Abstract
Deep sequencing of human tumours has uncovered a previously unappreciated role for epigenetic regulators in tumorigenesis. H3K4 methyltransferase KMT2C/MLL3 is mutated in several solid malignancies, including more than 10% of breast tumours. To study the tumour suppressor role of KMT2C in breast cancer, we generated mouse models of Erbb2/Neu, Myc or PIK3CA-driven tumorigenesis, in which the Kmt2c locus is knocked out specifically in the luminal lineage of mouse mammary glands using the Cre recombinase. Kmt2c knock out mice develop tumours earlier, irrespective of the oncogene, assigning a bona fide tumour suppressor role for KMT2C in mammary tumorigenesis. Loss of Kmt2c induces extensive epigenetic and transcriptional changes, which lead to increased ERK1/2 activity, extracellular matrix re-organization, epithelial-to-mesenchymal transition and mitochondrial dysfunction, the latter associated with increased reactive oxygen species production. Loss of Kmt2c renders the Erbb2/Neu-driven tumours more responsive to lapatinib. Publicly available clinical datasets revealed an association of low Kmt2c gene expression and better long-term outcome. Collectively, our findings solidify the role of KMT2C as a tumour suppressor in breast cancer and identify dependencies that could be therapeutically amenable.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer affects millions of women every year [1]. Despite the efficacy of therapies, there is a high degree of variability in response [2, 3]. Several studies [3,4,5,6,7] have performed deep sequencing analysis on breast cancer patient samples to reveal the landscape of somatic driver mutations, which could be associated with tumour aggressiveness and variation in response.
In these studies, among the most frequently mutated genes is the lysine methyltransferase KMT2C (also referred to as MLL3) [3, 6,7,8], an epigenetic modifier that methylates the histone 3 tail at lysine 4 (H3K4) at enhancers [9,10,11] and at specific promoters of genes in a cell-type specific manner [12] as part of the Complex of Proteins Associated with Set1 (COMPASS) [13]. KMT2C is thought to be functionally redundant with KMT2D [11, 12]. KMT2C is often found mutated not only in breast cancer but also in other solid tumours and haematological malignancies [14, 15].
In breast cancer, KMT2C plays a role in the transcriptional control of oestrogen-regulated genes [16,17,18]. Loss of KMT2C correlates with a reduction in proliferation in oestrogen-driven tumour cells but with a promotion of cell growth in oestrogen-depleted media [17]. Importantly, silencing of KMT2C in long-term oestrogen deprived cells associates with response to various therapeutic agents such as fulvestrant, AZD9496, ARN1917, GDC927 and RU58668 [17]. In other cancers, KMT2C has been implicated in the therapeutic response to DNA damaging factors or inhibitors of DNA repair proteins [19,20,21].
Despite the recent interest in KMT2C, its role in breast tumour initiation, progression and response to therapy remains poorly understood. Herein, we assessed the effect of Kmt2c loss in mammary tumorigenesis using tissue specific knockout of KMT2C locus in various murine models. Our results uncovered a bona fide tumour suppressor role for KMT2C in mammary tumorigenesis. Specifically, we show that mammary glands lacking KMT2C activity develop tumours with shorter latency than their wild-type (WT) counterparts. Moreover, inactivation of Kmt2c leads to an ERK/MAPK signalling imbalance, dysregulation of mitochondrial respiration and sensitivity to lapatinib.
Materials and methods
Mice
Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Neu, Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Myc and Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Myc;Pik3caH1047R mice were used in this study together with control mice that lack the KMT2Cfl/fl component or the Cre driver. The Kmt2cfl/fl mouse line was generated using CRISPR/cas9 in-house as follows: the W9.5 wild-type 129SV embryonic stem (ES) cells (kind gift from Colin L. Stewart) were cultured in KnockOut DMEM (Gibco, Cat No. 10829018), supplemented with 10% KnockOut serum replacement (Gibco, Cat No.10828028), penicillin/streptomycin (Corning, Cat No. 30-002-CI), nonessential amino acids (Corning, Cat No. 25–025-CI), L-glutamine (Corning, Cat No. 25-005-CI), β-Mercaptoethanol (Applichem, Cat No. A1108) and Leukemia inhibitory factor (LIF) (Santa Cruz, Cat No. sc-4989).
The targeting construct was linearized with Acc65I and phenol/chloroform purified. A mixture of the targeting construct (12.5 μg), the pX330 gRNA 1 (5 μg) and the pX330 gRNA 2 (5 μg) were electroporated into the ES cells that were selected with G418 for 8 days. We added the 2 gRNAs, designed for CRISPR, for increased efficiency.
After selection, a total of 288 resistant clones were picked and the genomic DNA was first screened with PCR genotyping. Thirty-three positive clones were further analyzed by Southern blot, using a 5’ external probe. Genomic DNA was digested with BglII. The targeted allele yielded a band of 2289 bp. Correctly targeted ES clones were injected in C57BL/6 blastocysts. Male chimeras were crossed to C57BL/6 females and offspring was genotyped to assess germline transmission.
The Kmt2cfl/fl were crossed with mice that carry a Krt8 CreERT2 driver as well as either the MMTV-Myc or the MMTV-Neu transgenes or the combination of MMTV-Myc;Pik3caH1047R (mentioned as MMTV-Myc;ex20 in the manuscript). CreERT2 was activated by administering tamoxifen (MCE) (a total of 15 mg/25gr distributed over 5 days and administered every other day) to 4 weeks old mice that had reached 18gr of body weight. Tamoxifen (20 mg/ml) was dissolved in corn oil using mild sonication for 30 min with 5 min intervals.
Animals were housed in individually ventilated cages under specific pathogen-free conditions in full compliance with FELASA (Federation of Laboratory Animal Science Associations) recommendations in the Animal House Facility of the Biomedical Research Foundation of the Academy of Athens (BRFAA, Greece). All procedures for the care and treatment of the animals were approved by the Institutional Committee on Ethics of Animal Experiments. The licence for the animal handling protocol for this project is: 1317/13-03-2019. All dissected tumors have comparable size. The size of the tumours was measured using caliper.
Generation of mouse cell lines
Mice were euthanized, and breast tumour was dissected and washed in ice-cold sterile PBS. Tumour was chopped into very fine pieces using razor blade under sterile conditions and it was incubated with 0.25% Trypsin–EDTA for 15 min at 37 °C. Trypsin was inactivated using DMEM high glycose with 10% FBS. The trypsinized tissue was passed through an 18 G syringe and was let to attach on an FBS-coated plate for 48 h. The plate was split once cells reached confluency. Cells were washed with PBS/5% FBS and stained with anti-CD24 (clone M1/69, Biolegend). CD24 + cells were sorted by flow cytometry (FACSAriaIII, BD) to mark luminal cells and to remove any fibroblasts present.
In vivo treatments
Lapatinib (MCE HY-50898) was administered by oral gavage daily (200 mg/kg daily) to genetically engineered mouse models (GEMMs). Lapatinib was dissolved in 0.5% carboxymethylcellulose/0.1% Tween-20 using mild sonication for 30 min. Tumour volume was measured twice a week with calliper and calculated as V = axb2/2, “a” being the largest diameter, “b” the smallest. Trametinib (MCE: HY-10999) was dissolved in 40% PEG400, 5% Tween-80, 45% saline and 10% DMSO and was administered orally (1 mg/kg/day).
Histopathological evaluation
Paraffin-embedded sections were deparaffinized and subjected to Hematoxylin and Eosin (H&E) staining using standard techniques for pathology assessment.
Immunohistochemistry (IHC) and immunofluorescence (IF)
IHC and IF were performed according to standard procedures. In brief, sections were deparaffinized, followed by heat-induced antigen retrieval with citrate buffer (pH 6) for 20 min using a pressure cooker. In IHC, endogenous peroxidase activity was blocked with hydrogen peroxide for 40 min followed by PBS washes. Sections were permeabilized with PBS/0.3% Triton X-100 and blocked with FBS/BSA/0.3% Triton X-100. Primary antibodies were added for overnight incubation at 4 °C in humidified chambers. The next day, secondary antibodies were added for 2 h at room temperature followed by washes. In IHC, the secondary antibodies (Cell Signaling Technology) were HRP-conjugated and the signal was detected with DAB (Vector Laboratories). Sections were counterstained with Mayer’s hematoxylin and mounted with DPX. In IF, secondary antibodies were purchased from the Jackson laboratory. Sections were counterstained with DAPI (MilliporeSigma) and mounted with mowiol. Primary antibodies: rabbit anti-Ki67 (Abcam ab15580, 1:150), rat anti-KRT8 (Troma I DSHB, 1:100), phospho-ERK1/2 (Cell Signaling Technology #4370, 1:100).
