Abstract
Sex chromosome aneuploidies are among the most common variations in human whole chromosome copy numbers, with an estimated prevalence in the general population of 1:400 to 1:1400 live births. Unlike whole-chromosome aneuploidies of autosomes, those of sex chromosomes, such as the 47, XXY aneuploidy that causes Klinefelter Syndrome (KS), often originate from the paternal side, caused by a lack of crossover (CO) formation between the X and Y chromosomes. COs must form between all chromosome pairs to pass meiotic checkpoints and are the product of meiotic recombination that occurs between homologous sequences of parental chromosomes. Recombination between male sex chromosomes is more challenging compared to both autosomes and sex chromosomes in females, as it is restricted within a short region of homology between X and Y, called the pseudo-autosomal region (PAR). However, in normal individuals, CO formation occurs in PAR with a higher frequency than in any other region, indicating the presence of mechanisms that promote the initiation and processing of recombination in each meiotic division. In recent years, research has made great strides in identifying genes and mechanisms that facilitate CO formation in the PAR. Here, we outline the most recent and relevant findings in this field. XY chromosome aneuploidy in humans has broad-reaching effects, contributing significantly also to Turner syndrome, spontaneous abortions, oligospermia, and even infertility. Thus, in the years to come, the identification of genes and mechanisms beyond XY aneuploidy is expected to have an impact on the genetic counseling of a wide number of families and adults affected by these disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction to meiosis: key concepts
A fundamental property of life is the ability to reproduce. In sexually reproducing higher eukaryotes, the generation of gametes, which are sperm and eggs, occurs through meiosis. Meiosis is a biological process in which germ cells, after a round of DNA replication, divide twice, halving DNA content. Meiosis not only grants the re-establishment of diploidy of the embryo after fertilization, but also allows for exchange of genetic material between maternal and paternal chromosomes. This genomic shuffling creates new genetic combinations that ultimately contribute to genetic diversity, leading to the creation of increasingly robust or specialized offspring. The key biological mechanism at the base of genetic reassortment is the homologous recombination (HR) of repair of DNA double strand breaks (DSBs), which are physiologically introduced into the genome during meiotic prophase I (Fig. 1A-B). Unlike mitosis, meiotic HR preferentially utilizes the homologous chromosome over the sister chromatid, as template for DSB repair. This may result in reciprocal exchange of genetic material between regions with strong similarity, forming crossovers (COs) [1]. In mice, DSBs are generated by SPO11 (the ortholog of subunit A of TopoVI DNA topoisomerase) [2,3,4,5,6,7] in complex with the TopoVI B-like subunit (TOPOVIBL) [7]. The Spo11 gene is conserved in humans [2, 5] and single nucleotide polymorphisms of SPO11 are associated with male infertility and decreased ovarian reserve [8,9,10,11]. Key TOPOVIBL domains are also conserved in humans [7], suggesting a shared mechanism for the formation of DSBs between species. In mouse spermatocytes, DSBs start to be made at leptonema, peak at zygonema, and decrease in number as cell progresses to pachynema. DSBs are distributed along the entire length of homologous autosomes (the homologous), each formed by two sister chromatids. Conversely, given that mouse (and human) sex chromosomes are heteromorphic, DSB formation and recombination is restricted within a short region of homology between them, the PAR, which genetically behave like autosomes. Mice have a single PAR (mPAR) [12], while human sex chromosomes have two PARs: PAR1 and PAR2. PAR1 corresponds to mPAR, as CO occurs only rarely in PAR2 [13].
In autosomes, only about 10–25% of DSBs is repaired by HR with the formation of COs [14,15,16]. COs produce new combinations of DNA sequences, resulting in enhanced genetic variation. The remaining DSBs are repaired by interhomolog recombination without reciprocal exchange, resulting in the formation of non-crossover (NCO) products [1, 15, 17, 18]. The latter, by allowing transmission distortion of genetic information and mutations, are also considered to provide a significant contribution to genome evolution [15, 19], and are essential to guarantee DSB-driven alignment and synapsis of the homologous (see below). Repair of DSBs with the formation of a CO occurs with reciprocal DNA exchange between parental chromosomes [15, 20]. By doing so, in addition to shuffling the genome, these events lead to the formation of interhomolog DNA links that are cytologically identifiable as chiasmata [21,22,23]. Chiasmata, by physically linking homologous, play a key role in the co-orientation of the sisters of each homolog to opposite spindle poles [24], counteracting forces exerted by the centromere–attached microtubules. This ensures proper alignment of homologous at the metaphase I spindle and segregation in daughter cells. If CO formation fails, meiotic chromosomes segregate randomly, with consequent meiotic arrest and infertility due to activation of the Spindle Assembly Checkpoint (SAC) mechanisms of selection [25, 26]. Hence, each homologous pair (bivalent) requires at least one CO for chromosomes to segregate properly [20], known as the “obligatory CO”. In male sex chromosomes, formation of the obligatory CO is restricted to the PAR. A failure in the generation of at least one DSB in the X-PAR and/or Y-PAR results in missegregation of the sex chromosomes. In mice, according to phenotypic penetrance, this can lead to infertility [27] or sub fertility [28], with generation of aneuploid sperm for the sex chromosomes [29, 30]. In men, infertile, oliogozospermic and oligoasthenoteratozoospermic patients manifest significantly higher levels of XY disomy [31,32,33], probably due to reduced recombination in the PAR [32, 34]. The lack of CO formation between the XY chromosomes also occurs in fathers with progeny of Klinefelter Syndrome (KS) (47, XXY) [35,36,37], which is the most common (1:500–1:1000 [38]) sex chromosome disorder of paternal origin. Nondisjunction of XY chromosomes also likely cause of fathering a progeny with Turner syndrome (TS). Therefore, understanding the genetics of XY recombination in mouse and humans is key to understand the etiology of sex chromosome disorders.
Recombination-independent mechanisms of XY chromosome pairing and synapsis
Meiotic prophase I differs substantially from mitotic prophase, not only because of the formation of COs but also because of the process that drives pairing and synapsis of parental chromosomes. Pairing between the homolog pairs (that is, approaching and juxtaposing of the chromosomes) begins after DNA duplication at preleptonema (prophase I), prior to SPO11-mediated DNA cleavage [39]. This recombination-independent mechanism requires an unknown DSB-independent activity of SPO11 and that of SUN1, a nuclear membrane protein that binds telomeres to the nuclear envelope [40]. The occurrence of pairing most likely facilitates the initiation of synapsis [39], that is, stable axial alignment of the homologous, which begins with the formation of DSBs and the assembly of the synaptonemal complex (SC) (see below). Interestingly, a recent study pointed out that homologous pairing at spermatogonia-early preleptotene stage occurs in over 70% of the mouse homologous, including the XY chromosomes. In the latter, pairing frequency further increases by mid-preleptotene stage occurring in over 85% of the cells [41]. This suggests that pairing promotes spatial proximity between the XY chromosomes, as proposed for non-sex chromosomes [39, 41]. However, it is unclear whether this proximity may play a role in the promotion of synapsis between the XY chromosomes, as synapsis does not occur until late zygotene [27]. Furthermore, whether the DSB independent function of SPO11 and that of SUN1 are required for the XY pairing has not been investigated. A key feature of meiotic cells is the assembly of the SC, a tripartite zipper-like proteinaceous structure that mediates homologous chromosome alignment and synapsis during prophase I [42,43,44]. Under physiological conditions the SC is formed along the full length of autosomes and at the PAR of bivalents that are undergoing recombination (Fig. 1A). Nevertheless, the SC also assembles in short stretches, between non-homologous chromosomes of DSB-deficient mutants, such as Spo11−/− [4, 6, 45], stabilizing synapsis, locally. Therefore, the SC also promotes recombination-independent synapsis, a function that lies in the ability of SC proteins to undergo self-assembly [46, 47]. An additional accessory mechanism that probably promotes spatial proximity between XY is the formation of the sex body. The sex body is a chromatin domain resulting From Meiotic Sex Chromosome Inactivation (MSCI), the process that silences XY-associated genes of heterologous (asynapsed) XY chromosomes regions at pachynema (see [48] and references therein). A key event in MSCI is phosphorylation of histone H2AX at serine 139 (γH2AX) (Fig. 1B and Fig. 2A), which attracts MDC1 (mediator of DNA damage checkpoint 1), a γH2AX binding partner. MDC1 promotes spreading of γH2AX to chromatin loops (Fig. 2B), for effective silencing of XY associated genes [48, 49]. In absence of H2ax X and Y are asynapsed (Fig. 2C) with a higher frequency compared to that of Mdc1−/− cells, in which the sex body formation fails only partially (Fig. 2B) [49,50,51]. Therefore, it is likely that nuclear compartmentalization of XY chromosomes elicited by the formation of the sex body plays a role in bringing the XY closer, promoting PARs proximity, pairing and synapsis [51]; possibly suppressing illegitimate recombination of non-homologous regions [52].
