Abstract
Objectives
The accurate detection and precise segmentation of lung nodules on computed tomography are key prerequisites for early diagnosis and appropriate treatment of lung cancer. This study was designed to compare detection and segmentation methods for pulmonary nodules using deep-learning techniques to fill methodological gaps and biases in the existing literature.
Methods
This study utilized a systematic review with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines, searching PubMed, Embase, Web of Science Core Collection, and the Cochrane Library databases up to May 10, 2023. The Quality Assessment of Diagnostic Accuracy Studies 2 criteria was used to assess the risk of bias and was adjusted with the Checklist for Artificial Intelligence in Medical Imaging. The study analyzed and extracted model performance, data sources, and task-focus information.
Results
After screening, we included nine studies meeting our inclusion criteria. These studies were published between 2019 and 2023 and predominantly used public datasets, with the Lung Image Database Consortium Image Collection and Image Database Resource Initiative and Lung Nodule Analysis 2016 being the most common. The studies focused on detection, segmentation, and other tasks, primarily utilizing Convolutional Neural Networks for model development. Performance evaluation covered multiple metrics, including sensitivity and the Dice coefficient.
Conclusions
This study highlights the potential power of deep learning in lung nodule detection and segmentation. It underscores the importance of standardized data processing, code and data sharing, the value of external test datasets, and the need to balance model complexity and efficiency in future research.
Clinical relevance statement
Deep learning demonstrates significant promise in autonomously detecting and segmenting pulmonary nodules. Future research should address methodological shortcomings and variability to enhance its clinical utility.
Key Points
-
Deep learning shows potential in the detection and segmentation of pulmonary nodules.
-
There are methodological gaps and biases present in the existing literature.
-
Factors such as external validation and transparency affect the clinical application.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
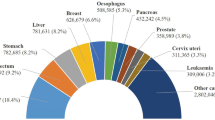
Lung cancer is responsible for 18.4% of all cancer deaths globally and is the leading cause of cancer-related mortality worldwide [1]. Pulmonary nodules, often an early indicator of lung cancer, do not always indicate malignancy [2]. Early detection and precise segmentation of lung nodules are crucial for accurately diagnosing and treating lung cancer [3,4,5,6].
Chest computed tomography (CT) is widely employed for pulmonary nodule detection [7, 8]. However, a single CT scan can yield hundreds of images, demanding substantial time and effort for radiologists to analyze [9]. Prolonged and arduous radiological analysis can compromise diagnostic accuracy [10, 11]. Image segmentation enables the analysis of phenotypic characteristics such as shape, size, and texture of regions of interest [12]. This examination can help determine the malignant potential. Such information is valuable for developing diagnostic and predictive models for personalized medicine in lung cancer research, leading to advancements in precision medicine within this field [13,14,15,16]. However, Image segmentation is predominantly a manual or semi-automated process, requiring labor while being prone to inter- and intra-rater discrepancies [17]. Consequently, the manual detection and segmentation of suspicious lesion regions in CT scans represent a laborious task for radiologists. In contrast, automatic detection and segmentation provide results directly from the input image, offering simplicity, speed, efficiency, and removing operator bias during segmentation [18].
Deep learning (DL), a subset of artificial intelligence (AI), leverages high-sensitivity detection, multifaceted information mining, and high-throughput computing, rendering it immensely promising in medicine [19,20,21,22,23]. DL has found applications in lung cancer imaging, encompassing tumor detection [24, 25], CT image segmentation [26], and classification [27, 28]. Nevertheless, most studies have focused on individual tasks, such as detection, segmentation, classification, or prognosis, and have exhibited varying degrees of quality. Additionally, no known analysis has been conducted on the quality of deep-learning studies for simultaneous pulmonary nodule detection and segmentation.
Therefore, this paper aims to conduct a comprehensive analysis and comparison of various published methods for lung nodule detection and segmentation. It seeks to summarize relevant publications to identify methodological gaps and biases. This paper will also serve as a reference for other researchers, enabling them to identify research gaps that require further investigation.
Materials and methods
The systematic review followed the guidelines of the Preferred Reporting Items for Systematic Reviews and Meta-Analysis (PRISMA) [29]. The review was registered on PROSPERO [30] before initiation (registration No. CRD < 42023454274 >).
Search strategy
This group searched PubMed, Embase, Web of Science Core Collection, and Cochrane Library databases up to May 10, 2023, to identify studies utilizing deep-learning techniques for detecting and segmenting lung nodules and lung cancer. Additionally, we manually searched the reference lists of relevant literature and included articles. Our search terms included “lung neoplasms,” “lung cancer,” “lung nodule,” solitary pulmonary nodule,” “multiple pulmonary nodules,” “artificial intelligence,” “deep learning,” and “image segmentation.” The detailed search criteria were described in the supplementary materials. The retrieval was performed without language and date restrictions.
