Abstract
APOEε4 is the major genetic risk factor for sporadic Alzheimer’s disease (AD). Although APOEε4 is known to promote Aβ pathology, recent data also support an effect of APOE polymorphism on phosphorylated Tau (pTau) pathology. To elucidate these potential effects, the pTau interactome was analyzed across APOE genotypes in the frontal cortex of 10 advanced AD cases (n = 5 APOEε3/ε3 and n = 5 APOEε4/ε4), using a combination of anti-pTau pS396/pS404 (PHF1) immunoprecipitation (IP) and mass spectrometry (MS). This proteomic approach was complemented by an analysis of anti-pTau PHF1 and anti-Aβ 4G8 immunohistochemistry, performed in the frontal cortex of 21 advanced AD cases (n = 11 APOEε3/ε3 and n = 10 APOEε4/ε4). Our dataset includes 1130 and 1330 proteins enriched in IPPHF1 samples from APOEε3/ε3 and APOEε4/ε4 groups (fold change ≥ 1.50, IPPHF1 vs IPIgG ctrl). We identified 80 and 68 proteins as probable pTau interactors in APOEε3/ε3 and APOEε4/ε4 groups, respectively (SAINT score ≥ 0.80; false discovery rate (FDR) ≤ 5%). A total of 47/80 proteins were identified as more likely to interact with pTau in APOEε3/ε3 vs APOEε4/ε4 cases. Functional enrichment analyses showed that they were significantly associated with the nucleoplasm compartment and involved in RNA processing. In contrast, 35/68 proteins were identified as more likely to interact with pTau in APOEε4/ε4 vs APOEε3/ε3 cases. They were significantly associated with the synaptic compartment and involved in cellular transport. A characterization of Tau pathology in the frontal cortex showed a higher density of plaque-associated neuritic crowns, made of dystrophic axons and synapses, in APOEε4 carriers. Cerebral amyloid angiopathy was more frequent and severe in APOEε4/ε4 cases. Our study supports an influence of APOE genotype on pTau-subcellular location in AD. These results suggest a facilitation of pTau progression to Aβ-affected brain regions in APOEε4 carriers, paving the way to the identification of new therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer’s disease (AD) is characterized by the extracellular deposition and self-aggregation of β-amyloid peptides (Aβ) into various types of Aβ deposits [22, 34], along with the intraneuronal accumulation and self-assembly of abnormally phosphorylated Tau proteins (pTau) into neurofibrillary tangles [14, 36, 37]. These neuropathological lesions affect successively different regions of the brain, following distinct stereotypical sequences described by the five Thal phases (0–5 for Aβ pathology [82]) and the six Braak stages (0-VI for Tau pathology [12]). A polymorphism of the apolipoprotein E gene (APOE) is the major genetic risk factor associated with sporadic AD. In comparison with the common APOEε3 allele, the APOEε4 allele is associated with an increased risk and an earlier onset for AD, in a dose-dependent manner (Odds Ratio APOEε4/ε4 = 14.2; www.alzgene.org; [19, 77]). In contrast, the rare APOEε2 allele confers a protection against the development of AD (Odds Ratio APOEε2/ε2 = 0.7; www.alzgene.org; [18]). In the brain, Apolipoprotein E (ApoE) is a glycoprotein predominantly secreted by astrocytes under physiologic conditions [11] and is involved in phospholipid and cholesterol transport: the C-terminus domain of ApoE binds with phospholipid packed into HDL-like particles conveying cholesterol [30, 68], while the N-terminus domain interacts with receptors of the LDLR family [41, 66].
ApoE is colocalized with parenchymal Aβ deposits in AD, as well as with vascular Aβ accumulation in a context of cerebral amyloid angiopathy (CAA) [57, 91]. The presence of the APOEε4 allele is strongly associated with an exacerbation of Aβ pathology [70]. Experimental data confirm the influence of APOE expression on Aβ accumulation [6, 31], unravelling a differential effect of ApoE isoforms on Aβ clearance impairment, aggregation and fibrillation (ApoE2 < ApoE3 < ApoE4) [15, 23, 39, 89]. In addition to these established effects of ApoE on Aβ pathology, emerging evidence suggests that ApoE also plays an important role in Tau pathology. The neuroanatomical progression of Tau pathology follows the expression gradients of APOE [54]. Furthermore, an exceptional resistance to an autosomal dominant familial AD mutation, associated with a massive parenchymal Aβ deposition but a surprisingly discrete Tau pathology, was recently attributed to the co-occurrence of a protective mutation in the APOEε3 sequence [1, 73]. Although ApoE is known to be colocalized with pTau within neurofibrillary tangles, their relationship remains elusive [57, 76]. In vivo experiments support an Aβ-independent effect of APOEε4 on pTau accumulation [44, 74], which may involve a promotion of Tau phosphorylation [13, 69] or a disruption of cholesterol metabolism and lysosomal functions [48]. Further investigations are needed in the AD brain to understand how APOE impacts Tau pathology, which correlates better than Aβ deposition with the cognitive status of AD cases [58].
The development of localized proteomics on post-mortem human brains, by our group, identified de novo proteins associated with Tau pathology [25, 26, 63, 64]. More particularly, the combination of anti-pTau pS396/pS404 immunoprecipitation with downstream proteomic analysis allowed the identification of proteins that specifically interact with pathologic pTau species involved in AD pathology [26]. In contrast, similar approaches focused on proteins interacting with total Tau, without discriminating its physiologic and pathologic forms [4, 9, 40, 49, 51, 53, 85, 87]. In this study, we took advantage of our unbiased strategy to fully uncover the effects of the AD risk factor APOEε4 on pTau metabolism: we combined anti-pTau immunoprecipitation with MS to map out, for the very first time, the pTau interactome in the AD brain of APOEε3 vs APOEε4 carriers.
Materials and methods
Cases
All procedures were performed under protocols approved by Institutional Review Boards at New York University Alzheimer’s Disease Research Center (NYU ADRC, NY, USA) and Columbia University Alzheimer’s Disease Research Center (CU ADRC, NY, USA). In all cases, written informed consent for research was obtained from the patient or legal guardian, and the material used had appropriate ethical approval for use in this project. All patients’ data and samples were coded and handled according to NIH guidelines to protect patients’ identities. A total of 25 cases of sporadic AD were included in this study. Cases were selected from donated brain tissue collected at the NYU ADRC and CU ADRC, based on their ABC score (A3, B3, C3; [55]), severity of Tau pathology in the frontal cortex and APOE genotype. The APOEε3/ε3 and APOEε4/ε4 groups were matched to the best of our ability in terms of age, sex and co-morbidities, as shown in Table 1. Our inclusion criteria involved indeed the absence of any additional primary tauopathy and of any major co-proteinopathy. For the neuropathological analysis, the presence of a concomitant Lewy Body disease of the amygdala-predominant type was tolerated for n = 2 cases per group to increase our number of cases, as this co-pathology is common in the elderly population and because its even distribution among our groups did not impact our comparative study design. Individual case information is detailed in Table 1 (age, sex, APOE genotype, post-mortem interval (PMI), ABC score, Braak stage, neuropathological findings, technical application).
Genotyping
APOE genotypes were provided by the NYU ADRC and CUMC brain banks for 15 out of 25 cases. For cases 1, 2, 4, 6, 7, 8, 14, 15, 16 and 18, APOE genotyping was performed as previously described [24]. A fragment of frozen frontal cortex was dissected (~ 25 mg), then collected into a 1.5 mL tube using single-use consumables in DNA-free experimental conditions. DNA was isolated using the DNeasy Blood & Tissue kit following the manufacturer’s instructions (#69,504, Qiagen). A single endpoint PCR was performed in a total volume of 25 µl containing 0.2 µM of each custom primer (Forward primer 5′ AGCCCTTCTCCCCGCCTCCCACTGT 3′; reverse primer 5′ CTCCGCCACCTGCTCCTTCACCTCG 3′; Millipore Sigma), 10 µl of DreamTaq Green PCR Master Mix 2X (#K1081, Thermo Scientific) and 4.2 µl of Betaine (#B0300, Millipore Sigma). Cycling conditions were set as follows: 98 °C for 4 min, 35 cycles at 98 °C/10 s, 63 °C/45 s and 72 °C/1 min 10 s, followed by 72 °C for 10 min. Unpurified PCR products were submitted to Genewiz for Sanger sequencing and the sequences were analyzed using the SnapGene 5.3.1 software.
Homogenization
Ten cases of sporadic AD were used for proteomic analysis (Table 1). The grey matter was dissected from the frontal cortex of archived fresh frozen human tissue samples stored at – 80 °C (~ 0.25 g per sample). Cortical tissue was homogenized as previously described [26]. Frozen tissue was enveloped into aluminum foil and pulverized on dry ice using a hammer. The powder was collected into a Dounce homogenizer, then homogenized on ice with ~ 25 strokes in a low salt homogenization buffer (50 mM HEPES pH 7.0, 250 mM sucrose, 1 mM EDTA) with inhibitors of proteases (cOmplete ULTRA Tablets, Mini, EDTA-free; #5,892,791,001, Millipore Sigma) and phosphatases (PhosphoSTOP EASYpack; #4,906,845,001, Millipore Sigma). The total protein concentration of homogenates was assessed with the Micro BCA Protein Assay Kit, following the supplier’s guidelines (#23,235, Thermo Scientific). Samples were stored at – 80 °C until use.