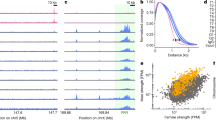
DSB formation and assembly of the SC during meiotic prophase I. A In leptonema of normal meiotic prophase, the sister chromatids of each chromosome develop a proteinaceous axis (the SYCP3-positive axial element), with chromatin extending out in loops. Associations form between the axes of homologous chromosomes and extend progressively during the zygotene stage, forming the tripartite Synaptonemal Complex (SC) that comprises both axial elements (red) and transverse filaments of the central element (orange) that connects them. Synapsis is completed by pachynema, so that the SC joins homologous chromosomes along their entire lengths. Meiotic recombination involves the formation and repair of DSBs (stars) that start to be made at leptonema up to zygonema on asynapsed autosomal homologous. B In the mouse, formation of the SC and synapsis of the homologous is monitored cytologically on prophase I meiotic chromosomes spreads, by staging them with an anti-SYCP3 antibody (red). DSBs are typically monitored by examining γH2AX [53], a phosphorylated form of histone H2AX, which appears on chromosomes in early meiotic prophase I (i.e., from leptonema to zygonema) in a SPO11-dependent fashion [4, 6]. At pachynema and diplonema γH2AX persists only on the sex chromosomes as the part of the MSCI mechanism [48]
Chromosome spreads of mouse spermatocytes at pachynema. A In wild type spermatocytes, the SC axial element SYCP3 (red) extends along fully paired autosomes, while the XY chromosomes only synapse at PAR. The X and Y chromosomes are embedded within the sex body, a chromosome domain identified by the phosphorylation of histone H2ax (γH2AX, green). B In Mdc1−/− spermatocytes sex body formation partially fails, due to inefficient spreading of γH2AX to chromatin loops [49]. XY chromosomes synapse in a large fraction of cells, although with reduced proficiency compared to wild type [51]. C In H2ax−/− spermatocytes, sex body formation fails [50], and the X and Y chromosomes (labelled with the antibody that recognizes the phosphorylation status of the HORMA [Hop1, Rev7 and Mad2] domain one protein [HORMAD1] [164]; white) are unsynapsed with a higher percentage compared to Mdc1−/− cells [51]. In A and B, white arrows head point to the PAR
Recombination-dependent mechanism of chromosome pairing and synapsis
In mice carrying the mutation of the catalytic tyrosine of Spo11 (Y/F substitution), synapsis of the homologous is disrupted, indicating that pairing achieved through recombination–independent mechanisms (see above) is dependent on the DSB activity of SPO11 [39]. In accordance, previous studies have demonstrated that DSB formation precedes synapsis of the homologous [53], and that when the expression level of SPO11 is reduced below a critical threshold, and DSB numbers decrease considerably, homolog synapsis fails, with the consequent elimination of defective spermatocytes by apoptosis [54, 55]. In a simplified view of the events driving the synapse between homologs, formation of DSBs by SPO11 is followed by processing of DSBs into 3’ ssDNA ends, which are required for the search of the complementary sequences in the homologous chromosome, leading to interhomolog interactions pairing and synapsis, with the assembly of the SC [1] (Fig. 1A).
Despite DSB numbers may vary considerably among spermatocytes [15, 56], proper establishment of synapsis between autosomes always requires that DSBs are initiated at multiple sites along chromosome length (Fig. 1A). It is believed that starting homology-dependent DNA interactions from multiple recombination events enforces homolog pairing during meiotic prophase [57], while suppressing interactions between non-allelic homologous sequences [58]. For this reason, the DSB numbers per cell (> 200 on average in mice and humans) [14, 15, 27, 55] substantially exceed those of COs. Under these circumstances, smaller autosomes and sex chromosomes which physiologically receive a low number of DSBs, are particularly vulnerable to DSBs decrement, engaging in non-homologous synapsis [54]. Thus, a robust wave of DSBs at leptonema and zygonema paves the way for the success of meiosis.
DSB formation in the PAR
Based on the average frequency of DSBs in mouse spermatocytes it is estimated that fewer than one DSB form per ten Megabase pairs [27]. The PAR is ~ 0.7 Megabase long [59, 60]. This would predict that the PAR receives fewer than one DSB for every ten meioses (that is, a > 90% failure rate). In contrast, PAR experiences one or two DSBs per meioses [27, 61, 62]. Therefore, the formation of DSBs in the PAR is 10–20 times higher than the average autosomal region [27]. It follows that there must be mechanisms in place that implement the probability that SPO11 is recruited and active in this region. In addition to the frequency, the timing with which DSBs are formed in autosomes and in the mPAR is also different. In autosomes, DSBs start to form at leptonema, peak in number at zygonema, and decrease by late zygonema and pachynema. Conversely, a DSB is detected in the X-mPAR and Y-mPAR, more frequently at late zygonema [27]. The different timings of DSB formation in autosomes and sex chromosomes correlate, in mice, with the expression of two splicing isoforms of Spo11: Spo11β and Spo11α [2, 5]. SPO11β is expressed earlier, by leptonema, when DSB formation starts nucleus-wide on autosomes, whereas SPO11α starts to be expressed (concomitantly with Spo11β) in late prophase I [2, 5, 63]. These two isoforms differ from each other for the inclusion (Spo11β) or skipping (Spo11α) of exon2 [2, 5, 63]. The ratio of Spo11β and Spo11α isoforms is dependent on modulation of RNA polymerase II and the recruitment of splicing factors [64, 65]. Consistent with their timing of expression, it has been proposed that SPO11β is required for DSB formation on autosomes, while SPO11α on the PAR. Using the mouse as a model system, Kauppi et al. demonstrated that transgenic expression of SPO11β in a Spo11−/− genetic background (Spo11β-only transgenic mouse), complements autosomal synapsis defects observed in Spo11−/− mice [4, 6], while XY chromosomes remain asynapsed, with consequent infertility [27]. However, a subsequent study using the same animal model showed that defective XY synapsis can be reduced to 50% in mice with a different genetic background, and males are fertile [28]. This indicates that, other than SPO11β, the expression of other Spo11 splice isoforms is not essential for male fertility. This calls into question what the function of SPO11α is, and why genetic background changes impact the proficiency of XY recombination and synapsis. Using a knock-in mouse model that expresses only SPO11β under its physiological promoter (Spo11βki-only mouse) we confirmed that DSB frequency in the PAR changes with genetic background. Moreover, we observed that it fails more frequently in the Y-PAR than the X-PAR [30]. Furthermore, we demonstrated using the Spo11αki mouse model, that concomitant expression of SPO11α with SPO11β strongly boost formation of DSBs in the Y-PAR, regardless of the genetic background [30]. Therefore, it can be concluded that the expression of the single SPO11β isoform increases the risk of XY asynapsis. To date, the exact molecular mechanism behind the cooperation of Spo11 splice isoforms remains unknown. According to in vitro protein–protein interaction experiments, SPO11α is unable to interact with TOPOVIBL [7]. Thus, it is assumed that in vivo, SPO11α cannot form heterotetramers with SPO11β and TOPOVIBL. It has been hypothesized that SPO11α may titrate an inhibitor of TOPOVIBL, for example via a protein–protein interaction, raising the possibility that the heterotetramers formed by SPO11β and TOPOVIBL [7] form the DSB on PAR, in late zygonema [66]. However, this remains speculation.
Epigenetic determination of DSBs hotspot
Mammalian meiotic DSBs are not randomly distributed but they occur preferentially in genomic regions called “hotspots”, which are positioned by the (widely conserved) mouse meiosis-specific methyltransferase PRDM9 (PR domain-containing 9) protein [67,68,69]. PRDM9 is a zinc finger protein that, through interaction with the HELLS chromatin remodeler binds DNA [70, 71] and trimethylates histone 3 in Lysine-4 (H3K4me3) and Lysine-36 (H3K36me3) in nearby nucleosomes [72,73,74], providing access to the DSB initiating complex in nucleosome-depleted regions [68, 69, 73, 75,76,77]. Meiotic DSBs form in normal numbers in mouse spermatocytes with inactivated Prdm9, but occur at “default sites”, which are PRDM9-independent H3K4me3 enriched regions (such as promoters and enhancers) that are rarely targeted in wild type mice [75, 76]. This relocation parallels a defect in the repair of DSBs, with a consequent failure of synapsis between homologues and may result in sterility [75, 76, 78, 79]. Heterozygosity of Prdm9 also leads to sterility in some hybrid mice [80], while genetic background shift in mice with Prdm9 loss of function mutation partially restores fertility [78, 81]. These results suggest the presence of genetic modifiers of PRDM9 function, for example the expression, in specific genetic contexts, of other chromatin modifiers partially substituting PRDM9 function, when the gene is deleted. Beyond H3K4me3 and H3K36me3, H3 lysine 9 acetylation (H3K9ac) is also enriched concurrently at recombination autosomal hotspots. H3K9ac also promotes chromatin openness, enabling DSB repair by homologous recombination [82]. On this regard The H3K4me3 and H3K36me3 reader ZCWPW1 (Zinc Finger CW-Type and PWWP Domain Containing 1) is recruited to recombination hotspots by PRDM9 and is essential for the execution of early repair steps at DSBs hotspots, by antagonizing histone deacetylase proteins [82,83,84,85]. In mouse, the PAR region contains a large H3K4me3 hotspot [75], which, however, is generated independently of PRDM9 [75]. To date, the methyltransferase responsible for H3K4me3 deposition in the mPAR has not been identified; hence, the function of H3K4me3 in the PAR remains to be experimentally validated. This contrasts with what has been found in humans, where PRDM9 does localize peaks of recombination in the PAR1 [86], and it is thus likely relevant for CO formation, in this region.
