Japan, Australia, and also Europe. Chen et al. characterized a type I ifn from this fish and determined the expression during nodavirus infection. Groupers infected with nodavirus had elevated levels of ifn and administration of recombinant IFN type I, which led to upregulated antiviral activity (Chen et al. 2014). In large yellow croaker (L. crocea), a type I group II interferon was identified by Ding and coworkers. The ifn was constitutively expressed in all examined tissues, spleen, liver, skin, head kidney, gills, blood, muscle, heart, brain, and intestine. The expression was rapidly upregulated in spleen and head kidney by poly I:C and Aeromonas hydrophila (Ding et al. 2019). A type I interferon gene was identified in Japanese eel (Anguilla japonica). The ifn was expressed constitutively in liver, spleen, intestine, gills, skin, kidney, heart, and muscle. After injection with LPS, poly I:C, and live A. hydrophila, expression levels increased in both liver, spleen, and kidney (Feng et al. 2017). A transgenic cell line for the detection of salmon interferons has been established. It is based on a CHSE-214 cell line containing a reporter construct expressing firefly luciferase under the control of a rainbow trout promoter for the IFN-induced mx1 gene. The mx promoter was shown to respond to both salmon IFN type I and trout IFN type II in a dose-dependent manner, while there was no response to recombinant tnfa and ilb (Jorgensen et al. 2007). Three distinct members of type I interferons were identified in the mandarin fish (Siniperca chuatsi) by Laghari et al. Fish injected intraperitoneally with poly I:C resulted in an enhanced expression of all three genes in the head kidney. The disease infectious spleen and kidney necrosis virus (ISKNV) caused an increased but delayed response of ifns (Laghari et al. 2018). Liu and coworkers studied ifn subgroups of salmonid species like rainbow trout (O. mykiss), chinook salmon (Oncorhynchus tshawytscha), coho salmon (Oncorhynchus kisutch), Atlantis salmon (Salmo salar), and Arctic charr (Salvelinus alpinus) and compared them with other species. The analysis confirmed that salmonids have a complex (in terms of ifn subgroups present) and (large number of genes) type I ifn repertoire relative to other teleost fish (Liu et al. 2020a). Milne et al. studied three distinct type I interferons in meagre (Argyrosomus regius), namely ifnc, ifnd, and ifnh. Constitutive expression was analyzed during larval development and in adult tissues (gills, midgut, head kidney, spleen). The spleen had high transcript levels of all three ifns. Ifnd and ifnh were also highly expressed in gills. The expression of each subgroup increased significantly across all four tissues following injection of poly I:C (Milne et al. 2018). In Atlantic salmon, Sun et al. identified an ifn multigene cluster encoding three ifn subtypes (ifna, ifnb, and ifnc). Each ifn subtype was constitutively expressed in head kidney. The three subtypes showed a striking difference in expression properties in response to stimulation with poly I:C. Both ifna and ifnc transcripts increased, while ifnb was only slightly induced by poly I:C (Sun et al. 2009). Type I interferon genes were cloned and characterized in the rock bream (Oplegnathus fasciatus). Their expression was upregulated in blood cells and head kidney by LPS, poly I:C, E. tarda, Streptococcus iniae, and iridovirus, and recombinant ifn I protein induced a rapid and transient expression of the mx gene in head kidney cells (Wan et al. 2012). An overview of type I ifns in different fish species with their activities is found in Table 2.23.
2.9.3 Interferon Type II
Interferon-gamma (ifng), the only type II interferon, is a pleiotropic pro-inflammatory and antiviral cytokine. In mammals, it is constitutively produced by NK cells, whereas T lymphocytes produce IFNG after activation or differentiation. IFNG is a key cytokine for innate and adaptive immunity against viral and intracellular bacterial infections and is involved in tumor control. An updated teleost interferon-gamma review has recently been published (Pereiro et al. 2019). Arts and coworkers made recombinant proteins of the carp (Cyprinus carpio) IFN-γ sequences of both clusters (ifng1 and ifng2) and tested their effects on expression of pro-inflammatory mediators (Arts et al. 2010). An interferon-responsive stable cell line RTG-3F7 has been developed for rainbow trout by modifying the RTG-2 cell line through transfection with a plasmid construct containing a promoter element from the IFN-γ responsive gene TAP2 linked to a luciferase reporter gene and a hygromycin resistance gene. The results indicate that the stable cell line RTG-3F7 is an excellent tool for monitoring the presence of trout ifng in biological samples (Castro et al. 2010). The large yellow croaker (L. crocea) is an important mariculture fish species in China, and the bacterium Vibrio harveyi and the ciliate protozoan C. irritans are the two major pathogens of this species. The nucleotide sequence of ifng was obtained, and expression studies were performed. Fish were challenged with V. harveyi and C. irritans, respectively. One day after injection with V. harvey, all 10 tissues investigated had a higher expression of ifng, while only spleen, muscle, intestine, heart, and skin had higher expression after infection with C. irritants (Chen et al. 2015). Jung and coworkers produced a recombinant ifng (rifng) from the olive flounder (P. olivaceus). Stimulation of kidney leukocytes in vitro with rinfg induced the gene expression of il1b, signal transducer and activator of transcription 1 (stat1), CXCL-13-like chemokine (cxcl13), and ifng. Intraperitoneal injection of a mixture of rifng and E. tarda into olive flounder resulted in a survival rate of 60% compared to 0% in the group treated with E. tarda only (Jung et al. 2012).
The extensive use of paraquat (PQ) in agricultural practice throughout the world may compromise the integrity of biological systems in fish. PQ toxicity has been found to be mediated by the production of free radicals, which cause oxidative damage to cells. In a study by Ma et al. (2014), the acute toxicity of PQ in common carp (C. carpio) was determined. The results suggest that PQ exposure may result in suppression or excessive activation of the immune system that leads to immune dysfunction and reduced immunity (Ma et al. 2014). In the report of Pereiro et al. (2016), an antiviral turbot (S. maximus) interferon-gamma gene was characterized, and its expression pattern under basal conditions, after type I ifn administration and viral and bacterial infections, was evaluated (Pereiro et al. 2016). The intramuscular injection of an expression plasmid encoding turbot ifn gene was not able to affect the transcription of numerous immune genes directly related to the activity of ifng. It was neither able to reduce the mortality caused by a VHSV nor A. salmonicida challenge. Shibasaki and coworkers cloned and characterized two ginbuna crucian carp (Carassius auratus langsdorfii)-specific isoforms of ifng called ifng rel1 and ifng rel2. Recombinant ifng rel1 and ifng rel2 showed high antiviral activities against the lethal crucian carp hematopoietic necrosis virus (Shibasaki et al. 2014).
The antiviral activity of ifn gamma against IPNV and salmonid alphavirus (SAV) was studied by Sun et al. (2011). The studies were performed in Atlantic salmon TO cells and Chinook salmon embryo cells (CHSE-214). Ifn-γ induced antiviral activity against both IPNV and SAV3 in salmon cells (Sun et al. 2011). The marine flatfish Atlantic halibut (Hippoglossus hippoglossus) is of great commercial interest. However, due to poorly developed larva at hatching and a long live-feed stage, aquacultural use of this species is limited. Øvergård and coworkers cloned and characterized the gene encoding the ifng. A constitutive expression was found in both lymphoid and non-lymphoid organs with relatively high expression in the thymus and gills (Overgard et al. 2012). An overview of type II ifng in different fish species is presented in Table 2.24.
2.9.4 Tnfa
Tumor necrosis factor-alpha (tnfa) is a cytokine involved in systemic inflammation, apoptosis, cell proliferation, and regulation of immune cells (Wiens and Glenney 2011). It is produced mainly by activated macrophages as a membrane or secreted form. The main pro-inflammatory effects are mediated through the activation of endothelial cells (Roca et al. 2008). In bony fish, tnfa was first discovered in Japanese Flounder (Hirono et al. 2000) rainbow trout (Laing et al. 2001) and has since been characterized in a number of species. Fish have 14 tumor necrosis family genes. Their genomic existence and location have been investigated in the Japanese pufferfish (fugu) (Takifugu rubripes) (Biswas et al. 2015). Fugu was found to possess nine tnf superfamily genes including seven newly identified and two that had been previously reported. Poly I:C caused an elevated expression of three fugu tnf superfamily 10 genes in head kidney cells. Tnfa is an important factor for bacterial pathogen killing. A. salmonicida subsp. salmonicida is highly pathogenic for turbot, an economically important cultured flatfish in Europe, China, and Chile. In A. salmonicida-infected fish, the number of tnfa immunopositive cells was significantly increased in the kidney and spleen (Coscelli et al. 2016). Immunoreactive cells were also present in the digestive tract, liver, heart, gills, and skin (Ronza et al. 2015).
The striped trumpeter (Latris lineata Forster) is a new species in Tasmanian waters. The tnfa was cloned, and the expression was analyzed in response to an ectoparasite Chondracanthus goldsmidi. A significant upregulation was found in the gills, which are the site of parasite attachment. Head kidney cells showed a significant upregulation of tnfa, but spleen cells did not (Covello et al. 2009). The European sea bass (D. labrax) is intensely aquacultured in the Mediterranean area. The bacterial pathogen V. anguillarum provokes the highest mortality among several pathogens of this species. Available vaccines do not achieve the desired protection. In a recent study, recombinant tnfa was used as adjuvant in a commercial sea bass oral vaccine against V. anguillarum. Tnfa significantly enhanced disease resistance and induced recruitment of gut intraepithelial lymphocytes (Galindo-Villegas et al. 2013). In rainbow trout, two tnfa genes have been described. Recently, a third tnfa (tnfa3) that has low identities to known trout molecules was reported. The constitutive expression of tnfa3 was generally lower than the other two genes in tissues and cell lines. Expression of all three tnfa isoforms could be modulated by crude LPS, peptidoglycan, poly I:C and recombinant Ifng in cell lines and primary macrophages, and bacterial and viral infections (Hong et al. 2013). The genomic location of the two tnfa genes in zebrafish (D. rerio) and medaka (Oryzias latipes) was recently determined. Zebrafish tnfa1 and tnfa2 were found on chromosomes 19 and 15, and medaka tnfa1 and tnfa2 on chromosomes 11 and 16, respectively. There was a constitutive expression of the genes in different tissues. An increased expression of both was induced in head kidney cells by LPS in vitro (Kinoshita et al. 2014). Li and Zhang studied a tnfa homologue from the Tongue sole (Cynoglossus semilaevis) named CsTNF1. Expression of CsTNF1 was detected in liver, spleen, kidney, blood, gill, brain, muscle, heart, and intestine, and was upregulated by experimental challenge with bacterial and viral pathogens (Li and Zhang 2016).
Meagre (Argyrosomus regius) is an emerging aquaculture species found in the Mediterranean area and Black Sea due to its large size, fast growth, low feed conversion ratio, and high processing yield. Two types of tnfa were expressed in meagre (type 1 and type 2). Tnfa1 was more highly expressed in head kidney and gills. Both isoforms increased in expression in head kidney following injection with LPS (Milne et al. 2017). Atlantic bluefin tuna (Thunnus thynnus) was introduced into Mediterranean aquaculture in the early nineties, and has become the most valuable finfish aquaculture, representing more than half of the world’s total production (Pleic et al. 2014). Tuna aquaculture is a capture-based activity, where wild-caught tuna is cultured in marine cages for a period of time in order to increase their protein and fat content. The full-length cDNA and gene sequences of Bluefin tuna tnfa1 and tnfa2 were determined, and expression studies showed that they were constitutively expressed in liver and head kidney at similar levels. Expression of both cytokines was examined in acute and chronic natural infection of the parasites Pseudocycnus appendiculatus and Didymosulcus katsuwonicola (Pleic et al. 2015). D. katsuwonicola-infected gills showed significantly higher expression of tnfa2, while tnfa1 showed no difference in expression with either Pseudocycnus appendiculatus- or Didymosulcus katsuwonicola-infected gills.
Rainbow trout red blood cells (RBCs) are able to endocytose nanostructured tnfa in vitro despite not being phagocytic cells, and in response to nanostructured tnfa, the expression of different immune genes could be modulated (Puente-Marin et al. 2019a).
Tnfa was cloned in large yellow croaker (Pseudosciaena crocea), mainly distributed in coastal regions of East Asia, and is one of the most important cultured marine fish in China. Vibrio parahemolyticus challenge demonstrated enhanced expression of tnfa in head kidney and blood (Xie et al. 2008). Tnfa was also identified in grass carp (Ctenopharyngodon idella), and its role in signaling was defined (Zhang et al. 2012). Additionally, tnfa is involved in the control of ovulation (Crespo et al. 2010, 2015) (Table 2.25). For more details, please see Chap. 10 (“cytokines”) by Dr. C. Secombes.
2.9.5 The Complement System
The mammalian complement system is composed of about 35 plasma and membrane-associated proteins. The main functions of the complement system are opsonization, inflammation, and formation of the cytolytic membrane attack complex. The proteins are mostly produced by liver hepatocytes and secreted to the blood, except for some like factor D and c1q. Several components of the teleost complement contain isoforms like c3, c4, c5, c7, factor B, factor I, and MBL. Most homologs of mammalian complement components are present in teleosts and have been shown to be expressed in a variety of tissues like the kidney, skin, and intestine. Several reviews on the complement system of teleosts are available (Zhang et al. 2013b; Nakao et al. 2011; Uribe et al. 2011; Boshra et al. 2004, 2006). Additional information of the complement system in fish is given in Chap. 9.
2.9.5.1 c3
c3 is the central complement component and has been isolated, purified, and characterized in many teleost species. Recently, four c3 isoforms were purified from the Nile tilapia (Oreochromis niloticus) serum and were shown to possess an intrachain thioester bond. All named c3-1, c3-2, c3-3, and c3-4 showed the two-chain polypeptide structure typical of c3 (Abdel-Salam et al. 2014). Forn-Cuni et al. confirmed the presence of three c3 genes and in addition identified five more c3 genes in the zebrafish (D. rerio) genome (Forn-Cuni et al. 2014). Maternal immunization of female zebrafish with formalin-killed A. hydrophila caused a significant increase in c3 and factor b contents in the mother, a corresponding rise in the offspring, and induced a remarkable increase in the hemolytic activities in both the mother and offspring (Wang et al. 2009). The dojo loach (Misgurnus anguillicaudatus) is one of the most commercially cultured fish species in Eastern Asian countries including China, Japan, and Korea. Three isoforms of c3 were discovered in M. anguillicaudatus named c3-1, c3-2, and c3-3, respectively. The expression of c3–1 and c3–3 was mainly detected in liver followed by spleen and gonad. The mRNA levels were upregulated in the gill, skin, liver, and spleen after bath infection with A. hydrophila (Xu et al. 2018b). Furthermore, the complete nucleotide sequence of c3 from two Antarctic teleosts Trematomus bernacchii (two isoforms) and Chionodraco hamatus (a single isoform) was determined (Melillo et al. 2015).
Rainbow trout c3a and c5a receptors were cloned and functionally characterized. Both anaphylatoxin receptors were expressed at considerable levels by B cells. Treatment by lipopolysaccharide led to a significant upregulation of both receptors, suggesting that B cells play a role in the development of an inflammatory response (Li et al. 2007).
2.9.5.2 Classical Pathway
The classical complement pathway involves c1, c2, and c4. The first complement component (c1) is activated by recognition of antigen-bound immunoglobulins, and proteolytically activates c4 and c2 into c4b and c2a, respectively. Rock bream (Oplegnathus fasciatus) is one of the most economically important marine fish species in South Korea, which geographically distribute in the coastal water, especially in coral beds of the Pacific and Indian Ocean. Rock bream complement components c1r and c1s were characterized, and homology analysis showed 73.4% and 58% amino acid identity with orthologs of Pundamilia nyererei of Lake Victoria and the Japanese rice fish, Oryzias latipes, respectively. c1r was highest expressed in blood and c1s in the liver. The transcription of both components was found to be upregulated in response to pathogenic bacteria E. tarda and S. iniae and virus (rock bream iridovirus) (Godahewa et al. 2015).
Grass carp (C. idella) is susceptible to A. hydrophila infections. In the study of Dang et al., grass carps were given intraperitoneal injections of live A. hydrophila and 4, 8, 12, 24, 48, and 72 h after RNA sequencing of spleen tissue was performed. Four to 72 h after infection, the complement system, represented by c2, c3, c4, c5, c8a, c1q, and mbl, was upregulated with a transitory downregulation at 12 h (Dang et al. 2016).
Tissue of C. idella was infected with A. hydrophila. Two cDNA sequences of c4 from the common carp (C. carpio) were isolated sharing only 32% identity of amino acid level and having distinct binding specificities (Mutsuro et al. 2005).
2.9.5.3 Alternative Pathway
Factor B and factor D are both components of the alternative pathway of complement. Following activation of the alternative pathway, factor B is cleaved into Ba and Bb fragments. In rainbow trout, factor B is known to act as a c3 convertase, but the function of the Ba fragment is unknown. The expression patterns of tongue sole (Cynoglossus semilaevis) factor B and the biological activity of the Ba fragment were studied by Li and Sun (2017). Expression of factor B was high in liver, muscle, and heart and low in intestine, blood, and kidney. Bacterial infection (E. tarda, Pseudomonas fluorescens, and V. harveyi) induced an expression in kidney, spleen, and liver in a time-dependent manner. For the first time, it was found that overexpression of Ba significantly reduced bacterial dissemination in fish tissues, indicating that Ba possesses antimicrobial activity and may inhibit bacterial infection in fish (Li and Sun 2017).
Rock bream (Oplegnathus fasciatus) complement factor D (Cfd) was characterized and expression was analyzed. Factor D encodes 277 amino acids for a 30 kDa polypeptide and was most highly expressed in the liver and spleen. Transcription of factor D was upregulated in the spleen by lipopolysaccharide, S. iniae, rock bream iridovirus, and poly I:C (Godahewa et al. 2016). Rainbow trout liver seems not to be an important transcription site of the genes c1q, factor B (cfb), and c7–2. The novel characterized factor D of rainbow trout had 253 amino acids with a molecular weight of 27.2 kDa and shared a sequence identity with its human ortholog of 45% (Kobis et al. 2015).
