Abstract
Late blight caused by Phytophthora infestans is an economically important disease of potato and tomato worldwide. In Canada, an increase in late blight incidence and severity coincided with changes in genetic composition of P. infestans. We monitored late blight incidence on tomato and potato in Pacific western and eastern Canada between 2019 and 2022, identified genotypes of P. infestans, and examined their population genetic diversity. We identified four major existing genotypes US11, US17, US8, and US23 as well as 25 new genotypes. The US11 genotype was dominant in Pacific western Canada, accounting for 59% of the total population. We discovered the US17 genotype for the first time in Canada. We revealed a higher incidence of late blight and quite diverse genotypes of P. infestans in Pacific western Canada than in eastern Canada. We found high genetic diversity of P. infestans population from Pacific western Canada, as evidenced by the high number of multilocus genotypes, high values of genetic diversity indices, and emergence of 25 new genotypes. Considering the number of disease incidence, the detection of diverse known genotypes, the emergence of novel genotypes, and the high number of isolates resistant to metalaxyl-m (95%) from Pacific western Canada, the region could play a role in establishing sexual recombination and diverse populations, which could ultimately pose challenges for late blight management. Therefore, continuous monitoring of P. infestans populations in Pacific western region and across Canada is warranted.
Key points
• Genotypes of P. infestans in Pacific western were quite diverse than in eastern Canada.
• We discovered US17 genotype for the first time in Canada and identified 26 novel genotypes.
• Approximately 95% of P. infestans isolates were resistant to metalaxyl-m.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potato is a valuable crop as a staple food source and for the production of textiles, pharmaceuticals, and therapeutics (Llorente et al. 2010). Potato is produced in all Canadian provinces with 40, 38, and 22% in area of potato production in the Prairie provinces and British Columbia (BC), Atlantic Canada, and Central Canada, respectively (Anonymous 2021a). Potato contributed $1.4 billion in farm cash receipts in Canada and $2 billion in exports of potato and potato products in 2020 (Anonymous 2021a). Tomato is also grown as an annual crop in most Canadian provinces, with the highest production in Ontario (ON), followed by BC and Quebec (QC) in 2020, and the summer months are the peak production periods (Anonymous 2021b). In Canada, tomato production ranks first among greenhouse vegetables and contributes about $665.9 million in farm gate values (Anonymous 2021b).
Several diseases cause adverse impacts on the yield and quality of potato and tomato in Canada and worldwide (Soylu et al. 2006; Birch et al. 2012; Arafa et al. 2022). Of these, late blight caused by Phytophthora infestans (Mont.) de Bary is the most destructive and economically important disease of potato and tomato. It can destroy an entire field within a week when severe epidemics occur favored by low to moderate temperature (15 to 22 °C), high relative humidity (> 90% RH), and rainfall (Haverkort et al. 2008; Fry 2020; Hansen et al. 2016a; Arafa et al. 2022), resulting in global losses up to $6.7 billion annually. The pathogen caused the Great Famine in Ireland in the nineteenth century, which resulted in the death of over a million people and the mass emigration of more than a million (Fry et al. 1993; Rekad et al. 2017). In Canada, late blight of potato and tomato is widespread and usually occurs yearly with the highest pressure in eastern provinces, while it is generally localized with low to moderate pressure in BC and Alberta (AB) (Anonymous 2021a, b). Millions of dollars have been spent on fungicide applications worldwide to protect potato and tomato from late blight (Haverkort et al. 2008; Fry et al. 2013). Studies from Finland showed that the number of fungicide sprays to manage late blight in potato increased eight-fold from 1983 (0.5 sprays) to 2002 (4 sprays) in one season (Hannukkala et al. 2007).
P. infestans has both asexual and sexual life stages. The pathogen is a heterothallic oomycete with two designated mating types A1 and A2. Sexual populations with diverse genetic diversity were reported from the highlands of central Mexico and northwest Europe (Grunwald and Flier 2005; Widmark et al. 2011). In contrast, asexual populations dominate in Canada, the USA, and the majority of other potato and tomato-growing areas worldwide (Fry 2016). Despite the predominance of asexual population, high genetic diversity was reported in North America (Kalischuk et al. 2012; Peters et al. 2014; Saville and Ristaino 2019; Saville et al. 2021), which could be due to mutation, mitotic recombination, and gene conversion (Abu-El Samen et al. 2003; Maurice et al. 2019). It is speculated that sexual recombination contributes little to disease epidemics in most of the regions of North America, though both mating types were found occasionally in a few locations (Fry 2016). For example, both mating types were found in ON in 2011 and BC in 2012 and 2013, indicating the potential of evolving sexual strains of P. infestans in Canada (Peters et al. 2014). However, further studies are required for verification. Nevertheless, late blight management will be difficult and complex if oospores exist in a field because the spores are hard and thick-walled and can tolerate adverse environmental conditions and survive several years in the soil (Mayton et al. 2000).
Several markers including allozymes of glucose-phosphate isomerase (Gpi), mating type, restriction fragment length polymorphism (RFLP) RG57 fingerprinting, and simple sequence repeats (SSR) or microsatellites have been used for characterization of P. infestans (Peters et al. 2014; Alkher et al. 2015). In North America, strains (genotypes) of P. infestans were traditionally identified using allozyme banding patterns and RFLP. RFLP fingerprinting is laborious, time-consuming, and technically difficult. Therefore, SSR markers have been preferred for genotyping P. infestans in recent years, as multiplexing of all pairs of SSR primers can be done in a single reaction, allowing genotyping in a short period of time (Saville and Ristaino 2019). In addition, SSR markers are highly polymorphic, affordable, reproducible, neutral, co-dominant, and easy to automate and score (Lees et al. 2006; Li et al. 2013; Arafa et al. 2018). Furthermore, SSR genotyping can identify the pathogen genotypes directly from infected tissues without isolating the pathogen (Guichoux et al. 2011; Li et al. 2013).
In Canada, displacement of previous populations of P. infestans by new populations was reported in 1994–1996 (Punja et al. 1998) and 2009–2011 (Peters et al. 2014). In 1994–1996, the mefenoxam-sensitive strain US1 was displaced by the mefenoxam-resistant strain US8. In 2009–2011, US8 strains were displaced by the US23 strains. New strains including US22 and US24 were also reported first in 2010 and later in 2011 (Alkher et al. 2015; Peters et al. 2014). Diverse strains including US7, BC11, BC13, BC14, and BC15 were found between 1993 and 1997 in BC (Punja et al. 1998); after that, no comprehensive studies were conducted to understand the population dynamics of P. infestans strains. From several regions of the world, it is reported that changes in strains or genotypes of P. infestans and their migration to new areas or regions lead to increased incidence and outbreaks of late blight of potato and tomato. For example, the 2009 pandemic of late blight in the northeast USA occurred due to the clonal lineage US22, and the 2014 late blight outbreak in West Bengal, India, occurred due to strain 13_A2 (Fry 2016; Saville et al. 2016). These epidemics underscore the importance of continuing the study of P. infestans populations to understand population dynamics and tailor late blight management programs and strategies. Therefore, the research objectives of this study were to (i) assess late blight occurrence in potato and tomato over multi-years from diverse regions in Pacific western Canada, (ii) identify the genotypes of P. infestans populations in Pacific western and eastern provinces of Canada, and iii) understand the genetic diversity and population structure of P. infestans isolates from the Pacific western Canada.
Materials and methods
Disease monitoring, sampling, and pathogen isolation and identification
Tomato and potato commercial fields, small farms, field plots in research centers, and community gardens from diverse locations in BC were visited during June to November between 2019 and 2021. Percent severity of late blight was recorded from the spots or plots where the disease was observed and plant tissue samples showing late blight symptoms were collected. Pathogen-infected samples collected from commercial potato fields included the potato varieties Kennebeck, AC Peregrine, Gemstar Russet, Russet Norkotah, and Warba. Potato samples were also collected from small farms, research centers, and a few community gardens, whereas most of the tomato samples were from community gardens and a few from small farms and home gardens. Potato leaf and stem samples and tomato leaf, stem, and fruit samples showing symptoms of late blight were collected from each field. The samples were packed in Ziploc plastic bags and stored at 4 °C before processing for pathogen isolation. Sampling details including numbers of samples, fields, host, locations, and years of collections are summarized in Table 1.
To isolate the pathogen, foliar tissue samples (leaves, stems) of potato and tomato, as well as fruits of tomato showing late blight symptoms, were washed with running tap water, and pieces (~ 3 cm × 4 cm) of the plant tissues with healthy and infected portions were cut and surface sterilized using 75% ethanol for 40 s, then immersed in 10% commercial bleach for 45 s, and finally rinsed three times with sterile water. These sterile plant tissues were kept in a moist chamber (Petri dishes with moist filter paper) and incubated at 18 °C for 3–5 days. Incubated plant samples were checked under a binocular microscope. Small pieces of mycelium with typical lemon-shaped sporangia were transferred onto either V8 rye PAR (Sapkota et al. 2023) or pea agar (120 g of frozen peas, 15 g of agar, 1 L of distilled water amended with 200 \(\mu\) l pimaricin, 250 mg ampicillin, 400 \(\mu\) l rifampicin). Plates were incubated for 2–4 weeks, and colony growth was examined and further purified as needed. Each isolate was obtained from individual lesions of infected plant samples, and the hyphal tip method was used to obtain each isolate after purification. Pathogen isolates were identified as P. infestans using colony morphology on V8 rye PAR and pea agar (white coenocytic non-septate mycelia), as well as sporangia morphology (oval-shaped, ellipsoidal to lemon-shaped, spindle-like in the base and semi-papillate) (Gallegly and Hong 2008). Reference isolates of P. infestans from previous studies (Peters et al. 2014) were also included as positive controls for identifying unknown isolates. All isolates recovered in this study were maintained on half-strength acidified V8 rye PAR or pea agar for short-term use. For long-term storage, isolates were grown on rye seeds placed on V8 rye PAR, and infected rye seeds were then placed in 2-mL vials with sterile water and stored at 10 °C. All the strains (Strain IDs: CAPi1- CAPi214) used in this study were deposited in the long-term storage of Plant Pathology Laboratory in Agassiz and Charlottetown Research Centres of Agriculture and Agri-Food Canada, which are public research institutes.
