Abstract
Clostridioides difficile (C. difficile) infection is associated with high morbidity and mortality. This study aimed to evaluate the protective effect of Lactiplantibacillus plantarum E51 (L. plantarum E51) on C. difficile infection using the Caco-2 monolayer in vitro model. Caco-2 cells were infected with C. difficile in the presence/absence of L. plantarum E51 or Lacticaseibacillus rhamnosus GG (LGG). Caco-2 intestinal barrier functions, such as monolayer integrity, IL-8 secretion, and tight junction protein expression, were quantified to investigate the extent to which L. plantarum E51 protected against C. difficile infection in vitro. Furthermore, inhibition of C. difficile adhesion to Caco-2 cells by L. plantarum E51 was explored using competition, exclusion, and displacement assays. The results indicated that L. plantarum E51 inhibited C. difficile growth, ameliorated C. difficile-caused decrease in transepithelial/ transendothelial electrical resistance, attenuated C. difficile-induced IL8 secretion, and upregulated claudin-1 protein expression that was inhibited by C. difficile. Moreover, L. plantarum E51 suppressed C. difficile adhesion to Caco-2 cells. In conclusion, these findings demonstrated that L. plantarum E51 substantially protected against C. difficile-induced damages on intestinal barrier functions in Caco-2 cells. The probiotic potential of L. plantarum E51 against C. difficile infection warrants further investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clostridioides difficile (C. difficile), an anaerobic spore-forming Gram-positive rod, is a major nosocomial pathogen that can cause diarrhea, antibiotic-associated pseudomembranous colitis, sepsis, and death (Paredes-Sabja et al. 2014). Colonization of the human intestinal tract by C. difficile occurs after alteration of the normal gut flora by antibiotic therapy. The pathogenesis of C. difficile infection is driven by the actions of its two exotoxins, Toxin A and Toxin B, which disrupt tight junctions in the intestinal epithelium and then cause inflammation (Di Bella et al. 2016; Simeon et al. 2019). C. difficile infection also results in cytotoxicity, apoptosis and necrosis (Tinoco-Veras et al. 2017); however, its treatments are still poorly understood (Elliott et al. 2017). Since the two main antibiotic treatments for C. difficile infection, metronidazole and vancomycin, have a 20% risk of reinfection, non-antibiotic treatments with sustained cure rates are being investigated (Wu and Wu 2012).
The FAO/WHO defined probiotic species as live microorganisms that confer a health benefit on the host when consumed in appropriate amounts (Hill et al. 2014). A variety of probiotic microorganisms were confirmed as beneficial to human health through their support of a healthy digestive system, such as multiple species of the lactic acid bacteria Lactobacillus and Bifidobacterium (Hill et al. 2014). Notably, 261 species of the genus Lactobacillus (at March 2020) were reclassified into 25 genera, including Lactiplantibacillus and Lacticaseibacillus (Zheng et al. 2020). Organisms suitable for probiotic use must be generally recognized as safe for ingestion (Sanders et al. 2010), able to survive and proliferate at gastric acid pH and in medium containing bile, and able to adhere to enterocytes (Capurso 2019).
A meta-analysis concluded that the use of confirmed probiotic species resulted in a decrease in C. difficile infections in patients taking antibiotics (Evans and Johnson 2015). A Cochrane Review of 31 trials (8672 participants) reported that probiotics reduced the risk of C. difficile-associated diarrhea by 60% (Goldenberg et al. 2017). Several lines of evidence indicated the promising effects of the probiotic agent Lacticaseibacillus rhamnosus GG (LGG) on C. difficile infection (Capurso 2019). First patented for use in 1989, LGG is the most widely studied probiotic strain (Capurso 2019). However, probiotic agents for the treatment of C. difficile infection are still scarce (Valdés-Varela et al. 2016). Thus, the search for organisms with the potential for clinical use as probiotic species agents is ongoing.
