Abstract
Current evidence suggests that some form of cellular organization arose well before the time of the last universal common ancestor (LUCA). Standard phylogenetic analyses have shown that several protein families associated with membrane translocation, membrane transport, and membrane bioenergetics were very likely present in the proteome of the LUCA. Despite these cellular systems emerging prior to the LUCA, extant archaea, bacteria, and eukaryotes have significant differences in cellular infrastructure and the molecular functions that support it, leading some researchers to argue that true cellularity did not evolve until after the LUCA. Here, we use recently reconstructed minimal proteomes of the LUCA as well as the last archaeal common ancestor (LACA) and the last bacterial common ancestor (LBCA) to characterize the evolution of cellular systems along the first branches of the tree of life. We find that a broad set of functions associated with cellular organization were already present by the time of the LUCA. The functional repertoires of the LACA and LBCA related to cellular organization nearly doubled along each branch following the divergence of the LUCA. These evolutionary trends created the foundation for similarities and differences in cellular organization between the taxonomic domains that are still observed today.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cellular organization is a defining characteristic of all life on Earth. A detailed and accurate understanding of the emergence of cellularity is, therefore, crucial to our broader account of early evolutionary history. The subject, however, remains somewhat contentious due to countervailing lines of evidence, some of which indicates an early evolution of cellularity, perhaps even coincident with the origin of life, itself, and some of which indicates a late evolution of cellularity, perhaps even following the divergence of the LUCA into the separate ancestors of bacteria and archaea.
Protocell experiments have demonstrated that membranes can form spontaneously from prebiotically available compounds such as decanoic acid (Namani and Walde 2005) as well as the lipid fractions of carbonaceous chondrite meteorites (Deamer and Pashley 1989), suggesting a possible role for membrane compartmentalization as early as the origin of life, itself. Artificial life simulations have shown that protocell encapsulation could have protected early replicator genomes from parasites (Hogeweg and Takeuchi 2003; Takeuchi and Hogeweg 2009; Shah, et al. 2019) and supported genomic stability in general (Takagi, et al. 2020). Other artificial life simulations have also shown that, in the kind of rich environment often associated with the origin of life, selection would have acted against protocell encapsulation even if that encapsulation was imposed by the environment rather than produced by the life form, but that protocell encapsulation would have eventually co-evolved along with the first metabolic pathways (Szathmary 2007; Takagi, et al. 2020).
Phylogenetic reconstructions of universal paralog protein families have shown that cell membrane-associated proteins such as ABC transporters, ATP synthase enzymes, and the signal recognition particle system, all underwent gene duplications prior to the time of the last universal common ancestor (LUCA) (Gribaldo and Cammarano 1998; Kollman and Doolittle 2000; Zhaxybayeva, et al. 2005; Harris and Goldman 2021). This evidence would indicate that cellular organization was well in place by the time of the LUCA. However, the primary constituents of cell membranes, phospholipids, are radically different between archaea and bacteria (Pereto, et al. 2004; Sojo 2019). Typical bacterial phospholipids contain fatty acid chains that are connected to the phosphate head group by an ester linkage, while typical archaeal phospholipids contain isoprenoid chains that are connected to the head group by an ether linkage. Though all bacterial and most archaeal phospholipids are composed of a head group and two hydrophobic tails that form a bilayer membrane, some archaeal phospholipids contain hydrophobic tails that bridge across the bilayer with a phosphate head group on either side. These differences in the main constituents of cell membranes, as well as the metabolic pathways that produce them, have been taken as evidence by some researchers that true cellular organization evolved after the time of the LUCA (Wachtershauser 1988; Koga, et al. 1998; Martin and Russell 2003; Weiss, et al. 2016).
Given the different lines of evidence, some researchers have argued for a central role of protocell compartmentalization as early as the origin of life itself (for example, (Saha and Chen 2015; West, et al. 2017; Damer and Deamer 2020; Nunes Palmeira, et al. 2022; Goldman 2023)) and that the last universal common ancestor was a fully cellular organism (Becerra, et al. 2007; Goldman, et al. 2023), while others have argued that even by the relatively later stage of the LUCA, life forms were still not fully cellular (Wachtershauser 1988; Koga, et al. 1998; Martin and Russell 2003; Koonin and Martin 2005; Weiss, et al. 2016).
