Abstract
Chromothripsis refers to massive genomic rearrangements developed during a catastrophic event. In total acute myeloid leukemia (AML), the incidence of chromothripsis ranges from 0 to 6.6%, in cases of complex karyotype AML, the incidence of chromothripsis ranges from 27.3 to 100%, whereas in cases of AML with TP53 mutations, the incidence ranges from 11.1 to 90%. For other types of malignancies, the incidence of chromothripsis also varies, from 0 to 10.5% in myelodysplastic syndrome to up to 61.5% in cases of myelodysplastic syndrome with TP53 mutations.
Chromothripsis is typically associated with complex karyotypes and TP53 mutations, and monosomal karyotypes are associated with the condition. ERG amplifications are frequently noted in cases of chromothripsis, whereas MYC amplifications are not. Moreover, FLT3 and NPM1 mutations are negatively associated with chromothripsis. Chromothripsis typically occurs in older patients with AML with low leukocyte counts and bone marrow blast counts. Rare cases of patients with chromothripsis who received intensive induction chemotherapy revealed low response rates and poor overall prognosis. Signal pathways in chromothripsis typically involve copy number gain and upregulation of oncogene gene sets that promote cancer growth and a concomitant copy number loss and downregulation of gene sets associated with tumor suppression functions.
Patients with chromothripsis showed a trend of lower complete remission rate and worse overall survival in myeloid malignancy. Large-scale studies are required to further elucidate the causes and treatments of the condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromothripsis refers to massive genomic rearrangements during a catastrophic event. The condition was first identified by Stephens et al., who used next-generation sequencing to identify chromothripsis in a man with chronic lymphocytic leukemia with 42 rearrangements involving the long arm of chromosome 4 [1]. They proposed that massive genomic rearrangements could occur during a single catastrophic event, causing cancer. Initially, chromothripsis was estimated to be present in only 2-3% of genomes of patients with cancer [1]. However, later studies have observed chromothripsis in the genomes of patients with several types of cancer [2,3,4,5,6] and patients with congenital anomalies [7,8,9]. Chromothripsis is thus one of the most common mechanisms involved in cancer development. The analysis of thousands of tumor samples has revealed that chromothripsis is prevalent in more than half of human cancer types [10, 11]. Chromothripsis occurs with a high frequency in liposarcomas, osteosarcomas, glioblastomas, and solid cancers such as esophageal cancer, lung cancer, and breast cancer. However, chromothripsis is relatively rare in myeloid malignancies [10].
Acute myeloid leukemia (AML) offers an excellent model to explore cancer mechanisms. The genetic evolution of AML is a dynamic process shaped by multiple cycles of mutation acquisition and clonal selection [12]. However, cases of AML developing suddenly without preceding history or abnormal laboratory results have also been reported. Chromothripsis in AML was first reported by Rausch et al. [13]. Chromothripsis has also been detected in other myeloid malignancies, but the incidence is heterogeneous (Table 1) [10, 13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28].
Chromothripsis is characterized by the aggressive clinical behavior of myelomas and melanomas and poor prognosis [2, 6]. Bochtler et al. reported chromothripsis detected or not which revealed no prognostic implication in AML patients with maker chromosome [18]. Fontana et al. reported that chromothripsis-positive patients had worse overall survival than patients with other cancer types [20]. However, the prognostic markers for AML are not well understood. The present study provides a review of the diagnostic criteria, incidence, cytogenetics, molecular findings, treatments, response, and prognosis of chromothripsis in myeloid malignancies.
Mechanism
Crasta et al. identified a mechanism by which errors in mitotic chromosome segregation generate DNA breaks, leading to the formation of micronuclei [29]. The newly generated micronuclei undergo defective and asynchronous DNA replication, resulting in DNA damage and extensive chromosome fragmentation in the micronucleus, which subsequently reintegrate into the genome. This fragmentation and reintegration is one explanation for the development of chromothripsis in cancer and developmental disorders [29]. Zhang et al. employed a combination of live-cell imaging and single-cell genome sequencing to demonstrate that micronucleus formation can generate a spectrum of genomic rearrangements, some of which mirror the features of chromothripsis [30].
