Abstract
N6-methyladenosine (m6A), the most prevalent internal mRNA modification, plays a critical role in physiological processes by regulating gene expression through modulation of mRNA metabolism at multiple stages. In recent years, m6A has garnered significant attention for a deeper understanding of the initiation, progression, and drug resistance of various cancers, including hematological malignancies. Dysregulation of m6A has been implicated in both cancer promotion and suppression. m6A methylation is a complex regulatory process involving methyltransferases (writers), demethylases (erasers), and proteins that recognize specific m6A modifications (readers). This intricate interplay presents challenges for precisely modulating m6A levels, either globally or at specific sites. This review specifically focuses on the role of m6A in chronic myeloid leukemia (CML), a blood cancer characterized by the BCR-ABL1 fusion. We emphasize its impact on leukemia cell survival and drug resistance mechanisms. Notably, inhibitors targeting m6A regulators show promise in preclinical models, suggesting a potential therapeutic avenue for CML. Integrating our understanding of m6A biology with current treatment strategies may lead to more effective therapies, especially for patients with advanced-stage or resistant CML.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
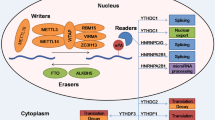
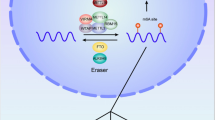
m6A is the most abundant internal modification of messenger RNAs (mRNAs) and long non-coding RNAs (lncRNAs) (reviewed in [1]). Despite its initial discovery in 1974, the field of m6A biology remained largely unexplored for decades until the development of sequencing methods for mapping m6A modification in RNA species [2, 3]. However, quantifying m6A abundance at specific positions within individual transcripts remains a significant challenge, despite the development of numerous methodologies for mapping m6A distribution [4,5,6,7]. Currently, it is undoubtedly the most extensively studied RNA modification with the greatest therapeutic potential. Methylation at the N6 position of the target adenosine residue in RNA occurs co-transcriptionally by the methyltransferase complex composed of methyltransferase-like protein 3 (METTL3, also known as MT-A70) and methyltransferase-like protein 14 (METTL14) (Fig. 1) (reviewed in [8]). Although METTL14 lacks catalytic activity itself, it plays a crucial role in recognizing the DRACH sequence (D = A, G, or U; R = A or G; H = A, C, or U; A methylated adenosine) in RNA, facilitating methylation of the adenosine within this motif by the catalytic subunit, METTL3. This complex is further regulated by a multiprotein assembly including Wilms tumor 1-associated protein (WTAP), Vir-like m6A methyltransferase-associated (VIRMA, also known as KIAA1429), Cbl proto-oncogene like 1 (CBLL1), RNA-binding motif 15 (RBM15), and zinc finger CCCH-type containing 13 (ZC3H13) [8]. m6A is not randomly distributed, but rather targeted to specific regions within RNA molecules [8]. These enriched regions often occur near the stop codon, within the 3' untranslated region (3'UTR), or in particularly long exons. Furthermore, it has also been found in intronic regions of pre-mRNAs and long noncoding RNAs [8]. Several factors influence the precise location of m6A and thus its stoichiometry. These include the recruitment of the modifying complex by transcription factors, epigenetic modifications, or RNA-binding proteins to specific regions on the RNA. Additionally, the presence of protein complexes, such as the exon junction complex, can sterically hinder methylation in certain areas [9,10,11]. Notably, the methyltransferase METTL16, though primarily targeting U6 snRNA, can also recognize and modify certain mRNA and lncRNA substrates in a DRACH-independent manner [1]. The m6A modification can be removed by two "erasers", ALKBH5 (alkB homolog 5) and FTO (fat-mass and obesity-associated protein) that belongs to the Fe2+ and 2-oxoglutarate (2OG)-dependent AlkB dioxygenase family [12, 13]. ALKBH5 exhibits high specificity for m6A in mRNA, while FTO acts on a broader range of modifications, including m6Am at the cap, internal m6Am within snRNAs, and m1A within tRNAs. FTO demethylase activity towards specific substrates is influenced by its subcellular localization and interaction with RNA binding proteins (Fig. 1) [14, 15].
Writers and erasers of m6A modification. m6A modification within RNA molecules is dynamically regulated by a balance between "writer" methyltransferase complexes and "eraser" demethylase enzymes. A In the nucleus, the METTL3-METTL14 complex acts as the primary writer, modifying adenines co-transcriptionally onto mRNA. Conversely, two demethylases, ALKBH5 and FTO, can remove m6A from nuclear RNA substrates. B Notably, nuclear FTO exhibits broader substrate specificity, demethylating internal m6A within U6 small nuclear RNA (snRNA) and m6Am in the cap region of U1 and U2 snRNAs. C Additionally, cytoplasmic FTO contributes to m6A regulation by demethylating both m6Am in the mRNA cap and internal m6A modifications within mature mRNAs. D Furthermore, METTL3 can indirectly promote translation by directly binding to mRNAs in the cytoplasm. Across all panels, DNA is depicted as a grey line, the nucleosome as a green barrel, RNA as a black line, the 5′ cap structure as a black dot, the poly(A) tail as An, the m6A modification as a red circle, and the m6Am modification as a yellow circle.
