Abstract
The Clostridium genus harbors compelling organisms for biotechnological production processes; while acetogenic clostridia can fix C1-compounds to produce acetate and ethanol, solventogenic clostridia can utilize a wide range of carbon sources to produce commercially valuable carboxylic acids, alcohols, and ketones by fermentation. Despite their potential, the conversion by these bacteria of carbohydrates or C1 compounds to alcohols is not cost-effective enough to result in economically viable processes. Engineering solventogenic clostridia by impairing sporulation is one of the investigated approaches to improve solvent productivity. Sporulation is a cell differentiation process triggered in bacteria in response to exposure to environmental stressors. The generated spores are metabolically inactive but resistant to harsh conditions (UV, chemicals, heat, oxygen). In Firmicutes, sporulation has been mainly studied in bacilli and pathogenic clostridia, and our knowledge of sporulation in solvent-producing or acetogenic clostridia is limited. Still, sporulation is an integral part of the cellular physiology of clostridia; thus, understanding the regulation of sporulation and its connection to solvent production may give clues to improve the performance of solventogenic clostridia. This review aims to provide an overview of the triggers, characteristics, and regulatory mechanism of sporulation in solventogenic clostridia. Those are further compared to the current knowledge on sporulation in the industrially relevant acetogenic clostridia. Finally, the potential applications of spores for process improvement are discussed.
Key Points
• The regulatory network governing sporulation initiation varies in solventogenic clostridia.
• Media composition and cell density are the main triggers of sporulation.
• Spores can be used to improve the fermentation process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a growing part of the world’s population is getting access to affordable energy, the global energy demand is increasing drastically. The energy consumed mainly originates from fossil resources, resulting in an acceleration of the depletion of natural resources and increased greenhouse gas (GHG) emissions. To inverse this trend, our societies are transitioning towards more sustainable economies, and countries worldwide are promoting the renewable energy sector. While substitutions to fossil energy generation processes such as hydrothermal, geothermal, solar, or wind energy are promoted, few alternatives to oil for freight, aviation, or the petrochemical sector are cost-effective. “Advanced biofuels,” defined by the International Energy Agency (IEA) as liquid or gaseous fuels derived from lignocellulosic (second-generation biofuel) or algal biomass (third-generation biofuel), are among the most promising substitutes to oil. These feedstocks have a relatively diverse composition and are available as complex polymeric structures (lignocellulose) that require dedicated enzymes to release the fermentable sugars. Some bacterial species from the Clostridium genus can hydrolyze these polymers, and ferment the carbohydrates to produce solvents. Clostridia are anaerobic and spore-forming Gram-positive bacteria, and the entire Clostridium genus currently comprises over 270 species (https://www.bacterio.net/), including pathogenic, probiotic, thermophilic, and benign soil bacteria. However, the Clostridium genus proposed by Prazmowski in 1880 (Prazmowski 1880) is not a monophyletic group (Collins et al. 1994; Jones 2001; Yutin and Galperin 2013), and only a subset of the 16S rRNA tree (cluster I), Clostridium sensu stricto is currently recognized as the genus’s representative (Cruz-Morales et al. 2019; Gupta and Gao 2009; Lawson and Rainey 2016). Several non-pathogenic clostridia have been studied for the production of advanced fuels and other biochemicals (Tracy et al. 2012). These species are commonly divided into acid-producing, solvent-producing, cellulolytic, and acetogenic species (Dürre 2005). These bacteria convert simple and complex carbon sources, from C1 compounds to cellulose, into a diverse range of metabolites, ranging from carboxylic acids such as acetate or butyrate to solvents like butanol and propanol. Ten Clostridium species are known to be solventogenic (Poehlein et al. 2017), see Table 1, with C. acetobutylicum, C. beijerinckii, C. pasteurianum, C. saccharobutylicum, and C. saccharoperbutylacetonicum being the most studied. These species have been used industrially during the twentieth century for acetone production through the ABE fermentation process (Berezina et al. 2012; Jones and Woods 1986; Jones 2001; Sauer 2016), but other solventogenic strains with high solvent productivity were isolated recently (Xin et al. 2018).
Despite many efforts, bioprocesses relying on these bacteria are not cost-effective due to high feedstock costs, expensive pretreatment of the feedstock, poor substrate use, and low solvent productivity (Green 2011; Tashiro et al. 2013). These issues need to be tackled to enable competitive biofuel prices. According to estimates, the product titer in a bioprocess aiming for the biofuel market needs to reach at least 50 g L-1, and the productivity should amount to 3 g L-1 h-1 to be commercially viable (Vees et al. 2020). While in standard batch conditions, solvent productivity has been reported to reach 10 g L-1 h-1 with clostridial fermentation, the clostridial maximum butanol titer in batch fermentation could not exceed 25 g L-1 without the application of in situ solvent removal techniques to prevent toxicity (Annous and Blaschek 1991; Qureshi and Blaschek 2001; Wang et al. 2019; Xu et al. 2015). It appears that both solvent toxicity and sporulation prevent the production of higher solvent titers (Cheng et al. 2019; Papoutsakis 2008). During the fermentation, bacteria are grown under stringent conditions, which are necessary for solvent production yet harsh for the cells. As a defense mechanism, the bacteria differentiate into highly resistant cells called endospores (henceforth designated as spores) while producing solvents. Spores are metabolically inactive, and their formation not only requires metabolic energy but also impairs solvent productivity (Tracy et al. 2012). Several efforts were made to engineer asporogenous solvent-producing strains, to prevent these undesirable effects (Al-Hinai et al. 2014; Bi et al. 2011; Scotcher and Bennett 2005). The regulation of sporulation and solventogenesis appears to be coupled, but the underlying regulatory networks remain unclear (Patakova et al. 2013). Due to the lack of efficient engineering tools for Clostridium, studies on the regulation of sporulation have been scarce. The recent development of markerless tools for clostridia (Atmadjaja et al. 2019; Cañadas et al. 2019; Diallo et al. 2020a; Huang et al. 2019; Joseph et al. 2018; Li et al. 2019; Seys et al. 2020; Wasels et al. 2020; Zhao et al. 2019) has made the genetic engineering of clostridia much more attainable. As a result, the number of engineered clostridia increased substantially and together with the rise of omics studies, the current knowledge on sporulation in solventogenic clostridia has expanded considerably.
Sporulation regulation in solventogenic clostridia
In response to changes in the environment, some bacteria produce spores to survive under unfavorable conditions. Depending on the formation mechanism and the structure, different spore types can be found in the environment (Paul et al. 2019). The spores, formed by Firmicutes, called endospores (Dürre 2014; Johnson 2019), are the most resilient. Endospores can survive harsh treatments such as high temperatures, the presence of oxygen (for anaerobic bacteria), desiccation, lysozyme incubation, ionizing radiation, and chemical solvents. The most studied sporulating bacteria belong to the Bacillus and Clostridium genera. The sporulation process was first described in Bacillus (Dawes and Mandelstam 1970; Kay and Warren 1968; Knaysi 1948; Tokuyasu and Yamada 1959), the model organism among spore formers, and the main features of its sporulation process are conserved in the Clostridium genus. Still, substantial differences in the spore morphology and sporulation initiation have been demonstrated between the two genera and within the Clostridium genus (Al-Hinai et al. 2015; Dürre 2014).
A sporulation model for the solventogenic clostridia (Al-Hinai et al. 2015) has been developed thanks to studies in C. acetobutylicum ATCC 824. Few studies were done on sporulation in other solventogenic clostridia strains to confirm the universality of this model. Although solventogenic clostridia are often presented as a homogenous group of bacteria, based on the first phylogenic studies on the Clostridium genus (Collins et al. 1994; Keis et al. 1995), this is not the case. Several strains were renamed and reclassified since 2000 (Keis et al. 2001), and recent phylogenic studies (Cruz-Morales et al. 2019; Yu et al. 2019) show that C. beijerinckii and C. acetobutylicum even belong to two different clades. Out of the seventeen clades dividing the Clostridium genus “sensu stricto,” solventogenic clostridia can be found in two groups, one harboring C. acetobutylicum and C. pasteurianum and another consisting of C. beijerinckii, C. saccharoperbutylacetonicum, and C. saccharobutylicum. Phylogenetically, C. beijerinckii is closer to the human pathogens C. perfringens and C. botulinum E than to the model solventogenic clostridia C. acetobutylicum, as depicted in Fig. 1. In line with what has been suggested for toxin genes (Cruz-Morales et al. 2019), solventogenesis genes might have been acquired by horizontal transfer. The localization of the sol operon, harboring the essential solvent genes, on a megaplasmid in C. acetobutylicum contrasting with the chromosomal sol operon in other clostridia supports this hypothesis. Thus, the regulation mechanisms described for C. acetobutylicum might not be identical in C. beijerinckii or other solventogenic species (Patakova et al. 2013).
Phylogenetic tree of the Clostridium sensu stricto group, amended from (Lawson and Rainey 2016) with permission of the Microbiology Society. The tree was reconstructed using the neighbor-joining method based on the pairwise comparison of approximately 1340 nt. Atopobium parvulum was used as the outgroup. Bootstrap values (> 90 %), expressed as a percentage of 1000 replications. Bar, 1 % sequence divergence. A blue disc next to the name of a The strains next to in the blue discs and purple boxes belong indicates that this strain belongs to the solventogenic group, while a green disk next to the name of a strain indicates that this strain belongs to the and the acetogenic clostridia
The sporulation cycle
The initiation of sporulation
The sporulation regulation network described in Bacillus has been used as a template for understanding the sporulation cascade in Clostridium (Davidson et al. 2018; Decker and Ramamurthi 2017; Paredes et al. 2005; Piggot and Hilbert 2004). However, several sporulation genes identified in Bacillus are absent in Clostridium, indicating a difference in the molecular regulation. Genome comparison studies were conducted to identify homologs to the sporulation genes studied in B. subtilis and the necessary set of genes required for sporulation (Galperin 2013; Galperin et al. 2012; Traag et al. 2013). Fifty-two genes were identified as essential for sporulation as they were found both in sporulating clostridia and bacilli by comparing more than 217 genomes of sporulating and non-sporulating Firmicutes. Even when gene homologs are found in Clostridium, their functions are not always identical to their homologs in Bacillus (Al-Hinai et al. 2015; Fimlaid and Shen 2015). Genome comparison, coupled with transcriptomic studies, enabled the identification of additional sporulation genes in solventogenic clostridia (Grimmler et al. 2011; Lee et al. 2015; Máté de Gérando et al. 2018; Sedlar et al. 2018). The role of the central sporulation regulators and sigma factors were studied in solventogenic clostridia through the generation of asporogenous mutants, mainly in C. acetobutylicum ATCC 824 and more recently in other solventogenic clostridia (Table 2).
The regulation model of sporulation in Clostridium is divided, like in Bacillus, into seven stages associated with morphological changes of the cell. In most clostridia, the sporulation process coincides with granulose accumulation and ends with the lysis of the mother cell and the release of the spores in the environment (Fig. 2). Sporulation is initiated at the end of vegetative growth and is reflected at the transcriptomic level by an increase in spo0A expression. Spo0A, the general regulator of the transition from vegetative to stationary growth, is conserved in all Firmicutes and has a central role in sporulation, toxin, and solvent production (Al-Hinai et al. 2015; Dürre 2014; Jones et al. 2008; Paredes et al. 2005; Ravagnani et al. 2000; Sauer et al. 1995).
Morphological changes and molecular regulation of sporulation in C. acetobutylicum modified from (Al-Hinai et al. 2015). The regulation of the sporulation process is mainly realized by the modulation of the transcription in each compartment. Post-translational regulation enables the activation of Spo0A and sporulation-specific transcription factors (σF, σG, σE). The activation mechanism of σG has not been investigated in C. acetobutylicum. Inactive transcriptional regulators are in grey, and active transcriptional regulators are in orange. DPA, dipicolinic acid; Sasps, small acid-soluble proteins. Black arrows indicate post-translational regulations, blue arrows transcriptional regulation. Arrows with short dashes indicate interactions described only in C. acetobutylicum, arrows with long dashes observed only in C. beijerinckii, and full arrows indicate interactions described in C. acetobutylicum and other clostridia
In Bacillus, at the end of vegetative growth, AbrB lifts its repression on σH; σH then promotes the transcription of spo0A (Piggot and Hilbert 2004). In solventogenic clostridia, the role of σH in spo0A regulation is unknown. The gene encoding σH is constitutively expressed in clostridia, which implies that the mechanism occurring in Clostridium differs from the one described in Bacillus. Several AbrB homologs were identified in solventogenic clostridia, in C. acetobutylicum ATCC 824, three homologs (cac0310, cac1941, and cac3647) were disrupted to study their role in the regulation of cellular events. The disruption of the most expressed AbrB homolog, cac0310, delayed sporulation and impaired solvent production (Scotcher et al. 2005). The disruption of cac3647 increased solvent production, while the solvent production of Δcac1941 cultures decreased by 6% compared to the wild type (Xue et al. 2016). No change in the sporulation of the Δcac1941 or Δcac3647 mutant was reported. According to these results, AbrB homologs belong to the sporulation and solvent regulation network in solventogenic clostridia. Unlike in Bacillus, these results indicate that AbrB (Cac0310) may promote sporulation in C. acetobutylicum. The Spo0A DNA binding motif, called 0A box, was found upstream of cac0310 and its homolog in C. beijerinckii NCIMB 8052, cbei4885, indicating that these abrB homologs belong to the Spo0A regulon. In fact, the disruption of this 0A box might impair solvent production and sporulation; one of the SNPs detected in asporogenous offspring of C. beijerinckii SA-1 (Table 2) was an SNP in the 0A box of cbei4885 (Seo et al. 2017a).
Mutations in the spo0A coding sequence of solventogenic clostridia have been shown to affect cell physiology considerably. Changes in growth, colony morphology, sporulation, and solvent productivity have been reported. Numerous spo0A mutants have been characterized (Table 2), and all have impaired sporulation with sometimes a change in solvent productivity (Atmadjaja et al. 2019; Harris et al. 2002; Seo et al. 2017b). The consequences for the phenotype seem to depend on the mutation location and the studied species. The Spo0A sequence harbors various domains that are conserved among Firmicutes and which are putatively involved in sporulation regulation (Fig. 3). Insertional mutations were shown to impair the protein function, blocking the central role of Spo0A in solventogenesis and sporulation regulation (Harris et al. 2002; Wilkinson and Young 1994 ). However, due to the gene engineering techniques used, the impacts of polar effects on the phenotype cannot be excluded (Bayat et al. 2018), but with the development of markerless gene engineering methods, precise mutations of spo0A were generated.
Mutations and associated phenotypes of markerless spo0A- strains. Five conserved regions are indicated by colored boxes; R: signal receiver domain, Green box: conserved region with no known function, HTM : helix turn motif, σA: putative σA activator region, σH: putative σH activator region. The ability to sporulate, to form heat resistant spores, and to produce solvent is indicated next to the scheme of the spo0A mutation present in each mutant strain; - : indicate a decrease compared to the wild type phenotype; -- : indicate the abolition of the feature in the mutant compared to the wild type phenotype; / : indicate that this characteristic could not be evaluated ; ++ indicate an increase compared to the wild type phenotype and NA stands for no data available
Mutations (deletions and single nucleotide modifications) in the putative σ factor activator domains did not cause a decrease in solvent production, as shown in Fig. 3. Like in Bacillus subtilis (Schmeisser et al. 2000), a mutation in the putative σH activator region disrupted the sporulation of C. saccharoperbutylacetonicum (I261T). In Bacillus, sporulation impairment was explained by a loss in the binding affinity of Spo0A for spoIIE and spoIIA promoters. Mutations in the region upstream from the DNA binding domain impaired solvent production and decreased sporulation efficiency in C. acetobutylicum and C. saccharobutylicum (Foulquier et al. 2019). In C. pasteurianum, a deletion in the σA activator region disrupted sporulation and increased solvent production (Sandoval et al. 2015). Surprisingly, in C. pasteurianum, the deletion of most of the gene (816 bp out of 822) led to the same phenotype (Schwarz et al. 2017). Spo0A seems to play a different role in sporulation regulation in C. pasteurianum.
