Abstract
Gender is an important factor influencing epidemiological and clinical features of Parkinson’s disease (PD). We aimed to evaluate gender differences in the expression of a panel of miRNAs (miR-34a-5p, miR-146a, miR-155, miR-29a, miR-106a) possibly involved in the pathophysiology or progression of disease. Serum samples were obtained from 104 PD patients (58 men and 46 women) never treated with levodopa. We measured levels of miRNAs using quantitative PCR. Correlations between miRNA expression and clinical data were assessed using the Spearman’s correlation test. We used STRING to evaluate co-expression relationship among target genes. MiR-34a-5p was significantly upregulated in PD male patients compared to PD female patients (fc: 1.62; p < 0.0001). No correlation was found with age, BMI, and disease severity, assessed by UPDRS III scale, in male and female patients. MiR-146a-5p was significantly upregulated in female as compared to male patients (fc: 3.44; p < 0.0001) and a significant correlation was also observed between disease duration and mir-146a-5p. No differences were found in the expression of miR-29a, miR-106a-5p and miR-155 between genders. Predicted target genes for miR-34a-5p and miR-146-5p and protein interactions in biological processes were reported. Our study supports the hypothesis that there are gender-specific differences in serum miRNAs expression in PD patients. Follow-up of this cohort is needed to understand if these differences may affect disease progression and response to treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Parkinson’s disease (PD) is the second most common neurodegenerative disorder and affects millions of individuals worldwide. Gender differences have been recognized in PD, both in epidemiological data, with men being 1.5 more likely to develop the disease compared to women, and in the incidence of motor and non-motor symptoms and the disease progression [1,2,3,4]. Since it is now widely known that PD is not a single disease, detection of biomarkers that may characterize patient groups and predict disease progression and outcome for the purpose of personalized medicine is the goal to strive for. A personalized therapeutic approach also considering gender differences would be advisable to improve PD patients’ quality of life [5].
MicroRNAs (miRNAs) are small non-coding RNAs which regulate gene expression at post-transcriptional level. MiRNAs can cross the blood–brain barrier and enter body fluids as blood, urine and saliva. In addition, miRNAs are stable, easily quantifiable and accessible by minimally-invasive procedures [6] Several studies have shown that specific panels of miRNAs are dysregulated in PD and other parkinsonian disorders [7, 8]. Such evidence shows that the use of miRNAs as biomarkers has great potential for early PD diagnosis.
Generally, deregulated miRNAs show consistent expression between men and women, but some miRNAs may be differentially expressed between genders in specific diseases [9]. These differentially expressed miRNAs have important roles in basic biological processes and contribute to the development and progression of diseases. So far, no gender-oriented analysis of miRNA panels has been performed in early PD.
Our aim was to evaluate gender differences in the expression of some miRNAs that, according to the literature, are possibly involved in the pathophysiology and progression of PD.
Materials and methods
Patients and sample collection
PD patients never treated with levodopa were recruited from 11 Italian centers. We enrolled 105 patients affected by PD (59 men and 45 women). Clinical features were reported in Table 1. The study protocol was approved by Ethics Committees at all participant centers (University of Salerno, Salerno; University of Campania “Luigi Vanvitelli”, Napoli; IRCCS San Raffaele, Roma; Fondazione IRCCS Istituto Neurologico Carlo Besta, Milan; AOU “Policlinico-San Marco”, Catania; IRCCS Mondino Foundation, Pavia; IRCCS Ca' Granda Ospedale Maggiore Policlinico, Milan; IRCCS San Raffaele Scientific Institute, Milan; University Federico II, Naples; IRCCS Istituto delle Scienze Neurologiche, Bologna; University San Raffaele, Roma). Written informed consent was obtained from all patients. Blood samples were obtained by vein puncture using dry vacutainer tubes (BD Biosciences, Italy). Each sample was processed for serum isolation within 2 h after withdrawal according to our previously published protocol [10, 11].
MIRNAS quantification
We assessed hemolysis grade according to Shah et al., which quantified the ratio of miR-451a and miR-23a-3p to determine the samples with low (miR ratio < 5), moderate (5 < miR ratio > 7) and severe (miR > 7) grade of hemolysis [12]. Then we excluded all samples with severe hemolysis. Serum miR-34a-5p and miR-146a-5p were quantified using LNATM enhanced microRNA assay (Exiqon) according to our previous published protocol [10, 11]. Each miRNA was quantified in duplicate and mean Ct values were used for fold change calculations.