Lactiplantibacillus plantarum E51 (L. plantarum E51) was isolated from paocai, a Chinese fermented vegetable (Chang et al. 2013). In a preliminary screening of these isolates, L. plantarum E51 survived at low pH and in bile and exhibited the greatest adhesion to Caco-2 cells, even greater than that of LGG (Chang et al. 2013). Caco-2 cells, an immortalized human colorectal adenocarcinoma cell line, were widely used to mimic intestinal barrier function to investigate the molecular mechanisms underlying C. difficile infection in vitro (Mehdizadeh Gohari et al. 2019; Ghosh et al. 2020). The aim of this study was to systematically assess the protective effect of L. plantarum E51 on C. difficile-induced damages on Caco-2 intestinal barrier functions, including monolayer integrity, IL-8 secretion, tight junction protein expression, and adhesion to Caco-2 cells. The anticipated results might provide preliminary evidence for the probiotic potential of L. plantarum E51 against C. difficile infection.
Methods
Bacterial strains and culture conditions
Clostridioides difficile of known ribotype (Ofori et al. 2018), positive for Toxins A and B (Di Bella et al. 2016) and binary toxin (Aktories et al. 2018), and LGG were obtained from the Bioresource Collection and Research Center (BCRC, Hsinchu City, Taiwan). C. difficile was cultured in reinforced Clostridial medium (RCM; Difco Laboratories, Franklin Lakes, NJ, USA) in an anaerobic chamber with continuous shaking at 37 °C. L. plantarum E51 was isolated from paocai as previously described (Chang et al. 2013). Most Lactiplantibacillus and Lacticaseibacillus species can grow under aerobic condition (Maresca et al. 2018), so both LGG and L. plantarum E51 were cultured in lactobacilli MRS broth (Difco Laboratories) at 37 °C in a regular orbital shaking incubator in this study. Notably, RCM was specifically designed for the cultivation and enumeration of Clostridia (Borriello and Barclay 1985). Consistently, we previously found that LGG and L. plantarum E51 were not able to grow in RCM, while C. difficile could grow in both RCM and MRS broth.
Acid and bile salts tolerance assays
An acid tolerance assay was performed as previously described (Corcoran et al. 2005) with minor modifications. Cultures of the isolated bacteria (LGG and L. plantarum E51) were grown overnight in 15 mL lactobacilli MRS broth. The cultures were centrifuged at 12,000 × g for 5 min and washed once in an equal volume of cold phosphate-buffered saline (PBS). The pellets were re-suspended in 5 mL PBS, and the volume equivalent to approximately 107–108 colony-forming units (CFU)/mL was further centrifuged and re-suspended in the appropriate volume of PBS at pH 2.0 or 3.0 to mimic gastric conditions. The acidic PBS buffers were prepared by dissolving 20.214 g of Na2HPO4∙7H2O or 3.394 g of NaH2PO4∙H2O in 800 mL of water, adjusting the solution to the desired pH with HCl, and adding water to a final volume of 1000 mL. Bacteria were incubated at various pH values in a 37 °C water bath with shaking at 80 rpm for 3 h. After serial dilution and Gram staining, surviving bacteria were spread on lactobacilli MRS agar plates, and the numbers of LGG and L. plantarum E51 were counted and the corresponding CFU/mL was calculated.
The effect of bile salts on the growth of lactic acid bacteria cells was determined as previously described (Daniel et al. 2006) with minor modifications. Fresh cultures were inoculated into MRS broth enriched with Oxgall at three concentrations (0.2, 0.3 and 0.4%, w/v) (Sigma, St. Louis, MO, USA) and incubated at 37 °C for 12 h. Growth curves were plotted, and bile salt tolerance was determined as the percentage growth rate in bile salts, which was the ratio of (increment of OD in MRS broth with 0.2%, 0.3% or 0.4% bile salts)/(increment of OD in MRS broth without bile salts) × 100.
