Abstract
Aims/hypothesis
Pancreatic polypeptide (PP) cells, which secrete PP (encoded by the Ppy gene), are a minor population of pancreatic endocrine cells. Although it has been reported that the loss of beta cell identity might be associated with beta-to-PP cell-fate conversion, at present, little is known regarding the characteristics of Ppy-lineage cells.
Methods
We used Ppy-Cre driver mice and a PP-specific monoclonal antibody to investigate the association between Ppy-lineage cells and beta cells. The molecular profiles of endocrine cells were investigated by single-cell transcriptome analysis and the glucose responsiveness of beta cells was assessed by Ca2+ imaging. Diabetic conditions were experimentally induced in mice by either streptozotocin or diphtheria toxin.
Results
Ppy-lineage cells were found to contribute to the four major types of endocrine cells, including beta cells. Ppy-lineage beta cells are a minor subpopulation, accounting for 12–15% of total beta cells, and are mostly (81.2%) localised at the islet periphery. Unbiased single-cell analysis with a Ppy-lineage tracer demonstrated that beta cells are composed of seven clusters, which are categorised into two groups (i.e. Ppy-lineage and non-Ppy-lineage beta cells). These subpopulations of beta cells demonstrated distinct characteristics regarding their functionality and gene expression profiles. Ppy-lineage beta cells had a reduced glucose-stimulated Ca2+ signalling response and were increased in number in experimental diabetes models.
Conclusions/interpretation
Our results indicate that an unexpected degree of beta cell heterogeneity is defined by Ppy gene activation, providing valuable insight into the homeostatic regulation of pancreatic islets and future therapeutic strategies against diabetes.
Data availability
The single-cell RNA sequence (scRNA-seq) analysis datasets generated in this study have been deposited in the Gene Expression Omnibus (GEO) under the accession number GSE166164 (www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE166164).
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
The islets of Langerhans consist of alpha, beta and delta cells and a fourth type of islet cell, namely, pancreatic polypeptide (PP) cells. PP cells are located at the periphery of the islets and secrete PP [1,2,3], encoded by the Ppy gene. PP is a member of the neuropeptide Y (NPY) family of peptides, which also includes peptide YY (PYY) and NPY, all reported to be involved in appetite regulation [4, 5]. However, the precise physiological functions of these peptides, including their roles in glucose homeostasis, remain poorly understood. A previous study demonstrated that Ppy-lineage cells were indispensable for the differentiation of a substantial fraction of endocrine cells by the diphtheria toxin (DT)-induced ablation of Ppy-lineage cells [6]. However, owing to the lack of specificity of the available PP antibody and the fidelity of the Ppy promoter used, the precise roles of Ppy-lineage cells have not yet been clarified.
Beta cell failure is the main hallmark of type 2 diabetes and it has been reported that inactivation of pancreatic and duodenal homeobox 1 (encoded by Pdx1) in mature beta cells results in the loss of beta cell identity and a beta-to-alpha cell fate conversion in vivo [7, 8]. Moreover, a reduction in the gene dose of Pdx1 by heterozygous deletion of its evolutionarily conserved enhancer region caused an increase not only in the number of alpha cells but also in the number of PP cells in the pancreatic islets of adult mice, together with insufficient beta cell development [9]. In another report, Rip-Cre-mediated deletion of Abcc8 in mice caused a sustained increase in intracellular Ca2+ concentrations, whereas beta cells underwent a fate switch to PP cells [10]. These findings suggest that unhealthy beta cells may shift not only towards alpha cells but also towards a PP cell fate.
Previous reports have highlighted the functional heterogeneity of beta cells. Many of these reports have particularly focused on the immature population of beta cells. For instance, they characterised immature beta cells located at the islet periphery, which are called ‘virgin beta cells’ [11], and low GLUT2-expressing immature beta cells with robust proliferative potential and resistance to streptozotocin (STZ)-induced cytotoxicity [12,13,14,15,16]. Other reports described Wnt-signal-regulated Fltp (also known as Cfap126)-positive beta cells [17] or NPY-positive immature beta cells [18]. The characterisation of beta cell subpopulations from the viewpoint of functionality and plasticity is especially important in the field of diabetes treatment for the further identification of beta cell sources with robust regenerative potential.
Methods
Animals
The study protocol was reviewed and approved by the Committee for Institutional Animal Care and Experimentation at Gunma University. All animals were housed in specific pathogen-free barrier facilities, maintained under a 12 h light/dark cycle, fed standard rodent food (CLEA Japan, Tokyo, Japan) and had access to water ad libitum. Male C57BL/6J mice (CLEA Japan, Tokyo, Japan) were used in all experiments, except for the single-cell RNA sequence (scRNA-seq) analysis and Ca2+ imaging experiments, in which male B6.Cg-Tg(Ins1-EGFP)1Hara/J (MIP-GFP) mice (stock number 006864; The Jackson Laboratory, Bar Harbor, ME, USA; https://www.jax.org/strain/006864) [19], which have a mixed background of CD-1 and C57BL/6J, were used. Ppy-Cre knockin mice were established previously (12-week-old male mice, weighing 23–26 g, were used), with the protein-coding region of Ppy in exon 2 being precisely replaced with that of NLS-Cre [20]. Ppy-DTA knockin mice, in which the diphtheria toxin A (DTA)-coding sequence was inserted in the Ppy locus, were also established by CRISPR/Cas9-mediated genome editing (12-week-old male mice, weighing 23–26 g, were used). Ins-TR1 (C57BL/6-human INS promoter-DTR TRECK) transgenic mice [21] have been described previously. B6.129X1-Gt(ROSA)26Sortm1(EYFP)Cos/J (Rosa26-YFP) mice (stock number 006148; The Jackson Laboratory; https://www.jax.org/strain/006148) and B6;129S6-Gt(ROSA)26Sortm14(CAG-tdTomato)Hze/J (Rosa26-tdTomato) mice (stock number 007908; The Jackson Laboratory; https://www.jax.org/strain/007908) were used as reporter mice. Randomisation and blinding were not carried out in this study. Mice outside of the indicated weight range were excluded from the experiments. We repeated all experiments at least twice, except for scRNA-seq analysis, which was validated by additional immunohistochemistry. To perform the IPGTT, 2 g/kg of glucose was injected intraperitoneally into mice and blood glucose levels were measured at the indicated times.