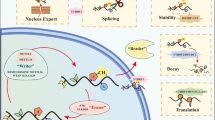
m6A plays a crucial role in RNA metabolism, regulating virtually all stages of its expression, including transcription, splicing, export, stability, and translation (Fig. 2) (reviewed in [16]). The effects of m6A on gene expression are mediated by the binding of "reader" proteins Among the direct readers, proteins of the YTH domain family play a prominent role, including, YTHDC1, the only nuclear member, YTHDC2, YTHDF1, YTHDF2, and YTHDF3. These readers possess a domain that specifically recognizes the m6A modification regardless of the RNA sequence [8, 16]. YTHDC1 plays a part in transcriptional regulation and contributes to various nuclear RNA processing events, including alternative splicing, polyadenylation, and nuclear export [8, 16]. Additionally, it regulates the function of nuclear lncRNAs, such as XIST and MALAT1, and participates in the formation of nuclear RNA condensates [8, 16]. Despite their high sequence homology, the YTHDF proteins were initially ascribed distinct functions. The prevailing view held that YTHDF1 and YTHDF3 functioned as translation activators, while YTHDF2 was thought to promote mRNA degradation. New evidence suggests that YTHDF proteins function independently, potentially exhibiting redundancy in their ability to promote mRNA degradation [17, 18]. YTHDC2, the only RNA helicase-containing YTH protein, can also stimulate mRNA translation and decay by binding to m6A-modified transcripts. YTHDC2 plays a crucial role in the development and maturation of germ cells, particularly during spermatogenesis. However, recent research indicates that its function in spermatogenesis is independent of m6A modification [19]. Additionally, RNA-binding proteins, such as members of the IGF2BP family (IGFBP1, IGFBP2 and IGFBP3), FMRP, FXR1, and FXR2, exhibit enhanced binding to their consensus sequences when m6A is present, although the exact mechanism remains unclear. Furthermore, the presence of m6A can induce alterations in the local three-dimensional structure of RNA, a phenomenon termed "m6A switch," favouring or impeding the binding of RNA-binding proteins [8, 16]. Among these proteins are heterogeneous nuclear ribonucleoproteins (hnRNPs), including hnRNPA2B1, hnRNPC, and hnRNPG, as well as Ras GTPase-activating protein-binding protein 1 (G3BP1) and G3BP2. The binding of the latter to RNA is disfavoured by the presence of m6A [8, 16].
Modulation of mRNA function by m6A modification. m6A modifications on RNA molecules exert widespread effects on various cellular processes. m6A reader proteins specifically recognize and bind to this mark, dictating the functional consequences of m6A on RNA expression. Within the nucleus, m6A can influence gene regulation through diverse mechanisms: A regulation of H3K9 Methylation, the YTHDC1 reader can influence transcription by regulating the methylation state of histone H3 at lysine 9 (H3K9); B Chromatin-Associated RNA (chrRNA) Targeting, YTHDC1 binds to and mediates the degradation of chrRNAs, thereby impacting chromatin structure and function; C nuclear mRNA condensates and stability, YTHDC1 can directly bind specific mRNAs and promote their liquid–liquid phase separation into condensates, potentially enhancing their stability; D alternative splicing regulation, the presence of m6A in pre-mRNAs can influence alternative splicing events by recruiting YTHDC1; E m6A switch, m6A can induce conformational changes in RNA structure, leading to altered binding of regulatory proteins and affecting splicing or other RNA processing events; F mRNA export, YTHDC1, along with other factors like FMRP, can stimulate the export of mature mRNAs from the nucleus to the cytoplasm. Within the cytoplasm, m6A modifications exert a multifaceted influence on mRNA metabolism. G The position of the m6A mark on the mRNA dictates its impact on translation. m6A located in the 5′-UTR can influence cap-independent translation. The ABCF1 and eIF3 readers can bind m6A present in the 5′-UTR of mRNAs and stimulate translation initiation, while the YTHDF1 and YTHDF3 readers can stimulate cap-dependent translation by binding m6A in the 3′ regions of mRNA. However, recent research has yielded conflicting results regarding the involvement of YTHDF1 and YTHDF3 in regulating translation. Furthermore, cytoplasmic METTL3 and METTL16 can regulate cap-dependent translation by a catalytic-independent mechanism. H mRNA stability, YTHDF1/2/3 readers can trigger mRNA decay pathways when they bind to m6A. Conversely, IGF2BP1/2/3 can enhance the stability of mRNAs containing m6A modifications. I Finally, YTHDFs readers can promote liquid–liquid phase separation into condensates of m6A-modified mRNAs. Across all panels, DNA is depicted as a grey line, the nucleosome as a green barrel, RNA as a black line, the 5' cap structure as a black dot, the poly(A) tail as An, the m6A modification as a red circle, the m6Am modification as a yellow circle, and the RNA degradation machinery as a red Pac-Man shape
Given its critical role in gene expression, dysregulation of m6A levels or its reader proteins can have profound consequences for cellular function. This is reflected in the altered expression of m6A levels, or its readers observed in various cancers, where they contribute to tumour initiation, progression, treatment response, and resistance (reviewed in [20]). In this review, we describe the role of m6A in chronic myeloid leukemia (CML), a hematological malignancy characterized by the clonal expansion of myeloid cells [21], and discuss how selective inhibitors targeting the writers and erasers of this modification could be incorporated into therapeutic treatments of CML.
Chronic myeloid leukemia (CML)
Hematological malignancies were among the first cancers where the oncogenic role of m6A was demonstrated, particularly acute myeloid leukemia (AML) [22,23,24]. Subsequently, m6A-linked oncogenic mechanisms have also been identified in CML [25,26,27]. CML is driven by the Philadelphia chromosome, resulting from a t(9;22)(q34;q11) translocation that creates the BCR-ABL1 fusion gene. The BCR-ABL1 fusion gene encodes a constitutively active tyrosine kinase, driving uncontrolled proliferation, impaired differentiation, and increased survival of myeloid progenitors by activating downstream pro-oncogenic signalling pathways including JAK/STAT, MAPK, and PI3K/AKT/mTOR pathways [21, 28, 29].