Weather data
Daily temperature and rainfall were recovered using weather stations at the Agassiz Research and Development Centre (ARDC) in Agassiz and substations at Abbotsford, Pitt Meadows and Vancouver, BC. The weather stations were equipped with temperature and relative humidity sensors (SEN-R Combisensor Temp/rh Adcon TR1) and rain gauges (Adcon Telemetry, Klosterneuburg, Austria) to log weather data in near real time at 15-min interval. Average rainfall and temperature data were calculated using daily data and weather stations that were in operation between 2019 and 2021.
Pathogen DNA extraction
Agar plugs of individual P. infestans isolates were taken from the edge of actively growing colonies on pea agar and transferred to pea broth supplemented with antibiotics (120 g of frozen peas, 1 L of distilled water amended with 200 \(\mu\) l pimaricin, 250 mg ampicillin, 400 \(\mu\) l rifampicin) in (15 mm × 15 mm) Petri plates. The Petri plates were incubated in a growth chamber at 16 °C for 2 to 3 weeks. Mycelia were harvested by filtering the broth through a Buchner funnel containing filter paper (Whatman no. 1, Kent, UK). Mycelia of isolates were freeze-dried overnight or for 2 days and stored at − 20 °C until further use. Genomic DNA was extracted using freeze-dried mycelia of each isolate as described by Goodwin et al. (1992) and Fry (2013). The quantity of the extracted DNA samples was measured using Qubit (Invitrogen, CA, USA).
Preparation of infected plant tissue for genotyping
Using Whatman’s FTA (Flinders Technology Associates) cards, late blight-infected potato tissues were prepared for SSR genotyping of P. infestans. This technique does not require culturing the pathogen, and infected plant tissue can be used directly. We processed six samples on FTA card for SSR genotyping in this study to determine the reliability of the method, as SSR genotyping using FTA card directly from infected late blight samples was not commonly used in Canada. To process the sample, the margins of actively spreading lesions of infected plant tissues were pressed onto FTA cards (Whatman Inc., Kent, UK) by applying moderate rubbing pressure using a pestle. The processed FTA cards were air-dried and stored in a desiccator until further use. Samples from FTA cards were processed and cleaned as described by Burlakoti et al. (2020). Briefly, two pieces of FTA disc (2 mm diameter size) were punched using a Harris Uni-Core™ (Ted Pella Inc., CA, USA), then were kept in a 1.5-ml microcentrifuge tube, and purified according to the manufacturer’s instructions (Whatman® FTA® Card Technology, Kent, UK). Purified pieces of card were placed directly as DNA templates into the PCR mix for the multiplex SSR genotyping.
Multiplexed SSR genotyping
Multiplex SSR genotyping of P. infestans isolates (n = 210) and pathogen-infected potato samples from Delta obtained in 2022 processed on FTA cards (n = 6) from this study and four reference isolates, US8, US11, US23, and US24 from Peters et al. (2014), were amplified using 12 sets of multiplex SSR primers as described in Li et al. (2013) and Cooke (2020) (Table 2). For PCR reaction, Qiagen (MD, USA) Microsatellite PCR kit and 20 to 25 ng template DNA were used and all other PCR reactions and thermal cycling conditions were used as described in Li et al. (2013), whereas concentration of primers was used as described by Cooke (2020). Amplified PCR products were diluted in distilled water to 1:10 ratio, and samples were genotyped at the Sequencing Center of the University of British Columbia, Vancouver, using the ABI 3730 capillary DNA sequencer (Applied Biosystems, CA, USA) according to the manufacturer’s instructions.
The peak size was determined against a GeneScan 500 LIZ standard (Applied Biosystems, CA, USA), and alleles were scored manually using the Peak Scanner v2 (Applied Biosystems, CA, USA) with fragment sizes rounded up to the nearest whole number for analysis. After that, the allelic banding patterns were compared with those of the reference isolates and other published data and assigned their nomenclature accordingly. Ten out of 12 SSR markers produced multiple alleles (2 to 11 alleles), whereas markers Pi04 and PinfSSR6 had 2 alleles (Table 2). For SSR genotyping data, all loci were considered triploid to include the third allele present in those isolates. When a third allele was not present in those individual isolates, it was considered a null allele. The SSR classifier tool (USABlight https://usablight.org/identify-ssr-genotype/, retrieved on 27 December 2022) was used to identify genotypes of isolates. The tool identifies the genotype of isolates by comparing allelic patterns of unknown isolates with reference isolates. Those isolates whose genotypes did not match with known genotypes in the SSR classifier tool at https://usablight.org were assigned as new genotypes (Table 1 and Fig. 1) based on the visual examination of allelic patterns of isolates and Bruvo’s genetic distance used in constructing a neighbor-joining (N-J) tree to visualize the clades (Olave-Achury et al. 2022). Using the Poppr v.2.8.3 (Kamvar et al. 2014) and R library adegenet v.2.1.1 (Jombart 2008), N-J tree was constructed based on Bruvo’s distance and a combination model (genome addition and genome loss). The data set was bootstrapped using 1000 replicates. In the N-J tree, we used P. infestans isolates from this study (204 isolates from BC, 2 isolates from ON, and 4 isolates from QC, Canada) and 34 reference isolates from the USA, Europe, and South America retrieved from the USABlight archive at https://usablight.org. A genetic distance of 0.28 was used as the cut-off value to determine the exact relatedness of two or more isolates. Nomenclatures of new genotypes were given as “CAC1 to CAC25” and “CAE1” (Table 1).
Neighbor-joining phylogram of Phytophthora infestans isolates from this study and reference isolates of US and European genotypes of P. infestans obtained from USABlight (https://usablight.org/identify-ssr-genotype/) and EuroBlight (https://agro.au.dk/forskning/internationale-platforme/euroblight) archives. P. infestans isolate codes with green color represent the reference isolates; blue represents British Columbia (BC) and Quebec potato isolates and Ontario (ON) tomato isolates; and red represents BC tomato isolates. Following letters in isolates code mean: US, United States; EC, Ecuador; BR, Brazil; T, tomato; P, potato; AB, Abbotsford; AG, Agassiz; CHI, Chilliwack; D, Delta; PM, Pitt meadows; SS, South Surrey; VAN, Vancouver Island; RCH, Richmond; ON, Ontario; QC Quebec; 19, 2019; 20, 2020; 21, 2021; 22, 2022; CAC, Canada coast; CAE, eastern Canada. AB, AG, CHI, D, PM, SS, VAN, and RCH are locations from BC
Population analyses
Using genotyping data of 12 SSR loci of P. infestans isolates (n = 204) from BC, the genetic diversity and population structure of those isolates were analyzed. P. infestans isolates were classified into populations based on host type, location, year, and major genotypes (Table 3). The R library Poppr (Kamvar et al. 2014) was used to calculate population statistics (genetic diversity). The population statistics included the following: MLGs, the number of multilocus genotypes; eMLG, the number of expected MLGs at the smallest sample size of at least 10 (Hurlbert 1971); H, Shannon-Weiner index of MLG diversity (Shannon 2001); G, Stoddart and Taylor index of MLG diversity (Stoddart and Taylor 1988); λ, Simpson index corrected for sample size by multiplying the index value by N/(N-1) (Wang et al. 2017); evenness (Grünwald et al. 2003); Hexp, Nei’s unbiased gene diversity; IA, index of association (Brown et al. 1980); and r¯d, the standardized index of association which removes sample size bias (Agapow and Burt 2001). Clone-corrected data were used in calculating the index of association (IA) and standardized index of association (r¯d), whereas all data were used to estimate other population diversity indices. R library Poppr v.2.8.1 (Kamvar et al. 2014) was used to generate clone-corrected data, where clonal isolates were removed such that each population contained only one representative of each haplotype.
The population structure of P. infestans isolates was analyzed using the software Structure Harvester v2.3.3 (Pritchard et al. 2000) and clone-corrected data were used. The data were run using 20,000 burn-in repeats and 1,000,000 Markov Chain Monte Carlo (MCMC) repeats under an admixture model to accommodate samples that may have originated from sexual recombination events. Independent runs of the model used K values from 1 to 10 with 20 replicate runs at each value of K. The optimal K was estimated using the Evanno method in the web tool Structure Harvester (Evanno et al. 2005; Earl and VonHoldt 2012). Also, the optimal K was inferred by directly observing the groupings of the samples by their assigned Q values. All runs for optimal K and non-optimal K values were averaged using CLUMPP v1.1.2 (Jakobsson and Rosenberg 2007) and visualized with the program DISTRUCT v.1.1 (Rosenberg 2004).
In addition, the genetic groupings were visualized by conducting a principal component analysis (PCA) and discriminant analysis of principal components (DAPC) using the R library adegenet (Jombart 2008; Beninal et al. 2022). PCA was constructed using the SSR data and isolates were grouped based on a combination of location, sampling year and host, whereas DAPC was constructed using SSR data from only BC to detect variation in BC isolates. To understand the variability in P. infestans populations between hosts, and among years, locations, and genotypes, a minimum spanning network (MSN) was constructed for samples from BC (n = 204), ON (n = 2), and QC (n = 4) based on Bruvo’s genetic distance using R library adegenet v2.1.1 (Jombart 2008) in the R package Poppr (Kamvar et al. 2014).
