Abstract
Key message
Extensive modulation of numerous ARF transcripts in the embryogenic culture of Arabidopsis indicates a substantial role of auxin signaling in the mechanism of somatic embryogenesis induction.
Abstract
Somatic embryogenesis (SE) is induced by auxin in plants and auxin signaling is considered to play a key role in the molecular mechanism that controls the embryogenic transition of plant somatic cells. Accordingly, the expression of AUXIN RESPONSE FACTOR (ARF) genes in embryogenic culture of Arabidopsis was analyzed. The study revealed that 14 of the 22 ARFs were transcribed during SE in Arabidopsis. RT-qPCR analysis indicated that the expression of six ARFs (ARF5, ARF6, ARF8, ARF10, ARF16, and ARF17) was significantly up-regulated, whereas five other genes (ARF1, ARF2, ARF3, ARF11, and ARF18) were substantially down-regulated in the SE-induced explants. The activity of ARFs during SE was also monitored with GFP reporter lines and the ARFs that were expressed in areas of the explants engaged in SE induction were detected. A functional test of ARFs transcribed during SE was performed and the embryogenic potential of the arf mutants and overexpressor lines was evaluated. ARFs with a significantly modulated expression during SE coupled with an impaired embryogenic response of the relevant mutant and/or overexpressor line, including ARF1, ARF2, ARF3, ARF5, ARF6, ARF8, and ARF11 were indicated as possibly being involved in SE induction. The study provides evidence that embryogenic induction strongly depends on ARFs, which are key regulators of the auxin signaling. Some clues on the possible functions of the candidate ARFs, especially ARF5, in the mechanism of embryogenic transition are discussed. The results provide guidelines for further research on the auxin-related functional genomics of SE and the developmental plasticity of somatic cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The developmental plasticity of plant somatic cells is widely exploited in green biotechnology in the micropropagation and production of transgenic plants. Among the regeneration pathways that can be induced in a culture of somatic cells/tissue of plants, the process of somatic embryogenesis (SE) is especially attractive for biotechnology purposes as an efficient and fast system for the clonal propagation of many plant species that are of commercial value (Karami et al. 2009). Beside its usefulness in biotechnology, SE was recommended as a model system with which to study the mechanism of embryogenic development in plants (Zimmerman 1993).
The SE process begins with embryonic induction, which is frequently preceded by the de-differentiation of explant tissue (Elhiti et al. 2013). The earliest stage of SE induction attracts the most research attention, because revealing the exo- and endogenous determinants of the embryonic switch contributes to our knowledge of the general mechanism that is involved in developmental cell plasticity and also supports the improvement of the plant regeneration systems that are used in biotechnology.
Among the factors that play a substantial role in SE induction, the exogenous hormone treatment of explant tissue was determined to be required in a culture of most plant species. In over 80% of the protocols on SE induction, auxin was used alone or in combination with other plant growth regulators (Gaj 2004; Karami and Saidi 2010). Thus, an auxin-related molecular mechanism is widely expected to operate during SE induction. As an additional support, a global analysis of SE-transcriptomes indicated that numerous auxin-related genes are transcribed in the embryogenic cultures of different species, including Picea sp. (van Zyl et al. 2003; Stasolla et al. 2004), Zea mays (Che et al. 2006), Glycine max (Thibaud-Nissen et al. 2003), Solanum tuberosum (Sharma et al. 2008), and Arabidopsis thaliana (Becker et al. 2014).
In auxin-related responses, the genetic components of the auxin-signaling pathway including the AUXIN RESPONSE FACTORs (ARFs) that control the target gene expression in response to auxin were indicated as playing a central role (Teale et al. 2006). ARF genes are specific to the plant kingdom and they were identified in the genomes of different plant species, including ferns, gymnosperms, monocots, and dicots (Wang et al. 2012). Twenty-three ARF genes were described in Arabidopsis, and except for one pseudogene (ARF23), they all encode the TFs that are involved in auxin-mediated responses (Riechmann et al. 2000). The proteins of ARFs contain a DNA-binding domain that is classified as a B3-type and specific to plants, which binds to the TCTCTC motif (AuxRE) found in the promoters of auxin-responsive genes (Guilfoyle et al. 1998). ARFs may activate or repress the target gene expression depending on the amino-acid sequence of the middle region in the functional domain that interacts with DNA. ARFs with domains that are rich in glutamine, serine, and leucine (ARF5, ARF6, ARF7, ARF8, and ARF19) activate transcription while an increased content of serine, proline, leucine, and glycine (ARF1, ARF2, ARF3, ARF4, and ARF9) results in the repression of target transcription (Guilfoyle and Hagen 2001; Tiwari et al. 2001). In addition, ARFs contain the PB1 (Phox and Bem1) domain that is required for the protein–protein interaction with other ARFs and Aux/IAA (Guilfoyle 2015). Recently, a second protein interaction module, the homodimerization DD domain, which is located in DBD, was revealed in at least ARF1 and ARF5 (Boer et al. 2014). The mechanism of auxin-induced gene activation has been well recognized. In the absence of auxin, the Aux/IAA protein interacts with its partner ARF, thereby inactivating any ARF activity; while in the presence of auxin, the Aux/IAA protein is degraded through ubiquitination by the SKP-Cullin-F-boxTIR1/AFB (SCFTIR1/AFBs) E3 ubiquitin ligase complex, which contains the auxin receptor TRANSPORT INHIBITOR RESPONSE1(TIR1)/AUXIN RECEPTOR F-BOX PROTEINS (AFBs) (Dharmasiri et al. 2005).
The involvement of different ARFs in the regulation of numerous auxin-controlled developmental processes including flowering, leaf senescence, gynoecium and seed formation, root development, vascular tissue formation, and abaxial identity of organs was indicated in Arabidopsis (Guilfoyle and Hagen 2007). An involvement of ARFs in the control of ZE is also evident (Rademacher et al. 2011), including an ARF-dependent suspensor (ARF1, ARF2, ARF6, ARF9, and ARF13) and proper embryo development (ARF1, ARF2, ARF5, ARF6, and ARF7). Consequently, mutations in ARF genes were found to strongly disturb zygotic embryo development (Rademacher et al. 2011, 2012).
In contrast to ZE, the role of ARFs in SE remains mostly unknown. In this study, we profiled the expression of all known ARF genes during SE that was induced in a culture of Arabidopsis immature zygotic embryo (IZE) explants. Gene expression profiling and analysis of insertional mutants, overexpressor, and reporter lines were used to identify SE-involved ARFs. The ARFs were found to significantly differ in their activity in the embryogenic culture and the impact on somatic embryo induction. The results confirm that numerous ARFs control the embryogenic transition that is induced in somatic cells. Of the ARFs that operate during embryogenic transition that is induced in vitro, ARF5 seems to be of particular importance for SE.
Materials and methods
Plant material
The Columbia (Col-0) genotype and the transgenic lines including insertional mutants, GFP reporter and overexpressing lines of Arabidopsis thaliana (L.) Heynh., were used (Supplementary Table 1). All seeds, except for 35S::ARF5, which were kindly provided by Dr. Ive de Smet (Department of Plant Systems Biology, VIB, Ghent, Belgium), were supplied by NASC (The Nottingham Arabidopsis Stock Centre).
Plant growth and in vitro culture conditions
The plants that were used as the source of IZE explants were grown in 42 mm diameter Jiffy-7 peat pots (Jiffy) in a ‘walk-in’ type phytotron under controlled condition at 22 °C under a 16 h photoperiod of 100 µM m− 2 s− 1 white, fluorescent light. Plant materials that were grown in sterile conditions were kept at 23 °C under a 16 h photoperiod of 40 µM m− 2 s− 1 white, fluorescent light.
Somatic embryogenesis
Immature zygotic embryos in the late cotyledonary stage of development were used as explants for the in vitro culture and the standard protocol was applied to induce SE in Arabidopsis (Gaj 2001). Ten IZEs were cultured in a Petri dish (35 mm) on an agar-solidified (8 g L− l) induction medium containing basal B5 micro- and macro-elements (Gamborg et al. 1968) and 20 g L− 1 sucrose. IZE explants were cultured on an SE-induction auxin medium (E5) with 5 µM 2,4-D (2,4-dichlorophenoxyacetic acid) and on a control, hormone-free (E0) medium. Ten explants were cultured in one Petri dish and thirty explants in three replicates in each culture combination were analyzed. The capacity for SE was evaluated in 3-week-old cultures. Two parameters of embryogenic potential were evaluated: SE efficiency—frequency of the explants that produced somatic embryos and SE productivity—the average number of somatic embryos that developed per embryogenic explant.
