Abstract
While microbial communities in limestone caves across the world are relatively understood, knowledge of the microbial composition in lava tubes is lagging behind. These caves are found in volcanic regions worldwide and are typically lined with multicolored microbial mats on their walls and ceilings. The Mount Etna (Sicily, S-Italy) represents one of the most active volcanos in the world. Due to its outstanding biodiversity and geological features, it was declared Natural Heritage of Humanity by the UNESCO in 2013. Despite the presence of more than 200 basaltic lava tubes, the microbial diversity of these hypogean systems has never been investigated so far. Here, we investigated bacterial communities in four lava tubes of Mount Etna volcano. Field emission scanning electron microscopy (FESEM) was carried out for the morphological characterization and detection of microbial features. We documented an abundant presence of microbial cells with different morphotypes including rod-shaped, filamentous, and coccoidal cells with surface appendages, resembling actinobacteria reported in other lava tubes across the world. Based on 16S rRNA gene analysis, the colored microbial mats collected were mostly composed of bacteria belonging to the phyla Actinomycetota, Pseudomonadota, Acidobacteriota, Chloroflexota, and Cyanobacteria. At the genus level, the analysis revealed a dominance of the genus Crossiella, which is actively involved in biomineralization processes, followed by Pseudomonas, Bacillus, Chujaibacter, and Sphingomonas. The presence of these taxa is associated with the carbon, nitrogen, and ammonia cycles, and some are possibly related to the anthropic disturbance of these caves. This study provides the first insight into the microbial diversity of the Etna volcano lava tubes, and expands on previous research on microbiology of volcanic caves across the world.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the last 20 years, considerable effort has been made to shed light on the microbial communities of karst caves, particularly speleothems and walls hosting wall paintings [1, 2]. In contrast, the microbiology of lava tubes has received much less attention.
Microbial communities in lava tubes grow forming extensive colored biofilms on speleothems and walls [3], similarly to what happens in karst caves [4, 5]. Recent studies conducted in lava tubes revealed a highly diverse microbiome, dominated by new microbial life forms and interactions differing from those occurring on the surface [3, 6]. They can thrive in these harsh oligotrophic environments by interacting with minerals and inducing biomineralization processes [7].
When considering the composition of such colored microbial mats, evidence available in the literature showed that these colonies are mainly composed of metabolically active Actinomycetota, as revealed by cDNA analysis of yellow colonies from a lava tube in La Palma, Canary Island, Spain [8]. In general, Actinomycetota and Proteobacteria are the two most abundant groups of microorganisms in lava tubes (e.g., Northup et al. [9]; Hathaway et al. [10]). One of the most complete studies to date on Actinomycetota was carried out by Riquelme et al. [3], in volcanic caves from the USA, Canada, and Portugal, highlighting the importance of caves as a source of new species of Actinomycetota [3, 11].
While we have some understanding on geomicrobiology of lava tubes from Hawaii, New Mexico and California, USA [6, 9, 10], Azores, Portugal [12], Easter Island, Chile [7, 13], British Columbia, Canada [3], Galapagos Islands, Ecuador [14,15,16], and La Palma Island, Spain [17, 18], no data are available on the microbial communities growing in lava tubes of Mount Etna (Sicily, S-Italy). Mount Etna represents one of the most active volcanos in the world [19], proclaimed a Natural Heritage Site by the UNESCO since 2013 due to its outstanding biodiversity and unique geological features. Its volcanic activity dates back more than 500,000 years ago. From about 57,000 years ago, the intense eruptive activity formed the 3600-m-high Ellittico stratovolcano, and from about 15,000 years ago, the Mongibello volcano, whose 357 lava flows cover 88% of the entire surface of Mount Etna [20].
Over 200 basaltic lava tubes are present around the Mount Etna volcano [21]. They generally form when the outer surface of lava channels cools more rapidly forming a hardened crust, while the inside continues to flow until it finishes draining and a tunnel is generated. The viscosity of the lava flow, which is related to the chemical composition of the molten rock, dissolved gases, temperature, and flow velocity, hence controls the formation of lava tubes [22]. The surface of “aa” lava flows is fragmental, spinose, and generally clinker, whereas “pahoehoe” lava flows are smooth. The formation of either is strongly influenced by the viscosity of the lava during eruptions [23]. Although it is believed that lava tubes only form on pahoehoe lavas, the formation of caves on Etna frequently occurs in “aa” lava, and some of the tubes that are considered to have formed on pahoehoe formed in large “aa” lava flows [24]. The composition of these caves corresponds mainly to basaltic lava, formed by silicates, such as clinopyroxene, plagioclase, and olivine, in addition to iron minerals, intercalated with carbonates and opal of biogenic origin [7, 13, 22].
Despite the great scientific interest determined by the frequent eruptive events occurring on the volcano, both on the summit and lateral flanks, and the peculiarity of its flora [25] and fauna (e.g., Caruso [26]; Magrini et al. [27]; Ebejer and Nicolosi [28]), research focusing on microorganisms is still lagging behind (but see Hopkins et al. [29]; Badalamenti et al. [30]), and study focused on microorganisms living on lava tubes has never been performed so far.
Here we aimed at providing the first microbiological assessment of biofilms coating the walls of four lava tubes located in Mount Etna Park (Sicily, Italy). The characterization of microbial communities from Etna lava tubes is fundamental to the understanding of this uncharted microbial diversity, and also contributes to the preservation of these unique geoheritage sites, recently receiving major interest from society both for their scientific and touristic value [31]. Therefore, reasonable protective and scientific measures should be applied to improve their value [32].
Materials and Methods
Studied Site and Sample Collection
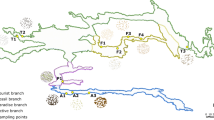
The Etna volcano is located 20 km north of the city of Catania (15° 0′ E, 37° 43.8′ N), in Sicily, Italy (Fig. 1).

Source: Google Maps [33]. B Entrance of “Grotta del Santo.” C Entrance of “Grotta Catanese I.” D General view of “Grotta Lunga.” E General view of “Grotta di Monte Corruccio”
Map of Mount Etna (Sicily, Italy) with the location of the sampled lava tubes (A).
In June 2018, we performed a sampling campaign in four lava tubes of the Etna volcano, namely “Grotta del Santo,” “Grotta Catanese I,” “Grotta Lunga,” and “Grotta di Monte Corruccio.” The main features of each cave are reported in Table S1. Replicate samples of colored microbial mats were aseptically collected using sterile scalpels and stored in sterile 1.5-ml microtubes. Each microbial mat sample was collected from an area of approximately 20 cm2. All samples were stored at 4 °C until transportation to the lab. Samples for DNA-based analysis were stored at − 80 °C until laboratory procedures were performed. Samples for microscopy observations were immediately processed upon arrival to the lab.