Next to the integrity of the spo0A gene, intracellular Spo0A levels appear to also play a role in sporulation regulation. In the non-sporulating strain, Clostridium sp. MF28, relatively low spo0A expression levels were detected, compared to other solventogenic clostridia, even though high solvent titers could be reached (Li and He 2016). In C. acetobutylicum, spo0A overexpression increased sporulation independently from the promoter region used (Ehsaan et al. 2016; Harris et al. 2000; Tracy et al. 2008). In contrast, in C. beijerinckii, spo0A overexpression led to a decrease in both sporulation and solvent production (Kolek et al. 2017). A slight change in Spo0A homeostasis might lead to a different regulation of sporulation and solventogenesis. No detailed study on the variation of active Spo0A and its impact on sporulation has been done in solventogenic clostridia to confirm this hypothesis. In Bacillus, depending on the intracellular concentration of phosphorylated Spo0A, differences in the expression of the Spo0A regulon were described (Fujita et al. 2005; Narula et al. 2012).
Once spo0A is transcribed and translated, Spo0A is activated by at least two post-translational modifications: acylation and phosphorylation, as illustrated in Fig. 4. Protein acylation is a post-translational modification that consists in adding an acyl group to a lysine residue. This reaction is reversible and does not need the intervention of an enzyme. Acylation neutralizes the negative charge of the lysine residue, altering the protein structure and its interaction with other proteins, cofactors, or substrates (Macek et al. 2019). Interestingly, Spo0A acylation was only reported in C. acetobutylicum (Xu et al. 2018). In Bacillus, acylation was also linked with sporulation (Kosono et al. 2015), but it was observed only for late-sporulation stage proteins (CotE, CotO, Cse15, SpoIVD, and SpoVR). Xu et al. showed in their study in C. acetobutylicum that several key proteins involved in metabolism and life cycle, such as Buk and Spo0A, were butyrylated during cultivation (Xu et al. 2018). Two butyrylation sites were detected in the Spo0A sequence, one close to the phosphorylated domain and another on the DNA binding domain, and replacing the lysine residue (K217) located in the DNA binding domain with glutamine decreased the DNA binding affinity of Spo0A. While the wild-type Spo0A could bind to its binding motif in the promoter region of Spo0A, the mutated Spo0A could not. This result suggests that Spo0A butyrylation is necessary for Spo0A activity and its autoregulation.
Transcriptional and post-translational regulation of Spo0A in C. acetobutylicum, C. beijerinckii, C. saccharoperbutylacetonicum. Figure adapted from (Al-Hinai et al. 2015) with results from (Feng et al. 2020; Kotte et al. 2020; X. Xin et al. 2020; J.-Y. Xu et al. 2018; Y. Yang et al. 2020). Blue arrows transcriptional regulation and black arrows post-translational regulation. The interrogation marks indicate interactions remain to be proven experimentally
In Clostridium, the phosphorylation mechanism activating Spo0A also contrasts with the phosphorelay system described in B. subtilis (Paredes et al. 2005), in that it relies on orphan histidine kinases, as illustrated in Fig. 4. Orphan histidine kinases are part of two-component quorum sensing systems (TCS). TCS usually consists of a sensor histidine kinase and a response regulator located in the same operon. However, isolated genes encoding sensor histidine kinases or response regulators were identified in the genome of numerous bacteria and especially in Firmicutes. Since these genes are not co-located with a gene encoding a specific sensor protein or response regulator, they are labeled orphans (Davidson et al. 2018; Williams and Whitworth 2010). Unlike regular TCS kinases, orphan histidine kinases can modulate several response regulators. In clostridia, orphan histidine kinases are integrated for signal transduction to the sporulation regulation network (Freedman et al. 2019; Paredes-Sabja et al. 2014; Steiner et al. 2011; X. Xin et al. 2020). In C. acetobutylicum, the impact of four histidine kinases (Cac0323, Cac0437, Cac0903 and Cac3319) on sporulation has been studied (Steiner et al. 2011). In vitro studies proved that two of these kinases, Cac0903 and Cac3319, are able to activate Spo0A. The disruption of cac0323, cac0903, and cac3319 resulted in a decreased sporulation, in contrast to cac0437 deficient strains hyper-sporulated (earlier sporulation and a fifteen-fold increase in heat resistant spores). In C. beijerinckii NCIM 8052, six homologs of these kinases were disrupted in a recent study (Xin et al. 2020). A significant decrease in sporulation efficiency (between 70 and 90%) coupled with an increase in solvent production of 38% and 14% were reported only for the ∆cbei2073 and ∆cbei4484 mutants, respectively. Interactions between these histidine kinases and Spo0A remain to be studied. Recently Seo et al. analyzed the genome of the hyper-butanol-producing mutant BA101 strain and identified mutations in the genome that could explain its reduced sporulation (Seo et al. 2021). Two mutations could explain their asporogenous phenotype; one in a gene encoding a histidine kinase (cbei_3078) and a second mutation in a gene coding for a serine/threonine protein phosphatase-like protein (cbei_4400). Similarly, the mutation in cbei_4400 was also reported in SA-1, another hyper-producing C. beijerinckii strain with delayed sporulation (Sandoval-Espinola et al. 2013).
Once phosphorylated, Spo0A binds to the 0A box to regulate the expression of sporulation- and solvent genes (Ravagnani et al. 2000; Zhao et al. 2002). Differences in the motif sequence (Ravagnani et al. 2000), the number (Sullivan and Bennett 2006), and the location of these 0A boxes (Patakova et al. 2013) between solventogenic clostridia were detected. These discrepancies further illustrate the variations in the role of Spo0A in solventogenic clostridia.
Stages of sporulation
In Fig. 2, a scheme of the stages of sporulation and phenotypic changes during sporulation is shown. Stage I of the sporulation process starts with the DNA replication and the positioning of Z-rings, close to the poles, to prepare for asymmetric division (Barák et al. 2019). Solvent production is initiated, and in most clostridia, Stage I coincides with a morphological change. The cell swells due to the accumulation of a starch-like polymer called granulose. In C. acetobutylicum, granulose and sporulation are regulated by an Agr quorum system (Steiner et al. 2012). In other solventogenic clostridia, only one study links granulose production and sporulation (Ravagnani et al. 2000). While investigating the role of Spo0A in C. beijerinckii, Ravagnani et al. discovered that Spo0A was essential for the accumulation of granulose. Still, granulose accumulation was not described in all solventogenic clostridia; C. tetanomorphum, for example, does not produce any granulose during the sporulation process (Patakova et al. 2014).
Once activated, Spo0A promotes the expression of spoIIE, triggering the entrance in stage II of the sporulation process. During Stage II, a septum forms on one pole of the cell dividing it into two compartments, the forespore and the mother cell. In Bacillus, this asymmetric division is orchestrated by SpoIIE and the cell division proteins (involved in binary fission) (Barák et al. 2019). SpoIIE simultaneously activates σF, the first sporulation-specific sigma factor, which is kept inactive by SpoIIAB. SpoIIE phosphorylates SpoIIAA, which binds the anti-sigma factor SpoIIAB which then releases σF in the forespore. In solventogenic clostridia, the role of SpoIIE was studied in C. acetobutylicum and C. beijerinckii through the generation of SpoIIE deficient mutants (Bi et al. 2011; Diallo et al. 2020b; Scotcher and Bennett 2005). Both mutants could no longer sporulate, but phenotypical differences between C. beijerinckii ΔspoIIE and C. acetobutylicum ΔspoIIE were noted. In C. acetobutylicum, spoIIE disruption prevented the formation of an asymmetric septum, while in C. beijerinckii, misplaced septa were observed. The morphology of the C. beijerinckii mutant corresponded to the spoIIE mutants described for Bacillus. This discordance in the mutants’ morphology indicates differences in the asymmetric septation mechanism of solventogenic clostridia. Another difference was observed; no critical change in the expression of sigF and sigE was detected in C. beijerinckii ΔspoIIE, contrasting with C. acetobutylicum ΔspoIIE, where sigF and sigE were downregulated.
The completion of asymmetric division marks the entrance in Stage III, during which the septum is prolonged and surrounds the whole forespore. This phenomenon, called engulfment, yields an isolated compartment surrounded by two membranes within the mother cell. Engulfment is coordinated by proteins belonging to σF (in the forespore) and σE regulons (in the mother cell). The roles of σF and σE were studied only in C. acetobutylicum ATCC 824, where sigF and sigE mutants were generated. Once again, these mutants’ cell morphology did not correspond to the morphology of sigF and sigE mutants generated in Bacillus or other clostridia (Al-Hinai et al. 2015). In both C. acetobutylicum mutants, the sporulation process was interrupted before the formation of the asymmetric septum, as observed for the spoIIE mutant. This suggests an earlier function of σF and σE in sporulation regulation (Jones et al. 2011; Tracy et al. 2011). The disruption of either sigF or sigE affected solventogenesis. Indeed, when mid to late exponential cells were inoculated for fermentation, solvent production decreased significantly.
After engulfment, the coat and the spore cortex are formed during Stage IV and V. Dipicolinic acid (DPA) is produced in the mother cell through the conversion of aspartate, and then transported into the forespore, in exchange for water, to bind Ca2+ in the forespore (Piggot and Hilbert 2004). Ca2+-DPA attaches to the forespore’s DNA to protect it against heat damages (Jamroskovic et al. 2016; Paidhungat et al. 2000; Paredes-Sabja et al. 2008). In the meantime, coat proteins assemble around the mother-cell-derived membrane of the forespore (Shen et al. 2019). All these events are coordinated by proteins regulated by σG in the prespore and σK in the mother cell. In contrast to pathogenic clostridia, these late stages of sporulation are barely studied in solventogenic clostridia. Nonetheless, σK and σG deficient mutants in C. acetobutylicum confirmed their crucial role in sporulation (Al-Hinai et al. 2014; Tracy et al. 2011). Disruption of sigG interrupted sporulation after engulfment as described in Bacillus and did not affect solventogenesis. The disruption of sigK, though, did not yield the same phenotype as the Bacillus mutant. In C. acetobutylicum, σK regulates sporulation initiation and sporulation maturation, as described in other clostridia (Al-Hinai et al. 2015). No Spo0A proteins were detected in sigK mutants of C. acetobutylicum, and the introduction of an extrachromosomal copy of spo0A under control of the ptb promoter led to the formation of heat-sensitive spores.
During Stages VI and VII, the spore matures as the size of both cortex and coat increases. Sporulation finishes with the lysis of the mother cell and the release of the spore in the environment. A study on an autolysin deficient mutant, C. acetobutylicum lyc::int(72), generated in C. acetobutylicum ATCC 824, showed that autolysins are needed to complete sporulation (Liu et al. 2015). The number of viable spores produced by C. acetobutylicum lyc::int(72) decreased by 30% compared to the wild-type strain. According to the authors, cell lysis might provide additional nutrients to sporulating cells and thus be required for successful sporulation.
Only a few studies on the molecular regulation of sporulation in solventogenic clostridia have been published (Al-Hinai et al. 2014; Bi et al. 2011; Diallo et al. 2020b; Jones et al. 2008, 2011; Ravagnani et al. 2000; Scotcher et al. 2005; Scotcher and Bennett 2005; Steiner et al. 2011, 2012; Tracy et al. 2011). Studies were mainly realized in C. acetobutylicum ATCC 824, and studies with other solventogenic strains show variations in the role of Spo0A and SpoIIE in the regulation of sporulation. Moreover, the post-translational regulation for sigma factor activation, observed in Bacillus and Clostridioides (former Clostridium) difficile, still needs to be investigated in solventogenic clostridia. Thus, several grey areas remain concerning the regulatory mechanisms controlling the sporulation cascade in solventogenic clostridia.
The clostridial endospore
At the end of sporulation, an endospore is released into the environment. The endospore is highly dehydrated and organized in proteinous layers protecting the core, which hosts the DNA. Five layers surround the core: the inner membrane, the germ cell wall, the cortex, the outer membrane, and the coat (Fig. 5b). These layers ensure a robust protection of the core against chemicals, oxygen, enzymes, and heat. In the core, the DNA is bound to small acid-soluble proteins (Sasps), ribosomes, enzymes, and DPA. The DPA content can reach up to 25% of the spore’s dry weight (Paredes-Sabja et al. 2014).
Morphology and composition of the clostridial endospore; a Transmission electron micrographs of mature endospores from three solventogenic clostridia. Depending on the species, the size of the cortex, as well as the location of the endospore in the mother cell, changes; b Possible location of the endospore in solventogenic clostridia; c Composition of the endospore. EX stands for exosporium
Spores are characterized by their size, shape, and location in cells (Dürre 2005, 2014). There is not a typical morphology for all clostridial spores, and few studies have been done to compare the spore morphology of clostridial strains. One study (Berezina et al. 2012) compared the spore morphology of the four main solventogenic species (C. acetobutylicum, C. beijerinckii, C. saccharobutylicum, and C. saccharoperbutylacetonicum) and showed that they all possess an oval shape. This characteristic cannot be generalized to all solventogenic clostridia since C. tetanomorphum yields round-shape spores (Patakova et al. 2014). The spore’s location may vary within the mother cell (Fig. 5a, b) from eccentric to terminal (Dürre 2005).
The spore coat (Long et al. 1983), the cortex, and even the core composition are species- and even sometimes strain-specific. For instance, differences in the spore’s DPA and Ca2+ content were detected, resulting in a difference in heat resistance and that the Ca2+/DPA ratio was not species but strain-specific (Jamroskovic et al. 2016) showed that a high Ca2+/DPA ratio correlates with more resistant spores. C. acetobutylicum ATCC 824 spores can sustain a long heat treatment (> 5 min) at 70 oC, while C. beijerinckii NCIMB 8052 spores germinate better with a short heat treatment (1 min) at higher temperatures (around 90 oC). The heat-shock treatment has to be adapted to the species to give the highest germination efficiency. Knowledge of these differences is crucial as it affects germination efficiency after a heat-shock treatment before fermentation (Jabbari et al. 2013; Li et al. 2011; Steiner et al. 2012).
Sporulation triggers
The sporulation is known to be a response to stressful conditions. The primary triggers of sporulation in Bacillus species have been shown to be nutrient starvation and high cell density (Awang et al. 1992; Basu et al. 2017). In contrast to bacilli, where the cessation of growth occurs due to a lack of nutrients, in Clostridium, an excess of a carbon or nitrogen source led to growth cessation (Dürre 2005; U. Sauer et al. 1995; Woods and Jones 1987). Furthermore, unlike bacilli, clostridia are anaerobic bacteria; thus, oxygen is a significant stress factor for the cells and triggers sporulation.
Several studies (Awang et al. 1992; Kolek et al. 2017; Long et al. 1983; Sedlar et al. 2021; Woods and Jones 1987) have reported the substantial impact of medium composition on sporulation initiation and sporulation efficiency (number of spore generated and the spores that can germinate) and suggested a link between carbon source, mineral content, external pH, and the number of spores produced.