Mating type and allozyme analyses
Mating types of the representative isolates of P. infestans (n = 66) were determined using the protocol described by Alkher et al. (2015). Strains US24 (A1) and US8 (A2) previously characterized by Peters et al. (2014) were used as tester strains. In brief, isolates forming oospores on plates paired with the A1 mating type (US24) were designated as the A2 mating type, while those that formed oospores on plates paired with the A2 mating type (US8) were designated as the A1 mating type.
Representative isolates of P. infestans (n = 56) were examined at the Gpi locus using cellulose acetate electrophoresis (CAE) to determine allozyme genotypes. The isolates were grown on pea agar (Peters et al. 2014) for 2 weeks, and mycelia were scrapped from the plates and placed in a 1.5-mL Eppendorf microcentrifuge tube and processed as described by Goodwin et al. (1995a) and Peters et al. (2014). Reference isolates (US 8, US11, US 22, US23, and US24) were included in the run, and the migration distances of Gpi proteins from the reference genotypes were compared with those from the uncharacterized isolates.
In vitro sensitivity to metalaxyl-m
Metalaxyl-m sensitivity of the P. infestans isolates (n = 183) was determined using technical grade metalaxyl-m (Syngenta, Basel, Switzerland). Two experiments were conducted with three replicates in each experiment and the data were pooled for analysis. Metalaxyl-m sensitivity assays were conducted using methods described previously (Peters et al. 2014; Alkher et al. 2015). Briefly the stock solution of metalaxyl-m was prepared as 100 mg ml−1 in dimethyl sulfoxide (DMSO). Final concentrations of 0, 1, 10, and 100 µg ml−1 by amending stock solution to clarified pea agar. Radial growth of each isolate was measured at 14 days after incubation. Isolates were categorized as: metalaxyl sensitive (MS), which had < 10% growth; metalaxyl moderately resistant (MMR), which had 10–60% growth; and metalaxyl highly resistant (MHR), which had > 60% growth at 100 µg ml−1 of metalaxyl-m.
Results
Prevalence of late blight and pathogen isolation
Late blight was found in BC in all 3 years (2019 to 2021); however, disease incidence timing and severity varied greatly over the years. Late blight was found in several commercial farms, research and small farms, and home and community gardens of eight diverse locations in BC (Table 1). In 2019, the disease was observed in field-grown tomatoes and potatoes on small farms in Chilliwack and Abbotsford during the second week of September and late blight severity was very high (~ 60 to 100%). During mid-September to the first week of October, the disease was also found in tomatoes and potatoes at several community and home gardens in Agassiz, Chilliwack, Abbotsford, and Pitt Meadows. Disease severity was also high (~ 40 to 50%) in three locations and completely wiped out in most of the plots in Pitt Meadows. In addition, late blight was found in a few patches of potato fields of the Agassiz research farm and commercial farms in Cloverdale, BC, in late September and early October; the limited spread of the disease in these plots could be due to routine application of fungicides. In 2020, late blight symptoms were observed early in the season (between mid-July and mid-August) in several commercial potato farms in BC (n = 10) including Delta, Richmond, Surrey, and Abbotsford. Late blight was found in a few patches of these fields with low to moderate disease severity (~ 15 to 25%). Late blight symptoms were also observed in tomatoes grown at several communities and home gardens in the Fraser Valley regions including Chilliwack, Abbotsford, and Pitt Meadows in August and September. Late blight severity in these communities and home gardens was high (~ 40% to 100%). In 2021, late blight symptoms were not found in commercial potato farms, but the disease was observed late in the season (early to mid-October) in community gardens in Abbotsford and Agassiz. Late blight was not found in most of the locations from BC in 2022, but the disease was reported in few farms in Delta. A total of 204 isolates of P. infestans (83 in 2019, 94 in 2020, and 29 in 2021) were obtained from pathogen-infected samples of potatoes (n = 57) and tomatoes (n = 147) from 28 fields in 8 locations in BC during 2019–2021 (Table 1). P. infestans cultures were isolated from infected leaves and stems of tomato and potato and fruits of tomato.
In contrast, late blight incidence was very low or absent in other provinces of Canada during 2019–2022 due to the unfavorable climatic conditions (persistent high temperature, low humidity, and dry weather) during the growing seasons (data not shown). However, the disease was sporadic in eastern Canada and two isolates of P. infestans were obtained from tomato samples from ON in 2021 and four isolates from potato samples were obtained from QC in 2022.
Identification of pathogen genotypes
Genotypes were identified directly from six infected tissue samples processed on FTA cards as well as DNA extracted from pure cultures of 210 isolates. Twelve SSR markers were able to amplify six samples processed directly on FTA cards and identified two genotypes, US11 (n = 5) and US17 (n = 1) of P. infestans. Among the SSR loci of 210 isolates, SSR4 had the highest number of alleles (n = 11), and Pi70 and SSR6 had the least number of alleles (n = 2) (Table 2). Among six potato isolates from eastern Canada, 5 were of US23 genotype and one isolate was a new genotype (CAE1). Among 204 isolates of P. infestans in BC, we identified US8 (3%), US11 (59%), US17 (7%), and 25 new genotypes “CAC1 to CAC25” (30%) (Table 1). The details of allelic patterns of 12 SSR makers of all genotypes were provided in Supplemental Table S1. Frequency distribution of pathogen genotypes varied with host, location, and year of collection. US8 was exclusively found in potato samples and only from three locations. US17 was detected for the first time in BC, and this genotype was found in both hosts and in multiple locations and years at low frequencies. US11 was the most dominant genotype across hosts, years, and locations. Approximately 67% of tomato isolates and 35% of potato isolates were of the US11 genotype. The frequency of US11 was very high in tomato samples collected from home and community gardens of Chilliwack (94%), Pitt Meadows (67%), and Vancouver Island (100%). The frequency of US11 was also high across years in these locations. In 2021, we only isolated P. infestans from tomato foliage from community gardens of Agassiz (n = 15) and Abbotsford (n = 12), where 58% of the isolates in Agassiz were new genotypes and 80% of the isolates from Abbotsford were US11. We discovered 25 new genotypes in BC, whose SSR banding patterns did not match with previously known genotypes. Phylogenetic analyses showed that most of these new genotypes were genetically close to US11, and a few of them were close to US17 and US8 (Fig. 1). A few new genotypes were in distinct clusters. New genotypes CAC5, CAC6, and CAC11 were recovered from both potato and tomato samples, whereas the other genotypes were found exclusively on either potatoes or tomatoes (Table 1 and Supplemental Table S1). Only a few new genotypes (CAC2 and CAC24) were detected from multiple locations, whereas the vast majority of them (> 95%) were only found in a single location and in the same year. The highest frequency of new genotypes in tomato samples was found in Abbotsford in 2019 (63%), followed by Agassiz in 2021 (58%), whereas the highest frequency in potato samples was observed in commercial farms in Delta (55%) and Richmond (86%).
Genetic diversity
High genetic diversity was found in P. infestans populations categorized based on host, location, and year (Table 3). Overall, 56 MLGs were detected from BC populations (n = 204) and five MLGs from isolates from eastern Canada (n = 6). Among the populations based on host and year, the 2021 tomato had the highest MLGs and diversity indices and closely followed by the 2020 potato population (Table 3). High values of H, G, λ, evenness, and Hexp were found in populations categorized based on the host in each year (Table 3). Among years, the 2019 population showed slightly lower numbers of MLGs (23 out of 83 isolates) than the populations from 2020 (30 out of 65) and 2021 (15 out of 27). Similar patterns of other genetic diversity indices (H, G, λ, evenness, and Hexp) were observed between the 2020 and 2021 populations, whereas the 2019 population had slightly lower indices of genetic diversity than the other 2 years (Table 3). Among locations, Abbotsford (17 out of 53 isolates), Agassiz (19 out of 32), and Pitt Meadows (20 out of 45) had higher MLGs than other locations. Values of H, G, λ, evenness, and Hexp were also high in the pathogen populations from these locations (Table 3). Among populations categorized based on genotype group, the US11 genotype had the highest genetic diversity indices (H = 1.698, G = 3.47, and λ = 0.712) (Table 3).
Values of IA and r¯d were further away from zero in potato population (IA = 3.275 and 1.731 for 2020 and 2019, respectively), indicating a greater level of clonality, whereas these values were close to zero in tomato populations, indicating a greater level of recombination. These values were close to zero in populations from Chilliwack and Pitt Meadows and most of the isolates from these locations were from tomatoes. Within a potato population, isolates obtained from Delta had an IA furthest from 0 (IA = 3.0704), indicating the greatest level of clonality, whereas isolates from Richmond showed the greatest level of recombination (IA = − 0.0556 and r¯d = − 0.0186) (Table 3).
Population structure
The N-J tree distinguished the European and American isolates by separating them into two major clades. Isolates from this study that matched with known genotypes (US8, US11, US17, and US23) clustered with corresponding genotypes. For example, the US23 reference isolates clustered with those obtained from eastern Canada and the US8 isolates from BC formed a clade with the US8 reference isolate. A few isolates from our study either formed a distinct cluster or clustered with either US8 or US17, whereas the majority of the new isolates clustered with US11 (Fig. 1).
Population structure analysis using Structure harvester determined the optimal K value of 8 based on the Evanno method (Fig. 2). At K = 8, the population clustered based on the identified genotypes. Isolates (US23 var and CAE) from ON and QC clustered together. Most of the isolates identified as the US11 formed a distinct cluster with slight subclonal variations among the US11 isolates. Isolates of US17 and US8 formed separate clusters, respectively (Fig. 2). New genotypes CAC11 and CAC22 formed another cluster and grouped closely with the US11 isolates (Fig. 2). Isolates belonging to the new genotypes CAC13, CAC14, CAC16, and CAC17 formed separate individual clusters. Likewise, new genotypes CAC20 and CAC21 formed a cluster (Fig. 2). In contrast, new genotypes CAC3, CAC12, CAC15, CAC10, CAC25, CAC5, CAC8, CAC1, and CAC19 are distinct individually and no clusters formed.