“Grotta del Santo” or “Grotta di San Nicola” (registered with reference SICT1032 at the “Catasto delle Grotte della Sicilia” of the “Federazione Speleologica Regionale Siciliana”) is a lava tube divided into several galleries reaching a total length of over 900 m. Lava flow here is attributed to a time interval of 15,000–3930 ± 60 years [20]. Results of bulk rock analyses for major (wt%) and trace (ppm) elements derived from Lanzafame and Ferlito [34] are presented in Table S1. The cave entrance has an altar erected in memory of Saint Nicola Politi, patron saint of Adrano (CT), who, according to tradition, lived in this place from 1134 to 1137. The cave is frequently visited, being a destination for religious pilgrimages (Fig. 1B). Although the presence of an iron gate, the cave is easily accessible. Within the lava cave galleries, four sampling sites comprising extensive yellow, white, and beige microbial mats coating the cave walls (designated GS_1A, GS_2A, GS_2B and GS_4; Table S2) were collected and analyzed.
“Grotta Catanese I” (SICT1037) is a lava cave that originated during the “Monte Arso” eruption, about 500 B.C. It is characterized by a large entrance hall, from which a narrow tunnel with about 70-m-long branches off laterally (Fig. 1C). The cave is freely accessible and occasionally frequented by local visitors. Grey and reddish colonies (designated GC1_1A and GC1_2; Table S2) were collected in the twilight zone.
“Grotta Lunga” or “Grotta di Monpeloso” (SICT1029) is a 55-m-long outflow tunnel that originated from the eruptive apparatus of “Monpeloso” formed in 300 ± 100 AD (Fig. 1D) [20]. Results of bulk rock analyses for major (wt%) and trace (ppm) elements derived from Matteoni [35] are presented in Table S1. Here we observed white and grey colonies (designated GL_1 and GL_3) along the main gallery (Table S2). The cave is freely accessible and frequently visited by tourists.
“Grotta di Monte Corruccio” (SICT1056) is an outflow tunnel partially contained in the effusive eruptive apparatus of the homonymous mount formed in a time interval of 15,000–3930 ± 60 years [20]. It has a total length of about 80 m (Fig. 1E). The cave is freely accessible and occasionally frequented by tourists. Here we observed yellow, white, and beige colonies (designated GMC_1, GMC_2, GMC_3, and GMC_4) along the cave galleries (Table S2).
Despite no information is available on the precise number of tourists entering each cave, based on an estimate of the tourist frequentation and the presence of waste due to visits, we can sort the four caves in two categories: those that experience low tourist use (“Grotta Catanese I” and “Grotta di Monte Corruccio”) and the ones experiencing higher tourist use (“Grotta del Santo” and “Grotta Lunga”).
Morphological Characterization by Microscopy Techniques
Small fragments of each sample collected in the four lava tubes were observed using a Zeiss Discovery V8 stereomicroscope (200 × maximum magnification) coupled to a MOTICAM 10.0 system to perform a detailed macroscopic examination of the sample surface.
Subsequently, samples were examined by field emission scanning electron microscopy (FESEM) using a high-resolution FEI Teneo SEM (FEI Company, Eindhoven, The Netherlands) equipped with an Oxford X-ray energy dispersive spectroscopy (EDS) detector for characterizing the microtopography of the samples and detect microbial cells.
Air-dried bulk samples with microbial mats were directly mounted on a sample stub and sputter coated with a thin platinum film, with an acceleration voltage of 5 kV, using the SE detector.
Statistical Analysis
The statistical analyses were performed in R [36]. The taxonomic richness measured at the order level for the four caves was tested by means of a generalized linear model (GLM) with a Poisson error distribution [37]. Differences among caves were then tested with the Tukey’s post hoc test, with the function “glht” from the multcomp package [38].
Taxonomic Characterization of Microbial Communities
Molecular biology techniques based on 16S rRNA gene analysis were conducted for the identification of the bacterial communities present in the Etna lava tube samples.
Genomic DNA was extracted from 12 samples collected in the Etna lava tubes using the DNeasy PowerLyzer PowerSoil Kit according to the manufacturer’s protocol (Qiagen), and quantified using a Qubit 4.0 fluorometer (Invitrogen). Sequencing libraries of the V3-V4 hypervariable region of the 16S rRNA gene were prepared according to the Illumina 16S Metagenomic Sequencing Library protocols. The gDNA input (2 ng) was amplified by PCR using the universal primer pair 314F (5′- Illumina overhang- CCTACGGGNGGCWGCAG -3′) and 805R (5′- Illumina overhang- GACTACHVGGGTATCTAATCC -3′) with Illumina adapter overhang sequences. The thermocycling conditions were as follows: 3 min at 95 °C, 30 s at 55 °C, and 30 s at 72 °C, followed by a 5-min final extension at 72 °C. The PCR products were then purified with AMPure beads (Agencourt Bioscience, Beverly, MA). Two microliters of each purified product was PCR amplified for final library construction containing the index using NexteraXT Indexed Primer, under the same thermocycling conditions as mentioned before, except for 10 cycles. After purification with AMPure beads, the final purified products were quantified using qPCR according to the qPCR Quantification Protocol Guide (KAPA Library Quantificatoin kits for Illumina Sequecing platforms) and qualified using the TapeStation D1000 ScreenTape (Agilent Technologies, Waldbronn, Germany). The purified amplicons were then sequenced using the MiSeq™ platform (Illumina, San Diego, USA) by Macrogen Sequencing Services (Korea).
The raw data obtained from Illumina platform MiSeq, for producing 300 PE reads, was initially quality checked, trimmed, and clustered in ASVs using QIIME2 [39] with DADA2 [40]. ASV is a higher-resolution analogue of the traditional OTU table, which records the number of times each exact amplicon sequence variant was observed in each sample. Taxonomic identification was carried out using SILVA database v.132. Alpha diversity metrics (Shannon’s index and Pielou’s evenness) of bacterial communities were also calculated to investigate community heterogeneity within sample diversity. The raw reads wear deposited into the NCBI Sequence Read Archive (SRA) database under the project id PRJNA914266.
Results and Discussion
Microscopy Observations
Under the stereomicroscope, samples showed colored stains coating the rock substrate or associated with mineral grains (Table S2). There were notable differences in colony morphology, texture, and size.