Diagnosis of chromothripsis
Korbel and Campbell described conceptual criteria for the inference of chromothripsis by ruling out the alternative hypothesis that stepwise rearrangements of genetic material occurred, including (A) clustering of breakpoints, (B) regularly oscillating copy number states, (C) interspersed regions of lost heterozygosity, (D) prevalent rearrangements affecting a specific haplotype, (E) random DNA segment order and fragment joints, and (F) the ability to walk the derivative chromosome [31]. Because of the high cost of genomic sequencing, chromothripsis mostly identified by examining oscillating copy number variations in subsequent studies as Rausch et al. [13].
Rausch et al. inferred chromothripsis in cases in which at least ten switches between two or three copy number states were apparent on an individual chromosome, such as a sequence of the states ‘2’ and ‘1’ (‘2; 1; 2; 1; 2; 1; 2; 1; 2; 1; 2’), or ten switches between ‘2’ and a highly amplified state such as ‘30’ (‘2; 30; 2; 30; 2; 30; 2; 30; 2; 30; 2’) [13]. Most studies have suggested that chromothripsis follows the diagnostic rules of oscillating copy number variations [15,16,17,18,19,20,21,22,23,24,25,26]. Mackinnon and Campbell reported two AML cases of chromothripsis illustrating different aspects of this phenomenon from a cytogenetic perspective, and they employed fluorescence in situ hybridization (FISH) analyses such as multicolor FISH and multicolor banding to reveal chromothripsis [14].
For determining the organization of highly rearranged genomes, Mackinnon described strategies combining single-nucleotide polymorphism array data with various metaphase FISH methods [32]. Additionally, Coccaro et al. applied cytogenetic analysis, multicolor FISH, and optical genome mapping to fully characterize chromothripsis at high resolution [27]. However, although FISH and optical genome mapping could demonstrate chromothripsis, FISH studies require fresh samples, whereas optical genome mapping requires high-quality, long genomic DNA. Most retrospective studies cannot provide adequate specimens for research. Each method has benefit and limitation to detect chromothripsis.
Incidence of chromothripsis in AML, MDS, and MPN
Chromothripsis has been reported in studies (Table 1) of patients with AML [10, 13,14,15,16, 18,19,20,21,22,23,24,25,26,27], myelodysplastic syndrome (MDS) [10, 16, 17, 25] and myeloproliferative neoplasms (MPNs) [10, 28]. The first study of chromothripsis in patients with AML was conducted by Rausch et al., who discovered that chromothripsis was associated with TP53 mutations in medulloblastomas [13]. Chromothripsis is more commonly detected in AML patients with TP53 mutations (8/17, 47.1%) than in AML patients with the TP53 wild type (1/91, 1.1%), and it is more commonly detected in cases of complex karyotype AML (22/56, 39.3%) than in noncomplex karyotype AML cases (0/255, 0%) [13]. Fontana et al. reported 26 of 395 (6.6%) patients with AML exhibited chromothripsis [20]. The incidence of chromothripsis in total patients with AML is generally low (0-6.6%).
Similar conclusions have been reached in other studies: Chromothripsis is the most common in cases involving TP53 mutations and complex karyotype AML and MDS [15,16,17,18,19,20,21,22,23,24,25,26] (Table 1). Only one study reported no chromothripsis in 16 patients with myeloid malignancy (8 patients with AML, 1 with MDS, and 7 with MPN); however, the karyotype and TP53 mutation status of the patients in that study were not reported [10]. Studies have found that the incidence of chromothripsis ranged from 0 to 10.5% in patients with MDS, and chromothripsis was detected in 8 of 13 (61.5%) patients with TP53 mutations, including 13 of 13 (100%) with complex karyotypes [10, 16, 17, 25]. By contrast, chromothripsis is rarely reported in patients with MPN [10, 28].
