Abstract
XY chromosome missegregation is relatively common in humans and can lead to sterility or the generation of aneuploid spermatozoa. A leading cause of XY missegregation in mammals is the lack of formation of double-strand breaks (DSBs) in the pseudoautosomal region (PAR), a defect that may occur in mice due to faulty expression of Spo11 splice isoforms. Using a knock-in (ki) mouse that expresses only the single Spo11β splice isoform, here we demonstrate that by varying the genetic background of mice, the length of chromatin loops extending from the PAR axis and the XY recombination proficiency varies. In spermatocytes of C57Spo11βki/− mice, in which loops are relatively short, recombination/synapsis between XY is fairly normal. In contrast, in cells of C57/129Spo11βki/− males where PAR loops are relatively long, formation of DSBs in the PAR (more frequently the Y-PAR) and XY synapsis fails at a high rate, and mice produce sperm with sex-chromosomal aneuploidy. However, if the entire set of Spo11 splicing isoforms is expressed by a wild type allele in the C57/129 background, XY recombination and synapsis is recovered. By generating a Spo11αki mouse model, we prove that concomitant expression of SPO11β and SPO11α isoforms, boosts DSB formation in the PAR. Based on these findings, we propose that SPO11 splice isoforms cooperate functionally in promoting recombination in the PAR, constraining XY asynapsis defects that may arise due to differences in the conformation of the PAR between mouse strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In eukaryotes, proper segregation of meiotic chromosomes and the production of balanced gametes require recombination between the homologous chromosomes (homologs), a process that is initiated by a programmed wave of double strand breaks (DSBs) introduced by the type IVA topoisomerase-like protein SPO11, along with TOPOVIBL [1,2,3,4,5,6,7,8,9,10]. Following formation of DSBs, DNA at the DSBs site is resected, resulting in single-stranded DNA (ssDNA) ends that become the binding site of DNA exchange factors that ultimately leads to the formation of cross-overs (COs) (see [11] and references therein). COs not only shuffle the genome, but also physically link homologs, which ensures they remain associated until segregation occurs at anaphase-I [12, 13]. In males of mouse and humans’ species, recombination between sex chromosomes is more challenging than between autosomes, as DSBs must occur within a short region of homology between them, the pseudoautosomal region (PAR). At least one DSB must form, to allow the generation of the so-called “obligatory CO”, which guarantees proper XY segregation. The haploid mouse genome averages less than one DSB/10Mb, whereas the < 1Mb PAR undergoes one to two DSBs, a frequency that is 10-20-fold higher than the genome average [14]. This indicates that there are mechanisms in place that increase SPO11 activity at the PAR or make it more conductive to the formation of DSBs. In recent years, studies on the mechanisms underlying XY recombination have revealed that proper expression of Spo11 splice isoforms is key to male sex chromosome recombination. In mammals, Spo11 has two major splice variants, which are developmentally regulated: Spo11β (44.5 kDa) and Spo11α (40.3 kDa; exon 2 skipped) both including exon 5, the one that encodes the catalytic tyrosine essential for the formation of DSBs [15,16,17]. By using a mouse transgenic model, it was shown that the expression of the single SPO11β variant causes XY segregation failure and sterility, due to the reduction of the formation of DSBs in the PAR [14]. More recently, it was unexpectedly found that the degree of XY recombination was partially rescued when the transgene was introduced into a different genetic background [18]. This indicates that although germ cells that express only SPO11β are vulnerable to XY recombination-failure, unknown genetic background-dependent factors shape this susceptibility.
The demonstration that the expression of Spo11β does not guarantee recombination at the PAR raised the question of whether, in certain genetic contexts, SPO11α is required to perform this function. In the germ cells, the latter is expressed later than SPO11β, approximately at the time when DSBs are made in the PAR and XY synapse [14], making it a perfect candidate as recombination initiator in the PAR. Nevertheless, no experimental proof of SPO11α role has been provided yet.
The initiation of meiotic recombination requires the expression, along with SPO11 and TOPOVIBL, of auxiliary proteins that are essential for the formation of DSBs in autosomes. In mammals, these include IHO1, MEI1, MEI4 and REC114 [19,20,21,22,23,24]. XY recombination has additional genetic requirements, demanding expression, and localization on the PAR of ANKRD31, a REC114 binding-protein [25, 26]. Several studies have shown that in yeast and mammals, SPO11-auxiliary proteins (also known as RMMAI proteins [23]) are loaded on the chromosome axis, prior to DSBs formation [19,20,21,22,23,24,25,26,27,28]. Nevertheless, according to the yeast model, meiotic DSBs are preferentially localized in the open region of the chromatin, within chromatin loops [29]. This observation has led to the theorization of the “tethering model” which predicts that SPO11 binds to chromatin loops and is successively tethered to the axis, where it is incorporated into the so-called DSB-promoting complex formed by the auxiliary factors [29, 30].
Studies in mice have shown that PAR axes are disproportionately long relative to DNA length (1Mb/mm of axis) compared to autosomes (10-13Mb/mm of axis). Since the density of the loop per millimeter is constant [31], this results in smaller chromatin loops, which according to the tethering model are thought to be more conducive to DSBs [14]. However, whether shorter PAR loops truly boost the formation of DSBs in the PAR is awaiting experimental proof.
By generating a Spo11β knock-in hemizygous mouse model (Spo11βki/-), we show that in mice with a mixed genetic background (C57BL/6 and 129Sv) the frequency of DSBs formation and recombination in the PAR is highly variable and that a shift to the C57 background greatly reduces such defects. Analysis of PAR ultrastructure revealed that rescue correlates with a shortening of PAR loops and an increased frequency of formation of DSBs. Furthermore, we provide experimental evidence that regardless of PAR structure characteristics, the hemizygous expression of the wild type allele of Spo11 limits the extent of XY synapsis defects. Finally, by generating Spo11α knock-in mice, we prove that SPO11α promotes the formation of DSBs in the PAR, upon concomitant expression of SPO11β.
Results
The testes weight of Spo11βki/- mice varies with the genetic background
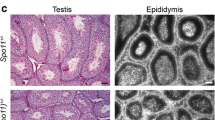
In male mammals, death of defective germ cells within the testis, causes an overall reduction in testis weight, so this can be used to quantify spermatogenesis performance (e.g., see [32]).To test how the expression of Spo11β affects spermatogenesis when the protein is expressed under normal physiological timing and at allelic dosage, we generated mice expressing a single knock-in allele of Spo11βb (thereafter named Spo11βki/-) under the control of the Spo11 promoter (Fig. S1). Mice were created with a mixed (C57BL/6 and 129Sv) genetic background (C57/129Spo11βki/−), see material and methods and Fig. S2A. Examination of relative testis weight (testis to body-weight ratio) revealed great variability among C57/129Spo11βki/− mice, compared to littermates C57/129Spo11+/-. Indeed, while some C57/129Spo11βki/− males had testes with visibly reduced weights, below the mean (i.e., small testis; ST), others appeared indistinguishable from Spo11+/- mice (i.e., with a het-like (HL) phenotype) (Fig. 1A). Nevertheless, relative testis weight of ST mice was greater than in Spo11−/− mice, in which progression of meiosis arrests at zygonema of the first meiotic division [5, 6, 32], indicating that in C57/129Spo11βki/− mice the arrest is either incomplete or it occurs beyond zygonema. Given that the mice were of mixed genetic background, we reasoned that the observed phenotypic variability could have been related to background variations. To test this interpretation, we introduced the Spo11βki allele into a pure C57/BL6 background (C57Spo11βki/− mice) (see Fig. S2B and material and methods); variability was greatly reduced, and testis to body weight ratio turned very similar to Spo11+/- (Fig. 1B). Next, to understand whether the phenotype would have worsened in the 129Sv background, we backcrossed C57/129Spo11βki/− mice into 129Sv for one generation (see Fig. S2C and material and methods). A single backcross shift was sufficient to worsen the phenotype (compare C57/129Spo11βki/− mice in Fig. 1A and Fig. 1C). This was also confirmed by backcrossing C57Spo11βki/− mice into 129Sv for one generation (Fig. S2D and S3A). We concluded that in males with the Spo11βki/- genotype, the performance of spermatogenesis changes with genetic background.
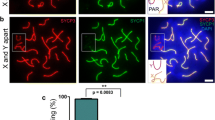
Variability of relative testis weight and XY asynapsis in Spo11βki/- mice with a different genetic background. A Testis to body weight ratio in mice with the indicated genotypes and genetic background. The dotted line indicates a testis to body weight ratio mean equal to 2.1 × 10–3 ± 0.6 × 10–3. B Testis to body-weight ratio in mice of the indicated genotypes upon seven backcrosses in C57BL/6 background. C Testis to body-weight ratio in mice of the indicated genotypes upon one backcross of mice with mixed background in 129/Sv background (HL = heterozygous-like; ST = small testis). In A-C, each dot represents a mouse. C.V. (%) = coefficient of variation. D Representative images of spermatocytes stained for the lateral element (SYCP3) and the central element (SYCP1) of the SC. X and Y indicate sex chromosomes; The white arrow points to the PAR. Magnification bar is 10 μm. E Frequency of XY asynapsis in nuclei at pachynema. Each dot is a mouse with the indicated genotype; n = total number of cells scored for each genotype. The error bars are the mean ± standard deviation (SD) of the mean; p = p value (two-tailed t-test, p < 0.05)
Reduced testis to body weight ratio in C57/129Spo11βki/− ST mice correlate with failure of sex chromosome synapsis and apoptotic elimination of spermatocytes at metaphase I
In mammals, synapsis of spermatocyte chromosomes occur in the context of the development of a zipper-like proteinaceous structure called synaptonemal complex (SC) [33]. Synapsis begins with the alignment of the homologs at leptonema and is completed by pachynema. Cytologically, cells in leptonema are identified by the appearance of SYCP3 positive stretches of the lateral elements of the SC; progression to zygonema is marked by the assembly of the SYCP1-positive central element of the SC, between pairs of synapsed homologues. At pachynema, autosomes are fully synapsed throughout their entire length and SYCP3 and SYCP1 signals overlap throughout. In contrast, synapsis between XY chromosomes occurs only at the PAR. Thus, a short stretch of SYCP1 forms between chromosomes, only in this region, while the rest of the chromosomes axes is marked by SYCP3.