Microbial cells and structures were imaged by high-resolution FESEM, as an effective and fast method for microbial life detection in complex samples. FESEM images showed abundant microbial cells with different morphotypes in all the samples (Figs. 2, 3, 4, and 5). The most common forms were rod-shaped and coccoidal cells with surface appendages, resembling the actinobacterial cells reported by [3] in lava tubes from USA, Canada, Portugal, and Spain. Samples from “Grotta del Santo,” GS_1A (Fig. 2A, B, C), GS_2A (Fig. 2D), GS_2B (Fig. 2E, F), GC1_1A (Fig. 3A, B), and from “Grotta di Monte Corruccio,” GMC_5 (Fig. 4G, H) showed to be the most abundant in actinobacteria-like cells, as revealed by FESEM.
Field emission scanning electron microscopy images of “Grotta del Santo” (GS) samples. Representative FESEM images of the studied samples, depicting A biofilm of filamentous actinobacteria-like cells (GS_1A); B close-up view of the spore chains with cell surface appendages (GS_1A); C fragments of reticulated filaments sparsely distributed within the sample (GS_1A); D biofilm of actinobacterial cells with spiny ornamentation (GS_2A); E dense network of actinobacteria cells with spiny ornamentations intermingled with Actinobacteria-like hyphae, and F spore chains with spiny protuberances on their surface (GS_2B); and G, H filamentous microbial structures with smooth surfaces in close association with the mineral substrate (GS_4)
Field emission scanning electron microscopy images of “Grotta Catanese I” (GC) samples. Representative FESEM images of the studied samples, depicting A Actinobacteria-like coccoid (with 1-μm diameter) and rod-shaped cells with spiny ornamentation (GC1_1A); B filaments with hairy ornamentation, intermingled with coccoid cells (GC1_1A); and C, D clusters of the diatom Orthoseira roeseana and Humidophila (GC1_2)
Field emission scanning electron microscopy images of “Grotta di Monte Corruccio” (GMC) samples. Representative FESEM images of the studied samples, depicting A rod-shaped cells embedded in the EPS layer (GMC_1); B actinobacteria-like spores with hairy filamentous forms (GMC_1); C filamentous cells associated with mineral grains (GMC_2); D Actinobacteria-like cells with appendages (GMC_2); E microbial imprints showing cell-like structures (GMC_4); F clusters of Actinobacteria-like cells with spiny ornamentation (left) and rod-shaped bacteria with spiny ornamentation impregnated in EPS (right) (GMC_4); G dense masses of bacterial cells, mainly rod-shaped mats showing filaments covered with pili/fimbrae with spheroid shapes emerging from the ends (GMC_5); and H Ca-rich spheroids closely associated with filamentous cells (GMC_5)
Field emission scanning electron microscopy images of GL samples. Representative field emission scanning electron microscopy images of the studied samples, depicting A, B reticulated filaments embedded in a matrix of extracellular polymeric substances (GL_1) and C, D Actinobacteria-like filamentous forms (GL_3)
FESEM observations showed an abundant presence of actinobacteria-like cells for the four samples collected in “Grotta del Santo,” mainly comprising spores with a hairy surface (Fig. 2A–C) or chains of spores with spiny surfaces (Fig. 2D–F). Actinobacteria have been frequently reported in karstic and volcanic caves worldwide (e.g., Riquelme et al. [3]; Porca et al. [41]; Axenov-Gribanov et al. [42]). These bacteria grow from the hyphal, which is an important basis for classification and comparable to filamentous fungi [43]. Other filamentous bacterial structures with smooth surfaces were also found, particularly in sample GS_4, in close association with the mineral substrate (Fig. 2G, H).
Sample collected in “Grotta Catanese I” (GC1_1A) revealed the presence of several actinobacteria-like coccoid (1-µm diameter) and rod-shaped cells (Fig. 3A) with spiny ornamentation, as well as filaments with hairy ornamentation, intermingled with coccoid cells (Fig. 3B). In contrast, sample GC1_2 from the same cave showed a prevalence of diatoms, specifically Orthoseira roeseana (Fig. 3C) and Humidophila (Fig. 3D), which is related to its location near the cave entrance. It is well documented that at the entrance of caves and on artificially illuminated cave walls, phototrophic organisms mainly cyanobacteria, green algae, and diatoms develop with increasing moisture [44]. In this study, samples of whitish, beige, grey, or yellow bacterial mats that often form on the middle or dark zones of caves were selected and collected, tentatively avoiding phototrophic-based biofilms located at the cave entrances.
Samples collected in “Grotta di Monte Corruccio” showed greater variety in microbial structures and cell morphologies (Fig. 4). In the yellow mat sample (GMC_1), a biofilm of rod-shaped cells, embedded in a matrix of extracellular polymeric substances (EPS) was clearly observed by FESEM (Fig. 4A). Biofilms are multicellular microbial populations that typically adhered to solid surfaces due to the release of EPS, which self-encapsulate the cells and provide structure to biofilms [45]. They comprise a survival strategy of microbial cells to thrive in these hostile environments. Actinobacteria-like spores with hairy ornamentation and spiny surfaces were also observed in all samples collected in this cave (Fig. 4B–H). Sample GMC_4 revealed the presence of microbial imprints, suggesting that the cell-like shapes occur within internal laminae (Fig. 4E), as well as the presence of some clusters of Actinobacteria-like cells and rod-shaped bacteria impregnated in EPS (Fig. 4F). A tangled mass of actinobacteria-like hyphae or archaeal-like cells with hami that protrude from their cell surfaces was observed by Perras et al. [46].
Observations conducted on sample GMC_5 revealed the presence of Ca-rich spheroids closely associated with filamentous cells (Fig. 4G, H), resembling the CaCO3 microspheres found in Kipuka Kanohina lava cave in Hawaii, USA [3]. Studies on cave microbial communities revealed that actinobacteria may induce biomineralization processes, promoting nucleation sites or changes in pH [3, 47]. Precipitation of carbonates can be induced by bacteria via urea hydrolysis [48] or through the uptake of carbon dioxide promoting changes in pH and followed by mineral precipitation and CaCO3 formation [47]. Numerous biogenic minerals have been reported also in lava caves. Calcite and Mg–silicate minerals were found associated with actinobacterial morphologies on coralloid type speleothems from Chile [7]. Calcium carbonate spheres closely related to dense networks of hairy filaments were observed within the colored microbial mats from Azorean, Canarian, and Hawaiian [3]. Silica microspheres embedded in EPS matrix were observed in Galapagos Islands [16].
Interestingly, sample from “Grotta Lunga” (GL_1) showed the presence of reticulated filaments embedded in a matrix of EPS (Fig. 5A, B). The presence of reticulated filaments has been frequently reported in caves worldwide, in both lava and limestone caves [49,50,51,52] but their nature still results enigmatic for microbiologists [54]. FESEM observations of the sample GL_3 revealed a variety of actinobacteria-like filamentous forms spread all over the sample (Fig. 5C, D).
