Abstract
Purpose
Hearing loss (HL) is often monogenic. The clinical importance of genetic testing in HL may further increase when gene therapy products become available. Diagnoses are, however, complicated by a high genetic and allelic heterogeneity, particularly of autosomal dominant (AD) HL. This work aimed to characterize the mutational spectrum of AD HL in Austria.
Methods
In an ongoing prospective study, 27 consecutive index patients clinically diagnosed with non-syndromic AD HL, including 18 previously unpublished cases, were analyzed using whole-exome sequencing (WES) and gene panels. Novel variants were characterized using literature and bioinformatic means. Two additional Austrian medical centers provided AD HL mutational data obtained with in-house pipelines. Other Austrian cases of AD HL were gathered from literature.
Results
The solve rate (variants graded as likely pathogenic (LP) or pathogenic (P)) within our cohort amounted to 59.26% (16/27). MYO6 variants were the most common cause. One third of LP/P variants were truncating variants in haploinsufficiency genes. Ten novel variants in HL genes were identified, including six graded as LP or P. In one cohort case and one external case, the analysis uncovered previously unrecognized syndromic presentations.
Conclusion
More than half of AD HL cases analyzed at our center were solved with WES. Our data demonstrate the importance of genetic testing, especially for the diagnosis of syndromic presentations, enhance the molecular knowledge of genetic HL, and support other laboratories in the interpretation of variants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hearing loss (HL) is among the most common disabilities and affects more than five percent of the world population (430 million people) to a significant degree [1]. The etiologies are diverse, but in many cases, the cause is genetic. Variants in more than 120 genes have been linked to non-syndromic HL following Mendelian inheritance patterns [2], and the list of ‘deafness genes’ is still expanding. The clinical presentation typically correlates with the mode of inheritance. Whereas most cases of autosomal recessive (AR) HL are congenital or manifest in early childhood, dominant (AD) HL is typically associated with a later onset and often progressive. Genetic diagnoses are critical for genetic counseling, family planning, and therapy choices [3]. This is particularly important for syndromic cases that might be misdiagnosed as non-syndromic if additional symptoms have been subclinical, have not developed yet, or if the link to the auditory phenotype has not been recognized. The identification of involved genes and variants continues to improve genetic diagnoses and provides insights into molecular inner-ear function and disease. Adding to the current significance, genetic HL has attracted interest as a candidate for treatment with gene therapy [4,5,6].
Genetic diagnoses are complicated by the genetic heterogeneity of the disease. Only two genes, GJB2 and STRC, have emerged that, in some populations, account for double-digit percentages of diagnoses in congenital severe-to-profound AR HL (50% [7, 8]) and early-onset moderate AR HL (up to 30% [9]), respectively. In AD HL, no gene accounting for comparable case numbers has been identified, and many causative variants are private to isolated families. Moreover, the pathogenicity assessment of novel variants is commonly hampered by a lacking availability of functional assays or family members for segregation analysis [10]. Efforts to standardize the interpretation of sequence variants [10,11,12], the continued publication of novel disease and candidate variants, and the deposition of genetic and patient data in public databases aim to improve consistent variant evaluations across laboratories and institutions [13, 14]. The growing pool of publicly available data further improves diagnosis by enabling the identification of mutational profiles and genotype–phenotype correlations of individual genes [10, 11, 15].
In a prospective genetic study, we have analyzed 27 Austrian families with clinically suspected non-syndromic AD HL. Here, we present and recapitulate the results from these efforts, including 18 unpublished cases screened by whole-exome sequencing (WES) and targeted analysis of AD HL genes. In about three quarters of the cohort cases (20/27), candidate causative variants could be identified, including 16 variants graded as likely pathogenic (LP) or pathogenic (P). Ten variants were novel (six graded as LP/P and four variants of unknown significance (VUS)). Novel missense variants were validated and characterized using literature research and prediction tools, and possible effects on the gene products were assessed using protein structures or models. Additional mutational data were gathered from two Austrian diagnostic centers and from literature to provide an overview of the Austrian mutational AD HL spectrum. Taken together, these results enhance the current understanding of AD HL genetic variation, contribute to an improved assessment of novel variants in the future, and provide insights into the genetic profile of AD HL in Austria.