To probe if variations in testis to body weight ratio in C57/129Spo11βki/− mice was related to the proficiency of XY synapsis, we quantified XY asynapsis in our genotypes of interest by staining surface spread chromosomes of C57/129Spo11βki/− ST, C57/129Spo11βki/− HL and C57Spo11βki/− males with anti-SYCP3 and anti-SYCP1 antibodies. While in C57/129Spo11βki/− ST, XY synapsis failed in ~ 55% of spermatocytes; the percentage was down to ~ 11% in C57/129Spo11βki/− HL and to 4% in C57Spo11βki/− mice (Fig. 1D–E), indicating that the reduced testis weight and frequency of XY asynapsis are closely correlated.
In male mice and humans, each seminiferous tubule cross section can be assigned to one of the 12 epithelial stages (numbered I-XII) based on the array of germ cell developmental stages it contains [34,35,36]. Elimination of MI spermatocytes that have achiasmate homolog pairs (non-exchange) occurs in stage XII by activating the spindle assembly checkpoint (SAC) [14, 37]. To evaluate the occurrence of germ cell loss by apoptosis at stage XII, we combined terminal deoxynucleotidyl transferase dUTP nick end labelling (TUNEL) and anti-H3Ser10 (pH3) staining in testis sections. The latter was used as a marker to identify metaphase I (MI) cells in stage XII. As shown in Fig. S3B and quantified in Fig. S3C, the frequency of MI cell apoptosis was higher in the tubules of C57/129Spo11βki/− ST males compared to those of C57/129Spo11βki/− HL and C57Spo11βki/− males. We concluded that in mice with a Spo11βki/- genotype testicular atrophy is related to failure of XY synapsis and apoptotic elimination of defective spermatocytes in stage XII.
C57/129Spo11βki/− ST spermatocytes are defective for the formation of DSBs in the PAR
To assess whether the defect of XY synapsis in C57/129Spo11βki/− ST mice was attributable to the lack of DSBs formation in the PAR, we combined the staining of the SC component SYCP3 and DMC1 (a surrogate marker of DSBs [12, 38, 39]) with that of PAR, using fluorescent in situ hybridization (FISH). The PAR probe recognizes a region at the boundary between the non-PAR region and the PARs of the X and Y chromosomes, and hybridizes with the tandem array of minisatellite mo-2 at the noncentromeric end of chromosomes 4, 9 and 13 [23]. This prevents iterated unequivocal identification of the X-PAR. On the contrary, the Y-PAR FISH signal has a distinctive pattern, as the FISH staining always extends from the Y chromosome axis to the chromatin loops, forming a distinguishable cloud around the Y-PAR (Fig. 2A). Under physiological conditions, DSB formation occurs with a comparable frequency in both the X-PAR and Y-PAR, mainly at the late zygotene stage, [14]. Therefore, since the Y-PAR is uniquely identified with the PAR FISH probe, we quantified the frequency of DMC1 foci in this region, in late zygonema spermatocytes from C57/129Spo11βki/− ST and C57Spo11βki/−mice. To enrich our samples for germ cells at late zygonema, we prepared chromosome spreads from juvenile (12 dpp) mice. At this time point, apoptosis selection of cells defective in XY synapsis had not yet occurred [32], therefore, the ST phenotype cannot be assessed. To overcome this problem, we evaluated the percentage of XY asynapsis and only included C57/129Spo11βki/− mice with at least 35% XY asynapsis in the analysis (henforth referred to as ST equivalent - STe) (Fig. 2B). This value was set according to the correlation between the frequency of XY asynapsis and testis to body weight ratio in adult C57/129Spo11βki/− mice (Fig. S3D). Alongside, with this, we analyzed DSBs formation in C57Spo11βki/− spermatocytes, in which the average XY asynapsis was less than 10% (Fig. 2B). The analysis of the presence of DMC1 foci in the PAR of late zygotene cells, revealed that the high degree of XY asynapsis correlates with a reduced frequency of the presence of DMC1 foci (Fig. 2C). However, the frequency of DMC1 in the Y-PAR was low compared to the percentage of XY asynapsis. This raised the question of whether DSBs form less frequently in the Y-PAR than in the X-PAR. To test this, we identified both PARs by immunolocalizing ANKRD31, which at the zygotene-pachytene transition and at the early pachytene stages aggregate on PARs (see [26] and below). We analyzed spermatocytes of C57/129Spo11βki/− STe (14 dpp) mice with an average XY asynapsis (estimated by SYCP3/SYCP1 staining) equal to 52.5 ± 8%. Of 42 cells with unsynapsed sex chromosomes, 25 (59.5%) had no foci on PARs (Fig. S3E), 10 (23,8%) showed a focus only on the X-PAR, 5 (12%) only in Y-PAR, and 2 (4.7%) in both PARs. The latter are likely cells in which foci are found on both chromosomes, upon release of one DSB from either PARs [14]. We concluded that in C57/129Spo11βki/− spermatocytes XY asynapsis occurs as a result of the lack/delayed formation of DSBs on PARs, confirming previous findings [14], and that the frequency of DSBs in the Y-PAR is about twice as low as in the X-PAR.
Quantification of the number of DSBs and the expression of SPO11. A Representative images of chromosome spreads of late zygotene stage spermatocytes of the indicated genotypes, stained with the anti- SYCP3 and DMC1 antibodies, and hybridized with the PAR FISH probe. Magnified views of the Y chromosomes are shown in the inset. Arrows point to the X-PAR and Y-PAR; *are heterochromatic mo-2 31-bp repeat of either ch4, ch9 or ch13, recognized by the PAR probe. Magnification bar is 10 μm. B Quantification of XY asynapsis in juvenile mice with the indicated genotypes and genetic backgrounds. C Frequency of the presence of a DMC1 focus on the Y-PAR of cells in A-B. In B and C, each dot is the frequency per mouse; n = total number of cells analyzed. D Immunoprecipitation (IP) and Western blot analysis of SPO11 expression in testes of mice with the indicated genotypes. Spo11−/− mice serve as negative controls. Each lane is the expression of one testicle of 4 different mice. The input is a Western blot analysis of the indicated protein markers in total testicular extracts used for IP. SYCP3 and tubulin were normalizers of the number of meiotic germ cells and proteins in the extracts, respectively. E Quantification of global DSB numbers in spermatocytes from 12 dpp mice with the indicated genotypes and genetic backgrounds. Each dot indicates the number of DMC1 foci per nucleus. Le leptonema; eZ/mZ early-mid zygonema; lZ late zygonema; eP early pachynema. Error bars are mean ± SD; p = p value (two-tailed t-test, p < 0.05); n = total number of cells scored at each stage (at least three mice per genotype)
Analysis of SPO11 expression in mice with different genetic backgrounds
In mice, the expression of the SPO11 protein below a critical amount may have an effect on DSB levels and chromosome synapsis proficiency [38, 39]. To test whether failure of XY synapsis in C57/129Spo11βki/− STe mice was related to faulty expression of SPO11β, we immunoprecipitated it from mouse testis extracts from juvenile mice at 12 dpp. Protein levels among Spo11+/- and Spo11βki/- mice were comparable (Fig. 2D and Fig. S3F). SPO11 protein levels were also comparable among C57/129Spo11+/-, C57Spo11βki/− and C57/129Spo11βki/− HL genotypes, in adults (Fig. S3G). Next, to investigate whether SPO11 function is normal in C57/129 Spo11βki/− STe spermatocytes, we quantified the number of DSBs nucleus wide by co-staining spermatocyte surface chromosome spreads with SYCP3 and DMC1. We did not observe a reduction in DMC1 foci number in C57/129Spo11βki/− STe cells compared to C57Spo11βki/− spermatocytes. Rather, the average number of foci at leptonema and early mid-zygonema increased slightly in C57/129 Spo11βki/− STe cells (Fig. 2E). We concluded that it is unlikely that the reduced frequency of DSB formation in the PAR of C57/129Spo11βki/− STe mice is due to defects of SPO11β expression or function.
Reduced DSB formation in the PAR of C57/129Spo11βki/− STe spermatocytes is not related to defects in the aggregation of the auxiliary proteins of SPO11.