Richness and Diversity of Cave Microbial Communities
We identified a total of 6310 unique ASVs for the 12 samples, with the samples collected in “Grotta Lunga” being richer than the others (GL_3 = 1190 ASVs; GL_1 = 1057 ASVs). These samples also showed the highest values in terms of alpha diversity indices, i.e., Shannon index [54] and Pielou’s evenness [55] (Table 1), calculated on the ASVs (Fig. 6).
When considering the taxonomic richness measured at the order level, the “Grotta Lunga” resulted the richest cave with 174 orders, followed by “Grotta Catanese I” with 150 orders, and then by “Grotta del Santo” and “Grotta di Monte Corruccio” caves with 148 and 147 orders, respectively (Fig. 7). The result of the GLM showed a significant effect of the cave identity on the taxonomic richness measured at the order level (Chi = 46.4; P < 0.001). By comparing the differences among caves with the Tukey’s post hoc test, we could highlight that most of this effect is due to the significant difference between the “Grotta Lunga” and the others, with the former being significantly richer than the others (Lunga – Catanese: z = 2.72, P = 0.034; Lunga – Santo: z = 6.41, P < 0.001; Lunga – Corruccio: z = 5.59, P < 0.001). Regarding the other pairwise comparisons, we could record a significant difference between “Grotta Catanese I” and “Grotta del Santo” (z = 3.15, P = 0.009), while no other significant differences were identified (|z|< 2.34; P > 0.05).
Due to the heterogeneity of the examined caves, the small number of samples and considering the pioneer nature of this study, specific patterns are hard to discern, as well as possible relations between the observed parameters of richness and abundance and any environmental factor characterizing the caves we studied. In general terms, major values of richness were observed in “Grotta Lunga” being characterized by low elevation (850 m) and high temperature (18.7 °C). On the contrary, lower values of richness were observed in “Grotta di Monte Corruccio,” which is located at 1365 m, with a temperature of 13.9 °C. Tentatively, differences among caves in terms of richness could also be explained by the different environmental conditions and biogeochemical processes operating in caves [56] as well as a possible response of the microbial communities to human disturbance [57]. Accordingly, richness was higher in “Grotta Lunga” and “Grotta Catanese I” experiencing low tourist use. Similar results were obtained by Ikner et al. [58] reporting lower diversity in caves highly exploited for touristic purposes.
Microbial Communities
Independent libraries of DNA sequences from each sample were built targeting the V3-V4 hypervariable regions of the 16S rRNA gene, with the objective of detecting the total bacteria present in these samples.
The relative abundances of the dominant phyla in each cave sample are shown in Fig. 8. Most of the identified bacteria belonged to Actinomycetota, Pseudomonadota, Acidobacteriota, Chloroflexota, and Cyanobacteria, followed by other phyla with lower representativeness. Taxonomic identification resulted in a prominent importance of the phylum Actinomycetota in almost all samples, ranging from 78.4% in GS_2B to 14.3% in GC1_2, corroborating the observations by FESEM.
The presence of Actinomycetota has been widely found in caves where are actively involved in the biomineralization processes (e.g., Riquelme et al. [3]; Barton et al. [59]). Moreover, members of the phylum Actinomycetota represent a promising source of bioactive metabolites for drug development [60, 61].
The phylum Pseudomonadota was also present in each cave, being the most representative group in “Grotta del Santo” and “Grotta Lunga” (GS_4, GL_1, and GL_3 with 48.97%, 44.43%, and 42.88%, respectively). The Pseudomonadota are well represented in lava caves across the world [62], including New Mexico, Hawaii, Azores, and Galapagos Islands [9, 11, 15].
A similar trend was observed by Gonzalez-Pimentel et al. [8] and Miller et al. [15], where the phyla Actinomycetota and Pseudomonadota were among the most representative groups in lava tubes from La Palma (Canary Islands) and Galápagos (Equador), respectively.
The phylum Acidobacteriota was equally distributed in all caves, ranging from 3.02% in “Grotta di Monte Corruccio” (sample GMC_5) and 12.39% in “Grotta del Santo” (sample GS_2A). Several studies have detected the presence of acidobacterial 16S rRNA gene sequences in caves (e.g., Holmes et al. [63]; Hutchens et al. [64]; Chelius and Moore [65]; Engel et al. [66]). However, their role is still poorly understood [67].
The phylum Cyanobacteria was abundant in Grotta Catanese I but poorly represented in the other caves. The presence of this taxon in sample GC1_2 is due to its proximity to the cave entrance, which receives sunlight allowing the development of photosynthetic-based biofilms on the cave wall. Accordingly, Cyanobacteria are mainly favored by the presence of light and thereby generally frequent near the cave entrance [68, 69], although the presence of artificial light can promote their growth in the innermost cave zones (e.g., Piano et al. [70]), which is not the case. Phyla with less representativeness, but with at least 5% of abundance, were Chloroflexota, Planctomycetota, Bacillota, Nitrospirota, Bacteroidota, Patescibacteria, and Verrucomicrobiota. They are also common at lower rates in other lava tubes [3, 6, 9]. Other phyla with relative abundances < 1% were also retrieved.
The bacterial communities identified at the class level showed differences between caves and sectors. Most bacteria belonged to the following class: Actinomycetes, Gammaproteobacteria, Alphaproteobacteria, Oxyphotobacteria, followed by the less abundant Bacilli, Bacteroidia, and Nitrospira (Fig. 9).
The relative abundance of the different classes varied in each sample, suggesting differences in the local environment and element composition of the volcanic substrate [71]. Actinomycetes was the most abundant class in “Grotta del Santo” (GS_2B: 78.21%) and “Grotta di Monte Corruccio” (GMC_5: 72.76%; GMC_1: 62.94%) sample, while Gammaproteobacteria was predominant in “Grotta Lunga” (GL_1: 28.60%) and “Grotta del Santo” (GS_4: 25.99%). Oxyphotobacteria were instead the most abundant class in “Grotta Catanese I” (GC1_2). The class Alphaproteobacteria was also well represented in all samples, ranging from 22.37% in “Grotta del Santo” GS_4 and 4.66% in “Grotta Catanese I” (GC1_2). The class Actinomycetes is common in caves and its studies in different locations have revealed the presence of several novels and rare taxon [3, 8, 57]. Investigations on this taxon have highlighted its biotechnological relevance [72,73,74] as well as their importance as potential pathogens (e.g., Jurado et al. [75]; Gonzalez-Pimentel et al. [8]; Buresova et al. [76]).