2.9.5.4 Lectin Pathway
The central components of the lectin pathway are MBL and MASPs. Teleost fish often possess several genes encoding different subtypes. Kania et al. (2010) characterized three homologs of mannan-binding lectin (named MBL H-1, MBL H-2, MBL H-3) (Gene: mbl and variants) in the rainbow trout. They were expressed in the spleen, anterior intestine, and liver. MBL H-1 and H-3 were also found in the vascular system. MBL H-1 had the highest expression level in the anterior intestine followed by gill, thymus, and skin, while the highest expression level of MBL H-2 and MBL H-3 occurred in the anterior intestine (Kania et al. 2010).
C. semilaevis mannan-binding lectin (Mbl)-associated protein 34 (MAP34) and Mbl-associated serine protease 1 (MASP1) are key factors involved in complement activation through MAPs’ ability to bind to M1 and MBL. Remarkably, in contrast to a negative regulatory role of mammalian MAP, the teleost C. semilaevis Map 34 exerts a positive effect on the activation of the lectin pathway (Li et al. 2016).
2.9.5.5 Terminal Pathway
c5, c6, c7, c8, and c9 are the components engaged in building the membrane attack complex. Native c5a of rainbow trout and recombinant infectious hematopoietic necrosis virus glycoprotein (G) fusion protein was constructed to test the adjuvant activity of rainbow trout c5a. At four to sixteen weeks postinjection, the serum Igm antibody levels were higher than those injected with G-protein alone, suggesting that c5a acts as molecular adjuvant in rainbow trout (Wu et al. 2014).
Grass carp (C. idella) is an economically important species, and its global production is more than 4.5 million tons per year making it the most highly consumed freshwater fish species in the world. A. hydrophila is the causative pathogen of intestinal hemorrhage, which has caused great economic loss in grass carp aquaculture. Fish were intraperitoneally injected with A. hydrophila, and the transcriptomic response was tested in the spleen. A total of 1591 genes were upregulated, and 530 were downregulated. c1, c7, and c8b were upregulated indicating activation of the classical pathway (Yang et al. 2016). c7 was isolated and characterized from grass carp. The predicted amino acid sequence of c7 cDNA exhibited 55.4% and 48.3% homology with rainbow trout c7–1 and zebrafish c7, respectively. c7 gene expression was detected in trunk kidney, liver, head kidney, skin, spleen, heart, and intestine. Significant changes in c7 transcript expression were detected following A. hydrophila infection, especially in head kidney and spleen (Shen et al. 2012).
Full-length c8a and c8b sequences from a cDNA library of rock bream (Oplegnathus fasciatus) and their genomic sequences were obtained. Quantitative real-time PCR analysis showed that both components were expressed in all examined tissues, with highest expression in the liver. Pathogen challenge, including E. tarda, S. iniae, and rock bream iridovirus, led to upregulation of both (Wickramaarachchi et al. 2013).
Complement component c9 is the last component that is involved in the formation of the membrane attack complex on the surface of target cells. The full-length c9 cDNA sequence was found in the southern catfish (Silurus meridionalis) and showed similarity with other teleost fish. The mRNA expression was highest in the liver and observed also in the spleen, head kidney, stomach, and intestine. Intraperitoneal injection of A. hydrophila gave upregulation of c9 in the liver, spleen, and intestine (Fu et al. 2019).
The large yellow croaker Larimichthys crocea is one of the most important marine fish in China and East Asian countries. Complement components c7 and c9 were characterized by Guo et al. (2016). c7 and c9 were mainly expressed in liver, but low levels were also constitutively expressed in most tissues. Fish challenged with Vibrio alginolyticus showed a rapidly upregulated response in the liver and head kidney (Guo et al. 2016). Miiuy croaker, Miichthys miiuy, belongs to the family Sciaenidae of the order Perciformes and mainly distributes from the western Japan Sea to the East China Sea. In China, it has been widely cultured since the late 1990s for its good taste and high nutritive and medicinal value. A truncated c9 cDNA sequence encoding 461 amino acids was cloned and characterized in the miiuy croaker (M. miiuy). The c9 of miiuy croaker shows the highest amino acid identity score with fugu c9 (61%) and the lowest with zebrafish c9 (36%). The highest levels of transcripts were detected in liver of both healthy and V. anguillarum-infected fish (Meng et al. 2012). Full-length c9 sequence was identified from a cDNA library of rock bream (O. fasciatus), and its genomic sequence was obtained. Quantitative real-time RT-PCR analysis confirmed that c9 was constitutively expressed in all the examined tissues, with highest expression occurring in the liver. Pathogen challenge including E. tarda, S. iniae, lipopolysaccharide endotoxin, and rock bream iridovirus led to upregulation of c9 in liver but resulted in no change in the peripheral blood cells (Wickramaarachchi et al. 2012). The transcriptional expression of central complement components during the ontogeny of the common sole (Solea solea) was studied by Ferraresso et al. (2016). The c2, c3, and fb showed a gradual increase in expression between 4 and 33 days post-hatch (dph). c4 and masp1 showed no differences in expression during the development, while c1qb showed a very high level of expression. Terminal components, c5, c6, c7, c8, and c9, showed an increase in expression until the onset of metamorphosis and a second increase after metamorphosis (Ferraresso et al. 2016).
2.9.5.6 Complement Regulation
Complement activation is controlled by both fluid phase and membrane inhibitors. Factor I regulates complement by proteolytic cleavage of components c3b and c4b. Factor H, the main cofactor of factor I, regulates the alternative pathway by acting in the breakdown of c3b to ic3b. Factor I (cfi) and factor H (cfh) of rainbow trout were cloned and characterized. The deduced amino acid sequences of factor I and factor H exhibited 42% and 32% identity with human orthologs, respectively (Anastasiou et al. 2011). The deduced amino acid sequence of factor H from large yellow croaker (Larimichthys crocea) showed 28% and 34% identity with human and rainbow trout orthologs, respectively. The highest expression levels were found in liver, kidney, and spleen. After injection with V. alginolyticus, the expression levels were upregulated in all three tested tissues (Qi et al. 2018a). Black rockfish (Sebastes schlegelii) is an important aquaculture species in the Republic of Korea. A c1 inhibitor gene from black rockfish was cloned and characterized by Nilojan et al. The c1 inhibitor was most highly expressed in the liver followed by the gills (Nilojan et al. 2018) (Table 2.26).
2.9.6 Acute-phase Component
During infection, stimulation with strong danger signals or stress, the fish may respond to produce acute-phase proteins (APPs). Especially, IL-1, IL-6, and tnfa are able to induce acute-phase response, as observed in higher vertebrates. The most common APPs are pentraxins such as serum amyloid A (SAA) and C-reactive protein (CRP). Dissimilar to many mammalian species, the fish show a modest acute-phase response when it comes to concentration of pentraxins in serum. CRP may be able to bind to (opsonize) certain bacteria, fungi, and parasites, activate the complement system, agglutinating particles, and may infer production of cytokines. There are two forms of SAA, one of them being acute-phase SAA. SAA may neutralize pathogen activity, reduce tissue damage, and restore homeostasis. Transferrin, haptoglobin, ceruloplasmin, alpha-2-macroglobulin, lectins, and complement component c3 are all considered to be AAPs. Most of these have regulatory activities limiting infection and restoring the physiological balance. Several reviews covering this topic are recommended (Roy et al. 2017; Bayne and Gerwick 2001; de Magalhaes et al. 2020; Magnadottir 2014; Nakao et al. 2011).
2.9.7 Chemokines and Their Receptors
The major function of chemokines is to guide the migration of cells. An example is chemokine-guided migration of leukocytes to inflammatory foci. Other functions involve immune surveillance where chemokines direct homing of leukocytes to lymphatic tissues. Some chemokines have function in growth of new blood vessels and wound healing. Chemokines are classified into four main subfamilies (CXC, CC, CX3C, and XC) dependent on the amino acid sequences (first two cysteine amino acid residues). Most of the chemokines bind to specific chemokine receptors on cells. Fish display numerous genes for different chemokines and chemokine receptors, suggested due to gene duplication events. Functional and significance studies of chemokine expression are generally not very well examined in fish. However, exceptions exist. We list the most recent findings within functional chemokine research. It has been shown that grass carp cxcl20b possessed antibacterial activity by attaching to the bacterial membrane (Xiao et al. 2020). In another study, it was shown that common carp Cxcb1 stimulated neutrophil extracellular trap formation—which was suggested to be an antipathogenic event (Pijanowski et al. 2020). Moreover, an ayu (Plecoglossus altivelis) CC-like chemokine was found to possess chemotactic activity against monocytes and neutrophils in vivo and in vitro (Yu et al. 2019). Chemotactic activity of Ccl4 has been shown in the golden pompano (Trachinotus blochii). This recombinant chemokine had also antimicrobial activity against E. tarda and Escherichia coli (Sun et al. 2019). Furthermore, a rainbow trout CC chemokine (Ck11) also displayed antimicrobial activity against different gram-positive and gram-negative bacteria by attaching to and disrupting their cell membranes (Munoz-Atienza et al. 2019).
2.9.8 Antibacterial Peptides (AMPs)
AMPs are a diverse class of highly conserved molecules that are produced as a first line of defense in all multicellular organisms, including fish. These small peptides (12–50 amino acids) are essential components of innate immunity capable of antimicrobial activity against a broad range of microbial pathogens (Semple and Dixon 2020; Zhang and Gallo 2016). Functionally, they can be described as either membrane disruptive AMPs, which induce membrane permeabilization, or they can be non-membrane disruptive where they can be internalized in cells and act on intracellular targets (Semple and Dixon 2020). In general, fish AMPs may be categorized into five different classes based on their structure: β-defensins, cathelicidins, hepcidins, histone-derived peptides, and piscidins (Brunner et al. 2020). As for chemokines and TLRs, fish possess numerous genes for antimicrobial peptides. Recently, reviews on the significance of AMPs are published (Brunner et al. 2020; Chaturvedi et al. 2020; Valero et al. 2020; Chen et al. 2020; Shabir et al. 2018).
2.10 Conclusion and Future Research
It is clear that fish are indeed equipped with an arsenal of defense mechanisms to prevent infection. An earlier report has used Rag knockout mutants (rag1−/− zebrafish), which possess no serum Igm, to assess the significance of the innate immune system in comparison with control fish with a rag1+/− genotype. An experimental challenge experiment revealed that the rag −/− zebrafish displayed similar protection as the controls (Tokunaga et al. 2017). This underscores the notion that the innate immune system alone may likely be as effective as a fully immune-equipped fish. However, immunized fish will normally acquire higher disease resistance than naïve fish. The concept of trained innate memory should be addressed as a trained innate immune system likely would add higher protection level during infection. Trained innate immunity involves activation of innate defense factors that in turn confer increased disease resistance to infection by homologous or heterologous pathogens. Trained immunity can be transferred to offspring as training induces heritable epigenetic changes.
The future will bring a vast more knowledge of innate immune factors through fish genomic and transcriptomic studies, and it is likely that many more innate immune factors will be revealed. To find their significance in the innate immune defense, these must be functionally examined.
Abbreviations
- AMP:
-
Antimicrobial peptide
- APC:
-
Antigen-presenting cell
- APP:
-
Acute-phase proteins. Resulting from acute inflammation
- c1, c2, c3, c4, c5, c6, c7, c8, and c9:
-
Complement components
- c1q:
-
A protein complex involved in the complement system
- Cathl1:
-
Cathelicidin gene 1. Antimicrobial peptide
- Cathl2:
-
Cathelicidin gene 2
- Ccl19:
-
Chemokine (C–C motif) ligand 19. A regulator of the induction of T-cell activation, immune tolerance, and inflammatory responses during continuous immune surveillance, homeostasis, and development
- Cd4:
-
Cluster of differentiation 4
- Chemokine:
-
A protein that can attract cells, toward a chemical gradient, having the specific receptor, and promote differentiation and multiplication of leukocytes, and cause tissue extravasation
- CircRNA:
-
Is a type of single-stranded RNA, which, unlike linear RNA, forms a covalently closed continuous loop. Can be protein coding and noncoding
- CpG DNA:
-
DNA that contains methylated nucleotides (CpG islands). Normally found in promoter regions, which modulate gene expression
- CRP:
-
C-reactive protein. An acute-phase protein
- Cxcl13:
-
Chemokine (C-X-C motif) ligand 13. Involved in chemotaxis of B lymphocytes
- Cxcl9:
-
Chemokine (C-X-C motif) ligand 9 is a small cytokine belonging to the CXC chemokine family. Plays role in chemotaxis
- DAMPs:
-
Damage-associated molecular patterns. Molecules released in a sterile inflammation or damage
- DC:
-
Dendritic cell
- DEGs:
-
Differentially expressed genes. From RNAseq data and bioinformatics
- DEPs:
-
Differential expressed proteins
- dsRNA:
-
Double-stranded RNA
- Epigenetics:
-
Is the study of heritable phenotype changes that do not involve alterations in the DNA sequence
- Factor B (c3 convertase):
-
Protein that is activated by cleavage, yielding Bb and Ba fragments. Factor B is cleaved only when it is bound to c3b
- Factor D:
-
A serine protease present in blood and tissue in an active sequence but self-inhibited conformation. The only known natural substrate of factor D is factor B. Alternative pathway of complement activation
- Factor H:
-
The main cofactor of factor I
- Factor I:
-
Protein of the complement system (c3b/c4b inactivator). Alternative pathway of complement activation
- Foxp3:
-
Forkhead box P3. Regulator of the regulatory pathway in the development and function of regulatory T cells
- Galectins:
-
Proteins that bind specifically to β-galactoside sugars
- Hepcidin:
-
An antimicrobial peptide
- Histone modifications and DNA methylation:
-
Both DNA methylation and histone modification are involved in establishing patterns of gene repression during development
- HSP1:
-
Heat shock protein I gene. Mitochondrial product
- HSP90b:
-
A chaperone protein that assists other proteins to fold properly, stabilizes proteins against heat stress, and aids in protein degradation
- Ifn:
-
Interferon (cytokine)
- Ifnrel:
-
Interferon-related
- Ifng:
-
Interferon-gamma
- IL:
-
Interleukin (cytokine)
- IRF:
-
Interferon regulatory factor
- Isoforms (subtypes):
-
Alternatively spliced genes
