Abstract
Cotton wool plaques (CWPs) have been described as features of the neuropathologic phenotype of dominantly inherited Alzheimer disease (DIAD) caused by some missense and deletion mutations in the presenilin 1 (PSEN1) gene. CWPs are round, eosinophilic amyloid-β (Aβ) plaques that lack an amyloid core and are recognizable, but not fluorescent, in Thioflavin S (ThS) preparations. Amino-terminally truncated and post-translationally modified Aβ peptide species are the main component of CWPs. Tau immunopositive neurites may be present in CWPs. In addition, neurofibrillary tangles coexist with CWPs. Herein, we report the structure of Aβ and tau filaments isolated from brain tissue of individuals affected by DIAD caused by the PSEN1 V261I and A431E mutations, with the CWP neuropathologic phenotype. CWPs are predominantly composed of type I Aβ filaments present in two novel arrangements, type Ic and type Id; additionally, CWPs contain type I and type Ib Aβ filaments. Tau filaments have the AD fold, which has been previously reported in sporadic AD and DIAD. The formation of type Ic and type Id Aβ filaments may be the basis for the phenotype of CWPs. Our data are relevant for the development of PET imaging methodologies to best detect CWPs in DIAD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer disease (AD) is a neurodegenerative disorder characterized by the extracellular deposition of the insoluble 4 kDa amyloid-β (Aβ) peptide and the intracellular aggregation of tau [13]. In AD, Aβ is the major component of amyloid plaques while the protein tau is the main constituent of neurofibrillary tangles (NFTs) [3, 11]. Different types of plaques may be observed in the various forms of AD, and their morphology may also be variable relative to anatomical brain areas [3, 5, 30]. The morphological classification of Aβ plaques as dense cored, neuritic, and diffuse has been redefined by immunohistochemical and biochemical methodologies as well as a detailed brain mapping of plaque distribution [3]. The most common Aβ plaques are dense cored and neuritic; the latter are characterized by a central amyloid core surrounded by a crown of dystrophic neurites [3, 5]. Diffuse plaques are not associated with a neuritic component and vary in shape and size [3, 28].
Approximately 10–15% of early onset AD (EOAD) cases are dominantly inherited Alzheimer disease (DIAD) and are associated with mutations in the Amyloid β Precursor Protein (AβPP), Presenilin 1 (PSEN1) and Presenilin 2 (PSEN2) genes [2, 6, 13]. The neuropathologic phenotypes of AD associated with PSEN1 mutations are not consistent. A distinct neuropathologic phenotype associated with several missense and deletion mutations in PSEN1, is characterized by the presence of cotton wool plaques (CWPs) as the predominant Aβ plaque [4, 15, 21]. CWPs are round eosinophilic plaques that lack a central core and are recognizable, but not fluorescent, in Thioflavin S (Th-S) preparations [1, 4, 15, 16]. Aβ is a 40 or 42/43-amino acid peptide generated by the successive proteolysis of AβPP by the β-site AβPP-cleaving enzyme 1 (BACE1) and the γ-secretase complex [33]. After β-cleavage, the carboxyl terminal fragment of AβPP, known as CTFβ, remains membrane associated and is further cleaved by γ-secretase releasing Aβ species of varying lengths [26, 33]. CWPs are composed predominantly of highly insoluble amino-terminally truncated Aβ peptide species, with or without a pyroglutamyl residue at positions Glu3 and Glu11, ending at position Ala42 or Thr43; full-length Aβ1–42 and Aβ1–43 peptide species are present at a significantly lower quantity and there is an absence of Aβ1–40 [15]. The absence of Aβ1–40 in CWPs suggests that they are not associated at all with vascular structures.
In AD, parenchymal deposition of Aβ has been linked to the development of cytosolic tau pathology in a causal relationship. NFTs are made of paired helical filaments (PHFs) and straight filaments (SF) that share a common structural subunit [8]. Both filaments contain all six tau isoforms (3R + 4R), having three isoforms with three microtubule-binding repeats (3R tau) and three isoforms with four microtubule-binding repeats (4R tau) [10]. In PHFs and SFs, residues 306–378 of tau make up the core of protofilaments which forms a base for the incorporation of both 3R and 4R tau into the filament [8].