Immunoprecipitation
For each case, two immunoprecipitation products were obtained: the first using the mouse antibody anti-pTau pS396/pS404 (PHF1, provided by Dr. Peter Davies, Albert Einstein University, NY, USA [35]) to enrich pTau and its binding partners, the second using a mouse isotype antibody to control non-specific binding (#400,202, BioLegend). As a result, 20 separate IP products were individually analyzed for proteomics. Each IP product required a total of six-reaction mixes to collect enough material for downstream biochemistry and proteomics analyses, using the Dynabeads Protein G Immunoprecipitation Kit and following the supplier’s guidelines with minor adjustments (#10007D, Thermo Scientific). For each reaction mix, brain homogenate (300 μg total proteins/mix) and antibodies (4 μg antibodies/mix) were incubated overnight at 4 °C with over-end rotation to allow antigen–antibody interaction. The next day, the samples containing antigen–antibody complexes were mixed with Dynabeads (1.5 mg/mix), then incubated overnight at 4 °C with over-end rotation. The antigen–antibody-Dynabeads complexes were recovered and washed using a DynaMag-2 magnet (#12321D, Thermo Scientific), then resuspended in 100 μl of phosphate buffered saline at pH 7.4. The six-reaction mixes were pooled into a new tube (600 μL total), to avoid the co-elution of proteins bound to the tube wall. A total of 500 μL of the IP product was kept at 4 °C until proteomics analysis. The remaining 100 μL were eluted by capturing the antigen–antibody-Dynabeads complexes with the magnet, before incubating the beads in 20 μL of a denaturing buffer (141 mM Tris base, 106 mM Tris HCl, 2% SDS, 0.51 mM EDTA, pH 8.5; 15 min at 70 °C and 1000 rpm). The eluted fractions were recovered on the magnet and stored at – 20 °C until analysis.
Biochemistry analysis
Western blotting was performed to confirm the enrichment of pTau in the IP products. The equivalent of 10% of the IP product submitted to proteomic analyses was mixed with DTT 100 mM and 4X Bolt LDS Sample Buffer (#B0007, Thermo Scientific), then boiled at 98 °C for 5 min. Proteins were resolved on 4–12% Bis–Tris gels (#NP0322BOX, Thermo Scientific), then transferred onto a 0.2 μm nitrocellulose membrane (#1,620,112, BioRad). Membranes were blocked with 5% milk in Tris-buffered saline with 0.1% Tween-20 for 1 h, then probed with an anti-pTau S199/S202 antibody (1:1500; #44-768G, Thermo Scientific) at room temperature for 1 h, before being incubated with an anti-rabbit horseradish peroxidase antibody (1:3000; #NA934, Cytiva). The signal was revealed using the Pierce ECL Western Blotting Substrate (#32,106, Thermo Scientific) and membranes were imaged with the ChemiDoc MP Imaging System (BioRad). Silver staining was conducted to confirm the presence of a sufficient amount of proteins in our samples for proteomics downstream analysis. The equivalent of 5% of the IP product used in proteomics was mixed with DTT 100 mM and 4X Bolt LDS Sample Buffer (#B0007, Thermo Scientific), then boiled at 98 °C for 5 min. Proteins were resolved on 4–12% Bis–Tris gels (#NP0322BOX, Thermo Scientific). The gels were extracted and the proteins were stained using the SilverQuest Silver Staining Kit following the supplier’s guidelines (#LC6070, Thermo Scientific), to confirm the presence of a sufficient amount of proteins for proteomic downstream analyses. Gels were imaged with the ChemiDoc MP Imaging System (BioRad).
Proteomic analysis
IP products were analyzed by liquid-chromatography and mass spectrometry (LC–MS/MS), as previously detailed with some adjustments [26].
On-bead digestion and protein extraction
The antigen–antibody-bead complexes were recovered on a magnet then washed twice with ammonium bicarbonate 100 mM. Samples were reduced with DTT 0.2 M at 57 °C for 1 h, then alkylated with iodoacetamide 0.5 M at RT in the dark for 45 min. Sequencing-grade modified trypsin (Promega) was added to the sample for overnight digestion on a shaker at room temperature (300 ng). The next day, samples were acidified to pH 2 using 10% trifluoroacetic acid, then loaded onto equilibrated Ultra-Micro SpinColumns (Harvard Apparatus) using a microcentrifuge, before being rinsed three times with 0.1% TFA. The extracted samples were further washed with 0.5% acetic acid. The peptides were eluted with 40% acetonitrile in 0.5% acetic acid, followed by the addition of 80% acetonitrile in 0.5% acetic acid. Organic solvent was removed using a SpeedVac concentrator, before reconstituting samples in 0.5% acetic acid.
LC–MS/MS analysis
A total of 1 μg of protein was analyzed for each sample. A liquid chromatography (LC) separation was performed online with MS using the autosampler of an EASY-nLC 1000 (Thermo Scientific). Peptides were gradient-eluted from the column into the Orbitrap Eclipse using an 85 min gradient (Thermo Scientific). Solvent A consisted of 2% acetonitrile in 0.5% acetic acid and solvent B of 80% acetonitrile in 0.5% acetic acid. The gradient was held at 5% solvent B for 5 min, ramped to 35% solvent B in 60 min, to 45% solvent B in 10 min and to 100% solvent B in another 10 min. High-resolution full MS spectra were acquired with a resolution of 120,000, an AGC target of 4e5, a maximum ion time of 50 ms and a scan range of 400 to 1,500 m/z. All MS/MS spectra were recorded in the orbitrap analyzer using the following instrument parameters: resolution of 30,000, AGC target of 2e5, maximum ion time of 200 ms, one microscan, 2 m/z isolation window and NCE of 27.
Data processing
The MS/MS spectra were searched against the UniProt human database using Sequest within Proteome Discoverer 1.4. The data were filtered to better than 1% peptide and protein FDR searched against a decoy database. Only proteins with at least two different peptides were considered for downstream analysis. To asses if there are any differences in phosphorylation between the APOE groups, we analyzed Tau-protein phosphorylation using Byos (ProteinMetrics). The phosphorylation assignment and area integration were manually verified using the Byos interface. The area under the curve for the same peptide, with and without phosphorylation, was integrated and the amount of phosphorylation reported as a percentage. Note: not all phospho-sites have been observed in all samples and their frequency of observation has been highlighted in Fig. 4.
Data analysis
Data were analyzed using the Significance Analysis of INTeractome express algorithm (SAINT), as previously detailed [26, 78]. All non-human proteins, introduced during sample preparation, were removed from the results. The proteins were ranked by SAINT score and proteins with a SAINT score ≥ 0.80, equivalent to a FDR of ≤ 5%, were considered as pTau interactors and further studied. Three lists of pTau interactors of interest were submitted to and analyzed with STRING 11.5 and Cytoscape 3.9.1, to investigate network functional enrichments based on the Gene Ontology (GO) terms “cellular component” and “biological process”, using the total genome as background and a redundancy cut-off of 0.7: (1) proteins identified as pTau interactors in both APOE groups (SAINT score ≥ 0.80 in APOEε3/ε3 and APOEε4/ε4 groups, n = 33 proteins), (2) proteins identified as pTau interactors associated with an APOEε3/ε3 genotype (SAINT score ≥ 0.80 in APOEε3/ε3 cases, SAINT score < 0.80 in APOEε4/ε4 cases, n = 47 proteins) and 3) proteins identified as pTau interactors associated with an APOEε4/ε4 genotype (SAINT score ≥ 0.80 in APOEε4/ε4 cases, SAINT score < 0.80 in APOEε3/ε3 cases, n = 35 proteins). Network images were extracted and enrichment tables were exported then analyzed using Excel and GraphPad Prism 9. 4. 1. In addition, proteins considered as pTau interactors associated with one APOE genotype or the other were further compared among APOE groups based on their fold change (FC), calculated as follows: FC = [mean #peptide spectral matches (group of interest) + 1]/[mean #peptide spectral matches (group of reference) + 1].
Data comparison with previous MS-based studies
Our data were systematically compared to previous AD-related proteomic studies, using two complementary approaches. First, the pTau interactors identified here were compared to our previous study using the same anti-pTau pS396/pS404 (PHF1) antibody and a similar experimental strategy, but different AD tissues and a less stringent SAINT score cut-off (SAINT score ≥ 0.65, FDR ≤ 10% [26]). Second, we interrogated our probable pTau interactors using the NeuroPro searchable database v1.12 [3]. NeuroPro is a website that compiled 38 experimental MS-based proteomic datasets designed to assess protein changes occurring specifically in the AD brain. The following filters were applied: conditions “AD” (proteins associated with the AD brain) and “AD/C” (proteins altered in AD vs control brains). The resulting tables were exported and analyzed using Excel.
Immunohistochemistry
Immunohistochemistry was performed on formalin-fixed paraffin-embedded 8 µm-thick sections of frontal cortex. Sections were deparaffinized and rehydrated through a series of xylene and ethanol washes. Antigen retrieval was performed by treatment with 88% formic acid for 7 min, followed by boiling in citrate buffer (10 mM sodium citrate, 0.05% Tween-20; pH 6). Sections were blocked with 10% normal goat serum, then incubated overnight at 4 °C with a primary antibody anti-pTau pS396/pS404 (1:200; mouse antibody PHF1; provided by Dr. Peter Davies, Albert Einstein University, NY, USA) or anti-Aβ 4G8 (1:1000; mouse antibody; #800,711, BioLegend), diluted in 4% normal goat serum. Sections were incubated for 1 h at room temperature with an anti-mouse secondary antibody (1:1000, #BA-2000, Vector Laboratories), revealed with an avidin–biotin complex HRP detection kit (#PK-6100, Vector Laboratories) in combination with a DAB substrate kit (#34,065, Thermo Scientific), counterstained with Mayer’s hematoxylin (#MHS16, Millipore Sigma) and coverslipped (#P36970, Thermo Scientific). This technique was applied on 21 cases (n = 11 APOEε3/ε3 cases, n = 10 APOEε4/ε4 cases; Table 1).
