Abstract
Here, we demonstrate the beneficial effect of surfactant-producing pseudomonads on Pantoea eucalypti 299R. We conducted a series of experiments in environments of increasing complexity. P. eucalypti 299R (Pe299R), and Pseudomonas sp. FF1 (Pff1) or Pe299R and surfactant-production deficient Pseudomonas sp. FF1::ΔviscB (Pff1ΔviscB) were co-inoculated in broth, on swarming agar plates, and on plants. In broth, there were no differences in the growth dynamics of Pe299R when growing in the presence of Pff1 or Pff1ΔviscB. By contrast, on swarming agar plates, Pe299R was able to co-swarm with Pff1 which led to a significant increase in Pe299R biomass compared to Pe299R growing with Pff1ΔviscB or in monoculture. Finally in planta, and using the single-cell bioreporter for reproductive success (CUSPER), we found a temporally distinct beneficial effect of Pff1 on co-inoculated Pe299R subpopulations that did not occur in the presence of Pff1ΔviscB. We tested three additional surfactant-producing pseudomonads and their respective surfactant knockout mutants on PE299R on swarming agar showing similar results. This led us to propose a model for the positive effect of surfactant production during leaf colonization. Our results indicate that co-motility might be common during leaf colonization and adds yet another facet to the already manyfold roles of surfactants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The microbial habitat that is presented by leaf surfaces, the phyllosphere, is densely colonized by microorganisms. Among these microorganisms, bacteria are the most abundant and prevalent group, establishing non-random patterns of colonization on leaves. Bacteria have a tendency to aggregate with one another in close proximity, rather than being distributed homogeneously along the leaf surface, suggesting the role of deterministic processes on leaf colonization [1].
In recent years, much attention has been paid to leaf colonizers and the factors that drive leaf colonization at the population and the community level. For example, the host plant species influences leaf colonization and bacterial community composition [2,3,4], leading to recurring patterns of bacterial communities on leaves, at least at low phylogenetic resolution, from year to year [5, 6].
At the same time, microbe-microbe interactions affect community assemblages on leaves [7,8,9]. However, many of the mechanisms driving these interactions remain unclear. In studies where bacterial communities are relatively simple, it appears that overlap in resource utilization ability of competitors has a limited impact on leaves [8, 10, 11]. This suggests that the highly segregated leaf environment constrains interspecies bacterial interactions, despite their tendency to co-aggregate [12, 13]. As proximity is a key factor for interaction, mobility might emerge as an important factor that is shaping interactions and, ultimately, bacterial communities. In general, mobility on leaves has received limited attention, with most studies being related to the presence of flagellar genes in Pseudomonas spp. [14,15,16]. Since roughly 5% of the leaf surface is colonized by bacteria [13] and not all leaf-colonizing bacteria are flagellated [17], other means of mobility must be present on leaves. This may include movement mediated by colony expansion or gliding motility [18, 19], or spread by dew [20, 21]. The genera Pantoea and Pseudomonas are known to be very common leaf colonizers [6, 22]. Both taxa interact with their plant host in various ways, ranging from pathogenicity to plant protection [23,24,25,26]. However, their interaction within communities has not received much attention, despite some initial efforts [12].
Phyllosphere-colonizing pseudomonads have previously been studied with regard to their ability to produce surfactants [27,28,29,30,31]. Interestingly, a fitness advantage of surfactant-producing pseudomonads over their surfactant-deficient counterparts was evident only under specific conditions such as fluctuating humidity [27]. Otherwise, surfactant production did not seem to affect the ability of pseudomonads to colonize leaves [29].
Surfactants have been suggested to have multiple roles in the phyllosphere, including increased spread of water by reducing surface tension, improved nutrient diffusion from the apoplast to the phyllosphere by increasing cuticle permeability [31], and increased drought resistance due to their hygroscopic nature [27, 28]. On semi-solid media, surfactants are necessary for swarming, and it has been shown that they can facilitate the mobilization of some bacteria in a process called co-swarming [32]. However, the effect of surfactant-producing bacteria on a second colonizer in the phyllosphere has, to our knowledge, not been studied.
Bacterial mobility relies on various mechanisms. Swimming motility is an active form of movement, powered by rotating flagella, occurring in liquid or low-viscosity conditions [33]. Gliding motility is movement on surfaces, independent of flagella or pili [34]. Another surface-associated motility mode is sliding motility, driven by excreted biosurfactants, preventing cells from forming thick biofilm layers and aiding dispersal on the surface they inhabit [35]. Twitching motility is mediated by type IV pili, which propels bacteria by retraction, while swarming motility is the coordinated movement of multiple bacteria across solid or semi-solid surfaces, requiring flagella, a functional quorum sensing system, and surfactant biosynthesis [36]. With this mechanism, bacteria co-migrate in side-by-side groups called rafts, instead of individually as in swimming motility [37].
Previously, we developed a bioreporter, CUSPER, to measure the number of divisions of individual cells of the bacterium Pantoea eucalypti 299R (formerly known as Erwinia herbicola 299R and Pantoea agglomerans 299R) after they arrive in new environments. This bioreporter for reproductive success is based on the dilution of green fluorescent protein (GFP) during cell division. Thereby, the GFP intensity becomes a direct proxy for the number of experienced divisions [38, 39]. In a previous study, we observed that Pseudomonas spp. increased the single-cell reproductive success of P. eucalypti 299R (Pe299R) in planta despite being strong competitors in vitro [11]. Given the common feature of surfactant production in pseudomonads [30, 40], we hypothesized that surfactant production increases the reproductive success of a second colonizer in the phyllosphere. To test this, we investigated the interactions between Pe299R, surfactant-producing Pseudomonas isolates, and their respective knockout mutants in various conditions, ranging from liquid medium, agar surfaces, and leaves. We observed contrasting results in liquid cultures compared to agar surfaces and leaves.