Study selection
Our study encompassed original research articles focusing on detecting and segmenting lung cancer or lung nodules using deep-learning techniques. Eligibility criteria included: (1) patients with lung cancer or lung nodules, (2) development or validation of deep-learning models for the detection and segmentation of lung nodules or cancer in CT images, and (3) the application of DL in the context of lung nodule or cancer detection and segmentation. We excluded case studies, editorials, letters, review articles, and conference abstracts. Additionally, non-deep-learning models were excluded, as well as studies exclusively employing animals or computer simulations.
Data extraction
Two authors (Chuan G. and L.W.) independently extracted data and discussed any discrepancies. The data extraction process focused on the study parameters: the first author’s name, the year of publication, the source of the dataset, the ground truth, the focused task, and the time used for detection and segmentation. The DL parameters were also considered, including the DL algorithm, data augmentation techniques, performance measures, code/data availability, and external validation or cross-validation.
Risk of bias assessment
Data from each included article underwent independent assessment by two radiologists (Chuan G. and L.W.). We employed the Quality Assessment of Diagnostic Accuracy Studies tool-2 (QUADAS-2) framework [31] to assess the risk of bias and applicability for each selected study, which was revised to incorporate elements from the Checklist for Artificial Intelligence in Medical Imaging (CLAIM) [32]. The detailed evaluation criteria were provided in supplementary materials. Conflict resolution occurred through consensus with a third reviewer (Chen G).
Results
Study selection
A total of 6793 articles were retrieved for this systematic review, and two researchers conducted a full-text review of 109 articles, excluding 16 records classified as reviews, case reports, meeting abstracts, or animal research. Additionally, 82 articles were excluded for solely focusing on segmentation without detection. Two other articles were excluded as they targeted the pulmonary lobes instead of nodules. Finally, nine studies were included in this systematic review [33,34,35,36,37,38,39,40,41] (Fig. 1). Detailed characteristics of the included articles are presented in Tables 1 and 2. A summary of the methodological overview of the included articles is presented in Supplementary Table 1.
Study characteristics
Table 1 summarizes the characteristics of the nine studies included in this analysis. These studies were published between 2019 and 2023, with one (1/9, 11%) [39] being a prospective in-silico clinical trial, while the rest (8/9, 89%) were retrospective in design. Among the nine articles, eight (8/9, 89%) [33,34,35,36,37,38, 40, 41] made use of publicly available datasets, with the Lung Image Database Consortium Image Collection and Image Database Resource Initiative (LIDC-IDRI) [42] dataset and its subsidiary, Lung Nodule Analysis 16 (LUNA16) [43], emerging as the most frequently employed datasets. Primary tasks addressed by most studies [33,34,35,36,37,38,39,40,41] encompassed detection and segmentation, while two (2/9, 22%) studies [34, 41] included three-dimensional (3D) visualization reconstruction. Two (2/9, 22%) [35, 40] studies incorporated classification, while another article (1/9, 11%) [39] incorporated prognosis prediction. Out of the nine studies, eight (8/9, 89%) [33,34,35,36,37,38, 40, 41] focused on the development of DL models or architectures, while one study (1/9, 11%) [39] focused on developing and building an application or a working prototype. Furthermore, six (2/3, 67%) [33, 34, 37, 39,40,41] of the nine studies reported the inference time for detection and segmentation tasks.
Ground truth definition
The definition of ground truth was ambiguous in three articles (1/3, 33%) [33, 37, 38]. One (1/9, 11%) [34] article defined raw images labeled with the label tool as ground truth, while another (1/9, 11%) [35] research mentioned the utilization of annotations in XML files to generate segmentation ground truth masks. One (1/9, 11%) article [36] considered intersection regions annotated by at least three radiologists as the ground truth. Similarly, another paper (1/9, 11%) [39] treated the contours segmented by experts as ground truth. One (1/9, 11%) article [40] also referred to the ground truth determined by standard physician annotations. Lastly, one article (1/9, 11%) [41] defined the ground truth as being manually prepared by expert clinicians and reviewed by thoracic surgeons.
External validation or cross-validation and data augmentation
Among the nine studies, two (2/9, 22%) [34, 39] conducted external validation, while one (1/9, 11%) [36] employed 5-fold cross-validation, and another (1/9, 11%) [38] utilized 10-fold cross-validation. Out of the nine articles, eight (8/9, 89%) [34,35,36,37,38,39,40,41] employed data augmentation technology, with only one (1/9, 11%) [33] article not utilizing it.