The carbon source
Solventogenic clostridia can ferment several carbohydrates (including C6 and C5 sugars and sugar polymers such as starch or xylan), yet, the solvent yield varies depending on the carbon source (Awang et al. 1992; Shaheen et al. 2000). Similarly, sporulation efficiency depends on the carbon source, as described for C. saccharobutylicum NCP 262 (Long et al. 1983). A complementary study with the same strain assessed the effect of 13 different carbon sources (carboxymethylcellulose, xylan, inulin, a starch/glucose mix, lactose, cellobiose, sucrose, maltose, glucose, mannose, fructose, galactose, and xylose) on both sporulation and solvent formation (Awang et al. 1992). Depending on the carbohydrate, sporulation frequency varied up to 44%, with glucose utilization leading to the most spores. Likewise, when grown on rhamnose, a decrease in spore formation was observed in C. beijerinckii cultures (Diallo et al. 2018). Recently Sedlar et al. observed that C. beijerinckii NCIMB 8052 and C. beijerinckii NRRLB 598 sporulated when glycerol was added to the medium instead of glucose (Sedlar et al. 2021). Surprisingly the opposite was observed in C. beijerinckii (formerly C. diolis) DSM 15410 cultures. The substrate may also accelerate sporulation initiation. For instance, xylose-fed cultures of C. acetobutylicum BOH3 sporulated earlier than glucose-fed cultures (Basu et al. 2017). In addition to the nature of the carbohydrate, its concentration also affects sporulation. High glucose concentrations doubled the number of endospores generated by C. saccharobutylicum NCP 262 (Long et al. 1984a).
These observations have been confirmed by studies on carbon metabolite repression in clostridia. The carbon catabolite repression protein A (CcpA) regulates the carbon catabolite repression in Firmicutes and is involved in triggering sporulation in pathogenic clostridia. CcpA activates (Varga et al. 2004) or represses sporulation (Antunes et al. 2011) depending on the species. Still, in C. acetobutylicum, CcpA positively regulates sporulation (Ren et al. 2012). In a CcpA deficient strain, sporulation was delayed, and the sporulation efficiency decreased (Table 2). Besides, transcriptome analysis of the mutant strain performed by Ren et al. suggested that CcpA represses abrB homologs and promotes the expression of the sporulation-specific sigma factors (sigE, sigG, sigK). This modification of the expression profile of those sporulation-related genes might have altered sporulation in the CcpA deficient C. acetobutylicum strain.
Other media components
Solventogenic clostridia are currently investigated for their potential to produce solvents from complex feedstock such a lignocellulosic and algal feedstocks. Pretreatment of these feedstocks is necessary to the utilization by the bacteria of the carbohydrates present. During the pretreatment, di- and monosaccharides, as well as inhibitory chemicals (salts, furfurans and phenolic compounds), are formed. Studies have been conducted to evaluate the impact of these toxic compounds on cell growth and solvent formation. Hence, few reports on their effect on sporulation can be found; still, three transcriptomic studies of C. beijerinckii and C. acetobutylicum cultures exposed to phenolic compounds detected changes in the expression of sporulation genes. Exposure to ferulic acid (Lee et al. 2015) and syringaldehyde (Ezeji et al. 2007) caused an upregulation of the late-stage sporulation genes C. beijerinckii. In C. acetobutylicum, a recent study showed through a gene coexpression network analysis (Liu et al. 2020) that exposure to vanillin and p-coumaric acid disturbed the transcription of early sporulation genes (spo0A, spoIIE, spoIIP) and sporulation specific sigma factors.
Acids, various metals and minerals, vitamins, and amino acids also affect both solvent production and sporulation in clostridia (List et al. 2019; Mukherjee et al. 2019; Nimbalkar et al. 2018, 2019; Reeve and Reid 2016), but few studies mentioned their impact on sporulation. Long et al. (1984a) investigated butyrate and acetate’s effect on sporulation in C. saccharobutylicum by adding them to the medium at the start of the fermentation in different concentrations. Even though the addition of acids was not necessary for sporulation, it increased the number of spores present in the culture by 40 to 100% for concentrations between 1 and 4 g L-1.
As for the impact of other media components on sporulation, one study reports that the addition of adenine in the media caused a 20-h delay in the onset of sporulation in C. saccharoperbutylacetonicum cultures (Kiyoshi et al. 2017). It is worth noting that, depending on the species, a compound can have an opposite impact on sporulation. For example, in C. perfringens, iron is necessary for sporulation (Lee et al. 1978), while its addition to the medium impairs sporulation in C. sporogenes (Mah et al. 2008).
Solventogenic clostridia harbor sporulation proteins requiring metal-containing cofactors; thus, the media’s metal content is expected to impact sporulation regulation. For instance, homologs of SpoIIE, SpoIIQ and CsfB, an anti-sigma factor of σE and σG, were identified in Clostridium. In Bacillus, their activity requires Mn2+and Zn2+ respectively (Król et al. 2017; Martínez-Lumbreras et al. 2018). In Bacillus, Mn2+ was proven to be crucial for SpoIIE’s phosphatase activity and the oligomerization of SpoIIE, and thus, asymmetric division (Król et al. 2017). Mn2+ has also been reported to be key for the development of heat-resistant spores in C. botulinum (Lenz and Vogel 2014). In C. difficile, Zn2+ is necessary for the formation of the SpoIIQ-SpoIIIAH complex, involved in engulfment and essential for the transit of molecules between mother cell and forespore (Serrano et al. 2016). Zn2+ was reported to promote sporulation in C. botulinum (Kihm et al. 1988) but to inhibit sporulation of C. sporogenes when the concentration in the medium exceeds 3.7 mM. (Lee et al. 2011). Ca2+ is also involved in spore formation since it forms together with dipicolinic acid (DPA), and several studies have shown that Ca2+is a crucial component for spore heat resistance (Church 1959; Huang et al. 2007; Jamroskovic et al. 2016; Mah et al. 2008; Paredes-Sabja et al. 2008).
Studies of the transcriptome of wild-type and mutant C. beijerinckii cultures during fermentation indicated changes in the expression of genes involved in ion- and amino acid transport at sporulation initiation. In C. beijerinckii NRRL B598, sporulation initiation was concomitant with an upregulation of the genes encoding a magnesium transporter, and an upregulation of genes encoding potassium, sodium, and iron transporters was detected during stationary phase (Vasylkivska et al. 2019). In cultures of the asporogenous C. beijerinckii ΔspoIIE strain, the expression of genes encoding iron transporters were downregulated during stationary phase (Diallo et al. 2020b), indicating a potential role of iron in sporulation.
Metabolite concentration
Metabolite stress has been suggested to trigger sporulation in solventogenic clostridia (Heluane et al. 2011; Sauer et al. 1995; Tomas et al. 2004; Zheng et al. 2009). When grown in batch reactors, solventogenic clostridia ferment the available carbohydrates into carboxylic acids, mainly acetate and butyrate, which accumulate in the culture and cause a drop in pH (Fig. 6).
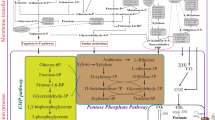
Simplified acetone-butanol-ethanol metabolic pathway in solventogenic clostridia. Some strains harbor a secondary alcohol dehydrogenase (s-adh) that enables the formation of isopropanol. In C. acetobutylicum, the acidogenic phase and solventogenic phase succeed each other during the fermentation. During exponential growth, the substrate is metabolized to form lactate, acetate, and butyrate during the acid phase. At stationary phase, the acids are reassimilated, and the culture produces ethanol, acetone (or isopropanol), and butanol. However, in other solventogenic species (C. beijerinckii for example), acidogenic and solventogenic phase occur concomitantly and solvent production starts during the exponential phase. The enzymes involved in the metabolic pathway are in grey boxes: pta, phosphotransacetylase; ack, acetate kinase; thl, thiolase; hbd, 3-hydroxybutyryl-CoA-dehydrogenase; crt, crotonase; bcd, butyryl-CoA-dehydrogenase; ctfA/B, CoA-transferase; buk, butyrate kinase; ptb, phosphotransbutyrylase; adh, aldehyde/alcohol dehydrogenase; edh, ethanol dehydrogenase; adc, acetoacetate decarboxylase; s-adh, secondary alcohol dehydrogenase; ald, butyraldehyde dehydrogenase; bdh, butanol dehydrogenase and -P stands for phosphate. As indicated by its title, this figure represents a simplified ABE pathway; indeed, studies have shown a diversity in the structure and number of enzymes involved depending on the solventogenic species
At the onset of solventogenesis and sporulation, acids are reassimilated and converted into solvents, resulting in a rise of pH in the culture. While solvent formation enables short-term relief from the pH stress, sporulation is regarded as a long-term stress response mechanism, protecting the cells from metabolic stress and interrupting sugar degradation (U. Sauer et al. 1995). Acetate and butyrate accumulation during exponential growth is proposed to trigger both solventogenesis and sporulation (Sauer et al. 1995; Thorn et al. 2013). In C. acetobutylicum, a peak in the intracellular undissociated acid concentration was observed at the start of the solventogenesis (Terracciano and Kashket 1986; Yang et al. 2013). Characterization of C. acetobutylicum recombinants deficient in phosphotransbutyrylase (Ptb), butyrate kinase (Buk) and acetate kinase (Ack) (Desai and Papoutsakis 1999; Green et al. 1996; Harris et al. 2000; Kuit et al. 2012) suggested that instead of the concentration of undissociated acids, the intracellular concentration of butyryl phosphate (BuP) might trigger both sporulation and solventogenesis. BuP is an intermediary metabolite in the ABE metabolic pathway (Fig. 6), formed during acidogenesis during butyrate formation. Studies evaluating the intracellular concentration of BuP during batch cultivation showed that BuP was indeed accumulated in the cell (Xu et al. 2018; Zhao et al. 2005). Two peaks in the cytoplasmic BuP concentration were detected inside the cells, one at the beginning of the cultivation and a second one coinciding with solventogenesis and sporulation initiation (Y. Zhao et al. 2005). It was suggested that BuP acted as a phosphate donor enabling the activation of Spo0A, the master regulator of sporulation and solventogenesis (Kuit 2013; Zhao et al. 2005), but recent data (Xu et al. 2018) present another post-translational regulation mechanism: protein butyrylation (see section on the initiation of sporulation).
Butanol has also been suspected of triggering sporulation (Zheng et al. 2009). Even though solventogenic clostridia naturally produce butanol, it affects cell growth when its concentration exceeds 0.5% v/v in the culture (Sedlar et al. 2018) and becomes lethal, around 1.5% v/v (Sedlar et al. 2019). Butanol concentration being a stress factor for the cells, researchers supposed that a rise in butanol concentration would initiate sporulation before it reaches toxic concentration. However, a decrease in granulose and spore number was observed in butanol stressed C. beijerinckii cultures (Sedlar et al. 2019). Transcriptional studies on butanol stressed cultures of C. acetobutylicum and C. beijerinckii showed no notable changes in the expression of the genes encoding the sporulation-specific sigma factors (sigF, sigE, sigG, sigK) (Patakova et al. 2019; Tomas et al. 2004). In C. acetobutylicum ATCC 824, no butanol-dependent impact on sporulation efficiency was described. Instead, a decrease in the expression of genes encoding small acid-soluble proteins was observed (Schwarz et al. 2012; Tomas et al. 2004). These proteins protect the DNA present in the spores and are crucial for their heat resistance (Leggett et al. 2012).
Secondary metabolites have also been reported to promote sporulation in solventogenic clostridia. Two categories of secondary metabolite biosynthesis gene clusters were identified in solventogenic clostridia (Letzel et al. 2013), polyketide- and ranthipeptide biosynthesis clusters. Polyketides have been studied in C. acetobutylicum and C. saccharoperbutylacetonicum. In C. saccharoperbutylacetonicum, polyketides involved in sporulation initiation, solvent formation, and tolerance were detected (Kosaka et al. 2007; Li et al. 2020a). In C. acetobutylicum, three polyketides were detected, and the structures of two of them, clostrienose and clostrienoic acid, were solved (Herman et al. 2017). In both species, the disruption of polyketide clusters decreased sporulation (Table 2). In C. beijerinckii, polyketides might also intervene in the regulation of sporulation; the interruption of sporulation in C. beijerinckii ΔspoIIE affected the expression of the polyketide gene cluster (Diallo et al. 2020b). Recently, the role of ranthipeptides, secondary metabolites belonging to the ribosomally synthesized and post-translationally modified peptide (RiPP) superfamily, was studied in C. beijerinckii and C. ljungdahlii (Chen et al. 2020). In C. beijerinckii, the genes encoding the precursor peptide and the radical SAM protein were disrupted, and the impact on the transcriptome was evaluated by RNA sequencing. In the mutant strain, sporulation genes were strongly downregulated and the agr locus encoding the Agr quorum sensing mechanism was upregulated. Secondary metabolites seem to play an important role in the initiation of sporulation, even so the interactions between the polyketides and ranthipeptides with sporulation regulators remain to be investigated.
Quorum sensing and cell density
As described for B. subtilis (Bischofs et al. 2009; Grossman and Losick 1988), cell density might regulate sporulation in solventogenic clostridia. In Bacillus, a minimum cell density was required for efficient sporulation (Grossman and Losick 1988; Hecker and Völker 2001). Similarly, in continuous cultures of Clostridium, where the specific dilution rate and cell morphology can be monitored, a decrease in the number of sporulating cells was observed when the dilution rate was raised (Heluane et al. 2011). Thanks to quorum sensing mechanisms, cells can monitor environmental changes such as cell density and launch their adaptation response when required. Two quorum sensing mechanism superfamilies were described in Gram-positive bacteria (Aframian and Eldar 2020): the membrane receptor family (TCS) and the cytoplasmic receptor family (RRNPP). Systems belonging to both families were found in solventogenic clostridia (Fig. 7). Two TCS, an Agr system and the BtrK/BtrR system, were described in C. acetobutylicum ATCC 824 (Steiner et al. 2012; Yang et al. 2020). The Agr system regulates both granulose formation and sporulation. In contrast, the BrtK/BtrR system seems to detect other environmental changes and regulates the growth rate, the start of solventogenesis and butanol tolerance. No role in sporulation regulation was described; still, the overexpression of the BtrK/BtrR operon changed the expression of genes involved in sporulation initiation (spo0J, spoIIE, spoIIR) and spore maturation (sigK, spoIVA).
RRNPP quorum-sensing systems positively regulating sporulation were recently identified in C. acetobutylicum (Kotte et al. 2020) and C. saccharoperbutylacetonicum (Feng et al. 2020). These systems named Qss are composed of two proteins: the receptor Qsr, which harbors a helix-turn-helix region (HTH) common to DNA binding domains, and the signal peptide precursor Qsp. In the genome of C. acetobutylicum, eight putative RRNPP systems were detected, but only two seem to intervene in sporulation regulation (Table 2). In C. saccharoperbutylacetonicum, four out of the five identified systems affected sporulation. The deletion of qssR1 and qssR2 increased sporulation efficiency, while sporulation frequency decreased in ΔqssR3 and ΔqssR5 mutants. Moreover, spo0A expression decreased in ΔqssR3 and ΔqssR5 mutants and qssR1/2 affected the expression of spo0E-like genes, putatively involved in the activation of Spo0A. However, no regulation mechanism was described in any of these species. In C. difficile, a similar RRNPP system (RstA) was described. RstA was reported to modulate the expression of sporulation genes through its DNA binding domain (Edwards et al. 2020). A similar regulation mechanism might take place in solventogenic clostridia. Changes in process parameters such as temperature or dilution rate also appear to trigger sporulation (Heluane et al. 2011; Kiyoshi et al. 2017). Sporulation is initiated by a considerable number of triggers (Fig. 7), and several quorum-sensing mechanisms and messengers interact with transcriptional regulators to integrate all the environmental cues and induce a cellular response.