Material and Methods
Bacteria, Strain Construction, and Growth Conditions
All bacteria used in this study are listed in Table 1. Bacteria were routinely grown on lysogeny broth agar (LB-Agar, HiMedia). Pantoea eucalypti 299R was equipped with a constitutively expressed red fluorescent mScarlet-I protein gene and the plasmid pCUSPER [11]. The resulting strain will be referred to as Pe299RCUSPER from here onwards. To maintain the plasmid carrying the reproductive success construct in Pe299RCUSPER, the agar was supplemented with 50 mg L−1 kanamycin (Roth). Constitutively cyan-fluorescent derivatives of strains Pseudomonas sp. Pff1 (Pff1cyan) and Pseudomonas sp. Pff1::ezTn5-viscB (Pff1ΔviscBcyan) used in this study were generated using plasmid pMRE-Tn7-141 [41] explained elsewhere using conjugation and the auxotrophic donor strain E. coli ST18 [42, 43]. Strains Pseudomonas sp. Pff2, Pseudomonas sp. Pff2::ezTn5-viscB, Pseudomonas sp. Pff3, Pseudomonas sp. Pff3::ezTn5-massB, Pseudomonas sp. Pff4, and Pseudomonas sp. Pff4::ezTn5-massB were characterized elsewhere and carry Tn5-knockout insertions in their respective surfactant production genes: viscosin B (viscB) or the massetolide B (massB) genes [29]. Those genes were previously shown to be responsible for surfactant production.
In Vitro Co-inoculation Assay
To investigate the co-colonization of Pe299RCUSPER, Pff1cyan or Pff1ΔviscBcyan in vitro, all strains were grown in 3 mL M9 minimal media (Na2HPO4•7H2O 64 g L−1, KH2PO4 15 g L−1, NaCl 2.5 g L−1, NH4Cl 5.0 g L−1) supplemented with 200 µM FeCl3 (to avoid production of autofluorescent pyoverdines by pseudomonads) and 0.13% glucose, fructose, and sorbitol each (M9 3C) overnight at 30 °C and 200 rpm. Five hundred microliters of each overnight culture were then used to inoculate 50 mL M9 3C, respectively. After 6 h, cultures were washed twice by centrifugation at 2000 g for 5 min and resuspended in PBS. Suspensions were diluted to an optical density (OD600nm) of 1 and the following treatments were prepared: Pe299RCUSPER, Pe299RCUSPER vs. Pff1cyan, Pe299RCUSPER vs. Pff1ΔviscBcyan, Pff1cyan, and Pff1ΔviscBcyan. These suspensions were diluted into either M9 supplemented with glucose 0.4% (M9gluc), M9 3C, or LB to a final OD600nm of 0.04 for each strain. Despite the lack of kanamycin, we did not expect any significant loss of plasmid during this experiment as the plasmid backbone of pCUSPER was described to be highly stable in Pe299R [45]. Furthermore, the fitness effects of near identical plasmids in Pe299R were described to be negligible [46]. Triplicate treatments were pipetted into a 96-well plate and placed into a CLARIOstar Plus microplate reader (BMG Labtech). The fluorescence of the mScarlet-I expressing Pe299RCUSPER was determined by excitation at 530–570 nm and measuring emission at 580–620 nm, while mTurquoise2, produced by pseudomonads, was excited at 400–440 nm and its emission was measured at 450–490 nm using the CLARIOstar software (version 5.70, BMG Labtech). Measurements were performed in 20-min intervals for 42 h. The plate was incubated at 30 °C, and the plate was shaken in a double orbital at 200 rpm between measurements.
The area under the curve of the red and cyan fluorescence kinetics of each sample was determined in Prism 10.0.1 (Graphpad). As cyan fluorescence kinetics exhibited a negative change in fluorescence that was media dependent, the data was corrected by a constant value as to move every datapoint above the baseline before the absolute area under the curve was determined.
Swarming Assays
To determine the swarming ability of Pe299RCUSPER, Pff1cyan, and Pff1ΔviscBcyan, each strain was inoculated onto soft agar. KB-based soft agar plates were prepared with 1.26% w/v King’s medium B, (LB-Agar, HiMedia) and 0.4% agarose. The center of the plate was inoculated with 10 µL of bacterial suspension (OD600nm = 1, prepared as above), and pictures were taken after 24 h of incubation at 30 °C using a dark field illumination stage [47] in a dark chamber (MultiImage Light Cabinet) with attached Axiocam 105 (Zeiss) and Zen Core (version 3.2, Zeiss).