Immunohistochemistry quantification
Tau pathology was quantified in the frontal cortex on anti-pTau pS396/pS404 PHF1 immunohistochemistry. Slides were scanned at a 40 × magnification with the Aperio VERSA 8 scanner and analyzed with Aperio ImageScope 12.4.3.5008 (Leica Biosystems). For each case, Tau pathology was quantified in the grey matter from three ROIs of 5.0 × 106 µm2 (± 0.5 × 106 µm2), encompassing all cortical layers and evenly spread over the cortical section. The total burden of PHF1 immunoreactive material was obtained by running the open source “Positive Pixel Count 2004-08-11” algorithm on each ROI, with the color saturation threshold set at 0.2. Raw data were exported on Excel to calculate the averaged percentage of immunopositive pixels out of the total number of pixels per case. Each neuropathological lesion composing Tau pathology was analyzed on their respective ROI: the first ROI was used to count the number of PHF1 positive neuronal profiles (pre-tangles, tangles and ghost tangles) per µm2; the second ROI was used to count the number of PHF1 positive neuritic crowns per µm2; in the third ROI, the burden of PHF1 positive neuropil threads was evaluated by assessing the averaged percentage of immunopositive pixels out of the total number of pixels from ten sub-ROIs of 1.0 × 104 µm2, covering neuropil areas evenly spread within the ROI and devoid of any neuronal profiles or neuritic crowns. Tau pathology was also quantified in the white matter from ten ROIs of 1.0 × 105 µm2 to assess the averaged percentage of immunopositive pixels out of the total number of pixels reflecting the burden of axonal threads. Unpaired non-parametric Mann–Whitney tests were performed on GraphPad Prism 9. 4. 1. to compare each of these ratios among APOE groups at a risk level of α = 0.05. Aβ pathology was quantified in the frontal cortex on anti-Aβ 4G8 immunohistochemistry. Slides were scanned at a 20 × magnification with the Aperio VERSA 8 scanner and analyzed with Aperio ImageScope 12.4.3.5008 (Leica Biosystems). For each case, Aβ pathology was quantified in the grey matter from three ROIs of 5.0 × 106 µm2 (± 0.5 × 106 µm2), encompassing all cortical layers and evenly spread over the cortical section. The total burden of 4G8 immunoreactive material was obtained by running the open source “Positive Pixel Count 2004-08-11” algorithm on each ROI, with the color saturation threshold set at 0.1. Raw data were exported on Excel to calculate the averaged percentage of immunopositive pixels out of the total number of pixels per case. CAA was analyzed in the parenchyma of the frontal cortex on anti-Aβ 4G8 immunohistochemistry, by attributing a semi-quantitative score to each case using the following criteria: 0 if none (no Aβ-positive vessel detected), 1 if sparse (< 25% of Aβ-positive vessels), 2 if moderate (about 50% of Aβ-positive vessels), 3 if severe (about 100% of Aβ-positive vessels). The type of CAA was evaluated by attributing a “Type 1” to cases presenting with capillary CAA along with CAA in larger vessels or a “Type 2” to cases presenting with CAA without capillary involvement [80].
Results
Proteomic overview
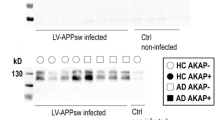
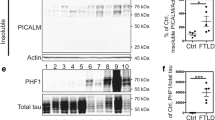
Immunoprecipitated fractions were obtained from frontal cortex homogenates of 10 advanced sporadic AD cases (n = 5 APOEε3/ε3 and n = 5 APOEε4/ε4 cases), using anti-pTau PHF1 (IPPHF1) or control IgG (IPIgG ctrl) antibodies. The enrichment of pTau in the IPPHF1 products was confirmed by western blot (Fig. 1a). A total of 1130 and 1330 proteins were detected in the IPPHF1 samples of the APOEε3/ε3 and APOEε4/ε4 groups (fold change ≥ 1.50, IPPHF1 vs IPIgG ctrl, Online Resource 1). Our dataset was filtered using the probabilistic SAINT score to identify the most likely pTau interactors, leading to the identification of 80 proteins of interest in the APOEε3/ε3 group and 68 in the APOEε4/ε4 group (SAINT scores ≥ 0.80, FDR ≤ 5%). Among these, 33 proteins were common to both APOE groups (Fig. 1b–d). They included 12/33 proteins previously identified as pTau interactors by our laboratory: AP3B2, ARMC8, KCNAB2, MAPT, PSMC1, PSMC2, PSMD13, PSMD2, PSMD3, RANBP9, SQSTM1 and WDR26 [26]. The remaining 21/33 proteins were identified as pTau interactors for the first time here, although they have been previously reported to have significantly altered protein levels in AD brain tissue (Fig. 1e and Online Resource 2; [3]). A significant network functional enrichment was associated with the 33 most probable pTau interactors common to APOEε3/ε3 and APOEε4/ε4 cases (PPI enrichment p value = 1.0 × 10–16; Online Resource 3a): they were predominantly associated with the proteasome system (GO terms “cellular component”, Fig. 1f) and involved in the regulation of mRNA metabolism, ubiquitin-dependent catabolic process or establishment of localization in cell (GO terms “biological process”, Fig. 1g).
Proteomic overview of the pTau interactome in sporadic AD cases of various APOE genotypes. a Western blot of the equivalent of 10% of the IPPHF1 or IPIgG ctrl products obtained from homogenates of frozen frontal cortex from 10 neuropathologically confirmed AD cases (cases 1–5 of APOEε3/ε3 genotype, cases 6–10 of APOEε4/ε4 genotype). b–c. One-sided volcano plots representing, for all proteins identified by LC–MS/MS, their SAINT score as a function of their fold change IPPHF1/IPIgG ctrl for the APOEε3/ε3 group (b) or for the APOEε4/ε4 group (c). d. Venn diagram representing the 80 and 68 proteins identified as probable pTau interactors in the APOEε3/ε3 and APOEε4/ε4 groups, respectively; 33 of these pTau interactors were common to both groups (SAINT score ≥ 0.80, FDR ≤ 5%). e Network representation of the 33 common pTau interactors. Each protein is represented by its gene ID as a node. The interacting nodes are connected by edges and their thickness indicates the strength of data support with a high confidence interaction score set at 0.7 (STRING). The node color reflects the protein status regarding previous AD-related proteomic studies, as detailed in the legend. f–g. Description of the functional enrichments associated with the 33 common pTau interactors as a function of –log10(FDR), using “genome” as background and a redundancy cut-off of 0.7. The incorporated bubble plots reflect the number of corresponding proteins in the network as a percentage, as detailed in the legend (PPI enrichment p value = 1.0 × 10–16; STRING and Cytoscape). The top panel details the top 5 GO terms “cellular component” (f) while the bottom panel shows the top 5 GO terms “biological process” (g). AD: Alzheimer’s disease; IP: Immunoprecipitation; FDR: false discovery rate
The pTau interactome in sporadic AD cases with an APOEε3/ε3 genotype
A total of 47 proteins were identified as most likely pTau interactors only for APOEε3/ε3 cases (SAINT score (APOEε3/ε3) ≥ 0.80, average SAINT score (APOEε3/ε3) ± SEM = 0.91 ± 0.01 vs SAINT score (APOEε4/ε4) < 0.80, average SAINT score (APOEε4/ε4) ± SEM = 0.52 ± 0.04). A significant network functional enrichment was attributed to these 47 proteins (PPI enrichment p value = 7.1 × 10–3; Online Resource 3b). In this network, 7/47 proteins were previously reported as probable pTau interactors by our group: KIF5C, LANCL2, PIN1, PIP4K2B, PSMC3, PSMD8 and PSMD11 [26]. The remaining 40/47 proteins were identified as pTau interactors for the first time here, although they included 38 proteins previously reported to have significantly altered protein levels in AD brain tissue (Fig. 2a and Online Resource 2; [3]). We detected four proteins particularly enriched in APOEε3/ε3 cases, in comparison to the APOEε4/ε4 group: NOP56, NOP58, PNN, TXNDC5 (fold change ≥ 1.50, IPPHF1(APOEε3/ε3) vs IPPHF1(APOEε4/ε4), ranked by relative abundance). The 47 proteins were predominantly enriched in proteins associated with the nucleoplasm (29/47 proteins; #1 GO term “cellular component”) and involved in RNA metabolic processes (19/47 proteins; #1 GO term “biological process”; Fig. 2a). A detailed analysis of the top significant functional enrichments associated with this network, ranked as per their FDR, emphasized a large predominance of functions associated with RNA binding and processing (Fig. 2b, top 5 GO terms “Cellular component”; Fig. 2c, top 5 GO terms “biological process”).
The pTau interactome associated with sporadic AD cases of APOEε3/ε3 genotype. a Network representation of the 47 proteins identified as probable pTau interactors associated with the APOEε3/ε3 group (SAINT score (APOEε3/ε3) ≥ 0.80 vs SAINT score (APOEε4/ε4) < 0.80). Each protein is represented by its gene ID as a node, which size reflects its relative abundance in comparison to the APOEε4/ε4 group (log10(fold change APOEε3/ε3/APOEε4/ε4)). The interacting nodes are connected by edges and their thickness indicates the strength of data support with a high confidence interaction score set at 0.7 (STRING). The node color reflects the protein status regarding previous AD-related proteomic studies, as detailed in the legend. These 47 proteins were mainly associated with the nucleoplasm (29/47; #1 GO term “cellular component”, orange outline) and involved in RNA metabolic processes (19/47; #1 GO term “biological process”, yellow outline). b–c Description of the functional enrichments associated with these 47 pTau interactors specific to the APOEε3/ε3 group as a function of –log10(FDR), using “genome” as background and a redundancy cut-off of 0.7. The incorporated bubble plots reflect the number of corresponding proteins in the network as a percentage, as detailed in the legend (PPI enrichment p value = 7.1 × 10–3; STRING and Cytoscape). The left panel details the top 5 GO terms “cellular component” (b), the right panel shows the top 5 GO terms “biological process” (c). All terms were ranked according to their FDR, the dashed lines materializing a significance threshold of FDR = 5%. FDR false discovery rate
The pTau interactome in sporadic AD cases with an APOEε4/ε4 genotype
By analogy, 35 proteins were identified as most likely pTau interactors only for APOEε4/ε4 cases (SAINT score (APOEε4/ε4) ≥ 0.80, average SAINT score (APOEε4/ε4) ± SEM = 0.88 ± 0.02 vs SAINT score (APOEε3/ε3) < 0.80, average SAINT score (APOEε3/ε3) ± SEM = 0.60 ± 0.02). A significant network functional enrichment was associated with these 35 proteins (PPI enrichment p value = 2.4 × 10–3; Online Resource 3c). In this network, 4/35 proteins were previously reported as probable pTau interactors by our laboratory: GLS, PSMC4, PSMC5 and SSBP1 [26]. The remaining 31/35 proteins corresponded to pTau interactors identified for the first time here, although they included 30 proteins previously reported to have significantly altered protein levels in AD brain tissue (Fig. 3a and Online Resource 2; [3]). We identified eight proteins particularly enriched in APOEε4/ε4 cases, in comparison to the APOEε3/ε3 group: ARRB1, SFXN5, GNL1, GRIA2, PP2R2A, SH3GL3, GLS, AP3B1 (fold change ≥ 1.50, IPPHF1 (APOEε4/ε4) vs IPPHF1 (APOEε3/ε3), ranked by relative abundance). This network of 35 proteins was predominantly associated with the synaptic compartment (14/35; #1 GO term “cellular component”) and involved in intracellular transport (21/35; #1 GO term “biological process”; Fig. 3a). A detailed analysis of the top significant functional enrichments associated with this network, ranked by FDR, depicted a majority of functions associated with synaptic transmission and cellular trafficking (Fig. 3b, top 5 GO terms “cellular component”; Fig. 3c, top 5 GO terms “biological process”).
