Abstract
By using gene editing technologies such as CRISPR/Cas, precise modifications can be made in the genome. CRISPR/Cas is especially valuable for targeted mutagenesis in polyploids, as it can induce mutations of multiple alleles simultaneously, to obtain regenerants that are homozygous for the desired mutation. A range of gene-edited traits have been developed in hexaploid bread wheat, including various nutrition and health-related traits, plant architecture, pest and disease resistance, tolerance to abiotic stress, and traits that enable more efficient breeding. Wheat is also known as a cause of some human diseases, particularly coeliac disease (CD), with a prevalence of 1–2% of the population. In the EU alone, at least 4.5 million people suffer from it. CD is a chronic inflammation of the small intestine, induced and maintained in genetically predisposed individuals by the consumption of gluten proteins from wheat, barley and rye. As there is no cure, patients must follow a life-long gluten-free diet. The dominant epitopes in gluten proteins that trigger the disease, have been characterized, but they cannot be removed by classical breeding without affecting baking quality, as it concerns over 100 gluten genes that occur partly as blocks of genes in the genome of wheat. Using gene editing, two studies have shown that it is possible to modify the epitopes in several alpha- and gamma-gliadins simultaneously, while deleting some of the genes completely. In some lines more than 80% of the alpha-gliadin genes were modified. These proof-of-principle studies show that it is feasible to use gene editing, along with other breeding approaches, to completely remove the CD epitopes from bread wheat. Gene-edited coeliac-safe wheat will have economic, social and environmental impact on food security, nutrition and public health, but the realisation will (partially) depend on new European legislation for plants produced by gene editing.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
1 Introduction
Bread wheat (Triticum aestivum, a hexaploid wheat species with the AABBDD genome) originated as a hybridization between tetraploid Triticum and (the) diploid Aegilops species during early agriculture in the Fertile Crescent. Triticum and Aegilops are closely related: Triticum is considered the domesticated form of Aegilops. Interspecific hybrids are found in nature and may also be produced artificially. Long before the onset of agriculture, a diploid wheat (T. urartu) carrying an AA genome hybridized with a grass-like wheat with a BB genome into the tetraploid T. turgidum with the combined AABB genome constitution. Near the Caspian Sea area, in an ancient agricultural field, a cultivated free-threshing tetraploid wheat (AABB) hybridized with a local wild diploid Aegilops tauschii with its DD genome, which led to the new hexaploid species T. aestivum (AABBDD). Spelt wheat is considered the result of one of the most recent natural hybridizations in which T. aestivum and a tetraploid T. turgidum (an emmer wheat type) were involved. Presently, synthetic hexaploid wheat lines are under production through various artificial hybridization programs, to increase the diversity in the D genome in hexaploid germplasm [1,2,3].
Wheat is a self-pollinating crop. Farmers can cultivate it by sowing seed material saved from the preceding year. Initially, cultivation practice applied mixtures of tetraploid and hexaploid genotypes. Through conscious or unconscious selection of spontaneous mutants, landraces gradually adapted to local environmental conditions, but there is a limit to the improvement of end-use quality that can be achieved in that way. Since the early twentieth century, genetics has been used in professional wheat breeding, including pure line selection and targeted breeding. Breeders are always interested in new genetic variation. This can be achieved through introgression (trait transfer through hybridization followed by back-crossing) from other wheat species, but this process also introduces many undesired traits that subsequently must be selected against. Alternatively, genetic variation can be induced within a cultivar through the application of mutagenic chemicals or ionizing irradiation (mutation breeding).
Presently, next to the hexaploid bread wheat and spelt wheat also diploid einkorn and various tetraploid T. turgidum subspecies (including emmer and durum wheat) are still being cultivated. However, almost 95% of the world’s annual volume of 700–750 million metric tons is bread wheat; durum wheat makes up only 5%, the other species have a minimal volume. Bread wheat and durum wheat are also referred to as soft wheat and hard wheat, respectively, due to the difference in hardness of the kernel and further technological qualities regarding milling and baking: bread, cakes and breakfast cereals are made from soft wheat, pasta from durum wheat.
Its three homoeologous genomes make bread wheat flexible and adaptive in cultivation and versatile in its applications, due to its high genetic and allelic redundancy. This versatility becomes apparent in the specific qualitative and quantitative protein composition of its water-insoluble gluten, as used in various food and non-food processing applications, and, together with the grain’s starch, in dough and derived baked foods. Remarkable is the interdependence of the gluten protein structure, the gluten quantity and the gluten applicability regarding genetic, environmental and processing factors [4,5,6].
In keeping with its wide cultivation and application, bread wheat is the species of choice for further improvement of agricultural and food technological traits through conventional breeding, using various breeding tools developed in the twentieth and twenty-first century, including marker-assisted breeding and speed breeding [7].
In wheat breeding, the main focus was traditionally directed to yield and quality trait improvements. This has culminated in the 1960s in the Green Revolution through the introduction of dwarf genes. The dwarf genes (actually, dwarfing mutations) reduce energy investment in stem growth and increase grain yield. These new varieties were highly recognized worldwide. Today’s breeding aims still include yield (especially starch quantity) and gluten and starch quality (for improved milling and baking quality), but adaptations to the biotic and abiotic environment, such as disease resistance genes, are receiving more attention because of the spread of major diseases and the threats of climate change, e.g., increased drought.
Impacts on export markets appeared to have played a role in the lack of commercialized GM wheat. The first GM (herbicide-tolerant) wheat applied for approval in the USA was abandoned in 2004 [8]. HB4 wheat, a GM wheat variety with drought tolerance has been approved for cultivation in Argentina in 2020 and subsequently for import in Brazil in 2021, which was important as for commercialization of GM events Argentina takes the impact on export markets into account. Since then, import has also been approved in the USA, Australia, New Zealand, Colombia, and Nigeria. Recently, Brazil has also approved cultivation of HB4 wheat.
2 Gene Editing in Polyploid Crops
To facilitate the time-consuming breeding of polyploid crops, alternative methods for introgression breeding and mutation breeding are important for polyploid crop improvement. Genome editing is such an alternative method. By using gene editing technologies such as CRISPR/Cas, precise modifications can be made to specific genes in the genome. CRISPR/Cas is especially valuable for targeted mutagenesis in polyploids, as it can induce mutations of multiple alleles simultaneously, so that regenerants homozygous for the desired mutation may be obtained. Moreover, the intrinsic specificity of CRISPR/Cas allows for the precise targeting of specific homoeologous genes. This enables allele-specific modifications, where only one copy of a gene is edited while leaving the others unchanged, or where all homoeologous copies are modified simultaneously. This ultimately allows for the introduction of desirable traits or the removal of undesirable ones without the need for laborious backcrossing to eliminate unwanted genetic material. Several studies have demonstrated the effectiveness and robustness of gene editing for targeted mutagenesis in auto- and allopolyploids [9], including wheat ([10]; see below and Table 13.1).