LB-based soft agar was prepared by diluting one part LB agar with two parts ddH2O and supplementing with 200 µM FeCl3. Two mL soft agar per well was then distributed into a 6-well microtiter plate (Greiner). After the agar was gelled, a 1.5 µL drop of washed bacterial suspensions (adjusted OD600nm = 0.5) was placed in the middle of each well. The following monocultures and mixtures were prepared: (i) Pe299RCUSPER, (ii) Pe299RCUSPER vs. Pff1cyan, (iii) Pe299RCUSPER vs. Pff1ΔviscBcyan, and (iv) Pff1cyan, and Pff1ΔviscBcyan. The plates were then incubated at 30 °C for 16 h. Afterwards, the plates were placed into a CLARIOstar Plus Microplate reader (BMG Labtech). The fluorescence of the mScarlet-I expressing Pe299RCUSPER was determined by exciting the sample at 530–570 nm and measuring emission at 580–620 nm while mTurquoise-producing pseudomonads were excited at 400–440 nm and its emission was measured at 450–490 nm. The plates were scanned using the 30 × 30 Matrix scan mode of the CLARIOstar software using bottom optics to obtain the spatial information of each strain. This resulted in a two-dimensional distribution of red and cyan fluorescence data. Background subtraction was performed individually for each datapoint. Data was visualized using the heatmap function of Prism. Total fluorescence of the red fluorescence channel was used as a proxy to determine the total biomass of Pe299RCUSPER. The experiment was performed four times on LB soft agar.
Additionally, to test if co-swarming also occurs when Pe299RCUSPER co-colonizes agar surfaces with other swarming pseudomonads, Pe299RCUSPER was co-inoculated with Pff2, Pff3, and Pff4, as well as their respective surfactant-deficient mutants on soft LB agar.
Plant Growth
Plants were prepared as explained in Miebach et al. [48]. Briefly, Arabidopsis thaliana Col-0 seeds were surface sterilized by vortexing in 70% v/v ethanol for 2 min. The ethanol was then removed by pipetting, and the seeds were treated with 50% v/v household bleach (NaOCl, 2.47%) and 0.02% v/v Tween-20 for 7 min. Afterwards, the seeds were washed three times with sterile water. Sterilized seeds were kept in water and stratified in the dark at 4 °C for 3 days. After stratification, the seeds were sown on cut pipette tips (∼5 mm in length) pre-filled with ½ strength Murashige-Skoog (½ MS) agar medium and placed in a Petri dish with ½ MS agar. Petri dishes were closed with parafilm (Bemis) and were placed in a M-5-Z growth cabinet (PolyKlima) with a 11/13 day/night interval (including 30 min dusk and 30 min dawn during which the lights slowly increase in intensity), 80% relative humidity, and 210 μmol s−1m−2 light intensity.
Gnotobiotic culture boxes were prepared following the methods described by [48]. In brief, Magenta Culture Boxes GA-7 were filled with 90-g-fine zeolite clay granulate (Klinoptilolith, 0.2–0.5 mm, Labradorit.de) and autoclaved with lids closed. Lids were previously perforated and holes were covered with a double layer of gas permeable tape (Micropore, 3M). After autoclaving and cooling down, 45 ml of sterile ¾ MS Medium was added per box under aseptic conditions. One week after sowing, seedlings were transferred, including the tips they were germinated on, into prepared Magenta boxes. Four seedlings were placed per box and grown under the same conditions described above.
Plant Inoculation
To prepare the inoculum of the different strains, a single colony was selected and used to prepare overnight cultures in LB medium (HiMedia, Pe299RCUSPER with 50 mg L−1 kanamycin, respectively). The overnight cultures were then used to inoculate fresh liquid cultures by adding 500 µl of cultures to 50 ml fresh medium. Those cultures were grown to mid exponential phase while shaking at 30 °C and 200 rpm. Pe299RCUSPER cultures were supplemented with kanamycin and 1 mM isopropylthio-β-galactoside (IPTG) to induce GFP expression [38]. When reaching log-phase after approximately 6 h of growth, cells were pelleted by 5 min of centrifugation at 4000 ✕ g, washed three times, and adjusted to an OD600nm of 0.1 with sterile phosphate buffered saline (PBS, 1.37 M NaCl, 27 mM KCl, 100 mM Na2HPO4, 18 mM KH2PO4, pH 7). Where appropriate, strains were mixed prior to inoculation, resulting in a relative OD600nm = 0.05 for each strain.
To inoculate A. thaliana plants, Magenta® box lids were temporarily replaced with a lid with only one central hole. Each box was then sprayed with 200 µl of inoculum with an airbrush paint gun (Ultra Spray gun, Harder & Steenbeck, Norderstedt, Germany). Then, the lids were replaced and plants were placed back into the growth chamber.
Plant Sampling and Bacterial Cell Recovery
Three-week-old plants were sampled after 0 h, 12 h, 18 h, and 24 h post inoculation. Plants were sampled by harvesting the total aboveground material using sterile forceps and scissors. Plant material was transferred into a 15-ml centrifuge tube, and fresh weight was determined before 1 ml PBS was added to recover leaf surface-attached bacteria. The samples were then vortexed for 15 s and sonicated for 5 min at 75% intensity in a sonication bath (Emmi 12 HC, EMAG). Leaf washes were recovered from the tubes and a 100 µl aliquot was used to determine colony-forming units (CFU) by serial dilution on LB agar supplemented with rifampicin (50 mg/L) and kanamycin (50 mg/L) to select for Pe299RCUSPER. The remaining leaf wash was centrifuged for 10 min at 15,000 × g at 4 °C, the supernatant was discarded, and the resulting bacterial pellet resuspended was fixed overnight in 4% w/v paraformaldehyde in PBS at 4 °C. After fixation, the PFA was removed by three washing steps with sterile PBS. After the last washing step, the pellets were resuspended in 50 µl PBS mixed with 50 µl 96% v/v ethanol and stored at − 20 °C until they were analyzed. All samples were analyzed within 2 weeks.