The class Gammaproteobacteria is among the most dominant group in habitats with either natural or anthropogenic organic inputs [77]. Accordingly, it seems to represent a good bioindicator to detect the presence of contaminants in soil [78,79,80] or caves [81]. This taxon was abundant in samples GL_1 (28.69%), GL_3 (24.65%) from “Grotta Lunga” and GS_4 (25.99) from “Grotta del Santo,” which represent “high tourist use” caves. The presence of pathogens in caves has rarely been discussed and evidence of contamination still results scarce (but see Luong et al. [82]). Among pathogens frequently detected in caves, considered indicators of human impact, there are Bacillus spp., Escherichia coli, and Staphylococcus aureus [81, 83, 84].
The bacterial communities identified at the genus level showed differences between caves and sectors. The genus Crossiella is well present in all samples, being the most abundant group in 8 out of 12 analyzed samples (Fig. 8). The taxon was abundant in sample GS_2B from “Grotta del Santo,” and GMC_1 and GMC_5 from “Grotta di Monte Corruccio” with a relative abundance respectively of 77.6%, 71.8%, and 62.54%. The genus is common in lava tube caves of Hawaii and Azores [3, 11, 85] and also in limestone caves [86]. Gonzalez-Pimentel [17] observed a high abundance of Crossiella in moonmilk from La Palma lava caves. Recent studies have hypothesized the influence of Crossiella on the nitrogen cycle and its possible contribution to calcium carbonate precipitation in caves [87, 88].
Some important groups after Crossiella were the genus Pseudomonas, Bacillus, Chujaibacter, and Sphingomonas. Nitrospira and the family groups Beijerinckiaceae and Nitrosococcaceae are respectively described as nitrite-oxidizing, nitrogen fixation, and ammonia-oxidizing bacteria, relatively common in caves [6, 9, 11, 89]. Chujaibacter is associated with heavy metal metabolism [90]. The information provided by the identification of these groups of bacteria could respond to the presence of the influence of vegetation and anthropic pressure on the studied caves.
The presence of Bacillus, Pseudomonas, and Sphingomonas in caves is often associated with the presence of high human impact (e.g., Lavoie and Northup [83]; Ikner et al. [58]; Bastian et al. [81]) and recent studies have showed their antimicrobial activity against pathogenic bacteria, as well as a potential source of new microorganism [91, 92]. Other genera identified were as follows: Alkanibacter, Flavobacterium, Steroidobacter, Sporosarcina, wb1-P19, Polaromonas, this last in clone libraries and culture collections from polar and high-elevation environments [93].
Conclusions
This study provides the first insight into the taxonomic groups constituting the microbial communities of Mount Etna lava tubes. Although the limited number of samples did not allow us to properly correlate these data with any environmental variables, we could detect the higher richness in “Grotta Lunga,” which is located at a lower elevation (850 m a.s.l.), whereas we observed a general lower richness at higher elevation (1365 m a.s.l.) in “Grotta di Monte Corruccio.” Accordingly, differences among caves in terms of richness could be explained by the different environmental conditions and biogeochemical processes operating in caves [55]. Further investigations would be therefore desirable to shed light on which factors drive the richness of microbial communities on Mount Etna lava caves.
Our results revealed the abundant presence of phylotypes previously detected in other lava tubes worldwide. The presence of a large number of unclassified bacteria in the 12 sampling sites suggests that these lava tubes have a great potential for the isolation of novel species. However, the land use overlying the cave, as well as the uncontrolled presence of visitors into these caves, may pose major threats to these ecosystems, especially those located at lower altitudes beyond the strictly protected areas of the volcano, generally exposed to a higher tourist frequentation. Further research is needed to ascertain the possible presence of novel bacteria in caves from the Etna lava caves, as well as the presence of contaminants and their potential risk associated with human health. Data of pathogens should be therefore considered not only for the conservation of these unique geosites, but also for the risk associated with human health.
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Saiz-Jimenez C, Cuezva S, Jurado V, Fernandez-Cortes A, Porca E, Benavente D, Cañaveras JC, Sanchez-Moral S (2011) Paleolithic art in peril: policy and science collide at Altamira Cave. Science 334:42–43
Saiz-Jimenez C, Miller AZ, Martin-Sanchez PM, Hernandez-Marine M (2012) Uncovering the origin of the black stains in Lascaux Cave in France. Environ Microbiol 14:3220–3231
Riquelme C, Hathaway JJM, Dapkevicius MLNE, Miller AZ, Kooser A, Northup DE, Jurado V, Fernandez O, Saiz-Jimenez C, Cheeptham N (2015) Actinobacterial diversity in volcanic caves and associated geomicrobiological interactions. Front Microbiol 6:1342. https://doi.org/10.3389/fmicb.2015.01342
Addesso R, Bellino A, D’Angeli IM, De Waele J, Miller AZ, Carbone C, Baldantoni D (2019) Vermiculations from karst caves: the case of Pertosa-Auletta system (Italy). CATENA 182:104178
Addesso R, Gonzalez-Pimentel JL, D’Angeli IM, De Waele J, Saiz-Jimenez C, Jurado V, Miller AZ, Cubero B, Vigliotta G, Baldantoni D (2021) Microbial community characterizing vermiculations from karst caves and its role in their formation. Microb Ecol 81:884–896
Lavoie KH, Winter AS, Read KJH, Hughes EM, Spilde MN, Northup DE (2017) Comparison of bacterial communities from lava cave microbial mats to overlying surface soils from Lava Beds National Monument, USA. Plos One 12:e0169339. https://doi.org/10.1371/journal.pone.0169339