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- lncRNAs:
-
A large and diverse class of transcribed RNA molecules with a length of more than 200 nucleotides that do not encode proteins
- LPS:
-
Bacterial lipopolysaccharide. A main constituent in gram-negative bacterial cell wall
- MAP kinase 1:
-
Mitogen-activated protein kinase I. Transcription factor
- MAP:
-
Mannan-binding lectin (MBL)-associated protein
- MASPs:
-
Serine proteases that function as a component of the lectin pathway of complement activation
- MBL:
-
Mannose-binding lectin (lectin pathway of complement activation)
- MHC I and II:
-
Major histocompatibility complex I and complex II. Function in antigen presentation
- MicroRNAs:
-
(miRNAs) are a family of small noncoding RNAs
- MPO:
-
Myeloperoxidase. An enzyme that catalyzes the formation of a number of reactive oxidant species
- MyD88:
-
Myeloid differentiation factor 88 (MyD88). A central component of the Toll-like receptor pathway
- Mx:
-
An interferon-induced GTP-binding protein
- NOD-like receptors (NLRs) and C-type lectin receptors (CLRs):
-
Belongs to RIG-I and PPR family
- PPRs:
-
Pattern recognition receptors. Expressed on many types of cells, especially on antigen-presenting cells. Recognize repeating molecular patterns often found in pathogens
- RNAseq:
-
RNA sequencing
- TLRs:
-
Toll-like receptors (belongs to PPR family)
- PAMPs:
-
Pathogen-associated molecular patterns
- Paraquat (PQ):
-
Toxic chemical that is widely used as an herbicide
- Pentraxins:
-
Family of acute-phase proteins produced during acute-phase response
- Poly I:C:
-
Polyinosinic:polycytidylic acid. Binds TLR3
- QPCR:
-
A real-time polymerase chain reaction, also known as quantitative polymerase chain reaction
- RAG:
-
Recombination-activating gene
- RIG-I-like receptors (RLRs):
-
Retinoic acid-inducible gene I-like receptors. Belongs to RIG-I and PPR family
- ROS:
-
Reactive oxygen species
- SAA:
-
Serum amyloid A. A protein formed during acute-phase response
- SETD3-OT:
-
Has function on histidine methylation which belongs to epigenetic occurrence
- ssRNA:
-
Single-stranded RNA
- STAT1:
-
Signal transducer and activator of transcription 1. STAT1 can be activated by several ligands such as IFN-α, IFN-γ, epidermal growth factor (EGF), platelet-derived growth factor (PDGF), IL-6, or IL-27
- TGF-ß:
-
Transforming growth factor-beta
- Th1 cells:
-
Pro-inflammatory T cells that are responsible for cell-mediated immunity and phagocyte-dependent protective responses
- Tnf:
-
Tumor necrosis factor
- Tnfr:
-
Tumor necrosis factor receptor
- TRIF:
-
TIR domain-containing adapter-inducing interferon-β. An adapter in responding to activation of Toll-like receptors (TLRs)
- Type I interferon:
-
A role in antiviral responses
- Type II interferon:
-
A role in adaptive and innate immunity
- Viperin:
-
Virus-inhibitory protein, endoplasmic reticulum-associated, interferon-inducible. An IFN-inducible gene
References
Aballai V, Aedo JE, Maldonado J, Bastias-Molina M, Silva H, Meneses C, Boltana S, Reyes A, Molina A, Valdes JA (2017) RNA-seq analysis of the head-kidney transcriptome response to handling-stress in the red tusk-eel (Genypterus chilensis). Comp Biochem Phys D 24:111–117. https://doi.org/10.1016/j.cbd.2017.09.002
Abdel-Salam SGR, Tsujikura M, Kondo M, Somamoto T, Nakao M (2014) Purification and functional characterization of complement C3 and a novel zymosan-binding protein in tilapia serum. Fish Sci 80(2):301–310. https://doi.org/10.1007/s12562-014-0700-7
Abouelmaatti RR, Algammal AM, Elfeil WMK, Elshaffy NM, Li XK, Ma JS, Wahdan A, El-Tarabili R, Shabana II (2020) Genetic characterization, cloning, and expression of Toll-like Receptor 1 mRNA Nile tilapia (Oreochromis niloticus). Vet Arhiv 90(2):185–196. https://doi.org/10.24099/vet.arhiv.0563
Ali A, Rexroad CE, Thorgaard GH, Yao JB, Salem M (2014) Characterization of the rainbow trout spleen transcriptome and identification of immune-related genes. Front Genet 5:348. https://doi.org/10.3389/fgene.2014.00348
Altmann S, Korytar T, Kaczmarzyk D, Nipkow M, Kuhn C, Goldammer T, Rebl A (2016) Toll-like receptors in maraena whitefish: evolutionary relationship among salmonid fishes and patterns of response to Aeromonas salmonicida. Fish Shellfish Immun 54:391–401. https://doi.org/10.1016/j.fsi.2016.04.125
Anastasiou V, Mikrou A, Papanastasiou AD, Zarkadis IK (2011) The molecular identification of factor H and factor I molecules in rainbow trout provides insights into complement C3 regulation. Fish Shellfish Immun 31(3):491–499. https://doi.org/10.1016/j.fsi.2011.06.008
Ao JQ, Mu YN, Wang KR, Sun M, Wang XH, Chen XH (2016) Identification and characterization of a novel Toll-like receptor 2 homologue in the large yellow croaker Larimichthys crocea. Fish Shellfish Immun 48:221–227. https://doi.org/10.1016/j.fsi.2015.11.002
Arts JAJ, Tijhaar EJ, Chadzinska M, Savelkoul HFJ, Verburg-van Kemenade BML (2010) Functional analysis of carp interferon-gamma: evolutionary conservation of classical phagocyte activation. Fish Shellfish Immun 29(5):793–802. https://doi.org/10.1016/j.fsi.2010.07.010
Asbakk K, Dalmo RA (1998) Atlantic salmon (Salmo salar L.) epidermal Malpighian cells - motile cells clearing away latex beads in vitro. J Mar Biotechnol 6(1):30–34
Austbo L, Aas IB, Konig M, Weli SC, Syed M, Falk K, Koppang EO (2014) Transcriptional response of immune genes in gills and the interbranchial lymphoid tissue of Atlantic salmon challenged with infectious salmon anaemia virus. Dev Comp Immunol 45(1):107–114
Bai JS, Li YW, Deng Y, Huang YQ, He SH, Dai J, Zhao SZ, Dan XM, Luo XC (2017) Molecular identification and expression analysis of TLR5M and TLR5S from orange-spotted grouper (Epinephelus coioides). Fish Shellfish Immun 63:97–102. https://doi.org/10.1016/j.fsi.2017.01.037
Bai HQ, Zhou T, Zhao J, Chen BH, Pu F, Bai YL, Wu YD, Chen L, Shi Y, Ke QZ, Yu XK, Xu P (2020) Transcriptome analysis reveals the temporal gene expression patterns in skin of large yellow croaker (Larimichthys crocea) in response to Cryptocaryon irritans infection. Fish Shellfish Immun 99:462–472. https://doi.org/10.1016/j.fsi.2020.02.024
Balla KM, Lugo-Villarino G, Spitsbergen JM, Stachura DL, Hu Y, Banuelos K, Romo-Fewell O, Aroian RV, Traver D (2010) Eosinophils in the zebrafish: prospective isolation, characterization, and eosinophilia induction by helminth determinants. Blood 116(19):3944–3954. https://doi.org/10.1182/blood-2010-03-267419
Baoprasertkul P, Peatman E, Somridhivej B, Liu ZJ (2006) Toll-like receptor 3 and TICAM genes in catfish: species-specific expression profiles following infection with Edwardsiella ictaluri. Immunogenetics 58(10):817–830. https://doi.org/10.1007/s00251-006-0144-z
Baoprasertkul P, Peatman E, Abernathy J, Liu ZJ (2007a) Structural characterisation and expression analysis of toll-like receptor 2 gene from catfish. Fish Shellfish Immun 22(4):418–426. https://doi.org/10.1016/j.fsi.2006.04.005
Baoprasertkul P, Xu P, Peatman E, Kucuktas H, Liu Z (2007b) Divergent toll-like receptors in catfish (Ictalurus punctatus): TLR5S, TLR20, TLR21. Fish Shellfish Immun 23(6):1218–1230. https://doi.org/10.1016/j.fsi.2007.06.002
Banchereau J, Briere F, Caux C, Davoust J, Lebecque S, Liu YJ, Pulendran B, Palucka K (2000) Immunobiology of dendritic cells. Annu Rev Immunol 18(1):767–811. https://doi.org/10.1146/annurev.immunol.18.1.767
Bassity E, Clark TG (2012) Functional identification of dendritic cells in the teleost model, rainbow trout (Oncorhynchus mykiss). PLoS One 7(3):e33196
Basu M, Swain B, Maiti NK, Routray P, Samanta M (2012a) Inductive expression of toll-like receptor 5 (TLR5) and associated downstream signaling molecules following ligand exposure and bacterial infection in the Indian major carp, mrigal (Cirrhinus mrigala). Fish Shellfish Immun 32(1):121–131. https://doi.org/10.1016/j.fsi.2011.10.031
Basu M, Swain B, Sahoo BR, Maiti NK, Samanta M (2012b) Induction of toll-like receptor (TLR) 2, and MyD88-dependent TLR- signaling in response to ligand stimulation and bacterial infections in the Indian major carp, mrigal (Cirrhinus mrigala). Mol Biol Rep 39(5):6015–6028. https://doi.org/10.1007/s11033-011-1415-9
Bayne CJ, Gerwick L (2001) The acute phase response and innate immunity of fish. Dev Comp Immunol 25(8–9):725–743. https://doi.org/10.1016/S0145-305x(01)00033-7
Belwal K, Thakuria D, Dighe V, Pande V, Pande A (2017) Molecular cloning and expression profile of toll-like receptor 3 from an Indian coldwater fish, Schizothorax richardsonii(Gray). Anim Biotechnol 28(2):144–147. https://doi.org/10.1080/10495398.2016.1217874
Bilodeau AL, Waldbieser GC (2005) Activation of TLR3 and TLR5 in channel catfish exposed to virulent Edwardsiella ictaluri. Dev Comp Immunol 29(8):713–721. https://doi.org/10.1016/j.dci.2004.12.002
Biswas G, Kinoshita S, Kono T, Hikima J, Sakai M (2015) Evolutionary evidence of tumor necrosis factor super family members in the Japanese pufferfish (Takifugu rubripes): comprehensive genomic identification and expression analysis. Mar Genom 22:25–36. https://doi.org/10.1016/j.margen.2015.03.003
Boltana S, Valenzuela-Miranda D, Aguilar A, Mackenzie S, Gallardo-Escarate C (2016) Long noncoding RNAs (lncRNAs) dynamics evidence immunomodulation during ISAV-Infected Atlantic salmon (Salmo salar). Sci Rep-Uk 6:22698. https://doi.org/10.1038/srep22698
Boltana S, Aguilar A, Sanhueza N, Donoso A, Mercado L, Imarai M, Mackenzie S (2018) Behavioral fever drives epigenetic modulation of the immune response in fish. Front Immunol 9:1241. https://doi.org/10.3389/fimmu.2018.01241
Boshra H, Gelman AE, Sunyer JO (2004) Structural and functional characterization of complement C4 and C1s-like molecules in teleost fish: Insights into the evolution of classical and alternative pathways. J Immunol 173(1):349–359. https://doi.org/10.4049/jimmunol.173.1.349
Boshra H, Li J, Sunyer JO (2006) Recent advances on the complement system of teleost fish. Fish Shellfish Immun 20(2):239–262. https://doi.org/10.1016/j.fsi.2005.04.004
Brietzke A, Arnemo M, Gjoen T, Rebl H, Korytar T, Goldammer T, Rebl A, Seyfert HM (2016) Structurally diverse genes encode Tlr2 in rainbow trout: The conserved receptor cannot be stimulated by classical ligands to activate NF-kappa B in vitro. Dev Comp Immunol 54(1):75–88. https://doi.org/10.1016/j.dci.2015.08.012
Brinchmann MF, Patel DM, Pinto N, Iversen MH (2018) Functional aspects of fish mucosal lectins interaction with non-self. Molecules 23(5):1119. https://doi.org/10.3390/molecules23051119
Brubaker SW, Bonham KS, Zanoni I, Kagan JC (2015) Innate immune pattern recognition: a cell biological perspective. Ann Rev Immunol 33(33):257–290. https://doi.org/10.1146/annurev-immunol-032414-112240
Brugman S (2016) The zebrafish as a model to study intestinal inflammation. Dev Comp Immunol 64:82–92. https://doi.org/10.1016/j.dci.2016.02.020
Brunner SR, Varga JFA, Dixon B (2020) Antimicrobial peptides of salmonid fish: from form to function. Biol Basel 9(8):233. https://doi.org/10.3390/biology9080233
Cao DY, Li JF, Huang BS, Zhang JD, Pan CH, Huang JS, Zhou H, Ma Q, Chen G, Wang ZL (2020) RNA-seq analysis reveals divergent adaptive response to hyper- and hypo-salinity in cobia, Rachycentron canadum. Fish Physiol Biochem 46(5):1713–1727. https://doi.org/10.1007/s10695-020-00823-7
Castro R, Martin SAM, Zou J, Secombes CJ (2010) Establishment of an IFN-gamma specific reporter cell line in fish. Fish Shellfish Immun 28(2):312–319. https://doi.org/10.1016/j.fsi.2009.11.010
Chatterjee A, Roy D, Patnaik E, Nongthomba U (2016) Muscles provide protection during microbial infection by activating innate immune response pathways in Drosophila and zebrafish. Dis Model Mech 9(6):697–705. https://doi.org/10.1242/dmm.022665
Chaturvedi P, Bhat RAH, Pande A (2020) Antimicrobial peptides of fish: innocuous alternatives to antibiotics. Rev Aquacult 12(1):85–106. https://doi.org/10.1111/raq.12306
Chaves-Pozo E, Zou J, Secombes CJ, Cuesta A, Tafalla C (2010) The rainbow trout (Oncorhynchus mykiss) interferon response in the ovary. Mol Immunol 47(9):1757–1764. https://doi.org/10.1016/j.molimm.2010.02.030
Chen XH, Wang Q, Yang CR, Rao YL, Li QM, Wan QY, Peng LM, Wu SQ, Su JG (2013) Identification, expression profiling of a grass carp TLR8 and its inhibition leading to the resistance to reovirus in CIK cells. Dev Comp Immunol 41(1):82–93. https://doi.org/10.1016/j.dci.2013.04.015
Chen YM, Kuo CE, Chen GR, Kao YT, Zou J, Secombes CJ, Chen TY (2014) Functional analysis of an orange-spotted grouper (Epinephelus coioides) interferon gene and characterisation of its expression in response to nodavirus infection. Dev Comp Immunol 46(2):117–128. https://doi.org/10.1016/j.dci.2014.04.004
Chen RN, Su YQ, Wang J, Liu M, Qiao Y, Mao Y, Ke QZ, Han KH, Zheng WQ, Zhang JS, Wu CW (2015) Molecular characterization and expression analysis of interferon-gamma in the large yellow croaker Larimichthys crocea. Fish Shellfish Immun 46(2):596–602. https://doi.org/10.1016/j.fsi.2015.07.008
Chen XY, Yi YH, Bian C, You XX, Shi Q (2020) Putative antimicrobial peptides in fish: using zebrafish as a representative. Protein Peptide Lett 27(11):1059–1067. https://doi.org/10.2174/0929866527666200517104610
Chi H, Sun L (2016) Neutrophils of Scophthalmus maximus produce extracellular traps that capture bacteria and inhibit bacterial infection. Dev Comp Immunol 56:7–12. https://doi.org/10.1016/j.dci.2015.11.005
Chirumbolo S (2012) State-of-the-art review about basophil research in immunology and allergy: is the time right to treat these cells with the respect they deserve? Blood Transfus-Italy 10(2):148–164. https://doi.org/10.2450/2011.0020-11
Collin M, Bigley V (2018) Human dendritic cell subsets: an update. Immunology 154(1):3–20. https://doi.org/10.1111/imm.12888
Coscelli G, Bermudez R, Ronza P, Losada AP, Quiroga MI (2016) Immunohistochemical study of inducible nitric oxide synthase and tumour necrosis factor alpha response in turbot (Scophthalmus maximus) experimentally infected with Aeromonas salmonicida subsp salmonicida. Fish Shellfish Immun 56:294–302. https://doi.org/10.1016/j.fsi.2016.07.022
Covello JM, Bird S, Morrison RN, Battaglene SC, Secombes CJ, Nowak BF (2009) Cloning and expression analysis of three striped trumpeter (Latris lineata) pro-inflammatory cytokines, TNF-alpha, IL-1 beta and IL-8, in response to infection by the ectoparasitic, Chondracanthus goldsmidi. Fish Shellfish Immun 26(5):773–786. https://doi.org/10.1016/j.fsi.2009.03.012
Crespo D, Bonnet E, Roher N, MacKenzie SA, Krasnov A, Goetz FW, Bobe J, Planas JV (2010) Cellular and molecular evidence for a role of tumor necrosis factor alpha in the ovulatory mechanism of trout. Reprod Biol Endocrin 8:34. https://doi.org/10.1186/1477-7827-8-34
Crespo D, Goetz FW, Planas JV (2015) Luteinizing hormone induces ovulation via tumor necrosis factor alpha-dependent increases in prostaglandin F-2 alpha in a nonmammalian vertebrate. Sci Rep-Uk 5:14210. https://doi.org/10.1038/srep14210
Cuesta A, Esteban MA, Meseguer J (2008) The expression profile of TLR9 mRNA and CpG ODNs immunostimulatory actions in the teleost gilthead seabream points to a major role of lymphocytes. Cell Mol Life Sci 65(13):2091–2104. https://doi.org/10.1007/s00018-008-8146-7
Dahle MK, Wessel O, Timmerhaus G, Nyman IB, Jorgensen SM, Rimstad E, Krasnov A (2015) Transcriptome analyses of Atlantic salmon (Salmo salar L.) erythrocytes infected with piscine orthoreovirus (PRV). Fish Shellfish Immun 45(2):780–790