Microscopical Analysis of CUSPER Signals and Image Cytometry
Recovered bacterial cells were analyzed using widefield fluorescence microscopy as described elsewhere [11]. Briefly, cells were drop spotted onto 1% w/v agarose slabs on a microscopy slide and covered with a cover slip. An AxioImager Z2 microscope (Zeiss) with an Axiocam 712 mono camera (Zeiss) and X-cite Xylis broad spectrum LED light source (Excelitas) were used. Images were acquired at 1000 × magnification (EC Plan-Neofluar 100 × /1.30 Ph3 Oil M27 objective) in phase contrast, green fluorescence, and red fluorescence (Zeiss filter sets 38HE (BP 470/40-FT 495-BP 525/50) and 43HE (BP 550/25-FT 570-BP 605/70), respectively) using the software Zen 3.3 (Zeiss). At least 120 cells were acquired per biological replicate which consisted of bacteria pooled from four different plants. Images were analyzed using FIJI and as described previously [11, 49]. Briefly, Pe299CUSPER cells were identified using their constitutive red fluorescence and the thresholding method “intermodes.” The resulting regions of interest were converted into a binary mask. Artifacts were excluded by analyzing only particle sizes from 0.5 to 2.5 µm and excluding cells touching the image edges. All images were manually curated using the phase contrast images to exclude false positive red fluorescent particles. The mask was then used to determine green fluorescence intensity of Pe299RCUSPER cells. In addition, the average background fluorescence was measured by randomly sampling the background area of each image. Background fluorescence was subtracted from the data. As previously described, the number of experienced divisions of every cell was determined by calculating log2(average cell’s GFP fluorescence at t = 0 divided by single cell’s fluorescence at time t) [38].
Results
Pseudomonads Negatively Affect Growth of Pe299RCUSPERIn Vitro
To investigate the interaction of Pe299RCUSPER with Pff1cyan or Pff1ΔviscBcyan in homogeneous conditions, they were grown in shaken liquid cultures. Under these conditions, there was a strong decrease of Pe299RCUSPER red fluorescence when it was co-inoculated with Pff1cyan or Pff1ΔviscBcyan (Fig. 1). Different media had slightly different effects on this interaction, and the decrease ranged from > 90% decrease in M9gluc or M9 3C to > 30% reduction in LB. This effect was not associated with the Pseudomonas red autofluorescence. By contrast, the effect of Pe299RCUSPER on Pff1cyan and Pff1ΔviscBcyan was much smaller. Only in rich LB growth medium, we could detect a significant negative effect of Pe299RCUSPER on the pseudomonads (Supplemental Fig. 1). In conclusion, the pseudomonads had a strong negative effect on Pe299RCUSPER in vitro, while Pe299RCUSPER is barely affecting the growth of the pseudomonad strains.
Impact of Pff1cyan and Pff1ΔviscBcyan on the growth of Pe299RCUSPER in different media. The effect on Pe299RCUSPER growth was measured by determining the area under the curve of the red fluorescence of Pe299RCUSPER monocultures and co-inoculations of Pe299RCUSPER and the pseudomonads. A) M9 supplemented with glucose B) M9 supplemented with glucose, fructose and sorbitol and C) LB. Note that the y-axis is on a log10 scale
Swarming on Solid Media
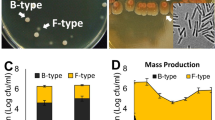
When inoculated onto soft KB agar medium alone, the different strains showed various swarming patterns. Pe299RCUSPER alone formed as a colony with entire edges that is slightly more slimy compared to growth on standard agar media and does not grow beyond the initial location of inoculation. Pff1cyan grew in a flower-shaped colony that is indicative for swarming, far beyond the initial site of inoculation. By contrast, Pff1ΔviscBcyan formed a colony similar to Pe299RCUSPER and was not able to move far beyond the initial site of inoculation (Fig. 2).
To investigate bacterial behavior after co-inoculating Pe299RCUSPER with the two different pseudomonads, we used their respective constitutively expressed fluorescent proteins to track their biomass on a swarming agar (Fig. 3). While monocultures behaved as expected (Fig. 3A), we found that co-inoculated bacteria affected each other in different fashions. In co-culture with the swarming Pff1cyan, Pe299RCUSPER had a much wider distribution on the agar surface (Fig. 3B). By contrast, in co-culture with the non-swarming Pff1ΔviscBcyan, Pe299RCUSPER was similarly restricted in its distribution as in a monoculture. By repeating the experiment four times, we were able to determine the total change in biomass during mono and co-cultures (Fig. 3C). As a result, the Pe299RCUSPER biomass, as measured by red fluorescence, was significantly higher in co-inoculations with surfactant-producing Pff1cyan compared to Pe299RCUSPER monocultures and co-inoculations with non-surfactant-producing Pff1ΔviscBcyan (p = 0.0001 and < 0.0001, respectively, one-way ANOVA Tukey’s multiple comparison test). By contrast, albeit not significant, Pe299RCUSPER biomass is slightly reduced during co-culture with Pff1ΔviscBcyan.