CML progresses through two distinct phases: the chronic phase and the acute phase, also known as blast crisis [30]. Some patients may have a transition between the two phases referred to as the accelerated phase, which is still lacking a precise biological definition. The chronic phase is the initial stage, characterized by relatively stable clinical manifestations and slower disease progression compared to later phases. Patients in chronic phase may be asymptomatic or have mild symptoms. Prior to the development of BCR-ABL1 tyrosine kinase inhibitors (TKIs), chronic phase CML patients invariably progressed to advanced stages of the disease, characterized by accelerated leukemic cell proliferation. This progression typically occurred within a median time frame of approximately 5 years. Therefore, timely diagnosis and initiation of targeted therapy with TKIs are paramount to achieving durable responses and preventing disease progression. The acute phase of CML is characterized by impaired differentiation and rapid proliferation of immature blasts (> 20% of bone marrow cells). While resembling AML in most cases, approximately 25% of patients exhibit pre-B lymphoblastic leukemia or, less frequently, T lymphoblastic transformation. Molecular and cytogenetic analyses may show additional chromosomal abnormalities and mutations in the epigenetic regulators such as ASXL1, DNMT3A, IDH1, and SETBP1 [31], indicative of clonal evolution and disease progression. This phase is associated with a dismal prognosis, with few treatment options available, and median survival of approximately 6 months. Early diagnosis and targeted therapy with TKIs are crucial for managing CML, with the advent of TKIs like imatinib dramatically transforming the prognosis for chronic phase patients, leading to deep molecular responses and prolonged survival. However, challenges remain, including persistence of leukemic stem cells that evade TKIs, development of TKI resistance, and long-term treatment-related toxicities. Aberrant m6A patterns have been implicated in the dysregulation of critical oncogenes and tumour suppressors in CML, thereby contributing to disease initiation, progression, and therapy resistance.
m6A roles in CML
Oncogenic roles for m6A regulators
METTL3 and METTL14 are upregulated in both primary chronic phase CML cells and established CML cell lines, including those derived from blast crisis [25]. Importantly, knocking down their expression leads to cell cycle arrest and decreased viability in both primary CML cells and CML cell lines [25]. Notably, this effect extends to imatinib-resistant CML cells, highlighting METTL3/METTL14 as a potential therapeutic target.
In CML, the interplay between global protein synthesis regulation and selective translation of specific mRNAs is crucial [32]. Similar to observations in AML, the METTL3-METTL14 complex promotes high levels of the MYC oncogene in CML [25]. MYC is one of the most common oncogenes in human cancers, exerting its growth-promoting effects primarily by enhancing ribosome biogenesis and mRNA translation [33]. The METTL3-METTL14 complex promotes MYC expression through a two-pronged approach: indirectly via SP1 regulation, a transcriptional activator of the MYC gene, and directly by enhancing MYC mRNA translation. Furthermore, CML exhibits cytoplasmic delocalization of METTL3. In this compartment, independent of its catalytic activity, METTL3 promotes the translation of PES1 and WTAP. PES1 is involved in ribosome biogenesis and cell cycle progression, while WTAP positively regulates the METTL3-METTL14 complex [24, 34]. Ultimately, the positive regulation of MYC and PES1 by METTL3 contributes to the aberrant protein translation that characterizes the leukemogenic activity of BCR-ABL1 (Fig. 3).
Oncogenic role for METTL3 in CML. A The METTL3-METTL14 complex promotes MYC expression through a two-pronged approach: indirectly by regulating SP1, a transcriptional activator of the MYC gene, and directly by enhancing MYC mRNA translation. B In CML cells, METTL3 is delocalized to the cytoplasm. Here, independent of its catalytic activity, METTL3 promotes the translation of PES1 and WTAP. PES1 is involved in ribosome biogenesis and cell cycle progression, while WTAP positively regulates the METTL3-METTL14 complex. Ultimately, the positive regulation of MYC and PES1 by METTL3 contributes to the aberrant protein translation that characterizes the leukemogenic activity of BCR-ABL1. Across all panels, RNA is depicted as a black line, the m7G at the 5' cap structure as a black dot, the poly(A) tail as An, and the m6A modification as a red circle.
VIRMA (KIAA1429), a regulator of the METTL3-METTL14 methyltransferase complex, is upregulated in acute-phase CML, contributing to elevated m6A levels [26]. Furthermore, VIRMA knockdown in CML cell lines recapitulates the effect of targeting the methyltransferase complex, leading to decreased proliferation and viability [26]. Importantly, VIRMA's m6A-stimulating action regulates RAB27B mRNA stability, a protein involved in imatinib efflux, through the YTHDF1 reader [26]. Reduced RAB27B levels due to VIRMA knockdown increase intracellular imatinib levels, consequently enhancing TKI sensitivity (Fig. 4A) [26].
VIRMA and YBX1: mediators of m6A-driven leukemogenesis in CML. A VIRMA promotes m6A modifications, which, through the YTHDF1 reader protein, stabilize RAB27B mRNA. RAB27B is a protein involved in imatinib efflux from cells. Reduced RAB27B due to VIRMA knockdown increases intracellular imatinib levels, enhancing TKI sensitivity. B YBX1 cooperates with IGF2BP in CML cells to regulate the m6A-dependent stability of YWHAZ mRNA, which activates the PI3K/AKT/mTOR signaling pathway, a pathway important for cell survival. This mechanism likely contributes to YBX1's role in CML LSC survival. Across all panels, RNA is depicted as a black line, the m7G at the 5' cap structure as a black dot, the poly(A) tail as An, and the m6A modification as a red circle.
An additional oncogenic mechanism mediated by m6A has recently emerged, centered on the RNA-binding protein YBX1 [27]. Upregulated in CML patients, particularly those in the acute phase, YBX1 cooperates with IGF2BP protein in CML cell lines to regulate the m6A-dependent stability of YWHAZ mRNA, an activator of the PI3K/AKT/mTOR signaling pathway. This mechanism likely contributes to the role of YBX1 in CML leukemia stem cells (LSCs) survival (Fig. 4B).
Interplay between m6A and lncRNAs
The lncRNA NEAT1 (Nuclear paraspeckle assembly transcript 1) exhibits progressive downregulation in CML patients, with the most pronounced decrease observed in the acute phase [35]. NEAT1 downregulation coincides with increased m6A levels due to CML-associated upregulation of the methyltransferase complex [24]. Notably, NEAT1 overexpression in CML cell lines (K562 and KCL22) suppresses cellular viability, enhances apoptosis, and inhibits tumour growth in xenograft models [35]. Mechanistically, NEAT1 acts as a competing endogenous RNA (ceRNA) for miR-766-5p, which downregulates the tumour suppressor CDKN1A (Yao 2021) (Fig. 5A). Furthermore, NEAT1 overexpression in K562 cells negatively regulates the expression of the ABCG2 transporter protein, which is involved in the efflux of drugs from the cell and the development of drug resistance [36].