Trans-acting factors of DSB formation in the PAR: the essential role of ANKRD31
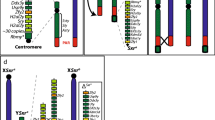
Functional activation of SPO11 at epigenetically marked hotspots requires the expression of several genes encoding auxiliary proteins of SPO11, namely: Iho1, Mei1, Mei4 Rec114, Ankrd31 (RMMAI proteins [62, 87,88,89,90]), which gene products form aggregates on the chromosome axes (the site where DSBs are made by SPO11) in advance of DSB formation [62, 87, 89, 91, 92]. Interestingly, mouse RMMAI aggregates form onto PAR much larger clusters (RMMAI blobs) than onto autosomes. RMMAI blobs also form onto autosomes at the (non-centromeric) telomeric regions of chromosomes 13, 9 and 4, which (like at the PAR) undergo DSB formation with at delayed timing compared to most autosomal hotspots [62]. Sequence analysis has revealed that the PAR and telomeres of these autosomes share the enrichment of tandem arrays of 31-bp repeats reach region known as mo-2 minisatellite [62]. In the mouse strains where mo-2 copy numbers in telomeres are lower, the aggregation of RMMAI factors (such as REC114) is lower [62]. Therefore, it has been concluded that the mo-2 minisatellite acts as a cis-acting determinant for RMMAI hyper aggregation [62]. Although the formation of DSBs in both autosomal and PAR hotspots require RMMAI aggregates, not all components of the complex are equally important for these regions. While expression of MEI1, MEI4 and REC114 is crucial for DSB formation across the entire genome [89], that of IHO1 (which is a direct binding partner of the axis associated protein HORMAD1 [87]), is obligate for autosomal DSB formation, while it is dispensable at the PAR [62, 93]. Conversely, the function of ANKRD31is critical in the PAR and PAR-like autosomal telomeres, and not essential at autosomal hotspots [90, 94]. In Ankrd31−/− spermatocytes near all of cells fails XY synapsis, and cells with achiasmata sex chromosomes arrest at MI, due to the activation of the SAC, leading to sterility [90, 94]. ANKRD31 is a direct REC114 interacting partner [90, 94]; therefore, it is speculated that ANKRD31 by recognising the cis-acting features of the PAR, recruits REC114 and other proteins by direct interaction, promoting DSB formation [94]. In this regard, it has recently been shown that REC114 also interacts directly with MEI1 ([93], Preprint) and TOPOVIBL [66] (Fig. 3). However, the interaction of TOPOVIBL and ANKRD31 with REC114 is mutually exclusive [66]. Therefore, given the essential function of ANKRD31 at the PAR [90, 94], it is not yet clear what is the exact interplay between these factors with TOPOVIB and SPO11. The interaction of ANKRD31 with REC114 is mediated by a conserved C-terminal region of ANKRD31, which wraps around the N-terminal PH domain (Pleckstrin Homology) of REC114, as shown by crystal structure studies [94]. Emphasizing the importance of such interaction, a recent study reported that in mice in which the REC114-ANKRD31 interaction is disrupted due to C-terminal truncation of ANKRD31, DSB formation at the PAR and XY synapsis is abolished, mimicking an Ankrd31 null phenotype [95]. The authors also demonstrated that when a missense mutation (E to A mutation at aa 1831) is introduced in the C-terminal of mouse ANKRD31, beside the ANKRD31-REC114 interaction was severely biodisrupted in Yeast 2-Hybrid assay, meiotic defects in homozygous mutants were much milder than in mice carrying the C-terminal truncation. This indicates that in vivo, the interaction is at least partially retained; perhaps strengthened by the network of interactions of ANKRD31 with other partners [95]. It is possible that ANKRD31 functions as a scaffold, interacting with multiple proteins at different times. In agreement with this concept, it has also demonstrated that ANKRD31 can also interact directly with MEI1 through one of its Ankyrin repeats [95], and with two more protein factors, namely: ZMYM3 (zinc finger, myeloproliferative, and mental retardation-type 3) and PTIP (Pax transactivation domain interacting protein; also known as PAXIP1) [62]. ZMYM3 is a chromatin interacting protein which promotes HR repair mediated by BRCA1 in somatic cells [96], and its deletion in mouse causes arrest of spermatocytes at MI [97]; which is compatible with a defect in CO formation between autosomes and/or sex chromosomes [25, 27]. PTIP, is an essential component of the activating H3K4me3 complex [98], and it is implicated in DNA damage repair [99]. Deletion of Ptip in testis causes arrest of spermatocytes at MI [100]. PTIP is not needed for global maintenance of H3K4me3 status [100], but its function might be required at specific subregions such as the mouse PAR, where the large H3K4me3 hotspot is PRDM9 independent [75]. Importantly, both ZMYM3 and PTIP are enriched at the PAR [62], which is a strong indication of their co-involvement in the formation of DSBs, and/or processing, in this region.

(adapted from [103])
Putative axis loop tethering model of the roles of FUS and EWS1 at the human PAR1 hotspot. FUS-PRDM9 interaction (might be indirect) links H3K4me3 K36me3-marked chromatin loops at the PAR1 with the SPO11 complex. The FUS-REC114 cooperation may enhance the proximity of the PAR1 hotspot with the axis and SPO11 auxiliary proteins. TOPOVIBL interacts directly with REC114 [66], possibly targeting the function of SPO11/TOPOVIBL onto axis. EWS1 may reinforce the role of FUS and TOPOVIBL via its interaction with PRDM9 and/or SPO11
Additional trans-acting factors with potential function in the PAR: EWS1 and FUS
The protein EWS1 (Ewing’s sarcoma breakpoint region 1) along with the gene products encoded by FUS (fused in sarcoma)/TLS (translocated in liposarcoma) and the TATA box binding associated factor 15 (Taf15), are RNA and DNA binding proteins that belong to the FET (FUS, EWS, TAF15) family of proteins [84]. Previous studies have demonstrated that Ewsr1−/− spermatocytes are deficient in synapsis of the autosomes and XY chromosomes, indicating their essential function during meiosis [101, 102]. By using the Spo11βki-only and Spo11αki-only models [30], we demonstrated that EWSR1 co-immunoprecipitates with SPO11β, SPO11α and REC114 [103]. Given the significance of SPO11α in DSB formation at the PAR [30], this observation supports a role for EWS1 in XY recombination. Intriguingly, homolog synapsis is also defective in spermatocytes lacking FUS/TLS expression [104]. FUS also co-immunoprecipitates with SPO11 splice isoforms and with REC114, and it localizes at the PAR hotspot [103]. Therefore, FUS/TLS is also a likely player in XY recombination.
The formation of DSBs occurs in the context of the spatial organisation of meiotic chromosomes, which form chromatin loops that extend from a linear protein axis. Based on the yeast model, which predicts that the DSB machinery assembled on the axis captures and breaks DNA loops, it has been proposed that in mammals, DSB occurs in PRDM9-marked regions on DNA loops, which are next attached to the axis, where DSBs are made and repaired by assembly of DNA repair factors [105, 106]. Both EWSR1 and FUS/TLS co-immunoprecipitates with PRDM9 in spermatocytes [103, 107] Therefore, it is speculated that these protein factors binding PRDM9 on chromatin loops are tethered to the axis (bound REC114) and with SPO11, promoting DSB formation [103]. In mice DSBs form in the PAR independently from PRDM9 [75]. However, in man, PRDM9 does localize at the PAR1 [86], therefore FET family proteins, with same mechanism hypothesized for the autosomes in mice, may facilitate DSB formation on the PAR1 chromosome axis (Fig. 3).
In addition to FET family proteins, in mice, PRDM9 also interacts directly with CXXC1 (CxxC finger protein 1), a H3K4me3 reader ortholog of S. cerevisiae Spp1 [108] in mammals [109], CDYL (chromodomain-containing Y chromosome-like) and EHMT2 (euchromatic histone methyltransferase 2). These protein factors are expressed in spermatocytes with the same timing as PRDM9 and are though to facilitate the association of putative hotspot sites in DNA loops with the chromosomal axis ([107], reviewed in [110]). To date, CDYL and EHMT2 function still await experimental validation, in vivo, while the function of CXXC1 at hotspots remains uncertain, as phenotypic characterization of Cxxc1 null mice with a C57BL/6 genetic background, provided contradictory results, likely due to differences in the experimental settings [109, 111].
Cis-acting factors of DSB formation in the PAR
In addition to the presence of mo2-minisatellites (see above and [62]), recent studies have shown that mPAR chromatin in spermatocytes forms relatively short loops on a long axis, compared to autosomes [27]. According to the tethering model [105, 106], this conformation is acquired to increase loops density, favoring DSB formation in the PAR [27]. Using high-resolution structured illumination microscopy, Acquaviva et al. demonstrated that in mice, PAR and telomeres of chromosomes with mo-2 minisatellite repetitive sequences undergo dynamic remodeling [62]. Specifically, the PAR axis elongates, and chromatin loops shorten, as cells progress from leptonema to zygonema. Moreover, as cells approach the late zygotene stage, the sister chromatids of X-PAR and Y-PAR undergo separation (splitting) (Fig. 4A).