Spatially resolved analysis of bacterial growth and swarming behavior on LB soft agar plates after 16 h. A Monocultures of Pe299RCUSPER, Pff1cyan, and Pff1ΔviscBcyan or B mixed cultures of Pe299RCUSPER in combination with Pff1cyan or Pff1ΔviscBcyan on LB soft agar plates. Bacterial growth and biomass were tracked with a fluorescent microtiter plate reader. Growth of Pe299RCUSPER was determined by measuring red fluorescence emission, and growth of pseudomonads was determined by measuring cyan fluorescence emission. Note that the color scales are presented as the decadic logarithm of arbitrary fluorescence units. Furthermore, the experiment was repeated four times on LB soft agar plates. C The fluorescence of Pe299RCUSPER was determined after 16 h, and the ratio of Pe299CUSPER growing in mixtures over its monoculture was determined
Co-inoculation In Planta Does Not Significantly Affect Pe299RCUSPER at the Population Scale but Does Affect Reproductive Success at the Single-Cell Resolution
To investigate the effect of a co-colonizer-producing surfactant on Pe299RCUSPER in planta, the strains were co-inoculated onto axenic A. thaliana plants. The two different co-colonizing pseudomonads affected the population size of Pe299RCUSPER similarly. Although we observed a trend that the population was slightly higher after 24 h in one of the experiments, this was not consistent between experiments (Supplemental Fig. 2).
By contrast, at the single-cell resolution, there are noteworthy differences in the population development during the co-colonization of leaves (Fig. 4). Generally, a proportionally larger subpopulation of Pe299RCUSPER experienced more cell divisions in the presence of the surfactant-producing Pff1cyan compared to the non-producing Pff1ΔviscBcyan. This effect is most apparent after 24 h and was reproduced in three independent experiments (Fig. 4 and Supplemental Fig. 3). Furthermore, we observed that after 18 h, the presence of the non-surfactant-producing strain had a positive effect on Pe299RCUSPER (Fig. 4).
Reproductive success of individual Pe299RCUSPER cells during co-colonization of leaves with Pff1cyan or Pff1ΔviscBcyan. In gray, the respective T0 fluorescence intensity of the Pe299RCUSPER population is depicted. Every increase in reproductive success depicts a cell division relative to the T0 population. Every sample is pooled from the bacteria recovered from four plants
Discussion
The potential roles of biosurfactants in plant–microbe interactions have been investigated from many different angles: their hygroscopic nature enhances the survival of pseudomonads [27] and increases water availability on leaves [28], they are suspected to increase diffusion of nutrients through the hydrophobic leaf cuticle [31], and they have been shown to increase alkane degradation by leaf-colonizing bacteria [29]. Generally, the abundance of biosurfactant-producing bacteria on leaves is proportionally high [30, 31, 50], indicating a potential ecological importance during life on leaves. Here, we show that surfactant-producing bacteria may facilitate the growth of co-colonizers on leaves in a surfactant production and surface colonization dependent manner.
Being both copiotrophic generalists that exhibit a wide range of nutrient utilization, it was expected that Pe299RCUSPER and both pseudomonads affected each other strongly in liquid culture. Indeed, it was mostly Pe299RCUSPER that was negatively affected in minimal media and less so in complex LB medium. In turn, the pseudomonads were almost not affected at all by the presence of Pe299RCUSPER in minimal media and to a smaller degree in LB medium. These results are in stark contrast to our observations on a spatially structured soft agar surface. Here, the non-surfactant-producing Pff1ΔviscBcyan negatively affected Pe299RCUSPER, but not Pff1cyan which instead increased the amount of Pe299RCUSPER fluorescence as a proxy for biomass by a factor of three and thereby increased its fitness dramatically. As this increase correlates to a larger spread of the Pe299RCUSPER, it is likely that this increase can be accredited to a mobilization of Pe299RCUSPER by Pff1cyan, similar co-swarming phenomena have previously been observed in Paenibacillus vortex and other Paenibacillus strains [51], as well as Pseudomonas aeruginosa and Burkholderia cenocepacia [32, 52]. Generally, it has been shown that the swarming conferring strain gains a benefit from the co-swarmer by for instance taking advantage of antibiotic resistances provided by the immobile strain [51], or by recovering mobility in case one strain loses mobility factors [32, 52]. In our study, there is no apparent benefit for the strains that mobilize Pe299RCUSPER. Instead, it is Pe299RCUSPER that seems to be able to take advantage of the swarming strains and increases its fitness as compared to homogeneous shaken liquid cultures where swarming does not confer any fitness advantages. This observation led us to test if this effect also pertains to colonization of leaf surfaces, which is the origin of isolation of all strains used in this study.
To test this, we tracked the changes of Pe299RCUSPER populations on A. thaliana and the ability of Pe299RCUSPER individual cells to divide on leaves, exploiting the reproductive success bioreporter CUSPER. At the population scale, the effect of co-colonization was minimal, although a trend of higher Pe299RCUSPER populations during co-colonization with Pff1cyan as compared to co-colonization with Pff1ΔviscBcyan could be observed (Supplemental Fig. 2). This is in contrast to single-cell observations, where we could detect an increased number of cells that experienced a notably higher number of divisions in the presence of Pff1cyan as compared to the presence of Pff1ΔviscBcyan (Fig. 4) after 24 h. Curiously, this effect was not visible after 18 h where Pe299RCUSPER seemed to have a minimal growth advantage in the presence of Pff1ΔviscBcyan. To explain this, we hypothesize the following model of interactions: During inoculation, it is more probable for individual strains to arrive on the leaf without a competitor in their local environment (Fig. 5(A)). In a scenario without swarming (Fig. 5(B) and (C)), the chance of bacterial strains interacting with each other is low as the minimal interaction distance between bacteria on leaves is about 10 µm [13, 53]. As a consequence, bacteria can initially grow rather unimpeded by competition. By contrast, in a scenario with swarming (Fig. 5(D) and (E)), the swarming strain will have higher chances to meet the non-swarming Pe299RCUSPER, which leads to high local competition and locally reduced populations of Pe299RCUSPER (compare Fig. 3, where the local biomass of Pe299RCUSPER is reduced during competition). However, as co-swarming leads to mobilization of Pe299RCUSPER and allows the exploration of new microenvironments, this should lead to an increased reproductive success of the strain despite the local competition (Fig. 5(E)). This is directly analogous to the scenario observed on the spatially structured agar surface (Fig. 3).