Dang YF, Xu XY, Shen YB, Hu MY, Zhang M, Li LS, Lv LQ, Li JL (2016) Transcriptome Analysis of the Innate Immunity-Related Complement System in Spleen Tissue of Ctenopharyngodon idella Infected with Aeromonas hydrophila. PLoS One 11(7):e0157413. https://doi.org/10.1371/journal.pone.0157413
Dash S, Das SK, Samal J, Thatoi HN (2018) Epidermal mucus, a major determinant in fish health: a review. Iran J Vet Res 19(2):72–81
Dawood MAO (2020) Nutritional immunity of fish intestines: important insights for sustainable aquaculture. Rev Aquac 13:642–663. https://doi.org/10.1111/raq.12492
de Magalhaes CR, Schrama D, Farinha AP, Revets D, Kuehn A, Planchon S, Rodrigues PM, Cerqueira M (2020) Protein changes as robust signatures of fish chronic stress: a proteomics approach to fish welfare research. BMC Genomics 21(1). https://doi.org/10.1186/s12864-020-6728-4
Dezfuli BS, Bosi G, DePasquale JA, Manera M, Giari L (2016) Fish innate immunity against intestinal helminths. Fish Shellfish Immun 50:274–287. https://doi.org/10.1016/j.fsi.2016.02.002
Ding X, Lu DQ, Hou QH, Li SS, Liu XC, Zhang Y, Lin HR (2012) Orange-spotted grouper (Epinephelus coioides) toll-like receptor 22: molecular characterization, expression pattern and pertinent signaling pathways. Fish Shellfish Immun 33(3):494–503. https://doi.org/10.1016/j.fsi.2012.05.034
Ding Y, Guan YY, Huang XH, Ao JQ, Chen XH (2019) Characterization and function of a group II type I interferon in the perciform fish, large yellow croaker (Larimichthys crocea). Fish Shellfish Immun 86:152–159. https://doi.org/10.1016/j.fsi.2018.11.036
Dong XY, Su BF, Zhou S, Shang M, Yan H, Liu FQ, Gao CB, Tan FH, Li C (2016) Identification and expression analysis of toll-like receptor genes (TLR8 and TLR9) in mucosal tissues of turbot (Scophthalmus maximus L.) following bacterial challenge. Fish Shellfish Immun 58:309–317. https://doi.org/10.1016/j.fsi.2016.09.021
Drzewina A (1909) Leucocytes with acidophilic granulations in blood of teleostian fishes (Preliminary note). Cr Soc Biol 66:514–516
Du XG, Li D, Li YK, Wu JY, Huang AQ, Bu GX, Meng FY, Kong FL, Cao XH, Han XF, Pan XF, Yu GZ, Yang SY, Zeng XY (2019a) Clone, identification and functional character of two toll-like receptor 5 molecules in Schizothorax prenanti. Fish Shellfish Immun 95:81–92. https://doi.org/10.1016/j.fsi.2019.10.027
Du XG, Wu JY, Li YK, Xi PZ, Li D, Yang XX, Yu GZ, Bu GX, Huang AQ, Meng FY, Kong FL, Cao XH, Han XF, Pan XF, Yang SY, Zeng XY (2019b) Multiple subtypes of TLR22 molecule from Schizothorax prenanti present the functional diversity in ligand recognition and signal activation. Fish Shellfish Immun 93:986–996. https://doi.org/10.1016/j.fsi.2019.08.042
Elvitigala DAS, Premachandra HKA, Yeo SY, Choi CY, Whang I, Lee J (2015) Molecular profile and expressional modulation of a Toll like receptor-1 homolog from rock bream (Oplegnathus fasciatus). Genes Genom 37(5):459–470. https://doi.org/10.1007/s13258-015-0275-4
Faeste CK, Tartor H, Moen A, Kristoffersen AB, Dhanasiri AKS, Anonsen JH, Furmanek T, Grove S (2020) Proteomic profiling of salmon skin mucus for the comparison of sampling methods. J Chromatogr B 1138:121965. https://doi.org/10.1016/j.jchromb.2019.121965
Fan ZJ, Jia QJ, Yao CL (2015) Characterization and expression analysis of Toll-like receptor 2 gene in large yellow croaker, Larimichthys crocea. Fish Shellfish Immun 44(1):129–137. https://doi.org/10.1016/j.fsi.2015.01.037
Fan YD, Zhou Y, Zeng LB, Jiang N, Liu WZ, Zhao JQ, Zhong QW (2018) Identification, structural characterization, and expression analysis of toll-like receptors 2 and 3 from gibel carp (Carassius auratus gibelio). Fish Shellfish Immun 72:629–638. https://doi.org/10.1016/j.fsi.2017.11.044
Fan HY, Wang LY, Wen HS, Wang KQ, Qi X, Li JF, He F, Li Y (2019) Genome-wide identification and characterization of toll-like receptor genes in spotted sea bass (Lateolabrax maculatus) and their involvement in the host immune response to Vibrio harveyi infection. Fish Shellfish Immun 92:782–791. https://doi.org/10.1016/j.fsi.2019.07.010
Fan GY, Song Y, Yang LD, Huang XY, Zhang SY, Zhang MQ, Yang XW, Chang Y, Zhang H, Li YX, Liu SS, Yu LL, Chu J, Seim I, Feng CG, Near TJ, Wing RA, Wang W, Wang K, Wang J, Xu X, Yang HM, Liu X, Chen NS, He SP (2020) Initial data release and announcement of the 10,000 Fish Genomes Project (Fish10K). Gigascience 9(8):giaa080. https://doi.org/10.1093/gigascience/giaa080
Feng J, Lin P, Wang Y, Guo S, Zhang Z, Yu L (2017) Identification of a type I interferon (IFN) gene from Japanese eel and its expression analysis in vivo and in vitro. Agric Gene 5:7. https://doi.org/10.1016/j.aggene.2017.07.003
Ferraresso S, Bonaldo A, Parma L, Buonocore F, Scapigliati G, Gatta PP, Bargelloni L (2016) Ontogenetic onset of immune-relevant genes in the common sole (Solea solea). Fish Shellfish Immun 57:278–292. https://doi.org/10.1016/j.fsi.2016.08.044
Fink IR, Pietretti D, Voogdt CGP, Westphal AH, Savelkoul HFJ, Forlenza M, Wiegertjes GF (2016) Molecular and functional characterization of Toll-like receptor (Tlr)1 and Tlr2 in common carp (Cyprinus carpio). Fish Shellfish Immun 56:70–83. https://doi.org/10.1016/j.fsi.2016.06.049
Flerova EA, Balabanova LV (2013) Ultrastructure of granulocytes of teleost fish (Salmoniformes, Cypriniformes, Perciformes). J Evol Biochem Phys 49(2):223–233. https://doi.org/10.1134/S0022093013020126
Forlenza M, Fink IR, Raes G, Wiegertjes GF (2011) Heterogeneity of macrophage activation in fish. Dev Comp Immunol 35(12):1246–1255. https://doi.org/10.1016/j.dci.2011.03.008
Forn-Cuni G, Reis ES, Dios S, Posada D, Lambris JD, Figueras A, Novoa B (2014) The evolution and appearance of C3 duplications in fish originate an exclusive teleost c3 gene form with anti-inflammatory activity. PLoS One 9(6):e99673. https://doi.org/10.1371/journal.pone.0099673
Franch R, Cardazzo B, Antonello J, Castagnaro M, Patarnello T, Bargelloni L (2006) Full-length sequence and expression analysis of Toll-like receptor 9 in the gilthead seabream (Sparus aurata L.). Gene 378:42–51. https://doi.org/10.1016/j.gene.2006.04.025
Fu YW, Zhu CK, Zhang QZ, Hou TL (2019) Molecular characterization, expression analysis, and ontogeny of complement component C9 in southern catfish (Silurus meridionalis). Fish Shellfish Immun 86:449–458. https://doi.org/10.1016/j.fsi.2018.11.069
Galbraith CG, Sheetz MP (1998) Ventral and dorsal traction forces generated by fish keratocytes. Mol Biol Cell 9:33a
Galindo-Villegas J, Mulero I, Garcia-Alcazar A, Munoz I, Penalver-Mellado M, Streitenberger S, Scapigliati G, Meseguer J, Mulero V (2013) Recombinant TNF alpha as oral vaccine adjuvant protects European sea bass against vibriosis: insights into the role of the CCL25/CCR9 axis. Fish Shellfish Immun 35(4):1260–1271. https://doi.org/10.1016/j.fsi.2013.07.046
Gao H, Wu L, Sun JS, Geng XY, Pan BP (2013) Molecular characterization and expression analysis of Toll-like receptor 21 cDNA from Paralichthys olivaceus. Fish Shellfish Immun 35(4):1138–1145. https://doi.org/10.1016/j.fsi.2013.07.027
Gao QX, Yue YF, Min MH, Peng SM, Shi ZH, Sheng WQ, Zhang T (2018) Characterization of TLR5 and TLR9 from silver pomfret (Pampus argenteus) and expression profiling in response to bacterial components. Fish Shellfish Immun 80:241–249. https://doi.org/10.1016/j.fsi.2018.06.014
Gao QX, Tang QY, Zhao M, Min MH, Wang Q, Peng SM, Ma LB (2020) Molecular characterization of TLR3 and TRIL in silvery pomfret (Pampus argenteus) and their expression profiles in response to bacterial components. Int J Biol Macromol 155:805–813. https://doi.org/10.1016/j.ijbiomac.2020.03.246
Gerdol M, Buonocore F, Scapigliati G, Pallavicini A (2015) Analysis and characterization of the head kidney transcriptome from the Antarctic fish Trematomus bernacchii (Teleostea, Notothenioidea): a source for immune relevant genes. Mar Genom 20:13–15. https://doi.org/10.1016/j.margen.2014.12.005
Godahewa GI, Bathige SDNK, Herath HMLPB, Noh JK, Lee J (2015) Characterization of rock bream (Oplegnathus fasciatus) complement components C1r and C1s in terms of molecular aspects, genomic modulation, and immune responsive transcriptional profiles following bacterial and viral pathogen exposure. Fish Shellfish Immun 46(2):656–668. https://doi.org/10.1016/j.fsi.2015.07.026
Godahewa GI, Perera NCN, Bathige SDNK, Nam BH, Noh JK, Lee J (2016) Complement factor D homolog involved in the alternative complement pathway of rock bream (Oplegnathus fasciatus): molecular and functional characterization and immune responsive mRNA expression analysis. Fish Shellfish Immun 55:423–433. https://doi.org/10.1016/j.fsi.2016.06.018
Gong YW, Feng SS, Li SQ, Zhang Y, Zhao ZX, Hu M, Xu P, Jiang YL (2017) Genome-wide characterization of Toll-like receptor gene family in common carp (Cyprinus carpio) and their involvement in host immune response to Aeromonas hydrophila infection. Comp Biochem Phys D 24:89–98. https://doi.org/10.1016/j.cbd.2017.08.003
Grayfer L, Hodgkinson JW, Belosevic M (2014) Antimicrobial responses of teleost phagocytes and innate immune evasion strategies of intracellular bacteria. Dev Comp Immunol 43(2):223–242. https://doi.org/10.1016/j.dci.2013.08.003
Grayfer L, Kerimoglu B, Yaparla A, Hodgkinson JW, Xie J, Belosevic M (2018) Mechanisms of fish macrophage antimicrobial immunity. Front Immunol 9:1105
Guo BY, Wu CW, Lv ZM, Liu CL (2016) Characterisation and expression analysis of two terminal complement components: C7 and C9 from large yellow croaker, Larimichthys crocea. Fish Shellfish Immun 51:211–219. https://doi.org/10.1016/j.fsi.2016.01.015
Han C, Li Q, Zhang ZP, Huang JR (2017) Characterization, expression, and evolutionary analysis of new TLR3 and TLR5M genes cloned from the spiny eel Mastacembelus armatus. Dev Comp Immunol 77:174–187. https://doi.org/10.1016/j.dci.2017.08.007
Han C, Li Q, Liu JM, Hao ZQ, Huang JR, Zhang Y (2019) Characterization, evolution, and expression analysis of TLR7 gene subfamily members in Mastacembelus armatus (Synbranchiformes: Mastacembelidae). Dev Comp Immunol 95:77–88. https://doi.org/10.1016/j.dci.2019.02.002
Haugarvoll E, Bjerkas I, Nowak BF, Hordvik I, Koppang EO (2008) Identification and characterization of a novel intraepithelial lymphoid tissue in the gills of Atlantic salmon. J Anat 213(2):202–209
Haugland GT, Jordal A-EO, Wergeland HI (2012) Characterization of small, mononuclear blood cells from salmon having high phagocytic capacity and ability to differentiate into dendritic like cells. PLoS One 7(11):e49260
Havixbeck JJ, Barreda DR (2015) Neutrophil development, migration, and function in teleost fish. Biology (Basel) 4(4):715–734. https://doi.org/10.3390/biology4040715
He LB, Wang H, Luo LF, Jiang SH, Liu LY, Li YM, Huang R, Liao LJ, Zhu ZY, Wang YP (2016) Characterization, expression analysis and localization pattern of toll-like receptor 1 (tlr1) and toll-like receptor 2 (tlr2) genes in grass carp Ctenopharyngodon idella. J Fish Biol 89(2):1434–1440. https://doi.org/10.1111/jfb.12997
Hirono I, Nam BH, Kurobe T, Aoki T (2000) Molecular cloning, characterization, and expression of TNF cDNA and gene from Japanese flounder Paralichthys olivaceus. J Immunol 165(8):4423–4427. https://doi.org/10.4049/jimmunol.165.8.4423
Hirono I, Takami M, Miyata M, Miyazaki T, Han HJ, Takano T, Endo M, Aoki T (2004) Characterization of gene structure and expression of two toll-like receptors from Japanese flounder, Paralichthys olivaceus. Immunogenetics 56(1):38–46. https://doi.org/10.1007/s00251-004-0657-2
Hodgkinson JW, Grayfer L, Belosevic M (2015) Biology of bony fish macrophages. Biology 4(4):881–906
Hogan SP, Rosenberg HF, Moqbel R, Phipps S, Foster PS, Lacy P, Kay AB, Rothenberg ME (2008) Eosinophils: biological properties and role in health and disease. Clin Exp Allergy 38(5):709–750. https://doi.org/10.1111/j.1365-2222.2008.02958.x
Hong S, Li RG, Xu QQ, Secombes CJ, Wang TH (2013) Two types of TNF-alpha exist in teleost fish: phylogeny, expression, and bioactivity analysis of type-II TNF-alpha 3 in rainbow trout oncorhynchus mykiss. J Immunol 191(12):5959–5972. https://doi.org/10.4049/jimmunol.1301584
Hu GB, Li XP, Liu DH, Liu QM, Zhang SC (2015a) A toll-like receptor 3 homologue that is up-regulated by poly I:C and DNA virus in turbot Scophthalmus maximus. J Fish Biol 86(2):431–447. https://doi.org/10.1111/jfb.12559
Hu GB, Zhang SF, Yang X, Liu DH, Liu QM, Zhang SC (2015b) Cloning and expression analysis of a Toll-like receptor 22 (tlr22) gene from turbot, Scophthalmus maximus. Fish Shellfish Immun 44(2):399–409. https://doi.org/10.1016/j.fsi.2015.03.001
Huang XN, Wang ZY, Yao CL (2011) Characterization of Toll-like receptor 3 gene in large yellow croaker, Pseudosciaena crocea. Fish Shellfish Immun 31(1):98–106. https://doi.org/10.1016/j.fsi.2011.04.009
Huang R, Dong F, Jang SH, Liao LJ, Zhu ZY, Wang YP (2012) Isolation and analysis of a novel grass carp toll-like receptor 4 (tlr4) gene cluster involved in the response to grass carp reovirus. Dev Comp Immunol 38(2):383–388. https://doi.org/10.1016/j.dci.2012.06.002
Huang WJ, Shen YB, Xu XY, Hu MY, Li JL (2015) Identification and characterization of the TLR18 gene in grass carp (Ctenopharyngodon idella). Fish Shellfish Immun 47(2):681–688. https://doi.org/10.1016/j.fsi.2015.09.052
Huang WJ, Yang XM, Shen YB, Xu XY, Li LS, Wang RQ, Li JL (2016) Identification and functional analysis of the toll-like receptor 20.2 gene in grass carp, Ctenopharyngodon idella. Dev Comp Immunol 65:91–97. https://doi.org/10.1016/j.dci.2016.06.019
Huo RX, Zhao XY, Han JL, Xu TJ (2018) Genomic organization, evolution and functional characterization of soluble toll-like receptor 5 (TLR5S) in miiuy croaker (Miichthys miiuy). Fish Shellfish Immun 80:109–114. https://doi.org/10.1016/j.fsi.2018.05.048
Hwang SD, Asahi T, Kondo H, Hirono I, Aoki T (2010) Molecular cloning and expression study on Toll-like receptor 5 paralogs in Japanese flounder, Paralichthys olivaceus. Fish Shellfish Immun 29(4):630–638. https://doi.org/10.1016/j.fsi.2010.06.011
Hwang SD, Kondo H, Hirono I, Aoki T (2011) Molecular cloning and characterization of Toll-like receptor 14 in Japanese flounder, Paralichthys olivaceus. Fish Shellfish Immun 30(1):425–429. https://doi.org/10.1016/j.fsi.2010.08.005
Hwang SD, Ohtani M, Hikima JI, Jung TS, Kondo H, Hirono I, Aoki T (2012) Molecular cloning and characterization of Toll-like receptor 3 in Japanese flounder, Paralichthys olivaceus. Dev Comp Immunol 37(1):87–96. https://doi.org/10.1016/j.dci.2011.12.004
Hwang JY, Kwon MG, Jung SH, Park MA, Kim DW, Cho WS, Park JW, Son MH (2017) RNA-Seq transcriptome analysis of the olive flounder (Paralichthys olivaceus) kidney response to vaccination with heat-inactivated viral hemorrhagic septicemia virus. Fish Shellfish Immun 62:221–226. https://doi.org/10.1016/j.fsi.2017.01.016
Inami M, Taverne-Thiele AJ, Schroder MB, Kiron V, Rombout JHWM (2009) Immunological differences in intestine and rectum of Atlantic cod (Gadus morhua L.). Fish Shellfish Immun 26(5):751–759. https://doi.org/10.1016/j.fsi.2009.03.007
Ishii A, Matsuo A, Sawa H, Tsujita T, Shida K, Matsumoto M, Seya T (2007) Lamprey TLRs with properties distinct from those of the variable lymphocyte receptors. J Immunol 178(1):397–406. https://doi.org/10.4049/jimmunol.178.1.397
Jault C, Pichon L, Chluba J (2004) Toll-like receptor gene family and TIR-domain adapters in Danio rerio. Mol Immunol 40(11):759–771. https://doi.org/10.1016/j.molimm.2003.10.001
Jayaramu PK, Tripathi G, Kumar AP, Keezhedath J, Pathan MK, Kurcheti PP (2017) Studies on expression pattern of toll-like receptor 5 (TLR5) in Edwardsiella tarda infected Pangasianodon hypophthalmus. Fish Shellfish Immun 63:68–73. https://doi.org/10.1016/j.fsi.2017.01.041