3 Improving Regeneration of Wheat in Tissue Culture
Efficient protocols for plant transformation and regeneration in tissue culture are essential prerequisites for utilizing gene editing techniques in wheat. However, the process of regeneration in tissue culture presents difficulties, particularly when dealing with recalcitrant species and is often dependent on identifying genotypes with capacity to regenerate. Overcoming these hurdles is crucial for wheat tissue culture as well. Promising advancements have been made in this area, such as the utilization of the developmental regulators that stimulate somatic embryogenesis and de novo meristem formation. Notably, recent studies have demonstrated that overexpressing a GRF4-GIF1 chimera construct and the TaWOX5 gene can substantially increase the regeneration capacity of wheat tissue cultures, while reducing the dependence on specific genotypes [11, 12]. These findings open new avenues for improving the regeneration efficiency of wheat in tissue culture and pave the way for more accessible and genotype-independent regeneration protocols. Continued research and exploration of innovative strategies, along with the advancement of gene editing techniques, can contribute to further enhancing the regeneration potential of wheat and accelerating the development of gene-edited varieties.
4 Current Research and Trials Using Gene Edited Traits in Wheat
A range of gene edited traits have been developed in wheat, largely in the last 5 years (Table 13.1). These include various nutrition and health-related traits such as low asparagine content, high fibre content, reduced coeliac disease epitopes in gluten, and biofortification. Next to these, plant architecture, yield improvement, pest and disease resistance, tolerance to abiotic stress, and herbicide tolerance are also investigated. The studies used Cas9 and Cpf1 [13,14,15,16] nucleases or prime editing [17] for targeted mutagenesis, as well as some base editors (see Table 13.1). Luo et al. [18] worked towards homology-directed repair enabling the introduction of larger sequences such as complete genes or promoters.
A series of breeding-related traits are also subject of gene editing research, including recombination, crossing over, haploid induction, and male sterility. These are related on one hand to the need to introgress traits, such as disease resistances and quality traits, from wild relatives, on the other hand to the desire to develop hybrid wheat varieties. When compared to traditionally inbred varieties, hybrid wheat is estimated to have a potential yield increase of 10% [19]. Furthermore, hybrid wheat, as with other hybrid crops, offers enhanced resilience and provides greater yield stability, even in the presence of environmental stresses and the extreme impacts of climate change. However, the breeding and widespread adoption of hybrid wheat is lagging behind. One reason is a costly production of hybrid seeds due to wheat’s inherent tendency for self-pollination and the production of only one seed per flower [19]. Overcoming these barriers and establishing effective breeding techniques for large-scale hybrid wheat breeding requires substantial research and development efforts. For example, male sterility provides the method to block self-fertilization, while modifying flower architecture can facilitate pollination.
Below we will in detail describe why and how gene editing and other approaches that target wheat gluten are used to reduce or remove epitopes that cause coeliac disease in susceptible people.
5 Wheat Gluten
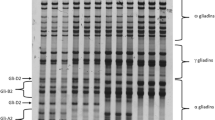
Flour obtained by milling of whole wheat grains can be separated into four major protein fractions: albumins plus globulins, gliadins, glutenins, and a group of residual proteins. The gliadins and the glutenins, both water-insoluble, can be further subdivided into subtypes (Table 13.2). Cysteine residues, although relatively low in number, are characteristic to most gluten molecules (except to the omega-gliadins) as they form intrachain bonds in the alpha- and gamma-gliadins, stabilizing their tertiary structure, and interchain bonds among LMW and HMW glutenins. Gluten polymerization, a unique characteristic of gluten from wheat, is crucial for dough-making and baking. The building of large gas-retaining gluten networks may even increase through opening of intrachain gliadin S-S bonds by heating, enabling their binding to glutenin polymers [5]. High hydrostatic pressure (extrusion) of isolated guten to increase further polymerization can be applied in the production of meat analogues [67].
Wheat flour plus water produces a viscoelastic mass (a starchy dough) in which the gliadins are especially responsible for viscosity, whereas the glutenins are responsible for elasticity of the network that retains the gas from the yeast or sourdough fermentation and makes the dough rise. The degree of viscosity relates to the ratio of gliadins to glutenins: a high ratio (around 3.0) results in a soft and less viscous dough; a low ratio (around 2.0) results in a highly viscous and strong dough [5]. Heating during baking makes the gluten rigid by the loss of its water to the starch, keeping the baked bread shape stable. The presence of glutenin in doughs is essential for baking of bread and cookies, and in batter for cakes. Gliadins may be exchanged for other proteins or even partly omitted without significant loss of baking quality [68]. Guzmán-López et al. [69] managed to bake bread from wheat lines in which most gliadins had been silenced by RNAi.
Removing the starch from unheated dough by repeated washings with water yields in a purified gluten mass, called vital wheat gluten (VWG), which is applied in the food (bakery) and non-food industry.
6 Health-Related Wheat Components in Food
Due to its food technological versatility, wheat and its isolated components (vital wheat gluten, wheat starch, and derivatives such as glucose syrups, maltodextrin, sorbitol) are applied in numerous food products. They appear in about 30% of the labelled and mostly highly processed food items [71]. Highly processed foods, including those with wheat-derived ingredients, are often classified as unhealthy as they often contain a too high content of non-wheat ingredients such as sugar, salt and fat, and a too low content of fibre. However, whole grain wheat foods are recognized as healthy for their proven contribution to the reduction of risk of several ‘western lifestyle’-related chronic diseases including obesity and diabetes, heart and vascular diseases, immune-related diseases, and certain forms of cancer. Therefore, consumption of whole grain (wheat) foods is stimulated by governmental food authorities in many countries [72, 73].
7 Wheat-Related Human Diseases
Wheat is also known as a cause of some human diseases: wheat allergy with a prevalence of 0.25% is relatively rare; non-coeliac wheat sensitivity (NCWS) with a self-reported prevalence of 10% and a clinically estimated prevalence of 1% has mostly mild symptoms; and coeliac disease (CD), the most severe but also the best-known and well-documented disease, has a prevalence of 1–2% [74, 75]. CD is a chronic inflammation of the small intestine, induced and maintained in genetically predisposed individuals by the consumption of gluten proteins from wheat, barley and rye. As there is no cure, patients must follow a life-long gluten-free diet. In the EU alone, at least 4.5 million people suffer from CD.
The general daily intake of gluten from the consumption of bread and related foods is about 15 g [76], whereas the safe daily amount of gluten in foods for individuals with CD is estimated between 10 and 100 mg gluten [77]. To stay below that load, food products should not surmount the 20 ppm threshold as is defined according to extensive research and established in EU regulation EC828/2014. Note the huge gap between current foods in a gluten containing and a gluten-free lifestyle, especially when considering the high number of food items in supermarkets and elsewhere processed with wheat and wheat-derived components. As a correct diagnosis of CD is difficult because of the great variety of occurring symptoms, it is estimated that 85% of the patients remain undiagnosed, which means that they daily consume gluten, unaware that their bad chronic health status is caused by this food ingredient.
8 Gluten-Free Lifestyle
During the last decades, a gradual but significant increase has been observed in the number of consumers that embrace a gluten-free or wheat-free diet. This gluten/wheat avoiding population varies globally according to geographic location between 3.7% and 17.2% [78]. The main incentive for this diet choice is the desire for better health. This trend creates a decreasing bread and wheat-product market with a negative impact on the whole wheat value chain on the one hand, and an increasing and diversifying gluten-free product market on the other hand, with an annual growth rate of 10.4% between 2014 and 2019 in Europe, and an estimated compound annual growth rate (CAGR) of 8,1%. It had a market value of 8.3 Billion US dollars in 2025 (https://www.statista.com/statistics/248467/global-gluten-free-food-market-size/). The global gluten-free packaged food market now represents about 3% of the total global packaged food market, reflecting the population share that is interested in gluten-free foods, which is 15–20 times higher than the diagnosed CD population.