Miller AZ, Pereira MF, Calaforra JM, Forti P, Dionísio A, Saiz-Jimenez C (2014) Siliceous speleothems and associated microbe-mineral interactions from Ana Heva Lava Tube in Easter Island (Chile). Geomicrobiol J 31:236–245
Gonzalez-Pimentel JL, Miller AZ, Jurado V, Laiz L, Pereira MFC, Saiz-Jimenez C (2018) Yellow coloured mats from lava tubes of La Palma (Canary Islands, Spain) are dominated by metabolically active Actinobacteria. Sci Rep 8:1–11. https://doi.org/10.1038/s41598-018-20393-2
Northup DE, Melim LA, Spilde MN, Hathaway JJM, Garcia MG, Moya M, Stone FD, Boston PJ, Dapkevicius MLNE, Riquelme C (2011) Lava cave microbial communities within mats and secondary mineral deposits: implications for life detection on other planets. Astrobiology 11:601–618
Hathaway JJM, Garcia MG, Balasch MM, Spilde MN, Stone FD, Dapkevicius MLNE, Amorim IR, Gabriele R, Borges PAV, Northup DE (2014) Comparison of bacterial diversity in Azorean and Hawaiian lava cave microbial mats. Geomicrobiol J 31:205–220
Riquelme C, Rigal F, Hathaway JJM, Northup DE, Spilde MN, Borges PAV, Gabriel R, Amorim IR, Dapkevicius MDLNE (2015) Cave microbial community composition in oceanic islands: disentangling the effect of different colored mats in diversity patterns of Azorean lava caves. FEMS Microb 91:fiv141
Ríos ADL, Bustillo MA, Ascaso C, Carvalho MDR (2011) Bioconstructions in ochreous speleothems from lava tubes on Terceira Island (Azores). Sediment Geol 236:117–128
Miller AZ, José M, Jiménez-Morillo NT, Pereira MF, González-Pérez JA, Calaforra JM, Saiz-Jimenez C (2016) Analytical pyrolysis and stable isotope analyses reveal past environmental changes in coralloid speleothems from Easter Island (Chile). J Chromatogr A 1461:144–152
Miller AZ, Garcia-Sanchez AM, Pereira MF, Gazquez F, Calaforra JM, Forti P, Toulkeridis T, Martínez-Frías J, Saiz-Jimenez C (2016b) Biomineralization and biosignatures of coralloid-type speleothems from lava tubes of Galapagos Islands: evidences on the fossil record of prokaryotes. EGU General Assembly Conference, 23–28 April 2017, Vienna, Austria. EPSC2016–16574
Miller AZ, García-Sánchez AM, Coutinho ML, Costa Pereira MF, Gázquez F, Calaforra JM, Forti P, Martínez-Frías J, Toulkeridis T, Caldeira AT, Saiz-Jimenez C (2020) Colored microbial coatings in show caves from the Galapagos Islands (Ecuador): first microbiological approach. Coatings 10:1134
Miller AZ, Jiménez-Morillo NT, Coutinho ML, Gazquez F, Palma V, Sauro F, Pereira MFC, Rull F, Toulkeridis T, Caldeira AT, Forti P, Calaforra JM (2022) Organic geochemistry and mineralogy suggest anthropogenic impact in speleothem chemistry from volcanic show caves of the Galapagos. iScience 25:104556. https://doi.org/10.1016/j.isci.2022.104556
Gonzalez-Pimentel JL (2019) Microorganismos de las cuevas volcánicas de La Palma (Islas Canarias). Diversidad y potencial uso biotecnológico [dissertation]. University Pablo de Olavide, Seville
Gonzalez-Pimentel JL, Martin-Pozas T, Jurado V, Miller AZ, Caldeira AT, Fernandez-Lorenzo O, Sanchez-Moral S, Saiz-Jimenez C (2021) Prokaryotic communities from a lava tube cave in La Palma Island (Spain) are involved in the biogeochemical cycle of major elements. PeerJ 9:e11386
Branca S, Del Carlo P (2004) Eruptions of Mt Etna during the past 3.200 years: a revised compilation integrating the historical and stratigraphic records. Mt. Etna: volcano laboratory. In: Bonaccorso A, Calvari S, Coltelli M, Del Negro C, Falsaperla S (eds) Etna volcano laboratory. AGU (Geophysical monograph series), Washington, DC 143:384
Branca S, Coltelli M, Groppelli G, Lentini F (2011) Geological map of Etna volcano, 1:50,000 scale. Ital J Geosci 130:265–291
Centro Speleologico Etneo (1999) Dentro il vulcano. Le grotte dell’Etna. Parco dell’Etna, Nicolosi, Italy
Forti P (2005) Genetic processes of cave minerals in volcanic environments: an overview. J Cave Karst Stud 67:3–13
Macdonald GA (1953) Pahoehoe, aa, and block lava. Am J Sci 251:169–191
Calvari S, Pinkerton H (1999) Lava tube morphology on Etna and evidence for lava flow emplacement mechanisms. J Volcanol Geotherm Res 90:263–280
Sciandrello S, Minissale P, Del Galdo GG (2020) Vascular plant species diversity of Mt. Etna (Sicily): endemicity, insularity and spatial patterns along the altitudinal gradient of the highest active volcano in Europe. Peer J 8:e9875
Caruso D (1999) La fauna delle grotte dell’Etna: descrizione e considerazioni. In: Proceedings of the IXth International Symposium on Vulcanospeleology, 11–19 September 1999, Catania, Italy
Magrini P, Baviera C, Taglianti AV (2006) Note sul genere Duvalius in Sicilia con descrizione di due nuove specie (Coleoptera, Carabidae). Fragm Entomol 38:33–53
Ebejer MJ, Nicolosi G (2022) New records of acalyptrate Diptera from Sicily (Brachycera, Muscomorpha: Asteiidae, Aulacigastridae, Carnidae, Lonchaeidae, Odiniidae, Pallopteridae, Periscelididae, Piophilidae, Sciomyzidae, Ulidiidae). Fragm Entomol 54:101–104
Hopkins DW, Badalucco L, English LC, Meli SM, Chudek JA, Ioppolo A (2007) Plant litter decomposition and microbial characteristics in volcanic soils (Mt Etna, Sicily) at different stages of development. Biol Fertil Soils 43:461–469
Badalamenti E, Catania V, Sofia S, Sardina MT, Sala G, La Mantia T, Quatrini P (2021) The root mycobiota of Betula aetnensis Raf., an endemic tree species colonizing the lavas of Mt Etna (Italy). Forests 12:1624
Cigna AA (2016) Tourism and show caves. Z Geomorphol 60:217–233
Antić A, Peppoloni S, Di Capua G (2020) Applying the values of geoethics for sustainable speleotourism development. Geoheritage 12:1–9. https://doi.org/10.1007/s12371-020-00504-0
Maps G (2018) Google maps. https://www.google.it/maps/. Accessed 5 July 2022
Lanzafame G, Ferlito C (2014) Degassing driving crystallization of plagioclase phenocrysts in lava tube stalactites on Mount Etna (Sicily, Italy). Lithos 206:338–347
Matteoni R (2013) Detecting small-scale geochemical and petrological variations within historical and modern lavas from Mt Etna [dissertation]. University of Pisa, Pisa