Immunoblotting
Tumours were lysed in 8 M UREA/50 mM TEAB with protease inhibitors (Calbiochem) using mild sonication on ice followed by homogenization with a 26 G syringe. Lysates were subjected to SDS-PAGE followed by immunoblot analysis. Primary antibodies: phospho-ERK1/2 (CST #4370), total ERK1/2 (CST #9102), phospho- p38 (CST #4511), total p-38 (CST # 8690), GAPDH (CST #5174), H3K4me1 (CST #5326), H3K4me3 (CST #9751), ATP5a (ab14748), NDUFSs3 (ab14711), phospho-c-JUN (sc-822), total c-JUN (CST #9165), phospho-JNK (sc-6254), H-RASs (Millipore), K-RAS (sc-30), KDM6A/UTX (CST #33510), ERα (sc-8002), KRT6α (Biolegend 905702), β-actin (MAB8929, Millipore).
Real-time PCR
RNA was isolated from snap frozen tumours in TRI reagent (MilliporeSigma) following the manufacturer’s instructions. To assist the RNA isolation, tissue was pulverized with liquid nitrogen using mortar and pestle. cDNA was prepared using PrimeScript RT Reagent Kit (Takara) and quantitative PCR was performed using KAPA SYBR FAST qPCR Master Mix (Kapa Biosystems), in a Roche LightCycler 96. Primer sequences are shown in supplementary Table 1.
RNA sequencing
RNA was isolated as described above. Initial raw fastq files were aligned to the Genome Reference Consortium Mouse Build 38 (GRCm38) using the ultrafast universal RNA-seq aligner (STAR) (Version 2.7.5b) [22]. Resulted raw counts were generated using the featureCounts (Version 2.0.1) [23]. The fastqc (Version 0.11.9) and the multiqc (Version 1.0) [24] tools were used to access the quality of the alignment and the quantification process. To perform differential expression analysis, DESeq2 [25] was applied. Functional enrichment analysis was performed by using the preranked tool of GSEA [26, 27] and Metascape [28].
ChIP sequencing
Tumours were dissociated into cells using a collagenase/hyalouronidase mix for 2 h at 37 °C, followed by brief incubation with 0.25% Trypsin–EDTA and then with Dispase as previously described (Prater and Stingl Methods). Cells were then fixed and subjected to chromatin immunoprecipitation as previously described [29] with an additional step of micrococcal nuclease incubation (CST #10011) prior to DNA sonication with metal probe. Antibodies: H3K4me1 (CST #5326), H3K4me3 (CST #9751), H3K27ac (CST #8173).
Raw fastq files were aligned to the reference mouse assembly (mm10) using the bowtie2 (Version 2.4.1) [30], while the filtering of the uniquely mapping reads was performed using the sambamba view (Version 0.6.6) [31]. The bam files of each histone modification were processed with deeptools [32], in order to inspect the degree of correlation between biological replicates, in each condition, genome-wide (3 kb bin counts). The pairs of biological replicates for each condition and histone modification were highly correlated (Pearson Correlation Coefficient > 0.94), thus bam files were down-sampled to the same level of mapped reads, and then merged using samtools [33]. For each IP-INPUT pair, peaks were called by using epic2 [34], an ultra-performant reimplementation of SICER peak caller. To compare the resulting peak sets between biological conditions in an unbiased manner, reads were pre-normalized to the same level of uniquely aligned reads. To scan the mouse genome for wide signal islands, a window of 400 bps with 1 gap was applied. Each read was extended to 200 bps, while the rest of the parameters were set as default. Additional filtering was applied to keep the most significant regions in terms of FDR and log2 Fold Change (FDR < 0.05, and log2 Fold Change > 1) and to exclude all the mm10 blacklisted regions by ENCODE. Peak overlap analysis was performed by using bedTools [35] and intervene python package [36]. Snapshots of bam signal distribution and peaks along particular genomic regions (Atp genes) was visualized using the Gviz package [37].
Peaks were annotated regarding their genomic position relative to genic regions, using ChiPSeeker [38]. To annotate peaks that were exclusively called in each biological condition, bedTools subtract-A was used. H3K4me3 peaks were assigned to gene promoters using bedtools intersect. To define active promoter and enhancer elements, MMTV-Neu H3K27ac peaks were intersected (bedtools) with Ensemble TSSs and FANTOM5 enhancer elements, respectively. H3K27ac, H3K4me3 and H3K4me1 signal was visualized along active promoters (2 kb regions around active TSSs) and active enhancers (2 kb region around enhancer TSSs/eTSSs) as heatmaps and average profiles, using seqMINER [39] and custom R scripts. Pathway enrichment analysis and motif enrichment analysis were performed using ClusterProfiler [40] and i-cisTarget [41], respectively. H3K4me1 exclusive peaks were assigned to candidate cis-regulatory elements (cCREs, The ENCODE Project Consortium, 2020) and were further processed by applying motif enrichment analysis, using HOMER [42].
Data availability
All the raw sequencing data has been submitted to NCBI SRA portal with the accession ID PRJNA787445.
Kaplan–Meier plots
Kaplan–Meier survival curves of tumour latency were calculated using the lifelines package (version 0.21.5) in python 3.7. p values were calculated using the Mann–Whitney U test in scipy package (version 1.2.1) and plots were created using matplotlib (version 3.0.3).
Clinical analysis
The online tool KM plotter [43] was used to observe statistical significance on RFS in breast cancer patients treated with any therapy. Separate graphs were generated based on the molecular subtype of the patients. The patients were split on median and results with log-rank p < 0.05 were assumed significant.
Mitochondria isolation and measurement of mitochondrial respiration
Mitochondria were isolated as described by [44]. Mitochondrial respiration was determined using a Clark-type O2 electrode connected to a computer operated Oxygraph control unit (Hansatech Instruments, Norfolk, U.K.) as previously described [44]. Freshly isolated mitochondria (150 μg of protein) were added to the respiration buffer (120 mM KCl, 5 mM KH2PO4, 3 mM HEPES, 1 mM EGTA, 1 mM MgCl2, 0.2% BSA, pH 7.2) containing 5 mM glutamate/2.5 mM malate. Basal O2 consumption was recorded (state 2) and after 2 min 500 μM ADP was added (State 3; indicates rate of ATP production, O2 consumption), followed by 6 μΜ oligomycin (State 4; denotes coupling) and 100 nM of the uncoupler (causes maximal respiration) carbonyl cyanide p-trifluoromethoxyphenylhydrazone (FCCP) (State FCCP). In all experiments, the temperature was maintained at 25 °C and the total reaction volume was 300 μl. The respiratory control ratio was calculated as the ratio of State 3 to State 4 (ST3/ST4).
Measurement of reactive oxygen species (ROS)
For the assessment of ROS production, tissues were incubated with 10 μM CM-H2DCFDA (Molecular Probes™/Thermo Fisher Scientific Inc.) dye in PBS for 30 min at 25 °C in the dark. Following dye removal, tissues were incubated for 15 min with PBS, then washed with PBS and lysed in NP-40 lysis buffer. The produced fluorescence was measured in cleared lysates using the Infinite 200 Tecan microtiter-plate photometer (Tecan Trading AG, Switzerland) at excitation and emission wavelengths of 490 and 540 nm, respectively. Obtained fluorescence values were normalized to total protein input.
Cell culture samples preparation and confocal laser scanning microscopy (CLSM) visualization
To label the mitochondria, cells grown on coverslips were incubated with the MitoTracker™ Green FM (Thermo Fisher Scientific Inc.) probe for 30 min at 37 °C as per manufacturer’s instructions. For visualizing nuclei, cells were counterstained with DAPI (Thermo Fisher Scientific Inc.). Samples were viewed in a Digital Eclipse C1 Nikon (Melville, NY, UAS) CLSM equipped with 20 × 0.50 NA differential interference contrast (DIC), 60 × 1.40 NA DIC Plan Apochromat objectives, using the EZC1 acquisition and analysis software (Nikon).
Results
KMT2C is a tumour suppressor in mammary glands of mice and humans
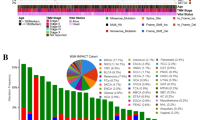
Sequencing data from human primary surgically resected breast tumours [7] identified mutations in the KMT2C locus, the majority of which are nonsense [45, 46]. Together with tumours carrying copy number alterations, mostly homozygous deletions, fusions and splice mutations, the cases showing genetic alterations in the KMT2C locus comprise 12% of total cases in breast cancer (Supplementary Fig. 1A). The nature of KMT2C alterations implies that it likely acts as a tumour suppressor in the mammary epithelium. In agreement with this notion, the disease-specific (DS) and overall survival (OS) of breast cancer patients with KMT2C alterations is significantly shorter (p = 0.009278 and 0.0299, respectively, Supplementary Fig. 1B).
