Abstract
Neonatal early-onset sepsis (EOS) has unfortunately been the third leading cause of neonatal death worldwide. The current study is aimed at discovering reliable biomarkers for the diagnosis of neonatal EOS through transcriptomic analysis of publicly available datasets. Whole blood mRNA expression profiling of neonatal EOS patients in the GSE25504 dataset was downloaded and analyzed. The binomial LASSO model was constructed to select genes that most accurately predicted neonatal EOS. Then, ROC curves were generated to assess the performance of the predictive features in differentiating between neonatal EOS and normal infants. Finally, the miRNA-mRNA network was established to explore the potential biological mechanisms of genes within the model. Four genes (CST7, CD3G, CD247, and ANKRD22) were identified that most accurately predicted neonatal EOS and were subsequently used to construct a diagnostic model. ROC analysis revealed that this diagnostic model performed well in differentiating between neonatal EOS and normal infants in both the GSE25504 dataset and our clinical cohort. Finally, the miRNA-mRNA network consisting of the four genes and potential target miRNAs was constructed. Through bioinformatics analysis, a diagnostic four-gene model that can accurately distinguish neonatal EOS in newborns with bacterial infection was constructed, which can be used as an auxiliary test for diagnosing neonatal EOS with bacterial infection in the future.
Conclusion: In the current study, we analyzed gene expression profiles of neonatal EOS patients from public databases to develop a genetic model for predicting sepsis, which could provide insight into early molecular changes and biological mechanisms of neonatal EOS.
What is Known: • Infants with suspected EOS usually receive empiric antibiotic therapy directly after birth. • When blood cultures are negative after 48 to 72 hours, empirical antibiotic treatment is often halted. Needless to say, this is not a short time. Additionally, because of the concern for inadequate clinical sepsis production and the limited sensitivity of blood cultures, the duration of antibiotic therapy for the kid is typically extended. | |
What is New: • We established a 4-gene diagnostic model of neonatal EOS with bacterial infection by bioinformatics analysis method. The model has better diagnostic performance compared with conventional inflammatory indicators such as CRP, Hb, NEU%, and PCT. |
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Early-onset sepsis (EOS) in newborns is a multiorgan system dysfunction that can lead to severe neonatal morbidity and mortality when pathogenic microbial strains are isolated from peripheral blood or cerebrospinal fluid within 7 days of birth [1]. Acute kidney injury (AKI) is one of the most common conditions presenting with neonatal EOS [2]. In practice, many newborns with suspected EOS are given intravenous broad-spectrum antibiotics for several days, which may interfere with early breastfeeding, lead to bacterial dysbiosis, and increase the risk of morbidity for many diseases, such as type I diabetes, asthma, and necrotizing enterocolitis (NEC), although the incidence of neonatal EOS is less than 6 per 1,0000 [3,4,5,6]. Early antibiotic treatment of very low birth weight preterm infants with suspected sepsis without blood cultures is associated with an increased risk of subsequent morbidity and mortality [7, 8]. In addition, no studies to date have shown that infants with EOS could benefit from antibiotic therapy. Due to the limited predictive power of routine laboratory tests such as complete blood count, erythrocyte sedimentation rate (ESR), procalcitonin (PCT), and C-reactive protein (CRP) to effectively distinguish neonatal EOS from suspected EOS, this may prolong the use of antibiotics in uninfected infants [9,10,11]. Hence, attempts to discover specific biomarkers of neonatal EOS and to reduce unnecessary antibiotic exposure in neonates with suspected sepsis are exceptionally important for clinical neonatal care. In the current study, we analyzed gene expression profiles of neonatal EOS patients from public databases to develop a genetic model for predicting sepsis, which could provide insight into early molecular changes and biological mechanisms of neonatal EOS.
Materials and methods
Obtainment of the GSE25504 cohort and identification of differentially expressed genes (DEGs)
Whole blood mRNA expression profiling of neonatal EOS patients in the GSE25504 dataset was downloaded from gene expression omnibus (GEO) databases (http://www.ncbi.nlm.nih.gov/geo/), which included microarray data from four platforms (GPL570, GPL6947, GPL13667, and GPL15158). Then, the Limma R package with cut-off criteria of p value less than 0.05 and |logFC|> 0.5 was used to identify DEGs between the sepsis and control groups on four platforms of the GSE25504 dataset. The overlapping DEGs among the four platforms were used to identify gene set enrichment with Gene set enrichment analysis (GSEA) in the Metascape database [12]. Results were considered statistically significant when the p value was less than 0.05.
