Abstract
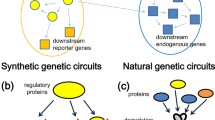
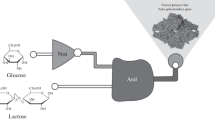
In the field of synthetic biology, genetic networks are designed by combining well-characterized genetic parts, similar to electronic circuits. Such gene networks are called synthetic genetic circuits. The design approach for synthetic genetic circuits is based on mathematical modeling and numerical simulation. The approach allows the realization of various cellular functions. However, unavoidable differences in the initial states or fluctuations of the gene expression in cells have prevented the precise prediction and control of cellular behavior. Therefore, the design of synthetic genetic circuits is not sufficient, and the dynamic control of the circuits is also required. In this report, we provide examples of synthetic circuit designs and the control of synthetic biological systems, as well as perspectives on design and control.
Chapter PDF
Similar content being viewed by others
References
Connor, M.R., Atsumi, S.: Synthetic biology guides biofuel production. J. Biomed. Biotechnol. 2010, 541698 (2010)
Ruder, W.C., Lu, T., Collins, J.J.: Synthetic biology moving into the clinic. Science 333, 1248–1252 (2011)
Elowitz, M.B., Leibler, S.: A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000)
Stricker, J., Cookson, S., Bennett, M.R., Mather, W.H., Tsimring, L.S., Hasty, J.: A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008)
Tigges, M., Marquez-Lago, T.T., Stelling, J., Fussenegger, M.: A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009)
Danino, T., Mondragón-Palomino, O., Tsimring, L., Hasty, J.: A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010)
Friedland, A.E., Lu, T.K., Wang, X., Shi, D., Church, G., Collins, J.J.: Synthetic gene networks that count. Science 324, 1199–1202 (2009)
Atsumi, S., Hanai, T., Liao, J.C.: Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451, 86–89 (2008)
Basu, S., Gerchman, Y., Collins, C.H., Arnold, F.H., Weiss, R.: A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005)
Tabor, J.J., Salis, H.M., Simpson, Z.B., Chevalier, A.A., Levskaya, A., Marcotte, E.M., Voigt, C.A., Ellington, A.D.: A Synthetic Genetic Edge Detection Program. Cell 137, 1272–1281 (2009)
Liu, C., Fu, X., Liu, L., Ren, X., Chau, C.K., Li, S., Xiang, L., Zeng, H., Chen, G., Tang, L.H., Lenz, P., Cui, X., Huang, W., Hwa, T., Huang, J.D.: Sequential establishment of stripe patterns in an expanding cell population. Science 334, 238–241 (2011)
Gardner, T.S., Cantor, C.R., Collins, J.J.: Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000)
Kramer, B.P., Viretta, A.U., Baba, M.D.-E., Aubel, D., Weber, W., Fussenegger, M.: An engineered epigenetic transgene switch in mammalian cells. Nat. Biotechnol. 22, 867–870 (2004)
Bratsun, D., Volfson, D., Tsimring, L.S., Hasty, J.: Delay-induced stochastic oscillations in gene regulation. Proc. Natl. Acad. Sci. U.S.A. 102, 14593–14598 (2005)
Brenner, K., You, L., Arnold, F.: Engineering microbial consortia: a new frontier in synthetic biology. Trends Biotechnol. 26, 483–489 (2008)
Dong, Y.H.: AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc. Natl. Acad. Sci. U.S.A. 97, 3526–3531 (2000)
Sekine, R., Yamamura, M., Ayukawa, S., Ishimatsu, K., Akama, S., Takinoue, M., Hagiya, M., Kiga, D.: Tunable synthetic phenotypic diversification on Waddington’s landscape through autonomous signaling. Proc. Natl. Acad. Sci. U.S.A. 108, 17969–17973 (2011)
Kambam, P.K.R., Sayut, D.J., Niu, Y., Eriksen, D.T., Sun, L.: Directed evolution of LuxI for enhanced OHHL production. Biotechnol. Bioeng. 101, 263–272 (2008)
Toettcher, J.E., Gong, D., Lim, W.A., Weiner, O.D.: Light-based feedback for controlling intracellular signaling dynamics. Nat. Methods 8, 837–839 (2011)
Milias-Argeitis, A., Summers, S., Stewart-Ornstein, J., Zuleta, I., Pincus, D., El-Samad, H., Khammash, M., Lygeros, J.: In silico feedback for in vivo regulation of a gene expression circuit. Nat. Biotechnol. 29, 1114–1116 (2011)
Morari, M., Lee, J.H.: Model predictive control: past, present and future. Comput. Chem. Eng. 23, 667–682 (1999)
Cantone, I., Marucci, L., Iorio, F., Ricci, M.A., Belcastro, V., Bansal, M., Santini, S., di Bernardo, M., di Bernardo, D., Cosma, M.P.: A yeast synthetic network for in vivo assessment of reverse-engineering and modeling approaches. Cell 137, 172–181 (2009)
Menolascina, F., di Bernardo, M., di Bernardo, D.: Analysis, design and implementation of a novel scheme for in-vivo control of synthetic gene regulatory networks. Automatica 47, 1265–1270 (2011)
Mukherji, S., van Oudenaarden, A.: Synthetic biology: understanding biological design from synthetic circuits. Nat. Rev. Gen. (2009)
Isaacs, F.J., Dwyer, D.J., Collins, J.J.: RNA synthetic biology. Nat. Biotechnol. 24, 545–554 (2006)
Yokobayashi, Y., Weiss, R., Arnold, F.H.: Directed evolution of a genetic circuit. Proc. Natl. Acad. Sci. U.S.A. 99, 16587–16591 (2002)
Haseltine, E.L., Arnold, F.H.: Synthetic gene circuits: design with directed evolution. Annu. Rev. Biophys. Biomol. Struct. 36, 1–19 (2007)
Collins, C.H., Leadbetter, J.R., Arnold, F.H.: Dual selection enhances the signaling specificity of a variant of the quorum-sensing transcriptional activator LuxR. Nat. Biotechnol. 24, 708–712 (2006)
Alper, H., Fischer, C., Nevoigt, E., Stephanopoulos, G.: Tuning genetic control through promoter engineering. Proc. Natl. Acad. Sci. U.S.A. 102, 12678–12683 (2005)
Wang, Q., Wu, H., Wang, A., Du, P., Pei, X., Li, H., Yin, X., Huang, L., Xiong, X.: Prospecting metagenomic enzyme subfamily genes for DNA family shuffling by a novel PCR-based approach. J. Biol. Chem. 285, 41509–41516 (2010)
Kim, S.H., Yamamoto, T., Fourmy, D., Fujii, T.: An electroactive microwell array for trapping and lysing single-bacterial cells. Biomicrofluidics 5, 24114 (2011)
Keshavarz, M., Barkhordari Yazdi, M., Jahed-Motlagh, M.R.: Piecewise affine modeling and control of a boiler–turbine unit. Applied Thermal Engineering 30, 781–791 (2010)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution-NonCommercial 2.5 International License (http://creativecommons.org/licenses/by-nc/2.5/), which permits any noncommercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2013 The Author(s)
About this paper
Cite this paper
Sekine, R., Yamamura, M. (2013). Design and Control of Synthetic Biological Systems. In: Suzuki, Y., Nakagaki, T. (eds) Natural Computing and Beyond. Proceedings in Information and Communications Technology, vol 6. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54394-7_9
Download citation
DOI: https://doi.org/10.1007/978-4-431-54394-7_9
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54393-0
Online ISBN: 978-4-431-54394-7
eBook Packages: Computer ScienceComputer Science (R0)