Materials and methods
Clinical testing and patient selection
Patients from 27 families with clinically suspected non-syndromic AD HL were recruited as part of the ongoing prospective genetic study. Affected study participants underwent a clinical audiologic examination and pure-tone audiometry. Medical and family histories were taken, including the age of onset, exposure to ototoxic substances or noise, and present or past additional audiologic complaints. Inclusion criteria were the presence of HL, a positive HL family anamnesis, and a presumed AD mode of inheritance as inferred from the pedigree. Ethical approval of the study protocol was received from the ethics committee of the Medical University of Vienna (approval number: ECS 198/2004; annual extensions to date). All patients and parents of minors gave informed consent for participation in the study and the publication of genetic and clinical data.
Targeted genetic analysis
Blood samples were taken and chromosomal DNA was extracted from fresh or frozen blood using a commercial isolation kit. The workflow of previously published cases is presented in detail in the respective publications [3, 15,16,17]. Panel sequencing of the previously unpublished cohort cases AD-7 and AD-27 was performed externally as part of a collaboration, as described in the supplementary file (Table S1). For the remaining index patients first presented in this study, WES on a NovaSeq 6000 device (Illumina, San Diego, CA, USA) and targeted analysis of all AD HL genes known at the time of the analysis were performed using a virtual gene panel (specified in Supplementary Table S2). Rare (≤ 0.001 in gnomAD v2.1, by reference to [10]) non-synonymous exonic and splice-site variants were validated by PCR and subsequent Sanger sequencing. In patients without exonic/splice-site variants graded as LP or P, the effect of all rare (by reference to gnomAD v3) synonymous or intronic variants on splicing within the covered regions of AD HL genes was assessed using SpliceAI [18]. In six of these families, additional family members could be recruited for parallel exome analysis or segregation analysis by PCR and Sanger sequencing. The pathogenicity of all novel variants was assessed by reference to the ClinGen-defined HL-specific recommendations for application of the ACMG/AMP criteria [10,11,12].
Protein alignments and models
Alignments of peptide sequences retrieved from UniProt or Ensembl were performed to assess the conservation levels of residues affected by novel candidate variants using Clustal Omega [19] or COBALT [16, 20]. In the present study, ACTG1, PTPRQ, WFS1, and MITF peptide sequences of species from multiple mammalian and non-mammalian classes were aligned. For the mammalian proteins NLRP3 and CEACAM16, representative peptide sequences were selected from major mammalian lineages. All species and accession numbers are specified in Supplementary Table S3. Protein structures of NLRP3 (6NPY) and ACTG1 5JLH) from the RCSB PDB database (rcsb.org/) and the human CEACAM16 (AF-Q2WEN9-F1), WFS1 (AF-O76024-F1), and MITF (AF-O75030-F1) AlphaFold models were used to illustrate the location of the respective variant using PyMOL or Jmol (jmol.org/) [15, 16, 21, 22]. Mutant residues were introduced with MISSENSE3D [23] and visualized in PyMOL (Schrodinger, LLC. 2010. Molecular Graphics System, Version 1.8).
Mutational spectrum
Retrospective genetic data from five additional patients with suspected AD HL based on the family history, clinical presentation, and/or genetic findings who had undergone standard diagnostic testing based on the suspicion of genetic HL were obtained from two medical centers in Austria, the Institute of Medical Genetics, Medical University of Vienna (Institution 1) and the Paracelsus Medical University, Salzburg (Institution 2). Clinical information and the respective analysis pipelines are contained in the Supplementary Table S4. Each variant was graded according to the ACMG/AMP guidelines. A literature search was performed to obtain previously published mutational information from Austrian AD HL patients.
Results
Clinical examination
Patients from 27 unrelated families from Austria were enrolled in the study, including 18 previously unpublished cases. The pedigrees and available audiograms are deposited in the Supplementary Fig. S1. The clinical presentation of previously published cases is described in the respective publications [3, 15,16,17].