Herein, we used cryogenic-electron microscopy (cryo-EM) to characterize Aβ and tau filaments extracted from the brain tissue of two individuals with DIAD with the CWP neuropathologic phenotype. Our work provides novel insights into the role that different types of Aβ filaments may play in the development of a particular plaque morphology and how this may affect binding of compounds for in vivo visualization and quantification of amyloid pathology.
Materials and methods
Clinical history and neuropathology
We studied two individuals with DIAD. Case #1 was a 55-year-old female carrying the PSEN1 V261I mutation, homozygous for the apolipoprotein E (APOE) ε3 allele [15]. The patient died with neuropathologically confirmed diagnosis of AD after a 7-year history of progressive dementia. A brain autopsy was carried out. Case #2 was a 50-year-old male carrying the PSEN1 A431E mutation and the APOE ε3/ε3 genotype who died with neuropathologically confirmed diagnosis of AD after a 9-year history of progressive dementia. A brain autopsy was carried out.
Tissue samples for neuropathological studies were obtained from representative brain regions. Eight-μm thick brain sections were used. Sections were stained with hematoxylin and eosin (H&E) and Luxol-fast blue (LFB). For immunohistochemistry of Aβ, primary antibodies were 21F12 (provided as gift by Elan, at 1:1000) and NAB228 (Thermo Fisher Scientific 37–4200, 1:300). For tau, the primary antibody was AT8 (Thermo Fisher Scientific MN1020, 1:300). The signal from the antibodies was visualized using avidin–biotin followed by horseradish peroxidase-conjugated streptavidin and the chromogen diaminobenzidine. Immunohistochemical sections were counterstained with hematoxylin to show nucleus and cytoplasmic structure.
Filament extraction
Sarkosyl-insoluble fractions were prepared from the cerebral cortex of freshly frozen frontal lobes as previously described [8, 11]. Briefly, ~ 4 g of gray matter was homogenized in A68 extraction buffer consisting of 10 mM Tris–HCl, pH 7.4, 0.8 M NaCl, 1 mM EGTA, 5 mM EDTA, and 10% sucrose with protease and phosphatase inhibitors. Samples were centrifuged at 20,000×g and the supernatants brought to 1% sarkosyl. Supernatants were incubated at room temperature (RT) while shaking. After centrifugation at 100,000×g/1 h/4 °C, the sarkosyl-insoluble pellets were resuspended in 10 μl/g tissue 50 mM Tris–HCl, pH 7.4. The pellet was diluted in A68 extraction buffer and centrifugated at 20,000×g/30 min/4 °C. The supernatant was centrifuged at 100,000×g/1 h/4 °C and the final pellet resuspended in 20 mM Tris–HCl, pH 7.4, with 100 mM NaCl, and stored at 4 °C.
Western blot analysis
Samples were sonicated for 1 min, boiled with gel 2 × Laemmli sample buffer (Bio-Rad) for 5 min at 100 °C and resolved on 4–12% Bis–Tris gels for Aβ amyloid or 10% Bis–Tris gels for tau (NUPAGE). For hexafluoroisopropanol (HFIP) treatment, samples were centrifuged at 200,000g for 30 min and HFIP added to the pellet, sonicated and left overnight at 37 °C. Samples were dried under nitrogen, washed 3 times with water and centrifuged at 200,000×g for 30 min. The final pellet was resuspended in loading buffer and resolved on 4–12% Bis–Tris gels. Proteins were transferred to nitrocellulose membranes and the membranes were incubated with blocking solution (5% non-fat milk in PBS with 0.1% Tween 20). Membranes were incubated for 1 h with primary antibody diluted in TBS. Antibodies used were 4G8 (Biolegend, 1:1000), 6E10 (Biolegend, 1:1000), GT622 (Thermo Fisher, 1:1000), AT8 (Thermo Fisher, 1:1000), and HT7 (Thermo Fisher, 1:1000). After incubation with secondary antibody for 45 min, proteins were visualized using a chemiluminescence kit (SuperSignal West Pico, ThermoFisher) according to the manufacturer's specifications.
Isolation of parenchymal amyloid peptides
Approximately 5 g of frontal cortex from case #1 (PSEN1 V261I mutation) was dissected free of any large vessel contamination. Amyloid peptides were isolated using a published procedure [15]. For matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF–MS) analysis, samples were resuspended with 10 uL of an isopropyl alcohol/water/formic acid (4:4:1) mixture and analyzed at NYU Langone’s Proteomics Laboratory.