Ultrastructure of the PAR before and after synapsis. A Spermatocyte chromosome spread stained with SYCP3 and IHO1, which allows for rapid identification of sex chromosomes [71]. Splitting of the X-PAR and Y-PAR in unsynapsed chromosomes is visible through Stimulated Emission Depletion (STED) super-resolution microscopy imaging (enlargement). B Splitting of PARs is lost following XY synapsis
These ultrastructural changes are closely correlated with the accumulation of RMMAI blobs and the formation of DSBs [62]. Similarly, telomeres with mo2-minisatellite sequences also undergo splitting paralleling changes in the PAR [62]. Once synapsis is formed, PAR sister chromatids collapse (Fig. 4B), with successive dissociation of RMMAI proteins, shortening of the axes, and elongation of chromatin loops [62]. Importantly, splitting at the PAR and telomeres with mo2-minisatellite is absent in Mei4−/− and Ankrd31−/− mutants that fail aggregation of RMMAI proteins [62]. This confirms that sister chromatids separation is somehow related to the formation of RMMAI protein clusters. Nevertheless, the function of PAR (and mo2 minisatellites enriched telomeres) splitting is not yet clear. It has been suggested that axis separation might suppress ineffective intersister recombination in favour of homologous recombination between chromosomes [62, 112]. An alternative (not mutually exclusive) interpretation is that the axes split to accommodate the RMMAI aggregates [112]. In this regard, we demonstrated that in Spo11βki-only mice with defective XY synapsis, splitting of the Y-PAR occurs with a slightly reduced frequency compared to wild type control, although apparent normal assembly of RMMAI proteins. Decreased frequency of Y-PAR splitting parallels that of DSB in the Y-PAR [30]. Therefore, we favor the hypothesis that splitting requires the suppression of inter-sister recombination. In addition, in accordance with the tethering model of formation of DSBs, we also observed that reduced formation of DSB in the Y-PAR at late zygonema, correlates with elongation of PAR loops compared to that of wild type. This underlines the importance of the ultrastructural conformation of PAR in the formation of DSBs [30]. It is interesting that the length of PAR loops varies with mouse genetic background [30]. It is therefore plausible that genetic polymorphisms in genes responsible for the ultrastructural organization of chromatin impacts the efficiency of DSB formation on PAR. Further studies will be necessary to explore these aspects in more detail.
Genetic factors of CO maturation
A DSB formation is the prerequisite but not the guarantee for a CO, which may form only after appropriate processing of DSBs. SPO11-mediated cleavage results in single-strand DNA overhangs that are subsequently coated by various recombination proteins that assembles onto chromosome axes as cytologically visible foci, including the strand exchange factors DMC1 and RAD51 [113,114,115]. Processing of DSBs allows for homology search, which in turn promotes homology pairing, synapsis, and DSB repair [116]. Repair of DSBs leads to the formation of COs or NCOs [1, 17]. At chromosome scale, the probability of receiving DSBs and resolving them as a CO is negatively correlated with the chromosome size [61]; moreover, the spacing between COs is regulated by a process called COs interference, which causes them be less spatially close than would be expected in a random distribution [1, 56, 117]. COs, subjected to interference are referred to as “Type I”, representing the major fraction (ranging between 90% and 95% in the mouse) of all COs [118]. A minor fraction of COs, not subjected to interference, also forms [1]. These are referred to as “Type II”, which involves structural specific endonucleases such as MUS81 [119]. On average, of all DSBs introduced into the genome only ~ 10–25% is converted into a Type I CO in mammals [16]. Designation of DSBs toward a Type I CO fate requires stabilization of specific DNA intermediates during HR-mediated repair, by pro-CO factors (i.e. directing DSB processing towards a CO fate), collectively known in budding yeast S. cerevisiae as ZMM proteins (an acronym of Zip1-4, Msh4-5, Mer3, Spo16). The orthologs and homologous of ZMM proteins in mammalian are SYCP1 (budding yeast Zip1), SHOC1/MZIP2 (ortholog of Zip2), RNF212 (ortholog of Zip3), TEX11 (ortholog of Zip4), MSH4, MSH5, HFM1 (ortholog of Mer3) [118], SPO16 [120]. One additional factor is the Human Enhancer of Invasion-10 (HEI10) (also known as CCNB1 interacting protein 1 [CCNB1IP1] in human), a RNF212 paralog with domain similarity, identified by mouse forward genetic screen [121]. Most of recombination intermediates (namely D-loop structures) stabilized by ZMMs are processed as COs in budding yeast. However, in mammals, ZMM foci outnumber the COs, indicating that DSB intermediates bound by some ZMM proteins at pre-CO sites (i.e. early selected but not yet designated COs) can still be unselected and resolved as NCOs [118], while few DSBs will become COs (designated-CO sites). In mammals, maturation of recombination intermediates toward a COs fate is established progressively, through successive assembly and dissociation of ZMM sub-complexes which partially colocalize with recombination foci defined by RAD51 and DMC1 (single-stranded DNA binding-proteins marking DSBs) [118]. In yeast the ZMM proteins Zip2-Zip4-Spo16 form a stable subcomplex (ZZS) with pro-CO activity [118]. This function and mutual proteins interaction is conserved in mammals, as shown by their mutual co-immunoprecipitation and reduced formation of COs and chiasmata in Shoc1−/− [122], Tex11−/− [123] and Spo16−/− [120] knockout mice, and association of ZZS [124,125,126,127] and other ZMM genes deleterious variants [128] with infertility in humans. SHOC1 co-localizes strongly with DMC1 at leptonema and zygonema, indicating a function in stabilizing the early recombination intermediates [122] (Fig. 5A), while co-localization with TEX11 is low at leptonema to early zygonema and increases by late zygonema/early pachynema [122] (Fig. 5B). MSH4-5 MutSγ proteins localize as foci at multiple sites in early synapsed regions (Fig. 5B) at most but not all DSBs, with foci number peaking at zygonema and decreasing at early and late pachynema [51, 122, 129]. This indicates a function in the commitment of a reduced pool of DSBs (i.e. pre-CO sites) toward a CO fate, as demonstrated in yeast [130]. Accumulation of a normal number of MSH4 foci requires HFM1 (helicases for meiosis 1). In the absence of HFM1, normal turnover of earlier recombination intermediates (i.e. RAD51) is impeded, indicating insufficient processing of DSBs [131], causing infertility [132]. Mouse RNF212 and HEI10 that function as small ubiquitin-like modifier (SUMO) and ubiquitin E3 ligases respectively, establish an early differentiation between CO/NCO sites. RNF212 is inferred to promote selective stabilization at a defined intermediate step, of a minority of TEX11 and MSH4-MSH5 bound recombination intermediates [16] beyond early pachynema (Fig. 5C, D), as indicated by their limited co-localization onto axis [16, 133], designating CO sites. The function of RNF212 is conserved in human, as indicated by genome-wide recombination rate changes and meiotic arrest associated with RNF212 sequence variants [134, 135]. The Cyclin-like cyclin N-Terminal Domain Containing 1 (CNTD1) and its binding partner Proline Rich 19 (PRR19) are the keys to narrow down pre-CO sites marked by MutSγ and RNF212, allowing successive loading of HEI10 [136, 137]. HEI10 is required for post-synapsis turnover of RNF212 and MutSγ co-complexes that culminates in its selective retention at designated CO sites [138] (Fig. 5D, E) and loading of MutLγ factors (MLH3, MLH1) and Cyclin-dependent Kinase-2 (CDK2), with consequent formation of COs [1, 16, 133, 138,139,140] (Fig. 5E).
Selection and maturation of COs. Block arrows represent the progression of meiotic recombination. Although four chromatids are present at this stage, only two are shown in the scheme for the sake of simplicity. Recombination proteins assembles at DSB sites as cytologically detectable foci. SPO16 foci are located on chromosome axes, most at synapsed regions; the number of foci increase progressively from leptonema to zygonema and slightly decreases at pachynema. The SPO16 binding partners SHOC1 and TEX11 form foci, which number peaks at zygonema and decreases as the cell progresses to pachynema. MSH4/MSH5 MutSγ proteins also localize at early synapsed regions in zygonema after the turnover of ssDNA binding proteins by HFM1. SPO16, SHOC1 and TEX11 co-localizes only with selected MutSγ positive sites at zygonema, while are lost by late pachynema, when few MSH4/MSH5 persist at designated CO sites. The RNF212 foci number increase as synapsis occurs, up to early pachynema, and subsequently decrease in number to disappear at late pachynema. In mid-pachynema, only one or two foci of RNF212 remain per synaptonemal complex and colocalize with selected MSH4-positive TEX11-positive foci and MutLγ proteins (MLH1 and MLH3) at mature CO sites. CNTD1 and PRR19 narrows down MutSγ-positive and RNF212-positive pre-CO sites with successive loading of HEI10. HEI10 is required for post-synapsis turnover of RNF212. Unlike RNF212, foci of HEI10 are rarely detected along nascent synaptonemal complexes during zygonema, while foci appear by early pachynema, pick at mid-pachynema, and decreases at late-pachynema, when RNF212 foci are already lost, colocalizing with MutLγ proteins at mature CO sites. CO crossover, NCO non-crossover
CO formation in the PAR
In mice and humans DSBs not only form more frequently in the PAR compared with autosomes but are also more likely to be processed toward a CO fate than autosomes [18, 27, 86]. It is estimated that in mouse there is ~ 570-fold higher CO density in the PAR compared to genome average, which translates to a ~ fivefold higher yield of COs per DSB compared to autosomes [61]. Therefore, specific mechanisms are expected to be in place to stabilize CO intermediates toward a CO fate. An important contribution to the formation of the obligatory CO in the PAR comes from ATM (Ataxia Telangiectasia Mutated). Atm null spermatocytes are defective in forming the obligate CO on the sex chromosomes [141]. This is in striking contrast to autosomes where the total number of DSBs onto autosomes (and non-homologous portion of sex chromosomes) and COs increases compared to the control [61, 141, 142]. Thus, ATM upgrades DSBs to a CO fate in the PAR, while constraining DSB formation and CO numbers in other chromosome regions. To date, the molecular mechanisms behind such regulatory control are largely unknown. ZMM proteins are certainly necessary for the maturation of DSB in the PAR, as demonstrated by the immunolocalization of MSH4 [140]; in addition, cytological observations show that XY synapsis fails with high frequency in the absence of Hfm1, indicating that, like in autosomes, HFM1 is required with MSH4 to initiate or maintain stable early recombination intermediates [131]. RNF212 also localizes at the PAR at early pachynema [133, 138] and is a dosage–sensitive regulator of XY synapsis in mice, likely stabilizing the nascent CO intermediates between PARs [16]. On the contrary, although HEI10 is also a dosage-sensitive regulator of CO formation in mice, no specific defect in HEI10 foci formation, CO maturation or maintenance of stable synapsis between XY chromosomes was reported in heterozygous mice mutants [138]. This could indicate that the stability and maturation of CO intermediates in the PAR is likely not dependent on the HEI10 function.