Here, we seek to reconcile the evidence for and against cellular organization by the time of the LUCA by identifying protein families associated with cellular organization in minimal proteome reconstructions of the LUCA as well as its successors, the last archaeal common ancestor (LACA) and the last bacterial common ancestor (LBCA). In doing so, we portray, in broad terms, the kinds of cellular functions that likely were and were not encoded in the LUCA and how these functions appear to have expanded along the first branches of the tree of life. By complementing an analysis of LUCA cellular functions with an analysis of the subsequent evolution of those functions following the LUCA, we aim to provide context for major differences in cellular organization between the bacteria and archaea that can still be observed today.
Methods
Previously published minimal proteomes representing the LUCA (Crapitto, et al. 2022), the LACA (Williams, et al. 2017), and the LBCA (Coleman, et al. 2021) were modeled as EggNOG clusters of proteins (Huerta-Cepas, et al. 2019), a proxy for families of homologous proteins. These reconstructions are referred to as minimal proteomes because some protein families present in the actual proteome of each ancestor may have undergone subsequent loss or non-orthologous gene displacement and therefore would be absent from the ancestral proteome reconstruction (Koonin, et al. 1996). The LUCA minimal proteome (Crapitto, et al. 2022) was inferred through a meta-analysis of eight previously published LUCA proteome studies (Harris, et al. 2003; Mirkin, et al. 2003; Delaye, et al. 2005; Yang, et al. 2005; Ranea, et al. 2006; Wang, et al. 2007; Srinivasan and Morowitz 2009; Weiss, et al. 2016) and represents the consensus predictions between all eight studies. The LACA and LBCA minimal proteomes were inferred from tree reconciliation studies (Williams, et al. 2017; Coleman, et al. 2021) using the Amalgamated Likelihood Estimation (ALE) algorithm (Szollosi, et al. 2013), which compares gene and species trees to estimate gene duplications and losses as well as horizontal gene transfer.
Data from the LUCA, LACA, and LBCA minimal proteome studies were acquired from the supplementary information associated with each of the three studies. The minimal LUCA proteome consisted of 366 EggNOG clusters, the minimal LACA proteome consisted of 174 EggNOG clusters, and the minimal LBCA proteome consisted of 397 EggNOG clusters. Gene Ontology (GO) terms (Ashburner, et al. 2000; Gene Ontology, et al. 2023) associated with each EggNOG cluster were identified using the UniProt ID Mapping database (Bairoch and Apweiler 1997; Huang, et al. 2011; UniProt 2023).
For each ancestral proteome, EggNOG clusters associated with cellular function were identified by searching for the following GO terms related to cellularity: “membrane” (GO:0016020), “phospholipid biosynthetic process” (GO:0008654), “phospholipid transfer to membrane” (GO:0006649), “protein targeting” (GO:0006605), “endocytosis” (GO:0006897), “exocytosis” (GO:0006887), “transmembrane transporter activity” (GO:0022857), “cell division” (GO:0051301), “cytoskeleton” (GO:0005856), “structural constituent of cytoskeleton” (GO:0005200), “FtsZ-dependent cytokinesis” (GO:0043093), “single organism reproductive process” (GO:0044702), “membrane fission” (GO:0090148), and “mitotic cell cycle process” (GO:1903047). After identifying EggNOG clusters associated with broadly defined cellular functions, we gathered all other GO terms associated with these EggNOG clusters (Supplementary Data 1) and removed non-cellular functions by hand. For each individual GO term, we also included all other GO terms that were more general (i.e., the parent terms of each term, the parent terms of those parent terms, etc.).
The Gene Ontology database contains annotations that are nested within more general annotations, or what the Gene Ontology database refers to as “parent” terms. For example, the GO term "SRP-dependent co-translational protein targeting to membrane" (GO:0006614) is nested within the more general parent GO term, “co-translational protein targeting to membrane” (GO:0006613), which itself is nested within the more general parent GO term "protein targeting to membrane" (GO:0006605). Different proteins within the LUCA, LBCA, and LACA, minimal proteomes were annotated at different levels of specificity and, overall, bacterial proteins tended to be annotated with more specific GO terms, while archaeal proteins tended to be annotated with more general GO terms.