Sporulation in another industrially relevant group: the acetogens
Acetogenic bacteria are C-1 compounds utilizing microorganisms. They can fix CO2, CO, or formate to produce acetate through the Wood-Ljungdahl pathway. Because of their unique metabolism and the development of biobased processes for chemical production, acetogens caught the interest of the biotechnological industry. Anaerobic acetogens are particularly interesting for industrial use since anaerobic conditions reduce flammability concerns linked to the use of CO and H2 and contamination risks (Liew et al. 2016). Fourteen acetogenic species have been identified within the Clostridium genus, and among which six (C. ljungdahlii, C. coskatii, C. drakei, C. carboxidivorans, C. ragsdalei, C. autoethanogenum) are also natural solvent producers. Numerous studies on these clostridial acetogens have been done in the last ten years and one company, Lanzatech, managed to scale up its gas fermenting process based on CO conversion to ethanol and operates a commercial plant (LanzaTech 2019).
Few data are available on the sporulation mechanism in acetogens. In fact, sporulation was rarely observed in the cultures of acetogenic clostridia. Still, some brief descriptions of the morphology of sporulating cells can be found for C. coskatii, C. drakei, C. carboxidivorans, C. ragsdalei, and C. autoethanogenum (Abrini et al. 1994; Huhnke et al. 2008; Liou et al. 2005; Zahn and Saxena 2011). As described for solventogenic clostridia, these species form sub-terminal and terminal spores. During sporulation, swelling of the cells was reported in C. coskatii (Zahn and Saxena 2011) and C. carboxidivorans (Liou et al. 2005), as described for other solventogenic clostridia. In contrast, no swelling of C. autoethanogenum (Abrini et al. 1994) and C. ragsdalei (Huhnke et al. 2008) cells was observed. A significant difference to most solventogenic clostridia is the absence of granulose in sporulating cells of acetogens.
To verify whether the reduced sporulation was due to a difference in the set of sporulation genes, we performed a blastp analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi) based on the minimal set of sporulation genes defined by Galperin et al. (Galperin et al. 2012). Homologs to the 52 core sporulation genes were found in the above-mentioned solvent producing acetogens. Further, we searched for homologs of genes known to be involved in the initiation of sporulation in solventogenic clostridia. In all the genomes analyzed, homologs of the Agr and RRNPP quorum-sensing systems, as well as homologs of orphan kinases, were identified. As for secondary metabolites, no polyketide encoding gene cluster (Letzel et al. 2013) was found; still, all strains harbor ranthipeptide producing enzymes. A recent study on the ranthipeptides produced by C. beijerinckii and C. ljungdahlii highlighted their importance in regulating cellular events (Chen et al. 2020); still, no link with sporulation in C. ljungdahlii could be made. Interestingly, homologs to the enzymes involved in granulose formation were found in C. drakei and C. carboxidivorans, but not in C. ljungdahlii, C. autoethanogenum or C. coskatii.
Therefore, it seems that the low sporulation frequency observed in acetogens is not due to the absence of sporulation genes in the genome. Differences in triggers, transcriptional or post-transcriptional regulations of sporulation proteins during fermentation could explain this low frequency. Using transcriptome analysis of C. ljungdahlii and C. autoethanogenum generated by Philips et al. 2017, Aklujkar et al. 2017, Whitham et al. 2015 and Diender et al. 2019, we looked at the expression profile of sporulation gene homologs under various conditions. When grown under salt stress conditions, C. ljungdahlii did not form spores despite the upregulation of spo0A (Philips et al. 2017). An interesting study on the variation of the transcriptome of C. ljungdahlii under lithotrophic and organotrophic growth conditions (CO2/H2 vs. fructose) reported the upregulation of an agr homolog and nine σE regulated sporulation genes during lithotrophic growth (Aklujkar et al. 2017). Surprisingly, no change in spo0A expression was observed. A similar change was observed when C. ljungdahlii was exposed to oxygen (Whitham et al. 2015). The homologs of abrB, sigE, and three late-stage sporulation genes were upregulated in the O2 challenged cultures. These results show that sporulation genes are expressed during the fermentation in C. ljungdahlii and that their expression varied in response to the environmental changes (changes in carbon source and presence of oxygen) as observed in other clostridia. A similar observation was made when studying the transcriptome analysis done on C. autoethanogenum (Diender et al. 2019). Diender et al. studied variations of the transcriptome of C. autoethanogenum in CO/H2 compared to CO grown-cultures and in mono-culture compared to co-culture with C. kluyveri. In CO/H2 grown-cultures, several stage II and III sporulation genes were upregulated, and a slight increase in the expression of the sporulation-specific sigma factors was detected. In the synthetic co-cultures, an up-regulation of spo0A and spoIIE homologs was observed, while sigF and some sporulation genes involved in cortex formation were down-regulated. According to these results, the regulation of the sporulation in C. ljungdahlii and C. autoethanogenum seems to have common characteristics with the systems described in other solventogenic clostridia. Like in solventogenic clostridia, carbon sources impact sporulation regulation in C. ljungdahlii and C. autoethanogenum. Indeed, their growth in lithotrophic conditions led to an upregulation of sporulation genes compared to organotrophic conditions. Moreover, the sigE and sigF regulons that have been described in C. acetobutylicum and C. beijerinckii seem to be conserved in C. ljungdahlii and C. autoethanogenum. Even so, these results are still not sufficient to claim that the regulatory network governing sporulation is identical in both acetogens and solventogenic clostridia.
Spore formation in fermentation: enemy or ally?
Connections between sporulation and solventogenesis
In solventogenic clostridia, sporulation and solvent production are both stress responses to unfavorable environmental conditions. Solventogenesis is thought to be triggered before sporulation (Patakova et al. 2014; Tracy et al. 2008), although several studies mention a link between sporulation and solvent production (Dürre 2014; Jones and Woods 1986; Long et al. 1984b; Rogers and Palosaari 1987; Schuster et al. 1998). Moreover, both cellular events can be lost simultaneously in a phenomenon named “strain degeneration,” which is observed after repeated batch fermentation or continuous fermentation (Clark et al. 1989; Cornillot et al. 1997; George et al. 1983; Jiao et al. 2016; Kashket and Cao 1995; Kosaka et al. 2007; Lv et al. 2016). Genomic and transcriptomic studies have been done on degenerated strains to unravel the mechanism of degeneration and find ways to prevent it. In C. acetobutylicum, it has been linked to the loss of the pSol megaplasmid, which contains genes encoding enzymes crucial for solvent production, an σX alternative sigma factor involved in sporulation (Behrens et al. 2000; Schuster et al. 2001) and SpoVD, a protein involved in the engulfment in Bacillus (Cornillot et al. 1997; Kashket and Cao 1995). In C. beijerinckii and C. saccharoperbutylacetonicum, which do not harbor the same type of megaplasmid, strain degeneration seems to be caused by mutations. Even though no consensus on the location, number, or type of mutations causing degeneration exists, the cell physiology of these degenerated strains is similar. Changes in medium composition were reported to prevent (Woo et al. 2018) or reestablish the solvent and sporulation ability, but the molecular mechanism is still unknown (Lv et al. 2016).
Nevertheless, common regulators of sporulation and solventogenesis have been identified. The study of cellular signaling pathways proved a close link between sporulation and solventogenesis initiation. The RRNPP quorum sensing systems seem to promote both cellular events. Several regulators enabling the transition from exponential phase to stationary phase are crucial to both cellular events. The disruption of general regulators such as CcpA and σL impaired both phenomena (Table 2). As for the role of Spo0A, it seems to differ depending on the species. In contrast to the phenotype described in C. acetobutylicum and C. beijerinckii (see section on the initiation of sporulation), the expression of spo0A is not required for solvent formation in C. pasteurianum (Schwarz et al. 2017). Proteins involved in Spo0A activation or inhibition, such as AbrB, orphan kinases, and σK, contribute to the regulation of sporulation and solvent production. Studies showed, nonetheless, that decoupling sporulation and solvent formation is possible. Several solvent-producing but asporogenous mutants (Table 2) and recombinant strains have been isolated and engineered (Jones and Keis 1995; Li and He 2016).
Spores in industrial processes
Sporulation in solventogenic clostridia is considered a drawback for several reasons. Once sporulation is initiated, cell growth stops, and the cell’s energy is used to generate metabolically inactive cells (Patakova et al. 2013). These events are undesirable in industrial settings as they negatively impact solvent productivity and cause cell wash-out in a continuous process (Li et al. 2020b; Papoutsakis 2008; Vees et al. 2020). Therefore, various attempts were made, like random mutagenesis, inactivation of early-stage sporulation proteins, and engineering of degenerated strain (Li et al. 2020b) to obtain asporogenous solventogenic strains or to control sporulation. No reports of inducible sporulation were made in solventogenic clostridia but in C. difficile (Dembek et al. 2017). Dembek et al. controlled sporulation by introducing an anhydrotetracycline sensitive promoter upstream from spo0A. Alternatively, modifications of the process can reduce the proportion of sporulating cells. For instance, in continuous culture, the dilution rate is controlled to keep the cells in the growing phase to reduce sporulation (Mutschlechner et al. 2000). Interestingly, continuous cultures can also select for asporogenous solvent-producing strains (Meinecke et al. 1984). Meinecke et al. isolated in continuous culture under phosphate limitation a stable asporogenous strain which produced acetone and butanol as main products. For instance, in continuous culture, the dilution rate is controlled to keep the cells in the growing phase to reduce sporulation (Mutschlechner et al. 2000).
In contrast, bacterial spores can have useful applications, e.g., the pharmaceutical and agroindustry (Ricca and Cutting 2003; Wolken et al. 2003; Zhang et al. 2020). Their high resistance to heat and radiation makes them excellent bio-dosimeters. B. subtilis spores, for instance, are used to test the UV disinfection performance during drinking water purification (Mamane-Gravetz and Linden 2004). B. subtilis spores are also known to be effective biopesticides. The use of spores for enzyme immobilization is actively explored (Ugwuodo and Nwagu 2020). Compared to other immobilization supports, spores are very cheap, and several studies have highlighted the positive effects of spore immobilization on enzyme activity, stability and recovery (Falahati-Pour et al. 2015; Hosseini-Abari et al. 2016; Peng et al. 2020; Song et al. 2019). Clostridium spores are already used in the food and feed industry in Japan and China. Indeed, C. butyricum spores have been used as probiotics for several years. In fact, a large number of acid-producing Clostridium species are found in the gut of healthy individuals. They metabolize nutrients that cannot be degraded by the host. Studies have also proven that C. butyricum can prevent antibiotic-associated diarrhea and prevent C. difficile infections (Cartman 2011; Guo et al. 2020). The use of clostridial spores in the pharmaceutical industry is in the pipeline. Spores of C. novyi and C. sporogenes are being investigated as potential carriers for enzymes involved in chemotherapy (Dürre 2014; Kubiak and Minton 2015; Theys and Lambin 2015).
Some research on the integration of sporulation in the bioprocess for butanol production has been done. Spores can be used for cell immobilization. Low biomass is one of the issues of continuous culture with clostridia, and cell immobilization may prevent wash-out at high dilution rates (Vees et al. 2020). Spore can be immobilized on porous carriers (Dolejš et al. 2014; Krouwel et al. 1983) and microencapsulation (Rathore et al. 2015) to prevent cell wash-out. Microencapsulated spores can even be reused several times, enabling the production of butanol at high yields.
Moreover, sporulation enables consistent conservation of the strain characteristics, protecting the strains from stressors and degeneration. Due to the oxygen sensitivity of vegetative cells and the appearance of degenerative changes after repeated subculturing, spores are usually preferred for strain storage (Jones and Woods 1986). According to documentations on the industrial ABE processes from the 1920s to the 1980s, strains were stored as spores in sterile soil or sand in commercial plants (Jones and Woods 1986; Jones 2001). In South Africa’s ABE commercial plant, multiple cycles of germination/sporulation were performed to increase solvent productivity (Jones 2001). More recently, Vrije and co-workers explored the possibility of integrating a heat treatment during the product recovery by gas stripping in repeated batch cultures (de Vrije et al. 2013). This treatment killed the remaining vegetative cells and triggered the germination of the spores present in the culture, allowing the fermentation to start again after removing the solvents without the need for a new inoculation of the culture. This procedure improved solvent recovery and prolonged strain stability.
Conclusion and perspectives for future studies
Next to solventogenesis, sporulation is a major stationary phase event occurring during ABE fermentation. Changes in carbon sources, media components, and cell density were described as the primary triggers of sporulation. Until recently, studies were mainly conducted with first-generation substrates, but with advanced biofuel production in mind, more studies on the effect of second and third-generation feedstocks on sporulation need to be realized.
Despite being a major part of the cell growth cycle, knowledge on its regulation mechanism in solvent-producing clostridia is scarce. Sporulation in Bacillus and pathogenic clostridia is well described, but noteworthy differences in the regulatory network exist between the two genera (Galperin 2013) and even within the clostridial pathogens (Shen et al. 2019). The model established for those species might not be applicable to solventogenic clostridia. As highlighted in a previous review (Patakova et al. 2013), most research is done on C. acetobutylicum ATCC 824, but with the advances in genome sequencing and gene engineering, more strains have become genetically accessible. Recent studies in other solventogenic strains have revealed differences in the transcriptional regulation of sporulation. Hence, more research in different solventogenic clostridia is required to understand better the complex regulation of sporulation and its interaction with other cellular events.
Furthermore, investigating the reasons behind the low sporulation frequency in cultures of acetogenic clostridia could give hints on strategies to take to prevent sporulation during the ABE fermentation. Knowledge in sporulation might be applied in the design of fermentation processes at different levels, by tailoring the medium composition to reduce/increase sporulation as desired or by integrating clostridial spore in the pretreatment of the substrate by displaying hydrolases on the spore surface or for cell immobilization during a continuous process. A better knowledge of sporulation in solventogenic and acetogenic clostridia would contribute to an improvement of the ABE and gas fermentation processes for the production of fuels and chemicals from renewable resources as a step towards a more sustainable industry.