Cytogenetic abnormalities, genetic mutations, and chromothripsis
Recurrent chromosomal abnormalities, such as t(8;21) and inv(16) in core binding factor leukemia or t(15;17) in acute promyelocytic leukemia, hold significant diagnostic and prognostic value for hematological malignancies. Cytogenetic abnormalities and genetic mutations associated with chromothripsis are shown in Table 2. 10–12% of all patients with AML have complex karyotypes, and the incidence of complex karyotypes increases with advanced age [33]. Complex karyotypes are associated with chromothripsis; the incidence of chromothripsis in patients with complex karyotype AML ranges from 27.3 to 100% [13, 15, 16, 18, 19, 22,23,24,25,26].
Rücker et al. reported chromothripsis in 39 of 112 patients with complex karyotype AML, and chromothripsis was associated with the chromosomal aberrations of − 5/del(5q), − 7/ del(7q), − 12/del(12p), − 16/del(16q), and − 17/del(17p) [19]. An abnormal chromosome of which no part can be identified through karyotyping is referred to as a marker chromosome. Bochtler et al. reported marker chromosomes in 165 of 1026 patients with noncore binding factor karyotype leukemia. Marker chromosomes were associated with chromothripsis, but not with the chromosomal aberrations of 5q, 7q, and 17p [18]. Additionally, Mackinnon and Campbell used multicolor FISH and cytogenetic banding analysis to diagram normal chromosomes before rearrangement, demonstrating that the normal chromosomes were broken in several segments before rejoining to form markers, rings, and derivative chromosomes in two cases of chromothripsis with complex karyotype AML [14].
Monosomal karyotypes are defined by the presence of a single autosomal monosomy (excluding the isolated loss of X or Y) in association with at least one additional autosomal monosomy or one structural chromosomal abnormality without core binding factor AML or acute promyelocytic leukemia [34]. Monosomal karyotypes are associated with poor prognoses in patients exhibiting complex karyotypes [34]. Monosomal karyotypes are associated with chromothripsis [18, 19].
TP53 mutations or loss associated with chromothripsis have been reported in several studies involving patients with AML [13, 15, 17,18,19,20, 23,24,25]. The incidence of chromothripsis in patients with AML having TP53 mutations ranges from 11.1 to 90% (Table 2) [13, 15, 17,18,19,20, 23,24,25]. ERG amplification is a secondary recurrent driver event in myeloid malignancies with complex karyotypes and TP53 mutations [23]. ERG amplification is associated with chromothripsis [23, 24]. L’Abatte et al. employed high-resolution genomic methods to investigate amplicons containing MYC in 23 patients with AML and revealed no chromothripsis [35].
Fontana et al. reported that FLT3 and NPM1 mutations were negatively associated with chromothripsis [20]. FLT3 and NPM1 mutations are commonly observed in patients with normal karyotype AML, which is also negatively associated with chromothripsis. Finally, IDH1/2, DNMT3A, CEBPA, RUNX1, and NRAS mutations did not significantly differ between patients with and without chromothripsis [20].
Clinical characteristics and chromothripsis
Bochtler et al. analyzed 49 patients with AML having marker chromosomes. Age, sex, prior MDS, leukocyte count, and lactate dehydrogenase (LDH) level did not differ between patients with and without chromothripsis [18].
Rücker et al. analyzed 39 AML patients with chromothripsis and 73 AML patients without chromothripsis [19]. Chromothripsis-positive patients were older (median age: 61 [range: 19–82] versus 56 [range: 21–76] years, p = 0.009) and had lower bone marrow blast counts (median 40% vs. 80%, p = 0.01). No differences were observed between patients in terms of sex, leukocyte count, platelet count, hemoglobin level, peripheral blood blast count, and serum LDH.
Fontana et al. analyzed 26 patients with AML with chromothripsis and 369 patients with AML without chromothripsis [20]. Significant differences were observed in age (67 vs. 60, p = 0.002) and leukocyte count (6,342 vs. 30,590, p = 0.04) between patients with chromothripsis and those without chromothripsis. Older patients had more complex karyotypes and TP53 mutations, lower bone marrow blast counts, and lower leukocyte counts. The clinical characteristics of chromothripsis are consistent with the genetic and cytogenetic findings.