Deep-learning algorithms
A diverse range of deep-learning algorithms incorporated by the articles included in the systematic review can be categorized into distinct types, including models tailored for object classification and detection, such as densely connected convolutional networks (DenseNet) [36], YOLOv4 and YOLOv5 [40, 41], as well as models designed for image segmentation tasks, like U-Net [37,38,39,40], Fast region-based convolutional neural network (R-CNN) [33, 37], and Mask R-CNN [34]. Furthermore, certain studies have employed general-purpose architectures, such as region-based convolutional neural networks (FCN) and feature pyramid networks (FPN) [33, 34], for various image analysis applications.
Code and Data Availability
Among the nine studies, 7 (7/9, 78%) [33,34,35,36,37,38, 40] used open-source public datasets. In one (1/9, 11%) study [39], a portion of the data was accessible; in another (1/9, 11%) study [41], it was mentioned that the data would be provided upon request. Out of the nine publications, only one (1/9, 11%) [39] author has uploaded the code to a public repository, while the remainder (8/9, 89%) does not mention the availability of code.
Performance metrics
Table 3 presents a summary of the performance metrics observed in the studies. The performance indicators for nodule detection include accuracy (ACC), sensitivity (SEN), precision, F-score, and area under the receiver operating characteristic curve (AUC). As for nodule segmentation, the reported performance metrics encompassed dice similar coefficient (DSC), Jaccard index (JI), Hausdorff distance (HD), and Intersection over Union (IOU). The accuracy of pulmonary nodule detection in these studies exceeded 90% [33, 36, 37], and the detection sensitivity was as high as 97% [39, 41]. In addition, the DSC of pulmonary nodule segmentation can be as high as 0.93 [38].
Risk of bias and quality assessment
Table 4 and Fig. 2 present the quality assessment of the included studies. Among the seven items of the QUADAS-2 tool, the most common item that could have been improved was the applicability concern regarding patient selection. In terms of the risk of bias, 7 out of 9 (7/9, 78%) studies were determined to have a low risk of bias in the domain of “index test” [33,34,35,36,37, 39, 40], and 8 out of 9 (8/9, 89%) studies were judged to have a low risk of bias in terms of “flow and timing” [33,34,35,36,37,38, 40, 41]. Additionally, 6 out of 9 (2/3, 67%) studies were found to have a low risk of bias in terms of “patient selection” [33,34,35,36,37, 40]. However, only 44% of the studies (4/9) [36, 39,40,41] were considered to have a low risk of bias in the “reference standard” domain. Only two (2/9, 22%) articles [36, 40] were determined to have a low risk of bias in all four domains.
Graphical representation of the Quality Assessment of Diagnostic Accuracy Studies version 2 (QUADAS-2) assessment results of included studies. Stacked bar charts show the results of the quality assessment for risk of bias and applicability of included studies. QUADAS-2 scores for methodologic study quality are expressed as the percentage of studies that met each criterion. For each quality domain, the proportion of included studies that were determined to have low, high, or unclear risk of bias and/or concerns regarding applicability is displayed in green, orange, and blue, respectively. QUADAS-2, Quality Assessment of Diagnostic Accuracy Studies 2
Discussion
This systematic review analyzed nine pertinent articles from four prominent databases and found that DL shows significant potential in detecting and segmenting pulmonary nodules concurrently instead of being restricted to a singular task [44,45,46,47,48,49] (Fig. 3). Nevertheless, the analysis identified methodological shortcomings and variations among the studies included.
Accurate detection is vital for precise segmentation. Some studies have successfully implemented multi-task learning [50,51,52], demonstrating the possibility of solving both aspects simultaneously. Human experts’ manual detection and annotation of lung cancer nodules is a time-consuming and inconsistent process [17, 53]. In contrast, deep-learning algorithms have demonstrated the ability to perform the same task rapidly and consistently [33, 34, 37, 39,40,41].
The quality of the generated automatic segmentation was evaluated against the corresponding reference segmentation, known as ground truths, demonstrating adherence to the requirements and best practices outlined in the AI Checklist [32]. The requirements included providing a detailed definition of the reference standard and a rationale for its selection [54, 55]. It was also important to mention the source of the ground truth annotation and the qualifications of the annotator. Additionally, it was necessary to measure annotation tools and internal variability and propose to mitigate and resolve any differences. However, the methodologies employed in the reviewed studies varied and were not standardized. Although four studies [36, 39,40,41] mentioned the involvement of physicians or experts in providing basic facts, most studies did not specify the professional background of these experts, such as whether they were experienced radiologists, trainees, or other non-clinical researchers. Most studies did not clearly state the number of relevant experts involved [33,34,35, 37,38,39,40] or the annotation tool used [33, 35,36,37,38, 40, 41]. Differences in the focus of the publishing journal and the authors’ background, such as computing researchers versus clinicians, may contribute to this variation. Furthermore, certain studies utilized public datasets where the ground truth is established by other research teams or experts, limiting the authors’ ability to provide detailed explanations.