The pTau interactome associated with sporadic AD cases of APOEε4/ε4 genotype. a Network representation of the 35 proteins identified as probable pTau interactors associated with the APOEε4/ε4 group (APOEε4/ε4) ≥ 0.80 vs SAINT score (APOEε3/ε3) < 0.80). Each protein is represented by its gene ID as a node, which size reflects its relative abundance in comparison to the APOEε3/ε3 group (log10(fold change APOEε4/ε4/APOEε3/ε3)). The interacting nodes are connected by edges and their thickness indicates the strength of data support with a high confidence interaction score set at 0.7 (STRING). The node color reflects the protein status regarding previous AD-related proteomic studies, as detailed in the legend. These 35 proteins were mainly associated with the synapse (14/35; #1 GO term “cellular component”, red outline) and involved in transport pathways (21/35; #1 GO term “biological process”, pink outline). b–c. Description of the functional enrichments associated with these 35 pTau interactors specific to APOEε4/ε4 cases as a function of –log10(FDR), using the total genome as background and a redundancy cut-off of 0.7. The incorporated bubble plots reflect the number of corresponding proteins in the network as a percentage, as detailed in the legend (PPI enrichment p value = 2.4 × 10–2; STRING and Cytoscape). The left panel details the top 5 GO terms “cellular component” (b), the right panel shows the top five GO terms “biological process” (c). All terms were ranked according to their FDR, the dashed lines materializing a significance threshold of FDR = 5%. FDR false discovery rate
Tau-phosphorylation landscape across APOE genotypes
The MS analysis of IPPHF1 products identified 30 phosphorylation sites associated with Tau proteins. Each site was mapped along Tau sequence based on the 2N4R Tau isoform of 441 amino acids, associated with its relative abundance and frequency of observation within each APOE group. MS analysis could not always identify the exact position of a phosphate group between T403 and S404 as the peptide backbone cleavage between these two amino acids was sometimes missing and because of an identical retention time under the LC conditions used. We therefore combined the information associated with the pT403 and pS404 sites in this analysis, referred to as pT403-pS404. The most abundant and frequently detected phosphorylation sites, alone or in combination on the same peptide, were the following: pT175 with pT181, pT181, pS202, pT212 with pT217, pT217, pT231, pT231 with pS235, pS262, pS289, pS396, pS396 with pS400, pS396 with pT403-pS404, pT403-pS404. Of note, the phosphorylation sites recognized by the anti-pTau PHF1 antibody, pS396 and pS404, were observed for all cases in both APOE groups, validating our success in enriching the targeted pTau proteins. As shown in Fig. 4 and Online Resource 4, the phosphorylation landscapes of Tau protein analyzed in our samples were similar between APOEε3/ε3 and APOEε4/ε4 groups, except for one observation: the combination of both pT175 and pT181 sites was only seen in the APOEε4/ε4 group, in which it was observed in three out of five cases on a unique peptide (IPAKpTPPAPKpTPPSSGEPPK). The unphosphorylated version of this peptide was not identified, which confirms this modification further as Trypsin can only cleave peptide bonds at the C-terminal side of lysine (K) and arginine (R) residues and will not cleave at the N-terminal side of a phosphorylated residue (Fig. 4 and Online Resource 4).
The phosphorylation landscape of Tau protein across APOE genotypes. Tau-phosphorylation sites were identified by mass spectrometry in anti-pTau PHF1 immunoprecipitated fractions (n = 5 APOEε3/ε3 cases, n = 5 APOEε4/ε4 cases). All detected sites were mapped along the Tau-protein sequence, based on the 2N4R Tau isoform of 441 amino acids. The relative abundance of each phosphorylation site was represented by the node color, as detailed in the legend. The node color-code reflects the average percentage of phosphorylated peptide normalized to all observed versions of the respective peptide, for each APOE group. Note that some phosphorylation sites were detected across multiple Trypsin cleavage products, hence the use of summed percentages for all peptides containing the same site(s). The frequency of observation of each phosphorylated modification within a group is represented separately by the node size, as shown in the legend. The node size reflects the number of cases out of 5 total presenting with the corresponding phosphorylation site(s), for each APOE group. The absence of any phosphorylation site is represented by a “- “
Characterization of Tau lesions among APOE groups
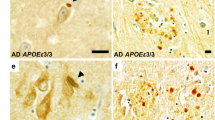
Our proteomic observations suggest that the expression of the AD risk factor APOEε4 mostly impacts pTau-subcellular location. To validate this hypothesis, a comparative quantification of Tau pathology across APOE genotypes was performed in the frontal cortex of 21 advanced sporadic AD cases, after an anti-pTau PHF1 immunohistochemistry (n = 11 APOEε3/ε3 and n = 10 APOEε4/ε4 cases). As expected, the pTau burden detected in the grey matter of the frontal cortex was similar between the APOEε3/ε3 and APOEε4/ε4 groups, composed of advanced AD cases (unpaired non-parametric Mann–Whitney test, p > 0.05; Fig. 5a). To assess the influence of APOEε4 on pTau-subcellular location, the distribution of pTau aggregates among the various neuronal compartment were further analyzed. All of the well-characterized subtypes of pTau pathologic lesions linked to AD were observed in both APOEε3/ε3 and APOEε4/ε4 groups, including: neurofibrillary tangles consisting of pTau aggregates in the neuronal soma (“tangles”, Fig. 5b and f), pTau-positive neuritic crowns consisting of degenerated axonal terminals and enlarged synapses which compose the neuritic amyloid plaques (“neuritic crowns”, Fig. 5c and g), neuropil threads consisting of the accumulation of pTau mostly in dendrites (“neuropil threads”, Fig. 5d and h), and axonal threads consisting of thin and fragmented threads observed in the white matter (“axonal threads”, Fig. 5e and i). The quantifications of tangles, neuropil threads and axonal threads did not show any differences between the APOEε3/ε3 and APOEε4/ε4 groups (unpaired non-parametric Mann–Whitney tests, p > 0.05, Fig. 5b, d–e). The density of pTau-positive neuritic crowns was, however, significantly higher in the APOEε4/ε4 cases, in comparison with the APOEε3/ε3 cases (unpaired non-parametric Mann–Whitney test, p = 0.0062, Fig. 5c); results were similar after data normalization with the pTau burden of each respective ROI (unpaired non-parametric Mann–Whitney test, p = 0.0159; not shown).
Influence of APOEε4 on Tau pathology in the frontal cortex of advanced AD cases. a–i. Histologic analysis of Tau pathology conducted on a total of n = 11 APOEε3/ε3 cases and n = 10 APOEε4/ε4 cases, presenting with advanced AD. Anti-pTau PHF1 immunohistochemistry with DAB revelation. Human brain, formalin-fixed paraffin-embedded frontal cortex, × 40 magnification. a Quantification of the total pTau burden in the frontal cortex as an averaged percentage of PHF1 positive pixels in n = 3 ROIs/case of 5.0 × 106 µm2 (± 0.5 × 106 µm2) each, encompassing all cortical layers and evenly spread over the cortical section. b Quantification of the density of tangles in the frontal cortex as a number of PHF1 positive neuronal profiles (pre-tangles, tangles, ghost tangles) per μm2. Analysis performed on n = 1 ROI/case of 5.0 × 106 µm2 (± 0.5 × 106 µm2) covering all the cortical layers of the grey matter. c Quantification of the density of neuritic crowns in the frontal cortex as a number of PHF1 positive neuritic crowns per μm2. Analysis performed on n = 1 ROI/case of 5.0 × 106 µm2 (± 0.5 × 106 µm2) covering all the cortical layers of the grey matter. d Quantification of the burden of PHF1 positive neuropil threads as an averaged percentage of PHF1 positive pixels from n = 10 sub-ROIs/case of 1.0 × 104 µm2, covering neuropil areas evenly spread over all cortical layers of the grey matter and devoid of any neuronal profiles or neuritic crowns, within n = 1 ROI/case of 5.0 × 106 µm2 (± 0.5 × 106 µm2). e. Quantification of the burden of PHF1 positive axonal threads as an averaged percentage of PHF1 positive pixels from n = 10 ROIs/case of 1.0 × 105 µm2 covering the white matter. f–h. Illustration of a tangle (f, arrows), neuritic crown (g, circle) and neuropil thread (h, arrowheads) observed in the grey matter. i Illustration of thin and fragmented axonal threads seen in the white matter (arrowheads). Scale bar: 20 μm. Unpaired non-parametric Mann–Whitney tests, ns p > 0.05, **p < 0.01. AD Alzheimer’s disease, ROI region of interest
CAA profile among APOE groups
A comparative analysis of Aβ pathology across APOE genotypes was performed in the frontal cortex of 21 advanced sporadic AD cases, after an anti-Aβ 4G8 immunohistochemistry (n = 11 APOEε3/ε3 and n = 10 APOEε4/ε4 cases, Fig. 6a–b). The Aβ burden detected in the grey matter of the frontal cortex was similar between the APOEε3/ε3 and APOEε4/ε4 groups, composed of advanced AD cases (unpaired non-parametric Mann–Whitney test, p > 0.05; Fig. 6c). It has been recently suggested that CAA interacts with neuritic plaques to enhance Tau pathology [65]. As the expression of ApoE4 is known to exacerbate CAA [71], in addition of being strongly associated with the presence of capillary CAA [80, 81], the CAA profile was analyzed in the parenchyma of the frontal cortex in our cohort (Fig. 6d). A semi-quantitative score was attributed to each case as follows: “none” when no Aβ-positive vessel was detected in the cortical parenchyma, “rare” when only a few scattered Aβ-positive vessels were seen, “moderate” if about 50% of vessels were Aβ-positive, “severe” if about 100% of vessels were Aβ-positive. The presence of CAA was detected in 36.4% of cases in the APOEε3/ε3 group, while this proportion reached 72.8% of cases in the APOEε4/ε4 group. When present, CAA was scored as “rare” in 100% of the CAA positive cases in the APOEε3/ε3 group. In contrast, the CAA scores observed in the APOEε4/ε4 group were either “rare” (25% of the CAA positive cases), “moderate” (37.5% of the CAA positive cases) or “severe” (37.5% of the CAA positive cases; Fig. 6). The distribution of CAA types was different among APOE groups: whereas no capillary CAA was observed among the CAA positive cases of the APOEε3/ε3 group (0% of CAA type 1, 100% of CAA type 2), the presence of capillary CAA was predominant among the CAA positive cases of the APOEε4/ε4 group (62.5% of CAA type 1, 37.5% of CAA type 2).