The formation of DSBs in the PAR by SPO11 occurs with the assistance of auxiliary proteins, including IHO1, MEI4, REC114, MEI1 and ANKRD31 (RMMAI complex) [19,20,21,22,23,24,25,26]. Aggregation of RMMAI proteins on the PAR occurs from the preleptotene stage, in advance of the formation of DSBs [23]. To investigate whether SPO11 auxiliary proteins localize normally in PARs of mice with increased XY asynapsis, we monitored the assembly of ANKRD31, MEI4, and REC114 from preleptonema to zygonema in C57/129Spo11βki/− STe mice. Spermatocytes from wild type C57 mice were used as a control. To identify their association with the PAR axis, surface chromosome spreads were stained with SYCP3 and the PAR probe. As shown in Fig. S4A-C and quantified in Fig. S4D-F, aggregation of these factors was comparable to that of the control. Furthermore, we immunolocalized aggregates of ANKRD31, MEI4 and REC114 at the zygotene/pachytene transition stage, the sub-stage when most DSBs form in the PAR [14]. To this end, we colocalized them with IHO1, which at this stage forms a blob signal only on X-PAR and Y-PAR [20]. In this case, we never observed cells without ANKRD31, MEI4, or REC114 aggregates in C57/129 Spo11βki/− STe mice (197, 168 and 231 cells analyzed respectively, from three mice per genotype) (Fig. S4G). From these results, we ruled out the possibility that a defective aggregation of RMMAI proteins is responsible for the XY asynapsis defects observed in C57/129Spo11βki/− STe spermatocytes.
Spermatocytes from C57/129Spo11βki/− STe and C57Spo11βki/− mice differ in the high-order chromatin structure of the PAR
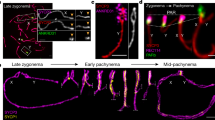
In mice, the formation of DSBs in the PAR is preceded by its ultrastructural remodeling that consists of the separation (splitting) by zygonema of the aligned sister chromatid axes, decorated with RMMAI proteins [23]. To monitor potential changes in PAR ultrastructure in C57/129Spo11βki/− STe spermatocytes, we analyzed the PAR of surface chromosome spreads of spermatocytes at the zygonema/pachynema transition using Stimulated Emission Depletion (STED) microscopy. To this end, the spermatocyte chromosome axis was stained with anti-SYCP3 antibody, while the sex chromosomes and PARs were identified by IHO1 stain [20]. PARs were also identified by using the anti-ANKRD31 antibody, which forms distinguishable large aggregates on both the X-PAR and Y-PAR [26] (Fig. 3A). By comparing STED images (insets in Fig. 3A), we found that the frequencies of X-PAR axis splitting in late zygonema were comparable between C57/129Spo11βki/− STe and C57Spo11βki/− mice we used as control (92%, n = 34 and 91%, n = 33, respectively), while Y-PAR splitting occurred less frequently in C57/129Spo11βki/− STe mice (C57/129Spo11βki/− ST 84%, n = 35; C57Spo11βki/− 96%, n = 41, p = 0.0004 Chi-Square test). Although the physiological role of PAR splitting is still unclear [23], this result suggested a small but noticeable defect in Y-PAR remodeling. In mouse, splitting of the PAR axes are strictly temporally correlated with the remodeling of the PAR chromatin loops and axis. The PARs loops are short at leptonema up to late zygonema, when DSBs are made in the PAR, and lengthen in early to mid-pachynema cells [23]. Correspondingly, the PAR axis is long as soon as it is detectable at leptonema and late zygonema/early pachynema and shortens in the mid-pachytene stage [23]. In our effort to understand the molecular basis of the defect of XY synapsis in C57/129Spo11βki/− STe spermatocytes, we sought to study the changes in PAR conformation by measuring the length of the loops and the axis during prophase I in surface spreads of spermatocytes stained with SYCP3 and the PAR FISH probe. We focused on the Y-PAR, as it is uniquely identifiable and its dynamic changes in wild- type cells are well characterized [23]. As a control, we employed C57Spo11βki/− males, which are more proficient in XY synapsis (Fig. 2C). The size of loops was defined as the axis-orthogonal extension of the PAR FISH signal, while the length of the PAR axis was determined as the distance from the PAR probe to the end of the SYCP3 axis (Fig. 3B–C) [14, 23, 40]. Comparing cells at late zygonema and early pachynema in C57/129Spo11βki/− STe spermatocytes, the average size of PAR loops at late zygonema was shorter than in early pachynema, confirming previous results [23]. This was true regardless of whether the XY synapses had just occurred at early pachynema (Fig. 3D). Similarly, the Y-PAR loops of C57Spo11βki/− spermatocytes at late zygonema were shorter compared to cells at early-pachynema with synapsed sex chromosomes. An upward trend in average loops length was also observed in early pachytene-stage cells with asynapsed XY, although the difference did not reach statistical significance. Remarkably, the comparison of FISH signals among cells of C57/129Spo11βki/− STe and C57Spo11βki/− mice indicated that the PAR loops of C57Spo11βki/− mice were constitutively more compact than those of C57/129Spo11βki/− STe cells (Fig. 3D), consistent with smaller loops. Side-by-side analysis of the length of the Y-PAR axis showed that it shortened slightly in early pachynema cells of C57Spo11βki/− mice, while no significant variations were found in C57/129Spo11βki/− STe cells (Fig. S5A). The latter was expected, as the shortening of the PAR axis is generally measurable by mid-pachynema [23]. We did not find mid-pachytene cells at the 12 dpp time point; therefore, shortening of the PAR axis at this more advanced stage could not be tested. From these experiments, we concluded that the spermatocytes of C57/129Spo11βki/− STe and C57Spo11βki/− mice differ for the high-order chromatin structure of the PAR.
Analysis of the Y-PAR conformation and the proficiency of the XY synapsis. A Representative images of cells from 12dpp C57Spo11βki/− and ST-equivalent C57/129 Spo11βki/− at late zygotene stage, stained with the indicated markers. IHO1 was used to identify asynapsed chromosomes, while ANKRD31 identifies PARs. Magnifications are STED images of the X and Y PARs. The analysis was performed in two mice per genotype. The numbers of X and Y chromosomes analyzed by STED are as follows: C57 Spo11βki/− mice 21 and 29 respectively; C57/129 Spo11βki/− STe spermatocytes, 26 chromosomes in either case. Arrows point to the X-PAR and Y-PAR. Magnification bar is 10 μm. B Schematic of the axis loop structure and the PAR FISH signal. Only one homolog is shown. The length of PAR loops is measured as the orthogonal extension of the FISH signal from the chromosome axis identified by SYCP3. The length of the axis is measured as the maximum distance from the PAR FISH signal to the distal end of the SYCP3-defined axis. C Representative image of a late zygotene cell used for the analysis. The dashed square encircles the Y chromosome, identified by the PAR FISH staining pattern (inset). Insets are magnifications showing the Y chromatin extension (green signal, top inset) and the axis extension (red tick signal, bottom inset). The white arrow points to the terminal end of the Y-PAR. In the insets, the white line indicates the length. Magnification bar is 10 μm. D Measurements of loop-axis extension from conventional immune-FISH images of cells at late zygonema and early pachynema, in mice with the indicated genotypes. Each dot represents the measurement of a single cell (three mice analyzed per genotype). Pachynema cells with synapsed or asynapsed sex chromosomes were separated into two groups. E Frequency of XY asynapsis in nuclei at pachynema, in C57Spo11βki/− and C57/129 Spo11βki/− ST mice. Each dot is a mouse; n is the total number of cells scored for each genotype. Error bars are SD, p indicates statistical significance (p < 0.05), two-tailed t-test
Interplay between PAR ultrastructure and expression of the Spo11 wild type allele
In mice carrying, a wild type allele of Spo11 in the mixed background (C57/129Spo11+/-), relative weight of the testes is high and less variable compared with that of C57/129Spo11βki/− mice (Figs. 1A, C and S3A). To investigate how these phenotypes correlate with the frequency of XY asynapsis, we quantified it in the genetic models of our interest. As shown in Fig. 3E, sex chromosome asynapsis was less frequent in C57/129 Spo11+/- mice compared to C57/129Spo11βki/− males. This indicates that the expression of the full set of Spo11 splice-isoforms by the wild type allele promotes XY recombination and synapsis better than the Spo11βki allele. The subsequent comparison of XY asynapsis in C57/129Spo11+/- and C57Spo11+/- males pointed out that the latter are the most proficient. To test whether this correlated with a shortening of the PAR loops length, we measured it in juvenile C57/129Spo11+/- and C57Spo11+/- mice. PAR loop length in spermatocytes with a C57 background were significantly shorter (Fig. S5B), confirming our previous results (Fig. 3D). Shortening of PAR loops also correlated with a recovery of XY asynapsis in cells from C57Spo11βki/− males (Fig. 3E). We concluded that reduced length of the PAR loops in the C57 background and the expression of a wild type set of Spo11 splice-isoforms, both impacts on XY recombination, likely by distinct mechanisms, in cooperation with each other.