In addition to the above, over the past 4 years, new genes with an impact on CO maturation have been identified, mainly by sequencing of the human genomes of patients categorized according to their fertility. Some gene functions have been shown to impact mainly recombination between male sex chromosomes, in some cases with no or little impact on autosomal recombination. These genes are described below.
-
a.
Genes upgrading DSB formation in the PAR
hnRNPH1: Coordinated regulation of alternative pre-mRNA splicing is essential for germ cell development. An RNA binding protein that has recently been found to play a key role in meiosis is the heterogeneous nuclear riboprotein (hnRNP) hnRNPH1, which absence in germ cells causes male and female sterility, due to altered gene expression and alternative splicing [65]. The lack of hnRNPH1 expression in spermatocytes compromises alternative splicing of genes related to meiosis including SPO11, by maintaining SPO11β expression at high level in 18 days post-partum testes, to detriment of the alternative transcript encoding for the SPO11α isoform [65]. This result agrees with a previous report that identified hnRNPH1 as a key regulator of Spo11α splicing in mouse spermatocytes [64]. Given that the PAR physiologically receives DSBs with a higher frequency than typical autosome segments [27, 61], and concomitant expression of SPO11β and SPO11α is the key for efficient formation of DSBs in the PAR [30], this is expected to have an impact on the initiation of XY recombination. Accordingly, the authors show that hnRNPH1-deficient mice display a tenfold increase of XY asynapsis compared to controls [65]. Furthermore, considering the ability of hnRNPH1 to interact with the splicing factors PTBP2 and SRSF3 in the testes [65], the authors also show that the Spo11 gene is regulated at the splicing level by PTBP2 and SRSF3c, which are recruited by hpRNPH1 [65].
-
b.
Genes upgrading the PAR CO program
USP26: In humans, meiotic XY missegregation can lead to KS offspring. However, to what extent genetic predisposes to paternal sex chromosome aneuploidy has remained long elusive. Liu et al. have demonstrated that deleterious mutations in the USP26 (ubiquitin-specific protease 26) gene increase the risk of fathering a KS progeny [143]. The study identified USP26 using a whole-exome sequencing (WES) in a cohort of KS patients, as well as KS family trios. By deleting Usp26 in mice, they demonstrated that USP26 de-ubiquitinate the ZZS protein TEX11, increasing its expression. Tex11 mutations have been often associated with male fertility defects [126, 144, 145], and its deletion in mice delayed resolution of DSB intermediates and decreased of CO numbers, with most spermatocytes arresting at pachynema [123, 143]. However, USP26 has several other substrates, including the androgen receptor. Therefore, Usp26 mutations are expected to impact spermatogenesis via multiple mechanisms [143, 146]. Importantly, some residual spermatogenesis still exists in Usp26-deficient mice; spermatozoa are aneuploid for the XY chromosomes [143], and sometime mice sired an XXY offspring [143], demonstrating USP26 potential function in the etiopathogenesis of KS.
RAD51AP2: RAD51-associated protein 2 is a meiosis-specific gene whose frameshift truncating mutations have been identified by WES, in patients with non-obstructive azoospermia. The modeling and phenotypic characterization of mutations in mice have shown that the lack of Rad51ap2 expression has no impact on processing of DSBs and synapsis of non-sex chromosomes, and although XY chromosomes synapse normally at the PAR in early and mid-pachynema, they separate precociously at late pachynema, before the completion of recombination [147]. Specifically, in the absence of RAD51AP2, the intermediate recombination markers MSH4 and TEX11 vanish precociously from the PAR (but not from autosomes). This demonstrates that RAD51AP2 stabilizes PAR recombination intermediates, playing a key role in maturing the obligatory CO in this chromosomal region. Importantly, the authors also show that RAD51AP2 co-immunoprecipitates with RAD51 (not with DMC1) and interact through the C-terminus of RAD51AP2. Consequently, any mutations occurring in the interaction domains of RAD51AP2 and/or RAD51 within the human genome could potentially have a negative impact on XY recombination. Further WES studies in individuals with KS or a different cohort of non-obstructive azoospermia patients are expected to uncover new variants of these genes, deleterious to recombination in the PAR.
M1AP: Meiosis 1 Arresting Protein is a vertebrate protein expressed only in female and male germ cells. Its mutation has been found to be associated with infertility both in men and in male mice [148,149,150,151], however, its molecular structure has remained unidentified. Recently Li et al. [151] modelled the M1ap c.1074+2T>C splicing mutation, equivalent to that found in patients with severe oligozoospermia in mice. The mutation causes an inactivating premature protein truncation, causing reduction of both CO formation and chromosome synapses in spermatocytes, particularly between XY chromosomes. Mechanistically, it has been demonstrated that M1AP form discrete foci on the chromosome axes of spermatocytes and that it interacts with the components of the ZZS complex SHOC1, TEX11 and SPO16, colocalizing with the foci TEX11 in a SPO16-dependent manner. Ablation of M1ap in mice reduces the recruitment of TEX11 without altering SHCO1 localization, thus altering the stability of early recombination intermediates that mostly affect recombination at the PAR [143, 151].
RNF212B: The ring Finger Protein 212B is a gene whose protein product is characterized by a ring finger domain commonly associated with E3 ubiquitin ligase activity. Serving as the closest paralog to RNF212, RNF212B shares functional significance in controlling recombination rate in mammals [134, 152, 153]. In a recent whole-exome sequency study in patients with severe male infertility Gershoni et al. [154] brought to light a pathogenic variant of the RNF212B gene causing a substitution at position 448 (C448T), that results in the conversion of the arginine-150 codon to a premature stop codon. This alteration predicts a truncation of the C-terminal half of the protein. The patients carrying the homozygous RNF212BC448T variant suffer of a severe chromosome nondisjunction defect in sperm cells, especially of the sex chromosomes [154]. This observation suggests that RNF212B has a functional role in the processing of DSBs formed in the PAR, although not strictly specific.
ATF7IP2: The Activating transcription factor 7 interacting protein 2, also called PMS2/MCAF2, is encoded by a gene preferentially expressed in the gonads, especially during the meiosis stage. The null mutation in Atf7ip2 causes male sterility, predominantly due to defective sex chromosome synapsis failure [155]. Shao et al. have reported that in the absence of ATF7IP2, the length of the chromosome axis increases in autosomes and in the PAR. This correlates with ~ 10% increase of CO frequency in autosomes, while PAR loses the obligatory CO [155]. They ruled out a defect in the occurrence of DSBs at the PAR, while the localization of the ZMM proteins stabilizer RNF212 [16] was impaired, as well that of MSH4. Therefore, the XY CO defect is probably caused by the instability of the recombination intermediates processed by MSH4 and RNF212. Therefore, ATF7IP2 seems to be a protein factor that boosts DSB to CO maturation in the PAR. More recently, a new Atf7ip2 null model was generated with the same genetic background, which shows no XY synapse defects [156]. The two models differ for the mutated exons (exons 3–6 deletion [155] and 17 bp exon 4 frameshift deletion [156]), which presumably causes the phenotypic difference. ATF7IP2 interacts with the H3K9 histone-lysine N-methyltransferase SETB1, to regulate SETB1 retention in the nucleus and H3K9 trimethylation of the chromatin of sex chromosomes [155, 156]. This function is required to regulate MSCI [155, 156], needed for successful spermatogenesis (see [48] and references therein).
Checkpoint mechanisms of selection of XY aneuploid germ cells
XY aneuploidy (but not autosomal aneuploidy) in human sperm increases with age, often in the face of even modest meiotic perturbations, with the risk of fathering a child with KS [157,158,159]. Based on recent experimental findings, the association between paternal age and XY aneuploidy lies, at least in part, in the weakening of Spindle Assembly Checkpoint (SAC) mechanisms that eliminate metaphase spermatocytes with misaligned chromosomes [160]. A limiting step in the study of SAC proteins during the meiotic progression of multicellular organisms is the identification of viable alleles since these proteins are essential for proper embryonic development. Therefore, only few studies have addressed whether the SAC mechanism is functional during meiotic progression in vivo, by genetically reducing the dosage of SAC protein [161]. Faisal et al., demonstrated that reduced MAD2 level dampens the apoptotic response in a mouse model with a high frequency of nonexchange XY chromosomes, with consequent generation of sperm aneuploid for the sex chromosomes [29]. Other genes whose heterozygosity led to aneuploid splenocytes (Bub3, Rae1, Bub3/Rae1 double heterozygotes and Rae1/Nup98 double heterozygotes) did not show such effect in spermatocytes [162], underlining the specificity of Mad2 function in these cells. In Usp26−/− mice XY diploid sperm (and more rarely sex chromosomes-nulliploid sperms) were produced only by aged 6-month-old Usp26−/− mice, when the level of SAC proteins (namely: MAD2, BUBR1, PLK1) was reduced; and aged mice sometime sired XXY offspring [143]. Therefore, it is likely that a combination between inefficient XY pairing and less stringent SAC surveillance allows XY sperm aneuploidy [163].