Eight out of nine studies utilized public datasets, the most frequently used ones being the LIDC-IDRI [42] and the LUNA16 dataset [43]. Public datasets offer abundant annotated data for training deep-learning models and enable researchers to validate and compare algorithms, enhancing research reproducibility [56, 57]. However, both LIDC/IDRI and LUNA16 originated from the United States and may not fully represent the global population. Additionally, these datasets often have an unbalanced male:female ratio, potentially hindering comprehensive gender-based analyses. While these datasets contain both standard-dose diagnostic and low-dose CT scans for lung cancer screening, the latter may compromise image quality and nodule contrast, presenting challenges in nodule detection and segmentation. The limitations of public datasets need to be carefully considered during model development and research to ensure the quality and representativeness of the data.
Furthermore, all studies have embraced convolutional neural networks (CNNs) and their derivatives, progressively enhancing network depth and complexity to bolster feature extraction capabilities [33,34,35,36,37,38,39,40,41]. Escalating model depth necessitates augmented computational resources, leading to longer training durations [58, 59]. To mitigate prolonged training, some algorithms have resorted to strategies that entail sacrificing substantial graphics processing unit (GPU) memory [60, 61]. However, this approach is unsustainable in the long run. Therefore, balancing network design, computation time, and cost is imperative.
Despite the growing interest in open science in current scientific research, only one study [39] has made its code openly accessible. Many research teams are reluctant to share their project code because of concerns about intellectual property and commercial interests. However, sharing project code can enhance the reproducibility of research findings and facilitate the validation of existing results, ultimately fostering collaboration within the scientific research community, and advancing the frontiers of knowledge.
Only two studies [34, 39] in our research utilized external test datasets, highlighting the inherent challenge of obtaining data from external collections. Some strategies have been employed to address this issue. For instance, two studies [36, 38] employed cross-validation methods, partially compensating for the lack of external testing datasets [62]. However, the absence of external validation datasets remains a significant limitation for the clinical applicability of the developed models [63]. This limitation arises from the inability of most studies to assess the risk of overfitting. Cross-validation helps mitigate this by evaluating model performance and detecting overfitting within existing data through repeated training and validation on subsets. However, its computational cost is high and there are problems such as sample imbalance. In addition, its results may be affected by the randomness of data partitioning. Different data partitioning may lead to different model performance evaluation results, thus affecting the accurate evaluation of the model. A combination of cross-validation and external validation was essential to achieve a more comprehensive and reliable evaluation of deep-learning models. Only one [33] article did not employ data augmentation techniques. Data augmentation is a highly effective method for enhancing data heterogeneity, preventing overfitting, and improving the robustness of CNN networks [64, 65].
The included studies have mostly concentrated on developing DL models and architectures [33,34,35,36,37,38, 40, 41]. However, only one study has examined the clinical application of these techniques [39]. This indicated the limited experience of non-medical researchers in selecting clinically relevant outcomes and their limited applicability in clinical practice.
Our study included only nine articles, while a wide variety of chest computer-aided diagnosis (CAD) systems are currently available on the market [66]. This disparity in numbers can be traced back to the progression of medical imaging CAD systems, transitioning from basic image-processing techniques to more advanced machine learning and deep-learning algorithms [67]. While some systems in the market still rely on traditional image-processing and machine-learning methods, the incorporation of DL has significantly enhanced the accuracy and efficiency of CAD systems [68]. Commercializing CAD systems involves various manufacturers and development teams, each capable of creating unique chest CAD systems based on their research and technological advancements [69]. These systems may consist of individual detection or segmentation modules customized to meet specific user requirements, leading to a diverse range of systems despite the limited number of articles.
This systematic review has several notable limitations. Firstly, the number of eligible studies was relatively small. Secondly, a meta-analysis of pooled outcomes could not be conducted as the median and standard deviation of the five articles were not provided in the results. Thirdly, the evaluation did not include individuals from engineering backgrounds, which may introduce a certain level of professional bias. Additionally, by excluding articles published in conference proceedings, there is a chance of excluding promising methods for lung nodule detection and segmentation.
Conclusions
In conclusion, this systematic review emphasizes the increasing significance of DL in detecting and segmenting lung nodules, which are crucial for early lung cancer diagnosis. Ensembles of deep-learning models demonstrate considerable potential in expediting the detection and segmentation processes, offering substantial advantages in clinical practice. However, certain challenges persist, including the need for diverse models, external validation, efficiency, and transparency. Future endeavors should address these challenges to foster the advancement of DL in lung cancer imaging, ultimately enhancing the accuracy and efficiency of early diagnosis.