Immunohistochemistry and cell quantification
Harvested pancreases were divided into the head and tail and fixed in 4% vol./vol. paraformaldehyde at 4°C overnight. Then, pancreases were immersed in sucrose solution for 24 h prior to embedding in O.C.T compound (Sakura Finetek Japan, Osaka, Japan). Frozen pancreas blocks were sectioned at 14 μm thickness and immunostained. The following primary antibodies were used at the stated dilutions: guinea pig anti-insulin (INS; 1:100; DAKO, Glostrup, Denmark; catalogue no. IR002); mouse anti-glucagon (GCG; 1:1000; Abcam, Cambridge, UK; catalogue no. ab10988); rabbit anti-GCG (1:1000; Abcam; catalogue no. ab92517); rabbit anti-somatostatin (SST; 1:1000; Peninsula Laboratories, San Carlos, CA, USA; catalogue no. T-4103), mouse anti-PP (1:1000; IBL, Gunma, Japan; catalogue no. 23-2D3 [20]); chicken anti-green fluorescent protein (GFP; 1:1000; Abcam; catalogue no. ab13970); rabbit anti-chromogranin-A (1:100; Abcam; catalogue no. ab15160); rabbit anti-GLUT2 (1:200; Abcam; catalogue no. ab54460); rabbit anti-urocortin 3 (UCN3; 1:500; Phoenix Pharmaceuticals, Burlingame, CA, USA; catalogue no. H-019-29); rat anti-tetraspanin 8 (TSPAN8; 1:50; R&D Systems, Minneapolis, MN, USA; catalogue no. MAB6524); rabbit anti-folate receptor 1 (FOLR1; 1:100; Thermo Fisher Scientific, Waltham, MA, USA; catalogue no. PA5-42004); and rabbit anti-secreted phosphoprorein 1 (SPP1; 1:500; Sigma-Aldrich;, St Louis, MO, USA; catalogue no. AB10910). Primary antibodies were detected with the secondary antibodies listed in electronic supplementary material (ESM) Table 1, all used at 1:1000 dilution. DAPI (Dojindo, Kumamoto, Japan) was used for nuclear counterstaining. Fluorescence images were captured using an FV1000-D confocal microscope (Olympus, Tokyo, Japan). The total number of hormone- and yellow fluorescent protein (YFP)-positive cells in five to ten islets per pancreas from four mice per genotype were manually counted using Adobe Photoshop 2021 (Adobe, San Jose, CA, USA). Beta cell mass was measured as described previously [22].
Diabetes induction
Diabetes was induced either by the injection of STZ (Sigma-Aldrich) into Ppy-Cre;Rosa26-YFP mice or by the injection of DT (Sigma-Aldrich) into Ppy-Cre;Rosa26-YFP;Ins-TR1 mice. Blood glucose levels were monitored using Glutest mint (Sanwa Kagaku Kenkyusyo, Aichi, Japan). STZ was dissolved in citrate buffer (pH 4.5) prior to injection, and a single dose of 200 mg/kg was injected intraperitoneally into 6-week-old male mice (18–22 g). DT was dissolved in PBS prior to injection and a single dose of 100 ng/kg was injected intraperitoneally into 6-week-old male Ppy-Cre;Rosa26-YFP;Ins-TR1 mice. Mice were killed at 7 days or 5 weeks after the injection.
Ca2+ imaging
Islets were isolated from the pancreases of 8- to 10-week-old MIP-GFP;Ppy-Cre;Rosa26-tdTomato mice (22–25 g), as described previously [23]. Isolated beta cells were seeded onto poly-l-lysine-coated glass-bottomed dishes (MatTek Corporation, Ashland, MA, USA) 24 h before observation. Cells were washed with modified KRB buffer. Cells were loaded with 2 μmol/l Fura 2-AM (Dojindo, Kumamoto, Japan) and 0.01% vol./vol. Cremophor EL (Sigma-Aldrich) in a humidified incubator (95% air and 5% CO2 at 37°C) for 30 min. After washing with modified KRB, the cells were visualised using an Olympus UPlanAPO 10× water objective lens (Olympus). To obtain fluorescence images, the AQUACOSMOS/ASHURA 3CCD-based fluorescence energy transfer imaging system (Hamamatsu Photonics, Tokyo, Japan) was used. The ratio of 340/380 nm fluorescence was calculated and values were normalised to F0.
Statistical analysis (other than for scRNA-seq analysis data)
Statistical analysis was performed using GraphPad Prism 9 software (GraphPad Software, San Diego, CA, USA). Data were expressed as means ± SEMs. Statistical comparisons between two groups were performed by the two-tailed Student’s t test, and one-way ANOVA followed by the Bonferroni post hoc test was used for comparisons between groups. A p value of less than 0.05 was considered to indicate a statistically significant difference between groups.
scRNA-seq analysis
Islet cells were isolated from an 8-week-old male MIP-GFP;Ppy-Cre;Rosa26-tdTomato mouse. Cell quality was checked under a microscope before loading cells onto the chip, and cell viability was confirmed to be 84%. Cells that met one or more of the following three criteria, including doublets, were excluded from further analysis: (1) cells with 200 or less, or 8000 or more detected genes per cell; (2) cells with 80,000 or more unique molecular identifiers (UMIs) per cell; and (3) cells with 5% or more mitochondrial gene UMIs / the number of total gene UMIs per cell. Isolated single cells were loaded for scRNA-seq analysis using the Chromium System (10x Genomics, Pleasanton, CA, USA), following the manufacturer’s protocol of Single Cell 3′ Library kit v3.1. RNA libraries were sequenced on a Nova Seq6000 platform (Illumina, San Diego, CA, USA) with the following sequencing parameters: 28 bp read 1; 8 bp index 1; 91 bp read 2.
Analysis of scRNA-seq data
Sequencing data were aligned to the mouse genome, Genecode release M25/GRCm38.p6 with GFP and tdTomato sequences, and UMI-collapsed with the Cell Ranger (v3.1.0) pipeline (10x Genomics).