The problem remains to the 85% undiagnosed CD population. This group would be really served with the development, global cultivation and processing of safe-gluten wheat varieties replacing current high CD-immunogenic wheat. Gluten safety of such varieties should be guaranteed quantitatively (elimination of all immunogenic gluten) or qualitatively (inactivation of CD-immunogenic fragments (epitopes) from individual gluten proteins). However, maintenance of the full food technological and health qualities combined with unimpaired agronomic quality in such varieties remains a prime prerequisite for a crop to be competitive on the (global) market.
9 Gluten Epitopes for Coeliac Disease
What makes gluten intolerable to certain genetically predisposed individuals developing CD? The high abundance of glutamine and proline amino acids in gluten proteins hinders degradation by the proteolytic enzymes in the digestive tract, leaving relatively long peptides in the small intestine. Several of these peptides contain a core sequence of nine amino acids that can be recognized by antigen-presenting cells and presented to T cells. Upon recognition, these T cells become activated, resulting in inflammation of the mucosa followed by degeneration of the mucosal villi with serious consequences for uptake of nutrients and leading to a variety of symptoms. Several epitopes are recognized by T cells from many patients, called ‘major’ epitopes. A comprehensive list of epitopes was recently updated [79].
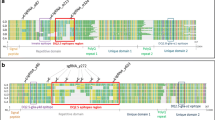
Gluten proteins are encoded by large gene families located at different sites on the wheat genome (Table 13.2) with the alpha-gliadin loci on the short arm of the homoeologous chromosomes 6A, 6B and 6D, and the gamma-gliadin genes on the short arm of 1A, 1B and 1D. A single bread wheat variety genome (e.g., from the variety Chinese Spring for which the genome has been sequenced [80]), may contain as many as ~100 different gluten genes of which about sixty are expressed into proteins, as was shown by mass spectrometry analysis [4, 81,82,83]. Comparing different wheat varieties, variation exists in the number of expressed gluten genes, in the sequences of the encoded proteins, and in the amount of protein produced per gene. Additional variation in the gluten composition (quantitatively and qualitatively) is induced by environmental factors in the field during the growing season, such as the temperature during certain stages of crop development, and nutrient availability from the soil, in particular nitrogen and sulphur [4].
Although both gliadins and glutenins harbour immunogenic epitopes that can trigger CD, the alpha-, gamma- and omega-gliadins contain by far the highest number and the most dominant (severe) epitopes. For alpha-gliadins the dominant epitopes are related to the genes from the D-genome, followed by the genes from the A-genome, whereas some genes from the B-genome are free from epitopes [84]. D-genome alpha-gliadins contain the so-called 33-mer peptide carrying up to six overlapping CD epitopes and is a strong binder and activator of human T cells; similarly, some gamma-gliadins have a highly immunogenic 26-mer peptide.
10 Breeding Methods for Coeliac-Safe Wheat
Due to its genetic flexibility, several advanced breeding techniques are being used in wheat [85]. Wide hybridizations may result in irregularities (translocations as well as deletions) in the genomes of embryos and their later plant gametes. Chromosomal deletion lines, with impacts regarding reduction of CD-immunogenic gliadin loci, have been found and tested [86]. Chromosomal translocations and deletions may be induced technologically (as was done in the varieties Chinese Spring and Paragon, amongst others) but they also occur naturally. They are interesting for genetic research, but often the deletions impact the performance of the plants in the field.
Synthetic hexaploids can be obtained from hybridizations of wheat varieties and species of different ploidy levels (e.g., a tetraploid durum female with AABB genome with a diploid Aegilops male with DD genome) requiring duplication of the originally allotriploid embryonic genome (ABD into AABBDD) to form a fertile hexaploid hybrid. An Aegilops line selected from the very wide D-genome diversity with a favourable CD profile may be hybridized with a low-immunogenic durum mother line. The resulting new hexaploids can be tested for their epitope profile at the gene and protein level, and applied in further breeding programs if desired [3]. This would be one way to eliminate the 33-mer in the D-genome alpha-gliadins.
EMS mutation breeding can generate large numbers of random mutations and could be applied to mutate gliadin genes, but it would be very resource-intensive to trace and combine mutations in multiple genes, from many plants, into one single, coeliac-safe and well-performing wheat plant [87]. Using γ-irradiation, large chromosomal deletions have been induced in two mutant lines of the wheat variety Paragon. Both mutants revealed a lack of sequence coverage in a large part of the gluten loci [25]. However, a clear need for more sophisticated approaches remains.
11 RNA Interference
RNA interference (RNAi) is a system that may be used to interfere with the synthesis of gluten proteins through their RNA transcripts, while the DNA still contains the intact genes. With a single RNAi construct designed for a conserved region that is common for many gluten genes, Gil-Humanes et al. [88] achieved up to 92% reduction of the gliadins, and a 10–100 fold reduction of epitopes as detected in T-cell tests. Similarly, the expression of twenty α-gliadin genes was decreased, although the production of other storage proteins increased [89]. Some of the wheat lines with reduced immunogenicity but with the baking quality largely intact [90] were sufficiently low in gliadins for successful clinical trials with patients [69]. Unfortunately, as RNAi requires stable genetic modification (GM) of the construct into wheat to silence the gliadins, and no transgenic wheat has been commercialised yet, it is unlikely that this transgenic line will be brought to the market.
12 Gene Editing of Gluten in Wheat
Two successful proof-of-concept studies with gene editing have been carried out in wheat. Sánchez-León et al. [24] targeted conserved sites in alpha-gliadin genes of a single wheat line. In this study, 47 offspring gene-edited plants were genotyped, and they found smaller and larger deletions at the target site, and plants with varying numbers of genes edited. In one plant 35 of the 45 alpha-gliadin genes were mutated, and seeds of this plant showed 85% reduction of total gluten protein as measured with the R5 gluten quantification assay, which is the type I method for gluten detection in the Codex Alimentarius. Guzmán-López et al. [91] developed a bioinformatics pipeline to show that in some offspring lines alpha-gliadin genes were deleted.
In the other study, Jouanin et al. [27] simultaneously targeted multiple sites in alpha-gliadin as well as gamma-gliadin genes, and confirmed 117 gene-edited plants, including plants with mutations in both gene families. Some gluten genes were deleted, as evidenced by droplet digital PCR (ddPCR) results for alpha-gliadins [27]. Evidence that in some regenerated plants blocks of genes had been deleted came from mapping of sequence data obtained with GlutEnSeq, a variant to RenSeq that targets gluten genes, to the Chinese Spring reference genome [25]. This is presumably the result of double-strand breaks in different gliadin genes that occur in tandem arrays in the wheat genome, leading to the deletion of tandem gene blocks in between the genes with double strand breaks.
As not all target sites in the gliadin genes will be mutated in a single progeny plant, gene editing will lead to populations of plants that each may contain a mosaic of unaffected, edited, and lost genes. The edits on different chromosomes will segregate in the next self-pollinated generations. Hence, it is useful to have a rigorous selection program aimed at maintaining only the few most promising plants (genotypes) for multiplication, cultivation and eventual application in coeliac-safe(r) food [92]. In this process, gene-edited loci may be combined with loci that are lost through smaller or large deletions.