Influence of APOEε4 on CAA frequency, severity and type in the frontal cortex of advanced AD cases. a–d. Histologic analysis of Aβ pathology conducted on a total of n = 11 APOEε3/ε3 cases and n = 10 APOEε4/ε4 cases, presenting with advanced AD. Anti-Aβ 4G8 immunohistochemistry with DAB revelation. Human brain, formalin-fixed paraffin-embedded frontal cortex, × 20 magnification. a–b Representative illustration of Aβ deposits seen in the frontal cortex of an APOEε3/ε3 case (a) and APOEε4/ε4 case (b). Note the presence of CAA in the parenchyma of the APOEε4/ε4 case, involving a capillary (arrow) and a larger vessel (arrowheads). Scale bar: 20 μm. c Quantification of the total Aβ burden in the frontal cortex as an averaged percentage of 4G8 positive pixels in n = 3 ROIs/case of 5.0 × 106 µm2 (± 0.5 × 106 µm2) each, encompassing all cortical layers and evenly spread over the cortical section. Unpaired non-parametric Mann–Whitney tests, ns p > 0.05. d Descriptive table and circle diagrams showing the frequency, severity and type of CAA among a total of n = 11 APOEε3/ε3 cases and n = 10 APOEε4/ε4 cases. A semi-quantitative CAA score was attributed to each case as follows: none (no Aβ-positive vessel detected), rare (scattered Aβ-positive vessels), moderate (about 50% of Aβ-positive vessels), severe (about 100% of Aβ-positive vessels). The CAA type was evaluated among CAA positive cases as follows: type 1 (capillary CAA associated with CAA in larger vessels) or type 2 (CAA without capillary involvement). AD Alzheimer’s disease, CAA Cerebral Amyloid Angiopathy, ROI region of interest
Discussion
By profiling the pTau interactome selectively in APOEε3/ε3 and APOEε4/ε4 carriers for the first time, we discovered that the APOE genotype significantly influences the pTau pS396/pS404 interactome. We determined that the pTau interactome reflects a different subcellular localization of pTau aggregates in APOEε3/ε3 and APOEε4/ε4 cases. We confirmed this result through our follow-up immunohistochemistry studies and propose that the AD risk factor APOEε4 facilitates Tau-pathology progression by enhancing the accumulation of pTau in axonal endings and synapses, particularly in Aβ-affected brain regions.
A total of 80 and 68 proteins were identified as probable pTau interactors in the APOEε3/ε3 and APOEε4/ε4 groups, including 33 proteins that interacted with pTau irrespective of APOE genotype. These 33 common pTau interactors showed a preserved interaction of pTau with proteins involved in the ubiquitin–proteasome system among APOEε3/ε3 and APOEε4/ε4 cases. This observation confirms the importance of the ubiquitin–proteasome system in a context of defective protein clearance and protein accumulation [56, 61], while highlighting some of its key members consistently observed among pTau interactors in the AD brain, such as: SQSTM1 (also known as p62), which is involved in the shuttle of polyubiquitinated Tau for proteasomal degradation; ubiquitin/polyubiquitin precursors such as RPS27A and UBC; ubiquitin protein ligases such as ARMC8 and UBR4; several members of the PSMC and PSMD families constituting the proteasome [5, 10, 26, 60]. The detected PSMC and PSMD members constitute only the lid and base of the 26S proteasome regulatory subunit involved in substrate recognition, deubiquitylation and unfolding, underlining the specificity of our approach [7]. Proteins involved in the regulation of RNA processing were also detected among the common pTau interactors, supporting an important role of RNA metabolism and translational stress response in Tau pathology as suggested by other studies [43]. These new results extend our previous analysis by bringing to light additional RNA-associated proteins of interest, which were previously detected as present in the neurofibrillary tangle proteome, although they were below the threshold to be considered as pTau interactors so far [26]: the splicing factors SRSF1 (also known as SF2/ASF) and TRA2B, involved in the regulation of the alternative splicing of Tau exon 10 impacting the ratio of Tau isoforms with three or four microtubule binding repeats domains (3R or 4R, with a 4R:3R ratio shifted from approximately 1:1 to 2:1 in AD) [17, 20, 33]. Interestingly, a large proportion of the proteins identified as probable pTau interactors in the APOEε3/ε3 and APOEε4/ε4 groups did not overlap. This observation cannot be explained by the inclusion of differing stages of AD pathology among APOE groups, as this study was conducted on an homogenous selection of sporadic cases diagnosed with an advanced AD pathology at autopsy (A3, B3, C3 scores [55]). This key result supports an influence of APOE expression on pTau metabolism in the AD brain.
A total of 47 proteins were identified as probable pTau interactors only for APOEε3/ε3 cases, while 35 proteins were detected as probable pTau interactors only for APOEε4/ε4 cases. A robust segregation of their associated functions and cell compartments was observed. The pTau interactome specific to APOEε3/ε3 cases contained a majority of nucleoplasmic proteins, involved in RNA binding and processing. PNN, a newly identified pTau interactor, was particularly enriched in APOEε3/ε3 cases and recently characterized in vitro as involved in the formation of RNA condensates defining subcellular sites of Tau aggregation [45, 46]. In vitro, Tau coacervates with polyanions, such as RNA, into liquid droplets [38, 47, 95]. The colocalization of Tau with RNA and RNA binding proteins in the cell can result in the formation of stress granules involved in protein-level regulation [2, 62], although experimental observations suggest that such complexes drive Tau aggregation [2, 38, 42, 52]. In AD neurons, pTau is accumulated at the nuclear envelope in a discrete manner, which correlates with nuclear component mis-localization [29, 38]. Our results support a predominance of these biologic events in APOEε3/ε3 cases. They can be protective in the early stages of the pathology, slowing down Tau pathology by sequestering pTau species at the nucleus while triggering the translational stress response. Over time, deleterious outcomes may take over including a promotion of pTau self-aggregation [2, 38, 42, 52] or a nucleocytoplasmic transport disruption [29, 94]. In contrast, the pTau interactome specific to APOEε4/ε4 cases contained a majority of synaptic proteins, involved in cell transport. ARRB1, a newly identified pTau interactor particularly enriched in the APOEε4/ε4 group, is a representative synaptic protein recently described as a promoter of Tau pathology [92]. Tau pathology in APOEε4/ε4 cases may be characterized by a predominant pool of dynamic pTau species, more prone to be transported to the synapse, which can facilitate the trans-synaptic progression of Tau pathology [88, 93] and Tau-mediated synaptic disruption [86]. We did not observe any major effect of APOEε4 on Tau-phosphorylation sites, as suggested by recent experimental observations [69]. Only the pT175/pT181 modification was specific to the APOEε4/ε4 group, although this result requires further investigations since this study is based on a selected fraction of pTau that is phosphorylated on pS396/pS404 and obtained from post-mortem material [32]. We suggest that the expression of the AD risk factor APOEε4 mostly impacts pTau-subcellular location.
A comprehensive analysis of the various subcellular lesions composing Tau pathology in APOEε3/ε3 and APOEε4/ε4 cases confirmed our hypothesis. The density of pTau-positive neuritic crowns was higher in APOEε4/ε4 vs APOEε3/ε3 cases, despite the confirmation of an even burden of Tau pathology among cases. Neuritic crowns are made of pTau-positive degenerated neurites wrapping the most mature type of Aβ deposits, constituting the neuritic amyloid plaque – or senile plaque [27]. The immunohistochemical signature of these pTau-positive neuritic crowns [72, 84], along with the observation of presynaptic vesicles [79], demonstrate their axonal nature. This observation aligns with our proteomics findings and further support an effect of APOEε4 on pTau cellular transport and relocation toward axonal endings and synapses. Tau pathology progresses from neuron to neuron through synaptic connections [16, 28, 83, 88, 93]; we hypothesize that the spreading of pathological pTau species is accelerated in APOEε4/ε4 carriers, in accordance with recent clinical observations [8, 75]. In vivo experiments support an Aβ-independent influence of APOEε4 on Tau-pathology spreading, by demonstrating an exacerbation of Tau pathology in PS19 mice expressing human APOEε4/ε4 [74], but a recent study questions this scenario [21]. These results suggest a direct consequence of APOE genotype itself on Tau pathology (e.g., the proteins identified in the present study as pTau interactors more associated with one APOE genotype or another could have different expression levels in control brains). This possibility is illustrated by the recent identification of 25 unique proteins defining the incipient AD proteomic signature, including 24 increased in young heterogenous APOEε4 carriers, while 1 protein was reduced in comparison to aged-matched APOEε4 non-carriers [67]. Although we could not identify any of our 47 APOEε3-associated and 35 APOEε4-associated pTau interactors in this list of 25 candidates, future studies are needed to address this possibility by better understanding how the AD risk factor APOEε4 shapes the basal metabolism of the brain. While they are not mutually exclusive, an alternative scenario involves an Aβ-mediated effect of APOE on Tau pathology. The expression of the APOEε4 allele is indeed strongly associated with an exacerbation of Aβ pathology, by promoting particularly the development of neuritic amyloid plaques and of CAA with a capillary involvement [70, 81, 90], as confirmed in our cohort. Recent neuropathological and clinical studies show that CAA interacts with neuritic amyloid plaques to enhance tau pathology and white-matter rarefaction [50, 59, 65]. We propose that the AD risk factor APOEε4 promotes neuritic degeneration, resulting in the accumulation of pTau in axonal endings and synapses which may facilitate Tau-pathology progression, particularly toward Aβ-affected brain regions.