For pre-processing of data, the Seurat (v3.1) R package for quality control filtering was used. Genes that were detected in at least three cells, and cells with more than 200 detected genes were selected. In addition, only cells with less than 8000 detected genes, less than 5% of mitochondrial genes, and less than 80,000 detected UMIs were retained.
Normalisation was performed using the global-scaling normalisation method, which normalises gene expression measurements for each cell by total gene expression, and then multiplies them by 10,000, and finally log-transforms the results. The variable genes were identified using the ‘vst’ method. Data were scaled to regress out the cell cycle score, the number of UMIs and the percentage of mitochondrial gene expression.
Based on the extracted variable genes, dimensionality reduction was performed by principal component analysis. A total of 50 dimensions were used for whole-cell analysis and 75 dimensions for beta cell analysis. Cell clustering was performed according to the Shared Nearest Neighbors method and was visualised in two-dimension by uniform manifold approximation and projection (UMAP).
Differentially expressed genes (DEGs) were identified by comparing the expression levels of the cluster cells with all other cells. For the analysis of statistical significance, the likelihood ratio test was performed by modelling as a two-part generalised regression model using model-based analysis of single-cell transcriptomics [24], and the Bonferroni method was used for multiple comparison adjustment. The mean fold change of expression was calculated by the following formula:
Results
Ppy-lineage cells contribute to the four major types of endocrine cells
The findings that the loss of beta cell identity may be associated with beta-to-PP cell-fate conversion led us to investigate the association between Ppy-lineage cells and beta cells. For this purpose, we analysed the fate of Ppy-expressing cells by developing a novel Ppy-knockin Cre driver, in which the coding region of the Ppy locus was replaced by that of Cre recombinase (Fig. 1a), and a PP-specific antibody (23-2D3) with no cross-reactivity to PYY or NPY [20]. In 12-week-old male Ppy-Cre;Rosa26-YFP mice, about 90% of the cells positive for PP encoded by the Ppy gene were positive for YFP, indicating high fidelity of this reporter (Fig. 1b). Notably, there were some YFP+ cells that were negative for PP (Fig. 1b, green arrowheads). Immunostaining analysis demonstrated that Ppy-lineage YFP+ cells contained cells that were costained for INS, GCG, SST or PP, indicating that Ppy-lineage cells contribute to all the four major types of endocrine cells (Fig. 1c–f). Some YFP+ GCG+ cells and YFP+ SST+ cells were also positive for PP, but there were very few YFP+ INS+ PP+ triple-positive cells (ESM Fig. 1a–c). The frequency of Ppy-lineage beta cells among total islet cells was similar between the head (14.9%) and tail (12.4%) of the pancreas, whereas the frequency of Ppy-lineage alpha (GCG+), delta (SST+) and PP cells differed between the two regions of the pancreas (Fig. 1g). We counted at least 1400 INS+ cells and 230 SST+ cells in the pancreatic head and tail of each mouse, and the number of each cell type was comparable between the two regions in each mouse. Importantly, we counted 365–489 GCG+ cells for each pancreatic tail but only 96–223 for each pancreatic head, and 258–436 PP+ cells for each pancreatic head but only 7–53 for each pancreatic tail. This difference in the population of endocrine cells between the head and tail is consistent with a previous report [25]. Ppy-lineage beta cells were mostly located at the peripheral region of the islets (81.2%), with a few cells in the central core of the islets (18.8%) (Fig. 1c and ESM Fig. 1a). As PP cells are much more abundant in the head of the pancreas, which is supposed to reflect the dynamics of Ppy-lineage and PP cells more accurately than the tail of the pancreas, we used this region for all subsequent analyses, unless otherwise stated.
Analysis of Ppy-lineage cells using novel Ppy-Cre mice. (a) Ppy-Cre mice were crossed with Rosa26-YFP mice to generate Ppy-Cre;Rosa26-YFP mice. (b) YFP+ PP+ cells (yellow arrowheads) and YFP single-positive cells (green arrowheads) in the head of the pancreas of adult Ppy-Cre;Rosa26-YFP mice. Representative images of n = 4 mice. Scale bar, 50 μm. (c–e) YFP+ INS+ cells (c), YFP+ GCG+ cells (d) and YFP+ SST+ (e) double-positive cells in the head of the pancreas of adult Ppy-Cre;Rosa26-YFP mice (yellow arrowheads for all). Scale bars, 50 μm. (f) The ratio of YFP+ INS+/GCG+/SST+/PP+ (YFP+ hormone+) cells to total YFP+ cells in the pancreas of adult Ppy-Cre;Rosa26-YFP mice (n = 4). (g) The ratio of YFP+ hormone+ cells to each type of hormone+ cell in the pancreas of adult Ppy-Cre;Rosa26-YFP mice (n = 4). Data are shown as the mean ± SEM. *p < 0.05, **p < 0.01, (two-tailed Student’s t test)
Ppy-DTA mice, which lack Ppy-lineage endocrine cells, demonstrate normal glucose tolerance
We next investigated the physiological roles of Ppy-lineage cells and PP cells using 12-week-old male Ppy-DTA mice (Fig. 2a). In Ppy-DTA mice, Ppy-lineage cells are designed to express DTA, and these cells eventually die owing to the accumulation of DTA. The composition of islet cells in Ppy-DTA mice was significantly different from that of wild-type (WT) mice, particularly regarding non-beta endocrine cells. PP cells were almost completely absent in 12-week-old Ppy-DTA mice (Fig. 2b), and the ratio of alpha and delta cells to total islet cells was significantly lower in the head of the pancreas of Ppy-DTA mice than in WT mice (Fig. 2c–h). These data are consistent with the results that a substantial number of alpha and delta cells have a history of Ppy activation (Fig. 1g). Although beta cell mass was also significantly smaller in the head of the pancreas of Ppy-DTA mice compared with WT mice (Fig. 2i), Ppy-DTA mice showed normal glucose tolerance (Fig. 2j). This indicates that Ppy-lineage cells, including Ppy-lineage beta cells and PP cells, are dispensable for maintaining glucose tolerance at least in physiological conditions.