Variant interpretation
All causative and candidate causative variants identified in AD HL genes are summarized in Table 1. Previously published variants in TECTA, COCH, MYO6 and TBC1D24 are discussed in detail in the respective publications [3, 15,16,17]. Variants in unpublished cohort cases included known AD HL variants in GSDME (c.991-15_991-13del, affecting splicing [24]; AD-4), ACTG1 (c.266C > T, p.Thr89Ile; AD-7), DIAPH1 (c.3637C > T, p.Arg1213Ter; AD-13), NLRP3 (c.778_780delinsTGG, p.Arg260Trp; AD-16) and WFS1 (c.2146G > A, p.Ala716Thr; in AD-26). The p.Arg1213Ter variant in DIAPH1 (patient AD-13) was accompanied by a novel VUS in PDE1C (Table 1) that was considered non-causative. Segregation of the GSDME (AD-4) and WFS1 (AD-26) variants was confirmed in 11 and five family members, respectively (Supplementary Fig. S1). The index patient AD-26 presented with low-frequency HL characteristic for WFS1 [10]. Novel AD HL variants included truncating alterations in the transcription factors EYA4 (c.1658delT, p.Ile553ThrfsTer12; AD-19) and POU4F3 (c.406G > T, p.Glu136Ter; in AD-20), which were classified as LP based on PVS1 (established loss-of-function mechanism) and PM2 (neither is listed in gnomAD). POU4F3 p.Glu136Ter segregated with HL in two family members (Supplementary Fig. S1). A VUS (PP3: REVEL score of 0.933, BP5) in MYH14 (c.866 T > C, p.Ile289Thr, Table 1) in the same two affected family members was considered non-causative. A novel LP missense variant in CEACAM16 (c.1045G > T, p.Ala349Ser, LP: PM2, PP1_Strong; AD-24) segregated in ten affected relatives. Missense variants of unknown clinical significance were identified in ACTG1 (c.1013C > T, p.Ser338Leu, VUS: PM2, PP3; AD-11), PTPRQ (p.3811G > C, p.Gly1271Arg, VUS: PM2; in AD-12), and NLRP3 (c.1904 T > C, p.Met635Thr, VUS: PM2; in AD-15), each as the sole candidate variant in the respective patient. In seven patients, no candidate variant (graded as VUS, LP, or P) was identified within coding nor splice regions, including two patients carrying likely benign (LB) variants in TECTA (c.3743C > T, p.Pro1248Leu, LB: BS1; in AD-8) and WFS1 (c.1371G > T, p.Arg457Ser, LB: BS1; AD-21). Following annotation with gnomAD v3, all very rare (MAF ≤ 0.001) variants (including synonymous and intronic variants) within the covered regions of AD HL genes were extracted for patients carrying either a VUS or no candidate (Table 1). As none of those variants had a predicted effect on splicing, they were no longer considered as candidates. Variants provided from external institutions included previously described pathogenic variants in GJB2 (c.224G > A, p.Arg75Gln; in Ext-2), MYH9 (c.2114G > A, p.Arg705His; in Ext-4), TMC1 (c.1714G > A, p.Asp572Asn; in Ext-5 [25]), a VUS in WFS1 (c.437G > A, p.Arg146His, VUS: PM2; in Ext-1), and a novel LP variant in MITF (c.1039C > T, p.Arg347Cys, LP: PM2, PS2_moderate, PP3, PP4; in Ext-3).