For gastrointestinal tract simulation, L. plantarum E51 was pre-treated with acid (pH 2.0 or 3.0) for 3 h, and the surviving bacteria were then collected by centrifugation (12,000 × g, 5 min) and washed with 0.1 M PBS (pH 7.4). The cell pellet was suspended in 3 mL MRS broth with 0.3% w/v Oxgall, and incubated at 37 °C for 12 h. After serial dilutions, the number of L. plantarum E51 on lactobacilli MRS agar plates was counted, and the CFU/mL was calculated. All assays were performed in triplicate.
Growth rate of C. difficile
C. difficile (108 CFU/mL) was incubated with or without in the presence or absence of LGG or L. plantarum E51 (108 CFU/mL) in lactobacilli MRS broth for 24 h at 37 °C in anaerobic chamber. Then, 0.1 ml MRS broth containing viable C. difficile was spread onto RCM agar plates at 0, 6, 10, 14, and 24 h and incubated anaerobically at 37 °C. After Gram staining, the number of C. difficile on each RCM agar plate was counted and the corresponding CFU/mL was calculated. All experiments were performed in triplicate.
Monolayer integrity assay
Caco-2 cells were routinely maintained in Dulbecco’s modified Eagle’s medium (DMED) supplemented with 10% fetal bovine serum (FBS), 1% non-essential amino acid, and penicillin–streptomycin (all from Sigma) in a 37 °C incubator with a humidified atmosphere of 5% CO2 in air. The extent of monolayer integrity was determined by measuring transepithelial/transendothelial electrical resistance (TEER). Briefly, Caco-2 monolayers were cultured on Millicell culture inserts (Merck Millipore, Burlington, MA, USA) in FBS-supplemented DMEM without antibiotics at 37 °C. Subsequently, C. difficile (108 CFU/mL) with or without either LGG or L. plantarum E51 (108 CFU/mL) were added to the apical compartment of Millicell cell culture inserts. TEER across the inner and outer compartments of the inserts was measured at 0, 4, 8, 12, 16, 20, and 24 h using the Millicell–ERS device (Merck Millipore).
IL-8 ELISA assay
Caco-2 monolayers with or without C. difficile in the presence or absence of LGG or L. plantarum E51 (each 108 CFU/mL), were cultured on the Millicell cell culture inserts (Merck Millipore) in FBS–supplemented DMEM without antibiotics at 37 °C. After a 15 h aerobic incubation, conditioned media from Caco-2 cells were harvested and centrifuged. The concentration of IL-8 in the supernatant was quantified using the human IL-8 ELISA kit (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instruction.
Western blotting
After a 15-h aerobic incubation with or without C. difficile in the presence or absence of LGG or L. plantarum E51 (each 108 CFU/mL), Caco-2 cells were lysed with RIPA buffer (Thermo Fischer Scientific), and the lysate proteins were separated by SDS–PAGE and transferred to a polyvinylidene fluoride (PVDF) membrane. The membrane was blocked in a solution of 5% skim milk, 1% bovine serum albumin, and 0.1% Tween 20 in Tris-buffered saline for 1 h at room temperature. The PVDF membrane was incubated with primary antibodies against claudin-1 (1:5000 dilution; Thermo Fisher Scientific) at 4 °C overnight, followed by incubation with secondary horseradish peroxidase (HRP) conjugated goat anti-rabbit IgG (1:2500 dilution; Thermo Fisher Scientific) for 1 h at room temperature. Proteins were visualized using Pierce ECL Western Blotting Substrate (Thermo Fisher Scientific). The claudin-1 band intensity was quantified using ImageJ software (National Institutes of Health, Bethesda, MD, USA) and normalized to that of β-actin in the same sample.