Analysis of Ppy-DTA mice. (a) Diagram of genotype of Ppy-DTA knockin mice. (b) Immunohistochemical analysis revealed almost complete deletion of PP cells in the head of the pancreas of adult Ppy-DTA mice. Representative images of n = 4 mice. CHGA, chromogranin-A. Scale bar, 50 μm. (c, d) Immunohistochemical analysis of INS+ and GCG+ cells in the head of the pancreas of adult WT mice (c) and Ppy-DTA mice (d). Representative images of n = 4 mice. Scale bars, 50 μm. (e) Ratio of GCG+ cells to total islet cells in adult WT vs Ppy-DTA mice (n = 4). (f, g) Immunohistochemical analysis of INS+ and SST+ cells in the head of the pancreas of adult WT mice (f) and Ppy-DTA mice (g). Representative images of n = 4 mice. Scale bars, 50 μm. (h) Ratio of SST+ cells to total islet cells in adult WT vs Ppy-DTA mice (n = 4). (i) Beta cell mass in the head of the pancreas of Ppy-DTA mice vs WT mice (n = 6). (j) Results of IPGTT performed on adult WT mice and Ppy-DTA mice. Data are shown as the mean ± SEM. *p < 0.05, **p < 0.01, (two-tailed Student’s t test)
scRNA-seq analysis clarified the heterogeneity of Ppy expression in beta cells
To investigate the molecular profiles of Ppy-lineage beta cells, we performed scRNA-seq analysis of islets (both the pancreatic head and tail) from an 8-week-old MIP-GFP;Ppy-Cre;Rosa26-tdTomato mouse. We profiled a total of 3949 cells using scRNA-seq analysis, and they were divided into 16 clusters (Fig. 3a–f). Each cluster was mapped to endocrine cells, exocrine cells or other known cell types. We also identified the recently described Procr+ progenitor cells, reproducing a recent study [26]. The PP cell population demonstrated the expression of unique signature genes, including the previously reported Ppy, Pyy, Tspan8 and Folr1 genes (ESM Fig. 2a, ESM Table 2) [26]. tdTomato was also detected as a marker gene for PP cells. Violin plots of tdTomato showed similar profiles to Ppy (Fig. 3e,f), corresponding to the Ppy-Cre-mediated activation of the tdTomato gene. Signals of tdTomato were detected not only in PP cells but also in alpha and delta cells (Fig. 3f). Therefore, the above data are consistent with the results that a significant fraction of alpha and delta cells have a history of Ppy activation (Fig. 1g).
Single-cell transcriptome analysis of Ppy-lineage beta cells. (a) UMAP visualisation (in two dimensions [UMAP_1 and UMAP_2] of 3949 islet cells from an 8-week-old male MIP-GFP;Ppy-Cre;Rosa26-tdTomato mouse. Cell counts of each cluster are presented in brackets. (b–f) Feature plot and violin plot of Ins2 (b), Gcg (c), Sst (d), Ppy (e) and tdTomato (f) mRNA expression in the various islet cell clusters. Relative expression index (arbitrary units) is shown on y-axes. (g) Reclustering of beta cells by UMAP visualisation identified seven subclusters (no. 0 to no. 6). (h–l) Feature plot and violin plot of Ins2 (h), Gcg (i), Sst (j), Ppy (k) and tdTomato (l) mRNA expression in the various beta cell clusters shown in (g). Relative expression index (arbitrary units) is shown on y-axes. (m, n) GO analysis based on DEGs between Ppy-lineage and non-Ppy-lineage beta cells was performed using ShinyGO v0.61 Gene Ontology Enrichment Analysis. GO biological process output to comma-separated values files were saved, and all pathways with an adjusted p value of less than 0.01 were selected for the analysis. Commonly recurring and highly ranked GO pathways, which were highly enriched in genes derived from the differential gene expression analysis were selected. p values were normalised using the following formula: –log10(p value). GO pathways upregulated (m) and downregulated (n) in Ppy-lineage beta cells are shown. P&S, Procr+ and stellate
To gain further insight into the heterogeneity of beta cells, transcriptomic characterisation of INS+ beta cells containing clusters beta-1, -2 and -3 was performed (see Fig. 3a). Refined clustering of beta cells demonstrated the presence of seven clusters (Fig. 3g–l, ESM Fig. 2b). In total, 24.6% of beta cells expressed Ppy mRNA and, importantly, such cells were enriched in clusters 4, 6 and 3, rather than being scattered among all beta cells, which is consistent with the enrichment of tdTomato mRNA (Fig. 3k,l). In cluster 3, however, the enrichment of Ppy and tdTomato mRNA was highly heterogeneous, in contrast to the homogeneous and clear enrichment of Ppy and tdTomato mRNA in clusters 4 and 6 (violin plots in Fig. 3k,l). For this reason, clusters 4 and 6 were annotated as Ppy-lineage beta cells and clusters 0, 1, 2 and 5 were annotated as non-Ppy-lineage beta cells in this study, and they were subjected to further analysis, including DEG and Gene Ontology (GO) analyses.
DEG analysis between Ppy-lineage and non-Ppy-lineage beta cells (ESM Table 3) demonstrated that Ppy-lineage beta cells show an upregulation of some of the signature genes for PP cells, including Tspan8, Folr1, Spp1 and Pyy in addition to Ppy, indicating that these beta cells share molecular characteristics with canonical PP cells. Moreover, these beta cells showed the downregulation of key genes for beta cell maturation and INS secretion (e.g. Slc2a2, Ucn3, Ins1, Ins2, MafA, Nkx6.1 [also known as Nkx6-1], Neurod1, G6pc2, Sytl4 and Ero1lb [also known as Ero1b]; ESM Table 3). GO analysis of the DEGs identified the upregulation of several cell pathways, including the cell proliferation pathway, and the downregulation of the transport and INS secretion pathway in Ppy-lineage beta cells compared with non-Ppy-lineage beta cells (Fig. 3m,n).
TSPAN8 is a marker for Ppy-lineage beta cells
Next, to validate the molecular profiles of Ppy-lineage beta cells shown by scRNA-seq analysis at the protein level, we performed immunohistochemical analysis of some of the upregulated and downregulated markers extracted from DEG analysis and compared them in Ppy-lineage and non-Ppy-lineage beta cells. Among the markers analysed, we found that TSPAN8 was specifically expressed in Ppy-lineage beta cells. TSPAN8 is a member of the tetraspanin transmembrane protein family and has been reported as a PP-cell signature gene [26]. Approximately 40% of Ppy-lineage beta cells expressed TSPAN8, and, importantly, no expression was observed in non-Ppy-lineage beta cells (Fig. 4a).