13 Impacts of Gene-Edited Low CD-Immunogenic (‘Low-Gluten’) Wheat
Producing low CD-immunogenic (hypo-immunogenic or ‘low-gluten’) wheat is technically feasible with gene editing using CRISPR/Cas technology. Such wheat varieties and the food products made with them have the potential to make a significant contribution to food security and public health for a group of individuals that suffer from coeliac disease, and for others that prefer to consume gluten-free products for health or diet related reasons. Such products could fit well in the gluten-free product market of packaged foods, which currently is growing fast.
Gene editing with CRISPR/Cas initially requires foreign DNA or proteins to be inserted into the plant genome or plant cell, as a temporary step to perform editing (read: to induce a directed and desired mutation) of target genes. In many jurisdictions such plants are not regulated as GM, provided that no foreign DNA is present in the final plant. In Europe, however, the process-based approach is applied, and because CRISPR/Cas includes a temporary GM step, gene-edited wheat will fall under the strict GM regulation (EC Directive 2001/18), whereas plants and derived products from chemically and irradiation induced mutagenesis are exempted from GM regulation according to EC Directive 2001/18, Annex 1B [93]. Presently, in 2023, the European Commission is preparing a proposal on changing the legislation for plants derived from cisgenesis or targeted mutagenesis. The possible impacts that gene-edited low CD-immunogenic wheat, once on the market, may have on the economy, society and the environment, were reviewed by Sánchez et al. (in press) [94].
One aspect of low-gluten wheat is that it will require the establishment of a separate supply chain under Hazard Analysis and Critical Control Points (HACCP) conditions [92]. Cross-contamination with regular wheat and other gluten-containing cereals (rye, barley) should be strictly avoided, requiring continuous control throughout the entire production chain. Further, labelling should emphasize the unique properties of the derived food products, particularly to CD-patients and NCWS patients. Importantly, the current legislations (EU 828/2014 and EU 41/2009) on gluten-free labelling requiring a threshold at 20 ppm gluten would need to be re-evaluated for low-gluten, low CD-immunogenic wheat products, because these regulations are based on quantitative measurement of the total amount of gluten in ‘gluten-free’ products which should be below the threshold for coeliac safety of 20 ppm. Gene-edited low CD-immunogenic wheat still will retain a gluten content far above this threshold and would require a separate ‘CD-safe gluten’ label.
Adoption of gene-edited coeliac-safe wheat might generate many new business opportunities for the entire wheat value chain, leading to diversification of production and products. In all, healthier and more safe foods for the CD patient population and reduced health care costs to society can be envisioned when gene-edited low-gluten CD-safe wheat would become mainstream on the food market ([94, 95] and refs therein).
References
Ogbonnaya, F.C., Abdalla, O., Mujeeb-Kazi, A., Kazi, A.G., Xu, S.S., Gosman, N., Lagudah, E.S., Bonnett, D., Sorrells, M.E., Tsujimoto, H.: Synthetic Hexaploids: harnessing species of the primary Gene Pool for wheat improvement. In: Janick, J. (ed.) Plant Breeding Reviews, pp. 35–122. Wiley, Hoboken (2013)
Rosyara, U., Kishii, M., Payne, T., Sansaloni, C.P., Singh, R.P., Braun, H.-J., Dreisigacker, S.: Genetic contribution of synthetic Hexaploid wheat to CIMMYT’s spring bread wheat breeding germplasm. Sci. Rep. 9, 12355 (2019). https://doi.org/10.1038/s41598-019-47936-5
Schaart, J.G., Salentijn, E.M.J., Goryunova, S.V., Chidzanga, C., Esselink, D.G., Gosman, N., Bentley, A.R., Gilissen, L.J.W.J., Smulders, M.J.M.: Exploring the alpha-gliadin locus: the 33-mer peptide with six overlapping coeliac disease epitopes in Triticum aestivum is derived from a subgroup of Aegilops tauschii. Plant J. 106, 86–94 (2021). https://doi.org/10.1111/tpj.15147
Shewry, P.: What is gluten—why is it special? Front. Nutr. 6, 101 (2019). https://doi.org/10.3389/fnut.2019.00101
Wieser, H., Koehler, P., Scherf, K.A.: Chemistry of wheat gluten proteins: qualitative composition. Cereal Chem. 100, 23–35 (2023). https://doi.org/10.1002/cche.10572
Wieser, H., Koehler, P., Scherf, K.A.: Chemistry of wheat gluten proteins: quantitative composition. Cereal Chem. 100, 36–55 (2023). https://doi.org/10.1002/cche.10553
Smulders, M.J.M.: Then and now: a scientific view on plant breeding and technological innovation. In: Institute on Science for Global Policy (ISGP) (ed.) Sustainable Agriculture: The Role of Plant Breeding Innovation. A program and conference organized, facilitated, moderated, and convened by the ISGP with support from the American Seed Trade Association and Euroseeds (Internet Format), Nov. 17—18, 2020. pp. 21–25. http://scienceforglobalpolicy.org/publication/sustainable-agriculture-the-role-of-plant-breeding-innovation-sa-pbi (2021)
Wulff, B.B.H., Dhugga, K.S.: Wheat—the cereal abandoned by GM. Science. 361, 451–452 (2018). https://doi.org/10.1126/science.aat5119
Schaart, J.G., van de Wiel, C.C.M., Smulders, M.J.M.: Genome editing of polyploid crops: prospects, achievements and bottlenecks. Transgenic Res. 30, 337–351 (2021). https://doi.org/10.1007/s11248-021-00251-0
Bansal, M., Jindal, S., Wani, S.H., Ganie, S.A., Singh, R.: Genome editing and trait improvement in wheat. In: Wani, S.H., Mohan, A., Singh, G.P. (eds.) Physiological, Molecular, and Genetic Perspectives of Wheat Improvement, pp. 263–283. Springer, Cham (2021)
Debernardi, J.M., Tricoli, D.M., Ercoli, M.F., Hayta, S., Ronald, P., Palatnik, J.F., Dubcovsky, J.: A GRF–GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat. Biotechnol. 38, 1274–1279 (2020). https://doi.org/10.1038/s41587-020-0703-0