High‑resolution cryo‑EM imaging
Cryo-EM grids of brain extracts of the two DIAD patients (Case #1 PSEN1 V261I and Case#2 PSEN1 A431E) were prepared in a biosafety level 2 cabinet while wearing appropriate personal protective equipment. 2–3 µl of the sample were applied on a graphene oxide coated EM grid, then washed with 10 mM Tris, pH 7.8, before vitrifying using a semi-automated Gatan CP3 cryo-plunger. High resolution cryo-EM movies were collected on a FEI Titan Krios at 300 kV with a Gatan K3 detector mounted on a Quantum energy filter with 20 eV slit width (Table S1). For case #1, we recorded 6,183 movies of consisting of 40 frames for a total accumulated dose of 57.43 electrons per Å2 at a pixel size of 1.054 Å. For case #2, we recorded 17,581 movies of 40 frames for a total accumulated dose of 57.79 electrons per Å2 at a pixel size of 1.054 Å. The datasets were collected with defocus values ranging from − 0.5 to − 1.5 μm. The movies were gain-corrected, motion-corrected and dose-weighted using RELION’s own implementation of motion correction.
The contrast transfer function (CTF) of all aligned and non-dose-weighted micrographs was estimated using CTFFIND-4.1.
Helical reconstruction
Image processing was performed in RELION 4.1 [22]. Filaments were picked manually and manually and automatically using RELION helical picker as end-to-end line segments. We extracted all the helical segments with a box size of 600 pixel (632 Å), down scaled to 200 pixels to speed up analysis and an inter-box distance of ~ 15 Å. Several rounds of reference-free 2D classifications were carried out to find homogeneous subsets using a regularization value of T = 1–2. Aβ and tau filaments were visually identified from the reference 2D class averages. For both, a new 256-pixel box size without downscaling was used to re-extract with inter-box distance of approximately 15 Å. The initial 3D reference maps were reconstructed de novo from best 2D class averages comprising a full helical crossover using relion_ini_model. The initial round of 3D classification was low-pass-filtered to 10 Å. Several rounds of 3D classification were carried out to obtain the best homogeneous subset. The final selected segments were used for final 3D auto-refinement with optimization of the helical twist and rise to yield a 3D map showing clearly visible beta-strand separation and side-chain densities. We used a 10% helical z percentage parameter for post-reconstruction application of helical symmetry. The final reconstructions were sharpened using the standard post-processing procedures in RELION. The overall resolution was calculated from Fourier shell correlations at 0.143 between two independently refined half-maps, employing phase randomization for the convolution effects correction of an optimized, soft-edged solvent mask as implemented in the trueFSC.py program in JSPR software (Fig. S4) [28].
Atomic modeling and structural analysis
The previously deposited model was fitted into the sharpened density maps using ChimeraX (https://doi.org/https://doi.org/10.1002/pro.3235, https://doi.org/https://doi.org/10.1002/pro.3943). The central chain of each model was manually adjusted in Coot (https://doi.org/https://doi.org/10.1107/S0907444904019158). The atomic positions of all models were refined with their respective helical symmetry parameters using Rosetta (https://doi.org/https://doi.org/10.7554/eLife.17219). Final atomic models were validated using MolProbity (https://doi.org/https://doi.org/10.1107/S0907444909042073) (Table S2). All models’ figures were generated in ChimeraX [18].
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession numbers 46417 (PHF tau), 46420 (SF tau), 46422 (Type Ic Aβ) and 46424 (Type Id Aβ). Refined atomic models have been deposited in the Protein Data Bank (PDB) under accession numbers 9CZI (PHF tau), 9CZL (SF tau), 9CZN (Type Ic Aβ) and 9CZP (Type Id Aβ).
Results
CWPs in two individuals affected by DIAD, carrying the PSEN1 V261I or the PSEN1 A431E mutations
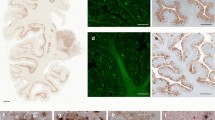
Case #1 was a 55-year-old female who is heterozygous for the PSEN1 V261I mutation and homozygous for the Apolipoprotein E (APOE) ε3 allele [15]. Case #2 was a 50-year-old male heterozygous for the PSEN1 A431E mutation and homozygous for the APOE ε3 allele. The PSEN1 A431E mutation is one of the most common mutations associated with DIAD in people of Mexican heritage [16, 20]. The neuropathologic phenotype of these cases was characterized by abundant CWPs and severe tau pathology [15, 16, 20]. CWPs were immunopositive using antibodies that recognize Aβ. Tau inclusions, including NFTs, dystrophic neurites, and neurites associated with the Aβ deposits in CWPs were immunopositive using an antibody against phosphorylated tau (Fig. 1).