Construction of the least absolute shrink and selection operator (LASSO) model
Based on these overlapping DEGs, the binomial LASSO model was constructed using the glmnet package on the GPL6947 platform. LASSO is a more refined model obtained by constructing a penalty function that makes it compress some coefficients while setting some coefficients to zero. Therefore, it retains the advantage of subset shrinkage and is a kind of biased estimation dealing with data with complex covariance. The characteristic of LASSO regression is that when building a generalized linear model, variables can be selectively put into the model to obtain better performance parameters, thus avoiding overfitting. Here, the generalized linear model contains one-dimensional continuous dependent variables, multidimensional continuous dependent variables, nonnegative count dependent variables, binary discrete dependent variables, and multivariate discrete dependent variables. The complexity of LASSO is controlled by λ, and the larger λ penalizes linear models with more variables more strongly, so as to finally obtain a good model with fewer variables. The score was at the last setup based on the premise of directly combining the equation underneath with the mRNA expression level duplicating the LASSO regression coefficient (β) when λmin was confirmed. Score = (βmRNA1 × mRNA1) + (βmRNA2 × mRNA2) + … + (βmRNAn × mRNAn). The accuracy of the predictive features was assessed by ROC examination. Receiver operating characteristic curve (ROC) analysis was used to examine the model’s ability to discriminate between EOS and normal infants.
Clinical specimens and quantitative real-time PCR (qRT-PCR) analysis
In the period from 1 Jan 2022 to 30 Jun 2022, 99 normal and 60 EOS infants from Children’s Hospital Affiliated to Zhengzhou University were included in this single-center retrospective case–control study. EOS infants presented with significant clinical signs and symptoms of sepsis (respiratory distress, anemia, fever, decreased absolute neutrophil count, or increased C-reactive protein) were confirmed with the results of positive blood culture. Peripheral blood mononuclear cells (PBMCs) were isolated from 1 mL of whole blood collected from each infant by density separation on a Ficoll-Paque. After total RNA was extracted from PBMCs, qRT-PCR was used to detect the mRNA levels of genes in the model [13]. Finally, the relative mRNA expression levels were normalized to β-ACTIN. Primer sequences are shown in Table 1.
Identification of miRNAs targeting genes in our diagnostic models
Perform miRNA target prediction analysis to identify miRNAs that target genes in our diagnostic model by binding to their 3′ UTRs with four online public websites, including TargetScan [14], mirDIP [15], miRWalk [16], and miRmap [17] databases. Subsequently, target miRNAs predicted concordantly by the four databases were chosen to construct a miRNA-mRNA network by Cytoscape software [18].
Statistical analysis
Categorical data were compared with the Pearson chi-square test or Fisher exact test whenever appropriate, and quantitative variables were analyzed using the independent-sample t-test. The clinical characteristics of 159 infants are shown in Table 2. Results were considered statistically significant when the p value was < 0.05.
Results
Identification of overlapped DEGs
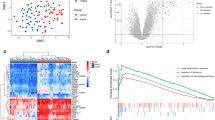
As shown in Fig. 1A, DEGs on the four platforms were screened with a cut-off criterion of p value < 0.05 and |logFC|> 0.5; 20 upregulated genes and 8 down-regulated genes were identified as common DEGs (Fig. 1B), which are shown in Fig. 1C. We also constructed a protein–protein interaction (PPI) network based on these DEGs (Fig. 1D). Twenty-eight nodes and 11 edges were acquired from the PPI network. The local clustering coefficient was 0.332, and the PPI enrichment p value was equal to 4.78e − 05. We found that CD247, CD3G, CD8B, and LCK might be the core genes. Finally, the results of the GSEA analysis showed that these overlapping DEGs are mainly associated with pathways related to infection, neutrophil degranulation, and cellular immune response (Fig. 1E).
Identification of critical genes involved in neonatal EOS
According to the binomial LASSO analysis, when λmin was equal to 0.02353066, eight genes (CST7, CD3G, CD8B, CD247, SIRPG, GPR84, MAL, and ANKRD22) were identified that most accurately predicted EOS (Fig. 2A). Score = (1.54278655 × CST7) − (0.02890854 × cd247) − (0.78324710 × CD3G) − (0.95299570 × CD8B) − (0.88133909 × SIRPG) + (0.33541335 × GPR84) + 0.11719602 × ANKRD22) − (0.55729858 × MAL). As shown in Fig. 2B, infants with EOS had significantly higher scores than these normal infants on the four platforms. What is more, the characterization of this eight-genes signature showed good diagnostic power with AUCs of 1, 1, 0.905, and 0.923 on the four platforms, respectively (Fig. 2C).