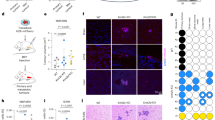
To investigate its presumptive tumour suppressor role in mice, we employed CRISPR/Cas9 to generate a conditional knockout of the Kmt2c locus (Kmt2cfl; Supplementary Fig. 2A and B). To delete the Kmt2c locus specifically in the luminal lineage of the mammary gland, we employed the Krt8-CreERT2 line [47] (hereafter K8-Cre), which drives the expression of the Tamoxifen-inducible CreERT2 recombinase in the luminal cells. We decided to study the KMT2C role in the context of mouse mammary tumours driven by the well-established breast cancer oncogenic drivers Erbb2/Neu, Myc and PIK3CAH1047R. Although mouse models of breast cancer do not faithfully recapitulate the molecular subtypes of the human disease, it has been reported that tumours arising in these models share molecular and histological features with the major human subtypes [48]. We thus generated pairs of cohorts of control and experimental female mice by crossing the Kmt2cfl allele and the K8-Cre transgene into the MMTV-Neu (YD) [49], (hereafter MMTV-Neu) transgenic line, and into bitransgenic mice carrying the MMTV-Myc [50] and the Pik3caH1047R conditional knock-in mutation (hereafter MMTV-Myc;ex20). In total, we generated three cohorts of experimental mice (K8-Cre;MMTV-Neu;Kmt2cfl/fl, K8-Cre;MMTV-Myc;Kmt2cfl/fl and K8-Cre;MMTV-Myc;aH1047R;Kmt2cfl/fl). As controls, we used littermates lacking either the K8-Cre or the Kmt2cfl alleles depending on the case (for details see Methods section). A reporter transgene (R26tdTomato) expressing the fluorescent protein Tomato in a Cre-dependent manner was introduced in all breeding programs employing the Kmt2cfl alleles to monitor recombination [51]. Mice were injected with Tamoxifen five consecutive days starting at the age of four weeks and once they reached a weight of 18 gr. After an 8-week recovery from the hormonal side effects of Tamoxifen, mice were bred for the rest of the monitoring period. Mice lacking KMT2C activity developed palpable tumours invariably earlier, with statistical significance, than the respective controls (Fig. 1A–C). Mice sometimes developed tumours in more than one mammary gland, although no difference in the number of tumours per animal or mammary gland was observed between control and KMT2C KO mice. H&E stainings did not indicate any gross differences between control and experimental mice (Fig. 1D, Supplementary Fig. 2C). Quantitative RT-PCR analysis of tumour samples indicated minimal amounts of Kmt2c transcripts, possibly reflecting contaminating stromal cells (Fig. 1E). Cytokeratin profiling of control and Kmt2c null tumours showed lack of Keratin 5 (Krt5) and maintenance of Keratin 8 abundance (Krt8) (Fig. 1F). Since the effect of Kmt2c loss on tumour latency was more conspicuous (p < 0.0001) in the MMTV-Neu mice (Fig. 1A), we focused our mechanistic studies in this model.
KMT2C acts as a tumour suppressor in breast cancer. A–C Tumour latency from various mouse models with or without Kmt2c. A MMTV-Neu n = 22, Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Neu (Kmt2c KO) n = 26, B Krt8CreERT2;R26tdTomato;MMTV-Myc;PIK3CAH1047R and Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Myc;PIK3CAH1047R (Kmt2c KO) n = 19 for both, C MMTV-Myc, n = 15, KMT2C KO n = 13, D H&E staining on MMTV-Neu and Kmt2c KO breast tumours, scale bar: 100 μm E Representative Kmt2c mRNA levels between control and KMT2C KO tumours presented as fold change, F Immunofluorescence against Krt5, Krt18 followed by Dapi to stain the nuclei on control and KO breast tumours
Loss of Kmt2c results in reduction of H3K4me1 and H3K4me3 levels globally
Although the individual roles of each KMT2 family member are not fully settled, it is generally considered that KMT2C and KMT2D which are closely related to the Drosophila methyltransferase Trr are involved in monomethylation of H3K4 at enhancer sites, while the closely associated trimethylation of H3K4 at transcription start sites is performed by KMT2A and KMT2B, which are homologous to Drosophila Trx [52]. To this end, we carried out ChIP-sequencing on histone 3 lysine 4-mono (H3K4me1) and tri-methylation (H3K4me3)—from MMTV-Neu (hereafter called control) and KMT2C knock out (KO) tumours. Global levels of H3K4me1 and H3K4me3 were significantly reduced in breast tumours from KO mice across all models (p = 0.027 and 0.006, respectively), as revealed by immunoblotting analysis (Fig. 2A and Supplementary Fig. 2D). We identified 9,512 active promoters and 4,451 active enhancers using the H3K27 acetylation (H3K27ac), a known mark for active enhancers (Fig. 2B) [53]. In agreement with the immunoblotting analysis, peak calling for the H3K4me3 and H3K4me1 marks showed fewer peaks in the KO compared to control mice. More specifically, we identified a total of 14,674H3K4me3 and 55,359H3K4me1 peaks for the control and 13,551H3K4me3 and 46,044 H3K4me1 peaks for the KO (Fig. 2C). By comparing the overlap of these peaks, we identified 934 unique H3K4me3 and 9,780 unique H3K4me1 in the Kmt2c KO (Fig. 2C). The signal around the TSS on the unique Kmt2c KO peaks was higher in the KO, while the signal around the TSS for the unique WT peaks was higher in the WT (Supplementary Fig. 3A), a trend also observed for the enhancer transcription start sites (eTSS) (Supplementary Fig. 3B). Furthermore, genomic peak coverage analysis showed that the decrease of H3K4me1 peak regions was corroborated by reduction of H3K4me1 genomic peak coverage in the KO tumours (10% for Neu vs 6% for KO), while the H3K4me1 mark remained essentially stable (1.8% for Neu vs 1.2% for KO).
Chromatin distribution of H3K4me3 and H3K4me1 upon KMT2C depletion. A Immunoblotting analysis of H3K4me3 and H3K4me1 on breast tumour lysates from MMTV-Neu and Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Neu (Kmt2c KO) mice. Loading normalization was performed by quantifying band intensity in ImageJ. The average of the three bands in control samples was set as 1, and the relative intensity for each antibody in Kmt2c KO was found at 0.11, 0.28 and 0.61 for H3K4me1, H3K4me3 and H3, respectively. Normalized signal intensity for H3K4me1 and HeK4me3 in Kmt2c KO was calculated at 0.18 (0.11:0.61) and 0.46 (0.28:0.61) for H3K4me1 and H3K4me3, respectively. Individual normalized H3K4me1 values are 0.27, 0.13 and 0.31 for controls, 0.023, 0.08 and 0.02 for knockout mice; for H3K4me3, the values are 1.81, 1.8 and 1.4 for controls and 0.47, 0.96 and 0.67 for knockout mice. Two-tailed t-test yields p values of 0.028 and 0.006 for H3K4me1 and H3K4me3, respectively. B Heatmap of H3K4me3 and H3K4me1 genomic peaks separated into active promoters and enhancers, respectively, by using the pattern of H3K27ac from the MMTV-Neu mouse tumors. C Venn diagrams depicting the overlap of H3K4me3 and H3K4me1 between control and Kmt2c KO breast tumours, D Genomic distribution of the unique H3K4me3 and H3K4me1 peaks for each condition
Genomic distribution of the unique peaks showed that H3K4me3 was enriched at the promoters and reduced at the distal intergenic regions in the KO versus the control. As expected, H3K4me1 peaks were reduced at the distal intergenic regions, possible enhancers, but had an enrichment at the promoters in the KO (Fig. 2D). Genomic distribution of all the peaks followed the same trend but to a lesser degree (Supplementary Fig. 3C). Since, KMT2C and KMT2D are believed to work redundantly, we looked at the levels of KMT2D and saw a reduction at the mRNA level in the KO mice. Additionally, mRNA levels of Wdr5 and Dpy30, common core subunits of all COMPASS complexes, were reduced in the KO mice (Supplementary Table 2). Whether this represents an en bloc regulation of the genes encoding COMPASS complex components or is merely a by-product of global chromatin remodelling and condensation is a question which warrants further investigation.