Ppy-lineage beta cells show higher expression of TSPAN8 and lower expression of GLUT2 and UCN3 than non-Ppy-lineage beta cells. (a) Immunohistochemical analysis of YFP+, TSPAN8+ and INS+ cells in the head of the pancreas of Ppy-Cre;Rosa26-YFP mice. Enlarged images of the inset are shown, with YFP+ TSPAN8+ INS+ cells indicated by arrowheads. Scale bar, 50 μm. (b) Immunohistochemical analysis of YFP+, GLUT2+ and INS+ cells in the head of the pancreas of Ppy-Cre;Rosa26-YFP mice. Enlarged images of the inset are shown, with YFP+ GLUT2− INS+ cells indicated by the dotted area. Scale bar, 50 μm. (c) Immunohistochemical analysis of YFP+, UCN3+ and INS+ cells in the head of the pancreas of Ppy-Cre;Rosa26-YFP mice. Enlarged images of the inset are shown, with YFP+ UCN3− INS+ cells indicated by the dotted area. Representative images of n = 4 mice. Scale bar, 50 μm
As a negative marker, we found a low expression of GLUT2/Slc2a2 in Ppy-lineage beta cells at the islet periphery (Fig. 4b). We also observed a lower expression of UCN3, a beta cell maturation marker, in Ppy-lineage beta cells compared with non-Ppy-lineage beta cells (Fig. 4c). These immunohistochemical results further confirm the PP-cell-like molecular characteristics and immaturity of Ppy-lineage beta cells indicated by scRNA-seq analysis.
Ppy-lineage beta cells demonstrate reduced glucose-stimulated Ca2+ responses
Our scRNA-seq analyses demonstrated lower expression of a group of genes implicated in beta cell maturation and INS secretion in Ppy-lineage beta cells compared with non-Ppy-lineage beta cells (ESM Table 3). We hence postulated that Ppy-lineage beta cells may demonstrate decreased glucose responsiveness. To assess this, islets from 8- to 10-week-old MIP-GFP;Ppy-Cre;Rosa26-tdTomato mice were dispersed into single cells and then the increase in intracellular Ca2+ concentrations ([Ca2+]i), which is the final trigger of INS exocytosis, was measured in isolated non-Ppy-lineage (GFP+) and Ppy-lineage (GFP+ tdTomato+) beta cells during exposure to basal (2.8 mmol/l) and high (25 mmol/l) glucose concentrations (Fig. 5a). Single-cell Ca2+ imaging demonstrated that 25 mmol/l glucose increased [Ca2+]i in both beta cell types (Fig. 5b). The peak Ca2+ response was smaller in Ppy-lineage beta cells than in non-Ppy-lineage beta cells (Fig. 5c,d). Accordingly, the AUC of [Ca2+]i was significantly decreased in Ppy-lineage beta cells. A similar increase in [Ca2+]i was observed in both beta cell types when cells were depolarised by a 40 mmol/l K+ stimulation (Fig. 5e). Taken together, the results of our physiological studies demonstrated that Ppy-lineage beta cells show a smaller glucose-stimulated Ca2+ response, supporting our molecular findings.
Comparison of glucose-stimulated Ca2+ influx between Ppy-lineage and non-Ppy-lineage beta cells. (a) Diagram of the method of identification of Ppy-lineage and non-Ppy-lineage beta cells using two reporters, GFP and tdTomato. (b) Ca2+ response to high-glucose stimulation in non-Ppy-lineage beta cells (green cells; n = 223) and Ppy-lineage beta cells (yellow cells; n = 84) from a total of n = 10 mice (n = 5 mice per group). Fura2-ratio is the ratio of 340/380 nm fluorescence. (c, d) Representative time course of Ca2+ responses in green cells (n = 111 cells) (c) and yellow cells (n = 44 cells) (d) from n = 1 mouse at glucose concentrations of 2.8 mmol/l (G2.8) and 25 mmol/l (G25). (e) Ca2+ response to 40 mmol/l K+ stimulation in non-Ppy-lineage beta cells (green cells; n = 62) and Ppy-lineage beta cells (yellow cells; n = 49) from a total of n = 8 mice (n = 4 mice per group). Data are shown as the mean ± SEM. **p < 0.01, (two-tailed Student’s t test)
Ppy-lineage beta cells with low GLUT2 expression become dominant after STZ administration to mice
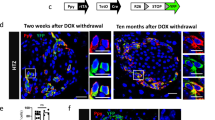
To characterise the behaviour of Ppy-lineage beta cells in diabetes, a single dose of 200 mg/kg STZ was administered to 6-week-old male Ppy-Cre;Rosa26-YFP mice. Mice were killed after 7 days and their pancreatic tissue was analysed. Mice became hyperglycaemic shortly after STZ injection and Ppy-lineage beta cells became relatively dominant (Fig. 6a,b), accounting for 48.8% of the remaining beta cells. TSPAN8+ INS+ beta cells also became dominant (ESM Fig. 3a–c), consistent with the results shown in Fig. 4a. However, the induction of hyperglycaemia by the administration of DT to 6-week-old male Ppy-Cre;Rosa26-YFP;Ins-TR1 mice, in which beta cells can be ablated by the administration of DT acting on the DT receptors that they express, did not change the proportion of Ppy-lineage beta cells among the remaining beta cells 7 days after DT administration (Fig. 6c,d). Considering that STZ is transported into beta cells via GLUT2, the increased population of Ppy-lineage beta cells in the islets of mice with STZ-induced diabetes is thought to correspond to the Ppy-lineage beta cells with low expression of Slc2a2/GLUT2 observed by scRNA-seq analysis and immunohistochemistry. Intriguingly, Ppy-lineage beta cells became dominant also in Ppy-Cre;Rosa26-YFP;Ins-TR1 mice 5 weeks after DT administration (Fig. 6e,f), suggesting that Ppy-lineage beta cells are more resistant to prolonged hyperglycaemia than non-Ppy-lineage beta cells.