Neurohistology and immunohistochemistry. Sections of the frontal cortex of case #1 (V261I, panels a, b, c) and case #2 (A431E, panels d, e, f), contralateral to the ones used for isolating amyloid fibrils. Numerous well-demarcated CWPs are seen (a, d) that are immunopositive using antibodies against the Aβ peptide (b, e) and contain tau-immunoreactive neurites (c, f). H&E stain (a), LFB (d), immunohistochemistry using antibodies 21F12 (b), nab228 (e) and the anti-tau antibody AT8 (c, f). Bar 100 μm
Structure of Aβ filaments from CWPs
Immunoblotting of sarkosyl-insoluble fractions from the gray matter of each of the two DIAD cases using Aβ antibodies reveals the presence of high molecular weight bands most likely representing aggregated Aβ filaments. Aβ monomers (4 kDa) and dimers (8 kDa) were evident after treatment of the samples with hexafluoro isopropanol (HFIP) to disaggregate Aβ (Fig. S1a). MALDI-TOF–MS of purified Aβ peptides from parenchymal deposits of the PSEN1 V261I case shows the presence of amino-terminally truncated Aβ3–42, Aβ3–43, Aβ11–42 and Aβ11-43 Aβ species and Aβ peptides with a pyroglutamyl residue at position Glu3 or Glu 11 as the predominant Aβ peptide species (Fig. S1b).
The structures of Aβ and tau filaments were identified by cryo-EM. 2D classifications revealed the presence of four types of Aβ filaments of variable widths and cross-over distances in each one of the samples (Figure S2). Type I and type Ib Aβ filaments have been previously reported in sAD cases [31]. In addition, we observed two novel arrangements of type I Aβ filaments, displaying unique packing of the protofilaments that we named type Ic and type Id Aβ filaments (Fig. 2).
Type I Aβ filaments represent approximately between 2 and 5% of the Aβ filaments. They are composed of two protofilaments arranged around a 21 screw symmetry axis with a crossover distance of ~ 280 Å. Each protofilament has vertically stacked S-shaped protomers with a spacing of 4.76 Å and consists of residues Gly9 to Ala42 (Fig. S3). Type Ib Aβ filaments were reported to be a small fraction of the total Aβ filaments in sAD [29]; in the two DIAD cases with CWPs, they represent ~ 2% of the Aβ filaments. Type Ib Aβ filaments are made of two type I Aβ filaments packed side by side with a twofold rotational symmetry (Fig. 2). The solvent-exposed negative charges of Glu22 and Asp23 in type I are now offset in type Ib through their interaction with Lys16 and His14, respectively, at the interface with the adjacent dimer. Type Ib Aβ filaments could only be reconstructed to a relatively low resolution, likely due to the low number of type Ib Aβ filaments in the samples and their intrinsic flexibility and heterogeneity (Fig. 2).