Pathway-involved chromothripsis
Fontana et al. reported that, in pathway enrichment analysis of patients with AML with chromothripsis, DNA repair, E2F-mediated regulation of DNA replication, signaling pathways involving PI3K, phospholipid biosynthesis and metabolism, and various growth factor signaling pathways were in the top 1% of pathways enriched for amplification events, indicating a high association of these events with chromothripsis [20]. Additionally, CTLA4 inhibitory signaling, synthesis of phosphatidylinositol phosphate at the late endosome membrane, the Fanconi anemia pathway, and genes regulating G0, the early G1 phase, pre-NOTCH transcription, and translation were in the best 1% of pathways enriched for deletion events, indicating the high association of these deletion events with chromothripsis [20]. Furthermore, Klever reported copy number gain and the upregulation of gene sets, including genes amplified in patients with cancer having chromothripsis. Among other cancer-related genes, copy number loss and the downregulation of gene sets affect many genes with confirmed tumor suppressor functions [26].
Treatment, response, and prognosis
Genetic and cytogenetic profiles have been thoroughly investigated for risk stratification in patients with AML. Mutational profiling can be used for risk stratification to inform prognostic and therapeutic decisions for patients with AML [36, 37]. The driver landscape of AML reveals distinct molecular subgroups reflecting discrete paths for the evolution of this cancer, enabling disease classification and prognostic stratification [38].
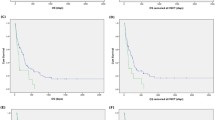
Fontana et al. reported that patients with chromothripsis had a poor response to induction chemotherapy, with only 3 of 10 patients with chromothripsis responding versus 152 of 229 patients without chromothripsis (p = 0.036). Additionally, patients with chromothripsis had worse overall survival (median of 120 days for patients with chromothripsis vs. 494 days for patients without chromothripsis, p < 0.001) [20].
Rücker et al. reported that chromothripsis was associated with a trend of lower complete remission rate (4/16 [25%] chromothripsis-positive vs. 21/45 [47%] chromothripsis-negative complex karyotype AML, p = 0.15) and thus predicted inferior survival. The estimated 2-year survival rates for chromothripsis-positive versus chromothripsis-negative patients were as follows: event-free survival, 0% versus 13% (log-rank; p < 0.001), relapse-free survival, 0% versus 34% (p < 0.008), and overall survival, 0% versus 28% (p < 0.001) [19].
Bochtler et al. reported cases from 2 large consecutive, prospective, randomized, multicenter intensive chemotherapy trials (AML96, AML2003) [18]. Marker chromosomes were detectable in 165 of 1026 (16.1%) patients with aberrant noncore binding factor karyotypes. Marker chromosomes were associated with poorer prognosis than other non-CBF aberrant karyotypes, leading to lower remission rates, inferior event-free survival, and inferior overall survival [18]. But chromothripsis detected or not which revealed no prognostic implication in a subgroup of AML patients with maker chromosome [18].
Chromothripsis is associated with poor prognosis in several malignancies [2, 6]. Only limited cases reported chromothripsis received intensive induction chemotherapy and showed fewer response rate and overall poor prognosis in AML [18,19,20]. Additionally, Fontana et al. reported 4 patients with chromothripsis receiving hypomethylating agents but did not report the outcomes [20]. Because most patients with chromothripsis have complex karyotypes and TP53 mutations, hypomethylating agents with or without venetoclax can be considered as alternative treatment for patients with AML and chromothripsis.
Further study
Most studies on patients with AML and chromothripsis follow the precedent of Rausch et al. and focus on complex karyotypes and TP53 mutations [13, 15, 17,18,19,20, 23,24,25]. Complex karyotype AML is a heterogeneous entity. Although only 10–12% of patients have complex karyotypes, effective treatment approaches should be developed for these myeloid malignancies. Hypomethylating agents with or without venetoclax and hemopoietic stem cell transplants are promising alternative treatments for these patients. However, large-scale studies of the prognosis and treatment of chromothripsis should be conducted.