Statistical analysis
We used miR-93-5p as reference miRNA according to our previously published protocol [8]. The data was checked for normality using the Anderson Darling test and analyzed using parametric or nonparametric tests accordingly. Correlations between miRNA expression and clinical features were assessed using the Spearman’s correlation test. We excluded samples with Ct values higher than 37 from the analysis. We calculated the fold changes (fc) using the 2 − 1CT method for miR-34a-5p and miR-146a-5p. All statistical analyses were performed using GraphPad Prism (GraphPad Software Inc., San Diego, CA, USA). A p < 0.05 was considered significant.
Target prediction
Target prediction of miRNAs that were found to be differentially expressed according to gender was obtained by querying the microRNA-target interactions using the miRTarBase, chosen due to its widespread use and completeness. We considered only strong mRNA-miRNA interactions experimentally confirmed by qRT-PCR, luciferase assays and Western Blots. Then, we used the search tool for retrieval of interacting genes (STRING) to evaluate co-expression relationships among target genes. We considered only the target genes with co-expression coefficients > 0.7. Furthermore, we used open target platform (https://platform.opentargets.org/) to evaluate if predicted targets were associated to PD.
Results
Serum MIR-34a-5p
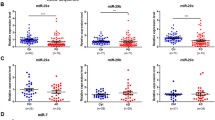
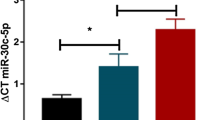
We found that miR-34a-5p was significantly upregulated in PD men patients compared with PD women patients (fc: 1.62; p < 0.0001) (Fig. 1A). We did not find any correlation with age, BMI, and disease severity, assessed by UPDRS III scale, in men and women. Using miRTarBase, we identified 85 target genes, confirming strong mRNA-miRNA interactions for miR-34a-5p. Many predicted target genes are involved in neurodegeneration. Using STRING, we found several interactions among target proteins (Fig. 2). We observed that 9 target genes are involved in aging [False Discovery Rate (FDR) = 0.00011]. In addition, 15 target genes are implicated in the regulation of neurogenesis (FDR = 0.000008) and 10 genes are involved in the regulation of neuronal death (FDR = 0.00084) (Table 2) Using open target platform, we identified 22 target genes associated to PD (Fig. 3).
Serum MIR-146a-5p
We observed that miR-146a-5p was significantly upregulated in PD men compared with PD women (fc: 3.44; p < 0.0001) (Fig. 1B). A weak but significant correlation was observed between disease duration and mir-146a-5p (r = 0.2744; p < 0.05) only in male patients. No correlation.was found between miR-146a-5p and age, BMI in PD patients of both genders.
Using miRTarBase, we identified 47 target genes confirming strong mRNA-miRNA interactions for miR-146a-5p. Many predicted target genes are involved in neurodegenerative processes. Using STRING, we found several interactions among target proteins (Fig. 4). We observed that 19 target genes are involved in neurogenesis (FDR = 0.000007) and 11 genes in the regulation of neurogenesis (FDR = 0.00013). In addition, 8 target genes are implicated in the process of axon guidance (FDR = 0.0000017) (Table 3). Using open target platform, we identified 17 target genes possibly associated to PD (Fig. 5).
No differences were found in the expression of miR-29a, miR-106a-5p and miR-155 between genders.