Wang, K., Shi, L., Liang, X., Zhao, P., Wang, W., Liu, J., Chang, Y., Hiei, Y., Yanagihara, C., Du, L., Ishida, Y., Ye, X.: The gene TaWOX5 overcomes genotype dependency in wheat genetic transformation. Nat. Plant. 8, 110–117 (2022). https://doi.org/10.1038/s41477-021-01085-8
Kumar, R., Kaur, A., Pandey, A., Mamrutha, H.M., Singh, G.P.: CRISPR-based genome editing in wheat: a comprehensive review and future prospects. Mol. Biol. Rep. 46, 3557–3569 (2019). https://doi.org/10.1007/s11033-019-04761-3
Kim, D., Hager, M., Brant, E., Budak, H.: Efficient genome editing in wheat using Cas9 and Cpf1 (AsCpf1 and LbCpf1) nucleases. Funct. Integr. Genomics. 21, 355–366 (2021). https://doi.org/10.1007/s10142-021-00782-z
Li, S., Zhang, C., Li, J., Yan, L., Wang, N., Xia, L.: Present and future prospects for wheat improvement through genome editing and advanced technologies. Plant Commun. 2, 100211 (2021). https://doi.org/10.1016/j.xplc.2021.100211
Smedley, M.A., Hayta, S., Clarke, M., Harwood, W.A.: CRISPR-Cas9 based genome editing in wheat. Curr. Protoc. 1, e65 (2021). https://doi.org/10.1002/cpz1.65
Lin, Q., Zong, Y., Xue, C., Wang, S., Jin, S., Zhu, Z., Wang, Y., Anzalone, A.V., Raguram, A., Doman, J.L., Liu, D.R., Gao, C.: Prime genome editing in rice and wheat. Nat. Biotechnol. 38, 582–585 (2020). https://doi.org/10.1038/s41587-020-0455-x
Luo, W., Suzuki, R., Imai, R.: Precise in planta genome editing via homology-directed repair in wheat. Plant Biotechnol. J. 21, 668–670 (2022). https://doi.org/10.1111/pbi.13984
ter Steeg, E.M.S., Struik, P.C., Visser, R.G.F., Lindhout, P.: Crucial factors for the feasibility of commercial hybrid breeding in food crops. Nat. Plant. 8, 463–473 (2022). https://doi.org/10.1038/s41477-022-01142-w
Raffan, S., Oddy, J., Mead, A., Barker, G., Curtis, T., Usher, S., Burt, C., Halford, N.G.: Field assessment of genome-edited, low asparagine wheat: Europe’s first CRISPR wheat field trial. Plant Biotechnol. J. 21, 1097–1099 (2023). https://doi.org/10.1111/pbi.14026
Raffan, S., Sparks, C., Huttly, A., Hyde, L., Martignago, D., Mead, A., Hanley, S.J., Wilkinson, P.A., Barker, G., Edwards, K.J., Curtis, T.Y., Usher, S., Kosik, O., Halford, N.G.: Wheat with greatly reduced accumulation of free asparagine in the grain, produced by CRISPR/Cas9 editing of asparagine synthetase gene TaASN2. Plant Biotechnol. J. 19, 1602–1613 (2021). https://doi.org/10.1111/pbi.13573
Anonymous: UK trials first CRISPR-edited wheat in Europe. Nat. Biotechnol. 39, 1174 (2021). https://doi.org/10.1038/s41587-021-01098-w
Zhang, S., Zhang, R., Gao, J., Song, G., Li, J., Li, W., Qi, Y., Li, Y., Li, G.: CRISPR/Cas9-mediated genome editing for wheat grain quality improvement. Plant Biotechnol. J. 19, 1684–1686 (2021). https://doi.org/10.1111/pbi.13647
Sánchez-León, S., Gil-Humanes, J., Ozuna, C.V., Giménez, M.J., Sousa, C., Voytas, D.F., Barro, F.: Low-gluten, nontransgenic wheat engineered with CRISPR/Cas9. Plant Biotechnol. J. 16, 902–910 (2018). https://doi.org/10.1111/pbi.12837
Jouanin, A., Borm, T., Boyd, L.A., Cockram, J., Leigh, F., Santos, B.A.C.M., Visser, R.G.F., Smulders, M.J.M.: Development of the GlutEnSeq capture system for sequencing gluten gene families in hexaploid bread wheat with deletions or mutations induced by γ-irradiation or CRISPR/Cas9. J. Cereal Sci. 88, 157–166 (2019). https://doi.org/10.1016/j.jcs.2019.04.008
Jouanin, A., Schaart, J.G., Boyd, L.A., Cockram, J., Leigh, F.J., Bates, R., Wallington, E.J., Visser, R.G.F., Smulders, M.J.M.: Outlook for coeliac disease patients: towards bread wheat with hypoimmunogenic gluten by gene editing of α- and γ-gliadin gene families. BMC Plant Biol. 19, 333 (2019). https://doi.org/10.1186/s12870-019-1889-5
Jouanin, A., Tenorio-Berrio, R., Schaart, J.G., Leigh, F., Visser, R.G.F., Smulders, M.J.M.: Optimisation of droplet digital PCR for determining copy number variation of α-gliadin genes in mutant and gene-edited polyploid bread wheat. J. Cereal Sci. 92, 102903 (2020). https://doi.org/10.1016/j.jcs.2019.102903
Rey, M.-D., Martín, A.C., Smedley, M., Hayta, S., Harwood, W., Shaw, P., Moore, G.: Magnesium increases Homoeologous crossover frequency during meiosis in ZIP4 (Ph1 gene) mutant wheat-wild relative hybrids. Front. Plant Sci. 9, 509 (2018). https://doi.org/10.3389/fpls.2018.00509
Hyde, L., Osman, K., Winfield, M., Sanchez-Moran, E., Higgins, J.D., Henderson, I.R., Sparks, C., Franklin, F.C.H., Edwards, K.J.: Identification, characterization, and rescue of CRISPR/Cas9 generated wheat SPO11-1 mutants. Plant Biotechnol. J. 21, 405–418 (2023). https://doi.org/10.1111/pbi.13961
Lv, J., Yu, K., Wei, J., Gui, H., Liu, C., Liang, D., Wang, Y., Zhou, H., Carlin, R., Rich, R., Lu, T., Que, Q., Wang, W.C., Zhang, X., Kelliher, T.: Generation of paternal haploids in wheat by genome editing of the centromeric histone CENH3. Nat. Biotechnol. 38, 1397–1401 (2020). https://doi.org/10.1038/s41587-020-0728-4
Budhagatapalli, N., Halbach, T., Hiekel, S., Büchner, H., Müller, A.E., Kumlehn, J.: Site-directed mutagenesis in bread and durum wheat via pollination by cas9/guide RNA-transgenic maize used as haploidy inducer. Plant Biotechnol. J. 18, 2376–2378 (2020). https://doi.org/10.1111/pbi.13415
Zhang, R., Zhang, S., Li, J., Gao, J., Song, G., Li, W., Geng, S., Liu, C., Lin, Y., Li, Y., Li, G.: CRISPR/Cas9-targeted mutagenesis of TaDCL4, TaDCL5 and TaRDR6 induces male sterility in common wheat. Plant Biotechnol. J. 21, 839–853 (2023). https://doi.org/10.1111/pbi.14000
Okada, A., Arndell, T., Borisjuk, N., Sharma, N., Watson-Haigh, N.S., Tucker, E.J., Baumann, U., Langridge, P., Whitford, R.: CRISPR/Cas9-mediated knockout of Ms1 enables the rapid generation of male-sterile hexaploid wheat lines for use in hybrid seed production. Plant Biotechnol. J. 17, 1905–1913 (2019). https://doi.org/10.1111/pbi.13106
Singh, M., Kumar, M., Albertsen, M.C., Young, J.K., Cigan, A.M.: Concurrent modifications in the three homeologs of Ms45 gene with CRISPR-Cas9 lead to rapid generation of male sterile bread wheat (Triticum aestivum L.). Plant Mol. Biol. 97, 371–383 (2018). https://doi.org/10.1007/s11103-018-0749-2