Shoot organogenesis
To induce shoot regeneration via organogenesis (ORG), the IZEs were incubated for 7 days in a liquid callus induction medium (CIM), then the cotyledons were cut off and cultured on solid shoot induction media (SIM-C) according to Kraut et al. (2011). The CIM medium contained a basal composition of a B5 medium (Gamborg et al. 1968), 0.5 g L− 1 MES, 20 g L− 1 glucose, 2.2 µM of 2,4-D, and 0.2 µM of kinetin (Kraut et al. 2011). The SIM-C medium contained micro-elements of MS (Murashige and Skoog 1962), macro-salts, vitamins of a B5 medium (Gamborg et al. 1968), and was supplemented with 30 g L− 1 sucrose, 0.5 µM of NAA (1-naphthaleneacetic acid) and 4.4 µM of BAP (6-benzylaminopurine). The explant capacity for ORG was evaluated in 3-week-old cultures. Two parameters of culture morphogenic potential were evaluated: ORG efficiency—frequency of the explants that produced shoots and ORG productivity—the average number of shoots that developed per explant.
RNA isolation and gene expression analysis
An RNAqueous Kit (AMBION) was used to isolate total RNA from the IZE explants that were induced on the SE induction (E5), callus induction (CIM), shoot induction (SIM), and the hormone-free (E0) medium on days 0, 3, 5, 10, and 15 of the culture. Depending on the age of the culture, from 250 (0 day) to 4 (15 days), explants were used for the isolation of RNA. The concentration and purity of RNA were evaluated with a ND-1000 spectrophotometer (NanoDrop). To prevent DNA contamination, the RNAs were treated with RQ1 RNase-free DNase I (Promega) following the manufacturer’s instructions. First-strand cDNA was produced in a 20 µL reaction volume using a RevertAid First-Strand cDNA Synthesis Kit (Fermentas).
The product of the reverse transcription was diluted with water in a 1:1 ratio and 1 µL of this solution was used for RT-PCR. To quantify the expression of ARF genes, Real-Time quantitative RT-PCR (RT-qPCR) method was employed. The product of the reverse transcription was diluted in 3:1 ratio than 2.5 µL was used for reactions. RT-qPCR was carried out in a 10 µL reaction volume using a LightCycler® 480 SYBR™ Green I Master (Roche) kit, a LightCycler® 480 Multiwell Plate 96, and MultiwellSealin Foil (Roche).
A LightCycler® 480 System (Roche) real-time detection system was used under the following reaction conditions: denaturation one repeat of 5 min at 95 °C, followed by 45 repeats of 10 s at 95 °C, 20 s at 58 °C, and 10 s at 72 °C. Denaturation for melt curve analysis was conducted at 95 °C followed by 5 s, at 65 °C by 1 min, and 98 °C (0.11ºC/s for fluorescence measurement). Cooling was at 40 °C for 10 s.
Relative RNA levels were calculated and normalized to internal controls, the AT1G07920 ELONGATION FACTOR1α (EF1α) gene encoded GTP-binding elongation factor (Reid et al. 2006) and AT4G27090 (TIN) gene encoded 60 S ribosomal protein (Thellin et al. 1999). Fold change values were calculated using the comparative 2− ΔΔCt method. The control gene exhibited a constant expression pattern with Cp = 24 ± 1 in all of the tissue samples that were analyzed. The primers (Supplementary Table 1) used for expression profiling of the genes were designed using the QuantPrime tool (http://www.quantprime.de/). The plant tissues for gene expression analysis were produced in three biological replicates and two technical replicates of each repetition were carried out. All of the information about the RNA and RT-qPCR quality and methodology are presented in Supplementary Table S1.
Microscopy
Analysis of GFP signal was carried out using a Nikon Eclipse Ni-E/Ni-U fluorescent microscope system. GFP fluorescence was excited using halogen lamphouses with a 100–240 VAC (Prior Lumen200) and a wavelength of 488 nm. Photographic documentation was created from images that were recorded with a Nikon Digital Sight DS-Fi2 with DS-U3 camera. Image processing was performed using the NIS-Elements F computer program version 4.0.
Statistical analysis
The student t test was used to calculate any significant differences (at P = 0.05) between the combinations that were being compared.
Results
Design of the experiment
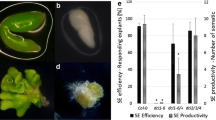
A culture of IZEs results in SE, shoot organogenesis (ORG), or seedling development in Arabidopsis, and the type of morphogenic response of explants mainly depends on the hormonal content of the media that is used (Kraut et al. 2011). To identify the ARFs that are differently expressed during SE, explant tissues that were induced on various media towards SE, ORG, and seedling development were sampled (Fig. 1). The analysis covered the different stages of the culture, including freshly isolated (0 day) and media-cultured (3–15 days) IZE explants. In the SE culture, samples taken on days 3 and 5 of the culture are related to the SE-induction stage, while the later culture stages, which are related to the formation and development of somatic embryos, correspond to days 10 and 15 of the culture, respectively.
Diverse morphogenic pathways that were induced in vitro in a culture of Arabidopsis IZE explants. IZE explants of Col-0 that were cultured on different media resulted in: somatic embryo development, SE (a), shoot regeneration, ORG (b), and seedling development, E0 (c); the tissue for the analysis of ARFs expression was sampled at different time points of the cultures, including 0, 3, 5, 10, and 15 days. Asterisk regenerated somatic embryos or shoots
Expression profiling of the ARFs transcribed in the embryogenic culture
The analysis of the ARFs expression based on qualitative RT-PCR reaction, which was monitored at different stages of SE, indicated that the majority (14) of the 22 analyzed genes were transcribed in the IZE explants and the embryogenic cultures that were derived, including ARF1, ARF2, ARF3, ARF5, ARF6, ARF7, ARF8, ARF9, ARF10, ARF11, ARF16, ARF17, ARF18, and ARF19 (Fig. S1). The RT-qPCR analysis revealed that the ARFs that were transcribed in the embryogenic culture displayed diverse patterns of expression and the genes differed distinctly in the levels and profiles of their transcription (Fig. S2). ARF5 and ARF10 were found to have the highest transcript accumulation in the SE culture, while in contrast, the activity of ARF7 and ARF11 was almost 7–9 threshold cycles lower. Most of the SE-transcribed ARFs displayed a substantial modulation of their expression level during SE in relation to 0 day and genes that displayed both increased and decreased transcription levels in specific SE-stages were indicated.
ARFs up-regulated during SE
Six of the ARFs were found to be significantly up-regulated at different (3–15 days) time points of the embryogenic culture and the genes with a stimulated transcription in the early (3–5 days) or late (10–15 days) stages of SE could be distinguished (Fig. 2). The majority (four) of the SE-stimulated ARFs including ARF5, ARF6, ARF10, and ARF16 displayed a distinct up-regulation during the inductive stage of SE (3–5 days), while two other genes, ARF8 and ARF17, showed a significantly increased level of expression in the advanced stages of SE that were associated with the formation and development of the somatic embryo (10–15 days).
ARF5 (a), ARF6 (b), ARF8 (c), ARF10 (d) ARF16 (e), and ARF17 (f) of up-regulated expression during the SE process. IZE explants of Col-0 were cultured on an E5 medium and the tissue was sampled on the 0, 3, 5, 10, and 15 days of the culture. The relative transcript level was normalized to the internal control (TIN gene) and calibrated to the 0 day culture. Asterisk expression level significantly different to that observed at 0 day at P < 0.05. Means and SD for three biological replicates are shown
Among the ARFs with an SE-stimulated expression, the ARF5 gene was found to be especially highly activated and its transcript level showed a distinguishably sharp increase in the early SE (up to 14-fold). This observation suggested that ARF5 strongly affects the embryogenic transition in explant cells.
ARFs down-regulated during SE
Five of the ARF genes including ARF1, ARF2, ARF3, ARF11, and ARF18 were observed to display a significant reduction in their activity in the early stage of SE induction (3–5 days), which also remained low in the advanced culture (10–15 days) (Fig. 3). Among these genes, the activity of ARF11 was found to be the most reduced in the embryogenic culture and transcripts of this gene were up to four-times less abundant in the SE-induced explants than in the freshly isolated explants.