Jiang Y, He LB, Ju CS, Pei YY, Ji M, Li YM, Liao LJ, Jang SH, Zhu ZY, Wang YP (2015) Isolation and expression of grass carp toll-like receptor 5a (CiTLR5a) and 5b (CiTLR5b) gene involved in the response to flagellin stimulation and grass carp reovirus infection. Fish Shellfish Immun 44(1):88–99. https://doi.org/10.1016/j.fsi.2015.01.024
Jin SZ, Zhao X, Wang HQ, Su JM, Wang JA, Ding CH, Li YG, Xiao TY (2018) Molecular characterization and expression of TLR7 and TLR8 in barbel chub (Squaliobarbus curriculus): responses to stimulation of grass carp reovirus Check for and lipopolysaccharide. Fish Shellfish Immun 83:292–307. https://doi.org/10.1016/j.fsi.2018.09.035
Johansen LH, Thim HL, Jorgensen SM, Afanasyev S, Strandskog G, Taksdal T, Fremmerlid K, McLoughlin M, Jorgensen JB, Krasnov A (2015) Comparison of transcriptomic responses to pancreas disease (PD) and heart and skeletal muscle inflammation (HSMI) in heart of Atlantic salmon (Salmo salar L). Fish Shellfish Immun 46(2):612–623. https://doi.org/10.1016/j.fsi.2015.07.023
Jorgensen JB, Johansen A, Hegseth MN, Zou J, Robertsen B, Collet B, Secombes CJ (2007) A recombinant CHSE-214 cell line expressing an Mx1 promoter-reporter system responds to both interferon type I and type II from salmonids and represents a versatile tool to study the IFN-system in teleost fish. Fish Shellfish Immun 23(6):1294–1303. https://doi.org/10.1016/j.fsi.2007.07.008
Jung TY, Hikima J, Ohtani M, Jang HB, del Castillo CS, Nho SW, Cha IS, Park SB, Aoki T, Jung TS (2012) Recombinant interferon-gamma activates immune responses against Edwardsiella tarda infection in the olive flounder, Paralichthys olivaceus. Fish Shellfish Immun 33(2):197–203. https://doi.org/10.1016/j.fsi.2012.04.015
Jurado C, Haserick JR, Lee J (2005) Slipping or gripping? Fluorescent speckle microscopy in fish keratocytes reveals two different mechanisms for generating a retrograde flow of actin. Mol Biol Cell 16(2):507–518. https://doi.org/10.1091/mbc.E04-10-0860
Kaitetzidou E, Crespo D, Vraskou Y, Antonopoulou E, Planas JV (2012) Transcriptomic Response of Skeletal Muscle to Lipopolysaccharide in the Gilthead Seabream (Sparus aurata). Mar Biotechnol 14(5):605–619. https://doi.org/10.1007/s10126-012-9469-9
Kania PW, Sorensen RR, Koch C, Brandt J, Kliem A, Vitved L, Hansen S, Skjodt K (2010) Evolutionary conservation of mannan-binding lectin (MBL) in bony fish: Identification, characterization and expression analysis of three bona fide collectin homologues of MBL in the rainbow trout (Oncorhynchus mykiss). Fish Shellfish Immun 29(6):910–920. https://doi.org/10.1016/j.fsi.2010.07.020
Kanwal Z, Wiegertjes GF, Veneman WJ, Meijer AH, Spaink HP (2014) Comparative studies of Toll-like receptor signalling using zebrafish. Dev Comp Immunol 46(1):35–52. https://doi.org/10.1016/j.dci.2014.02.003
Karlsen C, Sorum H, Willassen NP, Asbakk K (2012) Moritella viscosa bypasses Atlantic salmon epidermal keratocyte clearing activity and might use skin surfaces as a port of infection. Vet Microbiol 154(3–4):353–362. https://doi.org/10.1016/j.vetmic.2011.07.024
Kasamatsu J, Oshiumi H, Matsumoto M, Kasahara M, Seya T (2010) Phylogenetic and expression analysis of lamprey toll-like receptors. Dev Comp Immunol 34(8):855–865. https://doi.org/10.1016/j.dci.2010.03.004
Katzenback BA, Belosevic M (2009) Isolation and functional characterization of neutrophil-like cells, from goldfish (Carassius auratus L.) kidney. Dev Comp Immunol 33(4):601–611. https://doi.org/10.1016/j.dci.2008.10.011
Kelenyi G, Nemeth A (1969) Comparative histochemistry and electron microscopy of eosinophil leucocytes of vertebrates.1. A study of avian, reptile, amphibian and fish leucocytes. Acta Biol Acad Sci H 20(4):405
Kim YK, Lee JS, Jung JW, Hikima J, Ohtani M, Jang HB, Nho SW, Cha IS, Park SB, Lee JH, Aoki T, Jung TS (2017) Characterization of a specific monoclonal antibody against immunoglobulin light kappa/L1 chain in olive flounder (Paralichthys olivaceus). Fish Shellfish Immun 60:88–96. https://doi.org/10.1016/j.fsi.2016.11.023
Kinoshita S, Biswas G, Kono T, Hikima J, Sakai M (2014) Presence of two tumor necrosis factor (tnf)-alpha homologs on different chromosomes of zebrafish (Danio rerio) and medaka (Oryzias latipes). Mar Genom 13:1–9. https://doi.org/10.1016/j.margen.2013.10.004
Kobis JM, Rebl A, Kuhn C, Korytar T, Kollner B, Goldammer T (2015) Comprehensive and comparative transcription analyses of the complement pathway in rainbow trout. Fish Shellfish Immun 42(1):98–107. https://doi.org/10.1016/j.fsi.2014.10.032
Kordon AO, Karsi A, Pinchuk L (2018) Innate immune responses in fish: antigen presenting cells and professional phagocytes. Turk J Fish Aquat Sci 18(9):1123–1139. https://doi.org/10.4194/1303-2712-v18_9_11
Krol E, Noguera P, Shaw S, Costelloe E, Gajardo K, Valdenegro V, Bickerdike R, Douglas A, Martin SAM (2020) Integration of transcriptome, gross morphology and histopathology in the gill of sea farmed Atlantic Salmon (Salmo salar): lessons from multi-site sampling. Front Genet 11:610
Kumar G, Hummel K, Noebauer K, Welch TJ, Razzazi-Fazeli E, El-Matbouli M (2018) Proteome analysis reveals a role of rainbow trout lymphoid organs during Yersinia ruckeri infection process (vol 8, 13998, 2018). Sci Rep-Uk 8:15332. https://doi.org/10.1038/s41598-018-33154-y
Laghari ZA, Chen SN, Li L, Huang B, Gan Z, Zhou Y, Huo HJ, Hou J, Nie P (2018) Functional, signalling and transcriptional differences of three distinct type I IFNs in a perciform fish, the mandarin fish Siniperca chuatsi. Dev Comp Immunol 84:94–108. https://doi.org/10.1016/j.dci.2018.02.008
Lai RF, Liu H, Jakovlic I, Zhan FB, Wei J, Yang PH, Wang WM (2016) Molecular cloning and expression of toll-like receptor 4 (tlr4) in the blunt snout bream (Megalobrama amblycephala). Dev Comp Immunol 59:63–76. https://doi.org/10.1016/j.dci.2016.01.009
Lai RF, Jakovlic I, Liu H, Wei J, Zhan FB, Yang PH, Wang WM (2017a) Characterization and expression of Megalobrama amblycephala toll-like receptor 22 involved in the response to Aeromonas hydrophila. J Fish Biol 90(3):803–818. https://doi.org/10.1111/jfb.13199
Lai RF, Jakovlic I, Liu H, Zhan FB, Wei J, Wang WM (2017b) Molecular characterization and immunological response analysis of toll-like receptors. from the blunt snout bream (Megalobrama amblycephala). Dev Comp Immunol 67:471–475. https://doi.org/10.1016/j.dci.2016.09.005
Laing KJ, Wang TH, Zou J, Holland J, Hong SH, Bols N, Hirono I, Aoki T, Secombes CJ (2001) Cloning and expression analysis of rainbow trout Oncorhynchus mykiss tumour necrosis factor-alpha. Eur J Biochem 268(5):1315–1322. https://doi.org/10.1046/j.1432-1327.2001.01996.x
Lamas J, Ellis AE (1994) Atlantic Salmon (Salmo-Salar) neutrophil responses to Aeromonas-Salmonicida. Fish Shellfish Immun 4(3):201–219. https://doi.org/10.1006/fsim.1994.1019
Lee PT, Zou J, Holland JW, Martin SAM, Collet B, Kanellos T, Secombes CJ (2014) Identification and characterisation of TLR18-21 genes in Atlantic salmon (Salmo salar). Fish Shellfish Immun 41(2):549–559. https://doi.org/10.1016/j.fsi.2014.10.006
Lee FFY, Chuang HC, Chen NY, Nagarajan G, Chiou PP (2015) Toll-like receptor 9 alternatively spliced isoform negatively regulates TLR9 signaling in teleost fish. PLoS One 10(5):e0126388. https://doi.org/10.1371/journal.pone.0126388
Lee PT, Ho TH, Nguyen BT, Lin YL, Chiu PY (2020) Expression profile, subcellular localization and signaling pathway analysis of fish-specific TLR25 in Nile tilapia (Oreochromis niloticus). Fish Shellfish Immun 104:141–154. https://doi.org/10.1016/j.fsi.2020.05.028
Lewin HA, Robinson GE, Kress WJ, Baker WJ, Coddington J, Crandall KA, Durbin R, Edwards SV, Forest F, Gilbert MTP, Goldstein MM, Grigoriev IV, Hackett KJ, Haussler D, Jarvis ED, Johnson WE, Patrinos A, Richards S, Castilla-Rubio JC, van Sluys MA, Soltis PS, Xu X, Yang HM, Zhang GJ (2018) Earth BioGenome Project: sequencing life for the future of life. Proc Natl Acad Sci U S A 115(17):4325–4333. https://doi.org/10.1073/pnas.1720115115
Li XP, Sun L (2015) TLR7 is required for optimal immune defense against bacterial infection in tongue sole (Cynoglossus semilaevis). Fish Shellfish Immun 47(1):93–99. https://doi.org/10.1016/j.fsi.2015.08.025
Li X-P, Sun L (2017) A teleost complement factor Ba possesses antimicrobial activity and inhibits bacterial infection in fish. Dev Comp Immunol 71:49–58
Li MF, Zhang J (2016) CsTNF1, a teleost tumor necrosis factor that promotes antibacterial and antiviral immune defense in a manner that depends on the conserved receptor binding site. Dev Comp Immunol 55:65–75. https://doi.org/10.1016/j.dci.2015.10.010
Li J, Barreda DR, Zhang YA, Boshra H, Gelman AE, LaPatra S, Tort L, Sunyer JO (2007) Complement and B cell cooperation in teleost fish: role in phagocytosis and inflammation. Mol Immunol 44(1–3):205–205. https://doi.org/10.1016/j.molimm.2006.07.136
Li YW, Luo XC, Dan XM, Qiao W, Huang XZ, Li AX (2012) Molecular cloning of orange-spotted grouper (Epinephelus coioides) TLR21 and expression analysis post Cryptocaryon irritans infection. Fish Shellfish Immun 32(3):476–481. https://doi.org/10.1016/j.fsi.2011.11.021
Li MF, Li J, Sun L (2016) CsMAP34, a teleost MAP with dual role: A promoter of MASP-assisted complement activation and a regulator of immune cell activity. Sci Rep-Uk 6. https://doi.org/10.1038/srep39287
Li H, Yang GW, Ma F, Li T, Yang HT, Rombout JHWM, An LG (2017a) Molecular characterization of a fish-specific toll-like receptor 22 (TLR22) gene from common carp (Cyprinus carpio L.): evolutionary relationship and induced expression upon immune stimulants. Fish Shellfish Immun 63:74–86. https://doi.org/10.1016/j.fsi.2017.02.009
Li YJ, Li YL, Cao XC, Jin XY, Jin TC (2017b) Pattern recognition receptors in zebrafish provide functional and evolutionary insight into innate immune signaling pathways. Cell Mol Immunol 14(1):80–89. https://doi.org/10.1038/cmi.2016.50
Li FX, Wang PF, Zhao C, Fan SG, Yan LL, Wang CY, Qiu LH (2018a) Molecular cloning of sea perch (Lateolabrax japonicus) TLR1 and analysis of its expression pattern after stimulation with various bacteria. Aquac Res 49(7):2455–2465. https://doi.org/10.1111/are.13705
Li H, Li T, Guo YJ, Li YJ, Zhang Y, Teng N, Zhang FM, Yang GW (2018b) Molecular characterization and expression patterns of a non-mammalian toll-like receptor gene (TLR21) in larvae ontogeny of common carp (Cyprinus carpio L.) and upon immune stimulation. BMC Vet Res 14. https://doi.org/10.1186/s12917-018-1474-4
Li YK, Wu JY, Li D, Huang AQ, Bu GX, Meng FY, Kong FL, Cao XH, Han XF, Pan XF, Fan W, Yang SY, Wang J, Zeng XY, Du XG (2018c) Teleost-specific TLR25 identified from Schizothorax prenanti may recognize bacterial/viral components and activate NF-kappa B and type I IFNs signaling pathways. Fish Shellfish Immun 82:361–370. https://doi.org/10.1016/j.fsi.2018.08.007
Li JN, Zhao YT, Cao SL, Wang H, Zhang JJ (2020) Integrated transcriptomic and proteomic analyses of grass carp intestines after vaccination with a double-targeted DNA vaccine of Vibrio mimicus. Fish Shellfish Immun 98:641–652. https://doi.org/10.1016/j.fsi.2019.10.045
Liang YS, Ding X, Yu X, Wang Y, Zhou Y, He JA, Shi Y, Zhang Y, Lin HR, Lu DQ (2018) Identification and functional characterization of Toll-like receptor 13 from orange-spotted grouper (Epinephelus coioides). Fish Shellfish Immun 74:309–317. https://doi.org/10.1016/j.fsi.2017.12.054
Liao ZW, Wan QY, Su H, Wu CS, Su JG (2017) Pattern recognition receptors in grass carp Ctenopharyngodon idella: I. Organization and expression analysis of TLRs and RLRs. Dev Comp Immunol 76:93–104. https://doi.org/10.1016/j.dci.2017.05.019
Lieschke GJ, Trede NS (2009) Fish immunology. Curr Biol 19(16):R678–R682. https://doi.org/10.1016/j.cub.2009.06.068
Lin KB, Ge H, Lin Q, Wu JS, He LB, Fang QS, Zhou C, Sun MQ, Huang ZC (2013) Molecular characterization and functional analysis of Toll-like receptor 3 gene in orange-spotted grouper (Epinephelus coioides). Gene 527(1):174–182. https://doi.org/10.1016/j.gene.2013.06.014
Lindell K, Fahlgren A, Hjerde E, Willassen NP, Fallman M, Milton DL (2012) Lipopolysaccharide O-antigen prevents phagocytosis of vibrio anguillarum by rainbow trout (Oncorhynchus mykiss) skin epithelial cells. PLoS One 7(5):e37678. https://doi.org/10.1371/journal.pone.0037678
Liu DW, Mai KS, Ai QH (2015) Tumor necrosis factor alpha is a potent regulator in fish adipose tissue. Aquaculture 436:65–71. https://doi.org/10.1016/j.aquaculture.2014.10.030
Liu QN, Yang TT, Wang C, Jiang SH, Zhang DZ, Tang BP, Ge BM, Wang JL, Wang D, Dai LS (2019) A non-mammalian Toll-like receptor 26 (TLR26)gene mediates innate immune responses in yellow catfish Pelteobagrus fulvidraco. Fish Shellfish Immun 95:491–497. https://doi.org/10.1016/j.fsi.2019.11.005
Liu FG, Wang TH, Petit J, Forlenza M, Chen XH, Chen LB, Zou J, Secombes CJ (2020a) Evolution of IFN subgroups in bony fish-2. analysis of subgroup appearance and expansion in teleost fish with a focus on salmonids. Fish Shellfish Immun 98:564–573. https://doi.org/10.1016/j.fsi.2020.01.039
Liu RR, Hu XC, Lü AJ, Song YJ, Lian ZY, Sun JF, Sung YY (2020b) Proteomic profiling of zebrafish challenged by spring viremia of carp virus provides insight into skin antiviral response. Zebrafish 17(2):91–103. https://doi.org/10.1089/zeb.2019.1843
Loken OM, Bjorgen H, Hordvik I, Koppang EO (2020) A teleost structural analogue to the avian bursa of Fabricius. J Anat 236(5):798–808. https://doi.org/10.1111/joa.13147
Lv JJ, Huang R, Li HY, Luo DJ, Liao LJ, Zhu ZY, Wang YP (2012) Cloning and characterization of the grass carp (Ctenopharyngodon idella) Toll-like receptor 22 gene, a fish-specific gene. Fish Shellfish Immun 32(6):1022–1031. https://doi.org/10.1016/j.fsi.2012.02.024
Ma JG, Li XY, Li YY, Li Y, Niu DC (2014) Toxic effects of paraquat on cytokine expression in common carp, cyprinus carpio L. J Biochem Mol Toxic 28(11):501–509. https://doi.org/10.1002/jbt.21590
Magnadottir B (2014) The immune response of Atlantic cod, Gadus morhua L. Icel Agr Sci 27:41–61
Martin SA, Douglas A, Houlihan DF, Secombes CJ (2010) Starvation alters the liver transcriptome of the innate immune response in Atlantic salmon (Salmo salar). BMC Genomics 11:418. https://doi.org/10.1186/1471-2164-11-418
Medina-Gali R, Bello-Perez M, Martinez-Lopez A, Falco A, Ortega-Villaizan MM, Encinar JA, Novo B, Coll J, Perez L (2018) Chromatin immunoprecipitation and high throughput sequencing of SVCV-infected zebrafish reveals novel epigenetic histone methylation patterns involved in antiviral immune response. Fish Shellfish Immun 82:514–521. https://doi.org/10.1016/j.fsi.2018.08.056
Melillo D, Varriale S, Giacomelli S, Natale L, Bargelloni L, Oreste U, Pinto MR, Coscia MR (2015) Evolution of the complement system C3 gene in Antarctic teleosts. Mol Immunol 66(2):299–309. https://doi.org/10.1016/j.molimm.2015.03.247
Meng FX, Wang RX, Xu TJ, Sun YE, Cheng YZ, Shi G (2012) An unexpected loss of domains in the conservative evolution ninth complement component in a higher teleost, Miichthys miiuy. Fish Shellfish Immun 32(6):1171–1178. https://doi.org/10.1016/j.fsi.2012.02.010
Milne DJ, Campoverde C, Andree KB, Zou J, Secombes CJ (2017) Two types of TNF alpha in meagre (Argyrosomus regius): discovery, distribution and expression modulation. Mol Immunol 92:136–145. https://doi.org/10.1016/j.molimm.2017.10.007
Milne DJ, Campoverde C, Andree KB, Chen X, Zou J, Secombes CJ (2018) The discovery and comparative expression analysis of three distinct type I interferons in the perciform fish, meagre (Argyrosomus regius). Dev Comp Immunol 84:123–132. https://doi.org/10.1016/j.dci.2018.02.001