Model explaining behavior of Pe299CUSPER after co-inoculation with Pff1ΔviscBcyan (B, C) or Pff1cyan (D, E). (A) The initial distribution of non-motile, red-colored Pe299CUSPER cells and co-inoculated cyan-colored Pff1ΔviscBcyan or Pff1cyan cells. In the left scenario, cyan cells do not produce any surfactants and do not swarm. As a result red and cyan populations remain localized and rely on the locally available nutrients. In the right scenario, cyan cells produce surfactants and swarm. This leads to a higher probability of cyan cells to encounter red cells, which leads to a temporally high competition for nutrients and a reduction of red cell divisions (D). However, if red cells are mobilized by co-swarming as a result of close spatial proximity to cyan cells, this leads to the exploration of new sites and an increase of growth
Effect of Other Surfactant-Producing Pseudomonads on Pe299RCUSPER
Interestingly, in a previous study, we have observed a similar effect of increased reproductive success of Pe299RCUSPER when co-colonizing plant leaves with surfactant-producing P. koreensis P19E3 and P. syringae B728a despite high overlap in resource utilization abilities [11]. This effect can now likely be accredited to the production of surfactants and co-swarming. To further test if co-swarming of Pe299RCUSPER with pseudomonads can be generalized and accredited to surfactant production, we have tested the growth Pe299RCUSPER and three additional pseudomonads and their respective surfactant knockout mutants on soft agar. Indeed, we could observe that the Pseudomonas strains Pff2 and Pff4, but not their respective surfactant knockout mutants to co-swarming of Pe299RCUSPER (Supplemental Fig. 4). Intriguingly, Pff3 was not able to mobilize Pe299RCUSPER. Pff3 also exhibited a different style of swarming on soft agar plates which seemed to consist of a very thin layer of biomass (Supplemental Fig. 5), which might explain this difference in co-swarming. Exploring the reasons for those differences is beyond the scope of this study and might be growth medium dependent as well as strain dependent as bacteria have been shown to be diverse in their swarming behavior [54, 55].
Conclusion
While pseudomonads act as strong competitors in homogeneous environments, in spatially structured environments, they affect co-colonizing P. eucalypti 299R positively. While this effect cannot be observed on a population scale during leaf colonization, we provide evidence that P. eucalypti 299R takes advantage of the presence of surfactant-producing pseudomonads during leaf colonization. This stresses the importance of surfactants produced by pseudomonads as public goods during leaf colonization and implies that cheating of P. eucalypti 299R and possibly other taxa may be the rule rather than the exception. Our study highlights another pivotal role of surfactants in the phyllosphere and the implications of surfactant production in this environment. Particularly in the context of preparing microbial inocula for the application on plants, our findings provide additional traits, i.e., surfactant production and the ability for co-swarming, which should be considered during the formulation of products [].
Data Availability
The raw data of this article is available at https://doi.org/10.5281/zenodo.10610551.
References
Schlechter RO, Miebach M (2019) Remus-Emsermann MNP (2019) Driving factors of epiphytic bacterial communities: a review. J Adv Res 14(19):57–65
Bodenhausen N, Bortfeld-Miller M, Ackermann M, Vorholt JA (2014) A Synthetic community approach reveals plant genotypes affecting the phyllosphere microbiota. PLoS Genet 10(4):e1004283
Lajoie G, Kembel SW (2021) Plant-bacteria associations are phylogenetically structured in the phyllosphere. Mol Ecol 30(21):5572–5587
Mercier J, Lindow SE (2000) Role of leaf surface sugars in colonization of plants by bacterial epiphytes. Appl Environ Microbiol 66(1):369–374
Howe A, Stopnisek N, Dooley SK, Yang F, Grady KL, Shade A (2023) Seasonal activities of the phyllosphere microbiome of perennial crops. Nat Commun 14(1):1039
Vorholt JA (2012) Microbial life in the phyllosphere. Nat Rev Microbiol 10(12):828–840
Carlström CI, Field CM, Bortfeld-Miller M, Müller B, Sunagawa S, Vorholt JA (2019) Synthetic microbiota reveal priority effects and keystone strains in the Arabidopsis phyllosphere. Nat Ecol Evol 3(10):1445–1454
Schäfer M, Pacheco AR, Künzler R, Bortfeld-Miller M, Field CM, Vayena E, Hatzimanikatis V, Vorholt JA (2023) Metabolic interaction models recapitulate leaf microbiota ecology. Science 381(6653):5121