ARF1 (a), ARF2 (b), ARF3 (c), ARF11 (d), and ARF18 (e) of down-regulated expression during the SE process. IZE explants of Col-0 were cultured on an E5 medium and the tissue was sampled on the 0, 3, 5, 10, and 15 days of the culture. The relative transcript level was normalized to the internal control (TIN gene) and calibrated to the 0 day culture. Asterisk expression level significantly different to that observed at 0 day at P < 0.05. Means and SD for three biological replicates are shown
ARFs with a stable expression level during SE
The observed during SE changes in transcript level were found not to be significant for three genes: ARF7, ARF9, and ARF19, and thus, these ARFs were considered to have a stable expression during SE (Fig. S3).
Auxin treatment and ARFs expression
To reveal the ARFs with an auxin-regulated expression, in addition to the E5 auxin medium with a SE-promoting effect, the gene expression was also evaluated in IZEs that had been cultured on the auxin-free medium (E0) that resulted in seedling development. A comparison of the ARF expression profiles on the E5 versus E0 medium indicated that six ARF genes (ARF5, ARF6, ARF8, ARF9, ARF10, and ARF16) had a distinctly higher expression level under auxin treatment (Fig. 4). The majority of these genes (ARF5, ARF6, ARF8, ARF10, and ARF16) also showed a significantly increased transcript abundance in the embryogenic culture (Fig. 3). Thus, auxin treatment appears to be responsible for the distinct up-regulation of most of the ARF genes that are observed during SE. Among the ARFs that are auxin-stimulated and up-regulated during SE, ARF5 was ascertained to be the most highly auxin-responsive and its expression was up to 16-times higher (5 days) on E5 than on the E0 medium. ARF9 was also identified among auxin-stimulated ARFs; however, its activity was not significantly increased between 0–15 days of SE culture. By contrast, ARF17, which was observed to be up-regulated during SE (Fig. 2), was found not to be stimulated in response to auxin treatment, which implies that other factors besides auxin may be responsible for the activation of ARF genes during SE.
ARF5 (a), ARF6 (b), ARF8 (c), ARF9 (d), ARF10 (e), and ARF16 (f) genes of auxin-stimulated expression during the SE process. IZE explants of Col-0 were cultured on an E5 medium and the tissue was sampled on the 0, 3, 5, 10, and 15 days of the culture. Relative transcript level was normalized to the internal control (TIN gene) and calibrated to the gene expression in explants that had been induced on an auxin-free (E0) medium. Asterisk expression level significantly different to that observed on the E0 medium at P < 0.05. Means and SD for three biological replicates are shown
ARFs differentially expressed in SE versus ORG
To determine whether the patterns of ARFs expression that were observed in the embryogenic culture are unique for SE induction or whether they result from the general processes of de- and re-differentiation that are expected in hormone-treated explant tissue, IZE explants that had undergone ORG were also studied. The results revealed that all 14 ARFs that were transcribed in the embryogenic culture were also expressed in the ORG culture (Fig. S4). Comparison of the gene expression levels during SE and ORG, using the RT-qPCR method, resulted in the identification of ARF genes that displayed distinctly different levels of activity in these processes. Four of the ARFs (ARF5, ARF6, ARF8, and ARF9) were observed to show a lower expression level during ORG than in the SE culture (Fig. 5). Three of them (ARF5, ARF6, and ARF8) appear to be especially interesting in terms of their possible involvement in SE due to their considerably higher expression in the embryogenic culture.
ARF genes with a distinctly lower expression level in ORG than in the SE culture: ARF5 (a), ARF6 (b), ARF8 (c), and ARF9 (d). Relative transcript level was normalized to the internal control (TIN gene) and calibrated to the ARFs expression during the SE process. Asterisk expression level significantly different to that observed in the SE culture at P < 0.05. Means and SD for three biological replicates are shown
Summary of the results of the PCR/qPCR analysis
Comparison of the ARF expression profiles in the explants that were subjected to different culture conditions that promote SE and ORG showed that similar ARFs were transcribed in these processes and that the set of the SE/ORG-expressed genes included a majority (14 of 22) of the analyzed genes (Supplementary Table S2). The expression patterns of five of these ARFs (ARF6, ARF8, ARF9, ARF16, and ARF19) were distinctly different in SE compared to ORG and ARF5, ARF6, ARF8, and ARF9 displayed a significantly higher transcript accumulation in SE than in ORG. Except for three genes (ARF7, ARF9, and ARF19) that were transcribed at a stable level in the embryogenic culture, a predominant number of ARFs (11/14; 79%) displayed a significantly modulated expression profile during SE and a similar number of the up- (6) and down- (5) regulated ARFs were observed. In line with the key role of auxin in the SE induction mechanism, it was found that auxin treatment might have been the cause of the increased expression of almost all of the genes that were up-regulated during SE with the exception of ARF17.
GFP reporter analysis of ARF expression during SE-induced explants
The spatio-temporal pattern of ARFs expression in the IZE explants undergoing SE induction was analyzed using the GFP reporter lines. In particular, the explant areas that are involved in SE induction, i.e., the cotyledons and the vicinity of SAM (Kurczyńska et al. 2007) were inspected in terms of the GFP signal.
The GFP signal was identified in nine reporter lines that monitored expression of ARF1, ARF2, ARF3, ARF5, ARF6, ARF10, ARF16, ARF18, and ARF19, and in six of them, a reporter signal was detected in the IZE regions that are involved in SE induction (Fig. 6). In freshly isolated IZE explants (0 day), a GFP signal was detected for ARF2, ARF3, ARF5, and ARF10. During the early SE induction (3–5 days), ARF5, ARF6, ARF10, and ARF16 were observed to be expressed along the cotyledons and in the vicinity of SAM and an especially strong GFP signal was seen in the ARF5 reporter line. SE-related expression in the early culture (3–5 days) also displayed two others genes, ARF2 and ARF3, with the GFP signal localized exclusively in the vicinity of SAM. In the advanced embryogenic culture (10–15 days), a weak GFP signal was seen in the developing somatic embryos in some of the reporter lines (ARF2, ARF3, ARF6, and ARF16), while a strong fluorescence indicative of the expression of ARF2, ARF5, ARF10, and ARF16, was found to be associated with the callus tissue produced at the base of the somatic embryos and in the explant parts not engaged in SE (hypocotyl and root).
Functional test of SE-transcribed ARFs
arf mutants
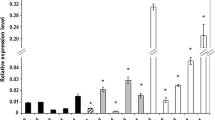
To further explore the involvement of ARF genes in SE, the arf insertional mutants were analyzed in terms of their capacity for SE induction. In total, 12 of the arf mutants (arf1, arf2, arf3, arf5, arf6, arf7, arf8, arf10, arf11, arf16, arf17, and arf19) were evaluated and the analysis indicated that most (seven) of the arf mutants were impaired in their embryogenic response (Fig. 7a). Three of the mutants, arf1, arf5, and arf7, displayed a significant reduction in both of the parameters of SE capacity, i.e., SE efficiency and SE productivity. The other mutants were defective in SE efficiency or SE productivity, and accordingly, arf3 and arf6 displayed a reduced frequency of SE-responsive explants, while arf8 and arf11 were observed to produce a significantly fewer number of somatic embryos per responsive explant.
Functional test of SE-transcribed ARFs. The embryogenic potential, which was measured by SE efficiency and SE productivity, was evaluated in cultures derived from: arf insertional mutants (a); arf5 explants of a weak (W), middle (M), and strong (S) mutant phenotype (b); 35S::ARF2 overexpressor line (c). Asterisk embryogenic capacity significantly different to that observed in the Col-0 control culture at P < 0.05. Means and SD for three biological replicates are shown
The arf5 mutant was found to be the most hampered in its embryogenic response and the severity of developmental defects that were displayed by the IZEs of the mutant (classified as a weak, medium, and strong mutant phenotype) was positively correlated with a reduction of the SE response (Fig. 7b). An extreme decrease in embryogenic capacity (over 80%) was observed in the culture of IZEs that had a strong, the so-called monopteros, mutant phenotype, which is characterised by a single cotyledon and the lack of the root meristem and hypocotyl (Odat et al. 2014).
ARF overexpressors
Further evidence of the considerable role of ARF5 in SE induction was provided by the analysis of the 35S::ARF5 line. Overexpression of ARF5 results in the strongly defective development of plants including distorted inflorescences, which may terminate in pin-shaped inflorescence tips and high plant sterility (Hardtke et al. 2004). Therefore, only 20 IZEs were collected from 12 plants and cultured on the SE-induction medium. None of the explant overexpressing ARF5 was capable of developing somatic embryos and instead only a callus was formed.