C. difficile adhesion to Caco-2 cells
Caco-2 monolayers were cultured in serum-supplemented DMEM in 24-well plates aerobically at 37 °C. Three assays for evaluating the inhibition of C. difficile adhesion to Caco-2 cells were conducted as previously described (Yu et al. 2011). First, competition assay: C. difficile (108 CFU/ml) was incubated on Caco-2 monolayers in the presence or absence of L. plantarum E51 or LGG (108 CFU/mL) for 1 h at 37 °C. Second, exclusion assay: Caco-2 monolayers were pre-incubated with L. plantarum E51 or LGG (108 CFU/mL) for 1 h, followed by the addition of C. difficile (108 CFU/mL) to the Caco-2 cultures and 1 h incubation at 37 °C. Third, displacement assay: Caco-2 monolayers were pre-incubated with C. difficile (108 CFU/mL) for 1 h, followed by addition of L. plantarum E51 or LGG (108 CFU/mL) and 1 h incubation at 37 °C. All three assays were conducted in aerobatic conditions due to the oxygen requirement of Caco-2 cells. At the end of each assay, the Caco-2 cultures were washed 3 times with sterile PBS to remove non-adherent bacteria, and were lysed with sterile distilled water for 5 min to release C. difficile that was adhered to Caco-2 cells. After tenfold dilution, cell lysates were spread on RCM agar plates followed by 6 h anaerobic incubation. Subsequently, the number of adhered C. difficile on each RCM agar plate was counted and the CFU/mL of adhered C. difficile was calculated. All experiments were performed in triplicate.
Statistical analysis
CFU values are presented as the mean and standard deviation (SD). The Kruskal–Wallis test was performed to examine the differences between groups. All data are expressed as the mean ± SD. The significance level was set as two-tailed p < 0.05. All statistical analyses were performed using IBM SPSS statistical software version 22 for Windows (IBM Corp., Armonk, NY, USA).
Results
L. plantarum E5 survived in acidic and bile-salt media
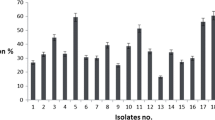
The survival of L. plantarum E51 in acidic medium and 0.2% bile salts was relatively better than that of LGG (Table 1). As the most widely studied probiotic strain, the survival capability of LGG under simulated gastrointestinal conditions and in the human gastrointestinal track has been demonstrated (Dommels et al. 2009; Karu and Sumeri 2016; Capurso 2019). Since LGG served as a positive control in this study, the survival of LGG in various pH conditions was not systematically examined. The survival rate of L. plantarum E51 was greater at pH 3.0 than pH 2.0. The survival of L. plantarum E51 was not affected by the concentration of bile salt. To mimic the environment of the gastrointestinal tract, L. plantarum E51 was first cultured in the acidic medium (pH 3.0 or 2.0), followed by the medium containing 0.3% bile salt. Consistently, the survival of L. plantarum E51 was higher at pH 3.0 than pH 2.0 (Table 1).
L. plantarum E51 inhibited C. difficile growth
The growth of C. difficile peaked after 8–10 h of co-culture with or without L. plantarum E51 or LGG in MRS broth and declined thereafter (Fig. 1). The maximum growth of C. difficile was slightly decreased by co-incubation with E51, compared to that of co-culture with LGG (Fig. 1).
L. plantarum E51 attenuated C. difficile-induced damage to Caco-2 monolayer integrity
TEER values of Caco-2 cells remained relatively steady over the 24 h incubation (Fig. 2). The integrity of the Caco-2 monolayer was disrupted in the presence of C. difficile, resulting in a gradual decrease in TEER. Both L. plantarum E51 and LGG attenuated the C. difficile-induced decrease in TEER, suggesting protective effects on Caco-2 cell membrane tight junctions or the inhibition of toxin production by C. difficile. This protective effect was greater for L. plantarum E51 than LGG (Fig. 2).
L. plantarum E51 reversed C. difficile-evoked changes in IL-8 secretion and claudin-1 protein expression
Clostridioides difficile significantly increased IL-8 secretion by Caco-2 cells (p < 0.01). In the presence of L. plantarum E51 or LGG, this upregulative effect of C. difficile on IL-8 secretion was significantly suppressed (both p < 0.01) (Fig. 3). On the other hand, C. difficile inhibited the protein expression of claudin-1 in Caco-2 cells. Both L. plantarum E51 and LGG reversed the inhibitory effect of C. difficile on claudin-1 expression, and significantly upregulated claudin-1 protein expression (both p < 0.01) (Fig. 4).