The function of Spo11β on PAR is boosted by the concomitant expression of Spo11α
SPO11α conserves the catalytically active tyrosine residue of Spo11 required for its DSB formation activity [15, 16]; therefore, it is a potentially catalytically active isoform. With the goal of testing the ability of this isoform to form DSBs, we generated a knock-in mouse model that expresses it under the control of the Spo11 promoter (Fig. S1). Mice homozygous for the Spo11αki allele were generated on a C57 background (C57Spo11αki/αki). Analysis of the morphology and relative testicular weight of these mice revealed that they phenocopied Spo11−/− mice [5, 6] (Fig. 4 A, B). Furthermore, histological observation of the ovaries of adult mice revealed that females were also phenotypically similar to Spo11−/− [5, 6], as primordial follicles could not be observed in the cortex (Fig. S5C). Consistent with these observations, staining of spermatocyte spread chromosomes with SYCP3 and SYCP1 antibodies, revealed that, just as Spo11−/− spermatocytes [5, 6], C57Spo11αki/αki cells were not able to progress beyond a zygotene-like stage (Fig. 4C). Successive quantification of the number of DSBs in spermatocytes using DMC1 as a surrogate marker, showed that the number of DSBs was extremely low in C57Spo11αki/αki cells compared to wild type mice, although slightly higher than in Spo11−/− spermatocytes (Fig. 4D, E). To confirm this result, we also quantified the number of γH2AX patches, which mark DSB sites regardless of the DMC1 assembly [41]. Again, numbers of γH2AX patches were slightly increased compared to Spo11−/− mice (Fig. S5D-E). Confirming the failure of proper formation of DSBs, the histological analyses of C57Spo11αki/αki testes revealed that, as previously demonstrated in Spo11−/− mice [5, 32], spermatocytes underwent massive cell death (Fig. S5F). Next, we went one step further by testing whether one of the few DSBs that form in C57Spo11αki/αki spermatocytes occur in the PAR. To this end, we immunolocalized DMC1 in the PAR of surface chromosome spreads of C57Spo11αki/αki cells in combination with SYCP3 and the PAR FISH probe (Fig. 4F). Of the three mice analyzed, we never observed DMC1 foci in the Y-PAR of cells in leptonema (n = 53) and found foci in 6/376 nuclei in the zygonema-like stage (1.2% ± 0.3). Conversely, DMC1 foci were never found in the Y-PAR of Spo11−/− cells at any stage (n = 218 cells, from three mice). We concluded that in C57Spo11αki/αki males, DSBs form with extremely low efficiency on both non-sex and sex chromosomes. To investigate whether such a phenotype was traceable to a low level of the protein, we immunoprecipitated SPO11 from C57 wild type, C57Spo11+/- and C57Spo11αki/αki testes. Samples were collected from 12 dpp mice to compare testes with similar progression of meiosis. SPO11α expression in C57Spo11αki/αki mice was visibly reduced compared to SPO11β in wild type and C57Spo11± spermatocytes (Fig. 5A). This suggests that the low frequency of DMC1 foci in C57Spo11αki/αki spermatocytes is at least in part attributable to the low protein level.
Characterization of C57Spo11αki/αki mice phenotype. A Histological analysis of testes from mice of the indicated genotypes; hematoxylin and periodic acid shift staining of testis sections from adult mice. Round spermatids and sperm are apparent in the wild type testes. In contrast, tubules in C57Spo11αki/αki and Spo11−/− mice, lacks haploid cells. Two to three mice were analyzed for each genotype. Magnification bar is 50μm. B Relative Testis to body weight ratio of mice with the indicated genotypes. C Representative images of surface-spread spermatocyte nuclei stained with antibodies recognizing SYCP3 and SYCP1. D Surface-spread spermatocytes stained with antibodies that recognize SYCP3 and DMC1. E Quantification of DMC1 foci in mice with the indicated genotypes; we analyzed three mice per genotype. Each dot on the graph represents a single cell. The error bars are SD, p indicates statistical significance (p < 0.05), two-tailed t-test. F Surface spread spermatocytes from mice of the indicated genotype, stained with antibodies recognizing SYCP3, DMC1 and with the PAR FISH probe. In C-D and F magnification bars are 10μm
Expression of SPO11and phenotypic characterization of the C57Spo11α/+ mice phenotype. A IP Western blot analysis of SPO11 expression in mice of the indicated genotypes. Each lane is the expression of one testis of four mice. Input is Western blot analysis of the indicated protein markers in total testicular extracts. SYCP3 and tubulin in input were normalizers of the amount of meiotic germ cells and proteins in the extracts, respectively. B Surface spread spermatocytes from mice of the indicated genotypes, stained with antibodies that recognize SYCP3, DMC1 and with the PAR FISH probe. Y-PAR was identified by the staining pattern of the PAR FISH probe. The inset is a magnification of the Y chromosome. The white arrows point to the PAR. Bar is 10 μm. C Quantification of the number of DMC1 foci in the Y-PAR at different substages of spermatogenesis, in mice with the indicated genotypes (three mice per genotype); n = number of cells analyzed per stage. The error bars are SD, p indicates statistical significance (p < 0.05); one tailed t-test. D Quantification of the global number of DMC1 foci, in spermatocytes of mice with the indicated genotypes. The error bars are SD, p indicates statistical significance (p < 0.05), two-tailed t-test
Considering that under physiological conditions, SPO11α is expressed in prophase I, later than SPO11β, [6, 15, 42, 43], we speculated that another reason why the proficiency of DSB formation in the PAR and autosomes of C57Spo11αki/αki spermatocytes is low is because it lacks SPO11β. As shown in Fig. 5A (left panel), in our Spo11α knock-in model, the protein is expressed with an early timing compared to wild type, as it is already well detected in testes of 12 dpp mice, when in wild type mice is only observed SPO11β. Taking advantage of this characteristic, we generated mice expressing one wild type allele of Spo11 in combination with the Spo11αki allele (i.e., C57Spo11αki/+ mice). After verifying the expression of both splice isoforms (Fig. 5A, right panel), we quantified the number of DSBs in the PAR, comparing it with C57 wild type and C57Spo11+/- spermatocytes, which by this age only express SPO11β. Our prediction was that if the function of SPO11β in the PAR is enhanced by concomitant expression of SPO11α, DSBs should form with greater efficiency in the PAR of C57Spo11αki/+ cells at leptonema and early zygonema compared to cells from control genotypes. This expectation was met. Quantification of DMC1 foci in the Y-PAR of leptotene stage cells revealed that the frequency of DSBs was increased by five folds in C57Spo11αki/+ spermatocytes compared to wild type C57 cells and by over 16 folds compared to cells from C57Spo11+/- mice. A smaller increase was also observed in the early/mid zygotene and early pachytene stages, compared to C57Spo11+/- cells (3.9 and 1.1, respectively) (Fig. 5B, C). From this observation, we concluded that SPO11β function in the PAR is augmented by the concomitant expression of SPO11α. Interestingly, quantification of DMC1 foci on whole chromatin of C57Spo11kiα/+ spermatocytes at leptonema and early/mid zygonema revealed that “global” DSBs increased less (1.4 and 1.1 folds, respectively) than in the PAR (Fig. 5D). This indicates that the expression of SPO11α is mainly functionally related to recombination initiation in the PAR.
C57/129Spo11βki/− ST mice are prone to sex chromosome aneuploidy in sperm
Previous studies have shown that in male mice prone to sex chromosome asynapsis, fertility and differentiation of aneuploid sperm are functions of the degree of XY asynapsis [44]. If XY pairing fails in not more than ∼50% of sperm, activation of the spindle assembly checkpoint (SAC) does not have an obvious impact on sperm production and mice are fertile [44]. Consistent with the fact that in C57/129Spo11βki/− ST mice XY synapsis fails in ~ 55% of cells, mice had reduced but still abundant spermatozoa in the cauda of the epididymis (Fig. S6A) and were fertile (Table S1). Next, to understand whether such mice generated sperm aneuploid for the sex chromosomes, we subjected cells collected from the cauda of the epididymis to FISH with probes against the X and Y chromosomes. A fluorescent in situ hybridization probe for chromosome 8 served as an internal control for correct identification of aneuploid sperm vs diploid ones (Fig. S6B). In C57/129Spo11βki/− ST mice, the percentage of sperm nuclei containing both X and Y or no sex chromosomes was increased (Fig. S6C). We conclude that C57/129Spo11βki/− ST males are prone to formation of aneuploid gametes.