Regulation of lncRNA activity by m6A in CML. A NEAT1 expression progressively decreases in CML patients, with the sharpest decline occurring during the acute phase. This decrease coincides with elevated cellular m6A levels, potentially linked to CML-associated upregulation of the m6A methyltransferase complex. NEAT1 acts as a ceRNA for miR-766-5p, preventing it from downregulating the tumor suppressor CDKN1A. B The oncogenic LINC00470 guides the m6A methyltransferase METTL3 to the mRNA of the tumor suppressor PTEN. This interaction increases m6A levels of PTEN, leading to its degradation and subsequent activation of the PI3K/AKT signaling pathway, which promotes cell survival and potentially contributes to TKI resistance in CML
LncRNAs can also act as regulators of METTL3 activity. A prime example is LINC00470, which guides METTL3 to the mRNA of the tumour suppressor PTEN [37]. This interaction leads to an increase in m6A modification of PTEN mRNA, followed by its subsequent degradation. This results in the activation of the oncogenic PI3K/AKT signalling pathway, leading to increased AKT activity and its downstream targets (Fig. 5B). Furthermore, LINC00470-mediated PTEN reduction promotes autophagy via AKT hyperactivation promoting TKI resistance. Notably, decreasing LINC00470 levels suppresses tumour growth in xenograft models, including those derived from imatinib-resistant K562 cells [37]. These findings suggest a potential role for LINC00470 in TKI resistance mechanisms of CML. Interestingly, depletion of METTL3 in K562 cells abrogated the LINC00470-induced downregulation and degradation of PTEN mRNA and protein, restoring normal m6A modification levels in PTEN [37].
m6A and DNA damage
Genomic instability is a hallmark of TKI-resistant chronic phase CML, leading to disease relapse and/or malignant progression. This phenomenon likely arises from an aberrant cellular response to elevated DNA damage, including high levels of ROS-induced oxidative damage and compromised DNA damage repair (DDR) mechanisms [38]. Recent studies have shed light on a potential role for m6A in this context. Upon DNA damage, ataxia telangiectasia mutated (ATM) phosphorylates METTL3 promoting its localization at damaged sites. Here, METTL3 methylates nascent RNA transcribed at double-strand breaks (DSBs), potentially facilitating the formation of DNA-RNA hybrids (R-loops) and enhancing DDR (Fig. 6) [39, 40]. R-loops additionally, m6A modifications may regulate the expression of DNA repair genes, adding another layer of complexity to this interplay. Consequently, METTL3 depletion sensitizes cancer cells to DNA damage-based therapies [39].
Regulation of DNA damage repair (DDR) by m6A modification. DNA damage triggers METTL3 recruitment to double-strand breaks (DSBs) via ATM phosphorylation or PARP1/2 interaction. METTL3 methylates RNA near the damage site, stabilizing these m6A-modified transcripts through YTHDC1 or YTHDF2 binding. Stabilized m6A-modified RNAs form R-loops with DNA, recruiting repair proteins like RAD51/BRCA1 for homologous recombination repair (HRR) or DNA Pol κ (Pol κ) for nucleotide excision repair (NER)
While a direct correlation between m6A and DNA damage in CML remains elusive, the emerging link between m6A signalling and DDR pathways holds promise for novel therapeutic strategies. This is particularly relevant because BCR-ABL1, the driver oncogene in CML, downregulates BRCA1, a critical protein for homologous recombination repair (HRR) [41]. Consequently, BCR-ABL1 activity leads to increased DNA damage. PARP inhibitors (PARPi), such as olaparib, exploit this vulnerability by further inhibiting base excision repair [41]. PARP1 plays a crucial role in repairing single-strand breaks (SSBs) in DNA. It detects and binds to damaged DNA, facilitating the recruitment of other repair proteins to the site of damage. These unrepaired SSBs can convert into single-ended DSBs during replication, a type of damage that is predominantly repaired by BRCA1-mediated HRR [41]. In HRR-deficient cells, such as CML, PARPi treatment in combination with METTL3 inactivation may induce a "synthetic lethality" effect, where the combined inhibition of two DNA repair pathways becomes lethal to the cancer cells. Notably, olaparib treatment has been shown to decrease the proliferation of CML cells in vitro and improve survival in a BCR-ABL1-dependent leukemia mouse model, highlighting its therapeutic potential in CML [42, 43]. Further research is warranted to explore the potential therapeutic effects of combining PARPi with METTL3 inhibitors in CML cells not responding to TKIs.
m6A and TKI resistance
Point mutations that reduce drug-binding affinity are a major cause of TKI resistance, although second and third generation TKIs are often designed to overcome these mutations (reviewed in [31]). Currently, five distinct TKIs against BCR-ABL1 fusion are available for the treatment of the chronic phase of CML: imatinib, dasatinib, nilotinib, bosutinib, ponatinib and asciminib. However, switching TKIs in resistant patients might inadvertently promote the emergence of complex mutations that render the cancer cells insensitive to most, if not all, available TKIs. Additionally, a subset of patients develops TKI resistance despite lacking identifiable resistance mutations. Patients harbouring TKI resistance mutations exhibit an increased propensity to transition into the blast crisis.