Noncomplex karyotype AML has been reported to be associated with chromothripsis [13, 18]. Thus, studies incorporating AML cases other than cases of complex karyotypes, MDS, and MPN are warranted to clarify the presentation of chromothripsis.
Double-minute karyotypes are associated with complex karyotypes characterized by frequent del(17p)/TP53 mutations, micronuclei formation, myelodysplastic features, and dismal prognosis in patients with AML; these factors, in the aggregate, are highly suggestive of chromothripsis [39, 40]. Garsed et al. isolated and analyzed cancer-associated neochromosomes from liposarcomas at single-nucleotide resolution, revealing ring chromosomes in cases of chromothripsis involving chromosome 12 [41]. Additionally, derivative chromosomes have been suggested to be by-products of chromothripsis [14, 42]. Because cytogenetic studies in such cases enable prognostic stratification and diagnosis, establishing the link between chromosomal aberrations and chromothripsis may enable the creation of a prognostic scale for complex karyotype AML.
Conclusions
Chromothripsis is commonly identified by examining oscillating copy number variations, and it is most common in cases of myeloid malignancies with complex karyotypes and TP53 mutations. The incidence of chromothripsis ranges from 0 to 6.6% in total AML but ranges from 27 to 100% in cases of complex karyotypes and from 11.1 to 90% in cases of AML with TP53 mutations. Chromothripsis is associated with monosomal karyotypes. ERG amplifications are frequently noted in cases of chromothripsis, but MYC amplifications are not. Moreover, FLT3 and NPM1 mutations are negatively associated with chromothripsis. Chromothripsis is characterized by low bone marrow blast counts and low leukocyte counts, and it most frequently occurs in older patients, which is consistent with genetic and cytogenetic findings. TP53 mutations are associated with the low bone marrow blast counts and leukocyte counts commonly found in older patients, who typically have more complex karyotypes.
Patients with chromothripsis showed a trend of lower complete remission rate and worse overall survival in myeloid malignancy. The patient numbers are limited in current studies, and additional large-scale studies are required to clarify.
Data availability
No datasets were generated or analysed during the current study.
References
Stephens PJ, Greenman CD, Fu B, Yang F, Bignell GR, Mudie LJ et al (2011) Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144(1):27–40
Magrangeas F, Avet-Loiseau H, Munshi NC, Minvielle S (2011) Chromothripsis identifies a rare and aggressive entity among newly diagnosed multiple myeloma patients. Blood 118(3):675–678
Kloosterman WP, Hoogstraat M, Paling O, Tavakoli-Yaraki M, Renkens I, Vermaat JS et al (2011) Chromothripsis is a common mechanism driving genomic rearrangements in primary and metastatic colorectal cancer. Genome Biol 12(10):R103
Northcott PA, Shih DJ, Peacock J, Garzia L, Morrissy AS, Zichner T et al (2012) Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 488(7409):49–56
Molenaar JJ, Koster J, Zwijnenburg DA, van Sluis P, Valentijn LJ, van der Ploeg I et al (2012) Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature 483(7391):589–593
Hirsch D, Kemmerling R, Davis S, Camps J, Meltzer PS, Ried T et al (2013) Chromothripsis and focal copy number alterations determine poor outcome in malignant melanoma. Cancer Res 73(5):1454–1460
Kloosterman WP, Guryev V, van Roosmalen M, Duran KJ, de Bruijn E, Bakker SC et al (2011) Chromothripsis as a mechanism driving complex de novo structural rearrangements in the germline. Hum Mol Genet 20(10):1916–1924