Structure analysis of Phytophthora infestans isolates from British Columbia, Ontario, and Quebec using allelic data generated from 12-plex simple sequence repeats (SSR) loci. The graph was generated from a single run of structure (K values 1 to 10; burn-in chain of 20,000 repeats; Markov chain Monte Carlo of 1,000,000 repeats; each K value repeated 20 times)
PCA revealed that P. infestans isolates from ON and QC (ON21T and QC22P) representing the genotypes US23 and CAE1 clustered together and were distinct from other genotypes (Fig. 3a). Likewise, BC isolate groups AB20P, AG19P, SS19P, and D20P consisting of the US8 and CAC12 formed a separate cluster, while the remaining isolates from BC including the US17, US11, and other novel isolates clustered together with varying subclonal variations observed among the groups (Fig. 3a). DAPC showed that the US17 and other new genotypes from Agassiz and Abbotsford in 2019 and 2021 from potato and tomato formed a distinct cluster (Fig. 3b). Another distinct cluster was formed by isolates from Abbotsford in 2019, alongside isolates originating from tomato samples in Pitt Meadows in 2020 (Fig. 3b).
a Principal component analysis (PCA) plot of Phytophthora infestans isolates from British Columbia, Ontario, and Quebec using allelic data generated from 12-plex simple sequence repeats (SSR) loci. All isolates from this study were included. T, tomato; P, potato; AB, Abbotsford; AG, Agassiz; CHI, Chilliwack; D, Delta; ON, Ontario; QC, Quebec; PM, Pitt meadows; SS, South Surrey; VAN, Vancouver Island; RCH, Richmond; and 19, 2019; 20, 2020; 21, 2021; and 22, 2022. b Discriminant analysis of principal components (DAPC) plot of P. infestans isolates from British Columbia
MSN showed the US8, CAC12, and J12 (AB20P, AG19P, SS19P, and D20P) isolates clustered together to form a unique cluster on the same branch on MSN (Figs. 5a and b). Also, P. infestans isolates from ON and QC (ON21T and QC22P) comprising the US23 and CAE1 formed a distinct branch. The US17 isolates clustered together and formed a branch (Fig. 4a and b) while CAC24 and CAC25 were located close to the center of the MSN as was CAC13. The clustering observed using the MSN is similar to those observed for other clustering analyses, but the MSN revealed more subclonal variations in US11 and some of the novel genotypes (Fig. 4a and b).
a Minimum spanning network (MSN) tree of Phytophthora infestans isolates from British Columbia, Ontario, and Quebec using allelic data generated from 12-plex simple sequence repeats (SSR) loci. Bruvo’s distance was used to generate MSN tree based on genotypes of P. infestans. Node size is proportional to the number of isolates of P. infestans. US, United States; CAC, Canada coast; CAE, eastern Canada. b MSN tree of P. infestans isolates from British Columbia, Ontario, and Quebec. Bruvo’s distance was used to generate MSN tree based on a combination of location, year, and host. The letters on the figure artwork indicate the following: T, tomato; P, potato; AB, Abbotsford; AG, Agassiz; CHI, Chilliwack; D, Delta; ON, Ontario; QC, Quebec; PM, Pitt meadow; SS, South Surrey; VAN, Vancouver Island; RCH, Richmond; 19, 2019; 20, 2020; 21, 2021
Mating type and allozyme analyses
P. infestans isolates from ON and QC (n = 6) and BC (n = 60) were of the A1 mating type. These include the US11, US17, US23, and a few of the new genotypes (Table 4). Allozyme analyses showed two Gpi banding pattern types were observed across the sampling years and locations consisting of the Gpi 100/100/111 and Gpi 100/100 (Goodwin et al. 1995a) belonging to the US11, US17, US23, and some new genotypes (Table 4). Overall, P. infestans isolates from BC banded as either 100/100/111 or 100/100 whereas all isolates from QC and ON banded as 100/100.
In vitro sensitivity to metalaxyl-m.
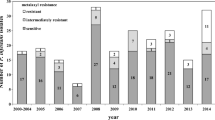
About 95% of P. infestans (n = 181) originating from BC between 2019 and 2021 were resistant (MMR or MHR) to metalaxyl-m (Table 5). The frequency of metalaxyl-resistant isolates was high across genotypes, years, and locations, whereas the US23 and CAE1 isolates (n = 2) originating from both ON and QC were sensitive to metalaxyl (Table 5).
Discussion
Late blight caused by P. infestans continues to impact the yield and marketability of potato and tomato in North America and elsewhere, as there has been an increase in disease severity, incidence, and pathogen diversity mainly because of the introduction of new and emerging genotypes (Alkher et al. 2015; Kalischuk et al. 2012; Hu et al. 2012; Fry et al. 2013). These new genotypes often pose disease management challenges because new genotypes may have different phenotypic traits, such as increased virulence and increased insensitivity to commonly used fungicides. In this context, we conducted a comprehensive study to monitor genotypes of P. infestans from tomato and potato in Pacific western Canada. Furthermore, we examined their genetic diversity and population structure and also compared these isolates with a few isolates from eastern Canada.
In the Pacific western Canada, we found a wide variability in first symptoms occurrence and intensity of late blight among multiple years and locations, which was associated with the variability in climate patterns across years. Late blight occurred on potato and tomato plants grown in home and community gardens in several locations of the Fraser Valley areas in 2019 and 2020 after mid-September, indicating that home and community gardens could be good reservoirs of P. infestans. Amount and frequencies of rainfall were high in the fall in both years, which afforded a conducive environment for late blight occurrence (Fig. 5). Interestingly, frequent rainfall occurred in summer months (June and July) in 2020, which led to early onset of late blight in several commercial potato farms in BC. In 2020, the greatest precipitation period was approximately 107 mm coupled with prolonged damp weather conditions characterized by high humidity favorable for P. infestans proliferation (Fig. 5). This is reflected in the number of isolates obtained in 2020 particularly because of high, multiple late blight incidence rates across BC. In contrast, a prolonged drought occurred from mid-June to late October in 2022 and 2021; therefore, a little incidence of late blight was found in 2022 in early summer, where the disease was found only after mid-October in 2021 in community gardens from two locations. In eastern Canada, dry, hot, and less humid conditions persisted (data not shown) resulting in less or no late blight incidences.
Over the years, various biochemical and molecular markers were used to characterize P. infestans populations in Canada and elsewhere to monitor the rapid displacement of common genotypes. More recently, microsatellite markers have been used in Canada (Peters et al. 2014; Alkher et al. 2015), extensively in the USA (Danies et al. 2013; Saville et al. 2016; Saville and Ristaino 2019, 2021) and elsewhere (Dey et al. 2018; Shimelash and Dessie 2020; Saville et al. 2021; Olave-Achury et al. 2022) to identify the genotypes of P. infestans populations as well as to study population genetic diversity. Multiplex microsatellite genotyping technique made it feasible to rapidly process a large number of isolates. In addition to genotyping several hundreds of P. infestans isolates, we also demonstrated that the pathogen genotypes can be identified directly from pathogen-infected potato leaf samples. FTA cards were used for genotyping P. infestans in the United States (Saville and Ristaino 2019, 2021), in Europe (Li et al. 2013; Mabon et al. 2021; Saville et al. 2021; Puidet et al. 2022), and Africa (Njoroge et al. 2019; Beninal et al. 2022). The use of FTA cards will help to eliminate the regulatory hurdle in importing pathogen live cultures from foreign countries and facilitate international collaborations. In addition, the time-consuming and laborious pathogen isolation is not required if genotyping pathogens directly from infected tissues processed on FTA cards performed well (Guichoux et al. 2011; Li et al. 2013). However, pathogen contamination and the amount of pathogen DNA could be issues in this technique. We also used allozyme analyses of (Gpi banding pattern) of representative isolates of P. infestans. Compared to the SSR allelic pattern, there was less variability in the Gpi banding pattern among isolates indicating that allozyme analyses are not sufficient to identify the genotypes of P. infestans.
We identified four known genotypes (US8, US11, US17, and US23) and 25 new genotypes of P. infestans in BC, Pacific western Canada, and frequency distributions of these genotypes varied among years, locations, and hosts (Table 1). The US11 genotype continued to dominate the BC population as we found approximately 60% of the isolates were of this genotype. The US11 genotype was reported in BC between 1994 and 1998 and 2011 and 2012 and in ON from 1996 to 1998 and 2011 (Punja et al. 1998; Peters et al. 1999, 2014; Daayf et al. 2000; Platt et al. 2002; Alkher et al. 2015). In the USA, US11 was reported in Washington in 1994 and 1996–1997, California and New York in 1995 (Goodwin et al. 1998; Dorrance et al. 1999), as well as in Taiwan from 1997 to 2002 (Jyan et al. 2004). Punja et al. (1998) and Peters et al. (1999) reported BC11 with similarities to US11 as the most prevalent genotype in 1995 and 1996 with 91% frequency. BC11 was the only genotype recovered early in the season in BC in 1997 (Punja et al. 1998). More recently, in 2012, US11 with intermediate resistance to metalaxyl, designated as an A1 mating type with 100/100/111 Gpi banding pattern, was reported in BC (Goodwin et al. 1998; Alkher et al. 2015). In our study, only one isolate out of 103 isolates of US11 genotype was sensitive to metalaxyl-m, while 102 isolates were resistant. Notably, US11, an important genotype in BC and some states in the USA was hypothesized as a more fit and competitive sexual recombinant of P. infestans (Goodwin et al. 1998; Fry 2020; Gavino et al. 2000; Peters et al. 2014; Saville and Ristaino 2019). This might be responsible for the displacement of existing strains such as US23 previously reported in BC in 2011 and the continued existence and predominance of US11 in BC. Furthermore, considering the close proximity of the sampled locations in the current study to AB and Washington State, there is a likelihood of transfer of inoculum and possible migration of this genotype through transportation of infected potato seed tubers and tomato transplants from other Canadian provinces and neighboring American states where US11 had been consistently reported (Saville and Ristaino 2019). This is not unexpected as there have been similar cases of the introduction of new genotypes through transportation of infected tubers and tomato transplants within and into Canada (Kawchuk et al. 2011; Kalischuk et al. 2012; Peters et al. 2014) and in the United States (Fry et al. 1992, 2013; Goodwin et al. 1994, 1998; Fry 2020; Gavino et al. 2000).