In addition to a highly expressed ARF5 gene, the effect of overexpression on an explant’s capacity for SE was also evaluated in the 35S::ARF2 line that overexpressed an ARF that had a substantially decreased transcription during SE. The analysis indicated that the increased activity of ARF2 resulted in a substantial reduction in embryogenic response and both SE efficiency and productivity were significantly reduced (Fig. 7c).
Candidate targets of ARFs in SE
We assumed that the targets of SE-modulated ARFs might be identified within the auxin-responsive TF genes that play a regulatory role in SE. Accordingly, using the Arabidopsis cis-regulatory element database (AGRIS; http://arabidopsis.med.ohio-state.edu/AtcisDB/), we found that an Auxin Response Element (AuxRE) is present in the promoters of the majority (60%) of SE-modulated TFs (Gliwicka et al. 2013) including the genes that were indicated to have an essential function in embryogenic transition such as LEAFY COTYLEDON (LEC1 and LEC2; Gaj et al. 2005), FUSCA3 (FUS3; Ledwoń and Gaj 2011), WUSCHEL (WUS; Zuo et al. 2002), BABY BOOM (BBM; Boutilier et al. 2002), and MYB115 (Wang et al. 2009). It remains to be revealed which of the candidate ARFs bind to the promoters of the key regulators of SE. Among the ARF-TF regulatory interactions that operate during SE, the direct regulation of LEC2 by ARF5/MP cannot be excluded due to the significantly reduced LEC2 transcript level that was found in an mp/arf5 mutant that had a seriously defective embryogenic response (Fig. S5).
To gain more insight into the ARF-mediated regulatory interactions that operate in SE, we also verified the possibility of an auto-feedback regulation of ARFs (Lau et al. 2011; Okushima et al. 2005). Accordingly, we used the AGRIS analytical tool to search for AuxREs in the promoters of the ARFs that were found to display a significantly and SE-modulated expression level in this study. The analysis showed the presence of AuxREs in the majority (85%) of the SE-modulated ARFs (Supplementary Table 3). Thus, the complexity of the ARF-controlled regulatory relations that operate in SE might be enhanced by the auto-feedback regulated transcription of the candidate ARFs.
Discussion
Numerous ARFs contribute to SE-transcriptome via distinctly different expression patterns
Auxin signaling is believed to play a pivotal role in plant development including morphogenic processes that are induced in in vitro cultured explants (reviewed in Ikeuchi et al. 2016; Weijers and Wagner 2016). Consistent with this belief, the global analysis of the SE-transcriptome of Arabidopsis indicated that components of the auxin signaling pathway, including AUX/IAA and ARF, which are the main regulators of the auxin signaling, together with auxin-responsive target genes are extensively modulated during SE induction (Gliwicka et al. 2013; Wickramasuriya and Dunwell 2015). This study, whose aim was to identify the candidate ARFs that are involved in SE induction indicated that most (14/23) of the ARFs encoded in Arabidopsis genome were transcribed in the embryogenic culture and that the great majority of these (11) displayed a significantly and SE-modulated expression level.
ARF proteins are believed to activate or repress the target genes (Weijers and Wagner 2016) and in the transient and heterologous assays, individual ARFs are classified as activators/repressors of the gene transcription (Tiwari et al. 2001). According to this classification, the ARFs of SE-modulated expressions that were identified in this study represent both the activators (ARF5, ARF6, ARF7, ARF8, and ARF19) and repressors (ARF1, ARF2, ARF3, and ARF9) of the target gene transcription. However, it is necessary to experimentally validate the regulatory relationship between the ARFs and their SE-targeted genes among which the auxin-responsive genes encoding the TFs that have a documented function in SE such as LEC2, FUS3, WUS, BBM, and MYB115 (reviewed in Nowak and Gaj 2016) might be considered.
It is also worth noting that more complex behaviours of ARFs in SE than simply the direct activation/repression of the target genes may exist including binding ARFs with repressor and activator functions to the same AuxREs in the same target gene (Vernoux et al. 2011; Bargmann and Estelle 2014; Chandler 2016). Consistent with this expectation, we observed a distinct up-regulation of several AuxRE-containing ARFs (ARF5, ARF6, ARF8, ARF10, and ARF16) in response to auxin treatment. In further support for a vital role of auxin treatment in the regulation of ARFs expression during SE, we observed that ARFs that lacked AuxRE were not transcribed (ARF14 and ARF22) or that their expression was not modulated (ARF7) in the embryogenic culture of Arabidopsis.
In addition to auxin, various abiotic stresses were also indicated as modulating the transcription of ARFs (Tian et al. 2004; Hannah et al. 2005; Jain and Khurana 2009; Blomster et al. 2011; Naser and Shani 2016). Thus, the stress conditions including ROS (Reactive Oxygen Species), which are actively produced under embryogenic culture, may control the expression of ARFs during SE induction (Zavattieri et al. 2010; Blomster et al. 2011).
ARF-mediated transcriptional responses are believed to be highly specific not only to the developmental process but also to tissue and cell type (Weijers et al. 2005), and this raises a question about the extent of the similarities of the ARF-mediated auxin responses that are activated in different SE systems. A comparison of the transcriptional profiles of ARFs that were identified in the embryogenic cultures of various plants infers that the contribution of the individual ARFs to the mechanism of SE induction may differ between embryogenic cultures (Thibaud-Nissen et al. 2003; Ooi et al. 2012; Yang et al. 2012; Xu et al. 2013; Lin et al. 2015; Indoliya et al. 2016; Zheng et al. 2016).
Considering the apparent molecular similarities of SE to ZE (Dodeman et al. 1997; Winkelmann 2016), we found the expression patterns of some ARFs to be convergent in both the embryogenic culture and in the ZE of Arabidopsis, and, e.g., a significant up-regulation of ARF5 and ARF6 in both of these processes is evident (present results versus http://www2.bri.nrc.ca/plantembryo). In ZE, these closely related ARFs act synergistically; however, a mutation in only in one of these genes, ARF5, causes substantial defects in the zygotic embryo (Rademacher et al. 2011). In contrast to ZE, we found the arf6 mutant to be significantly impaired in its capacity for SE and these inconsistent mutant phenotypes suggest different contributions of ARF6 in the regulation of SE and ZE. In particular, different roles and regulatory relations of individual ARFs can be expected in the very early stages of these processes. In contrast to the ZE that originates from a totipotent zygotic cell, SE induction involves the re-programming of already differentiated somatic cell/cells that respond to the auxin treatment. As a result, IAA is accumulated in the SE-induced cells (Michalczuk et al. 1992; Ribnicky et al. 2002; Kurczyńska et al. 2007; Wójcikowska and Gaj 2015) and in turn, a unique SE-specific set of ARFs might be triggered. To illustrate the assumptive differences in auxin responses in the early ZE and SE, we observed a distinct down-regulation of ARF2 and ARF3 during an early stage of SE that contrasted with the high expression of these genes in a zygotic cell of Arabidopsis (Rademacher et al. 2011; Xiang et al. 2011).
SE-involved candidate ARFs—implications from the mutant analysis
To validate the SE-associated functions of the ARFs that had an SE-modulated expression pattern, the arf mutants were evaluated in terms of their embryogenic potential. The analysis showed that in spite of the extensive functional redundancy of the ARFs that were indicated in the developmental processes (Hardtke et al. 2004; Guilfoyle and Hagen 2007), the majority of the arf mutants were found to be defective in the SE response. These distinct SE-phenotypes of individual arf mutants suggest that the SE-related functions of the ARF proteins are not simply interchangeable as was also postulated in ZE (Rademacher et al. 2012).
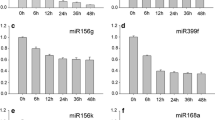
Several arf mutants including arf10 and arf16 were found not to be affected in embryogenic potential, although a significant and auxin-dependent accumulation of ARF10 and ARF16 transcripts was associated with SE induction (present results). Thus, it may be possible that similar to in planta development (Wang et al. 2005) the ARF10 and ARF16 genes also act redundantly in SE. In support of an ARF16 and ARF10, function in SE is the fact that these genes, together with IAA17/AXR3, act upstream of the PLETHORA (PLT) genes that regulate stem cell differentiation in Arabidopsis roots (Ding and Friml 2010) and have an assumed interaction with auxin in the control of the early SE (Horstman et al. 2014). Relevantly to this assumption, the significant accumulation of ARF16 and ARF10 transcripts (present result) was accompanied by an increased expression of the IAA17/AXR3, PLT1, and PLT2 in an embryogenic culture of Arabidopsis (Gliwicka et al. 2013; Wickramasuriya and Dunwell 2015). Further support for the engagement of ARF10 and ARF16 in SE is that the up-regulation of these genes in the SE of Arabidopsis and other plants is accompanied by the down-regulation of miR160, which is a post-transcriptional regulator of ARF10 and ARF16 expression in planta (Mallory et al. 2005; Zhang et al. 2012; Lin and Lai 2013; Szyrajew et al. 2017).