Discussion
Proficiency of XY recombination changes with mouse strain and correlates with variations of the ultrastructure of the PAR
Previous studies demonstrated that the expression of the single Spo11β splice-isoform in mouse predisposes defective XY recombination and synapsis [14]. However, the degree of XY recombination failure varies by mouse strain [18]. It remained unknown whether this occurred because a lack of concurrent expression of SPO11α, an altered recruitment of the RMMAI factors, alterations in the high-order chromatin structure of the PAR, or by other mechanisms. Herein, by generating a Spo11β knock-in model that expresses the protein under the control of its own promoter, we show that in mice with mixed genetic backgrounds (C57BL/6 and 129Sv), formation of DSBs in the PAR and XY synapsis is impaired with high variable frequency. On the contrary, the introduction of the knock-in allele in a pure C57 background greatly restored SPO11β function at the PAR and XY synapsis, providing a comparative model to investigate the mechanisms shaping the proficiency in PAR DSB formation. Comparison of SPO11β expression in C57/129Spo11βki/− ST and C57Spo11βki/− males revealed equal levels of protein expression and bulk of DSBs. This excluded any potential detrimental effect on XY recombination due to variable expression of the ki allele in different genetic backgrounds. Subsequent analysis of the presence and timing of aggregation of RMMAI factors in the PAR also ruled out the possibility of their defective recruitment as causative for XY recombination failure. In wild type, concomitantly with RMMAI proteins aggregation, the PAR undergoes notable ultrastructural rearrangements prior to DSB formation. These include the separation of the aligned sister chromatids from each other (splitting), the elongation of the PAR axis, and the shortening of the chromatin loops. These changes have been proposed to be essential for the recombination, pairing, and segregation of XY chromosomes [23]. However, whether alterations of the PAR ultrastructure correlate with XY recombination defects has never been experimentally tested. By analyzing PAR splitting in C57Spo11βki/− and C57/129Spo11βki/− ST spermatocytes, we observed in the latter, a small difference in the frequency of Y-PAR splitting, this suggested a defect in remodeling of the Y-PAR, which correlated with a more pronounced reduction of DSBs in the Y-PAR than in the X-PAR. To date, the functional significance of splitting of the PAR is unclear. Two strongly related hypotheses have been proposed. One is that separated axes would accommodate a considerable amount of SPO11 RMMAI proteins required for sufficient DSBs [23]. Alternatively, splitting could prevent unnecessary ineffective inter-sister recombination, to support repair of DSB by homologous recombination [45]. Given that in our model we did not observe substantial defects in hyperaccumulation of RMMAI proteins on the PAR, we favor the latter hypothesis. Next, deepening the analysis of the ultrastructure of the PAR, we analyzed loop/axis remodeling. We focused on the Y-PAR and found that in C57/129Spo11βki/− ST spermatocytes loops are considerably longer than those of C57Spo11βki/− cells. This is in line with the model that envisions short loops being more conducive to the formation of DSBs [14, 23]. We concluded that defective XY recombination in C57/129Spo11βki/− ST cells is likely due to the ultrastructural conformation of the PAR.
Interply between Spo11 splicing isoforms expression and PAR conformation
Comparison of XY recombination proficiency in mice expressing a single wild type Spo11 allele with that of mice with the Spo11βki/- genotype revealed that the former were more proficient in XY synapsis. This indicated that one wild type allele of Spo11 is superior to the Spo11βki allele, in promoting XY recombination. On the other hand, Spo11+/- with a C57 background were the most proficient in sex chromosome synapsis among Spo11+/- mice. This was also correlated with the presence of shorter PAR loops. We concluded that strain-dependent changes in PAR ultrastructure and the expression of a wild type Spo11 allele cooperate in promoting XY recombination, likely by different mechanisms. Based on these results we hypothesized that, in cases where the ultrastructure of the PAR is unfavorable for the formation of DSBs (i.e., long loops; for instance, for a constitutive 3D organization of PAR chromatin), the concomitant expression of other splicing isoforms of Spo11 additional to SPO11β may compensate for such characteristic. The Y-PAR is the one receiving DSBs with lowest frequency in C57/129Spo11βki/− mice with defective XY synapsis. Therefore, it is likely the one that benefits most from the expression of additional Spo11 splicing forms.
The concomitant expression of SPO11β with SPO11α promotes DSB formation in the Y-PAR
The major alternative splice isoform of Spo11 expressed in addition to Spo11β in Spo11+/- mice, is Spo11α. Thus, we took a step forward by generating a new knock-in model that expresses SPO11α under the control of Spo11 promoter. Phenotypic characterization of C57Spo11αki/αki mice showed that although proficiency of DSB formation in bulk chromatin and PAR was extremely low compared to normal mice, it was above the level in Spo11−/− cells. That said, SPO11α expression was surprisingly low in our model, reduced by approximately 2.5 folds of the level of SPO11β in age-matched Spo11+/- mice. The reason for this attenuated expression is unknown. It could be due to reduced expression of the knock-in allele. However, as the backbone construct is identical to that of the β isoform, it is unlikely. Therefore, we favor the hypothesis that the mRNA or the protein of the α isoform is less stable. Regardless of the underlying mechanism, this observation suggested that the inefficient formation of DSBs by SPO11α in our model might be linked to its reduced expression. However, comparison with a mouse model in which SPO11β is reduced at an apparently comparable level [38], indicated that in these cells the number of DMC1 foci was approximately 50-fold higher than in C57Spo11αki/αkicells (~ 100 DMC1 foci on average in early mid zygonema in Tg(Spo11β) ± vs. 2.2 on average in zygotene-like cells of C57Spo11αki/αki mice). This indicates that while SPO11α conserves the catalytic domain [15, 16] it has low DSB activity. Consistent with this interpretation, it has been shown that generation of DSBs by SPO11β requires its heterotetramerization with two TopoVIB-like (TOPOVIBL) subunits to adopt the structure required for DNA cleavage [10, 46]. TOPOVIBL is apparently unable to physically interact with SPO11α [10], which likely represents a limit in the activity of SPO11α. Given that SPO11α molecules can self-interact [43], we speculate that in C57Spo11αki/αki mice, protein complexes containing αα dimers form, and have no (or a very reduced) function in the formation of DSBs in autosomes and low DSB formation efficiency in the PAR. In wild type cells, the formation of DSBs in the PAR occurs at a time point when both SPO11β and SPO11α are expressed [14]. Therefore, it was predicted that the formation of DSBs in the PAR could be favored by their concomitant expression. To test this interpretation, we took advantage of the fact that SPO11α is expressed earlier than wild type in our knock-in mouse model, with the same timing as that of SPO11β. We asked if DSBs form with a greater efficiency in leptonema and early zygonema cells of juveniles Spo11αki/+ mice than in wild type and Spo11+/- controls. Beside the low level of SPO11α expression, the percentage of cells with a DMC1 focus on the Y-PAR at leptonema was increased considerably compared to Spo11+/- cells; an increase was also observed in cells at early mid-zygonema and early pachynema. This demonstrates that SPO11β function in the PAR is boosted by concomitant expression of SPO11α. Remarkably, the enhancement of DSB generation by this mechanism occurs in animals with a C57 genetic background, indicating that the implementation in DSB formation in the PAR due to splice isoforms co-expression is in addition to the presence of a favorable PAR ultrastructure. Successively, quantification of nucleus wide DSBs in Spo11αki/+ cells at leptonema and early-mid zygonema, revealed the DSBs increased only modestly, compared to the increased frequency of DSBs in the PAR. This underlines the functional specificity of SPO11α for XY chromosomes recombination. In a recent study it was demonstrated that a direct interaction of TOPOVIBL with REC114 is required in males for the formation of DSBs in the subtelomeric regions and at PAR and that the binding of REC114 to TOPOVIBL is mutually exclusive with ANKRD31 [47]. Given that Ankrd31 is essential for the formation of DSBs in the PAR [25, 26], we envision the possibility that the protein complex that leads to DSB formation at the PAR might involve the interaction of SPO11β with TOPOVIBL and REC114 and that of SPO11α with ANKRD31. The latter would possibly be mediated by a (TOPOVI type B-like) protein, which is perhaps expressed with the same timing of SPO11α and preferentially or exclusively binds to ANKRD31. Alternatively, SPO11α interacting with both ANKRD31 and REC114 could serve as an intermediary in the interaction of the SPO11β/TOPOVIBL heterotetramer with the PAR. In this regard, we recently demonstrated that SPO11α co-immunoprecipitates with REC114 in vivo [48], indicating that when this short form of SPO11 is expressed, it interacts with pre-DSB promoting factors, likely promoting DSB activity at the PAR. More studies will be needed to clarify how SPO11 splice isoforms interact dynamically with TOPOVIBL and/or additional type B-like proteins, as well as with RMMAI proteins while cells progress through prophase I.
Defective recombination initiation between XY chromosomes leads to differentiation of aneuploid sperms
One important output of our study is that alterations in the frequency of XY recombination initiation and synapsis closely correlates with the differentiation of XY aneuploid spermatozoa. Therefore, in the long term, understanding of the XY recombination mechanisms at the molecular level has the potential to illuminate the genetic origin of paternally-derived cases of Klinefelter syndrome (47, XXY) [49, 50] and male infertility when this is associated with high levels of XY aneuploidy [51].
Materials and methods
Targeting of Spo11 cDNA
cDNAs of Spo11β-bclI (Spo11βb) or Spo11α-bclI (Spo11αb) splice isoforms and a downstream pA sequence (from the SV40 TpA of pcDNA3.1 vector) were synthetized by Gene art (Thermo Fisher). Each cassette was then cloned into a pPGK-Keo vector, containing the kanamycin/neomycin resistance cassette (Keo), flanked by two lox-P (L) sites, downstream a hybrid intron (HI) structure. After retrieval of genomic DNA (BAC clone #RP23-20N4) into pDTA vector, the HI-cDNA-pA-LKL cassette was inserted into the genome, with deletion of the entire exon 1, to obtain the pDTA Spo11βb and pDTA Spo11αb vectors (Fig. S1A). Following linearization with AsiSI (New England Biolabs), DNA was electroporated in A9 ES cells (129 Sv and C57BL/6N background); mouse core facility, EMBL, Rome. Targeted cells (TA) were identified by southern blotting using the 5’ probe, following AflII (New England Biolabs) digestion and injected into 8 cell-stage C57BL/6N embryos. To remove the LKL cassette, the founder males carrying the TA allele were crossed with Deleter-CRE mice (C57BL/6N background) to obtain mice carrying either the Spo11βb Ki or Spo11αb Ki alleles (Spo11βki/ + or Spo11αki/ +). Cassette removal was verified by Southern blotting, after digestion of genomic DNA with the Afl II restriction enzyme and hybridization with the 5 'probe' (Fig. S1B).