Characteristics of Ppy-lineage beta cells under hyperglycaemic conditions. (a) Immunohistochemical analysis of YFP+ INS+ cells (arrowheads) in the head of the pancreas of Ppy-Cre;Rosa26-YFP mice 7 days after 200 mg/kg STZ injection. Scale bar, 50 μm. (b) Ratio of YFP+ INS+ cells to total INS+ cells in the head of the pancreas of Ppy-Cre;Rosa26-YFP mice 7 days after 200 mg/kg STZ injection compared with control mice (n = 4). Control mice were treated with citrate buffer. (c) Immunohistochemical analysis of YFP+ INS+ cells (arrowheads) in the head of the pancreas of Ppy-Cre;Rosa26-YFP;Ins-TR1 mice 7 days after 100 ng/kg DT injection. Scale bar, 50 μm. (d) Ratio of YFP+ INS+ cells to total INS+ cells in the head of the pancreas of Ppy-Cre;Rosa26-YFP;Ins-TR1 mice 7 days after 100 ng/kg DT injection compared with control mice (n = 4). Control mice were treated with PBS. (e) Immunohistochemical analysis of YFP+ INS+ cells (arrowheads) in the head of the pancreas of Ppy-Cre;Rosa26-YFP;Ins-TR1 mice 5 weeks after 100 ng/kg DT injection. Scale bar, 50 μm. (f) Ratio of YFP+ INS+ cells to total INS+ cells in the head of the pancreas of Ppy-Cre;Rosa26-YFP;Ins-TR1 mice 5 weeks after 100 ng/kg DT injection compared with control mice (n = 4). Control mice were treated with PBS. Ctrl, control mice. Data are shown as the mean ± SEM. **p < 0.01 (two-tailed Student’s t test)
Discussion
In this study, lineage tracing and scRNA-seq analysis using lineage tracers were performed to characterise PP cells and Ppy-lineage cells. YFP and tdTomato were detected in alpha and delta cells, in addition to beta cells (Figs 1c–e, 3f), suggesting that a substantial fraction of endocrine cells have a history of Ppy gene activation. Indeed, scRNA-seq analysis of adult WT islets showed that a subpopulation of alpha, beta and delta cells express Ppy mRNA, and the coexpression of PP and other endocrine hormones in these cells was confirmed at the protein level (ESM Fig. 1a–c). The substantial reduction of alpha and delta cell numbers in Ppy-DTA mice (Fig. 2c–h) also supports the idea that activation of the Ppy gene occurs in these endocrine cell types.
Ppy-lineage cells can contribute to all four major types of endocrine cells. The results of cell-lineage tracing may simply reflect the heterogeneity of beta cells, in which a subpopulation of beta cells express Ppy. Beta cell heterogeneity has attracted much attention and provides additional insight into the homeostatic regulation of islet function in the progression and treatment of diabetes. ‘Virgin beta cells’ [11] showed similar characteristics to the Ppy-lineage beta cells, with low levels of GLUT2 expression and their localisation at the islet periphery. However, these beta cells were shown to be induced via alpha-to-beta transdifferentiation, as assessed by Gcg-Cre-mediated lineage tracing. Therefore, ‘virgin beta cells’ were considered as a neogenic niche, a continuous supply of cells provided from alpha cells. There are reports of other beta cell subpopulations showing low levels of GLUT2 expression, robust proliferative capacity and resistance to STZ-induced cytotoxicity [12,13,14,15,16], possibly sharing the characteristics of Ppy-lineage beta cells shown in the present study (i.e. immaturity and impaired function confirmed by the low expression of GLUT2 and UCN3 and a low glucose-stimulated Ca2+ response). However, these previous studies lacked analysis using a molecular marker that identifies specific types of beta cells, and we propose that Ppy gene expression is a candidate for such a marker.
Additional analysis of the pathophysiological characteristics of Ppy-lineage beta cells in diabetic conditions using Ppy-CreERT2 knockin mice, which we generated together with Ppy-Cre knockin mice (data not shown), is expected to provide further useful information. However, as the pulse-and-chase labelling efficiency of these mice (particularly in their Ppy-lineage beta cells) is low, it is difficult to analyse the dynamics of Ppy-lineage beta cells within a specific time window in these mice. This insufficient labelling efficiency of Ppy-lineage beta cells might be owing to suppressed activity of the Ppy promoter during their differentiation into beta cells. The establishment of tools and techniques for detecting low levels of Ppy gene transcription in endocrine cells within a specific time window will resolve this limitation of our present study.
In this study, we found that 40% of the Ppy-lineage beta cells express TSPAN8 (Fig. 4a), a member of the tetraspanin transmembrane protein family that is mainly expressed in the gastrointestinal tract in both mice and humans [27]. Tspan8 has been reported as a PP-cell signature gene [26] that is highly expressed in human pancreatic ductal progenitors [28]. Other PP-cell signature markers, such as PP (ESM Fig. 1a), FOLR1 and SPP1 (ESM Fig. 4a,b), were rarely merged with Ppy-lineage beta cells [26, 29]. Some recent reports showed that TSPAN8 regulates cell proliferation, invasion and metastasis in various types of tumours, including pancreatic adenocarcinoma [30]. Further investigation of the physiological role of TSPAN8 in Ppy-lineage beta cells will also clarify the physiological role of Ppy-lineage beta cells, particularly those associated with their proliferative characteristics. Regarding cell proliferation, DEG analysis identified the upregulation of cell proliferation markers in Ppy-lineage beta cells (e.g. Jun, Junb, Fyn, Fgfr1, Pdgfb and Reg1). Of these enriched genes, Pdgfb and Reg1 are of particular interest, as they have been reported to regulate beta cell proliferation during ageing and in some models of diabetes [31,32,33,34]. The upregulation of these genes associated with beta cell proliferation indicates that Ppy-lineage beta cells are a promising therapeutic target in diabetes.
In summary, we found an unexpected degree of beta cell heterogeneity while investigating the characteristics of Ppy-lineage cells. High-resolution single-cell transcriptome analysis demonstrated that this subpopulation of beta cells shows unique functional characteristics and gene expression profile. We can speculate that Ppy-lineage beta cells with low levels of GLUT2 and UCN3 expression may be generated by the pancreas to survive conditions of metabolic stress under hyperglycaemia, at the expense of glucose-induced INS secretion. Identification of this unique subpopulation of beta cells is expected to provide valuable insight into the homeostatic regulation of islet function and contribute towards the development of novel therapeutic strategies to cure diabetes.