Cross-species alignment and protein structures
The conservation of VUS and novel missense variants in HL was assessed using cross-species alignments [15, 16]. ACTG1 encodes the cytoplasmic gamma-actin, which is highly conserved from human to yeast. The affected serine residue at position p.338 is fully conserved across all species (Fig. 1a) and human actins (data not shown). The protein structure illustrates its location in alpha-helix α14, where it engages in hydrogen bonds with p.Glu334 and p.Arg335. Modeling of the p.Ser338Leu mutant (AD-11) predicts a possible loss of the bond with p.Arg335, which may impact the contribution of p.Lys336 to the ADP binding site (Fig. 1a). For PTPRQ, no full-length model was available. The p.Gly1271 residue mutated in family AD-12 (p.Gly1271Arg) is highly conserved (Fig. 1b). NLRP3 (mutated in family AD-15; p.Met635Thr) belongs to the NLRP protein family that occurs in placental mammals, and the methionine at position p.635 is fully conserved across all major mammalian classes (Fig. 1c). The protein model illustrates location of the residue on the protein surface near an interaction site with the activator NEK7 (PDB-ID: 6NPY). CEACAM16 is a mammalian protein, and the peptide alignment shows full conservation of p.Ala349 across representatives from all major mammalian classes (Fig. 1d) except Xenarthra, which have no CEACAM16 orthologue. The residue p.Ala349 (AF-24; p.Ala349Ser) is located on the surface of the C-terminal IgV-like domain, and the change to serine has no predicted impact on conformation of the protein. However, due to its exposure on the surface, it might potentially be involved in interactions with other proteins. WFS1 p.Arg146 (altered in the external patient Ext-1, p.Arg146His) is evolutionary highly conserved. The residue is located in the cytosolic domain, and there is no predicted conformational impact caused by the substitution with histidine (Fig. 1e). Lastly, the mutated arginine at p.347 (external case Ext-3; p.Arg347Cys) in MITF is located in the loop of the bHLH (basic helix–loop–helix) domain involved in DNA binding. Importantly, most pathogenic variants reported for MITF HL syndromes affect the bHLH domain [26]. The p.Arg347 residue and adjacent residues are highly conserved across species (Fig. 1f).
Protein models and alignments. a The ACTG1 serine-to-leucine substitution at position p.338 (shown as spheres) is predicted to disrupt a structurally relevant hydrogen bond, potentially altering the spatial configuration of the nucleotide-binding site. The serine residue is invariant from human to yeast. b No full-length PTPRQ model is available on Alphafold, and the PHYRE2 search returned no homology-based protein model for the extracellular PTPRQ domain. The mutated p.Gly1271 residue is conserved from human to zebrafish. c The p.Met635Thr variant (spheres) is located near the NLRP3 interaction site with NEK7, indicating that the interaction might be altered. d The p.Ala349Ser variant (spheres) is located on the surface of the CEACAM16 C-terminal IgV-like domain (shown in red). The variant is predicted to have no effect on the protein structure. However, due to limited knowledge of binding partners, disruption of a protein-protein interaction cannot be ruled out. The affected residue is conserved across placental mammals. e The Wolframin (WFS1)-mutated p.Arg146 residue is highly conserved and located in the cytosolic domain of the protein (transmembrane region shown in green). There is no predicted conformational damage from the substitution with histidine. f The p.Arg347Cys variant in MITF is located in the loop of the bHLH (basic helix–loop–helix) domain of the protein (orange) that is required for DNA binding. The majority of pathogenic variants linked to MITF HL syndromes are located in this domain [26]. The affected arginine at p.347, as well as the adjacent residues, is highly conserved
Discussion
Genetic causes underlying monogenic HL have been identified in more than a hundred genes [2], making genetic diagnoses a challenging task. Standardized variant interpretation guidelines [10,11,12] and a growing pool of publicly available genetic and phenotypic data continue to facilitate and improve genetic diagnoses and counseling [13, 14]. The high clinical relevance of genetic diagnoses in HL may further increase in the future when gene-therapy options become available for treatment [27]. In a continuous effort to characterize the genetic landscape of AD HL in the Austrian population, patients from 27 families were recruited at our department and screened using targeted sequencing approaches.
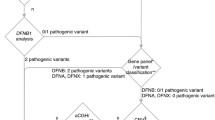
The solve rate within the total AD HL study cohort (n = 27) amounted to 59,26% (16/27), considering only variants classified as LP or P as per the ACMG guidelines (Fig. 2a). This percentage is slightly higher than the diagnostic yield of other larger cohort studies with a diagnostic yield ranging from 30 to up to 50% among AD HL patients (e.g., [9, 28, 29]) and was largely accounted for by known pathogenic variants reported in literature (10/18). Moreover, loss-of-function variants in established or suspected haploinsufficiency genes (EYA4, POU4F3, and MYO6) were found in one third of diagnoses in the total cohort and accounted for most of the novel diagnoses (Fig. 2b), allowing for a straightforward pathogenicity assessment based on rules PVS1 and PM2 [10, 12].