Type Ic Aβ filaments were the predominant type of Aβ filaments, representing between ~ 70 and 80% of the Aβ filaments (Table S1). Type Ic Aβ filaments consist of two type I Aβ filaments related by a twofold screw axis (Fig. 2). The two dimers appear to be arranged diagonally such that the side chain of Lys28 interacts with the terminal carboxyl group of Ala42 in the adjacent dimer (Fig. 3a). Notably, even though the hydrophobic cluster and amino acid configurations appear to be like those of type I Aβ filaments, there are subtle differences in the packing of the S-shaped monomers within each dimer. For example, in type I Aβ filaments, the Cα of residues Leu34 from one chain is ~ 11 Å from Val34 of the neighboring chain. However, in type Ic Aβ filaments they are at a distance of ~ 17 Å, suggesting minor differences in the packing of the S-shaped protomers (Fig. 4). These subtle changes in packing seem to result in a channel filled with non-proteinaceous densities, which are likely hydrophobic host factors co-assembled within type Ic Aβ filaments (Fig. 3a). In addition, the solvent exposed Glu22 and Lys16 in each of the 4 chains of type Ic Aβ filaments forms an intra-chain salt-bridge, which was not seen in type I Aβ filaments. Type Id Aβ filaments represent between 13 and 18% of the Aβ filaments (Table S1). The two Aβ dimer filaments are antiparallel, similar to the prion protein protofilaments observed in Gerstmann–Sträussler–Scheinker disease and in one of the lipidic alpha-synuclein polymorphs [9, 12], and arranged asymmetrically, in contrast to the twofold packing seen in type Ib and Ic Aβ filaments (Fig. 2). The two dimers interact through an inter-chain salt bridge between Asp23 and Lys28, as seen in the human Artic fold and in murine type III Aβ filaments [32, 34], and a hydrogen bond between interfacial residues His14 and Asp23. Like type Ic Aβ filaments, both dimers also have a central channel filled with non-proteinaceous densities (Fig. 3b). Type II Aβ filaments, the predominant type of Aβ filaments in two cases of DIAD which lack the presence of CWPs, and in cases in which diffuse deposits of Aβ are present as co-pathology [31], were not observed. At a single dimer level, type Id Aβ filaments appear similar to the tg-APPseArc structure [34]; however, the tg-APPseArc filament consist of a single dimer, while type Id filaments consist of two dimers packaged asymmetrically in a rarely observed anti-parallel fashion.
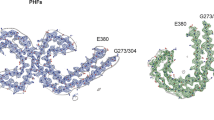
Aβ type Ic and Id filaments in DIAD brains. (a) Atomic model of type Ic Aβ filaments and (b) type Id Aβ filaments. Type Ic and Id Aβ filaments are made of two identical type I Aβ filaments protofilaments, extending from Gly9 to Ala42. Cartoon representation of amino acid residues: hydrophobic (white), positively charged (teal), polar (green), negatively charged (purple), and glycine (pink) residues
Comparison of Type I, Ic and Id Aβ filaments. Superposition of type I (orange), Ic (blue) and Id (yellow) Aβ filaments, based on the central layer of their S-shaped domains. The Cα of residues Leu34 from one chain is ~ 11 Å from Val34 of the neighboring chain in type I Aβ filaments, while in type Ic and Id Aβ filaments they are at a distance of ~ 17 Å and ~ 18 Å, respectively, suggesting minor differences in the packing of the S-shaped protomers
Structures of tau filaments
Western blot analysis of the sarkosyl-insoluble fractions from the gray matter of cases #1 and #2 using the tau antibody HT7 and the phosphorylation-dependent tau antibody AT8 shows the presence of tau bands with a migration pattern corresponding to 3 + 4R tau, with identical electrophoretic mobility on both cases (Fig. S1). The structure of tau PHFs and SFs (Fig. 5) in both cases were found to be identical to those reported in AD and other amyloidoses [7, 8, 11, 24]. We determined the structure of PHFs at a resolution of 3.0 Å and the structure of SFs at a resolution of 2.9 Å (Fig. 5). PHFs and SFs were similarly present in the sarkosyl-insoluble fractions from the gray matter of cases #1 and #2 (Table S1).
Cryo-EM reconstructions of tau filaments. Cryo-EM maps, depicted as central slices, of tau filaments from the two cases. The structures show identical pairs of C-shaped protofilaments and the symmetric inter-protofilament packing between PHFs, and asymmetric packing for SFs. The estimated resolution is shown on the bottom right. The scale bar, 10 nm, applies to all panels
Discussion
The neuropathologic phenotype of the two individuals affected by DIAD associated with either the V261I or V431E PSEN1 mutations is characterized by the presence of CWPs and neurofibrillary pathology in the cerebral cortex. We determined the cryo-EM structures of Aβ and tau filaments extracted from the cerebral cortex of both DIAD cases. We observed two novel types of Aβ filaments that we named type Ic and type Id Aβ filaments representing ~ 90% of the total Aβ filaments. Type I and type Ib Aβ filaments, previously seen in AD [31], were also present. Tau filaments had the AD fold and were identical to PHFs and SFs previously reported in sAD, DIAD, familial British dementia, familial Danish dementia, Gerstmann–Sträussler–Scheinker disease, Prion-Protein cerebral amyloid angiopathy and Down syndrome (DS) [7, 8, 11, 24, 32].