Knock out of Kmt2c leads to disruption of epithelial cell differentiation and an imbalance in ERK/MAPK signalling
Since both H3K4me3 and H3K4me1 are histone marks that are associated with active transcription [54], we wanted to assess the potential changes of Kmt2c KO in gene expression. Differential gene expression analysis by RNA-sequencing (RNA-seq) revealed 1562 genes upregulated and 1431 genes downregulated significantly (adjusted p < 0.05) in the KO as compared to controls (Supplementary Table 2). Pathway analysis using the gene set enrichment analysis (GSEA) and Metascape [28] showed upregulation of genes that promote epithelial to mesenchymal transition (EMT) and downregulation of genes that maintain epithelial differentiation in the KO breast tumours (Fig. 3A and B). We also observed that epithelial cells from KO tumours had more elongated morphology as compared to controls (Supplementary Fig. 4A). The EMT phenotype was corroborated by the higher expression of the EMT-activating transcription factors (Snai1, Snai2 and Zeb1) and of the cadherin member CDH2, whose expression is another hallmark of EMT [55, 56]. In addition, various types of collagen proteins (Col2a1, Col4a1, Col4a2, Col16a1, Col11a1), which disrupt normal epithelial cell differentiation and remodel the extracellular matrix [57, 58], were also upregulated, further promoting EMT (Fig. 3C and Supplementary Fig. 4B). The EMT transcription factors, Snai1 and Zeb1, were also upregulated in the MMTV-Myc and MMTV-Myc;ex20 Kmt2c KO mouse models (Supplementary Fig. 4C), further supporting the EMT phenotype in KMT2C KO background. Gene annotation and pathway analysis (Supplementary Fig. 5A and B) on the unique identified H3K4me3 peaks from KO tumours corroborated these findings by showing enrichment of Twist1, Twist2 (Supplementary Table 3) and extracellular matrix organization. On the contrary, similar analysis on unique H3K4me3 peaks in the control tumours showed enrichment of epithelial cell differentiation. Furthermore, motif analysis on unique peaks revealed a gain on motifs associated with Nanos1 and C2h2-type proteins. Nanos1 is a transcription factor that depicts inverse correlation with E-cadherin in literature [59]. C2h2-type proteins, include the EMT transcription factors Snai1, Snai2, Zeb1 and Zeb2. Additionally, motif analysis revealed loss of motifs bound by ESE1, an epithelium-specific ETS transcription factor 1, member of the Ets transcription factor family, which negatively regulates ZEBs [60]. Moreover, ESE1, EGR1 and C/Ebpb, which are associated with luminal cell fate commitment are enriched in MMTV-Neu tumors [61,62,63] (Supplementary Fig. 6).
KMT2C loss leads to an epithelial-to-mesenchymal (EMT) phenotype and an imbalance in ERK/MAPK signalling. A Enrichment of the EMT pathway in the Kmt2c KO tumours based on hallmark pathways with GSEA analysis, B Downregulated pathways in the Kmt2c KO as depicted by Metascape analysis, C Heatmap depicting the differential gene expression of genes associated with EMT and one representative qPCR on mRNA levels from Cdh1, Snai1 and Zeb1 from MMTV-Neu and Kmt2c KO breast tumours presented as fold change, D–F Immunohistochemical and immunoblotting analysis on phospho-ERK1/2 on various mouse models, G immunoblotting analysis on various signal transduction proteins. The GAPDH loading control from (G) was the same as in Fig. 2A
To investigate the potential changes in signal transduction pathways, we performed staining and immunoblotting analysis on various markers of core signalling cascades and observed increased levels of phospho-ERK1/2 in the KO mice (Fig. 3D–F). Interestingly, both ERK1 and ERK2 have been implicated in promoting EMT in cancer [64, 65].
In support, phospho-p38, another mitogen-activated protein kinase (MAPK), was significantly reduced in the KO tumours as compared to controls (Fig. 3G). Reportedly, p38 has been shown to maintain E-cadherin expression in a HER2 + mouse model [66] and to have opposing activity to ERK signalling in chondrogenesis [67] and neural development [68] along with a suppressive effect on RAS-mediated oncogenesis [69]. Nonetheless, we found no changes in the levels of H-Ras and K-Ras or of phospho-c-Jun and Jnk in our Kmt2c KO model (Fig. 3G).
Interestingly, Kmt2c KO tumours show higher mRNA and protein levels of the basal cytokeratin Krt6a, which is implicated in EMT [70] (Supplementary Table 2 and Fig. 3G). Moreover, Krt6a expression is associated with basal phenotypes [71], a subtype that has poor clinical outcomes in breast cancer [72, 73]. In line with the Krt6a upregulation, the basal marker Trp63 was also upregulated, while the expression levels of the luminal keratins Krt8/18 were lower in the KO tumours (Supplementary Fig. 5C). Finally, the abundance of oestrogen receptor (ER) was unchanged whereas KDM6A/UTX, a known specific component of the MLL3 COMPASS-like complex, was reduced in the Kmt2c KO (Fig. 3G).
KMT2C KO breast tumours present mitochondrial dysregulation and increased ROS levels
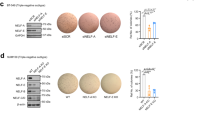
Our RNA sequencing analysis revealed that expression of ATPase subunits, including members involved in mitochondrial function and OXPHOS, was reduced in the Kmt2c KO (Fig. 4A). In agreement with this, our ChIP-seq data showed a remarkable reduction of H3K4me3 at the promoters for many of these genes in the Kmt2c KO (Fig. 4B, C and D). Further analysis by qPCR and Western blot of NDUFS3 (complex I) and ATP5a (complex V) showed reduced mRNA (Fig. 4E) and protein (Fig. 4F) levels in MMTV-Neu/KO tumours. Reduction of expression of these subunits was also observed in the MMTV-Myc and MMTV-Myc;ex20 KO mouse models (Fig. 4G and H).
Depletion of Kmt2c leads to mitochondrial dysfunction and upregulation of ROS levels. Α Differential gene expression on various ATPases also involved in mitochondrial function and oxidative phosphorylation (OXPHOS) from WT and Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Neu (Kmt2c KO) breast tumours, B–D H3K4me3 peak pattern on various genes related to the OXPHOS in WT and KMT2C KO breast tumours, E mRNA levels presented as fold change and F protein levels of ATP5a and NDUFS3 from breast tumours, mRNA expression of various OXPHOS-related genes in G MMTV-Myc;ex20 and H MMTV-Myc control and KO mice, I Measurement of mitochondrial respiration and ROS levels from MMTV-Neu and Kmt2c KO breast tumours
These findings prompted us to measure the mitochondrial respiration rate using freshly isolated mitochondria [44] from breast tumours of control and Kmt2c KO mice. Using a Clark-type oxygen electrode, we measured the state 3 (indicates rate of ATP production) and state 4 (indicates coupling) of mitochondrial respiration and calculated the respiratory control ratio (RCR) as the ratio of state 3 to state 4 (ST3:ST4), as previously described [44]. We found that the ST3:ST4 ratio was significantly reduced (p < 0.0001) in the Kmt2c KO tumours (Fig. 4J-left), which is an indication of mitochondrial dysfunction and damage [74]. Given that mitochondria are major sources of reactive oxygen species (ROS) production as by-product of oxidative phosphorylation [75], we assessed the levels of ROS in our samples and observed a significant increase (p < 0.0001) in ROS in the Kmt2c KO tumours, further supporting mitochondrial dysfunction, but also a likely deregulation of cellular antioxidant responses (Fig. 4J-right). In support, many genes with antioxidant activity [76], such as Gstm2, Gstt1 and Gsta3, were downregulated in the Kmt2c KO (Fig. 4G, H sand Supplementary Table 2).
Loss of Kmt2c is associated with better response to therapy
To understand whether Kmt2c KO tumours grow faster than the controls, we measured tumour growth rate once the tumours were palpable by using a calliper (Fig. 5A). We observed minimal differences in tumour growth rate and Ki67 staining (Fig. 5A-right) between the control and the Kmt2c KO tumours (Fig. 5A). Next, we investigated whether Kmt2c KO tumours were responsive to lapatinib and found that indeed they were (Fig. 5B). In fact, KMT2C KO mice were more sensitive to lapatinib than the MMTV-Neu control ones (p = 0.019 at day 19), followed by a marked reduction in Ki67 staining (p = 0.067) (Fig. 5B-right). This enhanced oncogene addiction most likely associates with the higher phospho-ERK1/2 levels presented earlier, indicating an increased dependence on the ERK1/2 signalling axis. Because Erbb2/Neu in our mouse model is expressed under the control of the MMTV LTR, and not the endogenous locus, it is unlikely that changes in the expression levels of the Erbb2/Neu oncogene are responsible for this. The downregulation of Met and Erbb3 in the KO tumours (Supplementary Table 2), on the other hand, might contribute to this phenotype. Immunoblotting analysis from mice treated with lapatinib (Fig. 5C) or the MEK inhibitor trametinib (Fig. 5D–E), showed reduction of phospho-ERK1/2 and Snai1 levels, further indicating that EMT is at least in part the result of increased ERK1/2 signalling.