Effects of L. plantarum E51 and LGG on C. difficile-mediated inhibition of claudin-1 protein expression in Caco-2 cells. Caco-2 monolayers were treated with or without C. difficile in the presence or absence of L. plantarum E51 or LGG for 15 h. A The representative Western blot images. B Histogram of western blot results. Data are presented as the mean ± SD of triplicate samples. **p < 0.01
L. plantarum E51 inhibited C. difficile adhesion to Caco-2 cells
The effect of L. plantarum E51 on the capacity of C. difficile to adhere to Caco-2 cells was evaluated using competition, exclusion, and displacement assays. In the competition assay, L. plantarum E51, but not LGG, significantly reduced the number of C. difficile adhered to Caco-2 cells (p < 0.01) (Fig. 5A). In the exclusion assay, both L. plantarum E51 and LGG significantly decreased the number of C. difficile adhered to Caco-2 cells (both p < 0.01) (Fig. 5B). The displacement assay showed that L. plantarum E51 significantly decreased the number of C. difficile adhered to Caco-2 cells (p < 0.01) (Fig. 5C).
Discussion
In this preliminary in vitro study, we observed that L. plantarum E51 exerted a variety of protective effects against C. difficile in Caco-2 cells. L. plantarum E51 tolerated low pH and high-concentration bile salts. Compared to LGG, L. plantarum E51 exhibited better inhibition of C. difficile growth. In addition, L. plantarum E51 attenuated C. difficile-induced damage to Caco-2 monolayers, enhanced claudin-1 protein expression, suppressed C. difficile-induced IL-8 secretion, and decreased the adhesion of C. difficile to Caco-2 cells. Together, L. plantarum E51 substantially protected against C. difficile-induced damages on Caco-2 intestinal barrier functions.
C. difficile Toxins A and B disrupt the actin cytoskeleton and tight junctions of intestinal epithelial cells (Czepiel et al. 2019), allowing the toxins to enter the laminar propria and submucosa, where they induce the production of proinflammatory and cytotoxic molecules, such as IL-8 (Boonma et al. 2014). Increased IL-8 secretion by intestinal epithelial cells causes a massive influx of neutrophils into the colonic mucosa, resulting in epithelial damage due to inflammatory edema (Rupnik et al. 2009). We observed multiple protective effects of L. plantarum E51 on C. difficile infection, suggesting that the mechanisms underlying the protective effects may involve multiple steps in C. difficile pathogenesis. Our findings suggest that L. plantarum E51 may inhibit C. difficile infection in at least three ways: (i) suppressing C. difficile growth, (ii) preserving Caco-2 monolayer integrity via upregulating tight junction protein, and (iii) inhibiting C. difficile adhesion to Caco-2 cells.
The distribution of mechanisms underlying probiotic effects shows that many are widespread among species, while others are species specific (Hill et al. 2014). Our proposed mechanism for the effects of L. plantarum E51 on C. difficile is supported by observations in other bacterial species. Yu et al. reported that Lactobacillus delbrueckii ssp. delbrueckii D11 and Levilactobacillus brevis inhibit the adhesion of pathogenic bacteria to Caco-2 cells and concluded that these strains may be useful for protecting against pathogenic infection (Yu et al. 2011). Similarly, Lactobacillus delbrueckii ssp. bulgaricus B-30892 was reported to inhibit the adhesion of C. difficile to Caco-2 cells (Banerjee et al. 2009). Upregulation of claudin-1 was observed to reverse the effect of aspirin-induced epithelial barrier disruption to improve the epithelial barrier function in vitro (Nishii et al. 2020). Another potential mechanism that was not investigated in this study was observed in several strains of Bifidobacterium spp. and Lactiplantibacillus spp., which exerted antagonistic effects on the production of C. difficile toxins A and B (Trejo et al. 2010).