Verification of the diagnostic model in a clinical cohort
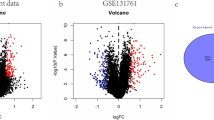
To verify the capability of the model in the diagnosing of neonatal EOS in actual clinical practice, qRT-PCR analysis was performed in a clinical cohort. There were no differences in gender, age, and NEU% levels between the EOS and normal infant groups (Table 2). The types of bacteria that the EOS infants were infected with are shown in Fig. 3. EOS infants had higher levels of PCT and CRP and lower levels of Hb when compared with those in the normal group. All genes in the model except CD8B, SIRPG, GPR84, and MAL were differentially expressed in the peripheral blood of EOS infants and normal infants (Fig. 4A). Four genes that were not significantly different were not included when calculating infant scores using the same formula. ROC analysis revealed that this diagnostic model performed well in our clinical cohort (Fig. 4B). In addition, when compared with conventional inflammatory indicators such as C-reactive protein (CRP), hemoglobin (Hb), neutrophil percentage (NEU%), and procalcitonin (PCT), the model has better diagnostic performance.
Verification of this eight-gene diagnostic model in a clinical dataset. A Difference analysis of eight genes expression between normal and EOS infants. B ROC analysis of the diagnostic model and other conventional inflammatory indicators in the clinical cohort. ns, not significant; **p < 0.01; ***p < 0.001
Construction of the miRNA-mRNA network
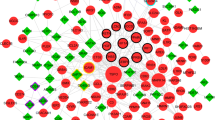
Due to the results of miRNA target prediction analysis within the four online public websites, as shown in Fig. 5A, a total of 5 miRNAs targeting the 3′ UTR of ANKRD22, 7 miRNAs targeting the 3′ UTR of CD3G, 2 miRNAs targeting the 3′ UTR of CST7, and 9 miRNAs targeting the 3′ UTR of CD247 were identified. Then, the miRNA-mRNA network consisted of genes in our diagnostic model, and potential target miRNAs were constructed (Fig. 5B).
Discussion
Despite breakthroughs in prenatal care and antibiotic prophylaxis, neonatal EOS, unfortunately, remains the third leading cause of neonatal death worldwide, due to a lack of reliability in identifying those infants who are infected [19]. Blood cultures remain the current gold standard for the diagnosis of EOS, however, its sensitivity is low in neonates, and diagnosis is delayed [20]. As a result, many newborns with suspected EOS, especially premature infants, are routinely treated with broad-spectrum intravenous antibiotics for several days, although empirical antibiotic administration may have a potential negative impact on the growth and development of newborns [9]. Hence, to minimize unnecessary antibiotic exposure in newborns with suspected EOS, there is an urgent need in clinical neonatal management for a more reliable method to diagnose EOS that utilizes neonatal blood or tissue.
In our current bioinformatics analysis study, potential diagnostic biomarkers between the sepsis and control groups were identified on the four platforms of the GSE25504 dataset. Then, we selected 28 common DEGs to construct a binomial LASSO model. Finally, a prognostic model consisting of eight genes was built and showed good diagnostic power on the four platforms. To verify the capability of the model in the diagnosing of neonatal EOS in actual clinical practice, qRT-PCR analysis was performed in a clinical cohort that consisted of peripheral blood samples from 99 normal and 60 EOS infants. All genes in the model except CD8B, SIRPG, GPR84, and MAL were differentially expressed in the peripheral blood of EOS infants and normal infants. After the score of infants was calculated with the same formula based on the four DEGs (CST7, CD3G, CD247, and ANKRD22), ROC analysis revealed that this diagnostic model performed well in our clinical cohort. In addition, when compared with conventional inflammatory indicators such as CRP, Hb, NEU%, and PCT, the model has better diagnostic performance. All the above results indicated that the diagnostic model constructed in our study could separate EOS infants from normal infants.
Most of the four genes in our diagnostic model are more or less associated with sepsis. Neutrophil-specific CST7 was significantly upregulated in the whole blood of patients with sepsis, and its encoded Cysteine F was involved in regulating the cytotoxicity of natural killer (NK) cells within the tumor microenvironment [21, 22]. CD3G is an upregulated gene involved in T cells and has been reported to be inversely correlated with sequential organ failure (SOFA) and mortality in sepsis [23]. CD247 has been reported to be involved in human and murine sepsis by many studies, and it can be involved in the occurrence and development of sepsis as a key gene of sepsis [24,25,26,27,28,29]. As for homo sapiens ankyrin repeat domain 22 (ANKRD22), which encoded a specific mitochondrial protein, it has been demonstrated to be involved in the progression of multiple tumors, including colorectal cancer [30], breast cancer [31], pancreatic cancer [32], prostate cancer [33], and nonsmall-cell lung cancer [34]; however, there are few relevant studies on sepsis and further investigation is required.