Previous cryo-EM studies of Aβ filaments from core plaques showed that type I Aβ filaments are the predominant type of filaments in sAD while type II Aβ filaments are found predominantly in cases of DIAD (AβPP V717F and PSEN1 F105L mutations) as well as in other neurodegenerative diseases in association with parenchymal Aβ deposition in the form of diffuse deposits [31]. No correlation between the type of Aβ filaments and the APOE genotype, the most established genetic risk factor for sAD [27], has been observed [31]. Recently, amyloid deposits in DS were shown to have type I and type II Aβ filaments, consistent with their presence in cored plaques and diffuse deposits in DS [7]. Type Ic and Id Aβ filaments are the predominant type of Aβ filaments in association with CWPs. Apart from some small differences on the packing of the S-shape monomers, the amino acid configuration of type Ic Aβ filaments is very similar to the one observed for type I Aβ filaments [31]. Type Id Aβ filaments are antiparallel and arranged asymmetrically, in contrast to the twofold parallel packing seen in type Ib and Ic Aβ filaments.
Formation of type Ic and type Id Aβ filaments in CWPs may be associated with the presence of a particular array of Aβ peptide species found in CWPs, which differs from the Aβ peptide species found in core plaques in sAD [15]. CWPs are composed predominantly of amino-terminally truncated Aβ peptide species with or without a pyroglutamyl residue at positions Glu3 and Glu11 and ending at positions 42 and 43. Full-length Aβ1–42 and Aβ1–43 peptide species are underrepresented. Aβ peptides starting with pyroglutamyl at residues Glu3 and Glu11, particularly AβN3pE-42, have been suggested to be early aggregating species in AD [14, 15, 17, 19, 21, 29] and to be more abundant in patients with PSEN1 mutations than in patients with sAD [15, 21]. The relative abundance of different Aβ peptide species may be the basis for the formation of the different types of Aβ filaments and the morphological diversity of amyloid deposits seen in AD (Fig. 6). Concerning tau pathology, the structure of tau filaments (i.e. the AD fold) present in individuals with DIAD and the CWP phenotype does not differ from the structure of tau filaments seen in all other forms of AD (Fig. 6) [8, 24].
Amyloid plaque morphologies and type of Aβ filaments. Core plaques (CP), as seen in sAD and DS, are predominantly associated with the presence of type I and type Ib Aβ filaments, while diffuse plaques (DP), as seen in DIAD cases as well as in DS, are predominantly associated with the presence of type II Aβ filaments. CWPs, seen in DIAD cases in association with a subset of PSEN1 mutations, are predominantly associated with the presence of type Ic and type Id Aβ filaments (a). Regardless of the type of Aβ filaments and independent of the type of amyloid subunit, we observe in the presence of different extracellular amyloid peptides the presence of intraneuronal NFTs composed of PHFs and SFs (b)
The characterization of Aβ filaments from CWPs, which are morphologically different from core plaques and diffuse deposits, is important for the understanding of the role that the structure and assembly of amyloid peptides may play in the morphological diversity of amyloid deposits, tau response, and disease process as well as for developing new diagnostic and therapeutic strategies for AD. While there is good correlation between PET using Pittsburg compound B (PiB) and amyloid load in sAD, underestimation of the total Aβ plaque burden in DIAD cases with CWPs using 11C-PiB PET has been reported [1]. Moreover, PET is negative in individuals with DIAD associated with the Arctic mutation in the AβPP gene, which shows Aβ filaments with a different fold (Arctic fold) and in a Japanese pedigree with an amino acid deletion (p.E693delta) in the same amino acid position as the Artic mutation [19, 23, 25, 32]. The absence or low levels of PiB reactivity in these cases seems to be associated with the particular type of amyloid plaque that lacks an amyloid core [19]. It is of interest to note that in the various forms of sporadic and hereditary AD, tau filaments that coexist with Aβ deposits have a consistent fold, the AD fold, regardless of the cryo-EM structure of the Aβ filament.