Solve rate and mutational spectrum. a Variants graded as pathogenic (P) or likely pathogenic (LP) were identified in 59,26% of our cohort patients (16/27). This percentage does not consider external variants or those gathered from literature. b Most LP and P variants identified in this cohort had previously been reported in other patients. Half of novel LP variants in our cohort were truncating variants in presumed haploinsufficiency genes. c Genes with more than one (candidate) causative variant, considering all Austrian patients from our cohort, external institutions and literature. As reported in [16], variants in MYO6 were the most common cause of AD HL, though all these cases were identified in our cohort
Among the novel variants first published in the present study were two truncating mutations classified as LP in the transcription factor genes EYA4 (c.1658delT, p.Ile553ThrfsTer12, in AD-19), encoding EYA Transcriptional Coactivator And Phosphatase 4, and POU4F3 (c.406G > T, p.Glu136Ter, AD-20), encoding POU domain class 4 transcription factor 3, that have both been linked to haploinsufficiency causing post-lingual, progressive AD HL. The hearing profiles with a predominant involvement of the mid (AD-19, EYA4, DFNA10) and high frequencies (AD-20, POU4F3, DFNA15), respectively, are consistent with previous reports [30, 31]. The POU4F3 variant in AD-20 was accompanied by a VUS (c.866 T > C, p.Ile289Thr) in MYH14, encoding myosin heavy chain 14, and both variants segregated with the disease in both tested family members. Since MYH14 is typically linked to early-onset or congenital AD HL, and due to the presence of a likely alternative cause, the variant was considered non-causative. A novel rare p.Ala349Ser missense variant was identified in CEACAM16, encoding Carcinoembryonic antigen-related cell adhesion molecule 16. Loss-of-function variants cause post-lingual AR HL [32], and missense variants within the N-terminal IgC domain and the C-terminal IgV domain have been linked to late-onset AD HL [33]. The p.Ala349 residue affected in our patient is located on the surface of the C-terminal IgV-like domain and may contribute to an interaction site. A p.Ala349Thr variant at the same position has been classified as LB based on its high allele frequency in some populations (MAF 0.0012 (EAS), gnomAD v2.1.1). However, segregation of the variant in the large family lead to a grading of p.Ala349Ser as LP (Fig. S1).
Interestingly, one cohort patient (AD-16) whose HL had clinically been considered non-syndromic was found to carry a known pathogenic p.Arg260Trp variant in NLRP3, linked to cryopyrin-associated autoinflammatory syndromes. A clinical reassessment confirmed additional inflammatory manifestations consistent with AD Muckle–Wells syndrome, which includes late-onset HL. Similarly, a patient (Ext-3) who had been referred for genetic testing on the suspicion of hereditary non-syndromic HL was found to carry a heterozygous suspected de novo (paternity not confirmed) p.Arg347Cys variant in MITF, a gene linked to AD Waardenburg syndrome, type 2A and AD Tietz albinism-deafness syndrome, characterized by pigment loss and congenital sensorineural HL. Indeed, the patient has mild pigmentation anomalies which corroborates the potential pathogenicity of the variant. These cases demonstrate the high importance of genetic testing for precise diagnoses and prognoses.
In conclusion, our results provide insight into the mutational spectrum of AD HL in Austria and expand the current pool of genetic and clinical data in AD HL, thereby supporting other research and diagnostic laboratories in the interpretation of variants. Further, the presented data indicate a high prevalence of presumed haploinsufficiency variants in our cohort and demonstrate the importance of genetic testing for precise diagnoses in HL.
Data availability
All data relevant to the conclusions of the study are presented or referenced within the article. Additional data will be made available upon request, provided that they are not compromised by ethical restrictions.
References
World Health Organization (2018) Deafness and hearing loss. Available at: http://www.who.int/news-room/fact-sheets/detail/deafness-and-hearing-loss. Accessed June 04 2023.