Our observation that L. plantarum E51 decreased C. difficile-induced secretion of IL-8 is consistent with the finding that L. rhamnosus L34 and L. casei L39 suppressed C. difficile-induced IL-8 production and inflammation (Boonma et al. 2014). Another L. planatarum species, Inducia (DSM 21379), was shown to decrease hamster gut colonization by C. difficile in vivo (Rätsep et al. 2017), and a combination of Lactiplantibacillus and Bifidobacterium strains conferred protection against C. difficile infection in antibiotic-treated mice (Kondepudi et al. 2014).
Because numerous mechanisms underlie the probiotic effects of studied species, differences in the type and extent of effects exerted by LGG and L. plantarum E51 on C. difficile are expected. LGG, a confirmed probiotic species, was widely used as a control strain to gauge the effects of L. plantarum E51 in this study. While LGG and L. plantarum E51 were similar overall in the types of effects exerted, the extent of these effects differed in some respects. Significant differences between LGG and L. plantarum E51 were observed in competition and displacement assays for inhibition of C. difficile adhesion to Caco-2 cells, with L. plantarum E51 demonstrating relatively strong inhibition.
Furthermore, an inhibitory effect of lactic acid produced by L. plantarum strain Inducia (DSM 21379) on the growth of C. difficile has been previously reported (Rätsep et al. 2017); however, the authors also suggest that this factor may not be the only mechanism underlying the observed inhibitory effect. In the present study, the pH values of the culture medium were not substantially decreased during the growth of LGG or L. plantarum E 51. Hence, the effect of lactic acid production by LGG and L. plantarum E51 in the inhibition of C. difficile growth appears to be minor in the present study.
This study has several limitations. First of all, since C. difficile Toxins A and B were not quantified in this study, the possibility that cellular damage by C. difficile observed may be at least in part caused by its produced toxins cannot be ruled out. In addition, because of aerobic respiration, Caco-2 cells constantly require oxygen. Hence, the experiments involving Caco-2 cells had to be performed in aerobic condition in the present study; however, the extent to which such aerobic condition affects the vitality and virulence of C. difficile is unclear. Therefore, although the Caco-2 monolayer model has been extensively used in the investigation C. difficile infection, in vivo studies are still warranted to assess the effectiveness of candidate probiotic agents against C. difficile infection.
In conclusion, the findings of this in vitro study demonstrated that L. plantarum E51 effectively reduced the harmful effects of C. difficile, thereby preserving Caco-2 intestinal barrier functions. Although L. plantarum E51 exhibited several necessary attributes of probiotic agents, further in vivo studies are warranted to investigate the probiotic potential of L. plantarum E51 against C. difficile infection.