Kumagai, Y., Liu, Y., Hamada, H., Luo, W., Zhu, J., Kuroki, M., Nagira, Y., Taoka, N., Katoh, E., Imai, R.: Introduction of a second “green revolution” mutation into wheat via in planta CRISPR/Cas9 delivery. Plant Physiol. 188, 1838–1842 (2022). https://doi.org/10.1093/plphys/kiab570
Pearce, S.: Towards the replacement of wheat ‘green revolution’ genes. J. Exp. Bot. 72, 157–160 (2021). https://doi.org/10.1093/jxb/eraa494
Liu, H., Wang, K., Tang, H., Gong, Q., Du, L., Pei, X., Ye, X.: CRISPR/Cas9 editing of wheat TaQ genes alters spike morphogenesis and grain threshability. J. Genet. Genomics. 47, 563–575 (2020). https://doi.org/10.1016/j.jgg.2020.08.004
Gupta, A., Hua, L., Zhang, Z., Yang, B., Li, W.: CRISPR-induced miRNA156 -recognition element mutations in TaSPL13 improve multiple agronomic traits in wheat. Plant Biotechnol. J. 21, 536–548 (2023). https://doi.org/10.1111/pbi.13969
Dixon, L.E., Pasquariello, M., Badgami, R., Levin, K.A., Poschet, G., Ng, P.Q., Orford, S., Chayut, N., Adamski, N.M., Brinton, J., Simmonds, J., Steuernagel, B., Searle, I.R., Uauy, C., Boden, S.A.: MicroRNA-resistant alleles of HOMEOBOX DOMAIN-2 modify inflorescence branching and increase grain protein content of wheat. Sci. Adv. 8, 5907 (2022). https://doi.org/10.1126/sciadv.abn5907
Fan, M., Zhang, X., Nagarajan, R., Zhai, W., Rauf, Y., Jia, H., Ma, Z., Yan, L.: Natural variants and editing events provide insights into routes for spike architecture modification in common wheat. Crop J. 11, 148–156 (2023). https://doi.org/10.1016/j.cj.2022.04.009
Wang, Y., Du, F., Wang, J., Wang, K., Tian, C., Qi, X., Lu, F., Liu, X., Ye, X., Jiao, Y.: Improving bread wheat yield through modulating an unselected AP2/ERF gene. Nat. Plant. 8, 930–939 (2022). https://doi.org/10.1038/s41477-022-01197-9
Kuzay, S., Lin, H., Li, C., Chen, S., Woods, D.P., Zhang, J., Lan, T., von Korff, M., Dubcovsky, J.: WAPO-A1 is the causal gene of the 7AL QTL for spikelet number per spike in wheat. PLoS Genet. 18, e1009747 (2022). https://doi.org/10.1371/journal.pgen.1009747
Kabir, M.R., Nonhebel, H.M.: Reinvestigation of THOUSAND-GRAIN WEIGHT 6 grain weight genes in wheat and rice indicates a role in pollen development rather than regulation of auxin content in grains. Theor. Appl. Genet. 134, 2051–2062 (2021). https://doi.org/10.1007/s00122-021-03804-3
Wang, W., Pan, Q., Tian, B., He, F., Chen, Y., Bai, G., Akhunova, A., Trick, H.N., Akhunov, E.: Gene editing of the wheat homologs of TONNEAU1-recruiting motif encoding gene affects grain shape and weight in wheat. Plant J. 100, 251–264 (2019). https://doi.org/10.1111/tpj.14440
Zhang, Y., Li, D., Zhang, D., Zhao, X., Cao, X., Dong, L., Liu, J., Chen, K., Zhang, H., Gao, C., Wang, D.: Analysis of the functions of TaGW2 homoeologs in wheat grain weight and protein content traits. Plant J. 94, 857–866 (2018). https://doi.org/10.1111/tpj.13903
Wang, W., Simmonds, J., Pan, Q., Davidson, D., He, F., Battal, A., Akhunova, A., Trick, H.N., Uauy, C., Akhunov, E.: Gene editing and mutagenesis reveal inter-cultivar differences and additivity in the contribution of TaGW2 homoeologues to grain size and weight in wheat. Theor. Appl. Genet. 131, 2463–2475 (2018). https://doi.org/10.1007/s00122-018-3166-7
Wang, W., Pan, Q.L., He, F., Akhunova, A., Chao, S.M., Trick, H., Akhunov, E.: Transgenerational CRISPR-Cas9 activity facilitates multiplex gene editing in allopolyploid wheat. CRISPR J. 1, 65–74 (2018). https://doi.org/10.1089/crispr.2017.0010
Zhang, Z., Hua, L., Gupta, A., Tricoli, D., Edwards, K.J., Yang, B., Li, W.: Development of an Agrobacterium -delivered CRISPR/Cas9 system for wheat genome editing. Plant Biotechnol. J. 17, 1623–1635 (2019). https://doi.org/10.1111/pbi.13088
Li, C., Zong, Y., Wang, Y., Jin, S., Zhang, D., Song, Q., Zhang, R., Gao, C.: Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol. 19, 59 (2018). https://doi.org/10.1186/s13059-018-1443-z
Hahn, F., Sanjurjo Loures, L., Sparks, C.A., Kanyuka, K., Nekrasov, V.: Efficient CRISPR/Cas-mediated targeted mutagenesis in spring and winter wheat varieties. Plan. Theory. 10, 1481 (2021). https://doi.org/10.3390/plants10071481
Kan, J., Cai, Y., Cheng, C., Jiang, C., Jin, Y., Yang, P.: Simultaneous editing of host factor gene TaPDIL5-1 homoeoalleles confers wheat yellow mosaic virus resistance in hexaploid wheat. New Phytol. 234, 340–344 (2022). https://doi.org/10.1111/nph.18002
Chen, H., Su, Z., Tian, B., Liu, Y., Pang, Y., Kavetskyi, V., Trick, H.N., Bai, G.: Development and optimization of a Barley stripe mosaic virus -mediated gene editing system to improve Fusarium head blight resistance in wheat. Plant Biotechnol. J. 20, 1018–1020 (2022). https://doi.org/10.1111/pbi.13819
Li, G.Q., Zhou, J.Y., Jia, H.Y., Gao, Z.X., Fan, M., Luo, Y.J., Zhao, P.T., Xue, S.L., Li, N., Yuan, Y., Ma, S.W., Kong, Z.X., Jia, L., An, X., Jiang, G., Liu, W.X., Cao, W.J., Zhang, R.R., Fan, J.C., Xu, X.W., Liu, Y.F., Kong, Q.Q., Zheng, S.H., Wang, Y., Qin, B., Cao, S.Y., Ding, Y.X., Shi, J.X., Yan, H.S., Wang, X., Ran, C.F., Ma, Z.Q.: Mutation of a histidine-rich calcium-binding-protein gene in wheat confers resistance to Fusarium head blight. Nat. Genet. 51, 1106–1112 (2019). https://doi.org/10.1038/s41588-019-0426-7
Su, Z.Q., Bernardo, A., Tian, B., Chen, H., Wang, S., Ma, H.X., Cai, S.B., Liu, D.T., Zhang, D.D., Li, T., Trick, H., St Amand, P., Yu, J.M., Zhang, Z.Y., Bai, G.H.: A deletion mutation in TaHRC confers Fhb1 resistance to Fusarium head blight in wheat. Nat. Genet. 51, 1099–1105 (2019). https://doi.org/10.1038/s41588-019-0425-8
Wang, Y., Cheng, X., Shan, Q., Zhang, Y., Liu, J., Gao, C., Qiu, J.-L.: Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014). https://doi.org/10.1038/nbt.2969
Li, S., Lin, D., Zhang, Y., Deng, M., Chen, Y., Lv, B., Li, B., Lei, Y., Wang, Y., Zhao, L., Liang, Y., Liu, J., Chen, K., Liu, Z., Xiao, J., Qiu, J.-L., Gao, C.: Genome-edited powdery mildew resistance in wheat without growth penalties. Nature. 602, 455–460 (2022). https://doi.org/10.1038/s41586-022-04395-9