Discussion
In this study, we evaluated gender differences in the expression of some miRNAs that, are possibly involved in the pathophysiology and progression of PD. MiR-34a-5p is abundantly expressed in the adult mammalian brain and we selected this miRNA because of its involvement in several neurodegenerative disorders, like Alzheimer's disease (AD), schizophrenia and major depression [13]. Specific deregulation of miR-34a-5p was found in cellular and animal models of PD but also in the blood and brain of PD patients [14, 15]. Recently, Grossi et al. found that miR-34a-5p was upregulated in plasmatic pure extracellular vesicles of PD patients. Furthermore, they observed that the levels of miR-34a-5p were associated with disease duration, Hoehn and Yahr and Beck Depression Inventory scores [16]. Mir-146a has a key role in inflammatory responses and is expressed within neurons, astrocytes and microglia [17]. Several reports have shown a downregulation of miR-146a-5p in PD patient-derived samples. Caggiu et al. found that miR-146a was downregulated in PD patients under levodopa treatment compared with healthy controls (HC) [18]. In another study, a group of 20 sporadic PD patients and 45 PD patients with mutations in the LRRK2 gene also found a decrease in levels of circulating miR146a in such patients [19]. MiR-155 is considered a pro-inflammatory mediator in the CNS and has a central role in the inflammatory response to α-syn in the brain [20]. Caggiu et al. reported that miRNA-155-5p was generally up-regulated in PD patients compared to HC, but the expression of miR-155-5p was modified by levodopa treatment, since a down-regulation of miR-155-5p in PD patients with the highest dosage was observed [18]. MiR-29a is involved in various neurodegenerative diseases, including AD and PD. Bai et al. reported that serum miR-29a was reduced in 80 PD patients compared with 80 HC and serum levels were higher in female patients and HC than male patients and HC [21]. Moreover, an upregulation of this miR was found in levodopa-treated PD patients compared to drug-naïve PD patients and healthy controls [22]. Finally, we also selected miR-106a-5p, that has been recently predicted to play a role in the pathogenesis of AD [23].
In our study, we found for the first time a significant increase of serum miR-34a-5p in PD male patients compared to PD female patients. miR-34a-5p was upregulated in plasmatic pure extracellular vesicles of 15 PD patients compared with healthy controls (HC) and its levels correlated with disease duration, Hoehn and Yahr and Beck Depression Inventory scores. Measuring miR-34a-5p levels in serum and not in extracellular vesicles in a larger sample of PD patients, we did not find correlations with clinical features. Differences in sample size, disease duration, antiparkinsonian treatment, in addition to the different methodological approach, may account for this discrepancy. [16]. Recently, Stefanik et al. found that hippocampal expression of miR-34a-5p was sex-dependent [24]. MiR-34a-5p is abundantly expressed in the brain and emerging evidence support its involvement in different neurodegenerative diseases as Alzheimer's disease and PD [13]. Moreover, Findeiss et al. observed that twelve miRNAS including miR-34a-5p were upregulated in α-synuclein-overexpressing Lund human mesencephalic neurons, a well-established cell model of PD, suggesting possible novel therapeutic targets for PD [25].
Neurodegeneration in PD results from a complex interplay of multiple immunological, inflammatory and genetic factors [26]. Certain genetic defects may contribute to microglial cell activation and production of inflammatory cytokines and chemokines, which finally lead to neurodegeneration [26]. Although dysregulation of miRNAs is only one of the disease-causing mechanisms that contribute to neurodegenerative disorders, evidence indicates that dysregulated miRNAs in NDs affect the severity and progression of neurodegenerative diseases [27]. MiRNAs not only affect gene expression inside the cells but also, when sorted into exosomes, systemically mediate the communication between different types of cells. Using miRTarBase, we predicted 85 target genes of miR-34a-5p and hypothesize that an increase of this miRNA can lead to the decreased expression of such predicted target genes. We used STRING to determine protein interactions among those genes involved in key biological processes such as regulation of neurogenesis, neuronal death and aging. Using open target platform, we identified 21 predicted target genes associated to PD. Among these target genes, AKT1, L1CAM and ATG5 may be particularly relevant. AKT1 is a serine/threonine-protein kinase responsible of the regulation of glucose uptake by mediating insulin-induced translocation of the SLC2A4/GLUT4 glucose transporter to the cell surface [28]. Sekar et al. observed a significant decrease of AKT1 in substantia nigra samples obtained from PD patients compared to the age-matched controls [28]. L1CAM is a neural cell adhesion molecule involved in multiple processes, including neuronal migration, axonal growth and synaptogenesis. In the mature brain, it plays a role in the dynamics of neuronal structure and function, including synaptic plasticity [29]. Recently, Cheng et al. found that L1CAM was downregulated in PD patients compared with HC. In addition, using LASSO model they observed that L1CAM was an immune hub gene for PD of their four-gene combined model [30]. Zhang et al. found that ATG5, a protein specifically required for autophagy, was downregulated in MPTP-induced PD mice model [31]. In addition, a significant downregulation of ATG5 in peripheral blood mononuclear cells (PBMCs) of PD patients compared with HC has been consistently reported [32, 33]. Moreover, Youn et al. reported significantly lower levels of the ATG5 protein in the cerebrospinal fluid samples of PD patients compared to HC [34].