Interestingly, we reported US17 for the first time in BC and Canada and recovered from both potato and tomato samples across multiple locations during 2019–2021. The US17 genotype was designated as A1 mating type which is highly resistant to metalaxyl (Goodwin et al. 1998). We also found all isolates of US17 were A1 mating type and 87% of isolates (n = 15) were resistant to metalaxyl in our study. The genotype was previously reported in the USA as a probable sexual recombinant between US6 and US8 and was isolated from tomatoes in Alabama, Florida, New Jersey, and New York during 1996 (Goodwin et al. 1998). The low frequency of the US8 genotype (3%) was recovered from two locations of BC in this study exclusively from potato samples in 2019 and 2020. This might indicate the host specificity of the genotype and correlates with those reported by Saville and Ristaino (2019) and Hu et al. (2012) in which host specificity was observed in some genotypes such as US14 specificity to potato and US21 exclusively isolated from tomato samples and loss or gain of avirulence genes was hypothesized for the observed host-specificity (Gilroy et al. 2011; Vleeshouwers et al. 2011). Previously, US8 was reported in BC from Creston in 1997 (Punja et al. 1998) and in New Brunswick (NB) and Prince Edward Island (PEI) from 2009 to 2010 (Kalischuk et al. 2012), and also in PEI and QC in 2011 (Peters et al. 2014). In addition, US8 was reported in neighboring states in the USA (Washington and California) until 2016 (Saville and Ristaino 2019). The US8 was the dominant genotype in Canada in 1996 apart from BC where the US11 dominated (Peters et al. 1998). The presence of US8 genotypes in our study indicated that this genotype has the ability to adapt to the microclimate of the Pacific Northwest. Only a few isolates of the US23 genotype were recovered from tomato and potato samples from eastern Canada in this study between 2021 and 2022. This genotype was previously reported in BC, AB, Saskatchewan (SK), Manitoba (MB), and NB between 2009 and 2012 (Kalischuk et al. 2012; Peters et al. 2014; Alkher et al. 2015). During this period, US23 displaced previously dominating genotypes of P. infestans from most of the provinces of Canada except BC (Peters et al. 2014). US23 was also predominated P. infestans population in Wisconsin between 2009 and 2010 and in Maryland, Virginia, and Pennsylvania in 2009 (Hu et al. 2012; Gevens and Seidl 2013). Although US23 has remained the predominant genotype in eastern Canada in recent years and the genotype was also reported in BC at low frequency in 2009 and 2010 (Alkher et al. 2015), we did not find any isolates of US23 from BC in our comprehensive survey between 2019 and 2022 (n = 204). The continued dominance of US23 in eastern Canada is most likely due to greater infection efficiency, more resilience to overwintering conditions, greater sporulation rates, and greater epidemic potential than other US lineages (Kalischuk et al. 2012; Fall et al. 2015; Seidl Johnson et al. 2015; Saville and Ristaino 2019). However, we do not know why the US23 genotype is absent in BC. The presence of three major genotypes (US11, US8, and US17) and the continued predominance of US11 in BC suggest local adaptation of these genotypes. Microclimate and cultivar type could have played a major role. However, in this study, we did not examine the impact of cultivar types or varieties on P. infestans genetic composition and diversity. In addition, changes to the pathogen population in the sensitivity of the fungicides could impact their fitness and competitive advantages to adapt to the local environment (Saville and Ristaino 2019; Matson et al. 2015).
We found 25 new genotypes (approximately 30% of total isolates from this study) of P. infestans and the majority of them were from tomato samples in community gardens over multiple years. However, new genotypes were also detected from potato samples in commercial potato fields from multiple locations and years. Approximately 94% of isolates of new genotypes (n = 53) were resistant to metalaxyl-m. In agreement with our study, new genotypes (such as BC1-BC11) were reported from BC between 1993 and 1997 (Punja et al. 1998) and from ON, QC, and NB (Peters et al. 2001, 2014; Danies et al. 2014; Alkher et al. 2015). Furthermore, new genotypes of P. infestans were reported from New York, USA (Danies et al. 2014); southern and northern Italy (Saville et al. 2021), and Colombia (Olave-Achury et al. 2022). It is hypothesized that the emergence of these new genotypes in North America could be due to recombination events or mutations accumulated over the years (Punja et al. 1998; Danies et al. 2014; Alkher et al. 2015; Saville and Ristaino 2019), whereas the emergence of the majority of the new isolates in Europe was due to recombination events resulting from sexual reproduction (Anderson et al. 2009; Sujkowski et al. 1994; Drenth et al. 1994; Yuen and Andersson 2013). These new genotypes were often short-lived, localized in a particular region, and only comprised a small percentage of the total samples (Danies et al. 2014; Peters et al. 2014; Alkher et al. 2015; Saville and Ristaino 2019). New genotypes could have better fit and are more competitive (Goodwin et al. 1998; Gavino et al. 2000; Saville and Ristaino 2019). The introduction of genotypes either via migration or evolved naturally in the region could pose new challenges in late blight management due to their genetic composition and phenotypic traits. The severe late bight epidemics in the USA and Canada occurred and caused significant losses for growers and total production abandonment for some tomato growers, which were due to the introduction and distribution of new genotypes from the shipment of tomato transplants to garden centers in retail stores in the northeast United States (Hu et al. 2012; Fry et al. 2013; Alkher et al. 2015; Saville and Ristaino 2019).
PCA, DAPC, Structure, and MSN analyses showed distinct subdivisions and groupings among the large populations of P. infestans in BC (Figs. 1, 2, 3, 4, and 5), which is due to variations in their genetic composition. The small number of isolates from eastern Canada was identified as US23, which formed a distinct cluster. Among BC population, US8 isolated exclusively from potato samples, also formed a distinct cluster. Furthermore, genetic sub-structuring was observed within the US11 isolates from various locations in BC and US8 and CAC12 also clustered away from other BC isolates (AB20P, AG19P, SS19P, and D20P). The structure results and other clustering methods used in this study showed an increase in subclonal variation, corroborated by high genetic diversity indices and the number of MLGs within the US11 lineage (Table 3). Similar results were reported for the US23 and US11 lineages in the USA where considerable subclonal variation and recombination were observed in the lineages and within an aggressive Blue 13 lineage in Indian populations (Dey et al. 2018; Saville and Ristaino 2019). Examination of DAPC, PCA, N-J tree, and structure revealed that the novel genotypes from BC shared some genetic similarities with US11 and US17, whereas US8 and US23 shared little genetic similarities and groupings with US11, US17, and the new genotypes. The new genotype CAE1 from eastern Canada clustered with the US23. This shows the close relatedness of these novel genotypes from BC with some known genotypes (US11, US17, and US23). In agreement with our findings, other studies reported that isolates of US23 shared little genetic similarities with other US genotypes but clustered more with isolates from Bolivia, Brazil, and some parts of Mexico. In addition, our phylogram showed the close relatedness of US23 isolated from this study with those obtained from the USA and Europe (23-A1). Previously, US23 had been reported in the USA and Europe (Gevens and Seidl 2013; Stroud et al. 2016; Kröner et al. 2017; Saville and Ristaino 2019). Likewise, the US11 and US17 lineages showed strong genetic similarity levels and clustered together in the phylogram, PCA, DAPC, and MSNs. These isolates share certain similarities including resistance to metalaxyl; both are A1 mating types, identical RFLP, and similar SSR banding patterns. Similar observations were previously made (Hu et al. 2012; Hansen et al. 2016b; Saville and Ristaino 2019).
We observed high genetic diversity among P. infestans populations in Pacific western Canada as indicated by higher numbers of MLGs and other genetic diversity indices (Table 3). Among genotypes, the US11 genotype had the highest genetic diversity indices, and the 2021 population was the most diverse followed by 2020. Interestingly among locations, Richmond, Agassiz, and Pitt Meadows had higher genetic diversity than other locations. The indices of association results showed that the tomato population was close to a random mating population implying greater potential for recombination, whereas the potato population was away from a random mating population which is a clonal population. In addition, most of the potato isolates were from commercial and research farms and might share common seed sources whereas tomato isolates were mostly from community and home gardens which are from diverse and isolated areas. Other researchers also reported the clonal population of P. infestans (based on values of indices of association) from Denmark and other Nordic European countries (Brurberg et al. 2011; Montes et al. 2016; Maurice et al. 2019) as well as from the USA and South American countries (Saville and Ristaino 2019). Other studies also reported high genetic diversities in P. infestans populations from Canada (Chycoski and Punja 1996; Punja et al. 1998; Kalischuk et al. 2012; Peters et al. 2014; Alkher et al. 2015), USA, Mexico, and other countries (Brurberg et al. 2011; Maurice et al. 2019; Saville and Ristaino 2019; Saville et al. 2021). High genetic diversity in BC could be due to mutations and mitotic recombination or gene conversion (Abu-El Samen et al. 2003; Maurice et al. 2019) because P. infestans populations in Canada still mainly consist of asexual populations. In our study, an extensive sampling was conducted in BC between 2019 to 2021 and sampling sites include several communities and home gardens from diverse locations with distinct microclimates. Since the pathogen isolates were from several community gardens with multiple plots (~ 40 to 200 micro plots), which could have high diversity in the varieties and types of tomato. For instance, Agassiz had samples from both hosts: potato samples from research farms and tomato samples from community gardens, and likewise, Pitt Meadows had samples from huge community gardens consisting of more than two hundred plots. Also, we had P. infestans isolates from infected tomato samples belonging to Roma and Cherry varieties. Therefore, there could be microclimatic variation and diversity in hosts and variety types. Transportation of infected plant materials (seed tubers and tomato transplants) can introduce new genotypes, which could also contribute to genetic diversity (Kalischuk et al. 2012; Peters et al. 2014; Alkher et al. 2015; Saville and Ristaino 2019). High precipitation and humidity characteristic of BC weather might have contributed to increased late blight occurrence and genotypic diversity. These community and home gardens are often managed by non-expert individuals who obtain their seeds or transplants from diverse locations. In some cases, when infected seeds from previous years are planted, the pathogen overwinters and serves as a new source of inocula in the season. This emphasizes the need for planting clean and disease-free seeds, tubers, and transplants. Moreover, these gardens and farms often act as reservoirs for P. infestans populations. These fields are not sprayed with pesticides thus fostering easy pathogen proliferation, and as a result, P. infestans populations in these locations might be the most-fit genotypes and endemic in some cases. Therefore, a culmination of these factors including migration of infected plant materials and favorable environmental conditions might have contributed to the increased number of genotypes found in BC and the emergence of novel genotypes recorded in this study.