Deng, X., Cao, X.: Reconciliation between high yield and disease resistance. Nat. Rev. Genet. 23, 262–263 (2022). https://doi.org/10.1038/s41576-022-00471-y
Zhang, Y., Bai, Y., Wu, G., Zou, S., Chen, Y., Gao, C., Tang, D.: Simultaneous modification of three homoeologs of TaEDR1 by genome editing enhances powdery mildew resistance in wheat. Plant J. 91, 714–724 (2017). https://doi.org/10.1111/tpj.13599
He, F., Wang, C., Sun, H., Tian, S., Zhao, G., Liu, C., Wan, C., Guo, J., Huang, X., Zhan, G., Yu, X., Kang, Z., Guo, J.: Simultaneous editing of three homoeologues of TaCIPK14 confers broad-spectrum resistance to stripe rust in wheat. Plant Biotechnol. J. 21, 354–368 (2023). https://doi.org/10.1111/pbi.13956
Corredor-Moreno, P., Minter, F., Davey, P.E., Wegel, E., Kular, B., Brett, P., Lewis, C.M., Morgan, Y.M.L., Macías Pérez, L.A., Korolev, A.V., Hill, L., Saunders, D.G.O.: The branched-chain amino acid aminotransferase TaBCAT1 modulates amino acid metabolism and positively regulates wheat rust susceptibility. Plant Cell. 33, 1728–1747 (2021). https://doi.org/10.1093/plcell/koab049
Abdallah, N.A., Elsharawy, H., Abulela, H.A., Thilmony, R., Abdelhadi, A.A., Elarabi, N.I.: Multiplex CRISPR/Cas9-mediated genome editing to address drought tolerance in wheat. GM Crops Food. (2022). https://doi.org/10.1080/21645698.2022.2120313
Zhang, J., Zhang, H., Li, S., Li, J., Yan, L., Xia, L.: Increasing yield potential through manipulating of an ARE1 ortholog related to nitrogen use efficiency in wheat by CRISPR/Cas9. J. Integr. Plant Biol. 63, 1649–1663 (2021). https://doi.org/10.1111/jipb.13151
Liu, Y., Luo, W., Linghu, Q., Abe, F., Hisano, H., Sato, K., Kamiya, Y., Kawaura, K., Onishi, K., Endo, M., Toki, S., Hamada, H., Nagira, Y., Taoka, N., Imai, R.: In planta genome editing in commercial wheat varieties. Front. Plant Sci. 12, 648841 (2021). https://doi.org/10.3389/fpls.2021.648841
Ran, Y., Patron, N., Kay, P., Wong, D., Buchanan, M., Cao, Y.-Y., Sawbridge, T., Davies, J.P., Mason, J., Webb, S.R., Spangenberg, G., Ainley, W.M., Walsh, T.A., Hayden, M.J.: Zinc finger nuclease-mediated precision genome editing of an endogenous gene in hexaploid bread wheat (Triticum aestivum) using a DNA repair template. Plant Biotechnol. J. 16, 2088–2101 (2018). https://doi.org/10.1111/pbi.12941
Qiu, F., Xing, S., Xue, C., Liu, J., Chen, K., Chai, T., Gao, C.: Transient expression of a TaGRF4-TaGIF1 complex stimulates wheat regeneration and improves genome editing. Sci. China Life Sci. 65, 731–738 (2022). https://doi.org/10.1007/s11427-021-1949-9
Zhang, R., Liu, J., Chai, Z., Chen, S., Bai, Y., Zong, Y., Chen, K., Li, J., Jiang, L., Gao, C.: Generation of herbicide tolerance traits and a new selectable marker in wheat using base editing. Nat. Plant. 5, 480–485 (2019). https://doi.org/10.1038/s41477-019-0405-0
Pietsch, V.L., Werner, R., Karbstein, H.P., Emin, M.A.: High moisture extrusion of wheat gluten: relationship between process parameters, protein polymerization, and final product characteristics. J. Food Eng. 259, 3–11 (2019). https://doi.org/10.1016/j.jfoodeng.2019.04.006
van den Broeck, H.C., Gilissen, L.J.W.J., Smulders, M.J.M., van der Meer, I.M., Hamer, R.J.: Dough quality of bread wheat lacking α-gliadins with celiac disease epitopes and addition of celiac-safe avenins to improve dough quality. J. Cereal Sci. 53, 206–216 (2011). https://doi.org/10.1016/j.jcs.2010.12.004
Guzmán-López, M.H., Sánchez-León, S., Marín-Sanz, M., Comino, I., Segura, V., Vaquero, L., Rivero-Lezcano, O.M., Pastor, J., Sousa, C., Vivas, S., Barro, F.: Oral consumption of bread from an RNAi wheat line with strongly silenced gliadins elicits no immunogenic response in a pilot study with celiac disease patients. Nutrients. 13, 4548 (2021). https://doi.org/10.3390/nu13124548
Köhler, P., Belitz, H.-D., Wieser, H.: Disulphide bonds in wheat gluten: further cystine peptides from high molecular weight (HMW) and low molecular weight (LMW) subunits of glutenin and from γ-gliadins. Z. Lebensm. Unters. Forsch. 196, 239–247 (1993). https://doi.org/10.1007/BF01202740
Atchison, J., Head, L., Gates, A.: Wheat as food, wheat as industrial substance; comparative geographies of transformation and mobility. Geoforum. 41, 236–246 (2010). https://doi.org/10.1016/j.geoforum.2009.09.006
Kromhout, D., Spaaij, C.J.K., de Goede, J., Weggemans, R.M.: The 2015 Dutch food-based dietary guidelines. Eur. J. Clin. Nutr. 70, 869–878 (2016). https://doi.org/10.1038/ejcn.2016.52
Gilissen, L.J.W.J., van den Broeck, H.C.: Breeding for healthier wheat. Cereal Foods World. 63, 132–136 (2018). https://doi.org/10.1094/CFW-63-4-0132
Gilissen, L.J.W.J., van der Meer, I.M., Smulders, M.J.M.: Reducing the incidence of allergy and intolerance to cereals. J. Cereal Sci. 59, 337–353 (2014). https://doi.org/10.1016/j.jcs.2014.01.005
Rustgi, S., Kashyap, S., Ankrah, N., von Wettstein, D.: Use of microspore-derived Calli as explants for biolistic transformation of common wheat. In: Rustgi, S., Luo, H. (eds.) Biolistic DNA Delivery in Plants. Methods and Protocols. Methods in Molecular Biology, pp. 263–279. Humana Press Inc., New York (2020)
Kasarda, D.D.: Can an increase in celiac disease be attributed to an increase in the gluten content of wheat as a consequence of wheat breeding? J. Agric. Food Chem. 61, 1155–1159 (2013). https://doi.org/10.1021/jf305122s
Hischenhuber, C., Crevel, R., Jarry, B., Maki, M., Moneret-Vautrin, D.A., Romano, A., Troncone, R., Ward, R.: Review article: safe amounts of gluten for patients with wheat allergy or coeliac disease. Aliment. Pharmacol. Ther. 23, 559–575 (2006). https://doi.org/10.1111/j.1365-2036.2006.02768.x
Perrin, L., Allès, B., Buscail, C., Ravel, C., Hercberg, S., Julia, C., Kesse-Guyot, E.: Gluten-free diet in French adults without coeliac disease: sociodemographic characteristics, motives and dietary profile. Br. J. Nutr. 122, 231–239 (2019). https://doi.org/10.1017/S0007114519001053