KMT2C KO mice are more responsive to lapatinib than their WT counterparts. A Tumour growth of untreated mice accompanied with Ki67 immunohistochemistry staining and concomitant quantification (unpaired t-test was applied), B Tumour growth of lapatinib-treated mice (200 mg/kg/day) along with Ki67 staining and quantification of the Ki67 positive nuclei (unpaired t-test was applied). Multiple t-test was applied in the lapatinib-treated samples (*p ≤ 0.05, **p ≤ 0.01), The y axis of the graphs corresponds to the average values of the fold changes of the tumour volumes for each animal, C Immunoblotting analysis of pERK1/2, total ERK and Snai1 on untreated and treated breast tumour lysates. D Immunoblotting analysis and (E) qPCR on trametinib-treated (1 mg/kg/day) KO mice
To assess the clinical significance of KMT2C, we looked at the relation of KMT2C gene expression and the long-term outcome of patients on adjuvant therapy using KM plotter [43]. Low expression of KMT2C (probe 232940_s_at) was strongly associated with better RFS on breast cancer patients treated with any therapy (Fig. 6A–D), in this way supporting our mouse data. Combined with literature reports implicating KMT2C in chemotherapy response in hemopoeitic malignancies [77] and solid tumors [20], our findings uncover a broader role for KMT2C in modulating therapeutic response in neoplasia.
Clinical relevance of KMT2C high or low gene expression in various breast cancer subtypes. RFS Data A presented from ER positive patients treated with any therapy, B ER positive HER2 negative patients treated with any therapy, C HER2-positive patients treated with any therapy, D ER negative patients treated with any therapy. Red, patients with higher gene expression; black, patients with lower gene expression
Discussion
It has become apparent that apart from mutations on PIK3CA and TP53 loci, which dominate the mutational landscape of breast cancer samples [6, 7], there is a handful of genes which are recurrently mutated in at least 10% of the cases [6]. One such gene is KMT2C, an epigenetic writer of H3K4 methylation. Despite the recent interest in epigenetic modifiers as promising druggable targets in cancer [78], our knowledge on the role of KMT2C in breast cancer is still limited.
In this study, we sought to investigate the role of KMT2C in breast tumorigenesis and response to therapy using genetically engineered mouse models. In agreement with several lines of evidence from various cancers [79], our study demonstrates that KMT2C acts invariably as a tumour suppressor in breast cancer, since tumours that lack KMT2C became palpable significantly earlier than the WT counterparts. Global levels of H3K4 mono and tri-methylation were reduced in the KO mice. While reduction of H3K4me1 levels is expected, the reduction in H3K4me3 abundance is a paradox, given that this modification is carried out by KMT2A/Menin [12]. This could be explained by the reduction of Wdr5 and Dpy30 mRNA levels seen in the Kmt2c KO mice, which are known common core subunits of all mammalian COMPASS complexes [80].
While H3K4me3 is associated with active [81] or poised promoters [82, 83], H3K4me1 associates with active or primed enhancers [84] and more recently it was found at the promoters of a subset of silenced genes in a cell-type specific manner [12, 85]. To determine if there is re-distribution of the H3K4 methylation pattern upon KMT2C depletion, we looked at the global genomic distribution of methylation by ChIP-sequencing on H3K4 mono and tri-methylation and observed more peaks at the promoters and fewer peaks at the enhancers for both modifications. The reduction of H3K4me1 at enhancer sites is expected upon KMT2C depletion, but the increase of H3K4me1 at the promoters is a paradox along with the increase in the H3K4me3. Since both H3K4me3 and H3K4me1 are associated with either active or poised states of chromatin, our data cannot exclude the possibility that loss of KMT2C leads to perturbation in the methylation states of the chromatin, which then promotes transition to more undifferentiated cellular states and tumour growth. The epigenetic reshaping observed in the Kmt2c KO tumours might not be necessarily a direct result of Kmt2c loss but could reflect an adaptation process during tumour evolution which is driven by tumour fitness.
To determine if there is dysregulation of transcription upon KMT2C depletion, we conducted gene expression profiling (RNA-seq). Our data revealed that Kmt2c KO breast tumours lose their epithelial identity and present an epithelial-to-mesenchymal transition (EMT) phenotype. Although the molecular mechanism is not clear, EMT is associated with downregulation of the transcription factor Grhl2, a known suppressor of EMT in breast cancer [62] and previously found to be enriched in KMT2C binding events [16], and upregulation of EMT-promoting transcription factors such as Snai1, Snai2 and Zeb1 [86,87,88] as well as EMT target genes such as Cdh2 and various collagens proteins. Pathway and motif analysis on our ChIP-seq data corroborated these findings. These data are reinforced by previous studies conducted in gastric adenocarcinoma [89] and breast [90, 91], where they also observed an association of Kmt2c loss with EMT and more aggressive tumour phenotypes. Intriguingly, EMT has been associated with less differentiated stem cells [92]. In concordance with the transition to more undifferentiated states, we found an alteration in the expression of cytokeratins, which serve as markers for the identification of cellular subtypes. Specifically, the mRNA levels of the luminal-specific keratins, Krt8 and Krt18, were reduced in the Kmt2c KO, while the levels of the basal-specific keratin, Krt6α were increased. Finally, the levels of Trp63, an important marker of basal identity, stemness and direct transcriptional regulator of basal keratins [93, 94], were increased in the Kmt2c KO tumours.
In light of evidence from several studies on the fundamental role of MAPK signalling pathways in cell transformation and cancer [95,96,97], our results demonstrate a higher abundance of activated ERK1/2 in the KO and a loss of phosphorylated p38. The hyperactivation of ERK in our KO mice could explain the shorter tumour latency, since it is well established that activated ERK is sufficient to initiate cellular transformation [95, 96]. Moreover, ERK1/2 has been shown to induce EMT in various cancer cells acting as the converging hub of upstream activating signals [64, 65, 98, 99]. The finding of phospho-p38 downregulation in our Kmt2c KO samples is interesting, since p38 maintains E-cadherin expression [100] and functions as a suppressor of EMT and early dissemination in a HER2 + mouse model [66]. In a recent study, Kmt2c−/− mammary outgrowths from organoids showed enrichment in stem-cell-basal markers (CD49hi, CD61+) and activation of the PI3K pathway [101], an observation we did not see in our mouse tumours.
Furthermore, we discovered that KM2TC regulates the transcription of many subunits related to mitochondrial function and OXPHOS. Upon KMT2C depletion, we observed a decrease in both the H3K4me3 signal and the mRNA expression for many of these genes (Ndufs3, Atp5a, Atp6v1b1). Consistently, we saw a reduction in the mitochondria respiration rate along with increased ROS levels. This finding is reinforced by a previous work on KMT2D, in which a positive regulation of the glutathione detoxification pathway was observed by KMT2D [102]. Additionally, it was recently found that loss of KMT2D leads to deregulation of mitochondrial respiration and enhanced generation of ROS levels [103]. There is a large body of evidence that cancer cells often dysregulate mitochondria and show elevated levels of ROS [104]. While ROS can damage the cell, if they are not properly sequestered in normal physiological conditions, there is evidence that ROS can promote signal transduction in many cancer types [104]. For example, ROS triggered a marked activation of ERK in human glioma cells [105] and mediated degradation of MKP3, a negative regulator of ERK1/2, in ovarian cancer cells [106]. Thus, we cannot exclude the possibility that high ROS levels contribute to the increased phospho-ERK1/2 signal in Kmt2c KO tumours.
Finally, our data indicated that Kmt2c KO tumours, showed better response to lapatinib. This finding may have therapeutic implications in the clinic, by providing evidence on advantageous treatments to patients carrying Kmt2c loss of function mutations. A possible explanation of this finding could be that the genetic ablation of Kmt2c does not allow an otherwise expected re-wiring and increased transcription of kinases involved with lapatinib resistance [107,108,109]. In agreement with this hypothesis, the transcripts of MET and ERBB3 kinases are decreased in the Kmt2c KO tumours compared to the controls (Supplementary Table 2). In silico analysis from publicly available clinical samples corroborated these results, showing that KMT2C low expression is associated with better long-term outcome.
Taken together, our data demonstrate that KMT2C acts as a strong tumour suppressor in breast cancer. Kmt2c KO mice develop breast cancer earlier than their WT counterparts, possibly through the hyperactivation of ERK1/2 and the concomitant suppression of phospho-p38 followed by an activation of the EMT transcriptional program. Kmt2c loss leads to downregulation of genes involved in OXPHOS and antioxidant activity, which in turn promotes mitochondrial dysfunction and ROS accumulation, respectively. Importantly, genetic ablation of KMT2C sensitised mice to lapatinib treatment. In this sense, it would be interesting to investigate the value of Kmt2c status as a biomarker for treatment with anti-HER2 antibodies and/or small molecule inhibitors.
Availability of data and materials
All the raw sequencing data has been submitted to NCBI SRA portal with the accession ID PRJNA787445.