Van Camp G, Smith RJH. Hereditary hearing loss homepage. https://hereditaryhearingloss.org
Parzefall T, Frohne A, Koenighofer M, Kirchnawy A, Streubel B, Schoefer C, Gstoettner W, Frei K, Lucas T (2018) Identification of a rare COCH mutation by whole-exome sequencing: implications for personalized therapeutic rehabilitation in an Austrian family with non-syndromic autosomal dominant late-onset hearing loss. Wien Klin Wochenschr 130(9–10):299–306. https://doi.org/10.1007/s00508-017-1230-y
Akil O, Dyka F, Calvet C, Emptoz A, Lahlou G, Nouaille S, Boutet de Monvel J, Hardelin JP, Hauswirth WW, Avan P, Petit C, Safieddine S, Lustig LR (2019) Dual AAV-mediated gene therapy restores hearing in a DFNB9 mouse model. Proc Natl Acad Sci USA 116(10):4496–4501. https://doi.org/10.1073/pnas.1817537116
Landegger LD, Pan B, Askew C, Wassmer SJ, Gluck SD, Galvin A, Taylor R, Forge A, Stankovic KM, Holt JR, Vandenberghe LH (2017) A synthetic AAV vector enables safe and efficient gene transfer to the mammalian inner ear. Nat Biotechnol 35(3):280–284. https://doi.org/10.1038/nbt.3781
Askew C, Rochat C, Pan B, Asai Y, Ahmed H, Child E, Schneider BL, Aebischer P, Holt JR (2015) Tmc gene therapy restores auditory function in deaf mice. Sci Transl Med 7(295):295ra108. https://doi.org/10.1126/scitranslmed.aab1996
Kelsell DP, Dunlop J, Stevens HP, Lench NJ, Liang JN, Parry G, Mueller RF, Leigh IM (1997) Connexin 26 mutations in hereditary non-syndromic sensorineural deafness. Nature 387(6628):80–83. https://doi.org/10.1038/387080a0
Ramsebner R, Volker R, Lucas T, Hamader G, Weipoltshammer K, Baumgartner WD, Wachtler FJ, Kirschhofer K, Frei K (2007) High incidence of GJB2 mutations during screening of newborns for hearing loss in Austria. Ear Hear 28(3):298–301. https://doi.org/10.1097/AUD.0b013e318047932d
Sloan-Heggen CM, Bierer AO, Shearer AE, Kolbe DL, Nishimura CJ, Frees KL, Ephraim SS, Shibata SB, Booth KT, Campbell CA, Ranum PT, Weaver AE, Black-Ziegelbein EA, Wang D, Azaiez H, Smith RJH (2016) Comprehensive genetic testing in the clinical evaluation of 1119 patients with hearing loss. Hum Genet 135(4):441–450. https://doi.org/10.1007/s00439-016-1648-8
Oza AM, DiStefano MT, Hemphill SE, Cushman BJ, Grant AR, Siegert RK, Shen J, Chapin A, Boczek NJ, Schimmenti LA, Murry JB, Hasadsri L, Nara K, Kenna M, Booth KT, Azaiez H, Griffith A, Avraham KB, Kremer H, Rehm HL, Amr SS, Abou Tayoun AN, ClinGen Hearing Loss Clinical Domain Working Group (2018) Expert specification of the ACMG/AMP variant interpretation guidelines for genetic hearing loss. Hum Mutat 39(11):1593–1613. https://doi.org/10.1002/humu.23630
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL, ACMG Laboratory Quality Assurance Committee (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17(5):405–424. https://doi.org/10.1038/gim.2015.30
Abou Tayoun AN, Pesaran T, DiStefano MT, Oza A, Rehm HL, Biesecker LG, Harrison SM, ClinGen Sequence Variant Interpretation Working Group (ClinGen SVI) (2018) Recommendations for interpreting the loss of function PVS1 ACMG/AMP variant criterion. Hum Mutat 39(11):1517–1524. https://doi.org/10.1002/humu.23626
Azaiez H, Booth KT, Ephraim SS, Crone B, Black-Ziegelbein EA, Marini RJ, Shearer AE, Sloan-Heggen CM, Kolbe D, Casavant T, Schnieders MJ, Nishimura C, Braun T, Smith RJH (2018) Genomic landscape and mutational signatures of deafness-associated genes. Am J Hum Genet 103(4):484–497. https://doi.org/10.1016/j.ajhg.2018.08.006