In CML cell lines, a decrease in global m6A levels in mRNAs has been linked to TKI resistance against imatinib and nilotinib [44]. FTO upregulation in resistant cells is thought to cause a reduction in m6A levels, leading to subsequent upregulation of mRNA expression for genes involved in cellular proliferation and survival, particularly the myeloid epithelial reproductive tyrosine kinase receptor (MERTK) and B-cell lymphoma 2 (BCL-2) (Fig. 7). Both MERTK and BCL-2 are well-established contributors to reduced apoptosis, increased metastasis, and drug resistance. Notably, resistance to apoptosis is a hallmark of CML LSCs. This link is further supported by similar findings in primary CML cells with induced nilotinib resistance. These cells also displayed increased FTO levels and decreased m6A levels in the MERTK and BCL-2 mRNA. This finding suggests that FTO inhibition could be employed in combination with TKI therapy to mitigate the risk of resistance in CML. Intriguingly, elevated FTO expression in AML is associated with resistance to both TKI therapy and chemotherapy. This finding suggests that FTO inhibition could be a valuable strategy to overcome multidrug resistance and prevent disease relapse in leukemia.
m6A and TKI resistance. TKI-resistant cells often exhibit upregulation of FTO and decreased m6A methylation levels. This reduction in cellular m6A levels leads to increased stability and expression of mRNAs encoding proteins involved in cell proliferation and survival, such as the MERTK and BCL-2 proteins. Consequently, TKI-resistant cells experience reduced apoptosis, increased metastasis, and ultimately, resistance to TKI therapy. Conversely, strategies that decrease FTO expression or inhibit its catalytic activity can restore m6A levels, potentially leading to decreased mRNA stability of these pro-survival genes and a reduced risk of TKI resistance. Across all panels, RNA is depicted as a black line, the m7G at the 5' cap structure as a black dot, the poly(A) tail as An, and the m6A modification as a red circle.
m6A as therapeutic target
Through a combination of virtual screening and rational design researchers have developed a repertoire of potent and selective m6A machinery inhibitors. Notably, some of these inhibitors exhibit high efficacy in pre-clinical models of AML. The first inhibitors of m6A regulators were developed against the enzyme FTO, whose crystal structure has been known since 2010 [45]. While elevated m6A levels are generally considered oncogenic, the m6A eraser FTO has emerged as a promising therapeutic target in certain AML subtypes and, notably, in acquired TKI resistance in both AML and CML [44]. The first selective FTO inhibitors were derived from the meclofenamic acid molecule. Particularly, FB23-2 exhibited potent antiproliferative activity against AML cells both in vitro and in vivo, but still with a IC50 in the micromolar range. Structure-based virtual screening recently identified CS2 (formerly brequinar) as a potent inhibitor of FTO demethylase activity, demonstrating potent antileukemic activity in vitro and in vivo with a nanomolar IC50. However, CS2 also potently inhibits human dihydroorotate dehydrogenase (hDHODH), an enzyme crucial for de novo uridine synthesis, and targeting hDHODH itself has independent antileukemic activity. This raises the question of whether CS2's primary mechanism in leukemia involves hDHODH or FTO inhibition. Notably, CML cells are particularly sensitive to hDHODH inhibition [46]. Therefore, CS2 could offer a dual benefit: antiproliferative activity through hDHODH inhibition and potential mitigation of resistance via FTO inhibition. Furthermore, the activity of FTO extends beyond demethylating m6A on mRNA. Its roles in demethylating other RNA substrates, including m6A on snRNAs, cap-associated m6Am on mRNAs and snRNAs, and m1A on tRNA, which have been largely neglected in recent studies, warrant further investigation to comprehensively understand the biological effect of FTO inhibition.
Given the oncogenic role of METTL3 in various malignancies, several inhibitors have been developed (reviewed in [47]). Early attempts to develop METTL3 inhibitors focused on designing competitive inhibitors of its cofactor S-adenosylmethionine (SAM). These inhibitors are therefore capable of suppressing METTL3 catalytic activity when in complex with METTL14 but not the independent catalytic action that has been described for the protein when localized in the cytoplasm. Among these, UZH2 and STM2457 stand out as the most selective and effective inhibitors in preclinical leukemia models. These inhibitors have demonstrated efficacy in a broad spectrum of acute myeloid leukemias, and mice and human PDX AML models, suggesting their potential effectiveness in CML blast crisis and TKI-resistant cells, where METTL3 knockdown exhibits a robust antiproliferative effect [25]. However, the cytoplasmic delocalization of METTL3 in CML, where it exerts an oncogenic role independent of its catalytic activity, could potentially limit the efficacy of these inhibitors. The development of degradative inhibitors, such as proteolysis targeting chimeras (PROTACs), like those recently derived from the UZH2 inhibitor, holds promise as a therapeutic strategy for CML patients in blast crisis.
Furthermore, inhibitors targeting FTO and METTL3 could be employed in combination therapy with clinically approved molecules. For instance, considering that FTO upregulation of MERTK and BCL2 contributes to CML TKI resistance, a combination of catalytic FTO inhibitors with MERTK-targeting monoclonal antibodies or selective BCL2 inhibitors like venetoclax could be explored. Similarly, given the recent demonstration that METTL3 catalytic inhibition sensitizes tumor cells to genotoxic agents, such as the PARPi olaparib [48], METTL3 inhibitors could be utilized to enhance the efficacy of these agents to eliminate LSCs and to induce synthetic lethality in CML patients that do not respond to standard TKI-based therapy.
Future directions
Understanding the role of m6A regulators in CML holds immense potential for novel therapeutic strategies. As discussed, even if CML and AML are characterized by different genetic alterations, both diseases can progress to a state of rapid blast cell proliferation, potentially sharing overlapping dysregulation in cellular pathways. Therefore, investigating m6A regulators and pathways implicated in AML is crucial to gain insights into their potential role in CML, particularly in blast crisis. Here, we explore key areas for future investigation. While the m6A demethylase ALKBH5 has been implicated in promoting leukemogenesis and poor prognosis in AML [49], its role in CML remains largely unexplored. Future studies should focus on understanding whether ALKBH5 expression or activity is dysregulated in CML patients compared to healthy controls. Additionally, investigating the impact of ALKBH5 activity on the response of CML cells to current therapies or its influence on leukemia development could shed light on its potential role in the disease process. These investigations hold promise for revealing whether targeting ALKBH5 could be a viable therapeutic strategy for CML. Furthermore, the role of m6A readers in CML remains largely unelucidated. YTHDF2 is upregulated in AML samples [50]. The protein is essential for LSC self-renewal, and its depletion compromises the ability of LSCs to expand and thus propagate AML. Therein, it will be interesting to analyze its downregulation in CML. In AML, the nuclear m6A reader YTHDC1 promotes the formation of m6A-containing mRNA condensates, stabilizing oncogenic transcripts like MYC mRNA and promoting cancer cell survival [51]. MYC also plays a critical role in CML, suggesting that this mechanism might be relevant in this disease as well. Here, further investigation is warranted to determine if YTHDC1 and m6A-mediated mRNA stabilization contribute to CML pathogenesis. Similarly, Insulin-like growth factor-2 mRNA-binding proteins (IGF2BP1/2/3) have been found to be overexpressed in AML, and to regulate the stability of specific mRNAs, including MYC, in an m6A-dependent manner to promote tumor progression [52,53,54]. Also, the functions of the newly identified m6A reader proteins, such as PRRC2A/B and FXR1, in the context of leukemia remain largely unexplored. Future investigations should explore the expression patterns of these readers in CML cells compared to healthy controls and their functional consequences in m6A-regulated pathways relevant to CML development and progression. Additionally, the potential of manipulating reader function (e.g., through small molecule inhibitors) as a viable therapeutic strategy for CML warrants further exploration.