Liu P, Erez A, Nagamani SC, Dhar SU, Kołodziejska KE, Dharmadhikari AV et al (2011) Chromosome catastrophes involve replication mechanisms generating complex genomic rearrangements. Cell 146(6):889–903
Nazaryan L, Stefanou EG, Hansen C, Kosyakova N, Bak M, Sharkey FH et al (2014) The strength of combined cytogenetic and mate-pair sequencing techniques illustrated by a germline chromothripsis rearrangement involving FOXP2. Eur J Hum Genet 22(3):338–343
Cortés-Ciriano I, Lee JJ, Xi R, Jain D, Jung YL, Yang L et al (2020) Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat Genet 52(3):331–341
Voronina N, Wong JKL, Hübschmann D, Hlevnjak M, Uhrig S, Heilig CE et al (2020) The landscape of chromothripsis across adult cancer types. Nat Commun 11(1):2320
Walter MJ, Shen D, Ding L, Shao J, Koboldt DC, Chen K et al (2012) Clonal architecture of secondary acute myeloid leukemia. N Engl J Med 366(12):1090–1098
Rausch T, Jones DT, Zapatka M, Stütz AM, Zichner T, Weischenfeldt J et al (2012) Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148(1–2):59–71
Mackinnon RN, Campbell LJ (2013) Chromothripsis under the microscope: a cytogenetic perspective of two cases of AML with catastrophic chromosome rearrangement. Cancer Genet 206(6):238–251
Jacoby MA, De Jesus Pizarro RE, Shao J, Koboldt DC, Fulton RS, Zhou G et al (2014) The DNA double-strand break response is abnormal in myeloblasts from patients with therapy-related acute myeloid leukemia. Leukemia 28(6):1242–1251
Kjeldsen E (2015) Oligo-based high-resolution aCGH analysis enhances routine Cytogenetic Diagnostics in Haematological Malignancies. Cancer Genomics Proteom 12(6):301–337
Abáigar M, Robledo C, Benito R, Ramos F, Díez-Campelo M, Hermosín L et al (2016) Chromothripsis is a recurrent genomic abnormality in high-risk myelodysplastic syndromes. PLoS ONE 11(10):e0164370
Bochtler T, Granzow M, Stölzel F, Kunz C, Mohr B, Kartal-Kaess M et al (2017) Marker chromosomes can arise from chromothripsis and predict adverse prognosis in acute myeloid leukemia. Blood 129(10):1333–1342
Rücker FG, Dolnik A, Blätte TJ, Teleanu V, Ernst A, Thol F et al (2018) Chromothripsis is linked to TP53 alteration, cell cycle impairment, and dismal outcome in acute myeloid leukemia with complex karyotype. Haematologica 103(1):e17–e20
Fontana MC, Marconi G, Feenstra JDM, Fonzi E, Papayannidis C et al (2018) Ghelli Luserna di Rorá A Chromothripsis in acute myeloid leukemia: biological features and impact on survival. Leukemia 32(7): 1609–1620
Tolomeo D, L’Abbate A, Lonoce A, D’Addabbo P, Miccoli MF, Lo Cunsolo C et al (2019) Concurrent chromothripsis events in a case of TP53 depleted acute myeloid leukemia with myelodysplasia-related changes. Cancer Genet 237:63–68
Gao J, Chen YH, Mina A, Altman JK, Kim KY, Zhang Y et al (2020) Unique morphologic and genetic characteristics of acute myeloid leukemia with chromothripsis: a clinicopathologic study from a single institution. Hum Pathol 98:22–31
Lee WY, Gutierrez-Lanz EA, Xiao H, McClintock D, Chan MP, Bixby DL et al (2022) ERG amplification is a secondary recurrent driver event in myeloid malignancy with complex karyotype and TP53 mutations. Genes Chromosomes Cancer 61(7):399–411
Schandl CA, Mazzoni S, Znoyko I, Nahhas GJ, Chung D, Ding Y et al (2023) Novel high-risk acute myeloid leukemia subgroup with ERG amplification and biallelic loss of TP53. Cancer Genet 272–273:23–28
Abel HJ, Oetjen KA, Miller CA, Ramakrishnan SM, Day RB, Helton NM et al (2023) Genomic landscape of TP53-mutated myeloid malignancies. Blood Adv 7(16):4586–4598
Klever MK, Sträng E, Hetzel S, Jungnitsch J, Dolnik A, Schöpflin R et al (2023) AML with complex karyotype: extreme genomic complexity revealed by combined long-read sequencing and Hi-C technology. Blood Adv 7(21):6520–6531