Availability of data and materials
All data generated or analysed during this study are included in this published article [and its supplementary information files].
Abbreviations
- ACC:
-
Accuracy
- AI:
-
Artificial intelligence
- AUC:
-
Area under the receiver operating characteristic curve
- CAD:
-
Computer-aided diagnosis
- CNN:
-
Convolutional neural network
- CNNS:
-
Convolutional neural networks
- CT:
-
Computed tomography
- DenseNet:
-
Densely connected convolutional network
- DL:
-
Deep learning
- DSC:
-
Dice similar coefficient
- GPU:
-
Graphics processing unit
- IOU:
-
Intersection over Union
- JI:
-
Jaccard index
- LIDC-IDRI:
-
The Lung Image Database Consortium Image Collection and Image Database Resource Initiative
- LUNA16:
-
Lung Nodule Analysis 2016
- QUADAS-2:
-
Quality Assessment of Diagnostic Accuracy Studies 2
- R-CNN:
-
Region-based convolutional neural network
- SEN:
-
Sensitivity
References
Sung H, Ferlay J, Siegel RL et al (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Reeves AP, Chan AB, Yankelevitz DF et al (2006) On measuring the change in size of pulmonary nodules. IEEE Trans Med Imaging 25:435–450. https://doi.org/10.1109/TMI.2006.871548
Okazaki S, Shibuya K, Takura T et al (2022) Cost-effectiveness of carbon-ion radiotherapy versus stereotactic body radiotherapy for non-small-cell lung cancer. Cancer Sci 113:674–683. https://doi.org/10.1111/cas.15216
Uramoto H, Tanaka F (2014) Recurrence after surgery in patients with NSCLC. Transl Lung Cancer Res 3:242–249. https://doi.org/10.3978/j.issn.2218-6751.2013.12.05
Henschke CI, McCauley DI, Yankelevitz DF et al (1999) Early Lung Cancer Action Project: overall design and findings from baseline screening. Lancet 354:99–105. https://doi.org/10.1016/S0140-6736(99)06093-6
Jones GS, Baldwin DR (2018) Recent advances in the management of lung cancer. Clin Med 18:s41–s46. https://doi.org/10.7861/clinmedicine.18-2-s41
Libby DM, Smith JP, Altorki NK et al (2004) Managing the small pulmonary nodule discovered by CT. Chest 125:1522–1529. https://doi.org/10.1378/chest.125.4.1522
National Lung Screening Trial Research Team, Aberle DR, Adams AM et al (2011) Reduced lung-cancer mortality with low-dose computed tomographic screening. N Engl J Med 365:395–409. https://doi.org/10.1056/NEJMoa1102873
Gurcan MN, Sahiner B, Petrick N et al (2002) Lung nodule detection on thoracic computed tomography images: preliminary evaluation of a computer-aided diagnosis system. Med Phys 29:2552–2558. https://doi.org/10.1118/1.1515762
Nihashi T, Ishigaki T, Satake H et al (2019) Monitoring of fatigue in radiologists during prolonged image interpretation using fNIRS. Jpn J Radiol 37:437–448. https://doi.org/10.1007/s11604-019-00826-2
Taylor-Phillips S, Stinton C (2019) Fatigue in radiology: a fertile area for future research. Br J Radiol 92:20190043. https://doi.org/10.1259/bjr.20190043
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Zhang R, Wei Y, Wang D, et al (2023) Deep learning for malignancy risk estimation of incidental sub-centimeter pulmonary nodules on CT images. Eur Radiol 10.1007/s00330-023-10518–1. https://doi.org/10.1007/s00330-023-10518-1
Maldonado F, Varghese C, Rajagopalan S et al (2021) Validation of the BRODERS classifier (benign versus aggressive nodule evaluation using radiomic stratification), a novel HRCT-based radiomic classifier for indeterminate pulmonary nodules. Eur Respir J 57:2002485. https://doi.org/10.1183/13993003.02485-2020
Liu A, Wang Z, Yang Y et al (2020) Preoperative diagnosis of malignant pulmonary nodules in lung cancer screening with a radiomics nomogram. Cancer Commun 40:16–24. https://doi.org/10.1002/cac2.12002
Saied M, Raafat M, Yehia S, Khalil MM (2023) Efficient pulmonary nodules classification using radiomics and different artificial intelligence strategies. Insights Imaging 14:91. https://doi.org/10.1186/s13244-023-01441-6
Erasmus JJ, Gladish GW, Broemeling L et al (2003) Interobserver and intraobserver variability in measurement of non-small-cell carcinoma lung lesions: implications for assessment of tumor response. J Clin Oncol 21:2574–2582. https://doi.org/10.1200/JCO.2003.01.144