The present results showed that mutations in ARF1 and ARF3 that had a distinctly down-regulated transcription during SE resulted in a significantly impaired embryogenic response. Similarly, a knockout mutation in the SE-engaged gene, ERF022, which had a highly repressed transcription during SE, caused a significantly reduced embryogenic response (Nowak et al. 2015). To interpret these unobvious effects of the SE-modulated genes with regulatory function, fine-tuning of the gene transcript level to that required for the promotion of the embryogenic transition might be considered. In support of this explanation, a specific level of PLT genes transcription was indicated to be associated with embryogenic response induced in culture of Arabidopsis explants (Horstman et al. 2015). Thus, SE-promoting effect of ARFs might be dependent on their expression level and relevantly to this assumption in flower development, a specific level of ARF5 was found to be required for transcriptional regulation of the LEAFY gene (Yamaguchi et al. 2016).
Analysis of the overexpressor lines might be more informative than mutant analysis in the identification of the redundant gene functions (Prelich 2012). In accordance with this expectation, a phenotype of the arf2 mutant, similar to ZE (Rademacher et al. 2011), was not affected in SE, while the overexpression of ARF2 resulted in a significant reduction of the embryogenic response (present results). Consistent with the general function of ARF2 that is annotated to the repression of cell divisions (Schruff et al. 2006), we indicated a distinct down-regulation of this gene transcripts during SE. Another line of evidence that supports ARF2 involvement in the SE was provided by reports on the contribution of this gene to hormone-related responses including the repression of auxin signaling (Lim et al. 2010) and the integration of the auxin and brassinosteroid pathways (Vert et al. 2008).
ARF5/MONOPTEROS contribution to SE
Consistent with the global SE-transcriptome analysis, we found ARF5, which encodes the MONOPTEROS (MP) protein, to be the most highly expressed member of the ARF gene family in the embryogenic culture of Arabidopsis. An increased activity level of MP was also reported in an embryogenic culture of soybean (Thibaud-Nissen et al. 2003). Thus, MP, similar to its fundamental role in the regulation of different aspects of ZE in Arabidopsis (Hardtke and Berleth 1998; Aida et al. 2002; Schlereth et al. 2010) seems to control the development of the somatic embryo. Evidence about a possible function of MP in auxin signaling during SE was provided recently by the indicated regulatory relation between MP and PHABULOSA (PHB), which is a positive regulator of LEC2 that has an essential function in SE induction (Tang et al. 2012; Wójcikowska et al. 2013; Müller et al. 2015). In support of the regulatory impact of MP on the PHB-LEC2, pathway in SE is the up-regulation of PHB that was observed in an embryogenic culture of Arabidopsis (A. Wójcik, MDG, data not published) as well as the significantly reduced LEC2 transcript level in an mp/arf5 mutant (present results).
MP might also contribute to the embryogenic development that is triggered in plant somatic cells by controlling other TF genes that have a documented role during in planta development, including HOMEOBOX GENE 8 (ATHB8) and TARGET OF MONOPTEROS3 (TMO3), TMO5, TMO6, and TMO7 (Donner et al. 2009; Schlereth et al. 2010). All of these MP targets were found to be up-regulated during SE in Arabidopsis (Gliwicka et al. 2013; Wickramasuriya and Dunwell 2015).
Similar to other ARFs, the developmental specificity of ARF5/MP function is generated by interactions with Aux/IAA proteins (Kieffer et al. 2010; Weijers et al. 2005), and many members of the Aux/IAA family were indicated to interact with MP in Arabidopsis (Krogan et al. 2014). Among the SE-involved AUX/IAA candidates is IAA30, which interacts with MP in ZE (Müller et al. 2015), and in support of its role in SE, a significantly impaired embryogenic response of the iaa30 mutant was indicated (Gliwicka et al. 2013). Moreover, a regulatory relation between MP and BDL/IAA12 might be of importance for the SE induction mechanism as this protein pair is a major effector of auxin action in the zygotic embryo that influences the auxin-mediated cell-fate decisions in the early embryogenesis (Lau et al. 2011).
The function of MP in SE may also involve the regulation of the polar auxin transport and disturbed auxin transport was indicated to distinctly impair the embryogenic response of cultured explants (Chen and Chang 2004; Cueva-Agila et al. 2016). The MP-mediated control of the PIN1 (PIN-FORMED1) gene that encoded the auxin efflux carrier was reported and a feedback regulatory loop that involves auxin, MP, and PIN1 was proposed (Wenzel et al. 2007; Krogan et al. 2016). Relevant to the assumed role of MP-mediated auxin transport pathway in SE, explants of a pin1-7 mutant were indicated to be defective in embryogenic induction in vitro (Su et al. 2009). Moreover, the 35S::ARF5 line that resembles the pin1 phenotypes (Hardtke et al. 2004) was found to be incapable of SE induction (present results). In conclusion, the strong inhibition of embryogenic potential that was observed in this study in the mp/arf5 mutant and in the 35S::ARF5 line possibly result from the disturbed auxin signaling and impaired auxin transport that are expected in these forms (Mattsson et al. 2003; Lau et al. 2011).
Besides auxin, the possible contributions of MP in SE also involve the regulation of the cytokinin response pathway due to the involvement of MP in the regulation of cytokinin responses during plant development in planta and in vitro (Zhao et al. 2010; Ckurshumova and Berleth 2015). Considering that the patterning and cell organization that are associated with de novo regenerated shoot meristems resemble the formation of the embryonic SAM (Gordon et al. 2007), a cytokinin signaling-related function of MP in the development of the somatic embryo might also be expected. The distinctly up-regulated expression of ARF10 in SE (this study), which positively regulates de novo shoot regeneration via the activation of the shoot meristem-specific genes, supports this assumption (Qiao et al. 2012).
In conclusion, ARF5/MP is assumed to control SE via versatile pathways (Fig. S6) and further analyses are needed to experimentally verify which of the MP-mediated regulatory interactions occur in an embryogenic culture.
Conclusions
The study provides several pieces of evidence that numerous ARFs, which are the core regulators of auxin response, contribute significantly to the embryogenic switch that is induced in plant somatic cells in vitro. The candidate ARFs provide guidelines for further research on the auxin-mediated regulation of SE. Among the SE-involved candidates, the ARF5 encoding MP protein that has a substantial role in the development of the zygotic embryo seems to significantly contribute to SE.
In the regulation of SE, similar to the in planta development including ZE, not only do the activities of individual ARFs need to be considered, but also the total ARF complement of a cell may create a pre-pattern that determines the SE-specific cellular response (reviewed in Chandler 2016). Therefore, one current challenge in deciphering the SE-related auxin response is the identification of the spatio-temporal and cell-specific sets of the ARFs and the interacting AUX/IAA elements that contribute to the embryogenic transition of somatic cells. To understand the biological functions of ARFs in SE, the ARF-targeted TF genes are required to be identified. In addition, the miRNAs and chromatin remodelers that control the SE-involved ARFs need to be uncovered considering the recently indicated key role of the post-transcriptional regulation of ARFs in plant development (Mallory et al. 2005; Oh et al. 2014).
Author contribution statement
Małgorzata Danuta Gaj and Barbara Wójcikowska: (i) conceived and designed the experiments, and (ii) wrote the manuscript. Barbara Wójcikowska: performed the experiments and analyzed the data. All of the authors read and approved the manuscript.