Finally, the intricate interplay between m6A RNA methylation and the unfolded protein response (UPR) pathway remains largely unexplored in CML. While the UPR pathway is a known therapeutic target in CML [55, 56], the interplay between UPR and m6A RNA methylation remains poorly understood in this context. Existing research suggests a complex and cell-type specific role for m6A regulators in the ER stress response. Studies in mouse liver cells and other cell types demonstrate opposing effects of METTL3 and METTL14 on ER stress, highlighting the context-dependent nature of this interaction [57,58,59]. Similarly, studies in breast cancer reveal opposing functions of m6A regulators YTHDF2 and VIRMA [60, 61]. YTHDF2 reader downregulation induces ER stress and apoptosis, while VIRMA overexpression increases UPR regulator expression under stress, possibly through m6A modification. Interestingly, ER stress can also influence m6A machinery by elevating METTL3/METTL14 levels, promoting mRNA stability for proteins involved in ER-phagy (a specific form of autophagy targeting ER components) [62]. Importantly, knocking out METTL3/METTL14 sensitizes breast cancer cells to ER stress-inducing drugs [62]. These findings suggest a complex loop between m6A and ER stress, where each component can influence the other. Furthermore, these observations suggest that leukemia cells with elevated m6A levels might be more sensitive to UPR inducers. However, the molecular mechanisms linking ER stress-mediated m6A regulation and its role in the UPR pathway remain largely unexplored in leukemia.
By addressing these future directions, researchers can gain a deeper understanding of the m6A landscape in CML.
Conclusions
In conclusion, the burgeoning field of m6A RNA modification provides a compelling framework to elucidate the molecular underpinnings of CML pathogenesis, progression and resistance. Overall, targeting the m6A machinery offers a novel and promising therapeutic approach for CML, with the potential to improve treatment outcomes, particularly in patients with advanced disease or TKI resistance. By integrating the knowledge of m6A biology with ongoing therapeutic advancements, we can strive for more effective and personalized treatment strategies for CML patients.
Across all panels, RNA is depicted as a black line, the m7G at the 5' cap structure as a black dot, the poly(A) tail as An, and the m6A modification as a red circle.
Availability of data and material
Not applicable.
References
Sendinc E, Shi Y (2023) RNA m6A methylation across the transcriptome. Mol Cell 83:428–441. https://doi.org/10.1016/j.molcel.2023.01.006
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149:1635–1646. https://doi.org/10.1016/j.cell.2012.05.003
Dominissini D et al (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485:201–206. https://doi.org/10.1038/nature11112
Linder B, Grozhik AV, Olarerin-George AO, Meydan C, Mason CE, Jaffrey SR (2015) Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods 12:767–772. https://doi.org/10.1038/nmeth.3453
Garcia-Campos MA et al (2019) Deciphering the “m6A Code” via antibody-independent quantitative profiling. Cell 178:731–747. https://doi.org/10.1016/j.cell.2019.06.013
Shu X et al (2020) A metabolic labeling method detects m6A transcriptome-wide at single base resolution. Nat Chem Biol 16:887–895. https://doi.org/10.1038/s41589-020-0526-9
Hu L et al (2022) (2022) m6A RNA modifications are measured at single-base resolution across the mammalian transcriptome. Nat Biotechnol 40:1210–1219. https://doi.org/10.1038/s41587-022-01243-z
Zaccara S, Ries RJ, Jaffrey SR (2019) Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell Biol 20:608–624. https://doi.org/10.1038/s41580-019-0168-5
Yang X, Triboulet R, Liu Q, Sendinc E, Gregory RI (2022) Exon junction complex shapes the m6A epitranscriptome. Nat Commun 13:7904. https://doi.org/10.1038/s41467-022-35643-1
Uzonyi A et al (2023) Exclusion of m6A from splice-site proximal regions by the exon junction complex dictates m6A topologies and mRNA stability. Mol Cell 83:237–251. https://doi.org/10.1016/j.molcel.2022.12.026
He PC et al (2023) Exon architecture controls mRNA m6A suppression and gene expression. Science 379:677–682. https://doi.org/10.1126/science.abj9090
Gerken T et al (2007) The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318:1469–1472. https://doi.org/10.1126/science.1151710
Zheng G et al (2013) ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49:18–29. https://doi.org/10.1016/j.molcel.2012.10.015
Wei J et al (2018) Differential m6A, m6Am, and m1A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol Cell 71:973-985.e5. https://doi.org/10.1016/j.molcel.2018.08.011
Song H, Wang Y, Wang R, Zhang X, Liu Y, Jia G, Chen PR (2020) SFPQ Is an FTO-binding protein that facilitates the demethylation substrate preference. Cell Chem Biol 27(3):283-291.e6. https://doi.org/10.1016/j.chembiol.2020.01.002