de Margerie-Mellon C, Chassagnon G (2023) Artificial intelligence: a critical review of applications for lung nodule and lung cancer. Diagn Inter Imaging 104:11–17. https://doi.org/10.1016/j.diii.2022.11.007
Jiang B, Li N, Shi X et al (2022) Deep learning reconstruction shows better lung nodule detection for ultra-low-dose chest CT. Radiology 303:202–212. https://doi.org/10.1148/radiol.210551
Deng K, Wang L, Liu Y et al (2022) A deep learning-based system for survival benefit prediction of tyrosine kinase inhibitors and immune checkpoint inhibitors in stage IV non-small cell lung cancer patients: a multicenter, prognostic study. EClinicalMedicine 51:101541. https://doi.org/10.1016/j.eclinm.2022.101541
Xu Y, Hosny A, Zeleznik R et al (2019) Deep learning predicts lung cancer treatment response from serial medical imaging. Clin Cancer Res 25:3266–3275. https://doi.org/10.1158/1078-0432.CCR-18-2495
Coiera E (2018) The fate of medicine in the time of AI. Lancet 392:2331–2332. https://doi.org/10.1016/S0140-6736(18)31925-1
Kleppe A, Skrede O-J, De Raedt S et al (2021) Designing deep learning studies in cancer diagnostics. Nat Rev Cancer 21:199–211. https://doi.org/10.1038/s41568-020-00327-9
Xu J, Ren H, Cai S, Zhang X (2023) An improved faster R-CNN algorithm for assisted detection of lung nodules. Comput Biol Med 153:106470. https://doi.org/10.1016/j.compbiomed.2022.106470
Liu W, Liu X, Li H et al (2021) Integrating lung parenchyma segmentation and nodule detection with deep multi-task learning. IEEE J Biomed Health Inf 25:3073–3081. https://doi.org/10.1109/JBHI.2021.3053023
Wang S, Zhou M, Liu Z et al (2017) Central focused convolutional neural networks: Developing a data-driven model for lung nodule segmentation. Med Image Anal 40:172–183. https://doi.org/10.1016/j.media.2017.06.014
Zhou J, Hu B, Feng W et al (2023) An ensemble deep learning model for risk stratification of invasive lung adenocarcinoma using thin-slice CT. NPJ Digit Med 6:119. https://doi.org/10.1038/s41746-023-00866-z
Wang D, Zhang T, Li M et al (2021) 3D deep learning based classification of pulmonary ground glass opacity nodules with automatic segmentation. Comput Med Imaging Graph 88:101814. https://doi.org/10.1016/j.compmedimag.2020.101814
Page MJ, McKenzie JE, Bossuyt PM et al (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. Syst Rev 10:89. https://doi.org/10.1186/s13643-021-01626-4
Booth A, Clarke M, Ghersi D et al (2011) An international registry of systematic-review protocols. Lancet 377:108–109. https://doi.org/10.1016/S0140-6736(10)60903-8
Whiting PF, Rutjes AWS, Westwood ME et al (2011) QUADAS-2: a revised tool for the Quality Assessment of Diagnostic Accuracy Studies. Ann Intern Med 155:529–536. https://doi.org/10.7326/0003-4819-155-8-201110180-00009
Mongan J, Moy L, Kahn CE Jr (2020) Checklist for Artificial Intelligence in Medical Imaging (CLAIM): a guide for authors and reviewers. Radiol Artif Intell 2:e200029. https://doi.org/10.1148/ryai.2020200029
Huang X, Sun W, Tseng T-LB et al (2019) Fast and fully-automated detection and segmentation of pulmonary nodules in thoracic CT scans using deep convolutional neural networks. Comput Med Imaging Graph 74:25–36. https://doi.org/10.1016/j.compmedimag.2019.02.003
Cai L, Long T, Dai Y, Huang Y (2020) Mask R-CNN-based detection and segmentation for pulmonary nodule 3D visualization diagnosis. IEEE Access 8:44400–44409. https://doi.org/10.1109/access.2020.2976432
Dutande P, Baid U, Talbar S (2021) LNCDS: A 2D-3D cascaded CNN approach for lung nodule classification, detection and segmentation. Biomed Signal Process Control 67:102527. https://doi.org/10.1016/j.bspc.2021.102527
Zhang X, Li S, Zhang B, Dong J, Zhao S, Liu X (2021) Automatic detection and segmentation of lung nodules in different locations from CT images based on adaptive α‐hull algorithm and DenseNet convolutional network. Int J Imaging Syst Technol 31:1882–1893. https://doi.org/10.1002/ima.22580
Banu SF, Sarker MDMK, Abdel-Nasser M et al (2021) AWEU-Net: an attention-aware weight excitation U-Net for lung nodule segmentation. Appl Sci 11:10132. https://doi.org/10.3390/app112110132