Morera D, Roher N, Ribas L, Balasch JC, Donate C, Callol A, Boltana S, Roberts S, Goetz G, Goetz FW, MacKenzie SA (2011) RNA-Seq reveals an integrated immune response in nucleated erythrocytes. PLoS One 6(10):e26998
Munoz I, Sepulcre MP, Meseguer J, Mulero V (2013) Molecular cloning, phylogenetic analysis and functional characterization of soluble Toll-like receptor 5 in gilthead seabream, Sparus aurata. Fish Shellfish Immun 35(1):36–45. https://doi.org/10.1016/j.fsi.2013.03.374
Munoz I, Sepulcre MP, Meseguer J, Mulero V (2014) Toll-like receptor 22 of gilthead seabream, Sparus aurata: molecular cloning, expression profiles and post-transcriptional regulation. Dev Comp Immunol 44(1):173–179. https://doi.org/10.1016/j.dci.2013.12.003
Munoz-Atienza E, Aquilino C, Syahputra K, Al-Jubury A, Araujo C, Skov J, Kania PW, Hernandez PE, Buchmann K, Cintas LM, Tafalla C (2019) CK11, a teleost chemokine with a potent antimicrobial activity. J Immunol 202(3):857–870. https://doi.org/10.4049/jimmunol.1800568
Mutsuro J, Tanaka N, Kato Y, Dodds AW, Yano T, Nakao M (2005) Two divergent isotypes of the fourth complement component from a bony fish, the common carp (Cyprinus carpio). J Immunol 175(7):4508–4517. https://doi.org/10.4049/jimmunol.175.7.4508
Nadal AL, Ikeda-Ohtsubo W, Sipkema D, Peggs D, McGurk C, Forlenza M, Wiegertjes GF, Brugman S (2020) Feed, microbiota, and gut immunity: using the zebrafish model to understand fish health. Front Immunol 11:114. https://doi.org/10.3389/fimmu.2020.00114
Nakao M, Tsujikura M, Ichiki S, Vo TK, Somamoto T (2011) The complement system in teleost fish: progress of post-homolog-hunting researches. Dev Comp Immunol 35(12):1296–1308. https://doi.org/10.1016/j.dci.2011.03.003
Nie L, Cai SY, Shao JZ, Chen J (2018) Toll-like receptors, associated biological roles, and signaling networks in non-mammals. Front Immunol 9:1523. https://doi.org/10.3389/fimmu.2018.01523
Nilojan J, Bathige SDNK, Thulasitha WS, Kwon H, Jung S, Kim MJ, Nam BH, Lee J (2018) Transcriptional profiling, molecular cloning, and functional analysis of C1 inhibitor, the main regulator of the complement system in black rockfish, Sebastes schlegelii. Fish Shellfish Immun 75:263–273. https://doi.org/10.1016/j.fsi.2018.02.018
Odaka T, Suetake H, Maeda T, Miyadai T (2018) Teleost basophils have IgM-Dependent and dual Ig-independent degranulation systems. J Immunol 200(8):2767–2776. https://doi.org/10.4049/jimmunol.1701051
Okimura C, Taniguchi A, Nonaka S, Iwadate Y (2018) Rotation of stress fibers as a single wheel in migrating fish keratocytes. Sci Rep-Uk 8:10615. https://doi.org/10.1038/s41598-018-28875-z
Ortega-Villaizan M, Chico V, Falco A, Perez L, Coll JM, Estepa A (2009) The rainbow trout TLR9 gene and its role in the immune responses elicited by a plasmid encoding the glycoprotein G of the viral haemorrhagic septicaemia rhabdovirus (VHSV). Mol Immunol 46(8–9):1710–1717. https://doi.org/10.1016/j.molimm.2009.02.006
Oshiumi H, Tsujita T, Shida K, Matsumoto M, Ikeo K, Seya T (2003) Prediction of the prototype of the human Toll-like receptor gene family from the pufferfish, Fugu rubripes, genome. Immunogenetics 54(11):791–800. https://doi.org/10.1007/s00251-002-0519-8
Overgard AC, Nepstad I, Nerland AH, Patel S (2012) Characterisation and expression analysis of the Atlantic halibut (Hippoglossus hippoglossus L.) cytokines: IL-1 beta, IL-6, IL-11, IL-12 beta and IFN gamma. Mol Biol Rep 39(3):2201–2213. https://doi.org/10.1007/s11033-011-0969-x
Page DM, Wittamer V, Bertrand JY, Lewis KL, Pratt DN, Delgado N, Schale SE, McGue C, Jacobsen BH, Doty A, Pao Y, Yang HB, Chi NC, Magor BG, Traver D (2013) An evolutionarily conserved program of B-cell development and activation in zebrafish. Blood 122(8):E1–E11. https://doi.org/10.1182/blood-2012-12-471029
Palic D, Ostojic J, Andreasen CB, Roth JA (2007) Fish cast NETs: neutrophil extracellular traps are released from fish neutrophils. Dev Comp Immunol 31(8):805–816. https://doi.org/10.1016/j.dci.2006.11.010
Palti Y (2011) Toll-like receptors in bony fish: from genomics to function. Dev Comp Immunol 35(12):1263–1272. https://doi.org/10.1016/j.dci.2011.03.006
Palti Y, Gahr SA, Purcell MK, Hadidi S, Rexroad CE, Wiens GD (2010a) Identification, characterization and genetic mapping of TLR7, TLR8a1 and TLR8a2 genes in rainbow trout (Oncorhynchus mykiss). Dev Comp Immunol 34(2):219–233. https://doi.org/10.1016/j.dci.2009.10.002
Palti Y, Rodriguez MF, Gahr SA, Purcell MK, Rexroad CE, Wiens GD (2010b) Identification, characterization and genetic mapping of TLR1 loci in rainbow trout (Oncorhynchus mykiss). Fish Shellfish Immun 28(5–6):918–926. https://doi.org/10.1016/j.fsi.2010.02.002
Paludan SR, Pradeu T, Masters SL, Mogensen TH (2020) Constitutive immune mechanisms: mediators of host defence and immune regulation. Nat Rev Immunol. https://doi.org/10.1038/s41577-020-0391-5
Paneru B, Al-Tobasei R, Palti Y, Wiens GD, Salem M (2016) Differential expression of long non-coding RNAs in three genetic lines of rainbow trout in response to infection with Flavobacterium psychrophilum. Sci Rep-Uk 6:36032. https://doi.org/10.1038/srep36032
Pang JC, Gao FY, Wang M, Zhao JL, Lu MX (2017) Isolation and characterization of Toll-like receptor 21 and 22 genes from Nile tilapia, Oreochromis niloticus (Linnaeus). Aquac Res 48(7):3528–3544. https://doi.org/10.1111/are.13179
Passantino L, Cianciotta A, Patruno R, Ribaud MR, Jirillo E, Passantino GF (2005) Do fish thrombocytes play an immunological role? Their cytoenzymatic profiles and function during an accidental piscine candidiasis in aquarium. Immunopharm Immunot 27(2):345–356. https://doi.org/10.1081/Iph-200067959
Pei YY, Huang R, Li YM, Liao LJ, Zhu ZY, Wang YP (2015) Characterizations of four toll-like receptor 4s in grass carp Ctenopharyngodon idellus and their response to grass carp reovirus infection and lipopolysaccharide stimulation. J Fish Biol 86(3):1098–1108. https://doi.org/10.1111/jfb.12617
Pereiro P, Forn-Cuni G, Figueras A, Novoa B (2016) Pathogen-dependent role of turbot (Scophthalmus maximus) interferon-gamma. Fish Shellfish Immun 59:25–35. https://doi.org/10.1016/j.fsi.2016.10.021
Pereiro P, Figueras A, Novoa B (2019) Insights into teleost interferon-gamma biology: an update. Fish Shellfish Immun 90:150–164. https://doi.org/10.1016/j.fsi.2019.04.002
Pereiro P, Lama R, Moreira R, Valenzuela-Munoz V, Gallardo-Escarate C, Novoa B, Figueras A (2020) Potential involvement of lncRNAs in the modulation of the transcriptome response to nodavirus challenge in European Sea Bass (Dicentrarchus labrax L.). Biology-Basel 9(7):165. https://doi.org/10.3390/biology9070165
Phelan PE, Mellon MT, Kim CH (2005) Functional characterization of full-length TLR3, IRAK-4, and TRAF6 in zebrafish (Danio rerio). Mol Immunol 42(9):1057–1071. https://doi.org/10.1016/j.molimm.2004.11.005
Pietretti D, Wiegertjes GF (2014) Ligand specificities of Toll-like receptors in fish: indications from infection studies. Dev Comp Immunol 43(2):205–222. https://doi.org/10.1016/j.dci.2013.08.010
Pietretti D, Scheer M, Fink IR, Taverne N, Savelkoul HFJ, Spaink HP, Forlenza M, Wiegertjes GF (2014) Identification and functional characterization of nonmammalian Toll-like receptor 20. Immunogenetics 66(2):123–141. https://doi.org/10.1007/s00251-013-0751-4
Pijanowski L, Golbach L, Kolaczkowska E, Scheer M, Verburg-van Kemenade BML, Chadzinska M (2013) Carp neutrophilic granulocytes form extracellular traps via ROS-dependent and independent pathways. Fish Shellfish Immun 34(5):1244–1252. https://doi.org/10.1016/j.fsi.2013.02.010
Pijanowski L, Verburg-van Kemenade BML, Chadzinska M (2020) Chemokine CXCb1 stimulates formation of NETs in trunk kidney neutrophils of common carp. Dev Comp Immunol 103:103521. https://doi.org/10.1016/j.dci.2019.103521
Pleic IL, Secombes CJ, Bird S, Mladineo I (2014) Characterization of three pro-inflammatory cytokines, TNF alpha 1, TNF alpha 2 and IL-1 beta, in cage-reared Atlantic. bluefin tuna Thunnus thynnus. Fish Shellfish Immun 36(1):98–112. https://doi.org/10.1016/j.fsi.2013.10.011
Pleic IL, Buselic I, Trumbic Z, Bocina I, Sprung M, Mladineo I (2015) Expression analysis of the Atlantic bluefin tuna (Thunnus thynnus) pro-inflammatory cytokines, IL-1 beta, TNF alpha 1 and TNF alpha 2 in response to parasites Pseudocycnus appendiculatus (Copepoda) and Didymosulcus katsuwonicola (Digenea). Fish Shellfish Immun 45(2):946–954. https://doi.org/10.1016/j.fsi.2015.06.008
Press CM, Evensen O (1999) The morphology of the immune system in teleost fishes. Fish Shellfish Immun 9(4):309–318. https://doi.org/10.1006/fsim.1998.0181
Puente-Marin S, Nombela I, Ciordia S, Mena MC, Chico V, Coll J, Ortega-Villaizan MDM (2018) In silico functional networks identified in fish nucleated red blood cells by means of transcriptomic and proteomic profiling. Genes (Basel) 9(4)
Puente-Marin S, Thwaite R, Mercado L, Coll J, Roher N, Ortega-Villaizan MD (2019a) Fish red blood cells modulate immune genes in response to bacterial inclusion bodies made of TNF alpha and a G-VHSV fragment. Front Immunol 10:1055. https://doi.org/10.3389/fimmu.2019.01055
Puente-Marin S, Thwaite R, Mercado L, Coll J, Roher N, Ortega-Villaizan MDM (2019b) Fish red blood cells modulate immune genes in response to bacterial inclusion bodies made of TNFalpha and a G-VHSV fragment. Front Immunol 10:1055
Qi DL, Xia MZ, Chao Y, Zhao YL, Wu RR (2017) Identification, molecular evolution of toll-like receptors in a Tibetan schizothoracine fish (Gymnocypris eckloni) and their expression profiles in response to acute hypoxia. Fish Shellfish Immun 68:102–113. https://doi.org/10.1016/j.fsi.2017.07.014
Qi PZ, Wu B, Guo BY, Zhang C, Xu KD (2018a) The complement factor H (CFH) and its related protein 2 (CFHR2) mediating immune response in large yellow croaker Larimichthys crocea. Dev Comp Immunol 84:241–249. https://doi.org/10.1016/j.dci.2018.02.017
Qi ZT, Wang SS, Zhu XZ, Yang YY, Han PP, Zhang QH, Zhang SH, Shao R, Xu QQ, Wei QW (2018b) Molecular characterization of three toll-like receptors (TLR21, TLR22, and TLR25) from a primitive ray-finned fish Dabry’s sturgeon (Acipenser dabryanus). Fish Shellfish Immun 82:200–211. https://doi.org/10.1016/j.fsi.2018.08.033
Qi DL, Chao Y, Zhang CF, Wang ZJ, Wang W, Chen QC, Zheng ZQ, Zhang Z (2019) Duplication of toll-like receptor 22 in teleost fishes. Fish Shellfish Immun 94:752–760. https://doi.org/10.1016/j.fsi.2019.09.067
Qi ZT, Xu Y, Wang X, Wang SS, Zhang QH, Wang ZS, Gao Q (2020) TLR13, TLR22, TRAF6, and TAK1 in the soiny mullet (Liza haematocheila): molecular characterization and expression profiling analysis. Dev Comp Immunol 112:103774. https://doi.org/10.1016/j.dci.2020.103774
Qian TL, Wang KR, Mu YN, Ao JQ, Chen XH (2013) Molecular characterization and expression analysis of TLR 7 and TLR 8 homologs in large yellow croaker (Pseudosciaena crocea). Fish Shellfish Immun 35(3):671–679. https://doi.org/10.1016/j.fsi.2013.05.019
Qin CJ, Gong Q, Wen ZY, Yuan DY, Shao T, Li HT (2018) Molecular characterization and expression of toll-like receptor 5 genes from Pelteobagrus vachellii. Fish Shellfish Immun 75:198–207. https://doi.org/10.1016/j.fsi.2018.02.002
Qiu HT, Fernandes JMO, Hong WS, Wu HX, Zhang YT, Huang S, Liu DT, Yu H, Wang Q, You XX, Chen SX (2019) Paralogues from the expanded Tlr11 gene family in mudskipper (Boleophthalmus pectinirostris) are under positive selection and respond differently to LPS/Poly(I:C) challenge. Front Immunol 10:343. https://doi.org/10.3389/fimmu.2019.00343
Quiniou SMA, Boudinot P, Bengten E (2013) Comprehensive survey and genomic characterization of Toll-Like Receptors (TLRs) in channel catfish, Ictalurus punctatus: identification of novel fish TLRs. Fish Shellfish Immun 34(6):1731–1731. https://doi.org/10.1016/j.fsi.2013.03.291
Rauta PR, Nayak B, Das S (2012) Immune system and immune responses in fish and their role in comparative immunity study: a model for higher organisms. Immunol Lett 148(1):23–33. https://doi.org/10.1016/j.imlet.2012.08.003
Rauta PR, Samanta M, Dash HR, Nayak B, Das S (2014) Toll-like receptors (TLRs) in aquatic animals: Signaling pathways, expressions and immune responses. Immunol Lett 158(1–2):14–24. https://doi.org/10.1016/j.imlet.2013.11.013
Ream RA, Theriot JA, Somero GN (2003) Influences of thermal acclimation and acute temperature change on the motility of epithelial wound-healing cells (keratocytes) of tropical, temperate and Antarctic fish. J Exp Biol 206(24):4539–4551. https://doi.org/10.1242/jeb.00706
Rebl A, Siegl E, Kollner B, Fischer U, Seyfert HM (2007) Characterization of twin toll-like receptors from rainbow trout (Oncorhynchus mykiss): evolutionary relationship and induced expression by Aeromonas salmonicida salmonicida. Dev Comp Immunol 31(5):499–510. https://doi.org/10.1016/j.dci.2006.08.007
Reite OB (1998) Mast cells eosinophilic granule cells of teleostean fish: a review focusing on staining properties and functional responses. Fish Shellfish Immun 8(7):489–513. https://doi.org/10.1006/fsim.1998.0162
Reite OB, Evensen O (2006) Inflammatory cells of teleostean fish: a review focusing on mast cells/eosinophilic granule cells and rodlet cells. Fish Shellfish Immun 20(2):192–208. https://doi.org/10.1016/j.fsi.2005.01.012
Resseguier J, Dalum AS, Pasquier LD, Zhang Y, Koppang EO, Boudinot P, Wiegertjes GF (2020) Lymphoid tissue in teleost gills: variations on a theme. Biology (Basel) 9(6)
Reyes-Becerril M, Angulo C, Ascencio F (2015) Humoral immune response and TLR9 gene expression in Pacific red snapper (Lutjanus peru) experimentally exposed to Aeromonas veronii. Fish Shellfish Immun 42(2):289–296. https://doi.org/10.1016/j.fsi.2014.11.002
Reyes-Becerril M, Ascencio-Valle F, Hirono I, Kondo H, Jirapongpairoj W, Esteban MA, Alamillo E, Angulo C (2016) TLR21’s agonists in combination with Aeromonas antigens synergistically up-regulate functional TLR21 and cytokine gene expression in yellowtail leucocytes. Dev Comp Immunol 61:107–115. https://doi.org/10.1016/j.dci.2016.03.012
Reyes-Becerril M, Alamillo E, Rosales-Mendoza S, Ascencio F, Esteban MA, Angulo C (2017) Molecular characterization and expression analyses of toll like receptor-5 induced by Vibrio parahaemolyticus antigens in Pacific red snapper. Fish Shellfish Immun 68:180–189. https://doi.org/10.1016/j.fsi.2017.07.022
Rieger AM, Barreda DR (2011) Antimicrobial mechanisms of fish leukocytes. Dev Comp Immunol 35(12):1238–1245. https://doi.org/10.1016/j.dci.2011.03.009
Rivera A, Siracusa MC, Yap GS, Gause WC (2016) Innate cell communication kick-starts pathogen-specific immunity. Nat Immunol 17(4):356–363
Roach JC, Glusman G, Rowen L, Kaur A, Purcell MK, Smith KD, Hood LE, Aderem A (2005) The evolution of vertebrate Toll-like receptors. Proc Natl Acad Sci U S A 102(27):9577–9582. https://doi.org/10.1073/pnas.0502272102
Robertsen B (2006) The interferon system of teleost fish. Fish Shellfish Immun 20(2):172–191. https://doi.org/10.1016/j.fsi.2005.01.010
Roca FJ, Mulero I, Lopez-Munoz A, Sepulcre MP, Renshaw SA, Meseguer J, Mulero V (2008) Evolution of the inflammatory response in vertebrates: fish TNF-alpha is a powerful activator of endothelial cells but hardly activates phagocytes. J Immunol 181(7):5071–5081. https://doi.org/10.4049/jimmunol.181.7.5071
Rodriguez MF, Wiens GD, Purcell MK, Palti Y (2005) Characterization of Toll-like receptor 3 gene in rainbow trout (Oncorhynchus mykiss). Immunogenetics 57(7):510–519. https://doi.org/10.1007/s00251-005-0013-1