References
Abd-Alla MH, Zohri ANA, El-Enany AWE, Ali SM (2017) Conversion of food processing wastes to biofuel using clostridia. Anaerobe 48:135–143. https://doi.org/10.1016/j.anaerobe.2017.08.011
Abrini J, Naveau H, Nyns E-J (1994) Clostridium autoethanogenum, sp. nov., an anaerobic bacterium that produces ethanol from carbon monoxide. Arch Microbiol 161(4):345–351. https://doi.org/10.1007/BF00303591
Aframian N, Eldar A (2020) A bacterial tower of babel: quorum-sensing signaling diversity and its evolution. Annu Rev Microbiol 74(1):annurev-micro-012220-063740. https://doi.org/10.1146/annurev-micro-012220-063740
Aklujkar M, Leang C, Shrestha PM, Shrestha M, Lovley DR (2017) Transcriptomic profiles of Clostridium ljungdahlii during lithotrophic growth with syngas or H2 and CO2 compared to organotrophic growth with fructose. Sci Rep 7(1):13135. https://doi.org/10.1038/s41598-017-12712-w
Al-Hinai MA, Jones SW, Papoutsakis ET (2014) σK of Clostridium acetobutylicum is the first known sporulation-specific sigma factor with two developmentally separated roles, one early and one late in sporulation. J Bacteriol 196(2):287–299. https://doi.org/10.1128/JB.01103-13
Al-Hinai MA, Jones SW, Papoutsakis ET (2015) The Clostridium sporulation programs: diversity and preservation of endospore differentiation. Microbiol Mol Biol Rev 79(1):19–37. https://doi.org/10.1128/MMBR.00025-14
Alsaker KV, Spitzer TR, Papoutsakis ET (2004) Transcriptional analysis of spo0A overexpression in Clostridium acetobutylicum and its effect on the cell’s response to butanol stress. J Bacteriol 186(7):1959–1971. https://doi.org/10.1128/JB.186.7.1959-1971.2004
Annous BA, Blaschek HP (1991) Isolation and characterization of Clostridium acetobutylicum mutants with enhanced amylolytic activity. Appl Environ Microbiol 57(9)
Antunes A, Martin-Verstraete I, Dupuy B (2011) CcpA-mediated repression of Clostridium difficile toxin gene expression. Mol Microbiol 79(4):882–899. https://doi.org/10.1111/j.1365-2958.2010.07495.x
Atmadjaja AN, Holby V, Harding AJ, Krabben P, Smith HK, Jenkinson ER (2019) CRISPR-Cas, a highly effective tool for genome editing in Clostridium saccharoperbutylacetonicum N1-4(HMT). FEMS Microbiol Lett 366. https://doi.org/10.1093/femsle/fnz059
Avrova NP, Zubko IK, Alexeyeva EG (1981) The products of fermentation and the activity of pectolytic enzymes in Clostridium felsineum strains differing in the rate of spore formation. Mikrobiologiya 50(3):458–466 https://europepmc.org/article/med/7278720
Awang GM, Ingledew WM, Jones GA (1992) The effect of fermentable carbohydrate on sporulation and butanol production by Clostridium acetobutylicum P262. Appl Microbiol Biotechnol 38(1):12–16. https://doi.org/10.1007/BF00169411
Barák I, Muchová K, Labajová N (2019) Asymmetric cell division during Bacillus subtilis sporulation. Future Microbiol 14(4):353–363. https://doi.org/10.2217/fmb-2018-0338
Basu A, Xin F, Lim TK, Lin Q, Yang KL, He J (2017) Quantitative proteome profiles help reveal efficient xylose utilization mechanisms in solventogenic Clostridium sp. strain BOH3. Biotechnol Bioeng 114(9):1959–1969. https://doi.org/10.1002/bit.26332
Bayat H, Modarressi MH, Rahimpour A (2018) The conspicuity of CRISPR-Cpf1 system as a significant breakthrough in genome editing. Curr Microbiol 75(1):107–115. https://doi.org/10.1007/s00284-017-1406-8
Behrens S, Meyer U, Schankin H, Lonetto MA, Fischer RJ, Bahl H (2000) Identification of two genes encoding putative new members of the ECF subfamily of eubacterial RNA polymerase sigma factors in Clostridium acetobutylicum. J Mol Microbiol Biotechnol 2(3):265–269
Berezina O, Zakharova N, Yarotsky C, Zverlov V (2012) Microbial producers of butanol. Appl Biochem Microbiol 48(7):625–638. https://doi.org/10.1134/S0003683812070022
Bi C, Jones SW, Hess DR, Tracy MBP, Papoutsakis ET (2011) SpoIIe is necessary for asymmetric division, sporulation, and expression of σF, σE, and σG but does not control solvent production in Clostridium acetobutylicum ATCC 824. J Bacteriol 193(19):5130–5137. https://doi.org/10.1128/JB.05474-11
Bischofs IB, Hug JA, Liu AW, Wolf DM, Arkin AP (2009) Complexity in bacterial cell-cell communication: quorum signal integration and subpopulation signaling in the Bacillus subtilis phosphorelay. Proc Natl Acad Sci 106(16):6459–6464. https://doi.org/10.1073/pnas.0810878106
Cañadas IC, Groothuis D, Zygouropoulou M, Rodrigues R, Minton NP (2019) RiboCas: a universal CRISPR-based editing tool for Clostridium. ACS Synth Biol 8(6):1379–1390. https://doi.org/10.1021/acssynbio.9b00075
Cartman ST (2011) Time to consider Clostridium probiotics? Future Microbiol 6(9):969–971. https://doi.org/10.2217/fmb.11.86
Chen Y, Zhou T, Liu D, Li A, Xu S, Liu Q, Li B, Ying H (2013) Production of butanol from glucose and xylose with immobilized cells of Clostridium acetobutylicum. Biotechnol Bioprocess Eng 18(2):234–241. https://doi.org/10.1007/s12257-012-0573-5
Chen Y, Yang Y, Ji X, Zhao R, Li G, Gu Y, Shi A, Jiang W, Zhang Q (2020) The SCIFF-Derived ranthipeptides participate in quorum sensing in solventogenic clostridia. Biotechnol J 15(10):1–8. https://doi.org/10.1002/biot.202000136
Cheng C, Bao T, Yang ST (2019) Engineering Clostridium for improved solvent production: recent progress and perspective. Appl Microbiol Biotechnol 103(14):5549–5566. https://doi.org/10.1007/s00253-019-09916-7
Church BD (1959) Dependence of the heat resistance of bacterial endospores on their dipicolinic acid content. Nature 183:124–125
Clark SW, Bennett GN, Rudolph FB (1989) Isolation and characterization of mutants of Clostridium acetobutylicum ATCC 824 deficient in acetoacetyl-coenzyme a:acetate/butyrate:coenzyme a-transferase (EC 2.8.3.9) and in other solvent pathway enzymes. Appl Environ Microbiol 55(4):970–976. https://doi.org/10.1128/AEM.55.4.970-976.1989
Collins M, Lawson P, Willems A, Cordoba JJ, Fernandez-Garayzabal J, Garcia P, Cai J, Hippe H, Farrow J (1994) The phylogeny of the genus Clostridium: proposal of five new genera and eleven new species combinations. Int J Syst Bacteriol 44(4):812–826. https://doi.org/10.1099/00207713-44-4-812
Cornillot E, Nair RV, Papoutsakis ET, Soucaille P (1997) The genes for butanol and acetone formation in Clostridium acetobutylicum ATCC 824 reside on a large plasmid whose loss leads to degeneration of the strain. J Bacteriol 179(17):5442–5447 http://jb.asm.org/content/179/17/5442.abstract
Cruz-Morales P, Orellana CA, Moutafis G, Moonen G, Rincon G, Nielsen LK, Marcellin E, Bapteste E (2019) Revisiting the evolution and taxonomy of clostridia, a phylogenomic update. Genome Biol Evol 11(7):2035–2044. https://doi.org/10.1093/gbe/evz096
Davidson P, Eutsey R, Redler B, Hiller NL, Laub MT, Durand D (2018) Flexibility and constraint: evolutionary remodeling of the sporulation initiation pathway in Firmicutes. PLoS Genet 14(9):e1007470. https://doi.org/10.1371/journal.pgen.1007470
Dawes IW, Mandelstam J (1970) Sporulation of Bacillus subtilis in continuous culture. J Bacteriol 103(3):529–535. https://doi.org/10.1128/jb.103.3.529-535.1970
de Vrije T, Budde M, van der Wal H, Claassen PAM, López-Contreras AM (2013) “In situ” removal of isopropanol, butanol and ethanol from fermentation broth by gas stripping. Bioresour Technol 137:153–159. https://doi.org/10.1016/j.biortech.2013.03.098
Decker AR, Ramamurthi KS (2017) Cell death pathway that monitors spore morphogenesis. Trends Microbiol 25(8):637–647. https://doi.org/10.1016/j.tim.2017.03.005
Dembek M, Willing SE, Hong HA, Hosseini S, Salgado PS, Cutting SM (2017) Inducible expression of spo0A as a universal tool for studying sporulation in Clostridium difficile. Front Microbiol 8:1793. https://doi.org/10.3389/fmicb.2017.01793
Desai RP, Papoutsakis ET (1999) Antisense RNA strategies for metabolic engineering of Clostridium acetobutylicum. Appl Environ Microbiol 65(3):936–945. https://doi.org/10.1128/aem.65.3.936-945.1999
Diallo M, Simons AD, van der Wal H, Collas F, Houweling-Tan B, Kengen SWM, López-Contreras AM (2018) L-Rhamnose metabolism in Clostridium beijerinckii strain DSM 6423. Appl Environ Microbiol 85(5). https://doi.org/10.1128/AEM.02656-18
Diallo M, Hocq R, Collas F, Chartier G, Wasels F, Wijaya HS, Werten MWT, Wolbert EJH, Kengen SWM, van der Oost J, Ferreira NL, López-Contreras AM (2020a) Adaptation and application of a two-plasmid inducible CRISPR-Cas9 system in Clostridium beijerinckii. Methods 172:51–60. https://doi.org/10.1016/j.ymeth.2019.07.022
Diallo M, Kint N, Monot M, Collas F, Martin-Verstraete I, van der Oost J, Kengen SWM, López-Contreras AM (2020b) Transcriptomic and phenotypic analysis of a spoIIE mutant in Clostridium beijerinckii. Front Microbiol 11(September):1–19. https://doi.org/10.3389/fmicb.2020.556064
Diender M, Parera Olm I, Gelderloos M, Koehorst JJ, Schaap PJ, Stams AJM, Sousa DZ (2019) Metabolic shift induced by synthetic co-cultivation promotes high yield of chain elongated acids from syngas. Sci Rep 9(1):18081. https://doi.org/10.1038/s41598-019-54445-y
Dolejš I, Krasňan V, Stloukal R, Rosenberg M, Rebroš M (2014) Butanol production by immobilised Clostridium acetobutylicum in repeated batch, fed-batch, and continuous modes of fermentation. Bioresour Technol 169:723–730. https://doi.org/10.1016/j.biortech.2014.07.039
Dürre P (2005) Handbook on clostridia. In: Duerre P (ed) Handbook on Clostridia. CRC Press. https://doi.org/10.1201/9780203489819
Dürre P (2014) Physiology and sporulation in Clostridium. In The Bacterial Spore: from Molecules to Systems (Vol. 2, Issue 4, pp. 315–329). American Society of Microbiology. https://doi.org/10.1128/microbiolspec.TBS-0010-2012
Edwards AN, Krall EG, McBride SM (2020) Strain-dependent RstA regulation of Clostridioides difficile toxin production and sporulation. J Bacteriol 202(2). https://doi.org/10.1128/JB.00586-19
Ehsaan M, Kuit W, Zhang Y, Cartman ST, Heap JT, Winzer K, Minton NP (2016) Mutant generation by allelic exchange and genome resequencing of the biobutanol organism Clostridium acetobutylicum ATCC 824. Biotechnol Biofuels 9(1):4. https://doi.org/10.1186/s13068-015-0410-0
Ezeji T, Qureshi N, Blaschek HP (2007) Butanol production from agricultural residues: Impact of degradation products on Clostridium beijerinckii growth and butanol fermentation. Biotechnol Bioeng 97(6):1460–1469. https://doi.org/10.1002/bit.21373
Falahati-Pour SK, Lotfi AS, Ahmadian G, Baghizadeh A (2015) Covalent immobilization of recombinant organophosphorus hydrolase on spores of Bacillus subtilis. J Appl Microbiol 118(4):976–988. https://doi.org/10.1111/jam.12744
Feng J, Zong W, Wang P, Zhang ZT, Gu Y, Dougherty M, Borovok I, Wang Y (2020) RRNPP-Type quorum-sensing systems regulate solvent formation, sporulation and cell motility in Clostridium saccharoperbutylacetonicum. Biotechnol Biofuels 13(1):84. https://doi.org/10.1186/s13068-020-01723-x
Fimlaid KA, Shen A (2015) Diverse mechanisms regulate sporulation sigma factor activity in the Firmicutes. Curr Opin Microbiol 24:88–95. https://doi.org/10.1016/j.mib.2015.01.006
Forsberg CW (1987) Production of 1,3-Propanediol from glycerol by Clostridium acetobutylicum and other Clostridium species. Appl Environ Microbiol 53(4):639–643
Foulquier C, Huang C-N, Nguyen N-P-T, Thiel A, Wilding-Steel T, Soula J, Yoo M, Ehrenreich A, Meynial-Salles I, Liebl W, Soucaille P (2019) An efficient method for markerless mutant generation by allelic exchange in Clostridium acetobutylicum and Clostridium saccharobutylicum using suicide vectors. Biotechnol Biofuels 12(1):31. https://doi.org/10.1186/s13068-019-1364-4
Freedman JC, Li J, Mi E, McClane BA (2019) Identification of an important orphan histidine kinase for the initiation of sporulation and enterotoxin production by Clostridium perfringens Type F strain SM101. MBio 10(1):e02674–e02618. https://doi.org/10.1128/mbio.02674-18
Fujita M, González-Pastor JE, Losick R (2005) High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis. J Bacteriol 187(4):1357–1368. https://doi.org/10.1128/JB.187.4.1357-1368.2005
Gallazzi A, Branska B, Marinelli F, Patakova P (2015) Continuous production of n-butanol by Clostridium pasteurianum DSM 525 using suspended and surface-immobilized cells. J Biotechnol 216:29–35. https://doi.org/10.1016/j.jbiotec.2015.10.008
Galperin MY (2013) Genome diversity of spore-forming Firmicutes. Microbiol Spectr 1(2). https://doi.org/10.1128/microbiolspectrum.TBS-0015-2012
Galperin MY, Mekhedov SL, Puigbo P, Smirnov S, Wolf YI, Rigden DJ (2012) Genomic determinants of sporulation in Bacilli and Clostridia : towards the minimal set of sporulation-specific genes. Environ Microbiol 14(11):2870–2890. https://doi.org/10.1111/j.1462-2920.2012.02841.x
George HA, Johnson JL, Moore WE, Holdeman LV, Chen JS (1983) Acetone, isopropanol, and butanol production by Clostridium beijerinckii (syn. Clostridium butylicum) and Clostridium aurantibutyricum. Appl Environ Microbiol 45(3):1160–1163 http://aem.asm.org/content/45/3/1160.abstract
Gong F, Bao G, Zhao C, Zhang Y, Li Y, Dong H (2016) Fermentation and genomic analysis of acetone-uncoupled butanol production by Clostridium tetanomorphum. Appl Microbiol Biotechnol 100(3):1523–1529. https://doi.org/10.1007/s00253-015-7121-0
Green EM (2011) Fermentative production of butanol—the industrial perspective. Curr Opin Biotechnol 22(3):337–343. https://doi.org/10.1016/j.copbio.2011.02.004
Green EM, Boynton ZL, Harris LM, Rudolph FB, Papoutsakis ET, Bennett GN (1996) Genetic manipulation of acid formation pathways by gene inactivation in Clostridium acetobutylicum ATCC 824. Microbiology 142(8):2079–2086. https://doi.org/10.1099/13500872-142-8-2079