Data availability
The scRNA-seq analysis datasets generated in this study have been deposited in the Gene Expression Omnibus (GEO) under the accession number GSE166164 (www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE166164). Other datasets are available from the corresponding author upon reasonable request.
Abbreviations
- [Ca2+]i:
-
Intracellular Ca2+ concentration
- DEG:
-
Differentially expressed gene
- DT:
-
Diphtheria toxin
- DTA:
-
Diphtheria toxin A
- FOLR1:
-
Folate receptor 1
- GCG:
-
Glucagon
- GFP:
-
Green fluorescent protein
- GO:
-
Gene ontology
- INS:
-
Insulin
- NPY:
-
Neuropeptide Y
- PP:
-
Pancreatic polypeptide
- PYY:
-
Peptide YY
- scRNA-seq:
-
Single-cell RNA sequence
- SPP1:
-
Secreted phosphoprotein 1
- SST:
-
Somatostatin
- STZ:
-
Streptozotocin
- TSPAN8:
-
Tetraspanin 8
- UCN3:
-
Urocortin 3
- UMAP:
-
Uniform manifold approximation and projection
- UMI:
-
Unique molecular identifiers
- WT:
-
Wild-type
- YFP:
-
Yellow fluorescent protein
References
Larsson LI, Sundler F, Hakanson R (1976) Pancreatic polypeptide - a postulated new hormone: identification of its cellular storage site by light and electron microscopic immunocytochemistry. Diabetologia 12(3):211–226. https://doi.org/10.1007/bf00422088
Ekblad E, Sundler F (2002) Distribution of pancreatic polypeptide and peptide YY. Peptides 23(2):251–261. https://doi.org/10.1016/s0196-9781(01)00601-5
Kloppel G, Anlauf M, Perren A, Sipos B (2014) Hyperplasia to neoplasia sequence of duodenal and pancreatic neuroendocrine diseases and pseudohyperplasia of the PP-cells in the pancreas. Endocr Pathol 25(2):181–185. https://doi.org/10.1007/s12022-014-9317-8
Upchurch BH, Aponte GW, Leiter AB (1994) Expression of peptide YY in all four islet cell types in the developing mouse pancreas suggests a common peptide YY-producing progenitor. Development 120(2):245–252. https://doi.org/10.1242/dev.120.2.245
Teitelman G, Alpert S, Polak JM, Martinez A, Hanahan D (1993) Precursor cells of mouse endocrine pancreas coexpress insulin, glucagon and the neuronal proteins tyrosine hydroxylase and neuropeptide Y, but not pancreatic polypeptide. Development 118(4):1031–1039. https://doi.org/10.1242/dev.118.4.1031
Herrera PL, Huarte J, Zufferey R et al (1994) Ablation of islet endocrine cells by targeted expression of hormone-promoter-driven toxigenes. Proc Natl Acad Sci U S A 91(26):12999–13003. https://doi.org/10.1073/pnas.91.26.12999
Ahlgren U, Jonsson J, Jonsson L, Simu K, Edlund H (1998) beta-cell-specific inactivation of the mouse Ipf1/Pdx1 gene results in loss of the beta-cell phenotype and maturity onset diabetes. Genes Dev 12(12):1763–1768. https://doi.org/10.1101/gad.12.12.1763
Gao T, McKenna B, Li C et al (2014) Pdx1 maintains beta cell identity and function by repressing an alpha cell program. Cell Metab 19(2):259–271. https://doi.org/10.1016/j.cmet.2013.12.002
Fujitani Y, Fujitani S, Boyer DF et al (2006) Targeted deletion of a cis-regulatory region reveals differential gene dosage requirements for Pdx1 in foregut organ differentiation and pancreas formation. Gened Dev 20(2):253–266. https://doi.org/10.1101/gad.1360106
Stancill JS, Cartailler JP, Clayton HW et al (2017) Chronic beta-cell depolarization impairs beta-cell identity by disrupting a network of Ca(2+)-regulated genes. Diabetes 66(8):2175–2187. https://doi.org/10.2337/db16-1355
van der Meulen T, Mawla AM, DiGruccio MR et al (2017) Virgin beta cells persist throughout life at a neogenic niche within pancreatic islets. Cell Metab 25(4):911–926.e916. https://doi.org/10.1016/j.cmet.2017.03.017
Smukler SR, Arntfield ME, Razavi R et al (2011) The adult mouse and human pancreas contain rare multipotent stem cells that express insulin. Cell Stem Cell 8(3):281–293. https://doi.org/10.1016/j.stem.2011.01.015
Razavi R, Najafabadi HS, Abdullah S, Smukler S, Arntfield M, van der Kooy D (2015) Diabetes enhances the proliferation of adult pancreatic multipotent progenitor cells and biases their differentiation to more beta-cell production. Diabetes 64(4):1311–1323. https://doi.org/10.2337/db14-0070
Beamish CA, Strutt BJ, Arany EJ, Hill DJ (2016) Insulin-positive, Glut2-low cells present within mouse pancreas exhibit lineage plasticity and are enriched within extra-islet endocrine cell clusters. Islets 8(3):65–82. https://doi.org/10.1080/19382014.2016.1162367
Beamish CA, Zhang L, Szlapinski SK, Strutt BJ, Hill DJ (2017) An increase in immature beta-cells lacking Glut2 precedes the expansion of beta-cell mass in the pregnant mouse. PLoS One 12(7):e0182256. https://doi.org/10.1371/journal.pone.0182256
Feng Y, Qiu WL, Yu XX et al (2020) Characterizing pancreatic β-cell heterogeneity in the streptozotocin model by single-cell transcriptomic analysis. Mol Metab 37:100982. https://doi.org/10.1016/j.molmet.2020.100982
Bader E, Migliorini A, Gegg M et al (2016) Identification of proliferative and mature beta-cells in the islets of Langerhans. Nature 535(7612):430–434. https://doi.org/10.1038/nature18624
Rodnoi P, Rajkumar M, Moin ASM, Georgia SK, Butler AE, Dhawan S (2017) Neuropeptide Y expression marks partially differentiated beta cells in mice and humans. JCI Insight 2(12):e94005. https://doi.org/10.1172/jci.insight.94005
Hara M, Wang X, Kawamura T et al (2003) Transgenic mice with green fluorescent protein-labeled pancreatic beta -cells. Am J Physiol Endocrinol Metab 284(1):E177–E183. https://doi.org/10.1152/ajpendo.00321.2002