Rombout JHWM, Huttenhuis HBT, Picchietti S, Scapigliati G (2005) Phylogeny and ontogeny of fish leucocytes. Fish Shellfish Immun 19(5):441–455. https://doi.org/10.1016/j.fsi.2005.03.007
Rombout JHWM, Abelli L, Picchietti S, Scapigliati G, Kiron V (2011) Teleost intestinal immunology. Fish Shellfish Immun 31(5):616–626. https://doi.org/10.1016/j.fsi.2010.09.001
Ronza P, Losada AP, Villamarin A, Bermudez R, Quiroga MI (2015) Immunolocalization of tumor necrosis factor alpha in turbot (Scophthalmus maximus, L.) tissues. Fish Shellfish Immun 45(2):470–476. https://doi.org/10.1016/j.fsi.2015.04.032
Rosales C (2018) Neutrophil: a cell with many roles in inflammation or several cell types? Front Physiol 9:113. https://doi.org/10.3389/fphys.2018.00113
Roy S, Kumar V, Kumar V, Behera BK (2017) Acute phase proteins and their potential role as an indicator for fish health and in diagnosis of fish diseases. Protein Peptide Lett 24(1):78–89. https://doi.org/10.2174/0929866524666161121142221
Rozas-Serri M, Pena A, Maldonado L (2019) Transcriptomic profiles of post-smolt Atlantic salmon challenged with Piscirickettsia salmonis reveal a strategy to evade the adaptive immune response and modify cell-autonomous immunity. Fish Shellfish Immun 91:406–406
Sahoo BR (2020) Structure of fish Toll-like receptors (TLR) and NOD-like receptors (NLR). Int J Biol Macromol 161:1602–1617. https://doi.org/10.1016/j.ijbiomac.2020.07.293
Salazar C, Haussmann D, Kausel G, Figueroa J (2016) Molecular cloning of Salmo salar Toll-like receptors (TLR1, TLR22, TLR5M and TLR5S) and expression analysis in SHK-1 cells during Piscirickettsia salmonis infection. J Fish Dis 39(2):239–248. https://doi.org/10.1111/jfd.12354
Samanta M, Swain B, Basu M, Panda P, Mohapatra GB, Sahoo BR, Maiti NK (2012) Molecular characterization of toll-like receptor 2 (TLR2), analysis of its inductive expression and associated down-stream signaling molecules following ligands exposure and bacterial infection in the Indian major carp, rohu (Labeo rohita). Fish Shellfish Immun 32(3):411–425. https://doi.org/10.1016/j.fsi.2011.11.029
Samanta M, Basu M, Swain B, Panda P, Jayasankar P (2013) Molecular cloning and characterization of toll-like receptor 3, and inductive expression analysis of type I IFN, Mx and pro-inflammatory cytokines in the Indian carp, rohu (Labeo rohita). Mol Biol Rep 40(1):225–235. https://doi.org/10.1007/s11033-012-2053-6
Samanta M, Swain B, Basu M, Mahapatra G, Sahoo BR, Paichha M, Lenka SS, Jayasankar P (2014) Toll-like receptor 22 in Labeo rohita: molecular cloning, characterization, 3D modeling, and expression analysis following ligands stimulation and bacterial infection. Appl Biochem Biotech 174(1):309–327. https://doi.org/10.1007/s12010-014-1058-0
Samanta M, Basu M, Swain B, Paichha M, Lenka SS, Das S, Jayasankar P, Maiti NK (2017) Molecular cloning and characterization of LrTLR4, analysis of its inductive expression and associated down-stream signaling molecules following lipopolysaccharide stimulation and Gram-negative bacterial infection. Fish Shellfish Immun 60:164–176. https://doi.org/10.1016/j.fsi.2016.11.028
Scapigliati G, Fausto AM, Picchietti S (2018) Fish lymphocytes: an evolutionary equivalent of mammalian innate-like lymphocytes? Front Immunol 9:971. https://doi.org/10.3389/fimmu.0018.00971
Schmidt JE (1905) An article on normal and pathological histology of several types of cells in the mucous membrane of the human intestinal canal. Arch Mikro Anat Entw 66:12–40. https://doi.org/10.1007/Bf02979202
Secombes CJ, Zou J (2017) Evolution of interferons and interferon receptors. Front Immunol 8:209. https://doi.org/10.3389/fimmu.2017.00209
Semple SL, Dixon B (2020) Salmonid antibacterial immunity: an aquaculture perspective. Biology-Basel 9(10):331. https://doi.org/10.3390/biology9100331
Shabir U, Ali S, Magray AR, Ganai BA, Firdous P, Hassan T, Nazir R (2018) Fish antimicrobial peptides (AMP’S) as essential and promising molecular therapeutic agents: a review. Microb Pathogenesis 114:50–56. https://doi.org/10.1016/j.micpath.2017.11.039
Shan SJ, Liu DZ, Liu RR, Zhu YY, Li T, Zhang FM, An LG, Yang GW, Li H (2018a) Non-mammalian Toll-like receptor 18 (Tlr18) recognizes bacterial pathogens in common carp (Cyprinus carpio L.): indications for a role of participation in the NF-kappa B signaling pathway. Fish Shellfish Immun 72:187–198. https://doi.org/10.1016/j.fsi.2017.09.081
Shan SJ, Liu RR, Jiang L, Zhu YY, Li H, Xing WX, Yang GW (2018b) Carp Toll-like receptor 8 (Tlr8): an intracellular Tlr that recruits TIRAP as adaptor and activates AP-1 pathway in immune response. Fish Shellfish Immun 82:41–49. https://doi.org/10.1016/j.fsi.2018.08.001
Shao T, Zhu LY, Nie L, Shi W, Dong WR, Xiang LX, Shao JZ (2015) Characterization of surface phenotypic molecules of teleost dendritic cells. Dev Comp Immunol 49(1):38–43. https://doi.org/10.1016/j.dci.2014.11.010
Shen YB, Zhang JB, Xu XY, Fu JJ, Li JL (2012) Expression of complement component C7 and involvement in innate immune responses to bacteria in grass carp. Fish Shellfish Immun 33(2):448–454. https://doi.org/10.1016/j.fsi.2012.05.016
Shibasaki Y, Yabu T, Araki K, Mano N, Shiba H, Moritomo T, Nakanishi T (2014) Peculiar monomeric interferon gammas, IFN gamma rel 1 and IFN gamma rel 2, in ginbuna crucian carp. FEBS J 281(4):1046–1056. https://doi.org/10.1111/febs.12666
Sitja-Bobadilla A, Estensoro I, Perez-Sanchez J (2016) Immunity to gastrointestinal microparasites of fish. Dev Comp Immunol 64:187–201. https://doi.org/10.1016/j.dci.2016.01.014
Solbakken MH, Torresen OK, Nederbragt AJ, Seppola M, Gregers TF, Jakobsen KS, Jentoft S (2016) Evolutionary redesign of the Atlantic cod (Gadus morhua L.) Toll-like receptor repertoire by gene losses and expansions. Sci Rep-Uk 6:25211. https://doi.org/10.1038/srep25211
Stosik MP, Tokarz-Deptula B, Deptula W (2018) Specific humoral immunity in Osteichthyes. Cent Eur J Immunol 43(3):335–340. https://doi.org/10.5114/ceji.2018.80054
Stosik M, Tokarz-Deptula B, Deptula W (2019) Characterisation of thrombocytes in Osteichthyes. J Vet Res 63(1):123–131. https://doi.org/10.2478/jvetres-2019-0017
Sudhagar A, Ertl R, Kumar G, El-Matbouli M (2019) Transcriptome profiling of posterior kidney of brown trout, Salmo trutta, during proliferative kidney disease. Parasite Vector 12(1):569. https://doi.org/10.1186/s13071-019-3823-y
Sudhagar A, El-Matbouli M, Kumar G (2020) Identification and expression profiling of toll-like receptors of brown trout (Salmo trutta) during proliferative kidney disease. Int J Mol Sci 21(11):3755. https://doi.org/10.3390/ijms21113755
Sugamata R, Suetake H, Kikuchi K, Suzuki Y (2009) Teleost B7 expressed on monocytes regulates T cell responses. J Immunol 182(11):6799–6806. https://doi.org/10.4049/jimmunol.0803371
Suljevic D, Martinovic-Jukic A, Focak M, Alijagic A, Rukavina D, Zahirovic A (2017) Hematological importance of pseudoeosinophilic granulocytes in acclimation of common carp (Cyprinus carpio Linnaeus, 1758). Maced Vet Rev 40(1):5–11. https://doi.org/10.1515/macvetrev-2016-0091
Sun BJ, Robertsen B, Wang ZQ, Bin L (2009) Identification of an Atlantic salmon IFN multigene cluster encoding three IFN subtypes with very different expression properties. Dev Comp Immunol 33(4):547–558. https://doi.org/10.1016/j.dci.2008.10.001
Sun BJ, Skjaeveland I, Svingerud T, Zou J, Jorgensen J, Robertsen B (2011) Antiviral activity of salmonid gamma interferon against infectious pancreatic necrosis virus and salmonid alphavirus and its dependency on type I interferon. J Virol 85(17):9188–9198. https://doi.org/10.1128/Jvi.00319-11
Sun M, Mu YN, Ding Y, Ao JQ, Chen XH (2016a) Molecular and functional characterization of Toll-like receptor 21 in large yellow croaker (Larimichthys crocea). Fish Shellfish Immun 59:179–188. https://doi.org/10.1016/j.fsi.2016.10.024
Sun Y, Huang Y, Li XF, Baldwin CC, Zhou ZC, Yan ZX, Crandall KA, Zhang Y, Zhao XM, Wang M, Wong A, Fang C, Zhang XH, Huang H, Lopez JV, Kilfoyle K, Zhang Y, Orti G, Venkatesh B, Shi Q (2016b) Fish-T1K (Transcriptomes of 1,000 Fishes) Project: large-scale transcriptome data for fish evolution studies. Gigascience 5:18. https://doi.org/10.1186/s13742-016-0124-7
Sun QX, Fan ZJ, Yao CL (2018) Subcellular localization of large yellow croaker (Larimichthys crocea) TLR21 and expression profiling of its gene in immune response. J Ocean U China 17(2):335–343. https://doi.org/10.1007/s11802-018-3361-9
Sun BM, Lei Y, Cao ZJ, Zhou YC, Sun Y, Wu Y, Wang SF, Guo WL, Liu CS (2019) TroCCL4, a CC chemokine of Trachinotus ovatus, is involved in the antimicrobial immune response. Fish Shellfish Immun 86:525–535. https://doi.org/10.1016/j.fsi.2018.11.080
Sundaram AYM, Consuegra S, Kiron V, Fernandes JMO (2012a) Positive selection pressure within teleost toll-like receptors tlr21 and tlr22 subfamilies and their response to temperature stress and microbial components in zebrafish. Mol Biol Rep 39(9):8965–8975. https://doi.org/10.1007/s11033-012-1765-y
Sundaram AYM, Kiron V, Dopazo J, Fernandes JMO (2012b) Diversification of the expanded teleost-specific toll-like receptor family in Atlantic cod, Gadus morhua. BMC Evol Biol 12:256. https://doi.org/10.1186/1471-2148-12-256
Sveen L, Karlsen C, Ytteborg E (2020) Mechanical induced wounds in fish - a review on models and healing mechanisms. Rev Aquac. https://doi.org/10.1111/raq.12443
Svingerud T, Solstad T, Sun BJ, Nyrud MLJ, Kileng O, Greiner-Tollersrud L, Robertsen B (2012) Atlantic Salmon Type I IFN subtypes show differences in antiviral activity and cell-dependent expression: evidence for high IFNb/IFNc-producing cells in fish lymphoid tissues. J Immunol 189(12):5912–5923. https://doi.org/10.4049/jimmunol.1201188
Syahputra K, Kania PW, Al-Jubury A, Marnis H, Setyawan AC, Buchmann K (2019) Differential immune gene response in gills, skin, and spleen of rainbow trout Oncorhynchus mykiss infected by Ichthyophthirius multifiliis. PLoS One 14(6):e0218630
Tacchi L, Musharrafieh R, Larragoite ET, Crossey K, Erhardt EB, Martin SAM, LaPatra SE, Salinas I (2014) Nasal immunity is an ancient arm of the mucosal immune system of vertebrates. Nat Commun 5:5205. https://doi.org/10.1038/ncomms6205
Takano T, Kondo H, Hirono I, Endo M, Saito-Taki T, Aoki T (2007) Molecular cloning and characterization of Toll-like receptor 9 in Japanese flounder, Paralichthys olivaceus. Mol Immunol 44(8):1845–1853. https://doi.org/10.1016/j.molimm.2006.10.018
Tanekhy M, Kono T, Sakai M (2010) Cloning, characterization, and expression analysis of Toll-like receptor-7 cDNA from common carp, Cyprinus carpio L. Comp Biochem Phys D 5(4):245–255. https://doi.org/10.1016/j.cbd.2010.07.001
Tang LL, Xiang XY, Jiang YH, Lv YN, Zhou Y, Zhong H, Xiao J, Zhang FY, Jiang HY, Yan JP (2016) Identification and characterization of a novel Toll-like receptor 4 homologue in blunt snout bream, Megalobrama amblycephala. Fish Shellfish Immun 57:25–34. https://doi.org/10.1016/j.fsi.2016.08.015
Tang R, Wang SS, Han PP, Zhang QH, Zhang SH, Xing XP, Shao R, Xu W, Xu QQ, Wei QW, Qi ZT (2020) Toll-like receptor (TLR) 2 and TLR13 from the endangered primitive-ray finned fish Dabry’s sturgeon (Acipenser dabryanus) and their expression profiling upon immune stimulation. Aquacult Rep 16:100247. https://doi.org/10.1016/j.aqrep.2019.100247
Tavares-Dias M (2006) Cytochemical method for staining fish basophils. J Fish Biol 69(1):312–317. https://doi.org/10.1111/j.1095-8649.2006.01106.x
Tokunaga Y, Shirouzu M, Sugahara R, Yoshiura Y, Kiryu I, Ototake M, Nagasawa T, Somamoto T, Nakao M (2017) Comprehensive validation of T- and B-cell deficiency in rag1-null zebrafish: implication for the robust innate defense mechanisms of teleosts. Sci Rep-Uk 7(1):7536
Tong C, Lin YQ, Zhang CF, Shi JQ, Qi HF, Zhao K (2015) Transcriptome-wide identification, molecular evolution and expression analysis of Toll-like receptor family in a Tibet fish, Gymnocypris przewalskii. Fish Shellfish Immun 46(2):334–345. https://doi.org/10.1016/j.fsi.2015.06.023
Tribouley J, Tribouley-Duret J, Appriou M (1978) [Effect of Bacillus Calmette Guerin (BCG) on the receptivity of nude mice to Schistosoma mansoni] Influence du bacille de Calmette et Guerin (BCG) sur la receptivite de la Souris nude vis-a-vis de Schistosoma mansoni. C R Seances Soc Biol Fil 172(5):902–904
Tsuchida MA, Theriot JA (2013) An elastic actomyosin network in motile fish keratocytes. Biophys J 104(2):318a. https://doi.org/10.1016/j.bpj.2012.11.1763
Tu X, Liu L, Qi XZ, Chen WC, Wang GX, Ling F (2016) Characterization of Toll-like receptor gene expression in goldfish (Carassius auratus) during Dactylogyrus intermedius infection. Dev Comp Immunol 63:78–83. https://doi.org/10.1016/j.dci.2016.05.019
Turvey SE, Broide DH (2010) Innate immunity. J Allergy Clin Immun 125(2):S24–S32. https://doi.org/10.1016/j.jaci.2009.07.016
Uribe C, Folch H, Enriquez R, Moran G (2011) Innate and adaptive immunity in teleost fish: a review. Vet Med-Czech 56(10):486–503. https://doi.org/10.17221/3294-Vetmed
Valenzuela CA, Zuloaga R, Poblete-Morales M, Vera-Tobar T, Mercado L, Avendano-Herrera R, Valdes JA, Molina A (2017) Fish skeletal muscle tissue is an important focus of immune reactions during pathogen infection. Dev Comp Immunol 73:1–9. https://doi.org/10.1016/j.dci.2017.03.004
Valenzuela-Miranda D, Gallardo-Escarate C (2016) Novel insights into the response of Atlantic salmon (Salmo salar) to Piscirickettsia salmonis: Interplay of coding genes and lncRNAs during bacterial infection. Fish Shellfish Immun 59:427–438. https://doi.org/10.1016/j.fsi.2016.11.001
Valenzuela-Miranda D, Boltana S, Cabrejos ME, Yanez JM, Gallardo-Escarate C (2015) High-throughput transcriptome analysis of ISAV-infected Atlantic salmon Salmo salar unravels divergent immune responses associated to head-kidney, liver and gills tissues. Fish Shellfish Immun 45(2):367–377
Valenzuela-Munoz V, Valenzuela-Miranda D, Gallardo-Escarate C (2018) Comparative analysis of long non-coding RNAs in Atlantic and Coho salmon reveals divergent transcriptome responses associated with immunity and tissue repair during sea lice infestation. Dev Comp Immunol 87:36–50. https://doi.org/10.1016/j.dci.2018.05.016
Valenzuela-Munoz V, Pereiro P, Alvarez-Rodriguez M, Gallardo-Escarate C, Figueras A, Novoa B (2019) Comparative modulation of lncRNAs in wild-type and rag1-heterozygous mutant zebrafish exposed to immune challenge with spring viraemia of carp virus (SVCV). Sci Rep-Uk 9:14174. https://doi.org/10.1038/s41598-019-50766-0
Valero Y, Morcillo P, Meseguer J, Buonocore F, Esteban MA, Chaves-Pozo E, Cuesta A (2015) Characterization of the IFN pathway in the teleost fish gonad against vertically transmitted viral nervous necrosis virus. J Gen Virol 96:2176–2187. https://doi.org/10.1099/vir.0.000164
Valero Y, Saraiva-Fraga M, Costas B, Guardiola FA (2020) Antimicrobial peptides from fish: beyond the fight against pathogens. Rev Aquacult 12(1):224–253. https://doi.org/10.1111/raq.12314
Van AP, de Haro NA, Bron JE, Desbois AP (2020) Chromatin extracellular trap release in rainbow trout, Oncorhynchus mykiss (Walbaum, 1792). Fish Shellfish Immun 99:227–238. https://doi.org/10.1016/j.fsi.2020.01.040
Varela M, Diaz-Rosales P, Pereiro P, Forn-Cuni G, Costa MM, Dios S, Romero A, Figueras A, Novoa B (2014) Interferon-induced genes of the expanded IFIT family show conserved antiviral activities in non-mammalian species. PLoS One 9(6):e100015. https://doi.org/10.1371/journal.pone.0100015