Hesamian MH, Jia W, He X et al (2020) Synthetic CT images for semi-sequential detection and segmentation of lung nodules. Appl Intell 51:1616–1628. https://doi.org/10.1007/s10489-020-01914-x
Primakov SP, Ibrahim A, van Timmeren JE et al (2022) Automated detection and segmentation of non-small cell lung cancer computed tomography images. Nat Commun 13:3423. https://doi.org/10.1038/s41467-022-30841-3
Zhou Z, Gou F, Tan Y, Wu J (2022) A cascaded multi-stage framework for automatic detection and segmentation of pulmonary nodules in developing countries. IEEE J Biomed Health Inf 26:5619–5630. https://doi.org/10.1109/JBHI.2022.3198509
Dlamini S, Chen Y-H, Jeffrey Kuo C-F (2023) Complete fully automatic detection, segmentation and 3D reconstruction of tumor volume for non-small cell lung cancer using YOLOv4 and region-based active contour model. Expert Syst Appl 212:118661. https://doi.org/10.1016/j.eswa.2022.118661
Armato 3rd SG, McLennan G, Bidaut L et al (2011) The Lung Image Database Consortium (LIDC) and Image Database Resource Initiative (IDRI): a completed reference database of lung nodules on CT scans. Med Phys 38:915–931. https://doi.org/10.1118/1.3528204
Challenge Grand (2016) Lung Nodule Analysis 2016. Grand Challenge. https://luna16.grand-challenge.org/. Accessed Nov 21 2023
Jung H, Kim B, Lee I et al (2018) Classification of lung nodules in CT scans using three-dimensional deep convolutional neural networks with a checkpoint ensemble method. BMC Med Imaging 18:48. https://doi.org/10.1186/s12880-018-0286-0
Gu Y, Lu X, Yang L et al (2018) Automatic lung nodule detection using a 3D deep convolutional neural network combined with a multi-scale prediction strategy in chest CTs. Comput Biol Med 103:220–231. https://doi.org/10.1016/j.compbiomed.2018.10.011
Zhao W, Yang J, Sun Y et al (2018) 3D deep learning from CT scans predicts tumor invasiveness of subcentimeter pulmonary adenocarcinomas. Cancer Res 78:6881–6889. https://doi.org/10.1158/0008-5472.CAN-18-0696
Jiang J, Hu Y-C, Liu C-J et al (2019) Multiple resolution residually connected feature streams for automatic lung tumor segmentation from CT Images. IEEE Trans Med Imaging 38:134–144. https://doi.org/10.1109/TMI.2018.2857800
Kim H, Goo JM, Lee KH et al (2020) Preoperative CT-based deep learning model for predicting disease-free survival in patients with lung adenocarcinomas. Radiology 296:216–224. https://doi.org/10.1148/radiol.2020192764
Ohno Y, Aoyagi K, Yaguchi A et al (2020) Differentiation of benign from malignant pulmonary nodules by using a convolutional neural network to determine volume change at chest CT. Radiology 296:432–443. https://doi.org/10.1148/radiol.2020191740
Hunter B, Chen M, Ratnakumar P et al (2022) A radionics-based decision support tool improves lung cancer diagnosis in combination with the Herder score in large lung nodules. EBioMedicine 86:104344. https://doi.org/10.1016/j.ebiom.2022.104344
Gugulothu VK, Balaji S (2023) An early prediction and classification of lung nodule diagnosis on CT images based on hybrid deep learning techniques. Multimed Tools Appl 1–21. https://doi.org/10.1007/s11042-023-15802-2
Dong Y, Hou L, Yang W et al (2021) Multi-channel multi-task deep learning for predicting EGFR and KRAS mutations of non-small cell lung cancer on CT images. Quant Imaging Med Surg 11:2354–2375. https://doi.org/10.21037/qims-20-600
Watanabe H, Kunitoh H, Yamamoto S et al (2006) Effect of the introduction of minimum lesion size on interobserver reproducibility using RECIST guidelines in non-small cell lung cancer patients. Cancer Sci 97:214–218. https://doi.org/10.1111/j.1349-7006.2006.00157.x
Willemink MJ, Koszek WA, Hardell C et al (2020) Preparing medical imaging data for machine learning. Radiology 295:4–15. https://doi.org/10.1148/radiol.2020192224
Lakhani P, Kim W, Langlotz CP (2012) Automated extraction of critical test values and communications from unstructured radiology reports: an analysis of 9.3 million reports from 1990 to 2011. Radiology 265:809–818. https://doi.org/10.1148/radiol.12112438