References
Torre LA, Siegel RL, Ward EM, Jemal A (2015) Global cancer incidence and mortality rates and trends--an update. Cancer Epidemiol Biomark Prevent. 25:1–12. http://cebp.aacrjournals.org/cgi/doi/10.1158/1055-9965.EPI-15-0578
Blows FM, Driver KE, Schmidt MK, Broeks A, Van LFE, Wesseling J et al (2010) Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10, 159 cases from 12 studies. PLoS Med 7:e1000279
Gellert P, Segal CV, Gao Q, López-Knowles E, Martin L, Dodson A et al (2016) Impact of mutational profiles on response of primary oestrogen receptor-positive breast cancers to oestrogen deprivation. Nat Commun. 7:13294. http://www.nature.com/articles/ncomms13294
Nik-Zainal S, Van Loo P, Wedge DC, Alexandrov LB, Greenman CD, Lau KW et al (2012) The life history of 21 breast cancers. Cell 149:994–1007
Nik-Zainal S, Davies H, Staaf J, Ramakrishna M, Glodzik D, Zou X et al (2016) Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534:1–20. http://www.nature.com/doifinder/10.1038/nature17676
Pereira B, Chin S-F, Rueda OM, Vollan H-KM, Provenzano E, Bardwell HA et al (2016) The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat Commun 7:11479. http://www.nature.com/doifinder/10.1038/ncomms11479
Koboldt DC, Fulton RS, McLellan MD, Schmidt H, Kalicki-Veizer J, McMichael JF et al (2012) Comprehensive molecular portraits of human breast tumours. Nature 490:61–70
Wang X, Fu L, Li X, Wu X, Zhu Z, Fu L et al (2011) Somatic mutations of the mixed-lineage leukemia 3 ( MLL3) gene in primary breast cancers. Pathol Oncol Res 3:429–433
Herz H, Mohan M, Garruss AS, Liang K, Takahashi Y, Mickey K et al (2012) Enhancer-associated H3K4 monomethylation by Trithorax-related, the Drosophila homolog of mammalian Mll3/Mll4. Genes Dev 2:2604–2620
Kaikkonen MU, Spann NJ, Heinz S, Romanoski CE, Allison KA, Stender JD et al (2013) Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol Cell 51:310–25. https://linkinghub.elsevier.com/retrieve/pii/S1097276513005091
Hu D, Gao X, Morgan MA, Herz H, Smith ER, Shilatifard A (2013) The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol Cell Biol 33:4745–4754
Cheng J, Blum R, Bowman C, Hu D, Shilatifard A, Shen S et al (2014) A role for H3K4 monomethylation in gene repression and partitioning of chromatin readers. Mol Cell 53:979–992. http://www.ncbi.nlm.nih.gov/pubmed/24656132
Miller T, Krogan NJ, Dover J, Tempst P, Johnston M, Greenblatt JF et al (2001) COMPASS: a complex of proteins associated with a trithorax-related SET domain protein. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.231473398
You JS, Jones PA (2012) Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell 22:9–20
Chen C, Liu Y, Rappaport AR, Kitzing T, Schultz N, Shroff AS et al (2014) MLL3 is a haploinsufficient 7q tumor suppressor in acute myeloid leukemia. Cancer Cell 25:652–665
Jozwik KM, Chernukhin I, Serandour AA, Nagarajan S, Carroll JS (2016) FOXA1 directs H3K4 monomethylation at enhancers via recruitment of the methyltransferase MLL3. Cell Rep 17:2715–2723. https://doi.org/10.1016/j.celrep.2016.11.028
Gala K, Li Q, Sinha A, Razavi P, Dorso M, Sanchez-Vega F et al (2018) KMT2C mediates the estrogen dependence of breast cancer through regulation of ERα enhancer function. Oncogene 37:4692–4710. https://doi.org/10.1038/s41388-018-0273-5
Stauffer KM, Elion DL, Cook RS, Stricker T (2021) MLL3 is a de novo cause of endocrine therapy resistance. Cancer Med 1–20. https://onlinelibrary.wiley.com/doi/10.1002/cam4.4285
Ray Chaudhuri A, Callen E, Ding X, Gogola E, Duarte AA, Lee J et al (2016) Replication fork stability confers chemoresistance in BRCA-deficient cells. Nature 535:382–387. https://doi.org/10.1038/nature18325
Rampias T, Karagiannis D, Avgeris M, Polyzos A, Kokkalis A, Kanaki Z et al (2019) The lysine‐specific methyltransferase KMT2C/MLL3 regulates DNA repair components in cancer. EMBO Reports 20:1–20. https://onlinelibrary.wiley.com/doi/10.15252/embr.201846821
Chang A, Liu L, Ashby JM, Wu D, Chen Y, O’Neill SS et al (2021) Recruitment of KMT2C/MLL3 to DNA damage sites mediates DNA Damage responses and regulates PARP inhibitor sensitivity in cancer. Cancer Res 81:3358–3373. http://cancerres.aacrjournals.org/lookup/doi/10.1158/0008-5472.CAN-21-0688
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930
Ewels P, Magnusson M, Lundin S, Kaeller M (2016) MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32:3047–3048
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21
Mootha VK, Lindgren CM, Eriksson K-F, Subramanian A, Sihag S, Lehar J et al (2003) PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273. http://www.nature.com/articles/ng1180
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Zhou Y, Zhou B, Pache L, Chang M, Benner C, Chanda SK et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1–10. https://doi.org/10.1038/s41467-019-09234-6
Schmidt D, Wilson MD, Spyrou C, Brown GD, Hadfield J, Odom DT (2009) ChIP-seq: using high-throughput sequencing to discover protein-DNA interactions. Methods (San Diego, Calif) [cited 2014 Mar 26];48:240–248. http://www.ncbi.nlm.nih.gov/pubmed/19275939
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359
Tarasov A, Vilella AJ, Cuppen E, Nijman IJ, Prins P (2015) Sambamba: fast processing of NGS alignment formats. Bioinformatics 31:2032–2034
Ramírez F, Dündar F, Diehl S, Grüning BA, Manke T (2014) DeepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res 42:187–191
Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO et al (2021) Twelve years of SAMtools and BCFtools. GigaScience 10:1–4
Stovner EB, Sætrom P (2019) Epic2 efficiently finds diffuse domains in ChIP-seq data. Bioinformatics 35:4392–4393
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics (Oxford, England) [cited 2014 Jul 9];26:841–842. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2832824&tool=pmcentrez&rendertype=abstract
Khan A, Mathelier A (2017) Intervene: a tool for intersection and visualization of multiple gene or genomic region sets. BMC Bioinformatics 18:1–8
Hahne F, Ivanek R (2016) Visualizing Genomic Data Using Gviz and Bioconductor. pp 335–351. http://springerlink.fh-diploma.de/10.1007/978-1-4939-3578-9_16
Yu G, Wang LG, He QY (2015) ChIP seeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31:2382–2383
Ye T, Krebs AR, Choukrallah M-A, Keime C, Plewniak F, Davidson I et al (2011) seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res 39:e35–e35. https://academic.oup.com/nar/article-lookup/doi/10.1093/nar/gkq1287
Yu G, Wang LG, Han Y, He QY (2012) ClusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287
Herrmann C, Van De Sande B, Potier D, Aerts S (2012) i-cisTarget: an integrative genomics method for the prediction of regulatory features and cis-regulatory modules. Nucleic Acids Res 40:1–17
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P et al (2010) Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38:576–589
Győrffy B (2021) Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput Struct Biotechnol J 19:4101–4109
Tsakiri EN, Gumeni S, Iliaki KK, Benaki D, Vougas K, Sykiotis GP et al (2019) Hyperactivation of Nrf2 increases stress tolerance at the cost of aging acceleration due to metabolic deregulation. Aging Cell. https://doi.org/10.1111/acel.12845
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data: figure 1. Cancer Discov 2:401–404. http://cancerdiscovery.aacrjournals.org/lookup/doi/10.1158/2159-8290.CD-12-0095
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:1–20. http://www.ncbi.nlm.nih.gov/pubmed/23550210%0A. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4160307
Van Keymeulen A, Rocha AS, Ousset M, Beck B, Bouvencourt G, Rock J et al (2011) Distinct stem cells contribute to mammary gland development and maintenance. Nature 479:189–193. http://www.ncbi.nlm.nih.gov/pubmed/21983963
Hollern DP, Swiatnicki MR, Andrechek ER (2018) Histological subtypes of mouse mammary tumors reveal conserved relationships to human cancers. PLoS Genet. https://doi.org/10.1371/journal.pgen.1007135