Concluding remarks
Male sex chromosome aneuploidy in sperm is a risk factor for infertility, subfertility and generation of a progeny with KS and TS. However, whether and to what extent genetic factors may predispose to XY aneuploidy has long remained elusive. In recent years important progress in understanding the genetics behind XY recombination failure, with the identification of genes whose products have a specific or predominant function in recombination in the PAR. The experimental approaches have varied, from basic research in the field of meiosis to those based on DNA sequencing from infertile patients or patients with KS and their parents. The genes for mutations impacting XY recombination range from those involved in the formation of DSBs and PAR remodeling to those required for the processing of DSBs and the apoptotic selection of cells with an unbalanced number of XY chromosomes. Where gene mutations have been found to be associated with infertility or KS, these mutations were present in a limited pool of affected individuals and parents. This indicates that the catalogue of genes with mutations related to KS and infertility is probably much larger than what we know now. Mutations in individual genes are expected to be genetically unselected and likely to cause XY aneuploidy when in homozygosity and/or expressed in combination with other gene mutations weakening either recombination, chromosome segregation, and/or checkpoint mechanisms efficiency. Therefore, the path to identify genetic risk factors for XY aneuploidy remains a challenge for the future, to be continued.
Data availability
All data are available through the cited bibliographic references.
References
Hunter N (2015) Meiotic recombination: the essence of heredity. Cold Spring Harb Perspect Biol 7(12):a016618
Keeney S et al (1999) A mouse homolog of the saccharomyces cerevisiae meiotic recombination DNA transesterase Spo11p. Genomics 61(2):170–182
Keeney S, Giroux CN, Kleckner N (1997) Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88(3):375–384
Baudat F et al (2000) Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell 6(5):989–998
Romanienko PJ, Camerini-Otero RD (1999) Cloning, characterization, and localization of mouse and human SPO11. Genomics 61(2):156–169
Romanienko PJ, Camerini-Otero RD (2000) The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell 6(5):975–987
Robert T et al (2016) The TopoVIB-like protein family is required for meiotic DNA double-strand break formation. Science 351(6276):943–949
Zhang J et al (2011) An association study of SPO11 gene single nucleotide polymorphisms with idiopathic male infertility in Chinese Han population. J Assist Reprod Genet 28(8):731–736
Karimian M et al (2015) SPO11-C631T gene polymorphism: association with male infertility and an in silico-analysis. J Family Reprod Health 9(4):155–163
Ren ZJ et al (2017) The SPO11-C631T gene polymorphism and male infertility risk: a meta-analysis. Ren Fail 39(1):299–305
Tran TN, Schimenti JC (2019) A segregating human allele of SPO11 modeled in mice disrupts timing and amounts of meiotic recombination, causing oligospermia and a decreased ovarian reserve. Biol Reprod 101(2):347–359
Perry J et al (2001) A short pseudoautosomal region in laboratory mice. Genome Res 11(11):1826–1832
Flaquer A, Fischer C, Wienker TF (2009) A new sex-specific genetic map of the human pseudoautosomal regions (PAR1 and PAR2). Hum Hered 68(3):192–200
Baudat F, de Massy B (2007) Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res 15(5):565–577
Cole F, Keeney S, Jasin M (2010) Comprehensive, fine-scale dissection of homologous recombination outcomes at a hot spot in mouse meiosis. Mol Cell 39(5):700–710
Reynolds A et al (2013) RNF212 is a dosage-sensitive regulator of crossing-over during mammalian meiosis. Nat Genet 45(3):269–278
Cole F et al (2014) Mouse tetrad analysis provides insights into recombination mechanisms and hotspot evolutionary dynamics. Nat Genet 46(10):1072–1080
Jeffreys AJ, May CA (2004) Intense and highly localized gene conversion activity in human meiotic crossover hot spots. Nat Genet 36(2):151–156
Hinch R, Donnelly P, Hinch AG (2023) Meiotic DNA breaks drive multifaceted mutagenesis in the human germ line. Science 382(6674):eadh2531
Veller C, Kleckner N, Nowak MA (2019) A rigorous measure of genome-wide genetic shuffling that takes into account crossover positions and mendel’s second law. Proc Natl Acad Sci USA 116(5):1659–1668
La Volpe A, Barchi M (2012) Meiotic double strand breaks repair in sexually reproducing eukaryotes: we are not all equal. Exp Cell Res 318(12):1333–1339
Guillon H et al (2005) Crossover and noncrossover pathways in mouse meiosis. Mol Cell 20(4):563–573
Yun Y et al (2021) Cytological monitoring of meiotic crossovers in spermatocytes and oocytes. Methods Mol Biol 2153:267–286
Hirose Y et al (2011) Chiasmata promote monopolar attachment of sister chromatids and their co-segregation toward the proper pole during meiosis I. PLoS Genet 7(3):e1001329
Edelmann W et al (1996) Meiotic pachytene arrest in MLH1-deficient mice. Cell 85(7):1125–1134
Lipkin SM et al (2002) Meiotic arrest and aneuploidy in MLH3-deficient mice. Nat Genet 31(4):385–390
Kauppi L et al (2011) Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331(6019):916–920
Faisal I, Kauppi L (2016) Sex chromosome recombination failure, apoptosis, and fertility in male mice. Chromosoma 125(2):227–235
Faisal I, Kauppi L (2017) Reduced MAD2 levels dampen the apoptotic response to non-exchange sex chromosomes and lead to sperm aneuploidy. Development 144(11):1988–1996
Giannattasio T et al (2023) The proper interplay between the expression of Spo11 splice isoforms and the structure of the pseudoautosomal region promotes XY chromosomes recombination. Cell Mol Life Sci 80(10):279
Rubio C et al (2001) Incidence of sperm chromosomal abnormalities in a risk population: relationship with sperm quality and ICSI outcome. Hum Reprod 16(10):2084–2092
Tempest HG et al (2004) The association between male infertility and sperm disomy: evidence for variation in disomy levels among individuals and a correlation between particular semen parameters and disomy of specific chromosome pairs. Reprod Biol Endocrinol 2:82
Aran B et al (1999) Screening for abnormalities of chromosomes X, Y, and 18 and for diploidy in spermatozoa from infertile men participating in an in vitro fertilization-intracytoplasmic sperm injection program. Fertil Steril 72(4):696–701
Codina-Pascual M et al (2005) Synapsis and meiotic recombination analyses: MLH1 focus in the XY pair as an indicator. Hum Reprod 20(8):2133–2139
Hassold TJ et al (1991) XY chromosome nondisjunction in man is associated with diminished recombination in the pseudoautosomal region. Am J Hum Genet 49(2):253–260
Jacobs PA et al (1988) Klinefelter’s syndrome: an analysis of the origin of the additional sex chromosome using molecular probes. Ann Hum Genet 52(2):93–109
Thomas NS, Hassold TJ (2003) Aberrant recombination and the origin of klinefelter syndrome. Hum Reprod Update 9(4):309–317
Bojesen A, Juul S, Gravholt CH (2003) Prenatal and postnatal prevalence of klinefelter syndrome: a national registry study. J Clin Endocrinol Metab 88(2):622–626
Boateng KA et al (2013) Homologous pairing preceding SPO11-mediated double-strand breaks in mice. Dev Cell 24(2):196–205
Ding X et al (2007) SUN1 is required for telomere attachment to nuclear envelope and gametogenesis in mice. Dev Cell 12(6):863–872
Sole M et al (2022) Time to match; when do homologous chromosomes become closer? Chromosoma 131(4):193–205
Westergaard M, von Wettstein D (1972) The synaptinemal complex. Annu Rev Genet 6:71–110
Yuan L et al (2000) The murine SCP3 gene is required for synaptonemal complex assembly, chromosome synapsis, and male fertility. Mol Cell 5(1):73–83
de Vries FA et al (2005) Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev 19(11):1376–1389
Barchi M, Jasin M (2003) Seeking new meiotic genes. Proc Natl Acad Sci USA 100(26):15287–15289
Dunne OM, Davies OR (2019) A molecular model for self-assembly of the synaptonemal complex protein SYCE3. J Biol Chem 294(23):9260–9275
Billmyre KK et al (2023) SYCP1 head-to-head assembly is required for chromosome synapsis in mouse meiosis. Sci Adv 9(42):eadi1562
Alavattam KG et al (2021) Meiotic sex chromosome inactivation and the XY body: a phase separation hypothesis. Cell Mol Life Sci 79(1):18
Ichijima Y et al (2011) MDC1 directs chromosome-wide silencing of the sex chromosomes in male germ cells. Genes Dev 25(9):959–971
Fernandez-Capetillo O et al (2003) H2AX is required for chromatin remodeling and inactivation of sex chromosomes in male mouse meiosis. Dev Cell 4(4):497–508
Testa E et al (2018) H2AFX and MDC1 promote maintenance of genomic integrity in male germ cells. J Cell Sci 131(6):jcs214411. https://doi.org/10.1242/jcs.214411
McKee BD, Handel MA (1993) Sex chromosomes, recombination, and chromatin conformation. Chromosoma 102(2):71–80