Limitations
While there were many consistencies with our previous pTau interactome study [26], our results did not completely replicate our previous findings. The inter-individual variability associated with post-mortem human brain studies, combined with a modified MS method used here, could explain these differences. In our current study, our MS protocol was adjusted to simplify our workflow. To prevent the excess of antibodies co-eluted in the immunoprecipitated product from hindering the detection of proteins with similar mass weight and elution times, immunoprecipitated products were previously run on a gel from which bands containing antibodies were excised and analyzed separately on the mass spectrometer [26]. Here, we opted for a different strategy to minimize sample processing prior to MS analysis: a proteolytic digestion was performed straight on the antigen–antibody-bead complexes without removing antibodies prior to downstream proteomic analysis, increasing the power of our study by allowing a better technical consistency. Furthermore, our designation of “pTau interactors” relied on the binarization of a continuum of probabilistic scores, based on the use of a stringent threshold corresponding to a FDR of 5%. Although this strategy allows for the reduction of false positives and a focus on the most probable pTau interactors for a more stringent analysis, it increases the risk of false negatives among the proteins that did not pass the threshold. Altogether, these limitations explain why some important pTau interactors may be missing in the present study and emphasize the need to generate more AD-related proteomic datasets, to counterbalance inter-experimental differences. Although pathologically relevant, these proteomic findings may reflect advanced biologic responses in AD, as they are associated with the late epitope pTau pS396/pS404 extracted from advanced AD cases. It is still unclear if a different subset of pTau species interacts with different proteins. These aspects need to be further addressed in future investigations by exploring the interactome of alternative pTau epitopes, especially early ones extracted from early and intermediate stage AD cases.
Conclusion and perspectives
This study provides evidence for an influence of APOE expression on pTau-subcellular location, suggesting a greater variation of Tau pathology across AD cases. Indeed, these new results emphasize the complexity of Tau studies, as factors such as genotype can modify the subcellular localization of pTau and therefore its interactome. Our results pave the way to the potential identification of new therapeutic targets specific for APOEε4 carriers.
Data availability
The mass spectrometric raw files are accessible at https://massive.ucsd.edu under accession MassIVE MSV000094757 and at www.proteomexchange.org under accession PXD052263.
References
Arboleda-Velasquez J, Lopera F, O’Hare M, Delgado-Tirado S, Marino C, Chmielewska N et al (2019) Resistance to autosomal dominant Alzheimer’s disease in an APOE3 Christchurch homozygote: a case report. Nat Med 25:1680–1683. https://doi.org/10.1038/S41591-019-0611-3
Ash PEA, Lei S, Shattuck J, Boudeau S, Carlomagno Y, Medalla M et al (2021) TIA1 potentiates tau phase separation and promotes generation of toxic oligomeric tau. Proc Natl Acad Sci USA. https://doi.org/10.1073/PNAS.2014188118/-/DCSUPPLEMENTAL
Askenazi M, Kavanagh T, Pires G, Ueberheide B, Wisniewski T, Drummond E (2023) Compilation of reported protein changes in the brain in Alzheimer’s disease. Nat Commun 141(14):1–15. https://doi.org/10.1038/s41467-023-40208-x
Ayyadevara S, Balasubramaniam M, Parcon PA, Barger SW, Griffin WST, Alla R et al (2016) Proteins that mediate protein aggregation and cytotoxicity distinguish Alzheimer’s hippocampus from normal controls. Aging Cell 15:924–939. https://doi.org/10.1111/ACEL.12501
Babu JR, Geetha T, Wooten MW (2005) Sequestosome 1/p62 shuttles polyubiquitinated tau for proteasomal degradation. J Neurochem 94:192–203. https://doi.org/10.1111/J.1471-4159.2005.03181.X
Bales KR, Verina T, Dodel RC, Du Y, Altstiel L, Bender M et al (1997) Lack of apolipoprotein E dramatically reduces amyloid β-peptide deposition. Nat Genet 173(17):263–264. https://doi.org/10.1038/ng1197-263
Bard JAM, Goodall EA, Greene ER, Jonsson E, Dong KC, Martin A (2018) Structure and function of the 26S proteasome. Annu Rev Biochem 87:697. https://doi.org/10.1146/ANNUREV-BIOCHEM-062917-011931
Benson GS, Bauer C, Hausner L, Couturier S, Lewczuk P, Peters O et al (2022) Don’t forget about tau: the effects of ApoE4 genotype on Alzheimer’s disease cerebrospinal fluid biomarkers in subjects with mild cognitive impairment-data from the Dementia competence network. J Neural Transm 129:477–486. https://doi.org/10.1007/S00702-022-02461-0
Betters RK, Luhmann E, Gottschalk AC, Xu Z, Shin MR, Ptak CP et al (2023) Characterization of the Tau interactome in human brain reveals isoform-dependent interaction with 14–3–3 family proteins. Neuro 10:1–11. https://doi.org/10.1523/ENEURO.0503-22.2023
Blaudin de Thé FX, Lassus B, Schaler AW, Fowler SL, Goulbourne CN, Jeggo R et al (2021) P62 accumulates through neuroanatomical circuits in response to tauopathy propagation. Acta Neuropathol Commun. https://doi.org/10.1186/S40478-021-01280-W
Boyles JK, Pitas RE, Wilson E, Mahley RW, Taylor JM (1985) Apolipoprotein E associated with astrocytic glia of the central nervous system and with nonmyelinating glia of the peripheral nervous system. J Clin Invest 76:1501–1513. https://doi.org/10.1172/JCI112130
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82:239–259
Brecht WJ, Harris FM, Chang S, Tesseur I, Yu GQ, Xu Q et al (2004) Neuron-specific apolipoprotein e4 proteolysis is associated with increased tau phosphorylation in brains of transgenic mice. J Neurosci 24:2527–2534. https://doi.org/10.1523/JNEUROSCI.4315-03.2004
Brion JP, Passareiro H, Nunez J, Flament-Durand J (1985) Mise en évidence immunologique de la protéine Tau au niveau des lésions de dégénérescence neurofibrillaire de la maladie d’Alzheimer. Arch Biol (Bruxelles) 95:229–235
Castellano JM, Kim J, Stewart FR, Jiang H, DeMattos RB, Patterson BW et al (2011) Human apoE isoforms differentially regulate brain amyloid-β peptide clearance. Sci Transl Med. https://doi.org/10.1126/SCITRANSLMED.3002156/SUPPL_FILE/3-89RA57_SM.PDF
Clavaguera F, Bolmont T, Crowther RAA, Abramowski D, Frank S, Probst A et al (2009) Transmission and spreading of tauopathy in transgenic mouse brain. Nat Cell Biol 11:909–913. https://doi.org/10.1038/ncb1901
Conrad C, Zhu J, Conrad C, Schoenfeld D, Fang Z, Ingelsson M et al (2007) Single molecule profiling of tau gene expression in Alzheimer’s disease. J Neurochem 103:1228–1236. https://doi.org/10.1111/J.1471-4159.2007.04857.X
Corder E, Saunders A, Risch N, Strittmatter W, Schmechep D, Gaskell P et al (1994) Protective effect of apolipoprotein E type 2 allele for late onset Alzheimer disease. Nat Gen 7:180–184
Corder E, Saunders A, Strittmatter W, Schmechel D, Gaskell P, Small G et al (1993) Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science (80-) 261:921–923
D’Souza I, Schellenberg GD (2006) Arginine/serine-rich protein interaction domain-dependent modulation of a tau exon 10 splicing enhancer: altered interactions and mechanisms for functionally antagonistic FTDP-17 mutations Delta280K AND N279K. J Biol Chem 281:2460–2469. https://doi.org/10.1074/JBC.M505809200
Davies C, Tulloch J, Yip E, Currie L, Colom-Cadena M, Wegmann S et al (2023) Apolipoprotein E isoform does not influence trans-synaptic spread of tau pathology in a mouse model. Brain Neurosci Adv. https://doi.org/10.1177/23982128231191046
Delaère P, Duyckaerts C, He Y, Piette F, Hauw JJ (1991) Subtypes and differential laminar distributions of beta A4 deposits in Alzheimer’s disease: relationship with the intellectual status of 26 cases. Acta Neuropathol 81:328–335. https://doi.org/10.1007/bf00305876
Dodart JC, Marr RA, Koistinaho M, Gregersen BM, Malkani S, Verma IM et al (2005) Gene delivery of human apolipoprotein E alters brain Aβ burden in a mouse model of Alzheimer’s disease. Proc Natl Acad Sci USA 102:1211. https://doi.org/10.1073/PNAS.0409072102
Drummond E, Kavanagh T, Pires G, Marta-Ariza M, Kanshin E, Nayak S et al (2022) The amyloid plaque proteome in early onset Alzheimer’s disease and Down syndrome. Acta Neuropathol Commun 101(10):1–24. https://doi.org/10.1186/S40478-022-01356-1