Abbreviations
- 2,4-D:
-
2,4-Dichlorophenoxyacetic acid
- ARF:
-
AUXIN RESPONSE FACTOR
- Aux/IAA:
-
AUXIN/INDOLE-3-ACETIC ACID
- CIM:
-
Callus induction medium
- Ct:
-
Threshold cycle
- E0:
-
Auxin-free induction medium
- E5:
-
Embryogenesis induction medium with auxin
- IZE:
-
Immature zygotic embryo
- NAA:
-
1-Naphthaleneacetic acid
- ORG:
-
Organogenesis
- PGR:
-
Plant growth regulator
- SAM:
-
Shoot apical meristem
- SE:
-
Somatic embryogenesis
- SIM:
-
Shoot induction medium
- TF:
-
Transcription factor
- ZE:
-
Zygotic embryogenesis
References
Aida M, Vernoux T, Furutani M, Traas J, Tasaka M (2002) Roles of PIN-FORMED1 and MONOPTEROS in pattern formation of the apical region of the Arabidopsis embryo. Development 129:3965–3974
Bargmann BO, Estelle M (2014) Auxin perception: in the IAA of the beholder. Physiol Plant 151:52–61
Becker MG, Chan A, Mao X, Girard IJ, Lee S, Mohamed E, Stasolla C, Belmonte MF (2014) Vitamin C deficiency improves somatic embryo development through distinct gene regulatory networks in Arabidopsis. J Exp Bot 65:5903–5918
Blomster T, Salojärvi J, Sipari N, Brosché M, Ahlfors R, Keinänen M, Overmyer K, Kangasjärvi J (2011) Apoplastic reactive oxygen species transiently decrease auxin signaling and cause stress-induced morphogenic response in Arabidopsis. Plant Physiol 157:1866–1883
Boer DR, Freire-Rios A, van den Berg WA, Saaki T, Manfield IW, Kepinski S, López-Vidrieo I, Franco-Zorrilla JM, de Vries SC, Solano R, Weijers D, Coll M (2014) Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 156:577–589
Boutilier K, Offringa R, Sharma VK, Kieft H, Ouellet T, Zhang L, Hattori J, Liu CM, van Lammeren AAM, Miki BLA, Custers JBM, van Lookeren Compagne MM (2002) Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14:1737–1749
Chandler JW (2016) Auxin response factors. Plant Cell Environ 39:1014–1028
Che P, Love TM, Frame BR, Wang K, Carriquiry AL, Howell SH (2006) Gene expression patterns during somatic embryo development and germination in maize Hi II callus cultures. Plant Mol Biol 62:1–14
Chen JT, Chang WC (2004) TIBA affects the induction of direct somatic embryogenesis from leaf explants of Oncidium. PCTOC 79:315–320
Ckurshumova W, Berleth T (2015) Overcoming recalcitrance-auxin response factor functions in plant regeneration. Plant Signal Behav 10:e993293
Cueva-Agila AY, Medina J, Concia L, Cella R (2016) Effects of plant growth regulator, auxin polar transport inhibitors on somatic embryogenesis and CmSERK gene expression in Cattleya maxima (Lindl.). In: Mujib A (ed) Somatic embryogenesis in ornamentals and its applications. Springer, Delhi, pp 255–267
Dharmasiri N, Dharmasiri S, Estelle M (2005) The F-box protein TIR1 is an auxin receptor. Nature 435:441–445
Ding Z, Friml J (2010) Auxin regulates distal stem cell differentiation in Arabidopsis roots. PNAS 107:12046–12051
Dodeman VL, Ducreux G, Kreis M (1997) Zygotic embryogenesis versus somatic embryogenesis. J Exp Bot 48:1493–1509
Donner TJ, Sherr I, Scarpella E (2009) Regulation of preprocambial cell state acquisition by auxin signaling in Arabidopsis leaves. Development 136:3235–3246
Elhiti M, Stasolla C, Wang A (2013) Molecular regulation of plant somatic embryogenesis. In vitro Cell Dev Biol Plant 49:631–642
Gaj MD (2001) Direct somatic embryogenesis as a rapid and efficient system for in vitro regeneration of Arabidopsis thaliana. PCTOC 64:39–46
Gaj MD (2004) Factors influencing somatic embryogenesis induction and plant regeneration with particular reference to Arabidopsis thaliana (L.) Heynh. Plant Growth Regul 43:27–47
Gaj MD, Zhang S, Harada JJ, Lemaux PG (2005) Leafy cotyledon genes are essential for induction of somatic embryogenesis of Arabidopsis. Planta 222:977–988
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirement of suspension culture of soybean root cells. Exp Cell Res 50:151–158
Gliwicka M, Nowak K, Balazadeh S, Mueller-Roeber B, Gaj MD (2013) Extensive modulation of the transcription factor transcriptome during somatic embryogenesis in Arabidopsis thaliana. PLoS One 8:e69261
Gordon SP, Heisler MG, Reddy GV, Ohno C, Das P, Meyerowitz EM (2007) Pattern formation during de novo assembly of the Arabidopsis shoot meristem. Development 134:3539–3548
Guilfoyle TJ (2015) The PB1 domain in auxin response factor and Aux/IAA proteins: a versatile protein interaction module in the auxin response. Plant Cell 27:33–43
Guilfoyle TJ, Hagen G (2001) Auxin response factors. Plant Growth Reg 10:281–291
Guilfoyle TJ, Hagen G (2007) Auxin response factors. Curr Opin Plant Biol 10:453–460
Guilfoyle T, Hagen G, Ulmasov T, Murfett J (1998) How does auxin turn on genes? Plant Physiol 118:341–347
Hannah MA, Heyer AG, Hincha DK (2005) A global survey of gene regulation during cold acclimation in Arabidopsis thaliana. PLos Genet 1:e26
Hardtke CS, Berleth T (1998) The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J 17:1405–1411
Hardtke CS, Ckurshumova W, Vidaurre DP, Singh SA, Stamatiou G, Tiwari SB, Hagen G, Guilfoyle TJ, Berleth T (2004) Overlapping and non-redundant functions of the Arabidopsis auxin response factors MONOPTEROS and NONPHOTOTROPIC HYPOCOTYL4. Development 131:1089–1100
Horstman A, Willemsen V, Boutilier K, Heidstra R (2014) AINTEGUMENTA-LIKE proteins: hubs in a plethora of networks. Trends Plant Sci 19:146–157
Horstman A, Heidmann I, Weemen M, Angenent G, Boutilier K (2015) BABY BOOM and PLETHORA2 induce somatic embryogenesis in a dose-and context-dependent manner via the LAFL pathway. Dissertation, Wageningen University
Ikeuchi M, Ogawa Y, Iwase A, Sugimoto K (2016) Plant regeneration: cellular origins and molecular mechanisms. Development 143:1442–1451
Indoliya Y, Tiwari P, Chauhan AS, Goel R, Shri M, Bag SK, Chakrabarty D (2016) Decoding regulatory landscape of somatic embryogenesis reveals differential regulatory networks between japonica and indica rice subspecies. Sci Rep 6:1–15
Jain M, Khurana JP (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. Febs J 276:3148–3162
Karami O, Saidi A (2010) The molecular basis for stress-induced acquisition of somatic embryogenesis. Mol Biol Rep 37:2493–2507
Karami O, Aghavaisi B, Pour AM (2009) Molecular aspects of somatic-to-embryogenic transition in plants. J Biol Chem 2:177–190
Kieffer M, Neve J, Kepinski S (2010) Defining auxin response contexts in plant development. Curr Opin Plant Biol 13:12–20
Kraut M, Wójcikowska B, Ledwoń A, Gaj MD (2011) Immature zygotic embryo cultures of Arabidopsis—a model system for molecular studies on morphogenic pathways induced in vitro. Acta Biol Cracov Bot 53:59–67
Krogan NT, Berleth T (2015) The identification and characterization of specific ARF-Aux/IAA regulatory modules in plant growth and development. Plant Signal Behav 10:e992748
Krogan NT, Yin X, Ckurshumova W, Berleth T (2014) Distinct subclades of Aux/IAA genes are direct targets of ARF5/MP transcriptional regulation. New Phytol 204:474–483
Krogan NT, Marcos D, Weiner AI, Berleth T (2016) The auxin response factor MONOPTEROS controls meristem function and organogenesis in both the shoot and root through the direct regulation of PIN genes. New Phytol 212:42–50
Kurczyńska EU, Gaj MD, Ujczak U, Mazur E (2007) Histological analysis of direct somatic embryogenesis in Arabidopsis thaliana (L.) Heynh. Planta 226:619–628
Lau S, De Smet I, Kolb M, Meinhardt H, Jürgens G (2011) Auxin triggers a genetic switch. Nat Cell Biol 13:611–615