Mahadevaiah SK et al (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27(3):271–276
Kauppi L et al (2013) Numerical constraints and feedback control of double-strand breaks in mouse meiosis. Genes Dev 27(8):873–886
Faieta M et al (2016) A surge of late-occurring meiotic double-strand breaks rescues synapsis abnormalities in spermatocytes of mice with hypomorphic expression of SPO11. Chromosoma 125(2):189–203
Cole F et al (2012) Homeostatic control of recombination is implemented progressively in mouse meiosis. Nat Cell Biol 14(4):424–430
Burgess SM (2002) Homologous chromosome associations and nuclear order in meiotic and mitotically dividing cells of budding yeast. Adv Genet 46:49–90
Goldman AS, Lichten M (2000) Restriction of ectopic recombination by interhomolog interactions during saccharomyces cerevisiae meiosis. Proc Natl Acad Sci USA 97(17):9537–9542
Soriano P et al (1987) High rate of recombination and double crossovers in the mouse pseudoautosomal region during male meiosis. Proc Natl Acad Sci USA 84(20):7218–7220
Shi Q et al (2001) Single sperm typing demonstrates that reduced recombination is associated with the production of aneuploid 24, XY human sperm. Am J Med Genet 99(1):34–38
Lange J et al (2016) The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167(3):695-708.e16
Acquaviva L et al (2020) Ensuring meiotic DNA break formation in the mouse pseudoautosomal region. Nature 582(7812):426–431
Bellani MA et al (2010) The expression profile of the major mouse SPO11 isoforms indicates that SPO11beta introduces double strand breaks and suggests that SPO11alpha has an additional role in prophase in both spermatocytes and oocytes. Mol Cell Biol 30(18):4391–4403
Cesari E et al (2020) Combinatorial control of Spo11 alternative splicing by modulation of RNA polymerase II dynamics and splicing factor recruitment during meiosis. Cell Death Dis 11(4):240
Feng S et al (2022) hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells. Nat Commun 13(1):3588
Nore A et al (2022) TOPOVIBL-REC114 interaction regulates meiotic DNA double-strand breaks. Nat Commun 13(1):7048
Baudat F, Imai Y, de Massy B (2013) Meiotic recombination in mammals: localization and regulation. Nat Rev Genet 14(11):794–806
Baudat F et al (2010) PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327(5967):836–840
Parvanov ED, Petkov PM, Paigen K (2010) Prdm9 controls activation of mammalian recombination hotspots. Science 327(5967):835
Imai Y et al (2020) PRDM9 activity depends on HELLS and promotes local 5-hydroxymethylcytosine enrichment. Elife 9:e57117. https://doi.org/10.7554/eLife.57117
Spruce C et al (2020) HELLS and PRDM9 form a pioneer complex to open chromatin at meiotic recombination hot spots. Genes Dev 34(5–6):398–412
Eram MS et al (2014) Trimethylation of histone H3 lysine 36 by human methyltransferase PRDM9 protein. J Biol Chem 289(17):12177–12188
Powers NR et al (2016) The meiotic recombination activator PRDM9 trimethylates both H3K36 and H3K4 at recombination hotspots in vivo. PLoS Genet 12(6):e1006146
Wu H et al (2013) Molecular basis for the regulation of the H3K4 methyltransferase activity of PRDM9. Cell Rep 5(1):13–20
Brick K et al (2012) Genetic recombination is directed away from functional genomic elements in mice. Nature 485(7400):642–645
Diagouraga B et al (2018) PRDM9 methyltransferase activity is essential for meiotic DNA double-strand break formation at its binding sites. Mol Cell 69(5):853-865.e6
Grey C et al (2011) Mouse PRDM9 DNA-binding specificity determines sites of histone H3 lysine 4 trimethylation for initiation of meiotic recombination. PLoS Biol 9(10):e1001176
Mihola O et al (2009) A mouse speciation gene encodes a meiotic histone H3 methyltransferase. Science 323(5912):373–375
Sun F et al (2015) Nuclear localization of PRDM9 and its role in meiotic chromatin modifications and homologous synapsis. Chromosoma 124(3):397–415
Davies B et al (2016) Re-engineering the zinc fingers of PRDM9 reverses hybrid sterility in mice. Nature 530(7589):171–176
Mihola O et al (2019) Histone methyltransferase PRDM9 is not essential for meiosis in male mice. Genome Res 29(7):1078–1086
Yuan S et al (2022) The histone modification reader ZCWPW1 promotes double-strand break repair by regulating cross-talk of histone modifications and chromatin accessibility at meiotic hotspots. Genome Biol 23(1):187
Wells D et al (2020) ZCWPW1 is recruited to recombination hotspots by PRDM9 and is essential for meiotic double strand break repair. Elife 9:e53392. https://doi.org/10.7554/eLife.53392
Huang T et al (2020) The histone modification reader ZCWPW1 links histone methylation to PRDM9-induced double-strand break repair. Elife 9:e53459
Mahgoub M et al (2020) Dual histone methyl reader ZCWPW1 facilitates repair of meiotic double strand breaks in male mice. Elife 9:e53360
Hinch AG et al (2014) Recombination in the human pseudoautosomal region PAR1. PLoS Genet 10(7):e1004503
Stanzione M et al (2016) Meiotic DNA break formation requires the unsynapsed chromosome axis-binding protein IHO1 (CCDC36) in mice. Nat Cell Biol 18(11):1208–1220
Libby BJ et al (2002) The mouse meiotic mutation mei1 disrupts chromosome synapsis with sexually dimorphic consequences for meiotic progression. Dev Biol 242(2):174–187
Kumar R et al (2018) Mouse REC114 is essential for meiotic DNA double-strand break formation and forms a complex with MEI4. Life Sci Alliance 1(6):e201800259
Papanikos F et al (2019) Mouse ANKRD31 regulates spatiotemporal patterning of meiotic recombination initiation and ensures recombination between X and Y sex chromosomes. Mol Cell 74(5):1069-1085.e11
Reinholdt LG, Schimenti JC (2005) Mei1 is epistatic to Dmc1 during mouse meiosis. Chromosoma 114(2):127–134
Kumar R, Bourbon HM, de Massy B (2010) Functional conservation of Mei4 for meiotic DNA double-strand break formation from yeasts to mice. Genes Dev 24(12):1266–1280
Dereli I et al (2023) Seeding the meiotic DNA break machinery and initiating recombination on chromosome axes. bioRxiv. https://doi.org/10.1101/2023.11.27.568863
Boekhout M et al (2019) REC114 partner ANKRD31 controls number, timing, and location of meiotic DNA breaks. Mol Cell 74(5):1053-1068.e8
Xu J et al (2023) Essential roles of the ANKRD31-REC114 interaction in meiotic recombination and mouse spermatogenesis. Proc Natl Acad Sci USA 120(47):e2310951120
Leung JW et al (2017) ZMYM3 regulates BRCA1 localization at damaged chromatin to promote DNA repair. Genes Dev 31(3):260–274
Hu X et al (2017) Gene knockout of Zmym3 in mice arrests spermatogenesis at meiotic metaphase with defects in spindle assembly checkpoint. Cell Death Dis 8(6):e2910
Patel SR et al (2007) The BRCT-domain containing protein PTIP links PAX2 to a histone H3, lysine 4 methyltransferase complex. Dev Cell 13(4):580–592
Munoz IM, Rouse J (2009) Control of histone methylation and genome stability by PTIP. EMBO Rep 10(3):239–245
Schwab KR, Smith GD, Dressler GR (2013) Arrested spermatogenesis and evidence for DNA damage in PTIP mutant testes. Dev Biol 373(1):64–71
Li H et al (2007) Ewing sarcoma gene EWS is essential for meiosis and B lymphocyte development. J Clin Invest 117(5):1314–1323
Tian H, Billings T, Petkov PM (2021) EWSR1 affects PRDM9-dependent histone 3 methylation and provides a link between recombination hotspots and the chromosome axis protein REC8. Mol Biol Cell 32(1):1–14
Giannattasio T et al (2023) The RNA-binding protein FUS/TLS interacts with SPO11 and PRDM9 and localize at meiotic recombination hotspots. Cell Mol Life Sci 80(4):107
Kuroda M et al (2000) Male sterility and enhanced radiation sensitivity in TLS(−/−) mice. EMBO J 19(3):453–462
Panizza S et al (2011) Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell 146(3):372–383
Claeys Bouuaert C et al (2021) DNA-driven condensation assembles the meiotic DNA break machinery. Nature 592(7852):144–149
Parvanov ED et al (2017) PRDM9 interactions with other proteins provide a link between recombination hotspots and the chromosomal axis in meiosis. Mol Biol Cell 28(3):488–499
Adam C et al (2018) The PHD finger protein Spp1 has distinct functions in the set1 and the meiotic DSB formation complexes. PLoS Genet 14(2):e1007223
Tian H, Billings T, Petkov PM (2018) CXXC1 is not essential for normal DNA double-strand break formation and meiotic recombination in mouse. PLoS Genet 14(10):e1007657
Damm E, Odenthal-Hesse L (2023) Orchestrating recombination initiation in mice and men. Curr Top Dev Biol 151:27–42
Jiang Y et al (2020) CXXC finger protein 1-mediated histone H3 lysine-4 trimethylation is essential for proper meiotic crossover formation in mice. Development 147(6):dev183764
Humphrey E, Cole F (2020) How sex chromosomes break up to get together. Nature 582(7812):346–347
de Massy B (2013) Initiation of meiotic recombination: how and where? Conservation and specificities among eukaryotes. Annu Rev Genet 47:563–599
Pittman DL et al (1998) Meiotic prophase arrest with failure of chromosome synapsis in mice deficient for Dmc1, a germline-specific RecA homolog. Mol Cell 1(5):697–705