Dankort D, Maslikowski B, Warner N, Kanno N, Kim H, Wang Z et al (2001) Grb2 and Shc adapter proteins play distinct roles in Neu (ErbB-2)-induced mammary tumorigenesis: implications for human breast cancer. Mol Cell Biol 21:1540–1551. http://mcb.asm.org/cgi/doi/10.1128/MCB.21.5.1540-1551.2001
Stewart TA, Pattengale PK, Leder P (1984) Spontaneous mammary adenocarcinomas transgenic mice that carry and express MTV/myc fusion genes. Cell 38:627–637
Madisen L, Zwingman TA, Sunkin SM, Oh SW, Zariwala HA, Gu H et al (2010) A robust and high-throughput Cre Repooting and characterization. Nat Neurosci 13:133–140
Meeks JJ, Shilatifard A (2017) Multiple roles for the mll/compass family in the epigenetic regulation of gene expression and in cancer. Ann Rev Cancer Biol 1:425–446
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci USA 107:21931–21936
Kouzarides T (2002) Histone methylation in transcriptional control. Curr Opin Genet Dev 12:198–209
Wang M, Ren D, Guo W, Huang S, Wang Z (2016) N-cadherin promotes epithelial-mesenchymal transition and cancer stem cell-like traits via ErbB signaling in prostate cancer cells. Int J Oncol 48:595–606
Hulit J, Suyama K, Chung S, Keren R, Agiostratidou G, Shan W et al (2007) N-cadherin signaling potentiates mammary tumor metastasis via enhanced extracellular signal-regulated kinase activation. Cancer Res 67:3106–3117
Neira RE, Salazar EP (2012) Native type IV collagen induces an epithelial to mesenchymal transition-like process in mammary epithelial cells MCF10A. Int J Biochem Cell Biol 44:2194–2203. https://doi.org/10.1016/j.biocel.2012.08.018
Caroline B, Chou J, Werb Z (2014) Remodelling the extracellular matrix in development and disease. Nat Rev Mol Cell Biol 15:786–801
De Keuckelaere E, Hulpiau P, Saeys Y, Berx G, van Roy F (2018) Nanos genes and their role in development and beyond. Cell Mol Life Sci 75:1929–1946. https://doi.org/10.1007/s00018-018-2766-3
Sinh ND, Endo K, Miyazawa K, Saitoh M (2017) Ets1 and ESE1 reciprocally regulate expression of ZEB1/ZEB2, dependent on ERK1/2 activity, in breast cancer cells. Cancer Sci 108:952–960. https://onlinelibrary.wiley.com/doi/10.1111/cas.13214
Spike AJ, Rosen JM (2020) C/EBPß isoform specific gene regulation: it’s a lot more complicated than you think! J Mammary Gland Biol Neoplasia 6:1–12
Cieply B, Farris J, Denvir J, Ford HL, Frisch SM (2013) Epithelial—mesenchymal transition and tumor suppression are controlled by a reciprocal feedback loop between ZEB1 and grainyhead-like-2. Cancer Res 73:6299–6310
Kim H-R, Kim YS, Yoon JA, Lyu SW, Shin H, Lim HJ et al (2014) Egr1 is rapidly and transiently induced by estrogen and bisphenol A via activation of nuclear estrogen receptor-dependent ERK1/2 pathway in the uterus. Reprod Toxicol 50:60–67. https://linkinghub.elsevier.com/retrieve/pii/S0890623814002639
Shin S, Dimitri CA, Yoon S-O, William D, Blenis J (2010) ERK2 but Not ERK1 induces epithelial-to-mesenchymal transformation. Mol Cell 38:114–127. https://doi.org/10.1016/j.molcel.2010.02.020
Buonato MJ, Lazzara JM (2015) ERK1/2 Blockade prevents epithelial-mesenchymal transition in lung cancer cells and promotes their sensitivity to Egfr inhibition. Cancer Res 74:309–319
Harper KL, Sosa MS, Entenberg D, Hosseini H, Cheung JF, Nobre R et al (2016) Mechanism of early dissemination and metastasis in Her2+ mammary cancer. Nature 540:588–592. http://www.nature.com/articles/nature20609
Oh CD, Chang SH, Yoon YM, Lee SJ, Lee YS, Kang SS et al (2000) Opposing role of mitogen-activated protein kinase subtypes, Erk-1/2 and p38, in the regulation of chondrogenesis of mesenchymes. J Biol Chem 275:5613–5619. https://doi.org/10.1074/jbc.275.8.5613
Xia Z, Dickens M, Raingeaud J, Davis RJ, Michael E, Xia Z et al (2016) Opposing effects of ERK and JNK-p38 MAP kinases on apoptosis. Science 270:1326–1331
Chen G, Hitomi M, Han J, Stacey DW (2000) The p38 pathway provides negative feedback for Ras proliferative signaling*. J Biol Chem 275:38973–38980. https://doi.org/10.1074/jbc.M002856200
Yang B, Zhang W, Zhang M, Wang X, Peng S, Zhang R (2020) KRT6A promotes EMT and cancer stem cell transformation in lung adenocarcinoma. Technol Cancer Res Treat 19:1–8
Perou CM, Sùrlie T, Eisen MB, Van De RM, Jeffrey SS, Rees CA et al (2000) Molecular portraits of human breast tumours. Nature 533:747–752
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. In: Proceedings of the national academy of sciences of the United States of America 98:10869–10874. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=11553815%5Cn. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC58566/pdf/pq010869.pdf
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A et al (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. In: Proceedings of the national academy of sciences of the United States of America 100:8418–8423. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=166244&tool=pmcentrez&rendertype=abstract
Silva AM, Oliveira PJ (2012) Evaluation of respiration with clark type electrode in isolated mitochondria and permeabilized animal cells. pp 7–24. http://springerlink.fh-diploma.de/10.1007/978-1-61779-382-0_2
Kumar B, Koul S, Khandrika L, Meacham RB, Koul HK (2008) Oxidative stress is inherent in prostate cancer cells and is required for aggressive phenotype. Can Res 68:1777–1785
Ye ZW, Zhang J, Townsend DM, Tew KD (2015) Oxidative stress, redox regulation and diseases of cellular differentiation. Biochim Biophys Acta Gen Subj 1850:1607–1621. https://doi.org/10.1016/j.bbagen.2014.11.010
Chen R, Okeyo-Owuor T, Patel RM, Casey EB, Cluster AS, Yang W et al (2021) Kmt2c mutations enhance HSC self-renewal capacity and convey a selective advantage after chemotherapy. Cell Rep 34:108751. https://doi.org/10.1016/j.celrep.2021.108751
Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O et al (2010) Selective inhibition of BET bromodomains. Nature 468:1067–1073
Rao RC, Dou Y (2015) Hijacked in cancer: the KMT2 (MLL) family of methyltransferases. Nature 15:334–346. https://doi.org/10.1038/nrc3929
Vermeulen M, Timmers HM (2010) Grasping trimethylation of histone H3 at lysine 4. Epigenomics 2:395–406. https://www.futuremedicine.com/doi/10.2217/epi.10.11
Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NCT et al (2002) Active genes are tri-methylated at K4 of histone H3. Nature 419:407–411
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J et al (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125:315–326
Lesch BJ, Page DC (2014) Poised chromatin in the mammalian germ line. Development (Cambridge) 141:3619–3626
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD et al (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39:311–318
Bae S, Lesch BJ (2020) H3K4me1 distribution predicts transcription state and poising at promoters. Front Cell Dev Biol 8:1–11
Batlle E, Sancho E, Francí C, Domínguez D, Monfar M, Baulida J et al (2000) The transcription factor Snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol 2:84–89. http://www.nature.com/articles/ncb0200_84
Cano A, Pérez-Moreno MA, Rodrigo I, Locascio A, Blanco MJ, del Barrio MG et al (2000) The transcription factor Snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat Cell Biol 2:76–83. http://www.nature.com/articles/ncb0200_76
Browne G, Sayan AE, Tulchinsky E (2010) ZEB proteins link cell motility with cell cycle control and cell survival in cancer. Cell Cycle 9:886–891
Cho S-J, Yoon C, Lee JH, Chang KK, Lin J, Kim Y-H et al (2018) KMT2C mutations in diffuse-type gastric adenocarcinoma promote epithelial-to-mesenchymal transition. Clin Cancer Res 24:6556–69. http://clincancerres.aacrjournals.org/lookup/doi/10.1158/1078-0432.CCR-17-1679
Wang L, Zhao Z, Ozark PA, Fantini D, Marshall SA, Rendleman EJ et al (2018) Resetting the epigenetic balance of Polycomb and COMPASS function at enhancers for cancer therapy. Nat Med 24:758–769. https://doi.org/10.1038/s41591-018-0034-6
Fagan RJ, Dingwall AK (2019) COMPASS ascending: emerging clues regarding the roles of MLL3/KMT2C and MLL2/KMT2D proteins in cancer. Cancer Lett. https://doi.org/10.1016/j.canlet.2019.05.024
Mani SA, Guo W, Liao M, Eaton EN, Ayyanan A, Zhou AY et al (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133:704–715