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, Gu B, Hart J, Hoffman D, Jang W, Karapetyan K, Katz K, Liu C, Maddipatla Z, Malheiro A, McDaniel K, Ovetsky M, Riley G, Zhou G, Holmes JB, Kattman BL, Maglott DR (2018) ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46(D1):D1062–D1067. https://doi.org/10.1093/nar/gkx1153
Parzefall T, Frohne A, Koenighofer M, Neesen J, Laccone F, Eckl-Dorna J, Waters JJ, Schreiner M, Amr SS, Ashton E, Schoefer C, Gstœttner W, Frei K, Lucas T (2020) A novel variant in the TBC1D24 lipid-binding pocket causes autosomal dominant hearing loss: evidence for a genotype-phenotype correlation. Front Cell Neurosci 14:585669. https://doi.org/10.3389/fncel.2020.585669
Frohne A, Koenighofer M, Liu DT, Laccone F, Neesen J, Gstoettner W, Schoefer C, Lucas T, Frei K, Parzefall T (2021) High prevalence of MYO6 variants in an Austrian patient cohort with autosomal dominant hereditary hearing loss. Otol Neurotol 42(6):e648–e657. https://doi.org/10.1097/MAO.0000000000003076
Ramsebner R, Koenighofer M, Parzefall T, Lucas T, Schoefer C, Frei K (2014) Despite a lack of otoacoustic emission, word recognition is not seriously influenced in a TECTA DFNA8/12 family. Int J Pediatr Otorhinolaryngol 78(5):837–842. https://doi.org/10.1016/j.ijporl.2014.02.025
Jaganathan K, Kyriazopoulou Panagiotopoulou S, McRae JF, Darbandi SF, Knowles D, Li YI, Kosmicki JA, Arbelaez J, Cui W, Schwartz GB, Chow ED, Kanterakis E, Gao H, Kia A, Batzoglou S, Sanders SJ, Farh KK (2019) Predicting splicing from primary sequence with deep learning. Cell 176(3):535-548.e24. https://doi.org/10.1016/j.cell.2018.12.015
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539. https://doi.org/10.1038/msb.2011.75
Papadopoulos JS, Agarwala R (2007) COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 23(9):1073–1079. https://doi.org/10.1093/bioinformatics/btm076
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596(7873):583–589. https://doi.org/10.1038/s41586-021-03819-2
Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, Yuan D, Stroe O, Wood G, Laydon A, Žídek A, Green T, Tunyasuvunakool K, Petersen S, Jumper J, Clancy E, Green R, Vora A, Lutfi M, Figurnov M, Cowie A, Hobbs N, Kohli P, Kleywegt G, Birney E, Hassabis D, Velankar S (2022) AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res 50(D1):D439–D444. https://doi.org/10.1093/nar/gkab1061
Ittisoponpisan S, Islam SA, Khanna T, Alhuzimi E, David A, Sternberg MJE (2019) Can predicted protein 3D structures provide reliable insights into whether missense variants are disease associated? J Mol Biol 431(11):2197–2212. https://doi.org/10.1016/j.jmb.2019.04.009
Yu C, Meng X, Zhang S, Zhao G, Hu L, Kong X (2003) A 3-nucleotide deletion in the polypyrimidine tract of intron 7 of the DFNA5 gene causes nonsyndromic hearing impairment in a Chinese family. Genomics 82(5):575–579. https://doi.org/10.1016/s0888-7543(03)00175-7
Roesch S, Bernardinelli E, Nofziger C, Tóth M, Patsch W, Rasp G, Paulmichl M, Dossena S (2018) Functional testing of SLC26A4 variants-clinical and molecular analysis of a cohort with enlarged vestibular aqueduct from Austria. Int J Mol Sci 19(1):209. https://doi.org/10.3390/ijms19010209
Grill C, Bergsteinsdóttir K, Ogmundsdóttir MH, Pogenberg V, Schepsky A, Wilmanns M, Pingault V, Steingrímsson E (2013) MITF mutations associated with pigment deficiency syndromes and melanoma have different effects on protein function. Hum Mol Genet 22(21):4357–4367. https://doi.org/10.1093/hmg/ddt285
Omichi R, Shibata SB, Morton CC, Smith RJH (2019) Gene therapy for hearing loss. Hum Mol Genet 28(R1):R65–R79. https://doi.org/10.1093/hmg/ddz129