Grimmler C, Janssen H, Krauße D, Fischer R-JJ, Bahl H, Dürre P, Liebl W, Ehrenreich A (2011) Genome-wide gene expression analysis of the switch between acidogenesis and solventogenesis in continuous cultures of Clostridium acetobutylicum. J Mol Microbiol Biotechnol 20(1):1–15. https://doi.org/10.1159/000320973
Grossman AD, Losick R (1988) Extracellular control of spore formation in Bacillus subtilis. Proc Natl Acad Sci U S A 85(12):4369–4373. https://doi.org/10.1073/pnas.85.12.4369
Guo P, Zhang K, Ma X, He P (2020) Clostridium species as probiotics: potentials and challenges. J Anim Sci Biotechnol 11(1):24. https://doi.org/10.1186/s40104-019-0402-1
Gupta RS, Gao B (2009) Phylogenomic analyses of clostridia and identification of novel protein signatures that are specific to the genus Clostridium sensu stricto (cluster I). Int J Syst Evol Microbiol 59(2):285–294. https://doi.org/10.1099/ijs.0.001792-0
Harris LM, Desai RP, Welker NE, Papoutsakis ET (2000) Characterization of recombinant strains of the Clostridium acetobutylicum butyrate kinase inactivation mutant: need for new phenomenological models for solventogenesis and butanol inhibition? Biotechnol Bioeng 67(1):1–11. https://doi.org/10.1002/(SICI)1097-0290(20000105)
Harris LM, Welker NE, Papoutsakis ET (2002) Northern, morphological, and fermentation analysis of spo0A inactivation and overexpression in Clostridium acetobutylicum ATCC 824. J Bacteriol 184(13):3586–3597. https://doi.org/10.1128/JB.184.13.3586-3597.2002
Hecker M, Völker U (2001) General stress response of Bacillus subtilis and other bacteria. In Advances in Microbial Physiology (Vol. 44, pp. 35–91). Academic Press. https://doi.org/10.1016/S0065-2911(01)44011-2
Heluane H, Dagher MRE, Bruno-Bárcena JM, Evans MR, Dagher SF, Bruno-Bárcena JM (2011) Meta-analysis and functional validation of nutritional requirements of solventogenic clostridia growing under butanol stress conditions and coutilization of D-glucose and D-xylose. Appl Environ Microbiol 77(13):4473–4485. https://doi.org/10.1128/AEM.00116-11
Herman NA, Kim SJ, Li JS, Cai W, Koshino H, Zhang W (2017) The industrial anaerobe Clostridium acetobutylicum uses polyketides to regulate cellular differentiation. Nat Commun 8(1):1514. https://doi.org/10.1038/s41467-017-01809-5
Hocq R, Bouilloux-Lafont M, Lopes Ferreira N, Wasels F (2019) σ54 (σL) plays a central role in carbon metabolism in the industrially relevant Clostridium beijerinckii. Sci Rep 9(1):7228. https://doi.org/10.1038/s41598-019-43822-2
Hosseini-Abari A, Kim B-G, Lee S-H, Emtiazi G, Kim W, Kim J-H (2016) Surface display of bacterial tyrosinase on spores of Bacillus subtilis using CotE as an anchor protein. J Basic Microbiol 56(12):1331–1337. https://doi.org/10.1002/jobm.201600203
Hu S, Zheng H, Gu Y, Zhao J, Zhang W, Yang Y, Wang S, Zhao G, Yang S, Jiang W (2011) Comparative genomic and transcriptomic analysis revealed genetic characteristics related to solvent formation and xylose utilization in Clostridium acetobutylicum EA 2018. BMC Genomics 12(1):93. https://doi.org/10.1186/1471-2164-12-93
Huang SS, Chen D, Pelczar PL, Vepachedu VR, Setlow P, Li YQ (2007) Levels of Ca2+-dipicolinic acid in individual Bacillus spores determined using microfluidic Raman tweezers. J Bacteriol 189(13):4681–4687. https://doi.org/10.1128/JB.00282-07
Huang H, Chai C, Yang S, Jiang W, Gu Y (2019) Phage serine integrase-mediated genome engineering for efficient expression of chemical biosynthetic pathway in gas-fermenting Clostridium ljungdahlii. Metab Eng 52(September 2018):293–302. https://doi.org/10.1016/j.ymben.2019.01.005
Huhnke RL, Lewis RS, Tanner RS (2008) Isolation and characterization of novel clostridial species (Patent No. US7704723 B2). In Patent (US7704723 B2). U.S. Patent and Trademark Office. https://shareok.org/handle/11244/15409
Jabbari S, Steiner E, Heap JT, Winzer K, Minton NP, King JR (2013) The putative influence of the agr operon upon survival mechanisms used by Clostridium acetobutylicum. Math Biosci 243(2):223–239. https://doi.org/10.1016/j.mbs.2013.03.005
Jain MK, Beacom D, Datta R, Lansing E (1994) Mutant strain of C. acetobutylicum and process for making butanol (Patent No. US5192673A). In Biotechnology Advances (No. US5192673A). U.S. Patent and Trademark Office. https://doi.org/10.1016/0734-9750(94)90895-8
Jamroskovic J, Chromikova Z, List C, Bartova B, Barák I, Bernier-Latmani R (2016) Variability in DPA and calcium content in the spores of Clostridium species. Front Microbiol 7(NOV):1791. https://doi.org/10.3389/fmicb.2016.01791
Jiao S, Zhang Y, Wan C, Lv J, Du R, Zhang R, Han B (2016) Transcriptional analysis of degenerate strain Clostridium beijerinckii DG-8052 reveals a pleiotropic response to CaCO3- associated recovery of solvent production. Sci Rep 6:38818. https://doi.org/10.1038/srep38818
Johnson EA (2019) Clostridia. Encycl Microbiol 690–695. https://doi.org/10.1016/B978-0-12-801238-3.98821-4
Jones DT (2001) Applied acetone-butanol fermentation. In H Bahl & P Dürre (eds) Clostridia (Vol. 2, Issue 1, pp. 125–168). Wiley-VCH Verlag GmbH. https://doi.org/10.1002/3527600108.ch5
Jones DT, Keis S (1995) Origins and relationships of industrial solvent-producing clostridial strains. FEMS Microbiol. Rev. 17(3):223–232. https://doi.org/10.1111/j.1574-6976.1995.tb00206.x
Jones DT, Woods DR (1986) Acetone-butanol fermentation revisited. Microbiol Rev 50(4):484–524 3540574
Jones SW, Paredes CJ, Tracy B, Cheng N, Sillers R, Senger RS, Papoutsakis ET (2008) The transcriptional program underlying the physiology of clostridial sporulation. Genome Biol 9(7):R114. https://doi.org/10.1186/gb-2008-9-7-r114
Jones SW, Tracy BP, Gaida SM, Papoutsakis ET (2011) Inactivation of σF in Clostridium acetobutylicum ATCC 824 blocks sporulation prior to asymmetric division and abolishes σE and σG protein expression but does not block solvent formation. J Bacteriol 193(10):2429–2440. https://doi.org/10.1128/JB.00088-11
Joseph RC, Kim NM, Sandoval NR (2018) Recent developments of the synthetic biology toolkit for Clostridium. Front Microbiol 9:154. https://doi.org/10.3389/fmicb.2018.00154
Kashket ER, Cao Z-Y (1995) Clostridial strain degeneration. FEMS Microbiol. Rev. 17(3):307–315. https://doi.org/10.1016/0168-6445(95)00012-2
Kay D, Warren SC (1968) Sporulation in Bacillus subtilis. Morphological changes. Biochem J 109(5):819–824. https://doi.org/10.1042/bj1090819
Keis S, Bennett CF, Ward VK, Jones DT (1995) Taxonomy and phylogeny of industrial solvent-producing clostridia. Int J Syst Bacteriol 45(4):693–705. https://doi.org/10.1099/00207713-45-4-693
Keis S, Shaheen R, Jones DT (2001) Emended descriptions of Clostridium acetobutylicum and Clostridium beijerinckii, and descriptions of Clostridium saccharoperbutylacetonicum sp nov and Clostridium saccharobutylicum sp nov. Int J Syst Evol Microbiol 51(2001):2095–2103. https://doi.org/10.1099/00207713-51-6-2095
Kihm DJ, Hutton MT, Hanlin JH, Johnson EA (1988) Zinc stimulates sporulation in Clostridium botulinum 113B. Curr Microbiol 17(4):193–198. https://doi.org/10.1007/BF01589451
Kiyoshi K, Kawashima S, Nobuki K, Kadokura T, Nakazato A, Suzuki KI, Nakayama S (2017) Adenine addition restores cell viability and butanol production in Clostridium saccharoperbutylacetonicum N1-4 (ATCC 13564) cultivated at 37°C. Appl Environ Microbiol 83(7):e02960–e02916. https://doi.org/10.1128/AEM.02960-16
Knaysi G (1948) The endospore of bacteria. Bacteriol Rev 12(1):19–77. https://doi.org/10.1128/mmbr.12.1.19-77.1948
Kolek J, Diallo M, Vasylkivska M, Branska B, Sedlar K, López-Contreras AM, Patakova P (2017) Comparison of expression of key sporulation, solventogenic and acetogenic genes in C. beijerinckii NRRL B-598 and its mutant strain overexpressing spo0A. Appl Microbiol Biotechnol 101(22):1–13. https://doi.org/10.1007/s00253-017-8555-3
Kosaka T, Nakayama S, Nakaya K, Yoshino S, Furukawa K (2007) Characterization of the sol operon in butanol-hyperproducing Clostridium saccharoperbutylacetonicum strain N1-4 and its degeneration mechanism. Biosci Biotechnol Biochem 71(1):58–68. https://doi.org/10.1271/bbb.60370
Kosono S, Tamura M, Suzuki S, Kawamura Y, Yoshida A, Nishiyama M, Yoshida M (2015) Changes in the acetylome and succinylome of Bacillus subtilis in response to carbon source. PLoS One 10(6):e0131169. https://doi.org/10.1371/journal.pone.0131169
Kotte A-K, Severn O, Bean Z, Schwarz K, Minton NP, Winzer K (2020) RRNPP-type quorum sensing affects solvent formation and sporulation in Clostridium acetobutylicum. Microbiology 166(6):579–592. https://doi.org/10.1099/mic.0.000916
Król E, De Sousa BA, Kopacz M, Scheffers DJ (2017) Metal-dependent SpoIIE oligomerization stabilizes FtsZ during asymmetric division in Bacillus subtilis. PLoS One 12(3):e0174713. https://doi.org/10.1371/journal.pone.0174713
Krouwel PG, Groot WJ, Kossen NWF, van der Laan WFM (1983) Continuous isopropanol-butanol-ethanol fermentation by immobilized Clostridium beijerinckii cells in a packed bed fermenter. Enzym Microb Technol 5:46–54. https://doi.org/10.1016/0141-0229(83)90064-9
Kubiak AM, Minton NP (2015) The potential of clostridial spores as therapeutic delivery vehicles in tumour therapy. Res Microbiol 166(4):244–254. https://doi.org/10.1016/j.resmic.2014.12.006
Kuit W (2013) Metabolic engineering of acid formation in Clostridium acetobutylicum Wouter Kuit [Wageningen University]. http://library.wur.nl/WebQuery/wurpubs/fulltext/241021
Kuit W, Minton NP, López-Contreras AM, Eggink G (2012) Disruption of the acetate kinase (ack) gene of Clostridium acetobutylicum results in delayed acetate production. Appl Microbiol Biotechnol 94(3):729–741. https://doi.org/10.1007/s00253-011-3848-4
LanzaTech (2019) LanzaTech | Capturing carbon. Fueling growth. LanzaTech. https://www.lanzatech.com/
Lawson PA, Rainey FA (2016) Proposal to restrict the genus Clostridium Prazmowski to Clostridium butyricum and related species. Int J Syst Evol Microbiol 66(2):1009–1016. https://doi.org/10.1099/ijsem.0.000824
Lee KY, Juang TC, Lee KC (1978) Effect of metal ions on growth and sporulation of Clostridium perfringens in a synthetic medium. Chin J Microbiol Immunol 11(2):50–61
Lee AJ, Byun BY, Kang D-H, Tang J, Kim Y-W, Hwang H-J, Mah J-H (2011) The ability of zinc to inhibit the sporulation and viability of Clostridium sporogenes and growth of other bacteria. Int J Food Sci Technol 46(7):1494–1501. https://doi.org/10.1111/j.1365-2621.2011.02644.x
Lee S, Lee JH, Mitchell RJ (2015) Analysis of Clostridium beijerinckii NCIMB 8052’s transcriptional response to ferulic acid and its application to enhance the strain tolerance. Biotechnol Biofuels 8(1):68. https://doi.org/10.1186/s13068-015-0252-9
Leggett MJ, Mcdonnell G, Denyer SP, Setlow P, Maillard JY (2012) Bacterial spore structures and their protective role in biocide resistance. J Appl Microbiol 113(3):485–498. https://doi.org/10.1111/j.1365-2672.2012.05336.x
Lenz CA, Vogel RF (2014) Effect of sporulation medium and its divalent cation content ontheheat and high pressure resistance of Clostridium botulinum typeE spores. Food Microbiol 44:156–167. https://doi.org/10.1016/j.fm.2014.05.010
Letzel A-C, Pidot SJ, Hertweck C (2013) A genomic approach to the cryptic secondary metabolome of the anaerobic world. Nat Prod Rep 30(3):392–428. https://doi.org/10.1039/C2NP20103H
Li T, He J (2016) Simultaneous saccharification and fermentation of hemicellulose to butanol by a non-sporulating Clostridium species. Bioresour Technol 219:430–438. https://doi.org/10.1016/j.biortech.2016.07.138
Li J, Chen J, Vidal JE, McClane BA (2011) The Agr-like quorum-sensing system regulates sporulation and production of enterotoxin and beta2 toxin by Clostridium perfringens type a non-food-borne human gastrointestinal disease strain F5603. Infect Immun 79(6):2451–2459. https://doi.org/10.1128/IAI.00169-11
Li Q, Chen J, Minton NP, Zhang Y, Wen Z, Liu J, Yang H, Zeng Z, Ren X, Yang J, Gu Y, Jiang W, Jiang Y, Yang S (2016) CRISPR-based genome editing and expression control systems in Clostridium acetobutylicum and Clostridium beijerinckii. Biotechnol J 11(7):961–972. https://doi.org/10.1002/biot.201600053
Li Q, Seys FM, Minton NP, Yang J, Jiang Y, Jiang W, Yang S (2019) CRISPR-Cas9 D10A nickase-assisted base editing in solvent producer Clostridium beijerinckii. Biotechnol Bioeng 116:1475–1483. https://doi.org/10.1002/bit.26949
Li JS, Barber CC, Herman NA, Cai W, Zafrir E, Du Y, Zhu X, Skyrud W, Zhang W (2020a) Investigation of secondary metabolism in the industrial butanol hyper-producer Clostridium saccharoperbutylacetonicum N1-4. J Ind Microbiol Biotechnol 47(3):319–328. https://doi.org/10.1007/s10295-020-02266-8
Li S, Huang L, Ke C, Pang Z, Liu L (2020b) Pathway dissection, regulation, engineering and application: lessons learned from biobutanol production by solventogenic clostridia. Biotechnol Biofuels 13(1):39. https://doi.org/10.1186/s13068-020-01674-3
Liew F, Martin ME, Tappel RC, Heijstra BD, Mihalcea C, Köpke M (2016) Gas Fermentation—a flexible platform for commercial scale production of low-carbon-fuels and chemicals from waste and renewable feedstocks. Front Microbiol 7(694). https://doi.org/10.3389/fmicb.2016.00694
Liou JS-C, Balkwill DL, Drake GR, Tanner RS (2005) Clostridium carboxidivorans sp. nov., a solvent-producing Clostridium isolated from an agricultural settling lagoon, and reclassification of the acetogen Clostridium scatologenes strain SL1 as Clostridium drakei sp. nov. Int J Syst Evol Microbiol 55(5):2085–2091. https://doi.org/10.1099/ijs.0.63482-0