Vidal R, Gonzalez R, Gil F (2015) Characterization and expression analysis of Toll-like receptor 3 cDNA from Atlantic salmon (Salmo salar). Genet Mol Res 14(2):6073–6083. https://doi.org/10.4238/2015.June.8.5
Wan Q, Wicramaarachchi WDN, Whang I, Lim BS, Oh MJ, Jung SJ, Kim HC, Yeo SY, Lee J (2012) Molecular cloning and functional characterization of two duplicated two-cysteine containing type I interferon genes in rock bream Oplegnathus fasciatus. Fish Shellfish Immun 33(4):886–898. https://doi.org/10.1016/j.fsi.2012.07.018
Wang ZP, Zhang SC, Tong Z, Li L, Wang GF (2009) Maternal transfer and protective role of the alternative complement components in Zebrafish Danio rerio. PLoS One 4(2):e4498. https://doi.org/10.1371/journal.pone.0004498
Wang KR, Mu YN, Qian TL, Ao JQ, Chen XH (2013a) Molecular characterization and expression analysis of Toll-like receptor 1 from large yellow croaker (Pseudosciaena crocea). Fish Shellfish Immun 35(6):2046–2050. https://doi.org/10.1016/j.fsi.2013.10.022
Wang WJ, Shen YB, Pandit NP, Li JL (2013b) Molecular cloning, characterization and immunological response analysis of Toll-like receptor 21 (TLR21) gene in grass carp, Ctenopharyngodon idella. Dev Comp Immunol 40(3–4):227–231. https://doi.org/10.1016/j.dci.2013.03.003
Wang JL, Zhang Z, Liu J, Li F, Chang F, Fu H, Zhao J, Yin DL (2015) Structural characterization and evolutionary analysis of fish-specific TLR27. Fish Shellfish Immun 45(2):940–945. https://doi.org/10.1016/j.fsi.2015.06.017
Wang YJ, Bi XY, Chu Q, Xu TJ (2016a) Discovery of toll-like receptor 13 exists in the teleost fish: Miiuy croaker (Perciformes, Sciaenidae). Dev Comp Immunol 61:25–33. https://doi.org/10.1016/j.dci.2016.03.005
Wang YJ, Li JR, Han JJ, Shu C, Xu TJ (2016b) Identification and characteristic analysis of TLR28: a novel member of the TLR1 family in teleost. Dev Comp Immunol 62:102–107. https://doi.org/10.1016/j.dci.2016.05.001
Wang M, Yu F, Wu W, Zhang Y, Chang WG, Ponnusamy M, Wang K, Li PF (2017) Circular RNAs: a novel type of non-coding RNA and their potential implications in antiviral immunity. Int J Biol Sci 13(12):1497–1506. https://doi.org/10.7150/ijbs.22531
Wang M, Jiang S, Wu W, Yu F, Chang WG, Li PF, Wang K (2018a) Non-coding RNAs function as immune regulators in teleost fish. Front Immunol 9:2801. https://doi.org/10.3389/fimmu.2018.02801
Wang PF, Zhao C, Wang CY, Fan SG, Yan LL, Qiu LH (2018b) TLR3 gene in Japanese sea perch (Lateolabrax japonicus): molecular cloning, characterization and expression analysis after bacterial infection. Fish Shellfish Immun 76:347–354. https://doi.org/10.1016/j.fsi.2018.01.013
Watanabe T, Kitayama K, Takagi T, Murata J, Kono M, Takase T, Furukawa K (1997) Heterogeneity of peritoneal cells in marine teleosts. Fish Sci 63(4):576–581. https://doi.org/10.2331/fishsci.63.576
Watson LJ, Shechmeister IL, Jackson LL (1963) The hematology of goldfish, Carassius auratus. Cytologia 28:118–130
Wei YC, Pan TS, Chang MX, Huang B, Xu Z, Luo TR, Nie P (2011) Cloning and expression of Toll-like receptors 1 and 2 from a teleost fish, the orange-spotted grouper Epinephelus coioides. Vet Immunol Immunop 141(3–4):173–182. https://doi.org/10.1016/j.vetimm.2011.02.016
Wei YC, Hu S, Sun BB, Zhang QH, Qiao G, Wang ZS, Shao R, Huang GQ, Qi ZT (2017) Molecular cloning and expression analysis of toll-like receptor genes (TLR7, TLR8 and TLR9) of golden pompano (Trachinotus ovatus). Fish Shellfish Immun 63:270–276. https://doi.org/10.1016/j.fsi.2017.02.026
Wentzel AS, Janssen JJE, de Boer VCJ, van Veen WG, Forlenza M, Wiegertjes GF (2020) Fish macrophages show distinct metabolic signatures upon polarization. Front Immunol 11:152. https://doi.org/10.3389/fimmu.2020.00152
Wessel O, Olsen CM, Rimstad E, Dahle MK (2015) Piscine orthoreovirus (PRV) replicates in Atlantic salmon (Salmo salar L.) erythrocytes ex vivo. Vet Res 46:26
Wickramaarachchi WDN, Wan Q, Lee Y, Lim BS, De Zoysa M, Oh MJ, Jung SJ, Kim HC, Whang I, Lee J (2012) Genomic characterization and expression analysis of complement component 9 in rock bream (Oplegnathus fasciatus). Fish Shellfish Immun 33(4):707–717. https://doi.org/10.1016/j.fsi.2012.06.019
Wickramaarachchi WDN, Whang I, Wan Q, Bathige SDNK, De Zoysa M, Lim BS, Yeo SY, Park MA, Lee J (2013) Genomic characterization and expression analysis of complement component 8 alpha and 8 beta in rock bream (Oplegnathus fasciatus). Dev Comp Immunol 39(3):279–292. https://doi.org/10.1016/j.dci.2012.09.005
Wiegertjes GF, Wentzel AS, Spaink HP, Elks PM, Fink IR (2016) Polarization of immune responses in fish: the ‘macrophages first’ point of view. Mol Immunol 69:146–156. https://doi.org/10.1016/j.molimm.2015.09.026
Wiens GD, Glenney GW (2011) Origin and evolution of TNF and TNF receptor superfamilies. Dev Comp Immunol 35(12):1324–1335. https://doi.org/10.1016/j.dci.2011.03.031
Wittamer V, Bertrand JY, Gutschow PW, Traver D (2011) Characterization of the mononuclear phagocyte system in zebrafish. Blood 117(26):7126–7135. https://doi.org/10.1182/blood-2010-11-321448
Worbs T, Hammerschmidt SI, Foerster R (2017) Dendritic cell migration in health and disease. Nat Rev Immunol 17(1):30–48. https://doi.org/10.1038/nri.2016.116
Workenhe ST, Rise ML, Kibenge MJT, Kibenge FSB (2010) The fight between the teleost fish immune response and aquatic viruses. Mol Immunol 47(16):2525–2536. https://doi.org/10.1016/j.molimm.2010.06.009
Wu N, LaPatra SE, Li J, Sunyer JO, Zhang YA (2014) Complement C5a acts as molecular adjuvant in fish by enhancing antibody response to soluble antigen. Fish Shellfish Immun 40(2):616–623. https://doi.org/10.1016/j.fsi.2014.08.013
Wu N, Song YL, Wang B, Zhang XY, Zhang XJ, Wang YL, Cheng YY, Chen DD, Xia XQ, Lu YS, Zhang YA (2016) Fish gut-liver immunity during homeostasis or inflammation revealed by integrative transcriptome and proteome studies. Sci Rep-Uk 6:36048. https://doi.org/10.1038/srep36048
Wu M, Guo L, Zhu KC, Guo HY, Liu B, Jiang SG, Zhang DC (2018) Genomic structure and molecular characterization of Toll-like receptors 1 and 2 from golden pompano Trachinotus ovatus (Linnaeus, 1758) and their expression response to three types of pathogen-associated molecular patterns. Dev Comp Immunol 86:34–40. https://doi.org/10.1016/j.dci.2018.04.022
Wu M, Guo L, Zhu KC, Guo HY, Liu BS, Zhang N, Jiang SG, Zhang DC (2019) Molecular characterization of toll-like receptor 14 from golden pompano Trachinotus ovatus (Linnaeus, 1758) and its expression response to three types of pathogen-associated molecular patterns. Comp Biochem Phys B 232:1–10. https://doi.org/10.1016/j.cbpb.2019.02.010
Wu LT, Qin ZD, Liu HP, Lin L, Ye JM, Li J (2020) Recent advances on phagocytic B cells in teleost fish. Front Immunol 11:824. https://doi.org/10.3389/fimmu.2020.00824
Xiao XQ, Qin QW, Chen XH (2011) Molecular characterization of a Toll-like receptor 22 homologue in large yellow croaker (Pseudosciaena crocea) and promoter activity analysis of its 5'-flanking sequence. Fish Shellfish Immun 30(1):224–233. https://doi.org/10.1016/j.fsi.2010.10.014
Xiao X, Zhang YQ, Liao ZW, Su JG (2020) Characterization and antimicrobial activity of the teleost chemokine CXCL20b. Antibiotics-Basel 9(2):78. https://doi.org/10.3390/antibiotics9020078
Xie FJ, Zhang ZP, Lin P, Wang SH, Zou ZH, Wang YL (2008) Cloning and infection response of tumour-necrosis factor alpha in large yellow croaker Pseudosciaena crocea (Richardson). J Fish Biol 73(5):1149–1160. https://doi.org/10.1111/j.1095-8649.2008.01945.x
Xiong Y, Dan C, Ren F, Su ZH, Zhang YB, Mei J (2020) Proteomic profiling of yellow catfish (Pelteobagrus fulvidraco) skin mucus identifies differentially-expressed proteins in response to Edwardsiella ictaluri infection. Fish Shellfish Immun 100:98–108. https://doi.org/10.1016/j.fsi.2020.02.059
Xiu YJ, Jiang GP, Zhou S, Diao J, Liu HJ, Su BF, Li C (2019) Identification of potential immune-related circRNA-miRNA-mRNA regulatory network in intestine of paralichthys olivaceus during edwardsiella tarda infection. Front Genet 10:731. https://doi.org/10.3389/fgene.2019.00731
Xu TJ, Meng FX, Zhu ZH, Wang RX (2013) Characterization and comprehensive analysis of the miiuy croaker TLR2 reveals a direct evidence for intron insert and loss. Fish Shellfish Immun 34(1):119–128. https://doi.org/10.1016/j.fsi.2012.10.008
Xu TJ, Wang YJ, Li JR, Shu C, Han JJ, Chu Q (2016) Comparative genomic evidence for duplication of TLR1 subfamily and miiuy croaker TLR1 perceives LPS stimulation via MyD88 and TIRAP. Fish Shellfish Immun 56:336–348. https://doi.org/10.1016/j.fsi.2016.07.024
Xu TJ, Chu Q, Cui JX, Bi DK (2018a) Inducible MicroRNA-3570 feedback inhibits the RIG-I-dependent innate immune response to rhabdovirus in teleost fish by targeting MAVS/IPS-1. J Virol 92(2):e01594-17. https://doi.org/10.1128/JVI.01594-17
Xu YS, Yu YY, Zhang XT, Huang ZY, Li HL, Dong S, Liu YZ, Dong F, Xu Z (2018b) Molecular characterization and expression analysis of complement component 3 in dojo loach (Misgurnus anguillicaudatus). Fish Shellfish Immun 72:484–493. https://doi.org/10.1016/j.fsi.2017.11.022
Xu HY, Dong F, Zhai X, Meng KF, Han GK, Cheng GF, Wu ZB, Li N, Xu Z (2020) Mediation of mucosal immunoglobulins in buccal cavity of teleost in antibacterial immunity. Front Immunol 11:562795. https://doi.org/10.3389/fimmu.2020.562795
Yang C, Su J (2010) Molecular identification and expression analysis of Toll-like receptor 3 in common carp Cyprinus carpio. J Fish Biol 76(8):1926–1939. https://doi.org/10.1111/j.1095-8649.2010.02624.x
Yang C, Su J, Zhang R, Peng L, Li Q (2012) Identification and expression profiles of grass carp Ctenopharyngodon idella TLR7 in responses to double-stranded RNA and virus infection. J Fish Biol 80(7):2605–2622. https://doi.org/10.1111/j.1095-8649.2012.03316.x
Yang Y, Yu H, Li H, Wang AL (2016) Transcriptome profiling of grass carp (Ctenopharyngodon idellus) infected with Aeromonas hydrophila. Fish Shellfish Immun 51:329–336. https://doi.org/10.1016/j.fsi.2016.02.035
Yang N, Wang BB, Yu ZX, Liu XL, Fu Q, Cao M, Xue T, Ren YC, Tan FH, Li C (2020) Characterization of a novel lncRNA (SETD3-OT) in turbot (Scophthalmus maximus L.). Fish Shellfish Immun 102:145–151. https://doi.org/10.1016/j.fsi.2020.04.010
Yao CL, Kong P, Wang ZY, Ji PF, Cai MY, Liu XD, Han XZ (2008) Cloning and expression analysis of two alternative splicing toll-like receptor 9 isoforms A and B in large yellow croaker, Pseudosciaena crocea. Fish Shellfish Immun 25(5):648–656. https://doi.org/10.1016/j.fsi.2008.07.006
You XX, Bian C, Zan QJ, Xu X, Liu X, Chen JM, Wang JT, Qiu Y, Li WJ, Zhang XH, Sun Y, Chen SX, Hong WS, Li YX, Cheng SF, Fan GY, Shi CC, Liang J, Tang YT, Yang CY, Ruan ZQ, Bai J, Peng C, Mu Q, Lu J, Fan MJ, Yang S, Huang ZY, Jiang XT, Fang XD, Zhang GJ, Zhang Y, Polgar G, Yu H, Li J, Liu ZJ, Zhang GQ, Ravi V, Coon SL, Wang J, Yang HM, Venkatesh B, Wang J, Shi Q (2014) Mudskipper genomes provide insights into the terrestrial adaptation of amphibious fishes. Nat Commun 5:5594. https://doi.org/10.1038/ncomms6594
Yu YY, Kong WG, Yin YX, Dong F, Huang ZY, Yin GM, Dong S, Salinas I, Zhang YA, Xu Z (2018) Mucosal immunoglobulins protect the olfactory organ of teleost fish against parasitic infection. PLoS Pathog 14(11):e1007251. https://doi.org/10.1371/journal.ppat.1007251
Yu L, Li CH, Chen J (2019) A novel CC chemokine ligand 2 like gene from ayu Plecoglossus altivelis is involved in the innate immune response against to Vibrio anguillarum. Fish Shellfish Immun 87:886–896. https://doi.org/10.1016/j.fsi.2019.02.019
Zhan FB, Tan KA, Song XR, Yu JY, Wang WM (2019) Isolation and expression of four Megalobrama amblycephala toll-like receptor genes in response to a bacterial infection. Fish Shellfish Immun 93:1028–1040. https://doi.org/10.1016/j.fsi.2019.08.051
Zhang Q, Cao XT (2019) Epigenetic regulation of the innate immune response to infection. Nat Rev Immunol 19(7):417–432. https://doi.org/10.1038/s41577-019-0151-6
Zhang LJ, Gallo RL (2016) Antimicrobial peptides. Curr Biol 26(1):R14–R19. https://doi.org/10.1016/j.cub.2015.11.017
Zhang YB, Gui JF (2012) Molecular regulation of interferon antiviral response in fish. Dev Comp Immunol 38(2):193–202. https://doi.org/10.1016/j.dci.2012.06.003
Zhang AY, Chen DY, Wei H, Du LY, Zhao TQ, Wang XY, Zhou H (2012) Functional characterization of TNF-alpha in grass carp head kidney leukocytes: induction and involvement in the regulation of NF-kappa B signaling. Fish Shellfish Immun 33(5):1123–1132. https://doi.org/10.1016/j.fsi.2012.08.029
Zhang JR, Liu SK, Rajendran KV, Sun LY, Zhang Y, Sun FY, Kucuktas H, Liu H, Liu ZJ (2013a) Pathogen recognition receptors in channel catfish: III phylogeny and expression analysis of Toll-like receptors. Dev Comp Immunol 40(2):185–194. https://doi.org/10.1016/j.dci.2013.01.009
Zhang SC, Wang ZP, Wang HM (2013b) Maternal immunity in fish. Dev Comp Immunol 39(1–2):72–78. https://doi.org/10.1016/j.dci.2012.02.009
Zhang HY, Hu GB, Liu QM, Zhang SC (2016) Cloning and expression study of a Toll-like receptor 2 (tlr2) gene from turbot, Scophthalmus maximus. Fish Shellfish Immun 59:137–148. https://doi.org/10.1016/j.fsi.2016.10.001
Zhang J, Wang L, Zhao YJ, Kong XH, Wu F, Zhao XL (2017a) Molecular characterization and expression analysis of toll-like receptors 5 and 22 from natural triploid Carassius auratus. Fish Shellfish Immun 64:1–13. https://doi.org/10.1016/j.fsi.2017.03.004
Zhang XT, Zhang GR, Shi ZC, Yuan YJ, Zheng H, Lin L, Wei KJ, Ji W (2017b) Expression analysis of nine Toll-like receptors in yellow catfish (Pelteobagrus fulvidraco) responding to Aeromonas hydrophila challenge. Fish Shellfish Immun 63:384–393. https://doi.org/10.1016/j.fsi.2017.02.021
Zhang ZB, Chi H, Dalmo RA (2019) Trained innate immunity of fish is a viable approach in larval aquaculture. Front Immunol 10:42. https://doi.org/10.3389/fimmu.2019.00042
Zhao ML, Chi H, Sun L (2017) Neutrophil extracellular traps of cynoglossus semilaevis: production characteristics and antibacterial effect. Front Immunol 8:290. https://doi.org/10.3389/fimmu.2017.00290
Zhou ZX, Lin ZJ, Pang X, Shan PP, Wang JX (2018) MicroRNA regulation of Toll-like receptor signaling pathways in teleost fish. Fish Shellfish Immun 75:32–40. https://doi.org/10.1016/j.fsi.2018.01.036
Zhou T, Gui L, Liu ML, Li WH, Hu P, Duarte DFC, Niu HB, Chen LB (2019) Transcriptomic responses to low temperature stress in the Nile tilapia, Oreochromis niloticus. Fish Shellfish Immun 84:1145–1156. https://doi.org/10.1016/j.fsi.2018.10.023
Zhu KC, Wu M, Zhang DC, Guo HY, Zhang N, Guo L, Liu BS, Jiang SG (2020) Toll-like receptor 5 of golden pompano trachinotus ovatus (Linnaeus 1758): characterization, promoter activity and functional analysis. Int J Mol Sci 21(16):5916. https://doi.org/10.3390/ijms21165916
Zou J, Secombes CJ (2011) Teleost fish interferons and their role in immunity. Dev Comp Immunol 35(12):1376–1387. https://doi.org/10.1016/j.dci.2011.07.001
Acknowledgements
The authors acknowledge UiT—the Arctic University of Tromsø, the Research Council of Norway (grant 301401). Emma Vogel, Filipe Figueiredo, and Prof. Kim Præbel (UiT, genetic research group) are acknowledged for valuable comments and edits.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this chapter
Cite this chapter
Dalmo, R.A., Bøgwald, J. (2022). Innate Immunity. In: Buchmann, K., Secombes, C.J. (eds) Principles of Fish Immunology . Springer, Cham. https://doi.org/10.1007/978-3-030-85420-1_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-85420-1_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-85419-5
Online ISBN: 978-3-030-85420-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)