In order to directly compare these datasets, we standardized the level of specificity across all three ancestral minimal proteomes so that the presence or absence of membrane-associated GO terms could be compared between the LUCA, LACA, and LBCA. To do so, we identified fifteen GO terms that appeared to be at a similar level of generality and then identified the direct child terms of these general GO terms. These general GO terms were “reproduction” (GO:0000003), “cytokinesis” (GO:0000910), “transporter activity” (GO:0005215), “cytoskeleton” (GO:0005856), “intracellular protein transport” (GO:0006886), “cytoskeleton organization” (GO:0007010), “cellular process” (GO:0009987), “asexual reproduction” (GO:0019954), “cell cycle process” (GO:0022402), “reproductive process” (GO:0022414), “reproduction of a single-celled organism” (GO:0032505), “secretion by cell” (GO:0032940), “transmembrane transport” (GO:0055085), “membrane organization” (GO:0061024), and “import into cell” (GO:0098657). These general GO terms and their direct child terms are used in the analysis described below. Several of these general GO terms produced redundant child terms and were combined, resulting in the ten general GO terms shown in Fig. 1.
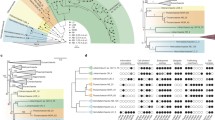
Cellular functions associated with the minimal proteomes of the LUCA, LACA, and LBCA. A The number of GO terms associated with cellular organization in each ancestor and the number of functions that are inferred to have been inherited, gained, and lost along each branch. Only GO terms that are direct child terms of the fifteen general GO terms relating to cellular organization were analyzed. B The total number of direct child GO terms within each category of general GO terms for each ancestor. Some of the original fifteen general GO terms produced redundant direct child GO terms and were combined. This figure was, in part, created with BioRender
Results
Within the minimal proteomes of the LUCA, LACA, and LBCA, we found 39, 27, and 45, EggNOG clusters associated with membrane functions, respectively (Supplemental File 1). Most of these EggNOG clusters were identified using the “membrane” GO term: 33/39 LUCA EggNOG clusters, 18/27 LACA EggNOG clusters, and 33/45 LBCA EggNOG clusters, representing 11%, 16%, and 11%, of the protein families in each ancestor’s proteome, respectively. Very few EggNOG clusters were associated with the “phospholipid biosynthetic process” GO term: 4/39 LUCA EggNOG clusters, 1/27 LACA EggNOG clusters, and 1/45 LUCA EggNOG clusters. The primarily eukaryotic GO terms, endocytosis, exocytosis, membrane fission, and mitotic cell cycle process, did not yield any EggNOG clusters in the reconstructed proteomes of the LUCA, LACA, and LBCA.
We hierarchically standardized the collected GO terms in order to compare them across ancestral proteomes (see Methods). This hierarchical analysis resulted in fifteen parent terms and fifty-seven direct child terms (Supplemental File S2). Comparing the non-redundant sets of direct child terms between the LUCA and the LACA or LBCA shows that 90% of GO terms associated with the LUCA were also present in the LBCA dataset while 62% of GO terms associated with the LUCA were present in the LACA dataset. The Jaccard similarity index between the LUCA and the LBCA GO terms was 43%, while the Jaccard similarity index between the LUCA and the LACA GO terms was 27%. The Jaccard similarity index between two datasets is calculated as the intersection, i.e., the number of shared items, divided by the union, i.e., the total number of items in both datasets. As a measure of similarity, it therefore takes into account both the number of shared items between two datasets and the sizes of both datasets. By both methods of comparison, i.e., the percentage of shared GO terms in the LUCA dataset and the Jaccard similarity index, the LBCA appears closer to the LUCA in terms of cellular functions, while the LACA appears more distinct from the LUCA.
The cellular functions associated with each ancestor and the differences between them are shown in Fig. 1. Most cellular functions associated with the LUCA proteome involve transport across the membrane, including the transport of amino acids, carbohydrates, and ions. The LUCA proteome also encodes the ability to target proteins to the membrane, a function that would be required for embedding transporter proteins into the membrane in the first place. Other functions associated with the reconstructed LUCA proteome are related to cell division, cell motility, cellular response to stimuli, cell wall organization or biogenesis, and cell aggregation.