List C, Hosseini Z, Lederballe Meibom K, Hatzimanikatis V, Bernier-Latmani R (2019) Impact of iron reduction on the metabolism of Clostridium acetobutylicum. Environ Microbiol 21(10):3548–3563. https://doi.org/10.1111/1462-2920.14640
Liu Z, Qiao K, Tian L, Zhang Q, Liu Z-Y, Li F-L (2015) Spontaneous large-scale autolysis in Clostridium acetobutylicum contributes to generation of more spores. Front Microbiol 6:950. https://doi.org/10.3389/fmicb.2015.00950
Liu H, Zhang J, Yuan J, Jiang X, Jiang L, Li Z, Yin Z, Du Y, Zhao G, Liu B, Huang D (2020) Gene coexpression network analysis reveals a novel metabolic mechanism of Clostridium acetobutylicum responding to phenolic inhibitors from lignocellulosic hydrolysates. Biotechnol Biofuels 13(1):163. https://doi.org/10.1186/s13068-020-01802-z
Long S, Jones DT, Woods DR (1983) Sporulation of Clostridium acetobutylicum P262 in a Defined Medium. Appl Environ Microbiol 45(4):1389–1393. https://doi.org/10.1128/AEM.45.4.1389-1393.1983
Long S, Jones D, Woods D (1984a) Initiation of solvent production, clostridial stage and endospore formation in Clostridium acetobutylicum P262. Appl Microbiol Biotechnol 20(4). https://doi.org/10.1007/BF00250635
Long S, Jones DT, Woods DR (1984b) The relationship between sporulation and solvent production in Clostridium acetobutylicum P262. Biotechnol Lett 6(8):529–534. https://doi.org/10.1007/BF00139997
Lund BM, Brocklehurst TF, Wyatt GM (1981) Characterization of strains of Clostridium puniceum sp.nov., a pink-pigmented, pectolytic bacterium. J Gen Microbiol 122(1):17–26. https://doi.org/10.1099/00221287-122-1-17
Lv J, Jiao S, Du R, Zhang R, Zhang Y, Han B (2016) Proteomic analysis to elucidate degeneration of Clostridium beijerinckii NCIMB 8052 and role of Ca2+ in strain recovery from degeneration. J Ind Microbiol Biotechnol 43(6):741–750. https://doi.org/10.1007/s10295-016-1754-6
Macek B, Forchhammer K, Hardouin J, Weber-Ban E, Grangeasse C, Mijakovic I (2019) Protein post-translational modifications in bacteria. Nat Rev Microbiol 17(11):651–664. https://doi.org/10.1038/s41579-019-0243-0
Mah JH, Kang DH, Tang J (2008) Effects of minerals on sporulation and heat resistance of Clostridium sporogenes. Int J Food Microbiol 128(2):385–389. https://doi.org/10.1016/j.ijfoodmicro.2008.10.002
Malaviya A, Jang YS, Lee SY (2012) Continuous butanol production with reduced byproducts formation from glycerol by a hyper producing mutant of Clostridium pasteurianum. Appl Microbiol Biotechnol 93(4):1485–1494. https://doi.org/10.1007/s00253-011-3629-0
Mamane-Gravetz H, Linden KG (2004) UV disinfection of indigenous aerobic spores: Implications for UV reactor validation in unfiltered waters. Water Res 38(12):2898–2906. https://doi.org/10.1016/j.watres.2004.03.035
Martínez-Lumbreras S, Alfano C, Evans NJ, Collins KM, Flanagan KA, Atkinson RA, Krysztofinska EM, Vydyanath A, Jackter J, Fixon-Owoo S, Camp AH, Isaacson RL (2018) Structural and functional insights into Bacillus subtilis sigma factor inhibitor, CsfB. Structure 26(4):640–648.e5. https://doi.org/10.1016/j.str.2018.02.007
Mate de Gerando H, Wasels F, Bisson A, Clement B, Bidard F, & Jourdier E. (2018). Genome sequence of the natural isopropanol producer Clostridium beijerinckii DSM 6423. https://www.ebi.ac.uk/ena/browser/view/PRJEB11626
Máté de Gérando H, Wasels F, Bisson A, Clement B, Bidard F, Jourdier E, López-Contreras AM, Lopes Ferreira N (2018) Genome and transcriptome of the natural isopropanol producer Clostridium beijerinckii DSM 6423. BMC Genomics 19(1):242. https://doi.org/10.1186/s12864-018-4636-7
Meinecke B, Bahl H, Gottschalk G (1984) Selection of an asporogenous strain of Clostridium acetobutylicum in continuous culture under phosphate limitation. Appl Environ Microbiol 48(5):1064–1065. https://doi.org/10.1128/aem.48.5.1064-1065.1984
Mukherjee M, Sarkar P, Goswami G, Das D (2019) Regulation of butanol biosynthesis in Clostridium acetobutylicum ATCC 824 under the influence of zinc supplementation and magnesium starvation. Enzym Microb Technol 129:109352. https://doi.org/10.1016/j.enzmictec.2019.05.009
Mutschlechner O, Swoboda H, Gapes JR (2000) Continuous two-stage ABE-fermentation using Clostridium beijerinckii NRRL B592 operating with a growth rate in the first stage vessel close to its maximal value. J Mol Microbiol Biotechnol 2(1):101–105
Narula J, Devi SN, Fujita M, Igoshin OA (2012) Ultrasensitivity of the Bacillus subtilis sporulation decision. Proc Natl Acad Sci 109(50):E3513–E3522. https://doi.org/10.1073/pnas.1213974109
Nimbalkar PR, Khedkar MA, Parulekar RS, Chandgude VK, Sonawane KD, Chavan PV, Bankar SB (2018) Role of trace elements as cofactor: an efficient strategy toward enhanced biobutanol production. ACS Sustain Chem Eng 6(7):9304–9313. https://doi.org/10.1021/acssuschemeng.8b01611
Nimbalkar PR, Khedkar MA, Chavan PV, Bankar SB (2019) Enhanced biobutanol production in folic acid-induced medium by using Clostridium acetobutylicum NRRL B-527. ACS Omega 4(7):12978–12982. https://doi.org/10.1021/acsomega.9b00583
Paidhungat M, Setlow B, Driks A, Setlow P (2000) Characterization of spores of Bacillus subtilis which lack dipicolinic acid. J Bacteriol 182(19):5505–5512. https://doi.org/10.1128/JB.182.19.5505-5512.2000
Papoutsakis ET (2008) Engineering solventogenic clostridia. Curr Opin Biotechnol 19(5):420–429. https://doi.org/10.1016/j.copbio.2008.08.003
Paredes CJ, Alsaker KV, Papoutsakis ET (2005) A comparative genomic view of clostridial sporulation and physiology. Nat Rev Microbiol 3(12):969–978. https://doi.org/10.1038/nrmicro1288
Paredes-Sabja D, Setlow B, Setlow P, Sarker MR (2008) Characterization of Clostridium perfringens spores that lack SpoVA proteins and dipicolinic acid. J Bacteriol 190(13):4648–4659. https://doi.org/10.1128/JB.00325-08
Paredes-Sabja D, Shen A, Sorg JA (2014) Clostridium difficile spore biology: sporulation, germination, and spore structural proteins. Trends Microbiol 22(7):406–416. https://doi.org/10.1016/j.tim.2014.04.003
Patakova P, Linhova M, Rychtera M, Paulova L, Melzoch K (2013) Novel and neglected issues of acetone-butanol-ethanol (ABE) fermentation by clostridia: Clostridium metabolic diversity, tools for process mapping and continuous fermentation systems. Biotechnol Adv 31(1):58–67. https://doi.org/10.1016/j.biotechadv.2012.01.010
Patakova P, Linhova M, Vykydalova P, Branska B, Rychtera M, Melzoch K (2014) Use of fluorescent staining and flow cytometry for monitoring physiological changes in solventogenic clostridia. Anaerobe 29:113–117. https://doi.org/10.1016/j.anaerobe.2013.10.006
Patakova P, Branska B, Sedlar K, Vasylkivska M, Jureckova K, Kolek J, Koscova P, Provaznik I (2019) Acidogenesis, solventogenesis, metabolic stress response and life cycle changes in Clostridium beijerinckii NRRL B-598 at the transcriptomic level. Sci Rep 9(1):1371. https://doi.org/10.1038/s41598-018-37679-0
Paul C, Filippidou S, Jamil I, Kooli W, House GL, Estoppey A, Hayoz M, Junier T, Palmieri F, Wunderlin T, Lehmann A, Bindschedler S, Vennemann T, Chain PSG, Junier P (2019) Bacterial spores, from ecology to biotechnology. Adv Appl Microbiol 106:79–111. https://doi.org/10.1016/bs.aambs.2018.10.002
Peng F, Zheng B, Zhang Y, Faheem A, Chai Y, Jiang T, Chen X, Hu Y (2020) Biocatalytic oxidation of aromatic compounds by spore-based system. ACS Sustain Chem Eng 8(37):14159–14165. https://doi.org/10.1021/acssuschemeng.0c04884
Philips J, Rabaey K, Lovley DR, Vargas M (2017) Biofilm formation by Clostridium ljungdahlii is induced by sodium chloride stress: experimental evaluation and transcriptome analysis. PLoS One 12(1):e0170406. https://doi.org/10.1371/journal.pone.0170406
Piggot PJ, Hilbert DW (2004) Sporulation of Bacillus subtilis. Curr Opin Microbiol 7(6):579–586. https://doi.org/10.1016/j.mib.2004.10.001
Poehlein A, Solano JDM, Flitsch SK, Krabben P, Winzer K, Reid SJ, Jones DT, Green E, Minton NP, Daniel R, Dürre P (2017) Microbial solvent formation revisited by comparative genome analysis. Biotechnol Biofuels 10(1):58. https://doi.org/10.1186/s13068-017-0742-z
Prazmowski A (1880) Untersuchungen über die Entwickelungsgeschichte und Fermentwirkung einiger Bacterien-Arten. Voigt.
Qureshi N, Blaschek HP (2001) Recent advances in ABE fermentation: hyper-butanol producing Clostridium beijerinckii BA101. J Ind Microbiol Biotechnol 27(5):287–291. https://doi.org/10.1038/sj.jim.7000114
Raedts J, Siemerink MAJ, Levisson M, van der Oost J, Kengen SWM (2014) Molecular characterization of an NADPH-dependent acetoin reductase/2,3-butanediol dehydrogenase from Clostridium beijerinckii NCIMB 8052. Appl Environ Microbiol 80(6):2011–2020. https://doi.org/10.1128/AEM.04007-13
Rathore S, Wan Sia Heng P, Chan LW (2015) Microencapsulation of Clostridium acetobutylicum ATCC 824 spores in gellan gum microspheres for the production of biobutanol. J Microencapsul 32(3):290–299. https://doi.org/10.3109/02652048.2015.1017617
Ravagnani A, Jennert KCB, Steiner E, Grünberg R, Jefferies JR, Wilkinson SR, Young DI, Tidswell EC, Brown DP, Youngman P, Gareth Morris J, Young M (2000) Spo0A directly controls the switch from acid to solvent production in solvent-forming clostridia. Mol Microbiol 37(5):1172–1185. https://doi.org/10.1046/j.1365-2958.2000.02071.x
Reeve BWP, Reid SJ (2016) Glutamate and histidine improve both solvent yields and the acid tolerance response of Clostridium beijerinckii NCP 260. J Appl Microbiol 120(5):1271–1281. https://doi.org/10.1111/jam.13067
Ren C, Gu Y, Wu Y, Zhang W, Yang C, Yang S, Jiang W (2012) Pleiotropic functions of catabolite control protein CcpA in Butanol-producing Clostridium acetobutylicum. BMC Genomics 13(1):349. https://doi.org/10.1186/1471-2164-13-349
Ricca E, Cutting SM (2003) Emerging applications of bacterial spores in nanobiotechnology. J Nanobiotechnol 1(1):6. https://doi.org/10.1186/1477-3155-1-6
Rogers P, Palosaari N (1987) Clostridium acetobutylicum mutants that produce butyraldehyde and altered quantities of solvents. Appl Environ Microbiol 53(12):2761–2766 http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=204194&tool=pmcentrez&rendertype=abstract
Sandoval NR, Venkataramanan KP, Groth TS, Papoutsakis ET (2015) Whole-genome sequence of an evolved Clostridium pasteurianum strain reveals Spo0A deficiency responsible for increased butanol production and superior growth. Biotechnol Biofuels 8(1):227. https://doi.org/10.1186/s13068-015-0408-7
Sandoval-Espinola WJ, Makwana ST, Chinn MS, Thon MR, Azcárate-Peril MA, Bruno-Bárcena JM (2013) Comparative phenotypic analysis and genome sequence of Clostridium beijerinckii SA-1, an offspring of NCIMB 8052. Microbiology 159(Pt_12):2558–2570. https://doi.org/10.1099/mic.0.069534-0
Sauer M (2016) Industrial production of acetone and butanol by fermentation-100 years later. FEMS Microbiol Lett 363(13). https://doi.org/10.1093/femsle/fnw134
Sauer U, Santangelo JD, Treuner A, Buchholz M, Dürre P (1995) Sigma factor and sporulation genes in Clostridium. FEMS Microbiol Rev 17(3):331–340. https://doi.org/10.1111/j.1574-6976.1995.tb00216.x
Schmeisser F, Brannigan JA, Lewis RJ, Wilkinson AJ, Youngman P, Barák I (2000) A new mutation in spo0A with intragenic suppressors in the effector domain. FEMS Microbiol Lett 185(2):123–128. https://doi.org/10.1111/j.1574-6968.2000.tb09049.x
Schuster KC, Van Den Heuvel R, Gutierrez NA, Maddox IS (1998) Development of markers for product formation and cell cycle in batch cultivation of Clostridium acetobutylicum ATCC 824. Appl Microbiol Biotechnol 49(6):669–676. https://doi.org/10.1007/s002530051230
Schuster KC, Goodacre R, Gapes J, Young M (2001) Degeneration of solventogenic Clostridium strains monitored by Fourier transform infrared spectroscopy of bacterial cells. J Ind Microbiol Biotechnol 27:314–321. https://springerlink.fh-diploma.de/content/pdf/10.1038/sj.jim.7000146.pdf
Schwarz KM, Kuit W, Grimmler C, Ehrenreich A, Kengen SWM (2012) A transcriptional study of acidogenic chemostat cells of Clostridium acetobutylicum — cellular behavior in adaptation to n-butanol. J Biotechnol 161(3):366–377. https://doi.org/10.1016/j.jbiotec.2012.03.018
Schwarz KM, Grosse-Honebrink A, Derecka K, Rotta C, Zhang Y, Minton NP (2017) Towards improved butanol production through targeted genetic modification of Clostridium pasteurianum. Metab Eng 40:124–137. https://doi.org/10.1016/j.ymben.2017.01.009
Scotcher MC, Bennett GN (2005) SpoIIE regulates sporulation but does not directly affect solventogenesis in Clostridium acetobutylicum ATCC 824. J Bacteriol 187(6):1930–1936. https://doi.org/10.1128/JB.187.6.1930-1936.2005
Scotcher MC, Rudolph FB, Bennett GN (2005) Expression of abrB310 and sinR, and effects of decreased abrB310 expression on the transition from acidogenesis to solventogenesis, in Clostridium acetobutylicum ATCC 824. Appl Environ Microbiol 71(4):1987–1995. https://doi.org/10.1128/AEM.71.4.1987-1995.2005
Sedlar K, Koscova P, Vasylkivska M, Branska B, Kolek J, Kupkova K, Patakova P, Provaznik I (2018) Transcription profiling of butanol producer Clostridium beijerinckii NRRL B-598 using RNA-Seq. BMC Genomics 19(1):415. https://doi.org/10.1186/s12864-018-4805-8
Sedlar K, Kolek J, Gruber M, Jureckova K, Branska B, Csaba G, Vasylkivska M, Zimmer R, Patakova P, Provaznik I (2019) A transcriptional response of Clostridium beijerinckii NRRL B-598 to a butanol shock. Biotechnol Biofuels 12(1):243. https://doi.org/10.1186/s13068-019-1584-7