Schäfer M, Vogel CM, Bortfeld-Miller M (2022) Mapping phyllosphere microbiota interactions in planta to establish genotype–phenotype relationships. Nat Microbiol 7(6):856–867
Remus-Emsermann MNP, Schlechter RO (2018) Phyllosphere microbiology: at the interface between microbial individuals and the plant host. New Phytol 218(4):1327–1333
Schlechter RO, Kear EJ, Bernach M, Remus DM (2023) Remus-Emsermann MNP (2023) Metabolic resource overlap impacts competition among phyllosphere bacteria. ISME J 9:1445–1454
Monier J-M, Lindow SE (2005) Spatial organization of dual-species bacterial aggregates on leaf surfaces. Appl Environ Microbiol 71(9):5484–5493
Remus-Emsermann MNP, Lücker S, Müller DB, Potthoff E, Daims H, Vorholt JA (2014) Spatial distribution analyses of natural phyllosphere-colonizing bacteria on Arabidopsis thaliana revealed by fluorescence in situ hybridization. Environ Microbiol 16(7):2329–2340
Delmotte N, Knief C, Chaffron S, Innerebner G, Roschitzki B, Schlapbach R, von Mering C, Vorholt JA (2009) Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc Natl Acad Sci USA 106(38):16428–16433
Haefele DM, Lindow SE (1987) Flagellar motility confers epiphytic fitness advantages upon Pseudomonas syringae. Appl Environ Microbiol 53(10):2528–2533
van der Wal A, Tecon R, Kreft J-U, Mooij WM, Leveau JHJ (2013) Explaining bacterial dispersion on leaf surfaces with an individual-based model (PHYLLOSIM). PLoS ONE 8(10):e75633
Bai Y, Müller DB, Srinivas G, Garrido-Oter R, Potthoff E, Rott M, Dombrowski N et al (2015) Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528(7582):364–369
Martínez A, Torello S, Kolter R (1999) Sliding motility in mycobacteria. J Bacteriol 181(23):7331–7338
Su P-T, Liao C-T, Roan J-R, Wang S-H, Chiou A, Syu W-J (2012) Bacterial colony from two-dimensional division to three-dimensional development. PLoS ONE 7(11):e48098
Beattie GA (2011) Water relations in the interaction of foliar bacterial pathogens with plants. Annu Rev Phytopathol 49:533–555
Van Stan, John T., II, Cindy E. Morris, Kyaw Aung, Yakov Kuzyakov, Donát Magyar, Eria A. Rebollar, Mitja Remus-Emsermann, Stéphane Uroz, and Philippe Vandenkoornhuyse. 2020. “Precipitation partitioning—hydrologic highways between microbial communities of the plant microbiome?” In Precipitation partitioning by vegetation: a global synthesis, edited by Van Stan, II, John T., Ethan Gutmann, and Jan Friesen, 229–52. Cham: Springer International Publishing.
Rastogi G, Sbodio A, Tech JJ, Suslow TV, Coaker GL, Leveau JHJ (2012) Leaf microbiota in an agroecosystem: spatiotemporal variation in bacterial community composition on field-grown lettuce. ISME J 6(10):1812–1822
Ramette A, Frapolli M, Saux M-L, Gruffaz C, Meyer J-M, Défago G, Sutra L, Moënne-Loccoz Y (2011) Pseudomonas protegens sp. Nov., widespread plant-protecting bacteria producing the biocontrol compounds 2,4-diacetylphloroglucinol and pyoluteorin. Syst Appl Microbiol 34(3):180–188
Vrancken K, Holtappels M, Schoofs H, Deckers T, Valcke R (2013) Pathogenicity and infection strategies of the fire blight pathogen Erwinia amylovora in Rosaceae: state of the art. Microbiology 159(Pt 5):823–832
Xin X-F, Kvitko B, He SY (2018) Pseudomonas syringae: what it takes to be a pathogen. Nat Rev Microbiol 16(5):316–328
Zengerer V, Schmid M, Bieri M, Müller DC, Remus-Emsermann MNP, Ahrens CH, Pelludat C (2018) Pseudomonas orientalis F9: a potent antagonist against phytopathogens with phytotoxic effect in the apple flower. Front Microbiol 9(February):145
Burch AY, Zeisler V, Yokota K, Schreiber L, Lindow SE (2014) The hygroscopic biosurfactant syringafactin produced by Pseudomonas syringae enhances fitness on leaf surfaces during fluctuating humidity. Environ Microbiol 16(7):2086–2098
Hernandez MN, Lindow SE (2019) Pseudomonas syringae increases water availability in leaf microenvironments via production of hygroscopic syringafactin. Appl Environ Microbiol 85(18):e01014-e1019
Oso S, Fuchs F, Übermuth C, Zander L, Daunaraviciute S, Remus DM, Stötzel I, Wüst M, Schreiber L, Remus-Emsermann MNP (2021) Biosurfactants produced by phyllosphere-colonizing pseudomonads impact diesel degradation but not colonization of leaves of gnotobiotic Arabidopsis thaliana. Appl Environ Microbiol 87:9
Oso S, Walters M, Schlechter RO, Remus-Emsermann MNP (2019) Utilisation of hydrocarbons and production of surfactants by bacteria isolated from plant leaf surfaces. FEMS Microbiol Lett 366:6
Schreiber L, Krimm U, Knoll D, Sayed M, Auling G, Kroppenstedt RM (2005) Plant-microbe interactions: identification of epiphytic bacteria and their ability to alter leaf surface permeability. New Phytol 166(2):589–594