Gatti V, Bongiorno-Borbone L, Fierro C, Annicchiarico-Petruzzelli M, Melino G, Peschiaroli A (2019) p63 at the crossroads between stemness and metastasis in breast cancer. Int J Mol Sci 20:2683. https://www.mdpi.com/1422-0067/20/11/2683
Choi W, Porten S, Kim S, Willis D, Plimack ER, Hoffman-censits J et al (2014) Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 25:152–165. https://doi.org/10.1016/j.ccr.2014.01.009
Cowley S, Paterson H, Kemp P, Marshall CJ (1994) Activation of MAP kinase kinase is necessary and sufficient for PC12 differentiation and for transformation of NIH 3T3 cells. Cell 77:841–852. https://linkinghub.elsevier.com/retrieve/pii/0092867494901333
Mansour SJ, Matten WT, Hermann AS, Candia JM, Rong S, Fukasawa K et al (1994) Transformation of mammalian cells by constitutively active MAP kinase kinase. Science 265:966–970. https://www.science.org/doi/10.1126/science.8052857
Zhang W, Liu HT (2002) MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Res 12:9–18. http://www.nature.com/articles/7290105
Strippoli R, Benedicto I, Lozano MLP, Cerezo A, López-Cabrera M, Del Pozo MA (2008) Epithelial-to-mesenchymal transition of peritoneal mesothelial cells is regulated by an ERK/NF-κB/Snail1 pathway. DMM Dis Models Mech 1:264–274
Lamouille S, Xu J, Derynck R (2014) Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 15:178–196. https://doi.org/10.1038/nrm3758
Strippoli R, Benedicto I, Foronda M, Perez-Lozano ML, Sánchez-Perales S, López-Cabrera M et al (2010) p38 maintains E-cadherin expression by modulating TAK1-NF-κB during epithelial-to-mesenchymal transition. J Cell Sci 123:4321–4331
Zhang Z, Christin JR, Wang C, Ge K, Oktay MH, Guo W (2016) Mammary-stem-cell-based somatic mouse models reveal breast cancer drivers causing cell fate dysregulation. Cell Rep 16:3146–3156. https://doi.org/10.1016/j.celrep.2016.08.048
Santos MA, Faryabi RB, Ergen AV, Day AM, Malhowski A, Canela A et al (2014) DNA-damage-induced differentiation of leukaemic cells as an anti-cancer barrier. Nature 514:107–111
Pacelli C, Adipietro I, Malerba N, Squeo GM, Piccoli C, Amoresano A et al (2020) Loss of function of the gene encoding the histone methyltransferase KMT2D leads to deregulation of mitochondrial respiration. Cells 9:1–20
Liou G-Y, Storz P (2010) Reactive oxygen species in cancer. Free Radic Res 44:479–496. http://www.tandfonline.com/doi/full/10.3109/10715761003667554
Lee WC, Choi CH, Cha SH, Oh HL, Kim YK (2005) Role of ERK in hydrogen peroxide-induced cell death of human glioma cells. Neurochem Res 30:263–270
Chan DW, Liu VWS, Tsao GSW, Yao KM, Furukawa T, Chan KKL et al (2008) Loss of MKP3 mediated by oxidative stress enhances tumorigenicity and chemoresistance of ovarian cancer cells. Carcinogenesis 29:1742–1750
Stuhlmiller TJ, Miller SM, Zawistowski JS, Nakamura K, Beltran AS, Duncan JS et al (2015) Inhibition of lapatinib-induced kinome reprogramming in ERBB2-positive breast cancer by targeting BET family bromodomains. Cell Rep 11:390–404. https://doi.org/10.1016/j.celrep.2015.03.037
Garrett JT, Olivares MG, Rinehart C, Granja-Ingram ND, Sánchez V, Chakrabarty A et al (2011) Transcriptional and posttranslational up-regulation of HER3 (ErbB3) compensates for inhibition of the HER2 tyrosine kinase. In: Proceedings of the national academy of sciences of the United States of America 108:5021–5026. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=3064360&tool=pmcentrez&rendertype=abstract
Amin DN, Sergina N, Ahuja D, McMahon M, Blair JA, Wang D et al (2010) Resiliency and Vulnerability in the HER2-HER3 Tumorigenic Driver. Science Translational Medicine 2. https://www.science.org/doi/10.1126/scitranslmed.3000389
Funding
Open access funding provided by HEAL-Link Greece. The work was funded by the Hellenic Foundation for Research and Innovation (H.F.R.I.) (Grant 236-EMPiBr) and the HORIZON2020 (Grant No. 801347).
Author information
Authors and Affiliations
Contributions
NS and AK designed the study and wrote the manuscript. NS performed most of the experiments. AC assisted with the animal and biochemical work. ADS and IPT designed and performed the ROS and mitochondrial respiration experiments as well as the confocal imaging of the cells. GP, EC and PM generated the KMT2C/MLL3 mouse. PM assisted also with the tamoxifen injections. AK assisted with the animal work. NP carried out the sorting of the cells. ISP performed the H&E imaging. EA and DK did the bioinformatic analysis.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Ethics approval and consent to participate
All procedures for the care and treatment of the animals were approved by the Institutional Committee on Ethics of Animal Experiments. The licence for the animal handling protocol for this project is: 1317/13–03-2019.
Consent for publication
All authors read and approved the final manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
18_2023_4734_MOESM2_ESM.xlsx
Supplementary file2 Supplementary Table 2: The statistically significant differentially expressed genes as revealed by the RNA-seq data (XLSX 376 KB)
18_2023_4734_MOESM3_ESM.xlsx
Supplementary file3 Supplementary Table 3: Summary of annotated genes to the unique identified peaks as revealed from the Venn diagram in H3K4me3 and H3K4me1 peaks from controls and Kmt2c KO (XLSX 90 KB)
18_2023_4734_MOESM4_ESM.pdf
Supplementary file4 Supplementary Figure 1: Occurrence of KMT2C mutations in humans. (A) Occurrence of KMT2C mutations in 122 out of 996 breast cancer patients taken from the TCGA PanCancer Atlas (cbioportal), (B) Survival data from TCGA PanCancer for breast cancer patients (KMT2C mutated patients=altered group, KMT2C non-mutated patients=unaltered group). The patient cohort is comprised of LumA, LumB, Her2 and basal subtypes. Supplementary Figure 2: Overview of the construct design for the Kmt2cfl/fl mice and H&E stainings from breast tumours. (A) Schematic representation of the construct for the Kmt2cfl/fl mice, (B) Representative southern blot image of the positive clone (c/+) accompanied with a wild type clone (+/+), (C) H&E stainings from breast tumours from MMTV-Myc and Krt8CreERT2;R26tdTomato;MMTV-Myc;Pik3caH1047R along with their Kmt2c KO counterparts, (D) immunoblotting analysis on H3K4me1 and H3K4me3 on MMTV-Myc;ex20, MMTV-Myc control and KO mice. Supplementary Figure 3: Average profiles on TSS and genomic distribution of ChIP-seq. (A) Average profiles at the TSS on unique H3K4me3 and (B) eTSS on unique H3K4me1 identified peaks (C) Genomic distribution of all the peaks from H3K4me3 and H3K4me1 ChIP-seq. Supplementary Figure 4: EMT phenotype in cells derived from MMTV-Neu and Kmt2c KO breast tumours. (A) Confocal images of cells isolated from MMTV-Neu and Krt8CreERT2;R26tdTomato;Kmt2cfl/fl;MMTV-Neu (Kmt2c KO) tumours stained with Dapi and Mitotracker showing the elongated phenotype of the KMT2C KO cells. Based on the cassette that was described in the materials ‘section, the KMT2C KO cells expressed Tomato upon tamoxifen administration, while the control mice not, (B) Snai1 and Snai2 abundances in cells isolated from breast tumours n=2 for each condition and representative qPCR showing mRNA levels of Snai1 in the cells, presented as fold change relative to control. Supplementary Figure 5: Pathway analysis and differential gene expression showing enrichment in extracellular matrix organization and basal markers in the Kmt2c KO tumours (A and B) Pathway analysis on the unique H3K4me3 peaks for each condition using GO and ClusterProfiler, (C) Heatmap showing the differential gene expression of Trp63, Krt8 and Krt18 in Kmt2c KO and control breast tumours as revealed by the RNA-seq data. Supplementary Figure 6: Motif analysis on the unique identified peaks. (A) Motif analysis on unique H3K4me3 peaks and (B) Motif analysis on unique H3K4me1 peaks. (Gained in Kmt2c KO-left, lost in Kmt2c KO-right). (PDF 25701 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Simigdala, N., Chalari, A., Sklirou, A.D. et al. Loss of Kmt2c in vivo leads to EMT, mitochondrial dysfunction and improved response to lapatinib in breast cancer. Cell. Mol. Life Sci. 80, 100 (2023). https://doi.org/10.1007/s00018-023-04734-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-023-04734-7