Generation of Spo11βki and Spo11αki mice models
C57/129Spo11βki/− and C57/129Spo11+/- matching controls were obtained by mating C57/129Spo11βki/+ founders with Spo11+/- mice with a mixed (C57BL/6 and 129 Sv) background [5] (Fig. S2A). C57Spo11βki/− mice and C57Spo11+/- matching controls were obtained by first crossing C57/129Spo11βki/+ mice with wild type C57BL/6 for seven generations. Next C57Spo11βki/+ were mated with C57Spo11+/- mice (Fig. S6B). The latter were obtained from C57/129Spo11+/- mice after seven backcrosses in the C57BL/6 background. The backcross of C57/129Spo11βki/− mice into 129 (one backcross), was achieved by first crossing C57/129Spo11βki/− females with wild type 129 males. Next, C57/129Spo11βki/+ and C57/129Spo11± of the F1 were mated with each other (Fig. S2C). C57Spo11αki/ αki mice were obtained by backcrossing C57/129Spo11αki/+ founders into C57 for seven generations. Then, C57Spo11αki/+ males and females were mated with each other. The phenotype of C57Spo11αki/ αki males was compared with that of C57Spo11−/−, obtained by mating mice with C57Spo11+/- genotype. C57Spo11αki/+ and C57Spo11+/+ controls were obtained by mating C57Spo11αki/+ mice. In all cases, to minimize variability from strain background, mice were compared with controls from the same litter or from the same mating involving closely related parents. Each analysis has been made for at least a minimum of 3 animals per genotype.
Genotyping
Genotyping was performed by conventional PCR, using 2X MyTaq Red Mix (Bioline Aurogene, BIO-25044) of tail tip DNA. Primer pairs (Integrated DNA Technologies, IDT) are indicated in supplementary table 2.
Morphometric analysis of the testes
Testis was collected from 45 to 60 dpp old mice. Each animal was euthanized and weighted; testes were removed and weighed as well. The mean between testis weight was calculated and normalized to body weight to minimize the difference in testis size due to mouse physiology.
Histology and immunostaining of tissues sections
The testes and ovaries were collected and fixed overnight (ON) at 4° C in 4% paraformaldehyde (PFA) or Bouin fixatives (Sigma, HT10132). The fixed samples were embedded in paraffin (Thermo SCIENTIFIC Histoplast, 6774006). Sections of 5 μm were stained with periodic acid–Schiff (PAS) (Schiff’s fuchsin sulfite reagent, Sigma, S5133) and with hematoxylin (VWR, 340374 T). Images were captured using a Zeiss Axioskop bright-field microscope equipped with a color CCD camera.
Terminal deoxynucleotidyl transferase dUTP nick end labelling (TUNEL) of testis sections
After deparaffinization and rehydration, sections were treated to unmask the antigenic epitope, using Tris–EDTA citrate buffer, pH 7.8 (UCS Diagnostic, TECH199) for 30 min in steam and subjected to the TUNEL assay, according to the manufacturer’s instructions, using the Roche In Situ Cell Death Detection Kit (POD) (cat. N. 11684817910). To identify stage XII, testis sections were co-stained with anti- pH3 antibody (see Table S3). For each genotype, we analyzed stages XII from at least three testis sections per mouse, cut at 50–80 μM distance from each other. The number of stages XII analyzed for each genotype are as follow: C57/129Spo11βki/− ST = 48; C57Spo11βki/− = 13; C57/129Spo11βki/− HL = 40.
Preparation of spermatocyte chromosome spreads, immunostaining, FISH hybridization and analysis of DMC1 foci in the PAR
The spermatocyte surface chromosome spreads were prepared and stained according to [40]. Primary and secondary antibodies used are listed in Supplementary Tables 3 and 4. Hybridization of the PAR by FISH was performed as previously described [40], labeling the X chromosome probe BAC RP24-500I4 which in mice strains under study hybridizes at the PAR boundary (~ 10.5 kb overlap with the X-PAR and Y-PAR) and extends into the non-homologous part of the X. The X and Y PARs were scored as positive for a DMC1 focus when the following criteria were fulfilled: the DMC1 focus co-localized with the SYCP3 signal and localized to the stretch of SYCP3 staining corresponding to the PAR, identified either by FISH or co-staining with ANKRD31. Images were captured using a Leica CTR6000 digital inverted microscope connected to a charge-coupled device camera and analyzed using the Leica software LAS-AF (Leica) for fluorescent microscopy. Super-resolution analysis was performed using the STEDYCON confocal microscope (Abberior Instruments).
Isolation of sperm and XY FISH
Spermatozoa have been collected from the cauda epididymis as described in [52]. Samples were stored at – 80° C. To perform FISH spermatozoa smears were obtained, fixed through washes in ethanol series, then 10 min in Methanol (Sigma, 32213)/Acetic Acid (VWR 20104.298) (3:1) on ice. Preparations were incubated in 10 mM DTT, 0.1 M Tris–HCl (pH 8.00) for 30 min on ice and air-dried. Mouse X, Y probes (MCEN-XY-10-GRRE, Empire Genomics) probes and chromosome 8 probe (BAC clone RP23126A1, BACPAC Genomics, CA USA) were mixed in the hybridization buffer provided with the Empire Genomics kit. Sex chromosome probes were labelled respectively with green and red fluorescent fluorophores, while the autosomal probe was either colabelled with Alexa Fluor-488 and Alexa Fluor-594 dUTP or labelled with Alexafluor-647 dUTP (Molecular Probes, Invitrogen), following a nick translation assay. Hybridization was performed accordingly to Empire Genomics instructions. Slides were mounted with antifade solution (Vectashield; Vector Laboratories, Newark, CA, USA) containing 1 μg/mL of 4′- 6- diamidino- 2- phenylindole (DAPI).
Slides were analyzed under a motorized fluorescence microscope (Zeiss Axio Imager.M1) equipped with a monochromatic CCD camera (Photometrix, Coolsnap HQ2). Analyses were carried out under a 100X oil immersion objective (N.A. = 1.30). For capture and image analysis the MetaMorph software (7.1.3.0, Molecular Device) and the MetaVue software (7.8.11.0, Molecular device) were respectively used.
Immunoprecipitation of SPO11 and Western blot analysis
Immunoprecipitation and Western blot have been performed according to [48, 53]. Briefly, testes from adult or juvenile mice were decapsulated and lysed using the Pierce IP Lysis Buffer (Thermo Fisher Scientific, 87787) complemented with proteases inhibitors 2X (Roche, cOmplete Tablets EDTA-free, 04693132001), phosphatases inhibitors 1X (Sigma-Aldrich, Phosphatase Inhibitor Cocktail 3, P0044) and benzonase (ChemCruz, sc-202391A) according to manufacturer instructions. Supernatants were incubated with Dynabeads Protein-A (Thermo Fisher Scientific, 1002D) loaded with the mouse monoclonal anti-SPO11-180 antibody (table S3) which recognizes specifically both SPO11β and SPO11α isoforms [14], in rotation at 4 °C. Mouse anti-IgG2A (table S3) served as a control. At the end of incubation, the dynabeads were washed three times with Lysis buffer and eluted with standard Laemmli buffer. The samples were fractionated on 8–12% SDS-PAGE and transferred to a PVDF membrane (GE Healthcare, Amersham Hybond P Western blotting membranes, GE10600023) using a semi-dry transfer system (Hoefer, TE22). For Western Blot (WB) analysis, membranes were probed with primary antibodies diluted in BSA 5%/TBS 0.1% Tween 20 (TBS-T). Secondary antibodies were diluted in 5% nonfat dry milk (AppliChem, A0830)/TBS-T. The primary and secondary antibodies used are indicated in supplementary tables S3 and S4. WB signals were detected using ECL reagent (BIO-RAD, Clarity Western ECL Substrate, #170–5061). Quantification of SPO11 protein level was performed by densitometry using ImageJ software. Values were normalized against SYCP3/tubulin or REC8/tubulin ratio in total extracts. SYCP3 or REC8 were used as markers for spermatocytes content in the testis.
Analysis of PAR ultrastructure
PAR loops and axis lengths were measured according to the method described by Acquaviva et al. [23].
Statistical analysis
Statistical analysis was performed using GraphPad Prism 9 for Macintosh (GraphPad Software, San Diego, CA). Data were expressed as mean ± SD or mean ± SEM, as detailed in the figure captions.
Artwork
The artwork was created with Adobe Photoshop and Illustrator 2022.
Availability of data and material
The authors confirm that the data supporting the findings of this study are available in the article and its supplementary materials.
References
Szostak JW, Orr-Weaver TL, Rothstein RJ, Stahl FW. The double-strand-break repair model for recombination. Cell. 1983;33(1):25–35. Epub 1983/05/01. doi: https://doi.org/10.1016/0092-8674(83)90331-8. PubMed PMID: 6380756.