When compared with the previous study, in which miRNAs obtained from umbilical cord plasma or umbilical cord tissue could well distinguish neonatal EOS from normal infants [35], this four-gene diagnostic model has a better discriminatory ability. In addition, the umbilical cord plasma or tissue may no longer be readily available when the newborn presents with signs of sepsis, making our model more practical. There is no doubt that our study has some limitations. The individual and geographic variability of EOS infants may affect the performance of this model. In addition, the small sample size in our clinical cohort limits the validation of the model, and future multicenter randomized controlled studies are needed to evaluate this model. Finally, our study did not include blood-culture-negative infants with EOS. Considering that blood-culture-positive and negative infants may have different peripheral blood transcriptome genetic changes, we need to collect blood-culture-negative infants with EOS and detect the expression changes of the four genes in their peripheral blood to determine whether the four-gene signature we constructed could identify septic infants with negative blood cultures in the future.
Conclusions
In summary, we constructed a four-gene diagnostic model that can accurately differentiate neonatal EOS with bacterial infection by bioinformatics analysis, which can be used as an ancillary test for the diagnosis of neonatal EOS with bacterial infection in the future.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- EOS:
-
Early-onset sepsis
- AKI:
-
Acute kidney injury
- NEC:
-
Necrotizing enterocolitis
- ESR:
-
Erythrocyte sedimentation rate
- PCT:
-
Procalcitonin
- CRP:
-
C-reactive protein
- DEGs:
-
Differentially expressed genes
- GEO:
-
Gene expression omnibus
- GSEA:
-
Gene set enrichment analysis
- LASSO:
-
Least absolute shrink and selection operator
- ROC:
-
Receiver operating characteristic curve
- qRT-PCR:
-
Quantitative real-time PCR
- PBMCs:
-
Peripheral blood mononuclear cells
- PPI:
-
Protein-protein interaction
- NEU:
-
Neutrophil
- Hb:
-
Hemoglobin
- NK:
-
Natural killer
- SOFA:
-
Sequential organ failure
- ANKRD22:
-
Homo sapiens ankyrin repeat domain 22
References
Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, Bellomo R, Bernard GR, Chiche JD, Coopersmith CM et al (2016) The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 315(8):801–810
Bagshaw SM, George C, Bellomo R (2008) Early acute kidney injury and sepsis: a multicentre evaluation. Crit Care 12(2):R47
Weitkamp JH (2020) The role of biomarkers in suspected neonatal sepsis. Clin Infect Dis
Schrag SJ, Farley MM, Petit S, Reingold A, Weston EJ, Pondo T, Hudson Jain J, Lynfield R (2016) Epidemiology of invasive early-onset neonatal sepsis, 2005 to 2014. Pediatrics 138(6)
Kuppala VS, Meinzen-Derr J, Morrow AL, Schibler KR (2011) Prolonged initial empirical antibiotic treatment is associated with adverse outcomes in premature infants. J Pediatr 159(5):720–725
Cho I, Yamanishi S, Cox L, Methé BA, Zavadil J, Li K, Gao Z, Mahana D, Raju K, Teitler I et al (2012) Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 488(7413):621–626
Ting JY, Synnes A, Roberts A, Deshpandey A, Dow K, Yoon EW, Lee KS, Dobson S, Lee SK, Shah PS (2016) Association between antibiotic use and neonatal mortality and morbidities in very low-birth-weight infants without culture-proven sepsis or necrotizing enterocolitis. JAMA Pediatr 170(12):1181–1187
Flannery DD, Ross RK, Mukhopadhyay S, Tribble AC, Puopolo KM, Gerber JS (2018) Temporal trends and center variation in early antibiotic use among premature infants. JAMA Netw Open 1(1):e180164
Kuzniewicz MW, Puopolo KM (2020) Antibiotic stewardship for early-onset sepsis. Semin Perinatol 44(8):151325
Lin GC, Küng E, Smajlhodzic M, Domazet S, Friedl HP, Angerer J, Wisgrill L, Berger A, Bingle L, Peham JR et al (2021) Directed transport of CRP across in vitro models of the blood-saliva barrier strengthens the feasibility of salivary CRP as biomarker for neonatal sepsis. Pharmaceutics 13(2)