Morin C, Landry M, Groleau Marie-Christine, Déziel E (2022) Surface motility favors codependent interaction between Pseudomonas aeruginosa and Burkholderia cenocepacia. mSphere 7(4):e0015322
Ha D-G, Kuchma SL, O’Toole GA (2014) Plate-based assay for swimming motility in Pseudomonas aeruginosa. Methods Mol Biol 1149:59–65
McBride MJ (2001) Bacterial gliding motility: multiple mechanisms for cell movement over surfaces. Annu Rev Microbiol 55:49–75
Wadhwa N, Berg HC (2022) Bacterial motility: machinery and mechanisms. Nat Rev Microbiol 20(3):161–173
Merz AJ, So M, Sheetz MP (2000) Pilus retraction powers bacterial twitching motility. Nature 407(6800):98–102
Kearns DB (2010) A field guide to bacterial swarming motility. Nat Rev Microbiol 8(9):634–644
Remus-Emsermann MNP, Leveau JHJ (2010) Linking environmental heterogeneity and reproductive success at single-cell resolution. ISME J 4(2):215–222
Remus-Emsermann MNP, Tecon R, Kowalchuk GA, Leveau JHJ (2012) Variation in local carrying capacity and the individual fate of bacterial colonizers in the phyllosphere. ISME J 6(4):756–765
Raaijmakers JM, De Bruijn I, Nybroe O, Ongena M (2010) Natural functions of lipopeptides from Bacillus and Pseudomonas: more than surfactants and antibiotics. FEMS Microbiol Rev 34(6):1037–1062
Schlechter RO, Jun H, Bernach M, Oso S, Boyd E, Muñoz-Lintz DA, Dobson RCJ, Remus DM, Remus-Emsermann MNP (2018) Chromatic bacteria - a broad host-range plasmid and chromosomal insertion toolbox for fluorescent protein expression in bacteria. Front Microbiol 9(December):3052
Schlechter R, Remus-Emsermann M (2019) Delivering ‘chromatic bacteria’ fluorescent protein tags to proteobacteria using conjugation. Bio-protocol 9:7
Thoma S, Schobert M (2009) An improved Escherichia coli donor strain for diparental mating. FEMS Microbiol Lett 294(2):127–132
Remus-Emsermann MNP, Kim EB, Marco ML, Tecon R, Leveau JHJ (2013) Draft genome sequence of the phyllosphere model bacterium Pantoea agglomerans 299R. Genome Announc 1:1
Miller WG, Leveau JH, Lindow SE (2000) Improved Gfp and inaZ broad-host-range promoter-probe vectors. Mol Plant-Microbe Interact: MPMI 13(11):1243–1250
Schlechter RO, Kear EJ, Remus DM, Remus-Emsermann MNP (2021) Fluorescent protein expression as a proxy for bacterial fitness in a high-throughput assay. Appl Environ Microbiol 87(18):e0098221
Parkinson JS (2007) A ‘bucket of light’ for viewing bacterial colonies in soft agar. Methods Enzymol 423:432–435
Miebach M, Schlechter RO, Clemens J, Jameson PE, Remus-Emsermann MNP (2020) Litterbox—A gnotobiotic Zeolite-Clay System to Investigate Arabidopsis–Microbe Interactions. Microorganisms 8(4):464. https://doi.org/10.3390/microorganisms8040464
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S et al (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7):676–682
Burch AY, Do PT, Sbodio A, Suslow TV, Lindow SE (2016) High-level culturability of epiphytic bacteria and frequency of biosurfactant producers on leaves. Appl Environ Microbiol 82(19):5997–6009
Finkelshtein A, Roth D, Jacob EB, Ingham CJ (2015) Bacterial swarms recruit cargo bacteria to pave the way in toxic environments. mBio 6(3):e00074-15
Venturi V, Bertani I, Kerényi A, Netotea S, Pongor S (2010) Co-swarming and local collapse: quorum sensing conveys resilience to bacterial communities by localizing cheater mutants in Pseudomonas aeruginosa. PLoS ONE 5(4):e9998
Esser DS, Leveau JHJ, Meyer KM, Wiegand K (2015) Spatial scales of interactions among bacteria and between bacteria and the leaf surface. FEMS Microbiol Ecol 91:3
Morris JD, Hewitt JL, Wolfe LG, Kamatkar NG, Chapman SM, Diener JM, Courtney AJ, Matthew Leevy W, Shrout JD (2011) Imaging and analysis of Pseudomonas aeruginosa swarming and rhamnolipid production. Appl Environ Microbiol 77(23):8310–8317
Wang Q, Frye JG, McClelland M, Harshey RM (2004) Gene expression patterns during swarming in Salmonella typhimurium: genes specific to surface growth and putative new motility and pathogenicity genes. Mol Microbiol 52(1):169–187
Acknowledgements
The authors like to thank Sandra Hirsch for providing technical assistance.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
MK, RS, and MRE conceived the work. MK performed the experimental work. MK and MRE analyzed the data. RS and MRE supervised the work. LS provided material. MK and MRE wrote the initial draft of the manuscript with major input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kunzler, M., Schlechter, R.O., Schreiber, L. et al. Hitching a Ride in the Phyllosphere: Surfactant Production of Pseudomonas spp. Causes Co-swarming of Pantoea eucalypti 299R. Microb Ecol 87, 62 (2024). https://doi.org/10.1007/s00248-024-02381-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02381-4