Sakuma N, Moteki H, Takahashi M, Nishio SY, Arai Y, Yamashita Y, Oridate N, Usami S (2016) An effective screening strategy for deafness in combination with a next-generation sequencing platform: a consecutive analysis. J Hum Genet 61(3):253–261. https://doi.org/10.1038/jhg.2015.143
Zazo Seco C, Wesdorp M, Feenstra I, Pfundt R, Hehir-Kwa JY, Lelieveld SH, Castelein S, Gilissen C, de Wijs IJ, Admiraal RJ, Pennings RJ, Kunst HP, van de Kamp JM, Tamminga S, Houweling AC, Plomp AS, Maas SM, de Koning Gans PA, Kant SG, de Geus CM, Frints SG, Vanhoutte EK, van Dooren MF, van den Boogaard MH, Scheffer H, Nelen M, Kremer H, Hoefsloot L, Schraders M, Yntema HG (2017) The diagnostic yield of whole-exome sequencing targeting a gene panel for hearing impairment in The Netherlands. Eur J Hum Genet 25(3):308–314. https://doi.org/10.1038/ejhg.2016.182
Vahava O, Morell R, Lynch ED, Weiss S, Kagan ME, Ahituv N, Morrow JE, Lee MK, Skvorak AB, Morton CC, Blumenfeld A, Frydman M, Friedman TB, King MC, Avraham KB (1998) Mutation in transcription factor POU4F3 associated with inherited progressive hearing loss in humans. Science 279(5358):1950–1954. https://doi.org/10.1126/science.279.5358.1950
Wayne S, Robertson NG, DeClau F, Chen N, Verhoeven K, Prasad S, Tranebjärg L, Morton CC, Ryan AF, Van Camp G, Smith RJ (2001) Mutations in the transcriptional activator EYA4 cause late-onset deafness at the DFNA10 locus. Hum Mol Genet 10(3):195–200. https://doi.org/10.1093/hmg/10.3.195
Booth KT, Kahrizi K, Najmabadi H, Azaiez H, Smith RJ (2018) Old gene, new phenotype: splice-altering variants in CEACAM16 cause recessive non-syndromic hearing impairment. J Med Genet 55(8):555–560. https://doi.org/10.1136/jmedgenet-2018-105349
Zheng J, Miller KK, Yang T, Hildebrand MS, Shearer AE, DeLuca AP, Scheetz TE, Drummond J, Scherer SE, Legan PK, Goodyear RJ, Richardson GP, Cheatham MA, Smith RJ, Dallos P (2011) Carcinoembryonic antigen-related cell adhesion molecule 16 interacts with alpha-tectorin and is mutated in autosomal dominant hearing loss (DFNA4). Proc Natl Acad Sci USA 108(10):4218–4223. https://doi.org/10.1073/pnas.1005842108
Löffler J, Nekahm D, Hirst-Stadlmann A, Günther B, Menzel HJ, Utermann G, Janecke AR (2001) Sensorineural hearing loss and the incidence of Cx26 mutations in Austria. Eur J Hum Genet 9(3):226–230. https://doi.org/10.1038/sj.ejhg.5200607
Janecke AR, Nekahm D, Löffler J, Hirst-Stadlmann A, Müller T, Utermann G (2001) De novo mutation of the connexin 26 gene associated with dominant non-syndromic sensorineural hearing loss. Hum Genet 108(3):269–270. https://doi.org/10.1007/s004390100484
Acknowledgements
The authors thank all the patients for their participation in this study.
Funding
Open access funding provided by Medical University of Vienna. This research was funded by the MedEl Corporation, Innsbruck, Austria (Ph.D. scholarship grant to Alexandra Frohne), by a grant from the Medical Scientific Fund of the Mayor of the City of Vienna (Project Number AP17107BGM) to Thomas Parzefall and by research grants provided by Paracelsus Medical University Salzburg to Silvia Dossena (2022-IiF-004-Dossena; FIZ RM&NT – Talent Pool/Senior Researcher 2023) and Sebastian Roesch (2022-IiF-004-Dossena).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Frohne, A., Vrabel, S., Laccone, F. et al. Mutational spectrum in patients with dominant non-syndromic hearing loss in Austria. Eur Arch Otorhinolaryngol 281, 3577–3586 (2024). https://doi.org/10.1007/s00405-024-08492-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-024-08492-5