In the present study, we also observed a significant up-regulation of miR-146a-5p in PD men compared with PD women. In addition, we observed a significant correlation between disease duration and miR-146a-5p specifically in male PD patients. The positive correlation between miR-146a-5p and disease duration may suggest a possible role of miR-146a-5p in the progression of disease in male patients. Using miRTarBase, we identified 47 target genes of miR-146a-5p and with STRING we determined protein interactions among those genes involved in neurogenesis, axon guidance and regulation of neurogenesis. Using an open target platform, we identified 17 predicted target genes associated with PD. Among these target genes, TGFB1, TLR2 and TLR4 may be of particular interest. Booth et al. found that TGFB1 was downregulated in iPSC-derived midbrain-patterned astrocytes from PD patients carrying the common LRRK2 G2019S missense mutation [35]. Several studies reported an upregulation of TLR2 and TLR4 in PD patients, but no studies are available regarding gender differences [36, 37]. Normal and aggregated a-syn have shown TLR2- or TLR4-mediated microglial cells activation and neuronal loss in PD and mouse models [38, 39]. As for TLR4, regulation of hippocampal neurogenesis is region-specific in adult male mice while broader changes in neurogenesis throughout the hippocampus are found in female mice [40].
As a possible limitation of the study, we recognize that we have included only PD patients at a very early disease stage and did not include more advanced PD patients. Indeed, in this study we enrolled a cohort of levodopa-naive PD patients, thus explaining the short disease duration and low H&Y scores. The same cohort of patients is being followed up to evaluate gender differences in serum miRNAs after levodopa start.
In conclusion, our study supports the hypothesis that there are gender-specific differences in PD for miR-34a-5p and miR-146a-5p. A follow-up study of this cohort is underway to establish the possible role of these biomarkers in predicting disease progression and response to anti-parkinsonian treatments according to gender.
Conflict of interest
The Authors declare no competing financial or non-financial interests directly or indirectly related to the work submitted for publication.
Data availability
Data are available as supplementary information.
Change history
08 May 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00415-023-11750-x
References
Wooten GF, Currie LJ, Bovbjerg VE, Lee JK, Patrie J (2004) Are men at greater risk for Parkinson’s disease than women? J Neurol Neurosurg Psychiatry 75:637–639. https://doi.org/10.1136/jnnp.2003.020982
Martinez-Martin P, Falup Pecurariu C, Odin P et al (2012) Gender-related differences in the burden of non-motor symptoms in Parkinson’s disease. J Neurol 259:1639–1647. https://doi.org/10.1007/s00415-011-6392-3
Dahodwala N, Shah K, He Y et al (2018) Sex disparities in access to caregiving in Parkinson disease. Neurology 90:e48-54. https://doi.org/10.1212/WNL.0000000000004764
Lyons KE, Hubble JP, Troster AI, Pahwa R, Koller WC (1998) Gender differences in Parkinson’s disease. Clin Neuropharmacol 21:118–121.
Espay AJ, Vizcarra JA, Marsili L et al (2019) Revisiting protein aggregation as pathogenic in sporadic Parkinson and Alzheimer diseases. Neurology 92(7):329–337. https://doi.org/10.1212/WNL.0000000000006926
Zhao Y, Gagliano Taliun SA (2022) Lipid-lowering drug targets and Parkinson's disease: A sex-specific Mendelian randomization study. Front Neurol 13:940118. doi: https://doi.org/10.3389/fneur.2022.940118
Yin X, Wang M, Wang W et al (2022) Identification of potential miRNA-mRNA regulatory network contributing to Parkinson’s disease. Parkinsons Dis 2022:2877728. https://doi.org/10.1155/2022/2877728
Zago E, Dal Molin A, Dimitri GM et al (2022) Early downregulation of hsa-miR-144-3p in serum from drug-naïve Parkinson’s disease patients. Sci Rep 12:1330. https://doi.org/10.1038/s41598-022-05227-6
Vallelunga A, Iannitti T, Capece S, et al. (2021) Serum miR-96–5P and miR-339–5P Are Potential Biomarkers for Multiple System Atrophy and Parkinson's Disease. Front Aging Neurosci 13:632891. doi: https://doi.org/10.3389/fnagi.2021.632891.