Although only A1 mating type was found in our study, occurrence of both mating types was reported previously in BC, ON, QC, NB, and PEI (Chycoski and Punja 1996; Kalischuk et al. 2012; Peters et al. 2014). Both mating types were found in close proximity in ON, QC, and PEI in 2011 (Peters et al. 2014). The presence of both mating types in close proximities could increase the chances of sexual recombination resulting in the production of oospores and the emergence of novel P. infestans genotypes in BC. These sexual spores can survive extreme weather conditions without the host (Medina and Platt 1999; Mayton et al. 2000). Oospores could allow the pathogen to overwinter without the host and serve as a source of inoculum in subsequent seasons, possibly resulting in the emergence of new recombinant genotypes (Goodwin et al. 1995b; Peters et al. 2014). However, there is little or no direct evidence of oospores in fields to date except for a few instances (Peters et al. 2001) and further epidemiological studies are required from different regions in Canada to understand the existence of oospores in the field. Kalischuk et al. (2012) postulated that sexual recombination contributed to increased genotypic characteristics of P. infestans populations in Canada. The emergence of g11 (US 11) might be the result of migration from Mexico (Miller et al. 1997; Peters et al. 2001). Furthermore, the complexity and large genome size of P. infestans detailed by whole genome sequencing could lead to higher gene diversity resulting from random genetic drift and mutations (Abu-El Samen et al. 2003; Haas et al. 2009). In addition, P. infestans populations can switch from triploid to diploid when exposed to chemical and environmental stresses, such as non-lethal doses of the fungicide metalaxyl and extremely cold winters (Li et al. 2017; Maurice et al. 2019). Therefore, exposure to such stresses could favor sexual reproduction because of the resistant nature of oospores in soil. However, some P. infestans populations were reported to self-fertilize (Maurice et al. 2019) and do not require both mating types to produce oospores. This observation indicates that oospores may not only be the result of sexual recombination but also an adaptation to all forms of stress (Li et al. 2017; Maurice et al. 2019).
Overall, the continued dominance of US11 in Pacific western Canada (BC) was found in this study. Our study identified US17 for the first time in Canada and revealed three major existing genotypes (US11, US17, and US8) and 25 new genotypes in BC. Interestingly, the vast majority of the P. infestans isolates (95%) from BC were resistant to metalaxyl-m. We found the sharp dichotomy in late blight incidence and prevalence between eastern and Pacific western Canada. We also found the continued dominance of US23 in eastern Canada, but this genotype was absent in BC. High genetic diversity was observed among P. infestans isolates originating from BC as evidenced by the number of MLGs, genetic diversity indices, and emergence of 25 new genotypes. This suggests that BC is a hotspot for sexual recombination and should be closely monitored for the establishment of a sexual population of P. infestans. Also, the emergence of new genotypes and the continued dominance of US11 in BC could pose management challenges. New recombinants may differ in both phenotypic and genotypic characteristics as compared to existing genotypes. Importation of infected plant materials from South America and Mexico was suggested as the potential source of new genotypes into the USA and consequent migration into Canada from the USA (Kalischuk et al. 2012; Peters et al. 2014; Saville and Ristaino 2019). Global movement of plant materials could have contributed to late blight epidemics; hence, there is a need for improved scrutiny of plant materials movement among regions and countries to avoid the migration of genotypes from one region to another. Considering the number of outbreaks each year, the detection of known genotypes such as US11, US17, and US8 and the emergence of novel genotypes from BC as described in this study and other surveys shows that BC is a hub for the establishment of diverse populations. Therefore, close monitoring of P. infestans populations in this region is useful. Furthermore, more in-depth monitoring of P. infestans populations in Canada, especially BC, is required to determine whether sexually reproducing populations are now established, although the recombinants detected in this study were short-lived and not widely spread. Lastly, the need to understand the sources of recombination events and the presence of an ephemeral sexual population as reported by Danies et al. (2014) needs to be studied extensively. In addition to the mechanisms of genetic diversity, gene flow, recombination, and ploidy analysis need to be holistically investigated and genome sequencing of new genotypes and lineages of P. infestans will be useful.
Data availability
All data generated or analyzed during the study is included in this published article and its supplementary information.
References
Abu-El Samen FM, Secor GA, Gudmestad NC (2003) Genetic variation among asexual progeny of Phytophthora infestans detected with RAPD and AFLP markers. Plant Pathol 52(3):314–325
Agapow PM, Burt A (2001) Indices of multilocus linkage disequilibrium. Mol Ecol Notes 1(1–2):101–102
Alkher H, Islam MR, Wijekoon C, Kalischuk M, Kawchuk LM, Peters RD, Al-Mughrabi KI, Conn KL, Dobinson KF, Daayf F (2015) Characterization of Phytophthora infestans populations in Canada during 2012. Can J Plant Pathol 37(3):305–314
Anonymous (2021a) Crop profile for potato in Canada, 2020. Catalogue No.: A118-10/22-2020E-PDF, 5th ed. Pest Management Program, Agriculture and Agri-Food Canada, Ottawa, ON, Canada. https://publications.gc.ca/collections/collection_2021/aac-aafc/A118-10-22-2020-eng.pdf. Accessed 20 Feb 2023
Anonymous (2021b) Crop profile for greenhouse tomato in Canada, 2020. Catalogue No.: A118-10/24-2020E-PDF, 5th ed. Pest Management Program, Agriculture and Agri-Food Canada, Ottawa, ON, Canada. https://publications.gc.ca/collections/collection_2021/aac-aafc/A118-10-24-2020-eng.pdf. Accessed 20 Feb 2023
Arafa RA, Soliman NE, Moussa OM, Kamel SM, Shirasawa K (2018) Characterization of Egyptian Phytophthora infestans population using simple sequence repeat markers. J Gen Plant Pathol 84:104–107
Arafa RA, Kamel SM, Taher DI, Solberg SØ, Rakha MT (2022) Leaf extracts from resistant wild tomato can be used to control late blight (Phytophthora infestans) in the cultivated tomato. Plants 11(14):1824 https://doi.org/10.3390/plants11141824
Beninal L, Bouznad Z, Corbiere R, Belkhiter S, Mabon R, Taoutaou A, Keddad A, Runno-Paurson E, Andrivon D (2022) Distribution of major clonal lineages EU_13_A2, EU_2_A1, and EU_23_A1 of Phytophthora infestans associated with potato late blight across crop seasons and regions in Algeria. Plant Pathol 71(2):458–469
Birch PR, Bryan G, Fenton B, Gilroy EM, Hein I, Jones JT, Prashar A, Taylor MA, Torrance L, Toth IK (2012) Crops that feed the world 8: potato: are the trends of increased global production sustainable? Food Secur 4:477–508
Brown AH, Feldman MW, Nevo E (1980) Multilocus structure of natural populations of Hordeum spontaneum. Genetics 96(2):523–536
Brurberg MB, Elameen A, Le VH, Nærstad R, Hermansen A, Lehtinen A, Hannukkala A, Nielsen B, Hansen J, Andersson B, Yuen J (2011) Genetic analysis of Phytophthora infestans populations in the Nordic European countries reveals high genetic variability. Fungal Biol 115(4–5):335–342
Burlakoti RR, Hsu CF, Chen JR, Sheu ZM, Bihon W, Kenyon L (2020) Capture of Ralstonia solanacearum species complex strains directly from plant tissue sampled on FTA cards for molecular characterization. J Plant Pathol 102:11–17
Chowdappa P, Brayford D, Smith J, Flood J (2003) Molecular discrimination of Phytophthora isolates on cocoa and their relationship with coconut, black pepper and bell pepper isolates based on rDNA repeat and AFLP fingerprints. Current Sci 10:1235–1238
Chycoski CI, Punja ZK (1996) Characteristics of populations of Phytophthora infestans from potato in British Columbia and other regions of Canada during 1993 to 1995. Plant Dis 80:579–589
Cooke (2020) EuroBlight 12-plex SSR genotyping of Phytophthora infestans. Available at: https://agro.au.dk/fileadmin/euroblight/Protocols/12plex_genotyping_Protocol_Apr_2020.pdf [Accessed July 27, 2022]
Daayf F, Platt HW, Peters RD (2000) Changes in mating types, resistance to metalaxyl, and Gpi-allozyme genotypes of Phytophthora infestans in Canadian provinces from 1996 to1998. Can J Plant Pathol 22:110–116
Danies G, Small IM, Myers K, Childers R, Fry WE (2013) Phenotypic characterization of recent clonal lineages of Phytophthora infestans in the United States. Plant Dis 97(7):873–881
Danies G, Myers K, Mideros MF, Restrepo S, Martin FN, Cooke DE, Smart CD, Ristaino JB, Seaman AJ, Gugino BK, Grünwald NJ (2014) An ephemeral sexual population of Phytophthora infestans in the northeastern United States and Canada. PLoS ONE 9(12):e116354
Deahl KL, Inglis DA (1995) Occurrence of Metalaxyl-insensitive Phytophthora infestans on Solanum sarachoides in Northwestern Washington. Plant Dis 79:540. https://doi.org/10.1094/PD-79-0540A
Deahl KL, Inglis DA, DeMuth SP (1993) Testing for resistance to metalaxyl in Phytophthora infestans isolates from northwestern Washington. Am Potato J 70:779–795
Dey T, Saville A, Myers K, Tewari S, Cooke DE, Tripathy S, Fry WE, Ristaino JB, Guha Roy S (2018) Large sub-clonal variation in Phytophthora infestans from recent severe late blight epidemics in India. Sci Rep 8(1):4429
Dorrance AE, Inglis DA, Derie ML, Brown CR, Goodwin SB, Fry WE, Deahl KL (1999) Characterization of Phytophthora infestans populations in western Washington. Plant Dis 83(5):423–428
Drenth A, Tas IC, Govers F (1994) DNA fingerprinting uncovers a new sexually reproducing population of Phytophthora infestans in the Netherlands. Eur J Plant Pathol 100:97–107
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620
Fall ML, Tremblay DM, Gobeil-Richard M, Couillard J, Rocheleau H, Van der Heyden H, Lévesque CA, Beaulieu C, Carisse O (2015) Infection efficiency of four Phytophthora infestans clonal lineages and DNA-based quantification of sporangia. PLoS ONE 10(8):e0136312
Flier WG, Grünwald NJ, Kroon LP, Sturbaum AK, van den Bosch TB, Garay-Serrano E, Lozoya-Saldaña H, Fry WE, Turkensteen LJ (2003) The population structure of Phytophthora infestans from the Toluca valley of central Mexico suggests genetic differentiation between populations from cultivated potato and wild Solanum spp. Phytopathology 93(4):382–390
Fry WE (2016) Phytophthora infestans: New tools (and old ones) lead to new understanding and precision management. Annu Rev Phytopathol 54:529–547
Fry WE (2020) Phytophthora infestans: the itinerant invader; “late blight”: the persistent disease. Phytoparasitica 48(1):87–94
Fry WE, Goodwin SB, Matuszak JM, Spielman LJ, Milgroom MG, Drenth A (1992) Population genetics and intercontinental migrations of Phytophthora infestans. Annu Rev Phytopathol 30(1):107–130
Fry WE, Goodwin SB, Dyer AT, Matuszak JM, Drenth A, Tooley PW, Sujkowski LS, Koh YJ, Cohen BA, Spielman LJ, Deahl KL, Inglis DA, Sandlan KP (1993) Historical and recent migrations of Phytophthora infestans: chronology, pathways, and implications. Plant Dise 77:653–661
Fry WE, McGrath MT, Seaman A, Zitter TA, McLeod A, Danies G, Small IM, Myers K, Everts K, Gevens AJ, Gugino BK (2013) The 2009 late blight pandemic in the eastern United States–causes and results. Plant Dis 97(3):296–306
Fry W (2013, May) “Biology of Phytophthora infestans and Management of Late Blight” Fry lab, Cornell University, http://www.plantpath.cornell.edu/Fry/Protocols-DNA-extraction.html. [Accessed 9 September 2021]
Gallegly ME, Hong CX (2008) Phytophthora: identifying species by morphology and DNA fingerprints. The APS Press. St. Paul, MN, USA
Gavino PD, Smart CD, Sandrock RW, Miller JS, Hamm PB, Lee TY, Davis RM, Fry WE (2000) Implications of sexual reproduction for Phytophthora infestans in the United States: generation of an aggressive lineage. Plant Dis 84(7):731–735
Gevens AJ, Seidl AC (2013) First report of late blight caused by Phytophthora infestans clonal lineage US-23 on tomato and potato in Wisconsin. United States Plant Dis 97(6):839–839
Gilroy EM, Breen S, Whisson SC, Squires J, Hein I, Kaczmarek M, Turnbull D, Boevink PC, Lokossou A, Cano LM, Morales J (2011) Presence/absence, differential expression and sequence polymorphisms between PiAVR2 and PiAVR2-like in Phytophthora infestans determine virulence on R2 plants. New Phytol 191(3):763–776
Goodwin SB, Drenth A, Fry WE (1992) Cloning and genetic analyses of two highly polymorphic, moderately repetitive nuclear DNAs from Phytophthora infestans. Curr Genet 22:107–115
Goodwin SB, Cohen BA, Deahl KL, Fry WE (1994) Migration from northern Mexico as the probable cause of recent genetic changes in populations of Phytophthora infestans. Phytopathology 84:553–558
Goodwin SB, Schneider RE, Fry WE (1995a) Use of cellulose-acetate electrophoresis for rapid identification of allozyme genotypes of Phytophthora infestans. Plant Dis 79(11):1181–1185
Goodwin SB, Sujkowski LS, Dyer AT, Fry BA, Fry WE (1995b) Direct detection of gene flow and probable sexual reproduction of Phytophthora infestans in northern North America. Phytopathology 85(4):473–479
Goodwin SB, Smart CD, Sandrock RW, Deahl KL, Punja ZK, Fry WE (1998) Genetic change within populations of Phytophthora infestans in the United States and Canada during 1994 to 1996: Role of migration and recombination. Phytopathology 88(9):939–949
Grünwald NJ, Goodwin SB, Milgroom MG, Fry WE (2003) Analysis of genotypic diversity data for populations of microorganisms. Phytopathology 93(6):738–746
Grunwald NJ, Flier WG (2005) The biology of Phytophthora infestans at its center of origin. Annu Rev Phytopathol 43:171–190
Guichoux E, Lagache L, Wagner S, Chaumeil P, Léger P, Lepais O, Lepoittevin C, Malausa T, Revardel E, Salin F, Petit RJ (2011) Current trends in microsatellite genotyping. Mol Ecol Resour 11(4):591–611
Haas BJ, Kamoun S, Zody MC, Jiang RH, Handsaker RE, Cano LM, Grabherr M, Kodira CD, Raffaele S, Torto-Alalibo T, Bozkurt TO (2009) Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans. Nature 461:393–398
Hannukkala AO, Kaukoranta T, Lehtinen A, Rahkonen A (2007) Late-blight epidemics on potato in Finland, 1933–2002; increased and earlier occurrence of epidemics associated with climate change and lack of rotation. Plant Pathol 56(1):167–176
Hansen ZR, Knaus BJ, Tabima JF, Press CM, Judelson HS, Grünwald NJ, Smart CD (2016a) Loop-mediated isothermal amplification for detection of the tomato and potato late blight pathogen. Phytophthora Infestans J Appl Microbiol 120(4):1010–1020
Hansen ZR, Everts KL, Fry WE, Gevens AJ, Grünwald NJ, Gugino BK, Johnson DA, Johnson SB, Judelson HS, Knaus BJ, McGrath MT (2016) Genetic variation within clonal lineages of Phytophthora infestans revealed through genotyping-by-sequencing and implications for late blight epidemiology. Plos One 3;11(11):e0165690 https://doi.org/10.1371/journal.pone.0165690
Haverkort AJ, Boonekamp PM, Hutten R, Jacobsen E, Lotz LA, Kessel GJ, Visser RG, Van der Vossen EA (2008) Societal costs of late blight in potato and prospects of durable resistance through cisgenic modification. Potato Res 51:47–57
Hu CH, Perez FG, Donahoo R, McLeod A, Myers K, Ivors K, Secor G, Roberts PD, Deahl KL, Fry WE, Ristaino JB (2012) Recent genotypes of Phytophthora infestans in the eastern United States reveal clonal populations and reappearance of mefenoxam sensitivity. Plant Dis 96(9):1323–1330
Hurlbert SH (1971) The non-concept of species diversity: a critique and alternative parameters. Ecology 52(4):577–586
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(14):1801–1806
Jombart T (2008) adegenet: an R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405
Jyan MH, Liou RF, Ann PJ, Tsai JN, Hsih SD, Chang TT (2004) Recent occurrence of Phytophthora infestans US-11 as the cause of severe late blight on potato and tomato in Taiwan. Can J Plant Pathol 26(2):188–192
Kalischuk ML, Al-Mughrabi KI, Peters RD, Howard RJ, Platt HW, Kawchuk LM (2012) Genetic composition of Phytophthora infestans in Canada reveals migration and increased diversity. Plant Dis 96(12):1729–1735
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281
Kawchuk LM, Howard RJ, Peters RD, Al-Mughrabi KI (2011) First report of Phytophthora infestans genotype US23 causing late blight in Canada. Plant Dis 95(7):873
Kröner A, Mabon R, Corbière R, Montarry J, Andrivon D (2017) The coexistence of generalist and specialist clonal lineages in natural populations of the Irish Famine pathogen Phytophthora infestans explains local adaptation to potato and tomato. Mol Ecol 26(7):1891–1901
Lees AK, Wattier R, Shaw DS, Sullivan L, Williams NA, Cooke DE (2006) Novel microsatellite markers for the analysis of Phytophthora infestans populations. Plant Pathol 55(3):311–319
Li Y, Cooke DE, Jacobsen E, van der Lee T (2013) Efficient multiplex simple sequence repeat genotyping of the oomycete plant pathogen Phytophthora infestans. J Microbiol Methods 92(3):316–322
Li Y, Shen H, Zhou Q, Qian K, van der Lee T, Huang S (2017) Changing ploidy as a strategy: the Irish potato famine pathogen shifts ploidy in relation to its sexuality. Mol Plant-Microbe Interact 24;30(1):45–52 https://doi.org/10.1094/MPMI-08-16-0156-R
Llorente B, Bravo-Almonacid F, Cvitanich C, Orlowska E, Torres HN, Flawia MM, Alonso GD (2010) A quantitative real-time PCR method for in planta monitoring of Phytophthora infestans growth. Lett Appl Microbiol 51(6):603–610
Mabon R, Guibert M, Corbière R, Andrivon D (2021) An improved PCR method for rapid and accurate identification of mating types in the late blight pathogen Phytophthora infestans. Plant Health Prog 22(3):362–367
Matson ME, Small IM, Fry WE, Judelson HS (2015) Metalaxyl resistance in Phytophthora infestans: Assessing role of RPA190 gene and diversity within clonal lineages. Phytopathology 105(12):1594–1600
Maurice S, Montes MS, Nielsen BJ, Bødker L, Martin MD, Jønck CG, Kjøller R, Rosendahl S (2019) Population genomics of an outbreak of the potato late blight pathogen, Phytophthora infestans, reveals both clonality and high genotypic diversity. Mol Plant Pathology 20(8):1134–1146
Mayton H, Smart CD, Moravec BC, Mizubuti ES, Muldoon AE, Fry WE (2000) Oospore survival and pathogenicity of single oospore recombinant progeny from a cross involving US-17 and US-8 genotypes of Phytophthora infestans. Plant Dis 84(11):1190–1196
Medina MV, Platt HW (1999) Viability of oospores of Phytophthora infestans under field conditions in northeastern North America. Can J Plant Pathol 21(2):137–143
Miller JS, Hamm PB, Johnson DA (1997) Characterization of the Phytophthora infestans population in the Columbia Basin of Oregon and Washington from 1992 to 1995. Phytopathology 87(6):656–660
Montes MS, Nielsen BJ, Schmidt SG, Bødker L, Kjøller R, Rosendahl S (2016) Population genetics of Phytophthora infestans in Denmark reveals dominantly clonal populations and specific alleles linked to metalaxyl-M resistance. Plant Pathol 65(5):744–753
Njoroge AW, Andersson B, Lees AK, Mutai C, Forbes GA, Yuen JE, Pelle R (2019) Genotyping of Phytophthora infestans in Eastern Africa reveals a dominating invasive European lineage. Phytopathology 109(4):670–680
Olave-Achury A, Cardenas D, Restrepo S, Lucca F, Fry WE, Myers KL, Danies G, Soto-Suarez M (2022) Phenotypic and genotypic characterization of Phytophthora infestans isolates associated with tomato and potato crops in Colombia. Phytopathology 112(8):1783–1794
Peters RD, Platt HW, Hall R (1998) Characterization of changes in populations of Phytophthora infestans in Canada using mating type and metalaxyl sensitivity markers. Can J Plant Pathol 20(3):259–273
Peters RD, Platt HW, Hall R (1999) Use of allozyme markers to determine genotypes of Phytophthora infestans in Canada. Can J Plant Pathol 21(2):144–153
Peters RD, Förster H, Platt HW, Hall R, Coffey MD (2001) Novel genotypes of Phytophthora infestans in Canada during 1994 and 1995. Am J Potato Res 78:39–45
Peters RD, Al-Mughrabi KI, Kalischuk ML, Dobinson KF, Conn KL, Alkher H, Islam MR, Daayf F, Lynn J, Bizimungu B, De Koeyer D (2014) Characterization of Phytophthora infestans population diversity in Canada reveals increased migration and genotype recombination. Can J Plant Pathol 36(1):73–82
Platt HW, Daayf F, MacPhail A (2002) Cross-Canada potato late blight survey in 2000. Can Plant Dis Surv 82:118–120
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Puidet B, Mabon R, Guibert M, Kiiker R, Soonvald L, Le VH, Eikemo H, Dewaegeneire P, Saubeau G, Chatot C, Aurousseau F (2022) Examining phenotypic traits contributing to the spread in northern European potato crops of EU_41_A2, a new clonal lineage of Phytophthora infestans. Phytopathology 112(2):414–421
Punja ZK, Förster H, Cunningham I, Coffey MD (1998) Genotypes of the late blight pathogen (Phytophthora infestans) in British Columbia and other regions of Canada during 1993–1997. Can J Plant Pathol 20(3):274–282
Rekad FZ, Cooke DE, Puglisi I, Randall E, Guenaoui Y, Bouznad Z, Evoli M, Pane A, Schena L, di San Lio GM, Cacciola SO (2017) Characterization of Phytophthora infestans populations in northwestern Algeria during 2008–2014. Fungal Biol 121(5):467–477
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4(1):137–138
Sapkota S, Burlakoti RR, Punja ZK (2023) Diversity in virulence and metalaxyl-m sensitivity of Phytophthora rubi isolates has implications for raspberry root rot and wilting complex management. Can J Plant Pathol. https://doi.org/10.1080/07060661.2023.2175912
Saville A, Ristaino JB (2019) Genetic structure and subclonal variation of extant and recent US lineages of Phytophthora infestans. Phytopathology 109(9):1614–1627
Saville AC, Ristaino JB (2021) Global historic pandemics caused by the FAM-1 genotype of Phytophthora infestans on six continents. Sci Rep 11(1):1–11
Saville AC, Martin MD, Ristaino JB (2016) Historic late blight outbreaks caused by a widespread dominant lineage of Phytophthora infestans (Mont.) de Bary. PLoS ONE 11(12):e0168381
Saville AC, La Spada F, Faedda R, Migheli Q, Scanu B, Ermacora P, Gilardi G, Fedele G, Rossi V, Lenzi N, Testa A (2021) Population structure of Phytophthora infestans collected on potato and tomato in Italy. Plant Pathol 70(9):2165–2178
Seidl Johnson AC, Frost KE, Rouse DI, Gevens AJ (2015) Effect of temperature on growth and sporulation of US-22, US-23, and US-24 clonal lineages of Phytophthora infestans and implications for late blight epidemiology. Phytopathol 8;105(4):449–459 https://doi.org/10.1094/PHYTO-03-14-0064-R
Shannon CE (2001) A mathematical theory of communication. ACM SIGMOBILE Mob Comput Commun Rev 5(1):3–55
Shimelash D, Dessie B (2020) Novel characteristics of Phytophthora infestans causing late blight on potato in Ethiopia. Curr Plant Biol 24:100172
Soylu EM, Soylu S, Kurt S (2006) Antimicrobial activities of the essential oils of various plants against tomato late blight disease agent Phytophthora infestans. Mycopathologia 161:119–128
Stoddart JA, Taylor JF (1988) Genotypic diversity: estimation and prediction in samples. Genetics 118(4):705–711
Stroud JA, Shaw DS, Hale MD, Steele KA (2016) SSR assessment of Phytophthora infestans populations on tomato and potato in British gardens demonstrates high diversity but no evidence for host specialization. Plant Pathol 65(2):334–341
Sujkowski LS, Goodwin SB, Dyer AT, Fry WE (1994) Increased genotypic diversity via migration and possible occurrence of sexual reproduction of Phytophthora infestans in Poland. Phytopathology 84(2):201–207
Vleeshouwers VG, Raffaele S, Vossen JH, Champouret N, Oliva R, Segretin ME, Rietman H, Cano LM, Lokossou A, Kessel G, Pel MA (2011) Understanding and exploiting late blight resistance in the age of effectors. Annu Rev Phytopathol 49:507–531
Wang J, Fernández-Pavía SP, Larsen MM, Garay-Serrano E, Gregorio-Cipriano R, Rodríguez-Alvarado G, Grünwald NJ, Goss EM (2017) High levels of diversity and population structure in the potato late blight pathogen at the Mexico centre of origin. Mol Ecol 26(4):1091–1107
Widmark AK, Andersson B, Sandström M, Yuen JE (2011) Tracking Phytophthora infestans with SSR markers within and between seasons–a field study in Sweden. Plant Pathol 60(5):938–945
Yuen JE, Andersson B (2013) What is the evidence for sexual reproduction of Phytophthora infestans in Europe? Plant Pathol 62(3):485–491
Acknowledgements
We acknowledge the Canada/New Brunswick Canadian Agricultural Partnership Program and the Canadian Agri-Science Cluster for Horticulture 3 (2810-ASC-18/19 Hort Cluster 13A) for funding this work. We would also like to thank Anne MacPhail and Taysia Nikaido-Landry for their technical assistance and all the growers and industry representatives who submitted diseased samples for this study.
Funding
Open Access funding provided by Agriculture & Agri-Food Canada. This research is supported by the Canada/New Brunswick Canadian Agricultural Partnership Program and the Canadian Agri-Science Cluster for Horticulture 3 (2810-ASC-18/19 Hort Cluster 13A).
Author information
Authors and Affiliations
Contributions
SB conducted experiments, analyzed data, contributed to new methods, wrote the paper, and reviewed the paper. RRB conceived, conceptualized, designed, and supervised research and data analyses and wrote and reviewed the paper. RP conceived, designed, and supervised research and reviewed the paper. AN conducted experiment and reviewed the paper. SS analyzed data and reviewed paper. KA designed and supervised research and reviewed the paper. BP supervised and reviewed the paper. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Babarinde, S., Burlakoti, R.R., Peters, R.D. et al. Genetic structure and population diversity of Phytophthora infestans strains in Pacific western Canada. Appl Microbiol Biotechnol 108, 237 (2024). https://doi.org/10.1007/s00253-024-13040-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13040-6