Sedlar K, Vasylkivska M, Musilova J, Branska B, Provaznik I, Patakova P (2021) Phenotypic and genomic analysis of isopropanol and 1,3-propanediol producer Clostridium diolis DSM 15410. Genomics 113(1):1109–1119. https://doi.org/10.1016/j.ygeno.2020.11.007
Seo SO, Janssen H, Magis A, Wang Y, Lu T, Price ND, Jin YS, Blaschek HP (2017a) Genomic, transcriptional, and phenotypic analysis of the glucose derepressed Clostridium beijerinckii mutant exhibiting acid crash phenotype. Biotechnol J 12(11):1700182. https://doi.org/10.1002/biot.201700182
Seo SO, Wang Y, Lu T, Jin YS, Blaschek HP (2017b) Characterization of a Clostridium beijerinckii spo0A mutant and its application for butyl butyrate production. Biotechnol Bioeng 114(1):106–112. https://doi.org/10.1002/bit.26057
Seo SO, Lu T, Jin Y-S, Blaschek HP (2021) A comparative phenotypic and genomic analysis of Clostridium beijerinckii mutant with enhanced solvent production. J Biotechnol 329(October 2020):49–55. https://doi.org/10.1016/j.jbiotec.2021.02.002
Serrano M, Crawshaw AD, Dembek M, Monteiro JM, Pereira FC, Pinho MG, Fairweather NF, Salgado PS, Henriques AO (2016) The SpoIIQ-SpoIIIAH complex of Clostridium difficile controls forespore engulfment and late stages of gene expression and spore morphogenesis. Mol Microbiol 100(1):204–228. https://doi.org/10.1111/mmi.13311
Seys FM, Rowe P, Bolt EL, Humphreys CM, Minton NP (2020) A gold standard, CRISPR/Cas9-based complementation strategy reliant on 24 nucleotide bookmark sequences. Genes (Basel) 11(4):458. https://doi.org/10.3390/genes11040458
Shaheen R, Shirley M, Jones DT (2000) Comparative fermentation studies of industrial strains belonging to four species of solvent-producing clostridia. J Mol Microbiol Biotechnol 2(1):115–124 10937496
Shen A, Edwards AN, Sarker MR, Paredes-Sabja D (2019) Sporulation and germination in clostridial pathogens. Microbiol Spectr 7(6). https://doi.org/10.1128/microbiolspec.gpp3-0017-2018
Shi Z, Blaschek HP (2008) Transcriptional analysis of Clostridium beijerinckii NCIMB 8052 and the hyper-butanol-producing mutant BA101 during the shift from acidogenesis to solventogenesis. Appl Environ Microbiol 74(24):7709–7714. https://doi.org/10.1128/AEM.01948-08
Sjolander NO, Langlykke AF, Peterson WH (1938) Butyl alcohol fermentation of wood sugar. Ind Eng Chem 30(11):1251–1255. https://doi.org/10.1021/ie50347a011
Song T, Wang F, Xiong S, Jiang H (2019) Surface display of organophosphorus-degrading enzymes on the recombinant spore of Bacillus subtilis. Biochem Biophys Res Commun 510(1):13–19. https://doi.org/10.1016/j.bbrc.2018.12.077
Steiner E, Dago AE, Young DI, Heap JT, Minton NP, Hoch JA, Young M (2011) Multiple orphan histidine kinases interact directly with Spo0A to control the initiation of endospore formation in Clostridium acetobutylicum. Mol Microbiol 80(3):641–654. https://doi.org/10.1111/j.1365-2958.2011.07608.x
Steiner E, Scott J, Minton NP, Winzer K (2012) An agr quorum sensing system that regulates granulose formation and sporulation in Clostridium acetobutylicum. Appl Environ Microbiol 78(4):1113–1122. https://doi.org/10.1128/AEM.06376-11
Sullivan L, Bennett GN (2006) Proteome analysis and comparison of Clostridium acetobutylicum ATCC 824 and Spo0A strain variants. J Ind Microbiol Biotechnol 33(4):298–308. https://doi.org/10.1007/s10295-005-0050-7
Tashiro Y, Yoshida T, Noguchi T, Sonomoto K (2013) Recent advances and future prospects for increased butanol production by acetone-butanol-ethanol fermentation. Eng Life Sci 13(5):432–445. https://doi.org/10.1002/elsc.201200128
Terracciano JS, Kashket ER (1986) Intracellular conditions required for initiation of solvent production by Clostridium acetobutylicum. Appl Environ Microbiol 52(1):86–91. https://doi.org/10.1128/aem.52.1.86-91.1986
Theys J, Lambin P (2015) Clostridium to treat cancer: dream or reality? Ann Transl Med 3(Suppl 1):S21. https://doi.org/10.3978/j.issn.2305-5839.2015.03.39
Thorn GJ, King JR, Jabbari S (2013) pH-induced gene regulation of solvent production by Clostridium acetobutylicum in continuous culture: parameter estimation and sporulation modelling. Math Biosci 241(2):149–166. https://doi.org/10.1016/J.MBS.2012.11.004
Tokuyasu K, Yamada E (1959) Fine structure of Bacillus subtilis. J Biophys Biochem Cytol 5(1):129–133. https://doi.org/10.1083/jcb.5.1.129
Tomas CA, Beamish J, Papoutsakis ET (2004) Transcriptional analysis of butanol stress and tolerance in Clostridium acetobutylicum. J Bacteriol 186(7):2006–2018. https://doi.org/10.1128/JB.186.7.2006-2018.2004
Traag BA, Pugliese A, Eisen JA, Losick R (2013) Gene conservation among endospore-forming bacteria reveals additional sporulation genes in Bacillus subtilis. J Bacteriol 195(2):253–260. https://doi.org/10.1128/JB.01778-12
Tracy BP, Gaida SM, Papoutsakis ET (2008) Development and application of flow-cytometric techniques for analyzing and sorting endospore-forming clostridia. Appl Environ Microbiol 74(24):7497–7506. https://doi.org/10.1128/AEM.01626-08
Tracy BP, Jones SW, Papoutsakis ET (2011) Inactivation of σE and σG in Clostridium acetobutylicum illuminates their roles in clostridial-cell-form biogenesis, granulose synthesis, solventogenesis, and spore morphogenesis. J Bacteriol 193(6):1414–1426. https://doi.org/10.1128/JB.01380-10
Tracy BP, Jones SW, Fast AG, Indurthi DC, Papoutsakis ET (2012) Clostridia: the importance of their exceptional substrate and metabolite diversity for biofuel and biorefinery applications. Curr Opin Biotechnol 23(3):364–381. https://doi.org/10.1016/j.copbio.2011.10.008
Ugwuodo CJ, Nwagu TN (2020) Stabilizing enzymes by immobilization on bacterial spores: A review of literature. Int J Biol Macromol 166:238–250. https://doi.org/10.1016/j.ijbiomac.2020.10.171
Varga J, Stirewalt VL, Melville SB (2004) The CcpA protein is necessary for efficient sporulation and enterotoxin gene (cpe) regulation in Clostridium perfringens. J Bacteriol 186(16):5221–5229. https://doi.org/10.1128/JB.186.16.5221-5229.2004
Vasylkivska M, Jureckova K, Branska B, Sedlar K, Kolek J, Provaznik I, Patakova P (2019) Transcriptional analysis of amino acid, metal ion, vitamin and carbohydrate uptake in butanol-producing Clostridium beijerinckii NRRL B-598. PLoS One 14(11):e0224560. https://doi.org/10.1371/journal.pone.0224560
Vees CA, Neuendorf CS, Pflügl S (2020) Towards continuous industrial bioprocessing with solventogenic and acetogenic clostridia: challenges, progress and perspectives. J Ind Microbiol Biotechnol 1–35:753–787. https://doi.org/10.1007/s10295-020-02296-2
Wang P, Feng J, Guo L, Fasina O, Wang Y (2019) Engineering Clostridium saccharoperbutylacetonicum for high level Isopropanol-Butanol-Ethanol (IBE) production from acetic acid pretreated switchgrass using the CRISPR-Cas9 system. ACS Sustain Chem Eng 7(21):18153–18164. https://doi.org/10.1021/acssuschemeng.9b05336
Wasels F, Chartier G, Hocq R, Lopes Ferreira N (2020) A CRISPR/Anti-CRISPR genome editing approach underlines the synergy of butanol dehydrogenases in Clostridium acetobutylicum DSM 792. Appl Environ Microbiol 86. https://doi.org/10.1128/aem.00408-20
Whitham JM, Tirado-Acevedo O, Chinn MS, Pawlak JJ, Grunden AM (2015) Metabolic response of Clostridium ljungdahlii to oxygen exposure. Appl Environ Microbiol 81(24):8379–8391. https://doi.org/10.1128/AEM.02491-15
Wilkinson SR, Young M (1994) Targeted integration of genes into the Clostridium acetobutylicum chromosome. Microbiology 140(1):89–95. https://doi.org/10.1099/13500872-140-1-89
Wilkinson SR, Young M, Goodacre R, Morris JG, Farrow JAE, Collins MD (1995) Phenotypic and genotypic differences between certain strains of Clostridium acetobutylicum. FEMS Microbiol Lett 125(2–3):199–204. https://doi.org/10.1111/j.1574-6968.1995.tb07358.x
Williams RHN, Whitworth DE (2010) The genetic organisation of prokaryotic two-component system signalling pathways. BMC Genomics 11(1):720. https://doi.org/10.1186/1471-2164-11-720
Wolken WAM, Tramper J, van der Werf MJ (2003) What can spores do for us? Trends Biotechnol 21(8):338–345. https://doi.org/10.1016/S0167-7799(03)00170-7
Woo JE, Lee SY, Jang YS (2018) Effects of nutritional enrichment on acid production from degenerated (non-solventogenic) Clostridium acetobutylicum strain M5. Appl Biol Chem 61(4):469–472. https://doi.org/10.1007/s13765-018-0372-6
Woods DR, Jones DT (1987) Physiological responses of Bacteroides and Clostridium strains to environmental stress factors. Adv Microb Physiol 28(C):1–64. https://doi.org/10.1016/S0065-2911(08)60236-2
Xin F, Wang C, Dong W, Zhang W, Wu H, Ma J, Jiang M (2016) Comprehensive investigations of biobutanol production by a non-acetone and 1,3-propanediol generating Clostridium strain from glycerol and polysaccharides. Biotechnol Biofuels 9(1):220. https://doi.org/10.1186/s13068-016-0641-8
Xin F, Yan W, Zhou J, Wu H, Dong W, Ma J, Zhang W, Jiang M (2018) Exploitation of novel wild type solventogenic strains for butanol production. Biotechnol Biofuels 11(1):252. https://doi.org/10.1186/s13068-018-1252-3
Xin X, Cheng C, Du G, Chen L, Xue C (2020) Metabolic engineering of histidine kinases in Clostridium beijerinckii for enhanced butanol production. Front Bioeng Biotechnol 8:214. https://doi.org/10.3389/fbioe.2020.00214
Xu M, Zhao J, Yu L, Tang I-C, Xue C, Yang S-T (2015) Engineering Clostridium acetobutylicum with a histidine kinase knockout for enhanced n-butanol tolerance and production. Appl Microbiol Biotechnol 99(2):1011–1022. https://doi.org/10.1007/s00253-014-6249-7
Xu M, Zhao J, Yu L, Yang S-T (2017) Comparative genomic analysis of Clostridium acetobutylicum for understanding the mutations contributing to enhanced butanol tolerance and production. J Biotechnol 263:36–44. https://doi.org/10.1016/J.JBIOTEC.2017.10.010
Xu J-Y, Xu Z, Liu X, Tan M, Ye B-C (2018) Protein acetylation and butyrylation regulate the phenotype and metabolic shifts of the endospore-forming Clostridium acetobutylicum. Mol Cell Proteomics 17(6):1156–1169. https://doi.org/10.1074/mcp.RA117.000372
Xue Q, Yang Y, Chen J, Chen L, Yang S, Jiang W, Gu Y (2016) Roles of three AbrBs in regulating two-phase Clostridium acetobutylicum fermentation. Appl Microbiol Biotechnol 100(21):9081–9089. https://doi.org/10.1007/s00253-016-7638-x
Yang X, Tu M, Xie R, Adhikari S, Tong Z (2013) A comparison of three pH control methods for revealing effects of undissociated butyric acid on specific butanol production rate in batch fermentation of Clostridium acetobutylicum. AMB Express 3(1):3. https://doi.org/10.1186/2191-0855-3-3
Yang Y, Lang N, Zhang L, Wu H, Jiang W, Gu Y (2020) A novel regulatory pathway consisting of a two-component system and an ABC-type transporter contributes to butanol tolerance in Clostridium acetobutylicum. Appl Microbiol Biotechnol 1–13:5011–5023. https://doi.org/10.1007/s00253-020-10555-6
Yu HY, Meade A, Liu SJ (2019) Phylogeny of Clostridium spp. Based on conservative genes and comparisons with other trees. Microbiology 88(4):469–478. https://doi.org/10.1134/s002626171904012x
Yutin N, Galperin MY (2013) A genomic update on clostridial phylogeny: Gram-negative spore formers and other misplaced clostridia. Environ Microbiol 15(10):2631–2641. https://doi.org/10.1111/1462-2920.12173
Zahn JA, Saxena J (2011) Novel ethanologenic Clostridium species, Clostridium coskatii. In Patent. http://www.google.com/patents/US20110229947
Zhang Y, Jiao S, Lv J, Du R, Yan X, Wan C, Zhang R, Han B (2017) Sigma factor regulated cellular response in a non-solvent producing Clostridium beijerinckii degenerated strain: a comparative transcriptome analysis. Front Microbiol 8(JAN):23. https://doi.org/10.3389/fmicb.2017.00023
Zhang X, Al-Dossary A, Hussain M, Setlow P, Li J (2020) Applications of Bacillus subtilis spores in biotechnology and advanced materials. Appl Environ Microbiol 86(17):1–22. https://doi.org/10.1128/AEM.01096-20
Zhao H, Msadek T, Zapf J, Madhusudan HJA, Varughese KI (2002) DNA complexed structure of the key transcription factor initiating development in sporulating bacteria. Structure 10(8):1041–1050. https://doi.org/10.1016/S0969-2126(02)00803-1
Zhao Y, Tomas CA, Rudolph FB, Papoutsakis ET, Bennett GN (2005) Intracellular butyryl phosphate and acetyl phosphate concentrations in Clostridium acetobutylicum and their implications for solvent formation. Appl Environ Microbiol 71(1):530–537. https://doi.org/10.1128/AEM.71.1.530-537.2005
Zhao R, Liu Y, Zhang H, Chai C, Wang J, Jiang W, Gu Y (2019) CRISPR-Cas12a-mediated gene deletion and regulation in Clostridium ljungdahlii and its application in carbon flux redirection in synthesis gas fermentation. ACS Synth Biol 8(10):2270–2279. https://doi.org/10.1021/acssynbio.9b00033
Zheng Y-N, Li L-Z, Xian M, Ma Y-J, Yang J-M, Xu X, He D-Z (2009) Problems with the microbial production of butanol. J Ind Microbiol Biotechnol 36(9):1127–1138. https://doi.org/10.1007/s10295-009-0609-9
Funding
The work presented in this article was financed by the European Union Marie Sklodowska Curie Innovative Training Networks (ITN) - Contract number 642068 (CLOSPORE) and the European Union’s Horizon 2020 research and innovation programme under Grant Agreement no. 761042 (BIOCON-CO2).
Author information
Authors and Affiliations
Contributions
MD designed the structure of the review, performed the literature search and data analysis, and wrote the manuscript. MD, SWMK, and AMLC revised the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diallo, M., Kengen, S.W.M. & López-Contreras, A.M. Sporulation in solventogenic and acetogenic clostridia. Appl Microbiol Biotechnol 105, 3533–3557 (2021). https://doi.org/10.1007/s00253-021-11289-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11289-9