Hara A, Nakagawa Y, Nakao K et al (2019) Development of monoclonal mouse antibodies that specifically recognize pancreatic polypeptide. Endocr J 66(5):459–468. https://doi.org/10.1507/endocrj.EJ18-0441
Matsuoka K, Saito M, Shibata K et al (2013) Generation of mouse models for type 1 diabetes by selective depletion of pancreatic beta cells using toxin receptor-mediated cell knockout. Biochem Biophys Res Commun 436(3):400–405. https://doi.org/10.1016/j.bbrc.2013.05.114
Takahashi M, Miyatsuka T, Suzuki L et al (2020) Biphasic changes in β-cell mass around parturition are accompanied by increased serotonin production. Sci Rep 10(1):4962. https://doi.org/10.1038/s41598-020-61850-1
Kitamura T, Kido Y, Nef S, Merenmies J, Parada LF, Accili D (2001) Preserved pancreatic beta-cell development and function in mice lacking the insulin receptor-related receptor. Mol Cell Biol 21(16):5624–5630. https://doi.org/10.1128/mcb.21.16.5624-5630.2001
Finak G, McDavid A, Yajima M et al (2015) MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol 16:278. https://doi.org/10.1186/s13059-015-0844-5
Baetens D, Malaisse-Lagae F, Perrelet A, Orci L (1979) Endocrine pancreas: three-dimensional reconstruction shows two types of islets of langerhans. Science 206(4424):1323–1325. https://doi.org/10.1126/science.390711
Wang D, Wang J, Bai L et al (2020) Long-term expansion of pancreatic islet organoids from resident Procr(+) progenitors. Cell 180(6):1198–1211.e1119. https://doi.org/10.1016/j.cell.2020.02.048
Champy MF, Le Voci L, Selloum M, Peterson LB, Cumiskey AM, Blom D (2011) Reduced body weight in male Tspan8-deficient mice. Int J Obes 35(4):605–617. https://doi.org/10.1038/ijo.2010.165
Qadir MMF, Álvarez-Cubela S, Klein D et al (2020) Single-cell resolution analysis of the human pancreatic ductal progenitor cell niche. Proc Natl Acad Sci U S A 117(20):10876–10887. https://doi.org/10.1073/pnas.1918314117
Sachs S, Bastidas-Ponce A, Tritschler S et al (2020) Targeted pharmacological therapy restores β-cell function for diabetes remission. Nat Metab 2(2):192–209. https://doi.org/10.1038/s42255-020-0171-3
Wang H, Rana S, Giese N, Büchler MW, Zöller M (2013) Tspan8, CD44v6 and alpha6beta4 are biomarkers of migrating pancreatic cancer-initiating cells. Int J Cancer 133(2):416–426. https://doi.org/10.1002/ijc.28044
Watanabe T, Yonemura Y, Yonekura H et al (1994) Pancreatic beta-cell replication and amelioration of surgical diabetes by Reg protein. Proc Natl Acad Sci U S A 91(9):3589–3592. https://doi.org/10.1073/pnas.91.9.3589
Gross DJ, Weiss L, Reibstein I et al (1998) Amelioration of diabetes in nonobese diabetic mice with advanced disease by linomide-induced immunoregulation combined with Reg protein treatment. Endocrinology 139(5):2369–2374. https://doi.org/10.1210/endo.139.5.5997
Chen H, Gu X, Liu Y et al (2011) PDGF signalling controls age-dependent proliferation in pancreatic β-cells. Nature 478(7369):349–355. https://doi.org/10.1038/nature10502
Cox AR, Beamish CA, Carter DE, Arany EJ, Hill DJ (2013) Cellular mechanisms underlying failed beta cell regeneration in offspring of protein-restricted pregnant mice. Exp Biol Med 238(10):1147–1159. https://doi.org/10.1177/1535370213493715
Acknowledgements
We thank Genble Inc., Fukuoka, Japan, for support with scRNA-seq and data analysis. We thank A. Suda, Y. Tamura, A. Fukaishi, W. Mizutani and Y. Miyazaki (from the Fujitani laboratory, Gunma University, Gunma, Japan) for their technical assistance, H. A. Popiel (Tokyo Medical University, Tokyo, Japan) for critical reading of the manuscript, and H. Mizukami (Hirosaki University, Aomori, Japan), H. Tatsuoka, S. Tokumoto (Kyoto University, Kyoto, Japan), J. Shirakawa (IMCR, Gunma University) and the members of Y. Kitamura’s laboratory (IMCR, Gunma University) for fruitful discussions. We thank A. Oue, T. Kakinuma and other members of Bioresource Center, Gunma University, and S. Takahashi and other members of Laboratory Animal Resource Center, University of Tsukuba, Ibaraki, Japan, for providing technical support.
Author’s relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Funding
This study was supported by grants from the Ministry of Education, Sports and Culture of Japan (JSPS KAKENHI; 16K09765 and 20H0364 to YF and 19K06435 to TS), and the Takeda Science Foundation (to YF), and by the Joint Usage and Joint Research Programs, the Institute of Advanced Medical Sciences, Tokushima University.
Author information
Authors and Affiliations
Contributions
YF and TF conceived and designed the study. TF, YN, AF and TS performed the animal experiments and acquired the data. AH, KN, MS, KK, TMi, MT, MM and TMa contributed to resource preparation and data acquisition. TF and YF wrote the original draft. YN, AF, TS, AH, KN, MS, KK, TMi, MT, MM, TMa, TY and HW interpreted the data and revised the manuscript for valuable intellectual content. All authors gave final approval of the version to be published. YF is the guarantor of this work, has full access to all the data in the study, and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fukaishi, T., Nakagawa, Y., Fukunaka, A. et al. Characterisation of Ppy-lineage cells clarifies the functional heterogeneity of pancreatic beta cells in mice. Diabetologia 64, 2803–2816 (2021). https://doi.org/10.1007/s00125-021-05560-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-021-05560-x