References
Abrahamson EE, Kofler JK, Becker CR, Price JC, Newell KL, Ghetti B et al (2022) 11C-PiB PET can underestimate brain amyloid-β burden when cotton wool plaques are numerous. Brain 145(6):2161–2176
Apostolova LG, Aisen P, Eloyan A, Fagan A, Fargo KN, Foroud T et al (2021) The longitudinal early-onset Alzheimer’s disease study (LEADS): framework and methodology. Alzheimers Dement 17(12):2043–2055
Beach TG (2022) A history of senile plaques: from Alzheimer to amyloid imaging. J Neuropathol Exp Neurol 81(6):387–413
Crook R, Verkkoniemi A, Perez-Tur J, Mehta N, Baker M, Houlden H et al (1998) A variant of Alzheimer’s disease with spastic paraparesis and unusual plaques due to deletion of exon 9 of presenilin 1. Nat Med 4(4):452–455
Dickson TC, Vickers JC (2001) The morphological phenotype of beta-amyloid plaques and associated neuritic changes in Alzheimer’s disease. Neuroscience 105(1):99–107
Di Fede G, Catania M, Morbin M, Rossi G, Suardi S, Mazzoleni G et al (2009) A recessive mutation in the APP gene with dominant-negative effect on amyloidogenesis. Science 323(5920):1473–1477
Fernandez A, Hoq MR, Hallinan GI, Li D, Bharath SR, Vago FS et al (2024) Cryo-EM structures of amyloid β and tau filaments in Down syndrome. Nat Struct Mol Biol 31(6):903–909
Fitzpatrick AWP, Falcon B, He S, Murzin AG, Murshudov G, Garringer HJ et al (2017) Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547(7662):185–190
Frieg B, Antonschmidt L, Dienemann C, Geraets JA, Najbauer EE, Matthes D et al (2022) The 3D structure of lipidic fibrils of α-synuclein. Nat Commun 13(1):6810
Goedert M, Spillantini MG, Cairns NJ, Crowther RA (1992) Tau proteins of Alzheimer paired helical filaments: abnormal phosphorylation of all six brain isoforms. Neuron 8:159–168
Hallinan GI, Hoq MR, Ghosh M, Vago FS, Fernandez A, Garringer HJ et al (2021) Structure of Tau filaments in Prion protein amyloidoses. Acta Neuropathol 142(2):227–241
Hallinan GI, Ozcan KA, Hoq MR, Cracco L, Vago FS, Bharath SR et al (2022) Cryo-EM structures of prion protein filaments from Gerstmann–Sträussler–Scheinker disease. Acta Neuropathol 144(3):509–520
Knopman DS, Amieva H, Petersen RC, Chételat G, Holtzman DM, Hyman BT et al (2021) Alzheimer disease. Nat Rev Dis Primers 7(1):33
Kuo YM, Emmerling MR, Woods AS, Cotter RJ, Roher AE (1997) Isolation, chemical characterization, and quantitation of Aβ 3-pyroglutamyl peptide from neuritic plaques and vascular amyloid deposits. Biochem Biophys Res Commun 237:188–191
Miravalle L, Calero M, Takao M, Roher AE, Ghetti B, Vidal R (2005) Amino-terminally truncated Aβ peptide species are the main component of cotton wool plaques. Biochemistry 44(32):10810–10821
Murrell J, Ghetti B, Cochran E, Macias-Islas MA, Medina L, Varpetian A et al (2006) The A431E mutation in PSEN1 causing familial Alzheimer’s disease originating in Jalisco State, Mexico: an additional fifteen families. Neurogenetics 7(4):277–279
Naslund J, Schierhorn A, Hellman U, Lannfelt L, Roses AD, Tjernberg LO et al (1994) Relative abundance of Alzheimer A beta amyloid peptide variants in Alzheimer disease and normal aging. Proc Natl Acad Sci USA 91:8378–8382
Pettersen EF, Goddard TD, Huang CC, Meng EC, Couch GS, Croll TI et al (2020) UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci 30:70–82
Philipson O, Lord A, Lalowski M, Soliymani R, Baumann M, Thyberg J et al (2012) The Arctic amyloid-β precursor protein (AβPP) mutation results in distinct plaques and accumulation of N- and C-truncated Aβ. Neurobiol Aging 33(5):1010.e1–13
Rogaeva EA, Fafel KC, Song YQ, Medeiros H, Sato C, Liang Y et al (2001) Screening for PS1 mutations in a referral-based series of AD cases: 21 novel mutations. Neurology 57(4):621–625
Russo C, Schettini G, Saido TC, Hulette C, Lippa C, Lannfelt L et al (2000) Presenilin-1 mutations in Alzheimer’s disease. Nature 405:531–532