Drummond E, Nayak S, Pires G, Ueberheide B, Wisniewski T (2018) Isolation of amyloid plaques and neurofibrillary tangles from archived alzheimer’s disease tissue using laser-capture microdissection for downstream proteomics. Methods Mol Biol 1723:319–334. https://doi.org/10.1007/978-1-4939-7558-7_18
Drummond E, Pires G, MacMurray C, Askenazi M, Nayak S, Bourdon M et al (2020) Phosphorylated tau interactome in the human Alzheimer’s disease brain. Brain 143:2803–2817. https://doi.org/10.1093/BRAIN/AWAA223
Duyckaerts C, Delatour B, Potier M-C (2009) Classification and basic pathology of Alzheimer disease. Acta Neuropathol 118:5–36. https://doi.org/10.1007/s00401-009-0532-1
Duyckaerts C, Uchihara T, Seilhean D, He Y, Hauw J-J (1997) Dissociation of Alzheimer type pathology in a disconnected piece of cortex. Acta Neuropathol 93:501–507
Eftekharzadeh B, Daigle JG, Kapinos LE, Coyne A, Schiantarelli J, Carlomagno Y et al (2018) Tau protein disrupts nucleocytoplasmic transport in alzheimer’s disease. Neuron 99:925-940.e7. https://doi.org/10.1016/J.NEURON.2018.07.039
Fagan AM, Holtzman DM, Munson G, Mathur T, Schneider D, Chang LK et al (1999) Unique lipoproteins secreted by primary astrocytes from wild type, apoE (-/-), and human apoE transgenic mice. J Biol Chem 274:30001–30007. https://doi.org/10.1074/JBC.274.42.30001
Fryer JD, Taylor JW, DeMattos RB, Bales KR, Paul SM, Parsadanian M et al (2003) Apolipoprotein E markedly facilitates age-dependent cerebral amyloid angiopathy and spontaneous hemorrhage in amyloid precursor protein transgenic mice. J Neurosci 23:7889–7896. https://doi.org/10.1523/JNEUROSCI.23-21-07889.2003
Gärtner U, Janke C, Holzer M, Vanmechelen E, Arendt T (1998) Postmortem changes in the phosphorylation state of tau-protein in the rat brain. Neurobiol Aging 19:535–543. https://doi.org/10.1016/S0197-4580(98)00094-3
Ginsberg SD, Che S, Counts SE, Mufson EJ (2006) Shift in the ratio of three-repeat tau and four-repeat tau mRNAs in individual cholinergic basal forebrain neurons in mild cognitive impairment and Alzheimer’s disease. J Neurochem 96:1401–1408. https://doi.org/10.1111/J.1471-4159.2005.03641.X
Glenner GG, Wong CW (1984) Alzheimer’s disease: initial report of the purification and characterization of a novel cerebrovascular amyloid protein. Biochem Biophys Res Commun 120:885–890. https://doi.org/10.1016/s0006-291x(84)80190-4
Greenberg SG, Davies P, Schein JD, Binder LI (1992) Hydrofluoric acid-treated tau PHF proteins display the same biochemical properties as normal tau. J Biol Chem 267:564–569. https://doi.org/10.1016/S0021-9258(18)48531-6
Grundke-Iqbal I, Iqbal K, Quinlan M, Tung Y, Zaidi M, Wisniewski H (1986) Microtubule-associated protein tau. A component of Alzheimer paired helical filaments. J Biol Chem 261:6084–6089
Grundke-Iqbal I, Iqbal K, Tung YC, Quinlan M, Wisniewski HM, Binder LI (1986) Abnormal phosphorylation of the microtubule-associated protein tau (tau) in Alzheimer cytoskeletal pathology. 83:4913–4917. https://doi.org/10.1073/pnas.83.13.4913
Hochmair J, Exner C, Franck M, Dominguez‐Baquero A, Diez L, Brognaro H, Kraushar ML, Mielke T, Radbruch H, Kaniyappan S, Falke S, Mandelkow E, Betzel C, Wegmann S (2022) Molecular crowding and RNA synergize to promote phase separation, microtubule interaction, and seeding of Tau condensates. EMBO J https://doi.org/10.15252/EMBJ.2021108882/SUPPL_FILE/EMBJ2021108882-SUP-0002-MOVIEEV1.ZIP
Holtzman DM, Bales KR, Tenkova T, Fagan AM, Parsadanian M, Sartorius LJ et al (2000) Apolipoprotein E isoform-dependent amyloid deposition and neuritic degeneration in a mouse model of Alzheimer’s disease. Proc Natl Acad Sci U S A 97:2892–2897. https://doi.org/10.1073/PNAS.050004797/ASSET/63D22B0F-2005-4799-9F0A-55761F108B24/ASSETS/GRAPHIC/PQ0500047005.GIF
Hsieh YC, Guo C, Yalamanchili HK, Abreha M, Al-Ouran R, Li Y et al (2019) Tau-mediated disruption of the spliceosome triggers cryptic RNA splicing and neurodegeneration in Alzheimer’s disease. Cell Rep 29:301-316.e10. https://doi.org/10.1016/J.CELREP.2019.08.104
Inneraritys TL, Friedlander EJ, Rall SC, Weisgraber KH, Mahley RW (1983) The receptor-binding domain of human apolipoprotein E. Binding of apolipoprotein E fragments. J Biol Chem 258:12341–12347. https://doi.org/10.1016/S0021-9258(17)44180-9
Kanaan NM, Hamel C, Grabinski T (2020) Combs B (2020) Liquid-liquid phase separation induces pathogenic tau conformations in vitro. Nat Commun 111(11):1–16. https://doi.org/10.1038/s41467-020-16580-3
Kavanagh T, Halder A, Drummond E (2022) Tau interactome and RNA binding proteins in neurodegenerative diseases. Mol Neurodegener. https://doi.org/10.1186/S13024-022-00572-6
Koutsodendris N, Blumenfeld J, Agrawal A, Traglia M, Grone B, Zilberter M, Yip O, Rao A, Nelson MR, Hao Y, Thomas R, Yoon SY, Arriola P, Huang Y (2023) Neuronal APOE4 removal protects against tau-mediated gliosis, neurodegeneration and myelin deficits. https://doi.org/10.1038/s43587-023-00368-3
Lester E, Van Alstyne M, McCann KL, Reddy S, Cheng LY, Kuo J et al (2023) Cytosolic condensates rich in polyserine define subcellular sites of tau aggregation. Proc Natl Acad Sci USA. https://doi.org/10.1073/PNAS.2217759120
Lester E, Ooi FK, Bakkar N, Ayers J, Woerman AL, Wheeler J et al (2021) Tau aggregates are RNA-protein assemblies that mislocalize multiple nuclear speckle components. Neuron 109:1675-1691.e9. https://doi.org/10.1016/J.NEURON.2021.03.026
Lin Y, McCarty J, Rauch JN, Delaney KT, Kosik KS, Fredrickson GH et al (2019) Narrow equilibrium window for complex coacervation of tau and RNA under cellular conditions. Elife. https://doi.org/10.7554/ELIFE.42571
Litvinchuk A, Suh JH, Guo JL, Lin K, Davis SS, Bien-Ly N et al (2023) Amelioration of Tau and ApoE4-linked glial lipid accumulation and neurodegeneration with an LXR agonist. Neuron. https://doi.org/10.1016/J.NEURON.2023.10.023
Liu C, Song X, Nisbet R, Götz J (2016) Co-immunoprecipitation with Tau Isoform-specific antibodies reveals distinct protein interactions and highlights a putative role for 2N tau in disease. J Biol Chem 291:8173–8188. https://doi.org/10.1074/JBC.M115.641902
Malek-Ahmadi M, Perez SE, Chen K, Mufson EJ (2020) Braak stage, cerebral amyloid angiopathy, and cognitive decline in early Alzheimer’s disease. J Alzheimers Dis 74:189. https://doi.org/10.3233/JAD-191151
Maziuk BF, Apicco DJ, Cruz AL, Jiang L, Ash PEA, da Rocha EL et al (2018) RNA binding proteins co-localize with small tau inclusions in tauopathy. Acta Neuropathol Commun 6:71. https://doi.org/10.1186/S40478-018-0574-5
McMillan PJ, Benbow SJ, Uhrich R, Saxton A, Baum M, Strovas T et al (2023) Tau-RNA complexes inhibit microtubule polymerization and drive disease-relevant conformation change. Brain 146:3206–3220. https://doi.org/10.1093/BRAIN/AWAD032
Meier S, Bell M, Lyons DN, Ingram A, Chen J, Gensel JC et al (2015) Identification of novel tau interactions with endoplasmic reticulum proteins in Alzheimer’s disease brain. J Alzheimers Dis 48:687–702. https://doi.org/10.3233/JAD-150298
Montal V, Diez I, Kim CM, Orwig W, Bueichekú E, Gutiérrez-Zúñiga R et al (2022) Network Tau spreading is vulnerable to the expression gradients of APOE and glutamatergic-related genes. Sci Transl Med. https://doi.org/10.1126/SCITRANSLMED.ABN7273
Montine TJ, Phelps CH, Beach TG, Bigio EH, Cairns NJ, Dickson DW et al (2012) National institute on aging-Alzheimer’s association guidelines for the neuropathologic assessment of Alzheimer’s disease: a practical approach. Acta Neuropathol 123:1–11. https://doi.org/10.1007/s00401-011-0910-3
Mori H, Kondo J, Ihara Y (1987) Ubiquitin is a component of paired helical filaments in Alzheimer’s disease. Science 235:1641–1644. https://doi.org/10.1126/SCIENCE.3029875
Namba Y, Tomonaga M, Kawasaki H, Otomo E, Ikeda K (1991) Apolipoprotein E immunoreactivity in cerebral amyloid deposits and neurofibrillary tangles in Alzheimer’s disease and kuru plaque amyloid in Creutzfeldt-Jakob disease. Brain Res 541:163–166
Nelson PT, Alafuzoff I, Bigio EH, Bouras C, Braak H, Cairns NJ et al (2012) Correlation of Alzheimer disease neuropathologic changes with cognitive status: a review of the literature. J Neuropathol Exp Neurol 71:362–381
Nichols JB, Malek-Ahmadi M, Tariot PN, Serrano GE, Sue LI, Beach TG (2021) Vascular lesions, APOE ε4, and tau pathology in Alzheimer disease. J Neuropathol Exp Neurol 80:240. https://doi.org/10.1093/JNEN/NLAA160
Ono M, Komatsu M, Ji B, Takado Y, Shimojo M, Minamihisamatsu T et al (2022) Central role for p62/SQSTM1 in the elimination of toxic tau species in a mouse model of tauopathy. Aging Cell. https://doi.org/10.1111/ACEL.13615
Perry G, Friedman R, Shaw G, Chau V (1987) Ubiquitin is detected in neurofibrillary tangles and senile plaque neurites of Alzheimer disease brains. Proc Natl Acad Sci USA 84:3033–3036. https://doi.org/10.1073/PNAS.84.9.3033