Guo L, Zhang Q, Ma X, Wang J, Liang T (2017) MiRNA and mRNA expression analysis reveals potential sex-biased miRNA expression. Sci Rep 7:39812. https://doi.org/10.1038/srep39812
Vallelunga A, Ragusa M, Di Mauro S et al (2014) Identification of circulating microRNAs for the differential diagnosis of Parkinson’s disease and multiple system atrophy. Front Cell Neurosci 8:156. https://doi.org/10.3389/fncel.2014.00156
Vallelunga A, Iannitti T, Dati G et al (2019) Serum miR-30c-5p is a potential biomarker for multiple system atrophy. Mol Biol Rep 46:1661–1666. https://doi.org/10.1007/s11033-019-04614-z
Shah JS, Soon PS, Marsh DJ (2016) Comparison of Methodologies to Detect Low Levels of Hemolysis in Serum for Accurate Assessment of Serum microRNAs. PLoS One 11: e0153200. doi: https://doi.org/10.1371/journal.pone.0153200
Chua CEL, Tang BL (2019) MiR-34a in Neurophysiology and Neuropathology. J Mol Neurosci 67:235–246. https://doi.org/10.1007/s12031-018-1231-y
Briggs CE, Wang Y, Kong B et al (2015) Midbrain dopamine neurons in Parkinson’s disease exhibit a dysregulated miRNA and target-gene network. Brain Res 1618:111–121. https://doi.org/10.1016/j.brainres.2015.05.021
Botta-Orfila T, Morató X, Compta Y et al (2014) Identification of blood serum micro-RNAs associated with idiopathic and LRRK2 Parkinson’s disease. J Neurosci Res 92:1071–1077. https://doi.org/10.1002/jnr.23377
Grossi I, Radeghieri A, Paolini L et al (2021) MicroRNA34a5p expression in the plasma and in its extracellular vesicle fractions in subjects with Parkinson’s disease: An exploratory study. Int J Mol Med 47:533–546. https://doi.org/10.3892/ijmm.2020.4806
Li YY, Cui JG, Dua P et al (2011) Differential expression of miRNA-146a regulated inflammatory genes in human primary neural, astroglial and microglial cells. Neurosci Lett 499:109–113. https://doi.org/10.1016/j.neulet.2011.05.044
Caggiu E, Paulus K, Mameli G, Arru G, Sechi GP, Sechi LA (2018) Differential expression of miRNA 155 and miRNA 146a in Parkinson's disease patients. eNeurologicalSci 13:1–4. doi: https://doi.org/10.1016/j.ensci.2018.09.002.
Oliveira SR, Dionísio PA, Correia Guedes L et al (2020) Circulating Inflammatory miRNAs associated with Parkinson’s disease pathophysiology. Biomolecules 10:945. https://doi.org/10.3390/biom10060945
Zingale VD, Gugliandolo A, Mazzon E (2022) MiR-155: an important regulator of neuroinflammation. Int J Mol Sci 23:90. https://doi.org/10.3390/ijms23010090
Bai X, Tang Y, Yu M et al (2017) Downregulation of blood serum microRNA 29 family in patients with Parkinson’s disease. Sci Rep 7:5411. https://doi.org/10.1038/s41598-017-03887-3
Serafin A, Foco L, Zanigni S et al (2015) Overexpression of blood microRNAs 103a, 30b, and 29a in L-dopa-treated patients with PD. Neurology 84:645–653. https://doi.org/10.1212/WNL.0000000000001258
Abyadeh M, Tofigh N, Hosseinian S et al (2022) Key Genes and Biochemical Networks in Various Brain Regions Affected in Alzheimer’s Disease. Cells 11:987. https://doi.org/10.3390/cells11060987
Štefánik P, Michalec J, Morová M, Olexová L, Kršková L (2022) Prenatal and perinatal phthalate exposure is associated with sex-dependent changes in hippocampal miR-15b-5p and miR-34a-5p expression and changes in testicular morphology in rat offspring. Arh Hig Rada Toksikol 73:191–199. https://doi.org/10.2478/aiht-2022-73-3641
Findeiss E, Schwarz SC, Evsyukov V, et al. (2021) Comprehensive miRNome-Wide Profiling in a Neuronal Cell Model of Synucleinopathy Implies Involvement of Cell Cycle Genes. Front Cell Dev Biol 9:561086. doi: https://doi.org/10.3389/fcell.2021.561086
Magnusen A F, Hatton S L, Rani R, & Pandey M K (2021). Genetic Defects and Pro-inflammatory Cytokines in Parkinson's Disease. Front Neurol, 12:636139. https://doi.org/10.3389/fneur.2021.636139