Comparisons between cellular functions associated with the reconstructed LUCA proteome and that of its successors, the LACA and LBCA, are largely characterized by the introduction of new cellular functions with a small number of functions lost along each branch (Fig. 1). These new functions acquired by the LACA and LBCA following the divergence of the LUCA suggest a parallel evolution of new cytoskeletal elements and cell reproduction processes as well as differentiation with respect to specific membrane transport functions. The LBCA ancestral proteome also includes functions associated with horizontal gene transfer (GO:0009292) as well as the killing of other cells (GO:0001906) through toxin activity (GO:0090729).
Discussion
Inferring the proteomes of organisms that lived approximately 3.5-4Gya is inherently difficult (Crapitto, et al. 2022). EggNOG clusters may be incorrectly included or excluded from one of the ancestral proteomes due to limitations of the methodologies. For example, a protein family may have been present in the LUCA but lost to such an extent in subsequent lineages that it cannot be reconstructed as such. For simplicity, we describe differences in the presence and absence of protein functions between the three ancestral proteomes as gains and losses, but these results can also be explained by methodological shortcomings. For this reason, we caution against interpreting the results of an individual protein family being present in one of the ancestral proteomes as definitive. Instead, we portray the evolution of cellular functions in the LUCA, LACA, and LBCA in broad terms that both address the competing hypotheses about when cellularity first evolved and also provide a roadmap for future research.
Taken together, these results suggest that the LUCA represents a population of cellular organisms. By the time of the LUCA, many different cellular functions had evolved, and these appear to have expanded during the subsequent evolution of the LACA and the LBCA. The small number of phospholipid biosynthesis enzyme families found in all three datasets agrees with the observation that bacterial (and eukaryotic) phospholipids differ chemically from archaea. Recent evidence suggests that phospholipid chemistry is diverse even within the archaeal (Caforio and Driessen 2017) and bacterial (Sohlenkamp and Geiger 2016) domains, which explains the lack of conserved phospholipid biosynthesis enzymes even within the LACA and LBCA proteomes. However, despite the lack of a clear signal of conserved phospholipid biosynthesis in any of these ancestors, other cellular systems were clearly in place at the time of the LUCA (Lombard, et al. 2012) and expanded upon during the evolution of the LACA and LBCA.
The cellular functions associated with the minimal LUCA proteome depict a cellular organism capable of embedding proteins within the membrane and controlling, to some extent, the translocation of ions and biomolecules across that membrane. Importantly, the LUCA also seems to have been capable of controlling its cell division rather than relying on spontaneous growth and division as is observed in protocells (Berclaz, et al. 2001; Hanczyc, et al. 2003). The LUCA also appears to have had at least some form of a cell wall even though cell wall composition is not universal across the bacteria, archaea, and eukaryotes. The LBCA and LACA appear to have evolved additional functions related to transmembrane transport and cell reproduction and exhibit a parallel evolution of cytoskeletal elements. However, it is also possible that any or all of these features were present in the LUCA as well, but were not reconstructed as such by our methods. The LBCA also evolved several other traits including cell killing through toxins and the ability to actively facilitate horizontal gene transfer, suggesting that it lived within a complex microbial ecology (Goldman and Kacar 2023).
Perhaps most intriguingly, LUCA appears to have more cellular functions in common with the LBCA than the LACA, suggesting that cellular organization in the LUCA was more like that of bacteria than archaea. If this trend is also true for phospholipid biosynthesis, it would imply that the LUCA membrane was composed of bacteria-like phospholipids, i.e., fatty acid tails and an ester-linked phosphate head group, and that the archaeal phospholipids with isoprenoid tails and an ether-linked phosphate head group were derived in the LACA lineage following the divergence of the LUCA. Future studies pairing phylogenetic analysis with ancestral reconstruction will provide greater detail about specific protein families present in the LUCA, LACA, and LBCA, as well as the molecular functions that those proteins were performing in ancient life.