Sun H, Treco D, Schultes NP, Szostak JW. Double-strand breaks at an initiation site for meiotic gene conversion. Nature. 1989;338(6210):87–90. Epub 1989/03/02. doi: https://doi.org/10.1038/338087a0. PubMed PMID: 2645528.
Cao L, Alani E, Kleckner N. A pathway for generation and processing of double-strand breaks during meiotic recombination in S. cerevisiae. Cell. 1990;61(6):1089–101. Epub 1990/06/15. doi: https://doi.org/10.1016/0092-8674(90)90072-m. PubMed PMID: 2190690.
Keeney S, Giroux CN, Kleckner N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell. 1997;88(3):375–84. Epub 1997/02/07. doi: https://doi.org/10.1016/s0092-8674(00)81876-0. PubMed PMID: 9039264.
Baudat F, Manova K, Yuen JP, Jasin M, Keeney S. Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell. 2000;6(5):989–98. Epub 2000/12/07. doi: https://doi.org/10.1016/s1097-2765(00)00098-8. PubMed PMID: 11106739.
Romanienko PJ, Camerini-Otero RD. The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell. 2000;6(5):975–87. Epub 2000/12/07. doi: https://doi.org/10.1016/s1097-2765(00)00097-6. PubMed PMID: 11106738.
Mahadevaiah SK, Turner JM, Baudat F, Rogakou EP, de Boer P, Blanco-Rodriguez J et al (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27(3):271–276. https://doi.org/10.1038/85830. (Epub 2001/03/10. PubMed PMID: 11242108)
Bergerat A, de Massy B, Gadelle D, Varoutas PC, Nicolas A, Forterre P (1997) An atypical topoisomerase II from archaea with implications for meiotic recombination. Nature 386(6623):414–417. https://doi.org/10.1038/386414a0. (Epub 1997/03/27. PubMed PMID: 9121560)
de Massy B (2013) Initiation of meiotic recombination: how and where? Conservation and specificities among eukaryotes. Annu Rev Genet 47:563–599. https://doi.org/10.1146/annurev-genet-110711-155423. (Epub 2013/09/21. PubMed PMID: 24050176)
Robert T, Nore A, Brun C, Maffre C, Crimi B, Bourbon HM et al (2016) The TopoVIB-like protein family is required for meiotic DNA double-strand break formation. Science 351(6276):943–949. https://doi.org/10.1126/science.aad5309. (Epub 2016/02/27. PubMed PMID: 26917764)
La Volpe A, Barchi M (2012) Meiotic double strand breaks repair in sexually reproducing eukaryotes: we are not all equal. Exp Cell Res 318(12):1333–1339. https://doi.org/10.1016/j.yexcr.2012.03.014. (Epub 2012/04/07. PubMed PMID: 22480866)
Handel MA, Schimenti JC (2010) Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nat Rev Genet 11(2):124–136. https://doi.org/10.1038/nrg2723. (Epub 2010/01/07. PubMed PMID: 20051984)
Hunter N (2015) Meiotic recombination: the essence of heredity. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a016618. (Epub 2015/10/30. PubMed PMID: 26511629; PubMed Central PMCID: PMCPMC4665078)
Kauppi L, Barchi M, Baudat F, Romanienko PJ, Keeney S, Jasin M (2011) Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331(6019):916–920. https://doi.org/10.1126/science.1195774. (Epub 2011/02/19. PubMed PMID: 21330546; PubMed Central PMCID: PMCPMC3151169)
Romanienko PJ, Camerini-Otero RD (1999) Cloning, characterization, and localization of mouse and human SPO11. Genomics 61(2):156–169. https://doi.org/10.1006/geno.1999.5955. (PubMed PMID: 10534401)
Keeney S, Baudat F, Angeles M, Zhou ZH, Copeland NG, Jenkins NA et al (1999) A mouse homolog of the saccharomyces cerevisiae meiotic recombination DNA transesterase Spo11p. Genomics 61(2):170–182. https://doi.org/10.1006/geno.1999.5956. (PubMed PMID: 10534402)
Carofiglio F, Inagaki A, de Vries S, Wassenaar E, Schoenmakers S, Vermeulen C et al (2013) SPO11-independent DNA repair foci and their role in meiotic silencing. PLoS Genet 9(6):e1003538. https://doi.org/10.1371/journal.pgen.1003538. (Epub 2013/06/12. PubMed PMID: 23754961; PubMed Central PMCID: PMCPMC3675022)
Faisal I, Kauppi L (2017) Reduced MAD2 levels dampen the apoptotic response to non-exchange sex chromosomes and lead to sperm aneuploidy. Development 144(11):1988–1996. https://doi.org/10.1242/dev.149492. (Epub 2017/05/17. PubMed PMID: 28506992)
Kumar R, Bourbon HM, de Massy B (2010) Functional conservation of Mei4 for meiotic DNA double-strand break formation from yeasts to mice. Genes Dev 24(12):1266–1280. https://doi.org/10.1101/gad.571710. (Epub 2010/06/17. PubMed PMID: 20551173; PubMed Central PMCID: PMCPMC2885662)
Stanzione M, Baumann M, Papanikos F, Dereli I, Lange J, Ramlal A et al (2016) Meiotic DNA break formation requires the unsynapsed chromosome axis-binding protein IHO1 (CCDC36) in mice. Nat Cell Biol 18(11):1208–1220. https://doi.org/10.1038/ncb3417. (Epub 2016/10/28. PubMed PMID: 27723721; PubMed Central PMCID: PMCPMC5089853)
Kumar R, Oliver C, Brun C, Juarez-Martinez AB, Tarabay Y, Kadlec J et al (2018) Mouse REC114 is essential for meiotic DNA double-strand break formation and forms a complex with MEI4. Life Sci Alliance. 1(6):e201800259. https://doi.org/10.26508/lsa.201800259. (Epub 2018/12/21. PubMed PMID: 30569039; PubMed Central PMCID: PMCPMC6288613)
Reinholdt LG, Schimenti JC (2005) Mei1 is epistatic to Dmc1 during mouse meiosis. Chromosoma 114(2):127–134. https://doi.org/10.1007/s00412-005-0346-4. (Epub 2005/06/02. PubMed PMID: 15928951)
Acquaviva L, Boekhout M, Karasu ME, Brick K, Pratto F, Li T et al (2020) Ensuring meiotic DNA break formation in the mouse pseudoautosomal region. Nature 582(7812):426–431. https://doi.org/10.1038/s41586-020-2327-4. (Epub 2020/05/29. PubMed PMID: 32461690)
Libby BJ, De La Fuente R, O’Brien MJ, Wigglesworth K, Cobb J, Inselman A et al (2002) The mouse meiotic mutation mei1 disrupts chromosome synapsis with sexually dimorphic consequences for meiotic progression. Dev Biol 242(2):174–187. https://doi.org/10.1006/dbio.2001.0535. (Epub 2002/02/01. PubMed PMID: 11820814)
Boekhout M, Karasu ME, Wang J, Acquaviva L, Pratto F, Brick K et al (2019) REC114 partner ANKRD31 controls number, timing, and location of meiotic DNA breaks. Mol Cell 74(5):1053–1688. https://doi.org/10.1016/j.molcel.2019.03.023. (Epub 2019/04/21. PubMed PMID: 31003867; PubMed Central PMCID: PMCPMC6555648)
Papanikos F, Clement JAJ, Testa E, Ravindranathan R, Grey C, Dereli I et al (2019) Mouse ANKRD31 regulates spatiotemporal patterning of meiotic recombination initiation and ensures recombination between X and Y sex chromosomes. Mol Cell 74(5):1069-8511 e11. https://doi.org/10.1016/j.molcel.2019.03.022. (Epub 2019/04/20. PubMed PMID: 31000436)
Lorenz A, Estreicher A, Kohli J, Loidl J (2006) Meiotic recombination proteins localize to linear elements in Schizosaccharomyces pombe. Chromosoma 115(4):330–340. https://doi.org/10.1007/s00412-006-0053-9. (Epub 2006/03/15. PubMed PMID: 16532353)
Miyoshi T, Ito M, Kugou K, Yamada S, Furuichi M, Oda A et al (2012) A central coupler for recombination initiation linking chromosome architecture to S phase checkpoint. Mol Cell 47(5):722–733. https://doi.org/10.1016/j.molcel.2012.06.023. (Epub 2012/07/31. PubMed PMID: 22841486)
Panizza S, Mendoza MA, Berlinger M, Huang L, Nicolas A, Shirahige K et al (2011) Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell 146(3):372–383. https://doi.org/10.1016/j.cell.2011.07.003. (Epub 2011/08/06. PubMed PMID: 21816273)
Blat Y, Protacio RU, Hunter N, Kleckner N (2002) Physical and functional interactions among basic chromosome organizational features govern early steps of meiotic chiasma formation. Cell 111(6):791–802. https://doi.org/10.1016/s0092-8674(02)01167-4. (Epub 2003/01/16. PubMed PMID: 12526806)
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754. https://doi.org/10.1146/annurev.genet.33.1.603. (Epub 2000/02/26. PubMed PMID: 10690419)
Barchi M, Mahadevaiah S, Di Giacomo M, Baudat F, de Rooij DG, Burgoyne PS et al (2005) Surveillance of different recombination defects in mouse spermatocytes yields distinct responses despite elimination at an identical developmental stage. Mol Cell Biol 25(16):7203–7215. https://doi.org/10.1128/MCB.25.16.7203-7215.2005. (Epub 2005/08/02. PubMed PMID: 16055729; PubMed Central PMCID: PMCPMC1190256)