Ledwoń A, Gaj MD (2011) LEAFY COTYLEDON1, FUSCA3 expression and auxin treatment in relation to somatic embryogenesis induction in Arabidopsis. Plant Growth Regul 65:157–167
Lim PO, Lee IC, Kim J, Kim HJ, Ryu JS, Woo HR, Nam HG (2010) AUXIN RESPONSE FACTOR 2 (ARF2) plays a major role in regulating auxin-mediated leaf longevity. J Exp Bot 61:1419–1430
Lin Y, Lai Z (2013) Comparative analysis reveals dynamic changes in miRNAs and their targets and expression during somatic embryogenesis in longan (Dimocarpus longan Lour.). PLoS One 8:e60337
Lin Y, Lin L, Lai R, Liu W, Chen Y, Zhang Z, XuHan X, Lai Z (2015) MicroRNA390-directed TAS3 cleavage leads to the production of tasiRNA-ARF3/4 during somatic embryogenesis in Dimocarpus longan Lour. Front Plant Sci 6:1–15
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17:1360–1375
Mattsson J, Ckurshumova W, Berleth T (2003) Auxin signaling in Arabidopsis leaf vascular development. Plant Physiol 131:1327–1339
Michalczuk L, Ribnicky DM, Cooke TJ, Cohen JD (1992) Regulation of indole-3-acetic acid biosynthetic pathways in carrot cell cultures. Plant Physiol 100:1346–1353
Müller CJ, Valdés AE, Wang G, Ramachandran P, Beste L, Uddenberg D, Carlsbecker A (2015) PHABULOSA mediates an auxin signaling loop to regulate vascular patterning in Arabidopsis. Plant Physiol 170:956–970
Murashige T, Skoog FA (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Naser V, Shani E (2016) Auxin response under osmotic stress. Plant Mol Biol 91:661–672
Nowak K, Gaj MD (2016) Transcription factors in the regulation of somatic embryogenesis. In: Somatic embryogenesis: fundamental aspects and applications. Springer International Publishing, Switzerland, pp 53–79
Nowak K, Wójcikowska B, Gaj MD (2015) ERF022 impacts the induction of somatic embryogenesis in Arabidopsis through the ethylene-related pathway. Planta 241:967–985
Odat O, Gardiner J, Sawchuk MG, Verna C, Donner TJ, Scarpella E (2014) Characterization of an allelic series in the MONOPTEROS gene of Arabidopsis. Genesis 52:127–133
Oh E, Zhu JY, Bai MY, Arenhart RA, Sun Y, Wang ZY (2014) Cell elongation is regulated through a central circuit of interacting transcription factors in the Arabidopsis hypocotyl. eLife 3:e03031. doi:10.7554/eLife.03031
Okushima Y, Overvoorde PJ, Arima K, Alonso JM, Chan A, Chang C, Ecker JR, Hughes B, Lui A, Nguyen D, Onodera C, Quach H, Smith A, Yu G, Theologis A (2005) Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17:444–463
Ooi SE, Choo CN, Ishak Z, Ong-Abdullah M (2012) A candidate auxin-responsive expression marker gene, EgIAA9, for somatic embryogenesis in oil palm (Elaeis guineensis Jacq.). PCTOC 110:201–212
Prelich G (2012) Gene overexpression: uses, mechanisms, and interpretation. Genetics 190:841–854
Qiao M, Zhao Z, Song Y, Liu Z, Cao L, Yu Y, Li S, Xiang F (2012) Proper regeneration from in vitro cultured Arabidopsis thaliana requires the microRNA-directed action of an auxin response factor. Plant J 71:14–22
Rademacher EH, Moller B, Lokerse AS, Llavata-Peris CI, Berg W, Weijers D (2011) A cellular expression map of the Arabidopsis AUXIN RESPONSE FACTOR gene family. Plant J 68:597–606
Rademacher EH, Lokerse AS, Schlereth A, Llavata-Peris CI, Bayer M, Kientz M, Rios AF, Borst JW, Lukowitz W, Jürgens G, Weijers D (2012) Different auxin response machineries control distinct cell fates in the early plant embryo. Dev Cell 22:211–222
Reid K, Olsson N, Schlosser J, Peng F, Lund ST (2006) An optimized grapevine RNA isolation procedure and statistical determination of reference genes for real-time RT-PCR during berry development. BMC Plant Biol 6:27–38
Ribnicky DM, Cohen JD, Hu WS, Cooke TJ (2002) An auxin surge following fertilization in carrots: a mechanism for regulating plant totipotency. Planta 214:505–509
Riechmann JL, Heard J, Martin G, Reuber L, Jiang CZ, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu GL (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Schlereth A, Moller B, Liu W, Kientz M, Flipse J, Rademacher EH, Schmid M, Jurgens G, Weijers D (2010) MONOPTEROS controls embryonic root initiation by regulating a mobile transcription factor. Nature 464:913–916
Schruff MC, Spielman M, Tiwari S, Adams S, Fenby N, Scott RJ (2006) The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signaling, cell division, and the size of seeds and other organs. Development 133:251–261
Sharma KS, Millam S, Hedley PE, McNicol S, Bryan GJ (2008) Molecular regulation of somatic embryogenesis in potato: an auxin led perspective. Plant Mol Biol 68:185–201
Stasolla C, Belmonte MF, van Zyl L, Craig DL, Liu W, Yeung EC, Sederoff RR (2004) The effect of reduced glutathione on morphology and gene expression of white spruce (Picea glauca) somatic embryos. J Exp Bot 55:695–709
Su YH, Zhao XY, Liu YB, Zhang CL, O’Neill SD, Zhang XS (2009) Auxin-induced WUS expression is essential for embryonic stem cell renewal during somatic embryogenesis in Arabidopsis. Plant J 59:448–460
Szyrajew K, Bielewicz D, Dolata J, Wójcik AM, Nowak K, Szczygiel-Sommer A, Szweykowska-Kulinska Z, Jarmolowski A, Gaj MD (2017) microRNAs are intensively regulated during induction of somatic embryogenesis in Arabidopsis. Front Plant Sci 8:18. doi:10.3389/fpls.2017.00018
Tang X, Bian S, Tang M, Lu Q, Li S, Liu X, Tian G, Nguyen V, Tsang EW, Wang A, Rothstein SJ, Chen X, Cui Y (2012) MicroRNA-mediated repression of the seed maturation program during vegetative development in Arabidopsis. PLoS Genet 8:e1003091
Teale WD, Paponov IA, Palme K (2006) Auxin in action: signaling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7:847–859
Thellin O, Zorzi W, Lakaye B, De Borman B, Coumans B, Hennen G, Grisar T, Igout A, Heinen E (1999) Housekeeping genes as internal standards: use and limits. J Biotechnol 75:291–295
Thibaud-Nissen F, Shealy RT, Khanna A, Vodkin LO (2003) Clustering of microarray data reveals transcript patterns associated with somatic embryogenesis in soybean. Plant Physiol 132:118–136
Tian CE, Muto H, Higuchi K, Matamura T, Tatematsu K, Koshiba T, Yamamoto KT (2004) Disruption and overexpression of AUXIN RESPONSE FACTOR 8 gene of Arabidopsis affect hypocotyl elongation and root growth habit, indicating its possible involvement in auxin homeostasis in light condition. Plant J 40:333–343
Tiwari SB, Wang XJ, Hagen G, Guilfoyle TJ (2001) Aux/IAA proteins are active repressors and their stability and activity are modulated by auxin. Plant Cell 13:2809–2822
van Zyl L, Bozhkov PV, Chacham D, Sederoff RR, von Arnold S (2003) Up, down and up again is a signature global gene expression pattern at the beginning of gymnosperm embryogenesis. Gene Expr Patterns 3:83–91
Vernoux T, Brunoud G, Farcot E, Morin V, Van den Daele H, Legrand J, Oliva M, Das P, Larrieu A, Wells D, Guédon Y, Armitage L, Picard F, Guyomarch S, Cellier C, Parry G, Koumproglou R, Doonan JH, Estelle M, Godin C, Kepinski S, Bennett M, De Veylder L, Traas J (2011) The auxin signaling network translates dynamic input into robust patterning at the shoot apex. Mol Syst Biol 7:508
Vert G, Walcher CL, Chory J, Nemhauser JL (2008) Integration of auxin and brassinosteroid pathways by AUXIN RESPONSE FACTOR 2. PNAS 105:9829–9834
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17:2204–2216
Wang X, Niu QW, Teng C, Li C, Mu J, Chua NH, Zuo J (2009) Overexpression of PGA37/MYB118 and MYB115 promotes vegetative-to-embryonic transition in Arabidopsis. Cell Res 19:224–235
Wang Y, Deng D, Shi Y, Miao N, Bian Y, Yin Z (2012) Diversification, phylogeny and evolution of auxin response factor (ARF) family: insights gained from analyzing maize ARF genes. Mol Syst Biol 39:2401–2415