Boulias K, Greer EL (2023) Biological roles of adenine methylation in RNA. Nat Rev Genet 24:143–160. https://doi.org/10.1038/s41576-022-00534-0
Lasman L et al (2020) Context-dependent functional compensation between Ythdf m6A reader proteins. Genes Dev 34:1373–1391. https://doi.org/10.1101/gad.340695.120
Zaccara S, Jaffrey SR (2020) A unified model for the function of YTHDF proteins in regulating m6A-modified mRNA. Cell 181:1582–1595. https://doi.org/10.1016/j.cell.2020.05.012
Li L, Krasnykov K, Homolka D, Gos P, Mendel M, Fish RJ, Pandey RR, Pillai RS (2022) The XRN1-regulated RNA helicase activity of YTHDC2 ensures mouse fertility independently of m6A recognition. Mol Cell 82:1678–1690. https://doi.org/10.1016/j.molcel.2022.02.034
Barbieri I, Kouzarides T (2020) Role of RNA modifications in cancer. Nat Rev Cancer 20:303–322. https://doi.org/10.1038/s41568-020-0253-2
Quintás-Cardama A, Cortes J (2009) Molecular biology of bcr-abl1-positive chronic myeloid leukemia. Blood 113:1619–1630. https://doi.org/10.1182/blood-2008-03-144790
Barbieri I et al (2017) Promoter-bound METTL3 maintains myeloid leukaemia by m6A-dependent translation control. Nature 552:126–131. https://doi.org/10.1038/nature24678
Weng H et al (2018) METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m6A modification. Cell Stem Cell 22:191–205. https://doi.org/10.1016/j.stem.2017.11.016
Vu LP et al (2017) The N6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat Med 23:1369–1376. https://doi.org/10.1038/nm.4416
Ianniello Z et al (2021) New insight into the catalytic -dependent and -independent roles of METTL3 in sustaining aberrant translation in chronic myeloid leukemia. Cell Death Dis 12:870. https://doi.org/10.1038/s41419-021-04169-7
Yao F et al (2023) The m6A regulator KIAA1429 stabilizes RAB27B mRNA and promotes the progression of chronic myeloid leukemia and resistance to targeted therapy. Genes Dis 11:993–1008. https://doi.org/10.1016/j.gendis.2023.03.016
Chai J, Wang Q, Qiu Q, Han G, Chen Y, Li W, Zhang H (2023) YBX1 regulates the survival of chronic myeloid leukemia stem cells by modulating m6A-mediated YWHAZ stability. Cell Oncol 46:451–464. https://doi.org/10.1007/s13402-022-00762-w
Van Etten RA (2004) Mechanisms of transformation by the BCR-ABL oncogene: new perspectives in the post-imatinib era. Leuk Res 28(Suppl 1):S21–S28. https://doi.org/10.1016/j.leukres.2003.10.005
El-Tanani M, Nsairat H, Matalka II, Lee YF, Rizzo M, Aljabali AA, Mishra V, Mishra Y, Hromić-Jahjefendić A, Tambuwala MM (2024) The impact of the BCR-ABL oncogene in the pathology and treatment of chronic myeloid leukemia. Pathol Res Pract 254:155161. https://doi.org/10.1016/j.prp.2024.155161
Rinaldi I, Winston K (2023) Chronic myeloid leukemia, from pathophysiology to treatment-free remission: a narrative literature review. J Blood Med 14:261–277. https://doi.org/10.2147/JBM.S382090
Braun TP, Eide CA, Druker BJ (2020) Response and resistance to BCR-ABL1-targeted therapies. Cancer Cell 37:530–542. https://doi.org/10.1016/j.ccell.2020.03.006
Perrotti D, Trotta R, Calabretta B (2003) Altered mRNA translation: possible mechanism for CML disease progression. Cell Cycle 2(3):177–180
van Riggelen J, Yetil A, Felsher DW (2010) MYC as a regulator of ribosome biogenesis and protein synthesis. Nat Rev Cancer 10:301–309. https://doi.org/10.1038/nrc2819
Sorci M, Ianniello Z, Cruciani S, Larivera S, Ginistrelli LC, Capuano E, Marchioni M, Fazi F, Fatica A (2018) METTL3 regulates WTAP protein homeostasis. Cell Death Dis 9:796. https://doi.org/10.1038/s41419-018-0843-z
Yao FY, Zhao C, Zhong FM, Qin TY, Wen F, Li MY, Liu J, Huang B, Wang XZ (2021) m(6)A Modification of lncRNA NEAT1 regulates chronic myelocytic leukemia progression via miR-766-5p/CDKN1A axis. Front Oncol 11:679634. https://doi.org/10.3389/fonc.2021.679634
Gao C, Zhang J, Wang Q, Ren C (2016) Overexpression of lncRNA NEAT1 mitigates multidrug resistance by inhibiting ABCG2 in leukemia. Oncol Lett 12:1051–1507. https://doi.org/10.3892/ol.2016.4738
Lai X, Wei J, Gu XZ, Yao XM, Zhang DS, Li F, Sun YY (2021) Dysregulation of LINC00470 and METTL3 promotes chemoresistance and suppresses autophagy of chronic myelocytic leukaemia cells. J Cell Mol Med 25:4248–4259. https://doi.org/10.1111/jcmm.16478
Bolton-Gillespie E et al (2013) Genomic instability may originate from imatinib-refractory chronic myeloid leukemia stem cells. Blood 121:4175–4183. https://doi.org/10.1182/blood-2012-11-466938
Zhang C et al (2020) METTL3 and N6-methyladenosine promote homologous recombination-mediated repair of DSBs by modulating DNA-RNA hybrid accumulation. Mol Cell 79:425–442. https://doi.org/10.1016/j.molcel.2020.06.017
Xiang Y et al (2017) RNA m6A methylation regulates the ultraviolet-induced DNA damage response. Nature 543:573–576. https://doi.org/10.1038/nature21671
Tarsounas M, Sung P (2020) The antitumorigenic roles of BRCA1-BARD1 in DNA repair and replication. Nat Rev Mol Cell Biol 21:284–299. https://doi.org/10.1038/s41580-020-0218-z