Sollid, L.M., Tye-Din, J.A., Qiao, S.-W., Anderson, R.P., Gianfrani, C., Koning, F.: Update 2020: nomenclature and listing of celiac disease–relevant gluten epitopes recognized by CD4+ T cells. Immunogenetics. 72, 85–88 (2020). https://doi.org/10.1007/s00251-019-01141-w
International Wheat Genome Sequencing Consortium (IWGSC): Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science. 361, eaar7191 (2018). https://doi.org/10.1126/science.aar7191
Huo, N., Zhang, S., Zhu, T., Dong, L., Wang, Y., Mohr, T., Hu, T., Liu, Z., Dvorak, J., Luo, M.-C., Wang, D., Lee, J.-Y., Altenbach, S., Gu, Y.Q.: Gene duplication and evolution dynamics in the Homeologous regions harboring multiple Prolamin and resistance gene families in Hexaploid wheat. Front. Plant Sci. 9, 673 (2018). https://doi.org/10.3389/fpls.2018.00673
Huo, N., Zhu, T., Altenbach, S., Dong, L., Wang, Y., Mohr, T., Liu, Z., Dvorak, J., Luo, M.-C., Gu, Y.Q.: Dynamic evolution of α-Gliadin Prolamin gene family in Homeologous genomes of Hexaploid wheat. Sci. Rep. 8, 5181 (2018). https://doi.org/10.1038/s41598-018-23570-5
Altenbach, S.B., Chang, H.-C., Simon-Buss, A., Mohr, T., Huo, N., Gu, Y.Q.: Exploiting the reference genome sequence of hexaploid wheat: a proteomic study of flour proteins from the cultivar Chinese Spring. Funct. Integr. Genomics. 20, 1–16 (2020). https://doi.org/10.1007/s10142-019-00694-z
van Herpen, T.W., Goryunova, S.V., van der Schoot, J., Mitreva, M., Salentijn, E., Vorst, O., Schenk, M.F., van Veelen, P.A., Koning, F., van Soest, L.J., Vosman, B., Bosch, D., Hamer, R.J., Gilissen, L.J., Smulders, M.J.: Alpha-gliadin genes from the a, B, and D genomes of wheat contain different sets of celiac disease epitopes. BMC Genomics. 7, 1 (2006). https://doi.org/10.1186/1471-2164-7-1
Gilissen, L.J.W.J., Smulders, M.J.M.: Low gluten and coeliac-safe wheat through gene editing. In: Ricroch, A., Chopra, S., Kuntz, M. (eds.) Plant Biotechnology, pp. 231–248. Springer, Cham (2021)
van den Broeck, H.C., van Herpen, T.W., Schuit, C., Salentijn, E.M., Dekking, L., Bosch, D., Hamer, R.J., Smulders, M.J., Gilissen, L.J., van der Meer, I.M.: Removing celiac disease-related gluten proteins from bread wheat while retaining technological properties: a study with Chinese Spring deletion lines. BMC Plant Biol. 9, 41 (2009). https://doi.org/10.1186/1471-2229-9-41
Jouanin, A., Gilissen, L.J.W.J., Boyd, L.A., Cockram, J., Leigh, F.J., Wallington, E.J., Van den Broeck, H.C., Van der Meer, I.M., Schaart, J.G., Visser, R.G.F., Smulders, M.J.M.: Food processing and breeding strategies for coeliac-safe and healthy wheat products. Food Res. Int. 110, 11–21 (2018). https://doi.org/10.1016/j.foodres.2017.04.025
Gil-Humanes, J., Pistón, F., Tollefsen, S., Sollid, L.M., Barro, F.: Effective shutdown in the expression of celiac disease-related wheat gliadin T-cell epitopes by RNA interference. Proc. Natl. Acad. Sci. 107, 17023–17028 (2010). https://doi.org/10.1073/pnas.1007773107
Becker, D., Wieser, H., Koehler, P., Folck, A., Mühling, K.H., Zörb, C.: Protein composition and techno-functional properties of transgenic wheat with reduced α-gliadin content obtained by RNA interference. J. Appl. Bot. Food Qual. 85, 23–33 (2012). https://ojs.openagrar.de/index.php/JABFQ/article/view/1995
Gil-Humanes, J., Pistón, F., Altamirano-Fortoul, R., Real, A., Comino, I., Sousa, C., Rosell, C.M., Barro, F.: Reduced-gliadin wheat bread: an alternative to the gluten-free diet for consumers suffering gluten-related pathologies. PLoS One. 9, e90898 (2014). https://doi.org/10.1371/journal.pone.0090898
Guzmán-López, M.H., Marín-Sanz, M., Sánchez-León, S., Barro, F.: A bioinformatic workflow for InDel analysis in the wheat multi-copy α-gliadin gene family engineered with CRISPR/Cas9. Int. J. Mol. Sci. 22, 13076 (2021). https://doi.org/10.3390/ijms222313076
Jouanin, A., Gilissen, L.J.W.J., Schaart, J.G., Leigh, F.J., Cockram, J., Wallington, E.J., Boyd, L.A., van den Broeck, H.C., van der Meer, I.M., America, A.H.P., Visser, R.G.F., Smulders, M.J.M.: CRISPR/Cas9 gene editing of gluten in wheat to reduce gluten content and exposure—reviewing methods to screen for coeliac safety. Front. Nutr. 7, 51 (2020). https://doi.org/10.3389/fnut.2020.00051
Jouanin, A., Boyd, L., Visser, R.G.F., Smulders, M.J.M.: Development of wheat with hypoimmunogenic gluten obstructed by the gene editing policy in Europe. Front. Plant Sci. 9, 1523 (2018). https://doi.org/10.3389/fpls.2018.01523
Sánchez, B., Barro, F., Smulders, M.J.M., et al.: Economic and social impacts of low-gluten coeliac-safe wheat developed by gene editing. JRC Tech. Rep. (in press) (2023)
Maaß, O., Consmüller, N., Kehlenbeck, H.: Socioeconomic impact of genome editing on agricultural value chains: the case of fungal-resistant and coeliac-safe wheat. Sustainability. 11, 6421 (2019). https://doi.org/10.3390/su11226421
Acknowledgements
The writing of this chapter was partially made possible by the Well on Wheat (WoW) project (https://www.um-eatwell.nl/wow/), an international research project under the umbrella of the Health Grain Forum and ICC-Vienna and partially funded by the Dutch Topsector AgriFood (project TKI 1601P01 to LJWJG), and partly by the Netherlands’ Ministry of Agriculture, Nature and Food Quality (BO-43-121-001 to MJMS and CMMvdW).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2024 The Author(s)
About this chapter
Cite this chapter
Smulders, M.J.M., Gilissen, L.J.W.J., Juranić, M., Schaart, J.G., van de Wiel, C.C.M. (2024). Gene Editing of Wheat to Reduce Coeliac Disease Epitopes in Gluten. In: Ricroch, A., Eriksson, D., Miladinović, D., Sweet, J., Van Laere, K., Woźniak-Gientka, E. (eds) A Roadmap for Plant Genome Editing . Springer, Cham. https://doi.org/10.1007/978-3-031-46150-7_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-46150-7_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-46149-1
Online ISBN: 978-3-031-46150-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)