Wang J, Sourlos N, Zheng S et al (2023) Preparing CT imaging datasets for deep learning in lung nodule analysis: insights from four well-known datasets. Heliyon 9:e17104. https://doi.org/10.1016/j.heliyon.2023.e17104
Goodman SN, Fanelli D, Ioannidis JPA (2016) What does research reproducibility mean? Sci Transl Med 8:341ps12. https://doi.org/10.1126/scitranslmed.aaf5027
Mayo RC, Leung J (2018) Artificial intelligence and deep learning - Radiology’s next frontier? Clin Imaging 49:87–88. https://doi.org/10.1016/j.clinimag.2017.11.007
Sharp G, Fritscher KD, Pekar V et al (2014) Vision 20/20: perspectives on automated image segmentation for radiotherapy. Med Phys 41:050902. https://doi.org/10.1118/1.4871620
Shen Z, Han X, Xu Z, Niethammer M (2019) Networks for Joint Affine and Non-parametric Image Registration. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2019. pp. 4219–4228. https://doi.org/10.1109/cvpr.2019.00435
de Vos BD, Berendsen FF, Viergever MA et al (2019) A deep learning framework for unsupervised affine and deformable image registration. Med Image Anal 52:128–143. https://doi.org/10.1016/j.media.2018.11.010
Baumann D, Baumann K (2014) Reliable estimation of prediction errors for QSAR models under model uncertainty using double cross-validation. J Cheminform 6:47. https://doi.org/10.1186/s13321-014-0047-1
Yu AC, Mohajer B, Eng J (2022) External validation of deep learning algorithms for radiologic diagnosis: a systematic review. Radiol Artif Intell 4:e210064. https://doi.org/10.1148/ryai.210064
Toda R, Teramoto A, Kondo M et al (2022) Lung cancer CT image generation from a free-form sketch using style-based pix2pix for data augmentation. Sci Rep 12:12867. https://doi.org/10.1038/s41598-022-16861-5
Wu L, Zhuang J, Chen W et al (2022) Data augmentation based on multiple oversampling fusion for medical image segmentation. PLoS One 17:e0274522. https://doi.org/10.1371/journal.pone.0274522
Jacobs C, van Rikxoort EM, Murphy K et al (2016) Computer-aided detection of pulmonary nodules: a comparative study using the public LIDC/IDRI database. Eur Radiol 26:2139–2147. https://doi.org/10.1007/s00330-015-4030-7
Chassagnon G, De Margerie-Mellon C, Vakalopoulou M et al (2023) Artificial intelligence in lung cancer: current applications and perspectives. Jpn J Radiol 41:235–244. https://doi.org/10.1007/s11604-022-01359-x
Pande NA, Bhoyar D (2022) A comprehensive review of Lung nodule identification using an effective Computer-Aided Diagnosis (CAD) System. 2022 4th International Conference on Smart Systems and Inventive Technology (ICSSIT). https://doi.org/10.1109/ICSSIT53264.2022.9716327
Benzakoun J, Bommart S, Coste J et al (2016) Computer-aided diagnosis (CAD) of subsolid nodules: evaluation of a commercial CAD system. Eur J Radiol 85:1728–1734. https://doi.org/10.1016/j.ejrad.2016.07.011
Funding
This work was supported by “Pioneer” and “Leading Goose” R&D Program of Zhejiang (Grant No. 2022C03046), National Natural Science Foundation of China (Grant No. 82102128), Zhejiang Provincial Natural Science Foundation of China (Grant No. LTGY24H180006, LTGY23H180001), Medical and Health Science and Technology Project of Zhejiang Province (Grant No. 2024KY129, 2024KY132, 2022KY230), Research Project of Zhejiang Chinese Medical University (Grant No. 2022JKJNTZ19). The study sponsors had no role in the study design, in the collection, analysis, and interpretation of data; in the writing of the manuscript; and in the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Chen Gao.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was not required for this study because this was a systematic review.
Ethical approval
Institutional Review Board approval was not required because this was a systematic review.
Study subjects or cohorts overlap
Not applicable.
Methodology
-
Retrospective
-
Systematic review
Additional information
Publisher’s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, C., Wu, L., Wu, W. et al. Deep learning in pulmonary nodule detection and segmentation: a systematic review. Eur Radiol (2024). https://doi.org/10.1007/s00330-024-10907-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00330-024-10907-0