Weijers D, Wagner D (2016) Transcriptional responses to the auxin hormone. Annu Rev Plant Biol 67:539–574
Weijers D, Benkova E, Jäger KE, Schlereth A, Hamann T, Kientz M, Wilmoth JC, Jürgens G (2005) Developmental specificity of auxin response by pairs of ARF and Aux/IAA transcriptional regulators. EMBO J 24:1874–1885
Wenzel CL, Schuetz M, Yu Q, Mattsson J (2007) Dynamics of MONOPTEROS and PIN-FORMED1 expression during leaf vein pattern formation in Arabidopsis thaliana. Plant J 49:387–398
Wickramasuriya AM, Dunwell JM (2015) Global scale transcriptome analysis of Arabidopsis embryogenesis in vitro. BMC Genomics 16:1
Winkelmann T (2016) Somatic versus zygotic embryogenesis: learning from seeds. In: Germana MA, Lambardi M (eds) In vitro embryogenesis in higher plants. Springer, New York, pp 25–46
Wójcikowska B, Gaj MD (2015) LEAFY COTYLEDON2-mediated control of the endogenous hormone content: implications for the induction of somatic embryogenesis in Arabidopsis. PCTOC 121:255–258
Wójcikowska B, Jaskóła K, Gąsiorek P, Meus M, Nowak K, Gaj MD (2013) LEAFY COTYLEDON2 (LEC2) promotes embryogenic induction in somatic tissues of Arabidopsis, via YUCCA-mediated auxin biosynthesis. Planta 238:425–440
Xiang D, Venglat P, Tibiche C, Yang H, Risseeuw E, Cao Y, Babic V, Cloutier M, Keller W, Wang E, Selvaraj G, Datla R (2011) Genome-wide analysis reveals gene expression and metabolic network dynamics during embryo development in Arabidopsis. Plant Physiol 156:346–356
Xu Z, Zhang C, Zhang X, Liu C, Wu Z, Yang Z, Zhou K, Yang X, Li F (2013) Transcriptome profiling reveals auxin and cytokinin regulating somatic embryogenesis in different sister lines of cotton cultivar CCRI24. J Integr Plant Biol 55:631–642
Yamaguchi N, Jeong CW, Nole-Wilson S, Krizek BA, Wagner D (2016) AINTEGUMENTA and AINTEGUMENTA-LIKE6/PLETHORA3 induce LEAFY expression in response to auxin to promote the onset of flower formation in Arabidopsis. Plant Physiol 170:283–293
Yang X, Zhang X, Yuan D, Jin F, Zhang Y, Xu J (2012) Transcript profiling reveals complex auxin signaling pathway and transcription regulation involved in dedifferentiation and redifferentiation during somatic embryogenesis in cotton. BMC Plant Biol 12:1
Zavattieri MA, Frederico AM, Lima M, Sabino R, Arnholdt-Schmitt B (2010) Induction of somatic embryogenesis as an example of stress-related plant reactions. Electron J Biotech 13:12–13
Zhang J, Zhang S, Han S, Wu T, Li X, Li W, Qi L (2012) Genome-wide identification of microRNAs in larch and stage-specific modulation of 11 conserved microRNAs and their targets during somatic embryogenesis. Planta 236:647–657
Zhao Z, Andersen SU, Ljung K, Dolezal K, Miotk A, Schultheiss SJ, Lohmann JU (2010) Hormonal control of the shoot stem-cell niche. Nature 465:1089–1092
Zheng Q, Zheng Y, Ji H, Burnie W, Perry SE (2016) Gene regulation by the AGL15 transcription factor reveals hormone interactions in somatic embryogenesis. Plant Physiol 172:2374–2387
Zimmerman JL (1993) Somatic embryogenesis: a model for early development in higher plants. Plant Cell 5:1411–1423
Zuo J, Niu QW, Frugis G, Chau NH (2002) The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J 30:349–359
Acknowledgements
This work was supported by Grant 2012/05/N/NZ2/01676 from the National Science Centre Poland. Author BW was supported by a scholarship within the framework of the UPGOW project of the European Social Fund, implemented at the University of Silesia, Katowice, Poland and ETIUDA Grant 2013/08/T/NZ2/00800 of the National Science Centre Poland. We thank Małgorzata Widuch for her excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Ray J. Rose.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2017_2114_MOESM1_ESM.xlsx
Table S1. The insertional mutants, reporter, overexpressor lines, the primers used in PCR analysis and a checklist outlining the RNA to RT-qPCR quality/methodology. (xlsx 28 kb)
299_2017_2114_MOESM4_ESM.tif
Figure S1. Qualitative RT-PCR analysis of ARF transcripts at different time points (0, 3, 5, 10, 15d) of the embryogenic Col-0 cultures induced on an E5 medium. RNA samples for the analysis were isolated from 0-, 3-, 5-, 10-, 15-day-old cultures. The TIN gene was used as the control for cDNA synthesis (n = 3). (TIFF 1024 kb)
299_2017_2114_MOESM5_ESM.tif
Figure S2. Transcript levels of ARF genes during the SE process that were analyzed using the RNA samples isolated from 0-, 3-, 5-, 10-, 15-day-old cultures. The expression level was calculated using the 45-Ct method. EF1α and TIN genes were used as the control for cDNA synthesis. Means and SD for three biological replicates are shown. (TIFF 4061 kb)
299_2017_2114_MOESM6_ESM.tif
Figure S3. Stable expression of ARF7 (a), ARF9 (b) and ARF19 (c) genes during the SE process. IZE explants of Col-0 were cultured on an E5 medium and the tissue was sampled on the 0, 3, 5, 10, 15d of the culture. Relative transcript level was normalized to the internal control (TIN gene) and calibrated to the 0d culture. * – expression level significantly different to that observed at 0d at P<0.05. Means and SD for three biological replicates are shown. (TIFF 3275 kb)
299_2017_2114_MOESM7_ESM.tif
Figure S4. Qualitative RT-PCR analysis of ARF transcripts in Col-0 IZE explants cultured on CIM/SIM medium to induce shoot organogenesis (ORG). RNA samples for the analysis were isolated on the 0, 3, 5, 10, 15d of the culture. The TIN gene was used as the control for cDNA synthesis (n = 3). (TIFF 1047 kb)
299_2017_2114_MOESM8_ESM.tif
Figure S5. Expression of LEC2 during the SE process that was induced in the culture of the arf5 mutant. IZE explants were cultured on an E5 medium and the tissue was sampled on the 0, 3, 5, 10, 15d. The relative transcript level was normalized to the internal control (TIN gene) and calibrated to the WT (Col-0) culture. * – expression level significantly different from that observed in the WT culture of the same age at P<0.05. Means and SD for three biological replicates are shown. (TIFF 4865 kb)
299_2017_2114_MOESM9_ESM.tif
Figure S6. A model of the ARF5-controlled pathways that are possibly involved in SE induction. It is proposed that auxin-stimulated ARF5 regulates the expression of numerous genes that are involved in SE induction including LEC2 (LEAFY COTYLEDON2) (presented results), which is an activator of the YUC1, YUC4, YUC10 (YUCCA) genes that are involved in auxin biosynthesis in SE (Wójcikowska et al. 2013) and PIN1 (PIN-FORMED1), which is involved in polar auxin transport and positively controls SE (Su et al. 2009). In addition, the involvement of other ARF5-controlled TFs, including ATHB8 (HOMEOBOX GENE8) and TMO3, TMO5, TMO6, TMO7 (TARGET OF MONOPTEROS) (Schlereth et al. 2010) might also be considered in SE (Gliwicka et al. 2013). Aux/IAA genes (IAA29, IAA30, IAA31), which interact with ARF5 via the regulatory feedback loop (Krogan and Berleth 2015) have also been indicated as contributing to SE induction (Gliwicka et al. 2013). Solid line – experimentally confirmed interaction. Dashed line – interaction that needs to be confirmed.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wójcikowska, B., Gaj, M.D. Expression profiling of AUXIN RESPONSE FACTOR genes during somatic embryogenesis induction in Arabidopsis . Plant Cell Rep 36, 843–858 (2017). https://doi.org/10.1007/s00299-017-2114-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-017-2114-3