Weng YT, Chang YM, Chern Y (2023) The impact of dysregulated microRNA biogenesis machinery and microRNA sorting on neurodegenerative diseases. Int J Mol Sci 24(4):3443. https://doi.org/10.3390/ijms24043443
Sekar S, Taghibiglou C (2018) Elevated nuclear phosphatase and tensin homolog (PTEN) and altered insulin signaling in substantia nigral region of patients with Parkinson’s disease. Neurosci Lett 666:139–143. https://doi.org/10.1016/j.neulet.2017.12.049
Appel F, Holm J, Conscience JF, Schachner M (1993) Several extracellular domains of the neural cell adhesion molecule L1 are involved in neurite outgrowth and cell body adhesion. J Neurosci 13(11):4764–4775. https://doi.org/10.1523/JNEUROSCI.13-11-04764.1993
Chen L, Wang Y, Huang J, Hu B, Huang W (2022) Identification of Immune-Related Hub Genes in Parkinson's Disease. Front Genet 13:914645. doi: https://doi.org/10.3389/fgene.2022.914645
Zhang L, Chen X, Chang M, Jiao B (2021) MiR-30c-5p/ATG5 Axis Regulates the Progression of Parkinson's Disease. Front Cell Neurosci 15:644507. doi: https://doi.org/10.3389/fncel.2021.644507.
Miki Y, Shimoyama S, Kon T et al (2018) Alteration of autophagy-related proteins in peripheral blood mononuclear cells of patients with Parkinson’s disease. Neurobiol Aging 63:33–43. https://doi.org/10.1016/j.neurobiolaging.2017.11.006
Sepúlveda D, Grunenwald F, Vidal A et al (2022) Insulin-like growth factor 2 and autophagy gene expression alteration arise as potential biomarkers in Parkinson’s disease. Sci Rep 12:2038. https://doi.org/10.1038/s41598-022-05941-1
Youn J, Lee SB, Lee HS et al (2018) Cerebrospinal fluid levels of autophagy-related proteins represent potentially novel biomarkers of early-stage Parkinson’s disease. Sci Rep 8:16866. https://doi.org/10.1038/s41598-018-35376-6
Booth HDE, Wessely F, Connor-Robson N et al (2019) RNA sequencing reveals MMP2 and TGFB1 downregulation in LRRK2 G2019S Parkinson’s iPSC-derived astrocytes. Neurobiol Dis 129:56–66. https://doi.org/10.1016/j.nbd.2019.05.006
Abdelmoaty MM, Machhi J, Yeapuri P, et al. (2022) Monocyte biomarkers define sargramostim treatment outcomes for Parkinson's disease. Clin Transl Med 12:e958. doi: https://doi.org/10.1002/ctm2.958
Chung LY, Lin YT, Liu C, et al. (2022) Neuroinflammation Upregulated Neuronal Toll-Like Receptors 2 and 4 to Drive Synucleinopathy in Neurodegeneration. Front Pharmacol 13:845930. doi: https://doi.org/10.3389/fphar.2022.845930
Su X, Maguire-Zeiss KA, Giuliano R et al (2008) Synuclein activates microglia in a model of Parkinson’s disease. Neurobiol Aging 29:1690–1701. https://doi.org/10.1016/j.neurobiolaging.2007.04.006
Alvarez-Erviti L, Couch Y, Richardson J, Cooper JM, Wood MJA (2011) Alpha-synuclein release by neurons activates the inflammatory response in a microglial cell line. Neurosci Res 69:337–342. https://doi.org/10.1016/j.neures.2010.12.02041
Connolly MG, Yost OL, Potter OV, Giedraitis ME, Kohman RA (2020) Toll-like receptor 4 differentially regulates adult hippocampal neurogenesis in an age- and sex-dependent manner. Hippocampus 30:958–969. https://doi.org/10.1002/hipo.23209
Acknowledgements
The study was funded by Agenzia Italiana del Farmaco-Ricerca Indipendente (Project: AIFA-2016-02364714). We wish to thank Professor Stefano Manfredini for the helpful discussion.
Funding
Open access funding provided by Università degli Studi di Salerno within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding author
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vallelunga, A., Iannitti, T., Somma, G. et al. Gender differences in microRNA expression in levodopa-naive PD patients. J Neurol 270, 3574–3582 (2023). https://doi.org/10.1007/s00415-023-11707-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-023-11707-0