Scheres SHW (2012) A Bayesian view on Cryo-EM structure determination. J Mol Biol 415(2):406–418
Schöll M, Wall A, Thordardottir S, Ferreira D, Bogdanovic N, Långström B et al (2012) Low PiB PET retention in presence of pathologic CSF biomarkers in Arctic APP mutation carriers. Neurology 79(3):229–236
Shi Y, Zhang W, Yang Y, Murzin AG, Falcon B, Kotecha A et al (2021) Structure-based classification of tauopathies. Nature 598(7880):359–363
Shimada H, Ataka S, Tomiyama T, Takechi H, Mori H, Miki T (2011) Clinical course of patients with familial early onset Alzheimer’s disease potentially lacking senile plaques bearing the e693delta mutation in amyloid precursor protein. Dement Geriatr Cogn Disord 32:45–54
Steiner H, Fukumori A, Tagami S, Okochi M (2018) Making the final cut: pathogenic amyloid β-peptide generation by γ-secretase. Cell Stress 2:292–310
Strittmatter WJ, Saunders AM, Schmechel D, Pericak-Vance M, Enghild J, Salvesen GS et al (1993) Apolipoprotein E: high-avidity binding to beta-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc Natl Acad Sci USA 90(5):1977–1981
Sun C, Gonzalez B, Vago FS, Jiang W (2021) High resolution single particle Cryo-EM refinement using JSPR. Prog Biophys Mol Biol 160:37–42
Tekirian TL (2001) Abeta N-Terminal Isoforms: critical contributors in the course of AD pathophysiology. J Alzheimer’s Dis 3:241–248
Walker LC (2020) Aβ plaques. Free Neuropathol 1:1–31
Yang Y, Arseni D, Zhang W, Huang M, Lövestam S, Schweighauser M et al (2022) Cryo-EM structures of amyloid-β 42 filaments from human brains. Science 375(6577):167–172
Yang Y, Zhang W, Murzin AG, Schweighauser M, Huang M, Lövestam S et al (2023) Cryo-EM structures of amyloid-β filaments with the Arctic mutation (E22G) from human and mouse brains. Acta Neuropathol 145(3):325–333
Zhang YW, Thompson R, Zhang H, Xu H (2011) APP processing in Alzheimer’s disease. Mol Brain 4:3
Zielinski M, Peralta Reyes FS, Gremer L, Schemmert S, Frieg B, Schäfer LU et al (2023) Cryo-EM of Aβ fibrils from mouse models find tg-APPArcSwe fibrils resemble those found in patients with sporadic Alzheimer’s disease. Nat Neurosci 26(12):2073–2080
Acknowledgements
We are grateful to the family of the patients for donating brain tissue. We thank G. Qi, M. Richardson, M. Jacobsen and K. Cox for technical support. This work was supported by the US National Institutes of Health (grants P30-AG010133, RF1-NS110437, RF1-AG071177), and by the Department of Pathology and Laboratory Medicine, Indiana University School of Medicine (to B.G. and R.V.). G.I.H was supported by K99-AG078500. K.A.O. was supported by T32-GM132024. We thank the Purdue Rosen Center for Advanced Computing for providing computing resources. We thank the Purdue Cryo-EM Facility (http://cryoem.bio.purdue.edu) for the use of the Titan Krios microscope. Mass spectrometry was provided by NYU Langone’s Proteomics Laboratory.
Author information
Authors and Affiliations
Contributions
B.G., W.J. and R.V. conceived and coordinated the study; A.F. performed protein purifications and biochemical studies; G.I.H., A.F., R.V. performed western blot analysis and mass spectrometry studies; H.J.G. and R.V carried out genetic studies; B.G. carried out neuropathologic studies, selected brain areas and prepared the tissue; M.R.H, K.A.O. and D.L. carried out helical reconstruction; M.R.H., K.A.O., F.S.V. and W.J. analyzed cryo-EM data; K.A.O, F.S.V and S.R.B. built the atomic models; M.R.H., A.F., B.G. and R.V. drafted the images for publication; M.R.H., A.F., G.I.H., B.G., W.J., and R.V. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hoq, M.R., Fernandez, A., Vago, F.S. et al. Cryo-EM structures of cotton wool plaques’ amyloid β and of tau filaments in dominantly inherited Alzheimer disease. Acta Neuropathol 148, 20 (2024). https://doi.org/10.1007/s00401-024-02786-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00401-024-02786-y