References
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Bairoch A, Apweiler R (1997) The SWISS-PROT protein sequence database: its relevance to human molecular medical research. J Mol Med (berl) 75:312–316
Becerra A, Delaye L, Islas S, Lazcano A (2007) The very early stages of biological evolution and the nature of the last common ancestor of the three major cell domains. Annu Rev Ecol Evol Syst 38:361–379
Berclaz N, Müller M, Walde P, Luisi PL (2001) Growth and transformation of vesicles studied by ferritin labeling and cryotransmission electron microscopy. J Phys Chem B 105:1056–1064
Caforio A, Driessen AJM (2017) Archaeal phospholipids: Structural properties and biosynthesis. Biochim Biophys Acta Mol Cell Biol Lipids 1862:1325–1339
Coleman GA, Davin AA, Mahendrarajah TA, Szantho LL, Spang A, Hugenholtz P, Szollosi GJ, Williams TA (2021) A rooted phylogeny resolves early bacterial evolution. Science. https://doi.org/10.1126/science.abe0511
Crapitto AJ, Campbell A, Harris AJ, Goldman AD (2022) A consensus view of the proteome of the last universal common ancestor. Ecol Evol 12:e8930
Damer B, Deamer D (2020) The hot spring hypothesis for an origin of life. Astrobiology 20:429–452
Deamer DW, Pashley RM (1989) Amphiphilic components of the murchison carbonaceous chondrite: surface properties and membrane formation. Orig Life Evol Biosph 19:21–38
Delaye L, Becerra A, Lazcano A (2005) The last common ancestor: what’s in a name? Orig Life Evol Biosph 35:537–554
Gene Ontology C, Aleksander SA, Balhoff J, Carbon S, Cherry JM, Drabkin HJ, Ebert D, Feuermann M, Gaudet P, Harris NL et al (2023) The gene ontology knowledgebase in 2023. Genetics 224:iyad031
Goldman AD (2023) How did life become cellular? Proc Biol Sci 290:20222327
Goldman AD, Kacar B (2023) Very early evolution from the perspective of microbial ecology. Environ Microbiol 25:5–10
Goldman AD, Weber JM, Larowe DE, Barge LM (2023) Electron transport chains as a window into the earliest stages of evolution. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.2210924120
Gribaldo S, Cammarano P (1998) The root of the universal tree of life inferred from anciently duplicated genes encoding components of the protein-targeting machinery. J Mol Evol 47:508–516
Hanczyc MM, Fujikawa SM, Szostak JW (2003) Experimental models of primitive cellular compartments: encapsulation, growth, and division. Science 302:618–622
Harris AJ, Goldman AD (2021) The very early evolution of protein translocation across membranes. PLoS Comput Biol 17:e1008623
Harris JK, Kelley ST, Spiegelman GB, Pace NR (2003) The genetic core of the universal ancestor. Genome Res 13:407–412
Hogeweg P, Takeuchi N (2003) Multilevel selection in models of prebiotic evolution: compartments and spatial self-organization. Orig Life Evol Biosph 33:375–403
Huang H, McGarvey PB, Suzek BE, Mazumder R, Zhang J, Chen Y, Wu CH (2011) A comprehensive protein-centric ID mapping service for molecular data integration. Bioinformatics 27:1190–1191
Huerta-Cepas J, Szklarczyk D, Heller D, Hernandez-Plaza A, Forslund SK, Cook H, Mende DR, Letunic I, Rattei T, Jensen LJ et al (2019) eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47:D309–D314
Koga Y, Kyuragi T, Nishihara M, Sone N (1998) Did archaeal and bacterial cells arise independently from noncellular precursors? A hypothesis stating that the advent of membrane phospholipid with enantiomeric glycerophosphate backbones caused the separation of the two lines of descent. J Mol Evol 46:54–63
Kollman JM, Doolittle RF (2000) Determining the relative rates of change for prokaryotic and eukaryotic proteins with anciently duplicated paralogs. J Mol Evol 51:173–181
Koonin EV, Martin W (2005) On the origin of genomes and cells within inorganic compartments. Trends Genet 21:647–654
Koonin EV, Mushegian AR, Bork P (1996) Non-orthologous gene displacement. Trends Genet 12:334–336
Lombard J, Lopez-Garcia P, Moreira D (2012) The early evolution of lipid membranes and the three domains of life. Nat Rev Microbiol 10:507–515
Martin W, Russell MJ (2003) On the origins of cells: a hypothesis for the evolutionary transitions from abiotic geochemistry to chemoautotrophic prokaryotes, and from prokaryotes to nucleated cells. Philos Trans R Soc Lond B Biol Sci 358:59–83 (discussion 83-55)