Nieborowska-Skorska M et al (2019) Non-NAD-like PARP1 inhibitor enhanced synthetic lethal effect of NAD-like PARP inhibitors against BRCA1-deficient leukemia. Leuk Lymphoma 60:1098–1101. https://doi.org/10.1080/10428194.2018.1520988
Hiroki H, Ishii Y, Piao J, Namikawa Y, Masutani M, Honda H, Akahane K, Inukai T, Morio T, Takagi M (2023) Targeting Poly(ADP)ribose polymerase in BCR/ABL1-positive cells. Sci Rep 13:7588. https://doi.org/10.1038/s41598-023-33852-2
Yan F, Al-Kali A, Zhang Z, Liu J, Pang J, Zhao N, He C, Litzow MR, Liu S (2018) A dynamic N6-methyladenosine methylome regulates intrinsic and acquired resistance to tyrosine kinase inhibitors. Cell Res 28:1062–1076. https://doi.org/10.1038/s41422-018-0097-4
Han Z et al (2010) Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464:1205–1209
Houshmand M et al (2022) Dihydroorotate dehydrogenase inhibition reveals metabolic vulnerability in chronic myeloid leukemia. Cell Death Dis 13:576. https://doi.org/10.1038/s41419-022-05028-9
Fiorentino F, Menna M, Rotili D, Valente S, Mai A (2023) METTL3 from target validation to the first small-molecule inhibitors: a medicinal chemistry journey. J Med Chem 66(3):1654–1677. https://doi.org/10.1021/acs.jmedchem.2c01601
Cesaro B et al (2024) Enhancing sensitivity of triple-negative breast cancer to DNA-damaging therapy through chemical inhibition of the m6A methyltransferase METTL3. Cancer Commun 44:282–286. https://doi.org/10.1002/cac2.12509
Shen C et al (2020) RNA demethylase ALKBH5 selectively promotes tumorigenesis and cancer stem cell self-renewal in acute myeloid leukemia. Cell Stem Cell 27:64-80.e9. https://doi.org/10.1016/j.stem.2020.04.009
Paris J et al (2019) Targeting the RNA m6A reader YTHDF2 selectively compromises cancer stem cells in acute myeloid leukemia. Cell Stem Cell 25:137-148.e6. https://doi.org/10.1016/j.stem.2019.03.021
Cheng YM et al (2021) N6-Methyladenosine on mRNA facilitates a phase-separated nuclear body that suppresses myeloid leukemic differentiation. Cancer Cell 39:958–972. https://doi.org/10.1016/j.ccell.2021.04.017
Feng MD et al (2021) YBX1 is required for maintaining myeloid leukemia cell survival by regulating BCL2 stability in an m(6)A-dependent manner. Blood 138:71–85. https://doi.org/10.1182/blood.2020009676
Weng H et al (2022) The m6A reader IGF2BP2 regulates glutamine metabolism and represents a therapeutic target in acute myeloid leukemia. Cancer Cell 40:1566-1582.e10
Zhang N et al (2022) The m6A reader IGF2BP3 promotes acute myeloid leukemia progression by enhancing RCC2 stability. Exp Mol Med 54:194–205
Kusio-Kobialka M, Podszywalow-Bartnicka P, Peidis P, Glodkowska-Mrowka E, Wolanin K, Leszak G, Seferynska I, Stoklosa T, Koromilas AE, Piwocka K (2012) The PERK-eIF2α phosphorylation arm is a pro-survival pathway of BCR-ABL signaling and confers resistance to imatinib treatment in chronic myeloid leukemia cells. Cell Cycle 11:4069–4078. https://doi.org/10.4161/cc.22387
Xu Z, Wang H, Wei S, Wang Z, Ji G (2018) Inhibition of ER stress-related IRE1α/CREB/NLRP1 pathway promotes the apoptosis of human chronic myelogenous leukemia cell. Mol Immunol 101:377–385. https://doi.org/10.1016/j.molimm.2018.07.002
Wei J et al (2021) HRD1-mediated METTL14 degradation regulates m6A mRNA modification to suppress ER proteotoxic liver disease. Mol Cell 81:5052–5065. https://doi.org/10.1016/j.molcel.2021.10.028
Kong Y, Zhang Y, Cai Y, Li D, Yi B, Xu Q (2022) METTL3 mediates osteoblast apoptosis by regulating endoplasmic reticulum stress during LPS-induced inflammation. Cell Signal 95:110335. https://doi.org/10.1016/j.cellsig.2022.110335
Du QY, Huo FC, Du WQ, Sun XL, Jiang X, Zhang LS, Pei DS (2022) METTL3 potentiates progression of cervical cancer by suppressing ER stress via regulating m6A modification of TXNDC5 mRNA. Oncogene 41:4420–4432. https://doi.org/10.1038/s41388-022-02435-2
Einstein JM et al (2021) Inhibition of YTHDF2 triggers proteotoxic cell death in MYC-driven breast cancer. Mol Cell 81:3048–3064. https://doi.org/10.1016/j.molcel.2021.06.014
Lee Q et al (2023) Overexpression of VIRMA confers vulnerability to breast cancers via the m6A-dependent regulation of unfolded protein response. Cell Mol Life Sci 80:157. https://doi.org/10.1007/s00018-023-04799-4
Wang J et al (2024) XBP1s activates METTL3/METTL14 for ER-phagy and paclitaxel sensitivity regulation in breast cancer. Cancer Lett 4:216846. https://doi.org/10.1016/j.canlet.2024.216846
Acknowledgements
Not applicable.
Funding
This work was supported by NextGenerationEU-PNRR M4C2-Investment 1.4 - CN00000041.
Author information
Authors and Affiliations
Contributions
G.F.R and M.T. revised the manuscript and prepared the figures; A.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fernandez Rodriguez, G., Tarullo, M. & Fatica, A. N6-methyladenosine (m6A) RNA modification in chronic myeloid leukemia: unveiling a novel therapeutic target. Cell. Mol. Life Sci. 81, 326 (2024). https://doi.org/10.1007/s00018-024-05379-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05379-w