References
Aktories K, Papatheodorou P, Schwan C (2018) Binary Clostridium difficile toxin (CDT)—a virulence factor disturbing the cytoskeleton. Anaerobe 53:21–29. https://doi.org/10.1016/j.anaerobe.2018.03.001
Banerjee P, Merkel GJ, Bhunia AK (2009) Lactobacillus delbrueckii ssp. bulgaricus B-30892 can inhibit cytotoxic effects and adhesion of pathogenic Clostridium difficile to Caco-2 cells. Gut Pathog. https://doi.org/10.1186/1757-4749-1-8
Boonma P, Spinler JK, Venable SF, Versalovic J, Tumwasorn S (2014) Lactobacillus rhamnosus L34 and Lactobacillus casei L39 suppress Clostridium difficile-induced IL-8 production by colonic epithelial cells. BMC Microbiol 14:177. https://doi.org/10.1186/1471-2180-14-177
Borriello SP, Barclay FE (1985) Protection of hamsters against Clostridium difficile ileocaecitis by prior colonisation with non-pathogenic strains. J Med Microbiol 19:339–350. https://doi.org/10.1099/00222615-19-3-339
Capurso L (2019) Thirty years of Lactobacillus rhamnosus GG: a review. J Clin Gastroenterol 53(Suppl 1):S1-s41. https://doi.org/10.1097/mcg.0000000000001170
Chang S-M, Tsai C-L, Wee W-C, Yan T-R (2013) Isolation and functional study of potentially probiotic Lactobacilli from Taiwan traditional paocai. Afr J Microbiol Res 7:683–691
Corcoran BM, Stanton C, Fitzgerald GF, Ross RP (2005) Survival of probiotic lactobacilli in acidic environments is enhanced in the presence of metabolizable sugars. Appl Environ Microbiol 71:3060–3067. https://doi.org/10.1128/aem.71.6.3060-3067.2005
Czepiel J et al (2019) Clostridium difficile infection: review. Eur J Clin Microbiol Infect Dis 38:1211–1221. https://doi.org/10.1007/s10096-019-03539-6
Daniel C, Poiret S, Goudercourt D, Dennin V, Leyer G, Pot B (2006) Selecting lactic acid bacteria for their safety and functionality by use of a mouse colitis model. Appl Environ Microbiol 72:5799–5805. https://doi.org/10.1128/aem.00109-06
Di Bella S, Ascenzi P, Siarakas S, Petrosillo N, di Masi A (2016) Clostridium difficile Toxins A and B: insights into pathogenic properties and extraintestinal effects. Toxins (Basel). https://doi.org/10.3390/toxins8050134
Dommels YE et al (2009) Survival of Lactobacillus reuteri DSM 17938 and Lactobacillus rhamnosus GG in the human gastrointestinal tract with daily consumption of a low-fat probiotic spread. Appl Environ Microbiol 75:6198–6204. https://doi.org/10.1128/aem.01054-09
Elliott B, Androga GO, Knight DR, Riley TV (2017) Clostridium difficile infection: evolution, phylogeny and molecular epidemiology. Infect Genet Evol 49:1–11. https://doi.org/10.1016/j.meegid.2016.12.018
Evans CT, Johnson S (2015) Prevention of Clostridium difficile infection with probiotics. Clin Infect Dis 60(Suppl 2):S122-128. https://doi.org/10.1093/cid/civ138
Ghosh C, AbdelKhalek A, Mohammad H, Seleem MN, Haldar J (2020) Aryl-alkyl-lysines: novel agents for treatment of C. difficile infection. Sci Rep 10:5624. https://doi.org/10.1038/s41598-020-62496-9
Goldenberg JZ et al (2017) Probiotics for the prevention of Clostridium difficile-associated diarrhea in adults and children. Cochrane Database Syst Rev 12:006095. https://doi.org/10.1002/14651858.CD006095.pub4
Hill C et al (2014) Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11:506–514. https://doi.org/10.1038/nrgastro.2014.66
Karu R, Sumeri IJJOFR (2016) Survival of Lactobacillus rhamnosus GG during simulated gastrointestinal conditions depending on food matrix. J Food Res 5:57–66. https://doi.org/10.5539/jfr.v5n5p57
Kondepudi KK, Ambalam P, Karagin PH, Nilsson I, Wadström T, Ljungh Å (2014) A novel multi-strain probiotic and synbiotic supplement for prevention of Clostridium difficile infection in a murine model. Microbiol Immunol 58:552–558. https://doi.org/10.1111/1348-0421.12184
Maresca D, Zotta T, Mauriello G (2018) Adaptation to aerobic environment of Lactobacillus johnsonii/gasseri strains. Front Microbiol 9:157. https://doi.org/10.3389/fmicb.2018.00157