RC Team (2020) R: a language and environment for statistical computing, R foundation for statistical computing, Vienna, Austria. https://www.R-project.org. Accessed 1 June 2022
Zuur AF, Ieno EN, Walker NJ, Savaliev AA, Smith GM (2009) Mixed effect models and extensions in ecology with R. Springer, Berlin, p 574
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biom J 50:346–363
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Porca E, Jurado V, Žgur-Bertok D, Saiz-Jimenez C, Pašić L (2012) Comparative analysis of yellow microbial communities growing on the walls of geographically distinct caves indicates a common core of microorganisms involved in their formation. FEMS Microbiol 81:255–266. https://doi.org/10.1111/j.1574-6941.2012.01383.x
Axenov-Gribanov DV, Voytsekhovskaya IV, Tokovenko BT, Protasov ES, Gamaiunov SV, Rebets YV et al (2016) Actinobacteria isolated from an underground lake and moonmilk speleothem from the biggest conglomeratic karstic cave in Siberia as sources of novel biologically active compounds. Plos One 11:e0149216. https://doi.org/10.1371/journal.pone.0149216
Li Q, Chen X, Jiang Y, Jiang C (2016) Morphological identification of actinobacteria. In: Dhanasekaran D, Jiang Y (eds) Actinobacteria - basics and biotechnological applications. InTech, India, pp 59–86. https://doi.org/10.5772/61461
Falasco E, Ector L, Isaia M, Wetzel CE, Hoffmann L, Bona F (2014) Diatom flora in subterranean ecosystems: a review. Int J Speleol 43:231–251. https://doi.org/10.5038/1827-806X.43.3.1
Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM (1995) Microbial biofilms. Annu Rev Microbiol 49:711–745. https://doi.org/10.1146/annurev.mi.49.100195.003431
Perras AK, Wanner G, Klingl A et al (2014) Grappling archaea: ultrastructural analyses of an uncultivated, cold-loving archaeon, and its biofilm. Front Microbiol 5:397
Cuezva S, Fernandez-Cortes A, Porca E et al (2012) The biogeochemical role of Actinobacteria in Altamira cave, Spain. FEMS Microbiol 81:281–290
Kosznik-Kwaśnicka K, Golec P, Jaroszewicz W et al (2022) Into the unknown: microbial communities in caves, their role, and potential use. Microorganisms 10:222
Northup DE, Barns SM, Yu LE, Spilde MN, Schelble RT, Dano KE, Crossey LJ, Connolly CA, Boston PJ, Natvig DO, Dahm CN (2003) Diverse microbial communities inhabiting ferromanganese deposits in Lechuguilla and Spider Caves. Environ Microbiol 5:1071–1086
Melim LA, Northup DE, Spilde MN, Jones B, Boston PJ, Bixby RJ (2008) Reticulated filaments in cave pool speleothems: microbe or mineral? J Caves Karst Stud 70:135–141
Jones B (2009) Cave pearls – the integrated product of abiogenic and biogenic processes. J Sediment Res 79:689–710
Jones B (2011) Biogenicity of terrestrial oncoids formed in soil pockets, Cayman Brac, British West Indies. Sediment Geol 236:95–108
Miller AZ, Hernández-Mariné M, Jurado A, Dionísio V, Barquinha P, Fortunato E, Afonso MJ, Chaminé HI, Saiz-Jimene C (2012) Enigmatic reticulated filaments in subsurface granite. Environ Microbiol Rep 4:596–603
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Pielou EC (1966) The measurement of diversity in different types of biological collections. J Theor Biol 13:131–144
Cuezva S, Sanchez-Moral S, Saiz-Jimenez C, Cañaveras JC (2009) Microbial communities and associated mineral fabrics in Altamira Cave. Spain Int J Speleol 38:9
Mammola S, Di Piazza S, Ziotti M, Badino G, Isaia M (2017) Human-induced alterations of the mycobiota in an alpine show cave (Italy, SW-Alps). Acta Carsologica 46(1):111–123. https://doi.org/10.3986/ac.v46i1.2531
Ikner LA, Toomey RS, Nolan G, Neilson JW, Pryor BM, Maier RM (2007) Culturable microbial diversity and the impact of tourism in Kartchner Caverns, Arizona. Microb Ecol 53:30–42
Barton HA, Spear JR, Pace NR (2001) Microbial life in the underworld: biogenicity in secondary mineral formations. Geomicrobiol J 18:359–368
Riquelme C, Dapkevicius MDLE, Miller AZ, Charlop-Powers Z, Brady S, Mason CT, Cheeptham N (2017) Biotechnological potential of Actinobacteria from Canadian and Azorean volcanic caves. Appl Microbiol Biotechnol 101:843–857
Rangseekaew P, Pathom-Aree W (2019) Cave actinobacteria as producers of bioactive metabolites. Front Microbiol 10:387
Saiz-Jimenez C (2015) The microbiology of show caves, mines tunnels and tombs: implications for management and conservation. In: Engel AS (ed) Microbial life of cave systems. DeGruiter, Berlin, pp 231–261
Holmes AJ, Tujula NA, Holley M, Contos A, James JM, Rogers P, Gillings MR (2001) Phylogenetic structure of unusual aquatic microbial formations in Nullarbor caves, Australia. Environ Microbiol 3:256–264
Hutchens E, Radajewski S, Dumont MG, McDonald IR, Murrell JC (2004) Analysis of methanotrophic bacteria in Movile Cave by stable isotope probing. Environ Microbiol 6:111–120
Chelius MK, Moore JC (2004) Molecular phylogenetic analysis of archaea and bacteria in Wind Cave, South Dakota. Geomicrobiol J 21:123–134
Engel AS, Porter ML, Stern LA, Quinlan S, Bennett PC (2004) Bacterial diversity and ecosystem function of filamentous microbial mats from aphotic (cave) sulfidic springs dominated by chemolithoautotrophic ‘Epsilonproteobacteria.’ FEMS Microbiol 51:31–53
Meisinger DB, Zimmermann J, Ludwig W et al (2007) In situ detection of novel Acidobacteria in microbial mats from a chemolithoautotrophically based cave ecosystem (Lower Kane Cave, WY, USA). Environ Microbiol 9:1523–1534
Popović S, Subakov Simić G, Stupar M, Unković N, Predojević D, Jovanović J, Ljaljević Grbić M (2015) Cyanobacteria, algae and microfungi present in biofilm from Božana Cave (Serbia). Int J Speleol 44:4
Popović S, Krizmanić J, Vidaković D, Jakovljević O, Trbojević I, Predojević D, Vidović M, Simić S (2020) Seasonal dynamics of cyanobacteria and algae in biofilm from the entrance of two caves. Geomicrobiol J 37:315–326
Piano E, Nicolosi G, Isaia M (2021) Modulating lighting regime favours a sustainable use of show caves: a case study in NW-Italy. J Nat Conserv 64:126075