Honore PM, Redant S, Kaefer K, Barreto Gutierrez L, Kugener L, Attou R, Gallerani A, De Bels D (2021) Procalcitonin is useful for antibiotic deescalation in sepsis and septic shock: beware of some confounders! Crit Care Med 49(6):e659
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, Benner C, Chanda SK (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10(1):1523
Zhang G (2020) Expression and prognostic significance of BANF1 in triple-negative breast cancer. Cancer Manag Res 12:145–150
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20
Tokar T, Pastrello C, Rossos AEM, Abovsky M, Hauschild AC, Tsay M, Lu R, Jurisica I (2018) mirDIP 4.1-integrative database of human microRNA target predictions. Nucleic Acids Res 46(D1):D360-d370
Sticht C, De La Torre C, Parveen A, Gretz N (2018) miRWalk: an online resource for prediction of microRNA binding sites. PLoS ONE 13(10):e0206239
Vejnar CE, Blum M, Zdobnov EM (2013) miRmap web: comprehensive microRNA target prediction online. Nucleic Acids Res 41:W165–168
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Liu L, Oza S, Hogan D, Perin J, Rudan I, Lawn JE, Cousens S, Mathers C, Black RE (2015) Global, regional, and national causes of child mortality in 2000–13, with projections to inform post-2015 priorities: an updated systematic analysis. Lancet 385(9966):430–440
Mithal LB, Palac HL, Yogev R, Ernst LM, Mestan KK (2017) Cord blood acute phase reactants predict early onset neonatal sepsis in preterm infants. PLoS ONE 12(1):e0168677
Sawyer AJ, Garand M, Chaussabel D, Feng CG (2021) Transcriptomic profiling identifies neutrophil-specific upregulation of cystatin F as a marker of acute inflammation in humans. Front Immunol 12:634119
Perišić Nanut M, Sabotič J, Švajger U, Jewett A, Kos J (2017) Cystatin F affects natural killer cell cytotoxicity. Front Immunol 8:1459
Almansa R, Heredia-Rodríguez M, Gomez-Sanchez E, Andaluz-Ojeda D, Iglesias V, Rico L, Ortega A, Gomez-Pesquera E, Liu P, Aragón M et al (2015) Transcriptomic correlates of organ failure extent in sepsis. J Infect 70(5):445–456
Zeng X, Feng J, Yang Y, Zhao R, Yu Q, Qin H, Wei L, Ji P, Li H, Wu Z et al (2021) Screening of key genes of sepsis and septic shock using bioinformatics analysis. J Inflamm Res 14:829–841
Gong FC, Ji R, Wang YM, Yang ZT, Chen Y, Mao EQ, Chen EZ (2020) Identification of potential biomarkers and immune features of sepsis using bioinformatics analysis. Mediators Inflamm 2020:3432587
Liang J, Wu W, Wang X, Wu W, Chen S, Jiang M (2022) Analysis of sepsis markers and pathogenesis based on gene differential expression and protein interaction network. J Healthcare Eng 2022:6878495
Kim KS, Jekarl DW, Yoo J, Lee S, Kim M, Kim Y (2021) Immune gene expression networks in sepsis: a network biology approach. PLoS ONE 16(3):e0247669
Chen M, Chen X, Hu Y, Cai X (2020) Screening of key genes related to the prognosis of mouse sepsis. Biosci rep 40(10)
Chen J, Lin M, Zhang S (2019) Identification of key miRNA-mRNA pairs in septic mice by bioinformatics analysis. Mol Med Rep 20(4):3858–3866
Pan T, Liu J, Xu S, Yu Q, Wang H, Sun H, Wu J, Zhu Y, Zhou J, Zhu Y (2020) ANKRD22, a novel tumor microenvironment-induced mitochondrial protein promotes metabolic reprogramming of colorectal cancer cells. Theranostics 10(2):516–536
Wu Y, Liu H, Gong Y, Zhang B, Chen W (2021) ANKRD22 enhances breast cancer cell malignancy by activating the Wnt/β-catenin pathway via modulating NuSAP1 expression. Bosn J Basic Med Sci 21(3):294–304
Luo L, Li Y, Huang C, Lin Y, Su Y, Cen H, Chen Y, Peng S, Ren T, Xie R et al (2021) A new 7-gene survival score assay for pancreatic cancer patient prognosis prediction. Am J Cancer Res 11(2):495–512
Qiu Y, Yang S, Pan T, Yu L, Liu J, Zhu Y, Wang H (2019) ANKRD22 is involved in the progression of prostate cancer. Oncol Lett 18(4):4106–4113
Yin J, Fu W, Dai L, Jiang Z, Liao H, Chen W, Pan L, Zhao J (2017) ANKRD22 promotes progression of non-small cell lung cancer through transcriptional up-regulation of E2F1. Sci Rep 7(1):4430
Ernst LM, Mithal LB, Mestan K, Wang V, Mangold KA, Freedman A, Das S (2021) