Abstract
Functional M cells are differentiated by receptor activator of NF-κB ligand (RANKL) and capture of luminal antigens to initiate immune responses. We aimed to use postbiotic-based recombinant chicken RANKL (cRANKL) to promote M cell differentiation and test the efficacy of oral vaccines. Chicks were divided into three groups that were administered phosphate-buffered saline (PBS), cell extracts of wild-type Lactococcus lactis subsp. lactis IL1403 (WT_CE), or cell extracts of recombinant L. lactis expressing cRANKL (cRANKL_CE). The expression of the M cell marker was measured, and the gut microbiome was profiled. The efficiency of the infectious bursal disease (IBD) vaccine was tested after 12 consecutive days of administering cRANKL_CE. The chickens that were administered cRANKL_CE (p = 0.038) had significantly higher Annexin A5 (ANXA5) mRNA expression levels than those in the PBS group (PBS vs. WT_CE, p = 0.657). In the gut microbiome analysis, no significant changes were observed. However, the relative abundance of Escherichia-Shigella was negatively correlated (r = − 0.43, p = 0.019) with ANXA5 mRNA expression in Peyer’s patches. cRANKL_CE/IBD (p = 0.018) had significantly higher IBD-specific faecal IgA levels than PBS/IBD (PBS/IBD vs. WT_CE/IBD, p = 0.217). Postbiotic-based recombinant cRANKL effectively improved the expression of M cell markers and the efficiency of oral vaccines. No significant changes were observed in the gut microbiome after administration of postbiotic-based recombinant cRANKL. This strategy can be used for the development of feed additives and adjuvants.
Key points
• Postbiotic-based recombinant cRANKL enhanced the expression of ANXA5 in chicken.
• The relative abundance of Escherichia-Shigella was negatively correlated with ANXA5 expression.
• Postbiotic-based recombinant cRANKL effectively improved the efficiency of oral vaccine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus defines postbiotics as “preparations of inanimate microorganisms and/or their components that confer a health benefit on the host”. It includes components such as cell wall fragments, metabolites, and other microbial derivatives (Salminen et al. 2021). Postbiotics contain a wide range of molecules, including peptidoglycans, surface proteins, cell wall polysaccharides, secreted proteins, bacteriocins, and organic acids, which have positive effects on the host, such as anti-inflammatory, immunomodulatory, antitumor, antimicrobial, and antioxidant effects (Nataraj et al. 2020; Teame et al. 2020). However, most studies have used wild-type postbiotics, which exhibit limitations with respect to the regulation of specific pathways.
Microfold (M) cells are specialised immune cells located in the gut-associated lymphoid tissue (GALT) and play an essential role in the initiation of the intestinal immune response that transports luminal antigens via the intestine toward GALT (Foussat et al. 2001). Receptor activator of NF-κB ligand (RANKL) can induce differentiation of functional M cells (Kunisawa et al. 2008; Knoop et al. 2009; Kanaya et al. 2012). Knoop et al. showed that purified recombinant mouse RANKL produced by recombinant E. coli differentiated M cells in the small intestine of RANKL-null mice (Knoop et al. 2009). In addition, Kim et al. showed that oral administration of recombinant L. lactis-secreting mouse RANKL significantly increased the number of mature M cells in the mouse small intestine (Kim et al. 2015). These studies demonstrate that recombinant RANKL produced by bacteria exhibits bioactivity. Moreover, owing to the characteristics of M cells, the promotion of M cell differentiation may be a viable strategy to increase the efficiency of oral vaccines (Yamamoto et al. 2012). Therefore, the use of postbiotic-based recombinant RANKL not only has the potential to increase the activity of M cells but also may demonstrate a synergistic effect of recombinant protein and postbiotics. Moreover, postbiotics influence the changes in the gut microbiome (Ozma et al. 2022). Therefore, it is necessary to determine the influence of postbiotic-based recombinant proteins on the gut microbiome.
In this study, the bioactivity of the postbiotic-based recombinant cRANKL was tested by measuring the expression of M cell markers in chickens, and changes in the intestinal microbial community were profiled. Therefore, we investigated the possibility of using postbiotic-based recombinant cRANKL as an adjuvant or additive.
Materials and methods
Microorganism strains and growth conditions
Wild-type Lactococcus lactis subsp. lactis IL1403 (Simon and Chopin 1988, strain collected from Seoul National University) was used as the host bacterium for target protein expression. Wild-type and recombinant strains were grown in M17 medium (MBcell, Korea) supplemented with 5 g/L of glucose (M17G) without antibiotics or with erythromycin (5 μg/mL) and chloramphenicol (5 μg/mL) at 30 °C, respectively.
Gene synthesis and plasmid construction
Two cRANKL amino acid (aa) sequences were obtained from the National Center for Biotechnology Information (NCBI): 400 aa (NCBI accession number: NP_001076830.1) and 318 aa (NCBI accession number: CDZ92724.1). These two proteins had the same extracellular domain sequences. Based on the results of alignment with the soluble form of mouse RANKL (mRANKL) (Knoop et al. 2009; Kim et al. 2015), the one of candidate gene fragments was selected, and the extracellular form of cRANKL was obtained from Sutton et al. (2015). To secrete the target protein from recombinant L. lactis, the signal peptide of USP45 (van Asseldonk et al. 1993) was added to the N-terminus of the target gene. The His-tag (His6x) was added to the C-terminus of the target gene to detect the target proteins. The restriction sites of NdeI and XhoI were located at the two ends for insertion and vector ligation. The vector construction is shown in Fig. 1A. The designed amino acid sequences were codon-optimised using DNAWorks v3.2.4 (Hoover and Lubkowski 2002) based on the L. lactis IL1403 codon usage table, and primers were used to synthesise the insert sequences using the overlap PCR method. The plasmid DNA pILPtuf.Mb (Kim et al. 2009) vector was used as the backbone. The insert and vector were ligated at NdeI and XhoI restriction sites and transformed into wild-type L. lactis IL1403 competent cells. All insert sequences are shown in Fig. S1.
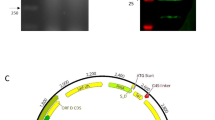
Production and secretion of recombinant cRANKL from recombinant L. lactis. A Schematic diagram for construction of pILPtuf.gExR.h vector. B Western blot for detecting recombinant cRANKL from cell extracts and cell-free culture supernatant. Lane 1, cell extracts of L. lactis IL1403; Lane 2, cell extracts of L. lactis IL1403 (pILPtuf.gExR.h); Lane 3, cell-free culture supernatant of L. lactis IL1403 (pILPtuf.gExR.h); Lanes 4–6: commercial His-tagged calmodulin (18 kDa) 1.5, 1, and 0.5 μg, respectively
SDS-PAGE and western blot assay
Wild-type and recombinant L. lactis were cultured in 10 mL of M17G medium without antibiotics or with antibiotics at 30 °C for 10 h, respectively. For the preparation of cell extracts, 10 mL of cultured cells was harvested by centrifugation 3,134 × g for 10 min at 4 °C, and cell pellets were washed by sterilised distilled water twice and resuspended in 200 µL of sterilised 1 × PBS. Subsequently, the solution containing the cell pellet was supplemented with 0.5 g of sterilised glass beads (0.5 mm), followed by disruption using a taco Prep Bead Beater (GeneReach, Taiwan) for 39 s. Cell debris was removed from the cell extracts by centrifugation at 12,300 × g for 5 min at 4 °C. For the preparation of secreted proteins, after centrifugation at 3134 × g for 10 min at 4 °C, the culture supernatant was filtered using the 0.2-µm filter (Minisart® Syringe Filter, Sartorius) and then precipitated with trichloroacetic acid (TCA, 4:1) at 4 °C for 1 h. The precipitates were washed twice with 1 mL of pre-chilled acetone, and centrifugation was performed at 12,300 × g for 5 min at 4 °C. The precipitated pellets were dried at 65 °C for 10 min and dissolved in 200 µL of sterilised 1 × PBS. To quantify the amount of target protein produced, commercial recombinant His-tagged human calmodulin (MERCK, Darmstadt, Germany) protein (18 kDa) was used for the construction of the standard curve with concentrations of 0.5, 1, and 1.5 µg.
The proteins from the total cell extracts or cell-free culture supernatant were separated by SDS-PAGE and transferred onto a nitrocellulose membrane (2 μm; Bio-Rad, Germany). The membrane was blocked with 2.5% (w/v) skim milk in 1 × tris-buffered saline (TBST, 0.1% Tween® 20 detergent) at 25 °C for 1 h. After blocking, the membrane was washed with 1 × TBST thrice for 10 min per wash and incubated with anti-His6x monoclonal antibody (1:500, R&D Systems, USA) at 4 °C for 12 h with shaking. After washing thrice with 1 × TBST, the membrane was visualised with ECL reagents (Bio-Rad, USA).
Validation of the bioactivity of recombinant cRANKL in vivo
One-day-old male ROSS 308 chicks were randomly assigned to three groups, namely PBS, WT_CE, and cRANKL_CE, with ten chicks in each group. The chicks in the PBS group were administered sterilised 1 × PBS. WT_CE and cRANKL_CE groups were administered cell extracts from wild-type L. lactis IL1403 and the recombinant strain L. lactis IL1403 (pILPtuf.gExR.h), respectively. For the preparation of cell extracts, 12-h-cultured wild-type or recombinant strain was harvested by centrifugation 3134 × g for 10 min at 4 °C, and cell pellets were washed by sterilised distilled water twice and resuspended in sterilised 1 × PBS. Subsequently, the solution containing the cell pellet was supplemented with 0.5 g of sterilised glass beads (0.5 mm), followed by disruption using a taco Prep Bead Beater (GeneReach, Taiwan) for 39 s. Glass beads were removed from the cell extracts by centrifugation at 12,300 × g for 5 min at 4 °C. Then, the extracted cell extracts were inoculated in M17G medium and found to contain no viable bacteria. The prepared cell extracts were administered orally to each chick using a 1 mL syringe without a needle. The dosage of cell extracts is shown in Table 1. All groups were fed for 12 consecutive days and sampled on the 13th day (Table 1, Fig. 2).
Peyer’s patches of ileum samples were extracted to determine the expression level of the M cell marker by measuring ANXA5 mRNA expression. Total RNA was extracted using TRIzol® (Thermo Fisher, Korea) according to the manufacturer’s instructions, and cDNA was synthesised using the PrimeScriptTM RT reagent Kit (Takara Bio, Japan). Quantitative real-time PCR (qRT-PCR) was conducted using TB Green® Premix Ex Taq™ (Tli RNaseH Plus, TAKARA, Japan) with specific primers ANXA5-F:5′-AGTATACAAGAGGCACCGTG-3′; ANXA5-R:5′-GTCTCATCAAAGATACCATC-3′, and primers for the housekeeping gene GAPDH-F:5′-GTGGTGCTAAGCGTGTTATCATC-3′; GAPDH-R:5′-GGCAGCACCTCTGCCATC-3′ (Olias et al. 2014). The mRNA level was presented as 2−ΔCt, where Ct, threshold cycle for target amplification, and ΔCt, Cttaget gene (specific genes for each sample) − Ctinternal reference (housekeeping gene for each sample).
16S rRNA amplicon sequencing
On the 13th day, genomic DNA was extracted from faecal samples using a NucleoSpin Soil kit (Macherey–Nagel, Düren, Germany), according to the manufacturer’s instructions. DNA samples (5 ng) were used to amplify the 16S ribosomal RNA V4 region using Takara Ex-Taq DNA polymerase (Takara Bio) with universal primer sets (forward:5′-GGACTACHVGGGTWTCTAAT-3′ and reverse:5′-GTGCCAGCMGCCGCGGTAA-3′) (Han et al. 2018). After amplification, all samples were normalised to 50 ng per sample. A DNA library was constructed and sequenced using the Illumina MiSeq platform (Illumina, San Diego, CA, USA), generating 2 × 300 bp paired-end reads.
Bioinformatic analysis of the gut microbiome
To analyse the microbial community, de-multiplexed and pre-processed sequence reads were imported into Quantitative Insights Into Microbial Ecology (QIIME2, version 2021.2) (Bolyen et al. 2019). Barcode and primer removal, quality control, amplicon sequence data correction, and de-replication were performed using the DADA2 software package (Callahan et al. 2016). Sequence reads were truncated to 200 bp using an in-house Perl script. Feature tables and representative sequence files were merged for downstream analysis using QIIME2. Taxonomic classification was assigned using the SILVA 132 database, with 99% identity based on the V4 16S region. All classifications were performed within QIIME2 and were assigned using the naïve Bayesian algorithm available in the Sklearn Python library. For phylogenetic diversity analysis, alpha and beta diversities were calculated using the q2-diversity plugin and included Faith’s phylogenetic diversity and weighted and unweighted UniFrac distances. Differential abundance analysis of microbiota was performed using an in-house Perl script. The relationship between relative abundance and ANXA5 mRNA expression was assessed by Pearson’s correlation efficient (r) and p values from simple linear regression. Statistical significance was set at p < 0.05.
Serum calcium assay
The serum calcium concentration was measured using a Calcium Colorimetric Assay Kit (BioVision Inc., USA). All serum samples were diluted to ten-folds (10 μL of serum and 90 μL of Chromogenic Reagent), and 60 μL of Calcium Assay Buffer was added to each well and mixed gently. The samples were incubated for 10 min at room temperature in the dark. After incubation, the optical density (OD) was analysed by measuring the absorbance of the samples at 575 nm.
Evaluation of oral vaccine efficiency
After 12 consecutive days of oral administration of postbiotic-based recombinant cRANKL, chickens were orally immunised with the infectious bursal disease (IBD) vaccine on the 13th day. The experimental schedule is shown in Fig. 2. The IBD vaccine (PoulShot® Gumboro) was purchased from the Central Vaccine Research Institute, Korea. The dosage of oral vaccination was based on vaccine programme guidelines. The relationship between faecal IgA and serum IgG was assessed by Pearson’s correlation efficient (r) and p values from simple linear regression. Statistical significance was set at p < 0.05.
Statistical analysis
Statistical analysis was performed using an in-house Perl script and R (v4.1.4). For significance analysis, the Kruskal–Wallis test was performed followed by Dunn’s posthoc test, and the data are expressed as follows: *p < 0.05, **p < 0.01, ***p < 0.001.
Results
Construction of recombinant strains
Although the correct insert sequences were cloned into L. lactis, some transformants did not form colonies, or mutations were found after sequencing (Figure S2). To confirm the expression of cRANKL, intracellular and secreted proteins were precipitated and analysed by western blotting. cRANKL was not detected in cell extracts of the wild-type strain. In the recombinant strain, only intracellular cRANKL was detected, with an expected size of 31.74 kDa (Fig. 1B). According to the standard curve, intracellular production of cRANKL was 2.2 μg/mL.
Validation of bioactivity of postbiotic-based recombinant cRANKL in vivo
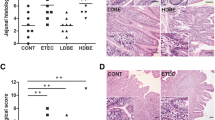
To validate the bioactivity of postbiotic-based recombinant RANKL from L. lactis cell extracts in ROSS 308 chickens, RT-PCR was performed to detect ANXA5-high M cells from the Peyer’s patches of chickens. As shown in Fig. 3, compared with the PBS group, the cRANKL_CE (p = 0.038) group showed significantly higher ANXA5 mRNA expression levels, and no significant difference was observed between the PBS and WT_CE groups (p = 0.657). Despite the absence of significant differences between the WT_CE and cRANKL_CE groups, the cRANKL_CE group had higher ANXA5 expression levels than the WT_CE group (p = 0.104). These results indicate that cRANKL from cell extracts of recombinant L. lactis affected ANXA5 expression in M cells. No significant differences were observed in body weight gain between the groups (Figure S3).
Gut microbial diversity
To compare the gut microbial diversity and communities, the alpha and beta diversities of the PBS, WT_CE, and cRANKL_CE groups were investigated from normalised microbiome sequencing reads. For alpha diversity, four indices that observed features, Evenness, Shannon index, and Faith’s phylogenetic diversity (Faith PD), were measured. No significant difference was observed in the observed features and Faith PD. However, the Evenness and Shannon indices of cRANKL_CE were significantly higher than those of the WT_CE group (Figure S4). For beta diversity, principal coordinate analysis (PCoA) of the unweighted and weighted UniFrac distances was performed. No significant differences were observed among the three groups (Figure S5).
Gut microbial composition and its correlation with the expression of ANXA5
To compare the differences in the major gut microbial taxa between the PBS, WT_CE, and cRANKL_CE groups, the microbial composition in these three groups was examined. The overall microbial composition in the gut was not significantly different among the three groups (Tables 1 and S1). However, at the phylum level, Firmicutes in the cRANKL_CE group were more abundant than those in the PBS (p = 0.058) and WT_CE groups (p = 0.244). At the genus level, Lactobacillus, Terrisporobacter, Enterococcus, Turicibacter, Staphylococcus, Tuzzerella, and Pseudomonas were significantly different. Compared to the WT_CE group, the abundance of Escherichia-Shigella decreased in the cRANKL_CE group (p = 0.062). Compared to the PBS group, the abundance of Lysinibacillus decreased in both the WT_CE (p = 0.070) and cRANKL_CE (p = 0.163) groups. Moreover, no significant differences were observed among the three groups with respect to the genus Lactococcus (Table 1). In correlation analysis, the relative abundance of Escherichia-Shigella was negatively correlated (r = − 0.43, p = 0.019) with host ANXA5 mRNA expression in Peyer’s patches (Fig. 5).
Serum calcium assay
To detect changes in serum calcium concentration attributed to bone metabolism disorder after the administration of recombinant cRANKL, serum calcium levels were measured. The results showed no significant difference in serum calcium levels among the three groups (Figure S6).
Evaluation of the effect of postbiotic-based recombinant cRANKL on adjuvanticity of oral vaccination
To investigate whether postbiotic-based recombinant cRANKL from cell extracts enhanced IBD-specific systemic and mucosal immune responses, anti-IBD serum IgG and faecal IgA levels were measured. As shown in Fig. 4 A and B, 2 weeks after immunisation, although no significant differences were observed, the levels of serum IgG in the WT_CE/IBD (p = 0.139) and cRANKL_CE/IBD (p = 0.073) groups were higher than those in the PBS/IBD group, and no significant difference was observed between the WT_CE/IBD and cRANKL_CE/IBD groups (p = 0.844). This result demonstrated that cell extracts of wild-type L. lactis IL1403 could play a role as an adjuvant to enhance systemic immunity in chickens. The faecal IgA level of the cRANKL_CE/IBD group was significantly higher than those in the PBS/IBD group (p = 0.018). A positive correlation was observed between serum IgG and faecal IgA production in each individual (Fig. 4C). This result demonstrated that cell extracts with cRANKL could play a role as an adjuvant to enhance mucosal immunity in chickens.
Discussion
Many wild-type postbiotics have been used for improving animal health owing to their effects, such as resistance to pathogens, benefits to gut barrier function, and immunomodulatory effects on the gut (Zhong et al. 2022). However, no recombinant proteins in their postbiotic form have been used in chickens.
Owing to the benefits of lactic acid bacteria (LAB) to animal health, LAB as probiotics are applicable as feed additives (Vieco-Saiz et al. 2019). In the past two decades, the application of L. lactis has been expanded from food additives to employment as successful microbial cell factories (Song et al. 2017). Kim et al. showed that recombinant mouse RANKL (mRANKL) secreted from recombinant L. lactis significantly increased the abundance of mature M cells in the mouse small intestine and improved the efficiency of oral vaccines (Kim et al. 2015). However, in this study, recombinant cRANKL was fed as a postbiotic to enhance the differentiation of M cells in the chicken small intestine. In a previous experiment, we successfully fed postbiotic-based recombinant mRANKL to mice to increase mature M cells (Xuan et al. 2022a). As non-existent organisms in the natural environment, once resealed into nature, living-modified organisms (LMOs) lead to ecological environment disorders (Prakash et al. 2011). Therefore, postbiotics can be safely used in animals. In addition, there is an advantage in that a recombinant protein purification process is not required.
In the process of producing recombinant bacteria, several strains were found to have mutations, and the transformants did not form colonies. Although the correct sequence was cloned into L. lactis, the gsR.h sequence appeared to be deleted or substituted, and the NS.gExR.h sequence appeared to be inserted (Figure S2). In some cases, after the translation of foreign proteins in recombinant bacterial cells, if the protein demonstrates “toxic effects” in the host, such as disruption of the normal metabolism and inhibition of cell proliferation, then bacteria will die or reduce their growth or even induce mutations to protect themselves.
With the increase in postbiotic research, some studies have confirmed that postbiotics affect the gut microbiome (Nataraj et al. 2020; Ozma et al. 2022). Our previous study showed that cell extracts from wild-type or recombinant L. lactis did not critically change the mouse gut microbiome; however, in the group that was administered mRANKL contained cell extracts, the composition of the gut microbiome was similar to that of patients with rheumatoid arthritis (Xuan et al. 2022a). In this study, cell extracts derived from wild-type or recombinant strains did not significantly change the gut microbiota of chickens, and only a few genera were significantly different (Table 2). The abundance of Lactobacillus in the WT_CE group was higher than in the PBS and cRANKL_CE groups. In mice, cell extracts of wild-type did not affect Lactobacillus abundance; however, in the chicken, the abundance of Lactobacillus was significantly increased. Unlike in mice, the abundance of Lactobacillus was reduced in the group fed RANKL (Xuan et al. 2022a). This may be attributed to species-specific differences. Other reasons for the decrease in Lactobacillus abundance induced by cRANKL require further investigation. No significant difference was observed between groups with respect to Escherichia-Shigella abundance; however, the lowest abundance was found in the group fed cRANKL_CE (Table 2). ANXA5 expressed by M cells can bind to LPS of gram-negative bacteria and block endotoxin activity, indicating that ANXA5 on M cells acts as an uptake receptor for gram-negative bacteria in mice (Rand et al. 2012). The expression of ANXA5 mRNA in the small intestine was negatively correlated with the abundance of Escherichia-Shigella (Fig. 5). M cells are considered to be the point of intestinal entry for antigens and invasive pathogens, such as Shigella, Salmonella, and Yersinia (McI Mowat et al. 2004). The abundance of Lysinibacillus decreased in the postbiotic groups of WT_CE and cRANKL_CE. Klaudia et al. showed that chickens infected with avian influenza had more Lysinibacillus in the intestine than uninfected chickens (Chrzastek et al. 2021). However, the mechanism underlying the relationship between Lysinibacillus and avian influenza remains unclear. In addition, the relationship between Lysinibacillus and immunity remains unclear. Similar to a previous study, no significant differences were observed with respect to Lactococcus among the three groups. This indicates that cell extracts of L. lactis did not elicit an immune response in the intestine (Xuan et al. 2022a, 2022b). If other strains of postbiotics were used, different changes may have been observed in the gut microbiome. In addition, no significant differences were observed in serum calcium levels among the three groups (Xuan et al. 2022a). This may indicate that recombinant cRANKL is insufficient to affect calcium metabolism in chickens.
IBD, also known as Gumboro disease, is an immunosuppressive and highly infectious disease caused by the IBD virus (IBDV), affecting young birds and causing significant economic losses in the poultry industry worldwide (Gómez et al. 2018). Birds aged 1–14 days are less susceptible to infection and are usually protected by maternal antibodies (Mahgoub 2012). In this study, after feeding cell extracts for 12 days, chickens were immunised with an attenuated IBDV oral vaccine to test whether wild-type postbiotic or postbiotic-based recombinant cRANKL could be used as an immune adjuvant. Although no significant differences were observed, our results showed that the WT_CE/IBD and cRANKL_CE/IBD groups had higher IBD-specific serum IgG levels than the PBS/IBD group. This result indicated that cell extracts from wild-type L. lactis IL1403 may interfere with IgG production by enhancing intestinal immunity. The levels of IBD-specific faecal IgA were significantly higher in the cRANKL_CE/IBD group than those in the PBS/IBD group. This suggests that M cell development by cRANKL may increase the antigen transcytosis of FAE, thereby enhancing the mucosal immune response. In a previous study, when mice were fed wild-type L. lactis IL1403-derived cell extracts, no significant changes were observed in the small intestinal transcriptome; however, when cell extracts containing mRANKL were fed, RANKL-related genes were up- or downregulated (Xuan et al. 2022a). More indicators need to be measured to determine how wild-type cell extracts or the combination of recombinant proteins and cell extracts affect chickens.
In conclusion, we found that postbiotic-based recombinant cRANKL effectively improved M cell differentiation and the efficiency of oral vaccines. In addition, feeding postbiotic-based recombinant cRANKL did not significantly alter the gut microbiome of chickens. This strategy may be useful for the development of other functional proteins and feed additives.
Data availability
Microbiome raw sequences were deposited in GenBank with the Accession Number PRJNA934978.
References
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Bin KK, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu YX, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Chrzastek K, Leng J, Zakaria MK, Bialy D, La Ragione R, Shelton H (2021) Low pathogenic avian influenza virus infection retards colon microbiota diversification in two different chicken lines. Anim Microbiome 3. https://doi.org/10.1186/s42523-021-00128-x
Foussat A, Balabanian K, Amara A, Bouchet-Delbos L, Durand-Gasselin I, Baleux F, Couderc J, Galanaud P, Emilie D (2001) Production of stromal cell-derived factor 1 by mesothelial cells and effects of this chemokine on peritoneal B lymphocytes. Eur J Immunol 31:350–359. https://doi.org/10.1002/1521-4141(200102)31:2%3c350::AID-IMMU350%3e3.0.CO;2-0
Gómez E, Lucero MS, Richetta M, Zoth SC, Berinstein A (2018) Infectious bursal disease virus. In: Prospects of Plant-Based Vaccines in Veterinary Medicine 169–187. https://doi.org/10.1007/978-3-319-90137-4_7
Han GG, Lee JY, Jin GD, Park J, Choi YH, Kang SK, Chae BJ, Kim EB, Choi YJ (2018) Tracing of the fecal microbiota of commercial pigs at five growth stages from birth to shipment. Sci Rep 8. https://doi.org/10.1038/s41598-018-24508-7
Hoover DM, Lubkowski J (2002) DNAWorks: an automated method for designing oligonucleotides for PCR-based gene synthesis. Nucleic Acids Res 30:2–7. https://doi.org/10.1093/nar/30.10.e43
Kanaya T, Hase K, Takahashi D, Fukuda S, Hoshino K, Sasaki I, Hemmi H, Knoop KA, Kumar N, Sato M, Katsuno T, Yokosuka O, Toyooka K, Nakai K, Sakamoto A, Kitahara Y, Jinnohara T, Mcsorley SJ, Kaisho T, Williams IR, Ohno H (2012) The Ets transcription factor Spi-B is essential for the differentiation of intestinal microfold cells. Nat Immunol 13:729–736. https://doi.org/10.1038/ni.2352
Kim EB, Piao DC, Son JS, Choi YJ (2009) Cloning and characterization of a novel tuf promoter from Lactococcus lactis subsp. lactis IL1403. Curr Microbiol 59:425–431. https://doi.org/10.1007/s00284-009-9455-2
Kim JI, Park TE, Maharjan S, Li HS, Lee H Bin, Kim IS, Piao D, Lee JY, Cho CS, Bok JD, Hong ZS, Kang SK, Choi YJ (2015) Soluble RANKL expression in Lactococcus lactis and investigation of its potential as an oral vaccine adjuvant. BMC Immunol 16. https://doi.org/10.1186/s12865-015-0132-x
Knoop KA, Kumar N, Butler BR, Sakthivel SK, Taylor RT, Nochi T, Akiba H, Yagita H, Kiyono H, Williams IR (2009) RANKL is necessary and sufficient to initiate development of antigen-sampling m cells in the intestinal epithelium. J Immunol 183:5738–5747. https://doi.org/10.4049/jimmunol.0901563
Kunisawa J, Nochi T, Kiyono H (2008) Immunological commonalities and distinctions between airway and digestive immunity. Trends Immunol 29:505–513. https://doi.org/10.1016/j.it.2008.07.008
Mahgoub HA (2012) An overview of infectious bursal disease. Arch Virol 157:2047–2057. https://doi.org/10.1007/s00705-012-1377-9
McI Mowat A, Millington OR, Chirdo FG (2004) Anatomical and cellular basis of immunity and tolerance in the intestine. J Pediatr Gastroenterol Nutr 3:S723–S724. https://doi.org/10.1097/00005176-200406003-00003
Nataraj BH, Ali SA, Behare P V, Yadav H (2020) Postbiotics-parabiotics: the new horizons in microbial biotherapy and functional foods. Microb Cell Fact 19. https://doi.org/10.1186/s12934-020-01426-w
Olias P, Adam I, Meyer A, Scharff C, Gruber AD (2014) Reference genes for quantitative gene expression studies in multiple avian species. PLoS One 9. https://doi.org/10.1371/journal.pone.0099678
Ozma MA, Abbasi A, Akrami S, Lahouty M, Shahbazi N, Ganbarov K, Pagliano P, Sabahi S, Köse Ş, Yousefi M, Dao S, Asgharzadeh M, Hosseini H, Kafil HS (2022) Postbiotics as the key mediators of the gut microbiota-host interactions. Infezioni in Medicina 30:180–193. https://doi.org/10.53854/liim-3002-3
Prakash D, Verma S, Bhatia R, Tiwary BN (2011) Risks and precautions of genetically modified organisms. ISRN Ecol 2011:1–13. https://doi.org/10.5402/2011/369573
Rand JH, Wu XX, Lin EY, Griffel A, Gialanella P, McKitrick JC (2012) Annexin A5 binds to lipopolysaccharide and reduces its endotoxin activity. mBio 3. https://doi.org/10.1128/mBio.00292-11
Salminen S, Collado MC, Endo A, Hill C, Lebeer S, Quigley EMM, Sanders ME, Shamir R, Swann JR, Szajewska H, Vinderola G (2021) The International Scientific Association of Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of postbiotics. Nat Rev Gastroenterol Hepatol 18:649–667. https://doi.org/10.1038/s41575-021-00481-x
Simon D, Chopin A (1988) Construction of a vector plasmid family and its use for molecular cloning in Streptococcus lactis. Biochimie 70(4):559–566. https://doi.org/10.1016/0300-9084(88)90093-4
Song AAL, In LLA, Lim SHE, Rahim RA (2017) A review on Lactococcus lactis: from food to factory. Microb Cell Fact 16. https://doi.org/10.1186/s12934-017-0669-x
Sutton KMC, Hu T, Wu Z, Siklodi B, Vervelde L, Kaiser P (2015) The functions of the avian receptor activator of NF-κB ligand (RANKL) and its receptors, RANK and osteoprotegerin, are evolutionarily conserved. Dev Comp Immunol 51:170–184. https://doi.org/10.1016/j.dci.2015.03.006
Teame T, Wang A, Xie M, Zhang Z, Yang Y, Ding Q, Gao C, Olsen RE, Ran C, Zhou Z (2020) Paraprobiotics and postbiotics of probiotic lactobacilli, their positive effects on the host and action mechanisms: a review. Front Nutr 7. https://doi.org/10.3389/fnut.2020.570344
van Asseldonk M, de Vos WM, Simons G (1993) Functional analysis of the Lactococcus lactis usp45 secretion signal in the secretion of a homologous proteinase and a heterologous a-amylase. Mol Gen Genet 240(3):428–434. https://doi.org/10.1007/BF00280397
Vieco-Saiz N, Belguesmia Y, Raspoet R, Auclair E, Gancel F, Kempf I, Drider D (2019) Benefits and inputs from lactic acid bacteria and their bacteriocins as alternatives to antibiotic growth promoters during food-animal production. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.00057
Xuan B, Park J, Lee GS, Kim EB (2022a) Oral administration of mice with cell extracts of recombinant Lactococcus lactis IL1403 expressing mouse receptor activator of NF-kB ligand (RANKL). Food Sci Anim Resour 42:1061–1073. https://doi.org/10.5851/kosfa.2022.e54
Xuan B, Park J, Yoo JH, Kim EB (2022b) Oral immunization of mice with cell extracts from recombinant Lactococcus lactis expressing SARS-CoV-2 spike protein. Curr Microbiol 79. https://doi.org/10.1007/s00284-022-02866-w
Yamamoto M, Pascual DW, Kiyono H (2012) M cell-targeted mucosal vaccine strategies. Curr Top Microbiol Immunol 354:39–52. https://doi.org/10.1007/82_2011_134
Zhong Y, Wang S, Di H, Deng Z, Liu J, Wang H (2022) Gut health benefit and application of postbiotics in animal production. J Anim Sci Biotechnol 13. https://doi.org/10.1186/s40104-022-00688-1
Funding
This work was supported by the National Research Foundation of Korea (NRF; 2019R1A2C1009406) and Yanbian University Doctoral Research Startup Fund (No. 602024010). Biao Xuan was supported by the BK21 Plus Program of the Ministry of Education.
Author information
Authors and Affiliations
Contributions
EBK conceived the original idea, funding acquisition, project administration, and supervision. BX, JP, and SJC were involved in DNA working and animal experiment. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC; Acceptance Number: KW-210611–1) of Kangwon National University.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xuan, B., Park, J., Choi, S. et al. Postbiotic-based recombinant receptor activator of NF-κB ligand enhanced oral vaccine efficiency in chicken. Appl Microbiol Biotechnol 108, 397 (2024). https://doi.org/10.1007/s00253-024-13237-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13237-9