Mehdizadeh Gohari I, Li J, Navarro M, Uzal F, McClane B (2019) Effects of claudin-1 on the action of Clostridium perfringens enterotoxin in Caco-2 cells. Toxins 11:582. https://doi.org/10.3390/toxins11100582
Nishii N et al (2020) Lubiprostone induces claudin-1 and protects intestinal barrier function. Pharmacology 105:102–108. https://doi.org/10.1159/000503054
Ofori E, Ramai D, Dhawan M, Mustafa F, Gasperino J, Reddy M (2018) Community-acquired Clostridium difficile: epidemiology, ribotype, risk factors, hospital and intensive care unit outcomes, and current and emerging therapies. J Hosp Infect 99:436–442. https://doi.org/10.1016/j.jhin.2018.01.015
Paredes-Sabja D, Shen A, Sorg JA (2014) Clostridium difficile spore biology: sporulation, germination, and spore structural proteins. Trends Microbiol 22:406–416. https://doi.org/10.1016/j.tim.2014.04.003
Rätsep M et al (2017) A combination of the probiotic and prebiotic product can prevent the germination of Clostridium difficile spores and infection. Anaerobe 47:94–103. https://doi.org/10.1016/j.anaerobe.2017.03.019
Rupnik M, Wilcox MH, Gerding DN (2009) Clostridium difficile infection: new developments in epidemiology and pathogenesis. Nat Rev Microbiol 7:526–536. https://doi.org/10.1038/nrmicro2164
Sanders ME et al (2010) Safety assessment of probiotics for human use. Gut Microbes 1:164–185. https://doi.org/10.4161/gmic.1.3.12127
Simeon R et al (2019) Selection and characterization of ultrahigh potency designed ankyrin repeat protein inhibitors of C. difficile toxin B. PLoS Biol 17:e3000311. https://doi.org/10.1371/journal.pbio.3000311
Tinoco-Veras CM et al (2017) Transforming growth factor β1/SMAD signaling pathway activation protects the intestinal epithelium from Clostridium difficile toxin A-induced damage. Infect Immun 85:e00430-e517. https://doi.org/10.1128/iai.00430-17
Trejo FM, Pérez PF, De Antoni GL (2010) Co-culture with potentially probiotic microorganisms antagonises virulence factors of Clostridium difficile in vitro. Antonie Van Leeuwenhoek 98:19–29. https://doi.org/10.1007/s10482-010-9424-6
Valdés-Varela L, Alonso-Guervos M, García-Suárez O, Gueimonde M, Ruas-Madiedo P (2016) Screening of Bifidobacteria and lactobacilli able to antagonize the cytotoxic effect of Clostridium difficile upon intestinal epithelial HT29 monolayer. Front Microbiol 7:577–577. https://doi.org/10.3389/fmicb.2016.00577
Wu HJ, Wu E (2012) The role of gut microbiota in immune homeostasis and autoimmunity. Gut Microbes 3:4–14. https://doi.org/10.4161/gmic.19320
Yu Q, Wang Z, Yang Q (2011) Ability of Lactobacillus to inhibit enteric pathogenic bacteria adhesion on Caco-2 cells. World J Microbiol Biotechnol 27:881–886. https://doi.org/10.1007/s11274-010-0530-4
Zheng J et al (2020) A taxonomic note on the genus Lactobacillus: description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int J Syst Evol Microbiol 70:2782–2858. https://doi.org/10.1099/ijsem.0.004107
Acknowledgements
The authors wish to thank Shiao-Ming Chang and Shi-Ming Chen for their assistance with the laboratory work of this study. We would like to thank Convergence CT for providing English editing to improve English grammar and language usage.
Funding
None.
Author information
Authors and Affiliations
Contributions
H–SJ and T-RY: conception and design. H–SJ: acquisition, analysis and interpretation of data, and drafting the manuscript. T-RY: critical revision of the manuscript. H–SJ and T-RY: final approval of the manuscript. T-RY: supervision.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jeng, HS., Yan, TR. Lactiplantibacillus plantarum E51 protects against Clostridioides difficile-induced damages on Caco-2 intestinal barrier functions. Arch Microbiol 204, 290 (2022). https://doi.org/10.1007/s00203-022-02837-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-02837-6