Coccaro N, Zagaria A, Anelli L, Tarantini F, Tota G, Conserva MR et al (2023) Optical genome mapping as a Tool to unveil New Molecular findings in Hematological patients with Complex chromosomal rearrangements. Genes (Basel) 14(12):2180
Brierley CK, Yip BH, Orlando G, Goyal H, Wen S, Wen J et al (2023) Chromothripsis orchestrates leukemic transformation in blast phase MPN through targetable amplification of DYRK1A. bioRxiv. 2023.12.08.570880
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y et al (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482(7383):53–58
Zhang CZ, Spektor A, Cornils H, Francis JM, Jackson EK, Liu S et al (2015) Chromothripsis from DNA damage in micronuclei. Nature 522(7555):179–184
Korbel JO, Campbell PJ (2013) Criteria for inference of chromothripsis in cancer genomes. Cell 152(6):1226–1236
MacKinnon RN (2018) Analysis of Chromothripsis by Combined FISH and microarray analysis. Methods Mol Biol 1769:53–77
Mrózek K (2008) Cytogenetic, molecular genetic, and clinical characteristics of acute myeloid leukemia with a complex karyotype. Semin Oncol 35(4):365–377
Breems DA, Van Putten WL, De Greef GE, Van Zelderen-Bhola SL, Gerssen-Schoorl KB, Mellink CH et al (2008) Monosomal karyotype in acute myeloid leukemia: a better indicator of poor prognosis than a complex karyotype. J Clin Oncol 26(29):4791–4797
L’Abbate A, Tolomeo D, Cifola I, Severgnini M, Turchiano A, Augello B et al (2018) MYC-containing amplicons in acute myeloid leukemia: genomic structures, evolution, and transcriptional consequences. Leukemia 32(10):2152–2166
Patel JP, Gönen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J et al (2012) Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med 366(12):1079–1089
Döhner H, Weisdorf DJ, Bloomfield CD (2015) Acute myeloid leukemia. N Engl J Med 373(12):1136–1152
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND et al (2016) Genomic classification and prognosis in Acute myeloid leukemia. N Engl J Med 374(23):2209–2221
Villa O, Salido M, Pérez-Vila ME, Ferrer A, Arenillas L, Pedro C et al (2008) Blast cells with nuclear extrusions in the form of micronuclei are associated with MYC amplification in acute myeloid leukemia. Cancer Genet Cytogenet 185(1):32–36
Huh YO, Tang G, Talwalkar SS, Khoury JD, Ohanian M, Bueso-Ramos CE et al (2016) Double minute chromosomes in acute myeloid leukemia, myelodysplastic syndromes, and chronic myelomonocytic leukemia are associated with micronuclei, MYC or MLL amplification, and complex karyotype. Cancer Genet 209(7–8): 313 – 20
Garsed DW, Marshall OJ, Corbin VD, Hsu A, Di Stefano L, Schröder J et al (2014) The architecture and evolution of cancer neochromosomes. Cancer Cell 26(5):653–667
Li Y, Schwab C, Ryan S, Papaemmanuil E, Robinson HM, Jacobs P et al (2014) Constitutional and somatic rearrangement of chromosome 21 in acute lymphoblastic leukaemia. Nature 508(7494):98–102
Acknowledgements
We thank the staff of the Eighth Core Lab., Department of Medical Research, National Taiwan University Hospital, for their support during the study. We thank the team of Wallace Academic Editing for English Editing.
Funding
The work was supported by the grant of National Taiwan University Hospital 111-S0097.
Author information
Authors and Affiliations
Contributions
C.C. conceived the idea, wrote, reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare there they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, CY. Chromothripsis in myeloid malignancies. Ann Hematol (2024). https://doi.org/10.1007/s00277-024-05814-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00277-024-05814-9