Gomez-Alvarez V, King GM, Nüsslein K (2007) Comparative bacterial diversity in recent Hawaiian volcanic deposits of different ages. FEMS Microbiol 60:60–73
Bhullar K, Waglechner N, Pawlowski A, Koteva K, Banks ED, Johnston MD, Barton HA, Wright GD (2012) Antibiotic resistance is prevalent in an isolated cave microbiome. Plos One 7:e34953
Hamedi J, Kafshnouchi M, Ranjbaran M (2019) A study on actinobacterial diversity of Hampoeil cave and screening of their biological activities. Saudi J Biol Sci 26:1587–1595
Syiemiong D, Jha DK (2019) Antibacterial potential of actinobacteria from a limestone mining site in Meghalaya, India. J Pure Appl Microbiol 13:789–802
Jurado V, Laiz L, Rodriguez-Nava V, Boiron P, Hermosin B, Sanchez-Moral S, Saiz-Jimenez C (2010) Pathogenic and opportunistic microorganisms in caves. Int J Speleol 39:2
Buresova A, Kopecky J, Sagova-Mareckova M, Alonso L, Vautrin F, Moenne-Loccoz Y, Nava VR (2022) Comparison of Actinobacteria communities from human-impacted and pristine karst caves. Microbiologyopen 11:e1276
Jones D, Lavoie K, Barton HA et al (2015) Microbial life of cave systems, vol 3. Walter de Gruyter GmbH & Co KG, Boston
Lors C, Ryngaert A, Périé F, Diels L, Damidot D (2010) Evolution of bacterial community during bioremediation of PAHs in a coal tar contaminated soil. Chemosphere 81:1263–1271
Lors C, Damidot D, Ponge JF, Périé F (2012) Comparison of a bioremediation process of PAHs in a PAH-contaminated soil at field and laboratory scales. Environ Pollut 165:11–17
Niepceron M, Martin-Laurent F, Crampon M, Portet-Koltalo F, Akpa-Vinceslas M, Legras M, Bru D, Bureau F, Bodilis J (2013) GammaProteobacteria as a potential bioindicator of a multiple contamination by polycyclic aromatic hydrocarbons (PAHs) in agricultural soils. Environ Pollut 180:199–205
Bastian F, Alabouvette C, Saiz-Jimenez C (2009) Bacteria and free-living amoeba in the Lascaux Cave. Res Microbiol 160:38–40
Luong M-L, Békal S, Vinh DC, Lauzon D, Leung V, Al-Rawahi GN, Ng B, Tamara Burdz T, Bernard K (2008) First report and characterization of Aurantimonas altamirensis in clinical samples. J Clin Microbiol 46:2435e2437
Lavoie KH, Northup DE (2006) Bacteria as indicators of human impact in caves. In: Proceedings 17th National Cave and Karst Management Symposium, 31 October – 4 November 2016, 2005NICKMS Steering Committee Albany, New York, pp 40–47
Davis MC, Messina MA, Nicolosi G, Petralia S, Baker MD, Mayne CKS, Dinon CM, Moss CJ, Onac BP, Garey JR (2020) Surface runoff alters cave microbial community structure and function. Plos One 15:e0232742
Spilde MN, Northup DE, Caimi NA, Boston PJ, Stone FD, Smith S (2016) Microbial mat communities in Hawaiian lava caves. In: Proceedings of the XVII International Symposium on Vulcanospeleology, Ocean View Community Center, Ocean View, Hawaii, 6–12 February 2016
Wiseschart A, Mhuantong W, Tangphatsornruang S, Chantasingh D, Pootanakit K (2019) Shotgun metagenomic sequencing from Manao-Pee cave, Thailand, reveals insight into the microbial community structure and its metabolic potential. BMC Microbiol 19:144
Omoregie AI, Ong DEL, Nissom PM (2019) Assessing ureolytic bacteria with calcifying abilities isolated from limestone caves for biocalcification. Lett Appl Microbiol 68:173–181
Enyedi NT, Makk J, Kótai L, Berényi B, Klébert S, Sebestyén Z, Molnár Z, Borsodi AK, Leél-Össy S, Demény A, Németh P (2020) Cave bacteria-induced amorphous calcium carbonate formation. Sci Rep 10:8696. https://doi.org/10.1038/s41598-020-65667-w
Tomczyk-Żak K, Zielenkiewicz U (2016) Microbial diversity in caves. Geomicrobiol J 33:20–38
Kim SJ, Ahn JH, Weon HY, Hong SB, Seok SJ, Kim JS, Kwon SW (2015) Chujaibacter soli gen. nov., sp. nov., isolated from soil. J Microbiol 53:592–597
Yasir M (2018) Analysis of bacterial communities and characterization of antimicrobial strains from cave microbiota. Brazilian J Microbiol 49:248–257
Nakaew N, Pathom-aree W, Lumyong S (2009) Generic diversity of rare actinomycetes from Thai cave soils and their possible use as new bioactive compounds. Actinomycetologica 23:21–26 ([Google Scholar])
Darcy JL, Lynch RC, King AJ, Robeson MS, Schmidt SK (2011) Global distribution of Polaromonas phylotypes-evidence for a highly successful dispersal capacity. Plos One 6(8):e23742
Acknowledgements
We thank the “Ente Parco dell’Etna” that authorized and allowed the carrying out of the scientific research activity in the Park.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was supported by the Spanish Ministry of Science and Innovation (MCIN) under the research project TUBOLAN PID2019-108672RJ-I00 funded by MCIN/AEI/ 10.13039/501100011033. In addition, this work received support from the Portuguese Foundation for Science and Technology (FCT) under the MICROCENO project (PTDC/CTA-AMB/0608/2020), and from the Spanish National Research Council (CSIC) through the intramural project PIE_20214AT021. A.Z.M. was supported by the CEECIND/01147/2017 contract from FCT and the Ramón y Cajal contract (RYC2019-026885-I) from the MCIN.
Author information
Authors and Affiliations
Contributions
GN and AZM conceived the study and performed the sampling activities. Funding was secured by AZM. Laboratory protocols were conducted by GN and AZM. JGP, EP, AZM, and GN analyzed the data. The first draft of the manuscript was written by GN along with significant contributions from JGP, EP, MI, and AZM. AZM and MI contributed to the review and editing of this manuscript. All authors contributed critically to the drafts and gave final approval for publication.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nicolosi, G., Gonzalez-Pimentel, J.L., Piano, E. et al. First Insights into the Bacterial Diversity of Mount Etna Volcanic Caves. Microb Ecol 86, 1632–1645 (2023). https://doi.org/10.1007/s00248-023-02181-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02181-2