Page SL, Hawley RS (2004) The genetics and molecular biology of the synaptonemal complex. Annu Rev Cell Dev Biol 20:525–558. https://doi.org/10.1146/annurev.cellbio.19.111301.155141. (Epub 2004/10/12. PubMed PMID: 15473851)
Russell LD, Ettlin R, Hikim AP, Cledgg ED (1990) Histological and histopatological evaluation of the testis. Cache River Press, Clerwater, FL
Ahmed EA, de Rooij DG (2009) Staging of mouse seminiferous tubule cross-sections. Methods Mol Biol 558:263–277. https://doi.org/10.1007/978-1-60761-103-5_16. (PubMed PMID: 19685330)
Muciaccia B, Boitani C, Berloco BP, Nudo F, Spadetta G, Stefanini M et al (2013) Novel stage classification of human spermatogenesis based on acrosome development. Biol Reprod 89(3):60. https://doi.org/10.1095/biolreprod.113.111682. (Epub 2013/08/16. PubMed PMID: 23946533)
Edelmann W, Cohen PE, Kane M, Lau K, Morrow B, Bennett S et al (1996) Meiotic pachytene arrest in MLH1-deficient mice. Cell 85(7):1125–1134. https://doi.org/10.1016/s0092-8674(00)81312-4. (Epub 1996/06/28. PubMed PMID: 8674118)
Kauppi L, Barchi M, Lange J, Baudat F, Jasin M, Keeney S (2013) Numerical constraints and feedback control of double-strand breaks in mouse meiosis. Genes Dev 27(8):873–886. https://doi.org/10.1101/gad.213652.113. (PubMedPMID:23599345;PubMedCentralPMCID:PMCPMC3650225)
Faieta M, Di Cecca S, de Rooij DG, Luchetti A, Murdocca M, Di Giacomo M et al (2016) A surge of late-occurring meiotic double-strand breaks rescues synapsis abnormalities in spermatocytes of mice with hypomorphic expression of SPO11. Chromosoma 125(2):189–203. https://doi.org/10.1007/s00412-015-0544-7. (Epub 2015/10/07. PubMed PMID: 26440409; PubMed Central PMCID: PMCPMC4830894)
Testa E, Nardozi D, Antinozzi C, Faieta M, Di Cecca S, Caggiano C et al (2018) H2AFX and MDC1 promote maintenance of genomic integrity in male germ cells. J Cell Sci. https://doi.org/10.1242/jcs.214411. (Epub 2018/02/14. PubMed PMID: 29437857)
Pittman DL, Schimenti KJ, Hanneman WH, Schimenti JC (1999) Transgenic and mutational analyses of meiotic recombination in mice. Ann N Y Acad Sci 870:220–222. https://doi.org/10.1111/j.1749-6632.1999.tb08882.x. (Epub 1999/07/23. PubMed PMID: 10415485)
Neale MJ, Pan J, Keeney S (2005) Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature 436(7053):1053–1057. https://doi.org/10.1038/nature03872. (Epub 2005/08/19. PubMed PMID: 16107854; PubMed Central PMCID: PMCPMC1262668)
Bellani MA, Boateng KA, McLeod D, Camerini-Otero RD (2010) The expression profile of the major mouse SPO11 isoforms indicates that SPO11beta introduces double strand breaks and suggests that SPO11alpha has an additional role in prophase in both spermatocytes and oocytes. Mol Cell Biol 30(18):4391–4403. https://doi.org/10.1128/MCB.00002-10. (PubMedPMID:20647542;PubMedCentralPMCID:PMCPMC2937527)
Faisal I, Kauppi L (2016) Sex chromosome recombination failure, apoptosis, and fertility in male mice. Chromosoma 125(2):227–235. https://doi.org/10.1007/s00412-015-0542-9. (Epub 2015/10/07. PubMed PMID: 26440410)
Humphrey E, Cole F (2020) How sex chromosomes break up to get together. Nature 582(7812):346–347. https://doi.org/10.1038/d41586-020-01483-6. (Epub 2020/06/17. PubMed PMID: 32541910)
Robert T, Vrielynck N, Mezard C, de Massy B, Grelon M (2016) A new light on the meiotic DSB catalytic complex. Semin Cell Dev Biol 54:165–176. https://doi.org/10.1016/j.semcdb.2016.02.025. (Epub 2016/03/21. PubMed PMID: 26995551)
Nore A, Juarez-Martinez AB, Clement J, Brun C, Diagouraga B, Laroussi H et al (2022) TOPOVIBL-REC114 interaction regulates meiotic DNA double-strand breaks. Nat Commun 13(1):7048. https://doi.org/10.1038/s41467-022-34799-0. (Epub 2022/11/18. PubMed PMID: 36396648; PubMed Central PMCID: PMCPMC9671922)
Giannattasio T, Testa E, Palombo R, Chellini L, Franceschini F, Crevenna A et al (2023) The RNA-binding protein FUS/TLS interacts with SPO11 and PRDM9 and localize at meiotic recombination hotspots. Cell Mol Life Sci 80(4):107. https://doi.org/10.1007/s00018-023-04744-5. (Epub 2023/03/27. PubMed PMID: 36967403; PubMed Central PMCID: PMCPMC10040399)
Arnedo N, Templado C, Sanchez-Blanque Y, Rajmil O, Nogues C (2006) Sperm aneuploidy in fathers of Klinefelter’s syndrome offspring assessed by multicolour fluorescent in situ hybridization using probes for chromosomes 6, 13, 18, 21, 22. X and Y. Hum Reprod. 21(2):524–528. https://doi.org/10.1093/humrep/dei321. (Epub 2005/10/04. PubMed PMID: 16199425)
Thomas NS, Hassold TJ (2003) Aberrant recombination and the origin of Klinefelter syndrome. Hum Reprod Update 9(4):309–317. https://doi.org/10.1093/humupd/dmg028. (Epub 2003/08/21. PubMed PMID: 12926525)
Templado C, Uroz L, Estop A (2013) New insights on the origin and relevance of aneuploidy in human spermatozoa. Mol Hum Reprod 19(10):634–643. https://doi.org/10.1093/molehr/gat039. (Epub 2013/05/31. PubMed PMID: 23720770)
Adler ID, Pacchierotti F, Russo A (2012) The measurement of induced genetic change in mammalian germ cells. Methods Mol Biol 817:335–375. https://doi.org/10.1007/978-1-61779-421-6_16. (Epub 2011/12/08. PubMed PMID: 22147580)
Parvanov ED, Tian H, Billings T, Saxl RL, Spruce C, Aithal R et al (2017) PRDM9 interactions with other proteins provide a link between recombination hotspots and the chromosomal axis in meiosis. Mol Biol Cell 28(3):488–499. https://doi.org/10.1091/mbc.E16-09-0686. (Epub 2016/12/10. PubMed PMID: 27932493; PubMed Central PMCID: PMCPMC5341731)
Acknowledgements
The authors are grateful to Elisa Palumbo (University of Padova, Italy) for her technical assistance in the early stages of aneuploidy analysis in spermatozoa. We are also grateful to Scott Keeney and Maria Jasin (Memorial Sloan-Kettering Cancer Centre, USA) for providing Spo11+/- mice and the anti SPO11-180 antibody, Bernard De Massy (Centre National de la Recherche Scientifique, France) and Attila Toth (Technische Universität Dresden, Germany) for the generous gift of antibodies, and Willy Baarends (Erasmus University Medical Center, Netherland) for critical reading of the first draft of the manuscript.
Funding
Open access funding provided by Università degli Studi di Roma Tor Vergata within the CRUI-CARE Agreement. This work was supported by the Telethon grant n. GGP12189 (MB). Additional sources came from the “Mission Sustainability” grant n. 141, from the University of Rome Tor Vergata (MB), grant “Gruppi di Ricerca 2020” from Regione Lazio, Italy (n. A0375-2020–36618) to MB, and Fondo di Beneficenza Intesa Sanpaolo (n. B/2021/0228) to MB. The founders had no role in study design, data collection and analysis, decision to publish, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
TG, ET, and MB contributed to the study conception and design. Material preparation, data collection, and analysis were performed by TG, ET, MF, ML, DN, SC and AR. The work was carried out under the supervision of MB. The first draft of the manuscript was written by MB and was reviewed by all authors. MB made the changes to the manuscript after peer review. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval and consent to participate
The study was approved by the National Institute of Health of Italy (Istituto Superiore di Sanità). All procedures involving animals were in compliance with the European Community Council Directive of 24 November 1986, and the approval by the research ethics committee of the University of Rome Tor Vergata.
Consent for publication
The research did not involve human participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Giannattasio, T., Testa, E., Faieta, M. et al. The proper interplay between the expression of Spo11 splice isoforms and the structure of the pseudoautosomal region promotes XY chromosomes recombination. Cell. Mol. Life Sci. 80, 279 (2023). https://doi.org/10.1007/s00018-023-04912-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-023-04912-7