Tarsounas M et al (1999) RAD51 and DMC1 form mixed complexes associated with mouse meiotic chromosome cores and synaptonemal complexes. J Cell Biol 147(2):207–220
Inagaki A, Schoenmakers S, Baarends WM (2010) DNA double strand break repair, chromosome synapsis and transcriptional silencing in meiosis. Epigenetics 5(4):255–266
de Boer E et al (2006) Two levels of interference in mouse meiotic recombination. Proc Natl Acad Sci USA 103(25):9607–9612
Pyatnitskaya A, Borde V, De Muyt A (2019) Crossing and zipping: molecular duties of the ZMM proteins in meiosis. Chromosoma 128(3):181–198
de los Santos T et al (2003) The Mus81/Mms4 endonuclease acts independently of double-holliday junction resolution to promote a distinct subset of crossovers during meiosis in budding yeast. Genetics 164(1):81–94
Zhang Q et al (2019) SPO16 binds SHOC1 to promote homologous recombination and crossing-over in meiotic prophase I. Sci Adv 5(1):eaau9780
Ward JO et al (2007) Mutation in mouse hei10, an e3 ubiquitin ligase, disrupts meiotic crossing over. PLoS Genet 3(8):e139
Guiraldelli MF et al (2018) SHOC1 is a ERCC4-(HhH)2-like protein, integral to the formation of crossover recombination intermediates during mammalian meiosis. PLoS Genet 14(5):e1007381
Adelman CA, Petrini JH (2008) ZIP4H (TEX11) deficiency in the mouse impairs meiotic double strand break repair and the regulation of crossing over. PLoS Genet 4(3):e1000042
Krausz C et al (2020) Genetic dissection of spermatogenic arrest through exome analysis: clinical implications for the management of azoospermic men. Genet Med 22(12):1956–1966
Wang W et al (2022) Bi-allelic variants in SHOC1 cause non-obstructive azoospermia with meiosis arrest in humans and mice. Mol Hum Reprod 28(6):gaac015
Song J et al (2023) Novel mutations of TEX11 are associated with non-obstructive azoospermia. Front Endocrinol 14:1159723
Qi Y et al (2023) Pathogenic bi-allelic variants of meiotic ZMM complex gene SPO16 in premature ovarian insufficiency. Clin Genet 104(4):486–490
Yang F et al (2015) TEX11 is mutated in infertile men with azoospermia and regulates genome-wide recombination rates in mouse. EMBO Mol Med 7(9):1198–1210
Moens PB et al (2007) Initiation and resolution of interhomolog connections: crossover and non-crossover sites along mouse synaptonemal complexes. J Cell Sci 120(Pt 6):1017–1027
Argueso JL et al (2004) Competing crossover pathways act during meiosis in saccharomyces cerevisiae. Genetics 168(4):1805–1816
Guiraldelli MF et al (2013) Mouse HFM1/Mer3 is required for crossover formation and complete synapsis of homologous chromosomes during meiosis. PLoS Genet 9(3):e1003383
Xie X et al (2022) Biallelic HFM1 variants cause non-obstructive azoospermia with meiotic arrest in humans by impairing crossover formation to varying degrees. Hum Reprod 37(7):1664–1677
Rao HB et al (2017) A SUMO-ubiquitin relay recruits proteasomes to chromosome axes to regulate meiotic recombination. Science 355(6323):403–407
Kong A et al (2008) Sequence variants in the RNF212 gene associate with genome-wide recombination rate. Science 319(5868):1398–1401
Riera-Escamilla A et al (2019) Sequencing of a ‘mouse azoospermia’ gene panel in azoospermic men: identification of RNF212 and STAG3 mutations as novel genetic causes of meiotic arrest. Hum Reprod 34(6):978–988
Bondarieva A et al (2020) Proline-rich protein PRR19 functions with cyclin-like CNTD1 to promote meiotic crossing over in mouse. Nat Commun 11(1):3101
Holloway JK et al (2014) Mammalian CNTD1 is critical for meiotic crossover maturation and deselection of excess precrossover sites. J Cell Biol 205(5):633–641
Qiao H et al (2014) Antagonistic roles of ubiquitin ligase HEI10 and SUMO ligase RNF212 regulate meiotic recombination. Nat Genet 46(2):194–199
Gray S, Cohen PE (2016) Control of meiotic crossovers: from double-strand break formation to designation. Annu Rev Genet 50:175–210
Palmer N et al (2020) A novel function for CDK2 activity at meiotic crossover sites. PLoS Biol 18(10):e3000903
Barchi M et al (2008) ATM promotes the obligate XY crossover and both crossover control and chromosome axis integrity on autosomes. PLoS Genet 4(5):e1000076
Lange J et al (2011) ATM controls meiotic double-strand-break formation. Nature 479(7372):237–240
Liu C et al (2021) Paternal USP26 mutations raise klinefelter syndrome risk in the offspring of mice and humans. EMBO J 40(13):e106864
Yu XC et al (2021) A new TEX11 mutation causes azoospermia and testicular meiotic arrest. Asian J Androl 23(5):510–515
Sha Y et al (2018) A novel TEX11 mutation induces azoospermia: a case report of infertile brothers and literature review. BMC Med Genet 19(1):63
Tian H et al (2019) Disruption of ubiquitin specific protease 26 gene causes male subfertility associated with spermatogenesis defects in mice. Biol Reprod 100(4):1118–1128
Ma H et al (2022) RAD51AP2 is required for efficient meiotic recombination between X and Y chromosomes. Sci Adv 8(2):eabk1789
Arango NA et al (2013) Meiosis I arrest abnormalities lead to severe oligozoospermia in meiosis 1 arresting protein (M1ap)-deficient mice. Biol Reprod 88(3):76
Tu C et al (2020) An M1AP homozygous splice-site mutation associated with severe oligozoospermia in a consanguineous family. Clin Genet 97(5):741–746
Wyrwoll MJ et al (2020) Bi-allelic mutations in M1AP are a frequent cause of meiotic arrest and severely impaired spermatogenesis leading to male infertility. Am J Hum Genet 107(2):342–351
Li Y et al (2023) M1AP interacts with the mammalian ZZS complex and promotes male meiotic recombination. EMBO Rep 24(2):e55778
Johnston SE et al (2016) Conserved genetic architecture underlying individual recombination rate variation in a wild population of soay sheep (Ovis aries). Genetics 203(1):583–598
Kadri NK et al (2016) Coding and noncoding variants in HFM1, MLH3, MSH4, MSH5, RNF212, and RNF212B affect recombination rate in cattle. Genome Res 26(10):1323–1332
Gershoni M et al (2023) A pathogenic variant in the uncharacterized RNF212B gene results in severe aneuploidy male infertility and repeated IVF failure. HGG Adv 4(3):100189
Shao Q et al (2023) ATF7IP2, a meiosis-specific partner of SETDB1, is required for proper chromosome remodeling and crossover formation during spermatogenesis. Cell Rep 42(8):112953
Alavattam KG et al (2024) ATF7IP2/MCAF2 directs H3K9 methylation and meiotic gene regulation in the male germline. Genes Dev 38(3–4):115–130. https://doi.org/10.1101/gad.351569.124
Lowe X et al (2001) Frequency of XY sperm increases with age in fathers of boys with klinefelter syndrome. Am J Hum Genet 69(5):1046–1054
Arnedo N et al (2006) Sperm aneuploidy in fathers of klinefelter’s syndrome offspring assessed by multicolour fluorescent in situ hybridization using probes for chromosomes 6, 13, 18, 21, 22. X and Y Hum Reprod 21(2):524–528
De Souza E, Morris JK, EUROCAT Working Group (2010) Case-control analysis of paternal age and trisomic anomalies. Arch Dis Child 95(11):893–897
Eaker S et al (2001) Evidence for meiotic spindle checkpoint from analysis of spermatocytes from robertsonian-chromosome heterozygous mice. J Cell Sci 114(Pt 16):2953–2965
Malmanche N, Maia A, Sunkel CE (2006) The spindle assembly checkpoint: preventing chromosome mis-segregation during mitosis and meiosis. FEBS Lett 580(12):2888–2895
Jeganathan KB, van Deursen JM (2006) Differential mitotic checkpoint protein requirements in somatic and germ cells. Biochem Soc Trans 34(Pt 4):583–586
Kauppi L (2021) USP26: a genetic risk factor for sperm X-Y aneuploidy. EMBO J 40(13):e108552
Fukuda T et al (2012) Phosphorylation of chromosome core components may serve as axis marks for the status of chromosomal events during mammalian meiosis. PLoS Genet 8(2):e1002485
Acknowledgements
Not applicable.
Funding
Open access funding provided by Università degli Studi di Roma Tor Vergata within the CRUI-CARE Agreement. This work was supported “Gruppi di Ricerca 2020” from Regione Lazio, Italy (no. A0375-2020-36618) to MB, and Fondo di Beneficenza Intesa Sanpaolo (no. B/2021/0228) to MB. The founders had no role in study design, data collection and analysis, decision to publish, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
The first draft of the manuscript was written by Matteo Lampitto and Marco Barchi. Marco Barchi made the changes to the manuscript after peer review. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no financial or non-financial interests to disclose.
Ethics approval and consent to participate
Not applicable.
Consent for publication
The research did not involve human participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lampitto, M., Barchi, M. Recent advances in mechanisms ensuring the pairing, synapsis and segregation of XY chromosomes in mice and humans. Cell. Mol. Life Sci. 81, 194 (2024). https://doi.org/10.1007/s00018-024-05216-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05216-0