Mirkin BG, Fenner TI, Galperin MY, Koonin EV (2003) Algorithms for computing parsimonious evolutionary scenarios for genome evolution, the last universal common ancestor and dominance of horizontal gene transfer in the evolution of prokaryotes. BMC Evol Biol 3:2
Namani T, Walde P (2005) From decanoate micelles to decanoic acid/dodecylbenzenesulfonate vesicles. Langmuir 21:6210–6219
Nunes Palmeira R, Colnaghi M, Harrison SA, Pomiankowski A, Lane N (2022) The limits of metabolic heredity in protocells. Proc Biol Sci 289:20221469
Pereto J, Lopez-Garcia P, Moreira D (2004) Ancestral lipid biosynthesis and early membrane evolution. Trends Biochem Sci 29:469–477
Ranea JA, Sillero A, Thornton JM, Orengo CA (2006) Protein superfamily evolution and the last universal common ancestor (LUCA). J Mol Evol 63:513–525
Saha R, Chen IA (2015) Origin of life: protocells red in tooth and claw. Curr Biol 25:R1175-1177
Shah V, de Bouter J, Pauli Q, Tupper AS, Higgs PG (2019) Survival of RNA replicators is much easier in protocells than in surface-based spatial systems. Life 9:65
Sohlenkamp C, Geiger O (2016) Bacterial membrane lipids: diversity in structures and pathways. FEMS Microbiol Rev 40:133–159
Sojo V (2019) Why the lipid divide? Membrane proteins as drivers of the split between the lipids of the three domains of life. BioEssays 41:e1800251
Srinivasan V, Morowitz HJ (2009) The canonical network of autotrophic intermediary metabolism: minimal metabolome of a reductive chemoautotroph. Biol Bull 216:126–130
Szathmary E (2007) Coevolution of metabolic networks and membranes: the scenario of progressive sequestration. Philos Trans R Soc Lond B Biol Sci 362:1781–1787
Szollosi GJ, Rosikiewicz W, Boussau B, Tannier E, Daubin V (2013) Efficient exploration of the space of reconciled gene trees. Syst Biol 62:901–912
Takagi YA, Nguyen DH, Wexler TB, Goldman AD (2020) The coevolution of cellularity and metabolism following the origin of life. J Mol Evol 88:598–617
Takeuchi N, Hogeweg P (2009) Multilevel Selection in models of prebiotic evolution II: a direct comparison of compartmentalization and spatial self-organization. PLoS Comput Biol 5:e1000542
UniProt C (2023) UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res 51:D523–D531
Wachtershauser G (1988) Before enzymes and templates: theory of surface metabolism. Microbiol Rev 52:452–484
Wang M, Yafremava LS, Caetano-Anolles D, Mittenthal JE, Caetano-Anolles G (2007) Reductive evolution of architectural repertoires in proteomes and the birth of the tripartite world. Genome Res 17:1572–1585
Weiss MC, Sousa FL, Mrnjavac N, Neukirchen S, Roettger M, Nelson-Sathi S, Martin WF (2016) The physiology and habitat of the last universal common ancestor. Nat Microbiol 1:16116
West T, Sojo V, Pomiankowski A, Lane N (2017) The origin of heredity in protocells. Philos Trans R Soc Lond B Biol Sci 372:20160419
Williams TA, Szollosi GJ, Spang A, Foster PG, Heaps SE, Boussau B, Ettema TJG, Embley TM (2017) Integrative modeling of gene and genome evolution roots the archaeal tree of life. Proc Natl Acad Sci U S A 114:E4602–E4611
Yang S, Doolittle RF, Bourne PE (2005) Phylogeny determined by protein domain content. Proc Natl Acad Sci U S A 102:373–378
Zhaxybayeva O, Lapierre P, Gogarten JP (2005) Ancient gene duplications and the root(s) of the tree of life. Protoplasma 227:53–64
Acknowledgements
ADG gratefully acknowledges support from NASA grant 80NSSC19M0069, the Joint NASA-NSF Ideas Lab on the “Origins of Life” (NSF Solicitation 16-570), and the National Science Foundation grant MRI1427949.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
ADG is an unpaid member of the Journal of Molecular Evolution Editorial Board (Senior Editor for Reviews and Perspectives). Otherwise, the authors have no other conflicts of interest to declare.
Additional information
Handling editor: Arturo Becerra.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kailing, F., Lieberman, J., Wang, J. et al. Evolution of Cellular Organization Along the First Branches of the Tree of Life. J Mol Evol (2024). https://doi.org/10.1007/s00239-024-10188-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00239-024-10188-7