Piatnitskaia S, Takahashi M, Kitaura H, Katsuragi Y, Kakihana T, Zhang L et al (2019) Fujii M (2019) USP10 is a critical factor for Tau-positive stress granule formation in neuronal cells. Sci Rep 91(9):1–15. https://doi.org/10.1038/s41598-019-47033-7
Pires G, McElligott S, Drusinsky S, Halliday G, Potier MC, Wisniewski T et al (2019) Secernin-1 is a novel phosphorylated tau binding protein that accumulates in Alzheimer’s disease and not in other tauopathies. Acta Neuropathol Commun. https://doi.org/10.1186/S40478-019-0848-6
Pires G, Ueberheide B, Wisniewski T, Drummond E (2023) Use of affinity purification-mass spectrometry to identify phosphorylated tau interactors in Alzheimer’s disease. Methods Mol Biol 2561:263–277. https://doi.org/10.1007/978-1-0716-2655-9_14
Rabin JS, Nichols E, La Joie R, Casaletto KB, Palta P, Dams-O’Connor K et al (2022) Cerebral amyloid angiopathy interacts with neuritic amyloid plaques to promote tau and cognitive decline. Brain 145:2823–2833. https://doi.org/10.1093/BRAIN/AWAC178
Rebeck WG, Reiter JS, Strickland DK, Hyman BT (1993) Apolipoprotein E in sporadic Alzheimer’s disease: allelic variation and receptor interactions. Neuron 11:575–580. https://doi.org/10.1016/0896-6273(93)90070-8
Roberts JA, Varma VR, An Y, Varma S, Candia J, Fantoni G et al (2021) A brain proteomic signature of incipient Alzheimer’s disease in young APOE ε4 carriers identifies novel drug targets. Sci Adv 7:8178. https://doi.org/10.1126/SCIADV.ABI8178
Saito H, Dhanasekaran P, Baldwin F, Weisgraber KH, Lund-Katz S, Phillips MC (2001) Lipid binding-induced conformational change in human apolipoprotein E. Evid Two Lipid-Bound States Spher Particles. https://doi.org/10.1074/jbc.M106337200
Saroja SR, Gorbachev K, Tcw J, Goate AM, Pereira AC (2022) Astrocyte-secreted glypican-4 drives APOE4-dependent tau hyperphosphorylation. Proc Natl Acad Sci USA. https://doi.org/10.1073/PNAS.2108870119
Schmechel DE, Saunders AM, Strittmatter WJ, Crain BJ, Hulette CM, Joo SH et al (1993) Increased amyloid beta-peptide deposition in cerebral cortex as a consequence of apolipoprotein E genotype in late-onset Alzheimer disease. Proc Natl Acad Sci 90:9649–9653. https://doi.org/10.1073/PNAS.90.20.9649
Schmechel DE, Saunders AM, Strittmatter WJ, Crain BJ, Hulette CM, Joo SH et al (1993) Increased amyloid beta-peptide deposition in cerebral cortex as a consequence of apolipoprotein E genotype in late-onset Alzheimer disease. Proc Natl Acad Sci U S A 90:9649. https://doi.org/10.1073/PNAS.90.20.9649
Schmidt ML, Lee VM, Trojanowski JQ (1991) Comparative epitope analysis of neuronal cytoskeletal proteins in Alzheimer’s disease senile plaque neurites and neuropil threads. Lab Invest 64:352–357
Sepulveda-Falla D, Sanchez JS, Almeida MC, Boassa D, Acosta-Uribe J, Vila-Castelar C et al (2022) Distinct tau neuropathology and cellular profiles of an APOE3 Christchurch homozygote protected against autosomal dominant Alzheimer’s dementia. Acta Neuropathol 144:589–601. https://doi.org/10.1007/S00401-022-02467-8
Shi Y, Yamada K, Liddelow SA, Smith ST, Zhao L, Luo W et al (2017) ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 549:523–527. https://doi.org/10.1038/nature24016
Steward A, Biel D, Dewenter A, Roemer S, Wagner F, Dehsarvi A et al (2023) ApoE4 and connectivity-mediated spreading of Tau pathology at lower amyloid levels. JAMA Neurol 80:1295–1306. https://doi.org/10.1001/JAMANEUROL.2023.4038
Strittmatter SA, Goedert M, Weisgraber K, Dong L, Jakes R, Huang D et al (1994) Isoform-specific interactions of apolipoprotein E with microtubule-associated protein tau: implications for Alzheimer disease. Proc Natl Acad Sci 91:11183–11186. https://doi.org/10.1073/PNAS.91.23.11183
Strittmatter W, Saunders A, Schmechel D, Pericak-vance M, Enghild J, Salvesen G et al (1993) Apolipoprotein E: high-avidity binding to, B-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.90.5.1977
Teo G, Liu G, Zhang J, Nesvizhskii AI, Gingras AC, Choi H (2014) SAINTexpress: improvements and additional features in Significance Analysis of Interactome software. J Proteom 100:37. https://doi.org/10.1016/J.JPROT.2013.10.023
Terry RD, Gonatas NK, Weiss M (1964) Ultrastructural studies in Alzheimer’s presenile dementia. Am J Pathol 44:269–297
Thal DR, Ghebremedhin E, Rüb U, Yamaguchi H, Del Tredici K, Braak H (2002) Two types of sporadic cerebral amyloid angiopathy. J Neuropathol Exp Neurol 61:282–293. https://doi.org/10.1093/jnen/61.3.282
Thal DR, Papassotiropoulos A, Saido TC, Griffin WST, Mrak RE, Kölsch H et al (2010) Capillary cerebral amyloid angiopathy identifies a distinct APOE epsilon4-associated subtype of sporadic Alzheimer’s disease. Acta Neuropathol 120:169–183. https://doi.org/10.1007/S00401-010-0707-9
Thal DR, Rüb U, Orantes M, Braak H (2002) Phases of Aβ-deposition in the human brain and its relevance for the development of AD. Neurology 58:1791–1800
Thierry M, Boluda S, Delatour B, Marty S, Seilhean D, Letournel F et al (2020) Human subiculo-fornico-mamillary system in Alzheimer’s disease: Tau seeding by the pillar of the fornix. Acta Neuropathol. https://doi.org/10.1007/s00401-019-02108-7
Thierry M, Marty S, Boluda S, Duyckaerts C (2017) Alzheimer’s senile plaque as shown by microcryodissection, a new technique for dissociating tissue structures. J Neural Transm 124:685–694. https://doi.org/10.1007/s00702-017-1718-7
Tracy TE, Madero-Pérez J, Swaney DL, Chang TS, Moritz M, Konrad C et al (2022) Tau interactome maps synaptic and mitochondrial processes associated with neurodegeneration. Cell 185:712-728.e14. https://doi.org/10.1016/J.CELL.2021.12.041
Wang C, Xiong M, Gratuze M, Bao X, Shi Y, Andhey PS et al (2021) Selective removal of astrocytic APOE4 strongly protects against tau-mediated neurodegeneration and decreases synaptic phagocytosis by microglia. Neuron 109:1657-1674.e7. https://doi.org/10.1016/J.NEURON.2021.03.024
Wang P, Joberty G, Buist A, Vanoosthuyse A, Stancu IC, Vasconcelos B et al (2017) Tau interactome mapping based identification of Otub1 as Tau deubiquitinase involved in accumulation of pathological Tau forms in vitro and in vivo. Acta Neuropathol 133:731–749. https://doi.org/10.1007/S00401-016-1663-9
Wang Y, Balaji V, Kaniyappan S, Krüger L, Irsen S, Tepper K et al (2017) The release and trans-synaptic transmission of Tau via exosomes. Mol Neurodegener 12:5. https://doi.org/10.1186/s13024-016-0143-y
Wisniewski T, Castaño EM, Golabek A, Vogel T, Frangione B (1994) Acceleration of Alzheimer’s fibril formation by apolipoprotein E in vitro. Am J Pathol 145:1030
Wisniewski T, Drummond E (2020) APOE-amyloid interaction: therapeutic targets. Neurobiol Dis. https://doi.org/10.1016/j.nbd.2020.104784
Wisniewski T, Frangione B (1992) Apolipoprotein E: a pathological chaperone protein in patients with cerebral and systemic amyloid. Neurosci Lett 135:235–238. https://doi.org/10.1016/0304-3940(92)90444-C
Woo JA, Yan Y, Kee TR, Cazzaro S, Percy KCMG, Wang X, Liu T, Liggett SB, Kang DE (2021) β-arrestin1 promotes tauopathy by transducing GPCR signaling, disrupting microtubules and autophagy. Life Sci Alliance https://doi.org/10.26508/LSA.202101183
Wu JW, Hussaini SA, Bastille IM, Rodriguez GA, Mrejeru A, Rilett K et al (2016) Neuronal activity enhances tau propagation and tau pathology in vivo. Nat Neurosci 19:1085–1092. https://doi.org/10.1038/nn.4328
Zhang K, Daigle JG, Cunningham KM, Coyne AN, Ruan K, Grima JC et al (2018) Stress granule assembly disrupts nucleocytoplasmic transport. Cell 173:958-971.e17. https://doi.org/10.1016/J.CELL.2018.03.025
Zhang X, Lin Y, Eschmann NA, Zhou H, Rauch JN, Hernandez I et al (2017) RNA stores tau reversibly in complex coacervates. PLOS Biol 15:e2002183. https://doi.org/10.1371/JOURNAL.PBIO.2002183
Acknowledgements
This manuscript was supported by NIH grants P01AG060882 and P30AG066512 (to T.W.), and funding from the Bluesand Foundation (to E.D.). We thank the Columbia University Alzheimer’s Disease Research Center, funded by NIH grant P30AG066462 to S.A. Small (P.I.), and A. Teich for providing biological samples and associated information. We also thank A. Cardiel and J. Xie for their technical assistance with this project.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thierry, M., Ponce, J., Martà-Ariza, M. et al. The influence of APOEε4 on the pTau interactome in sporadic Alzheimer’s disease. Acta Neuropathol 147, 91 (2024). https://doi.org/10.1007/s00401-024-02744-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00401-024-02744-8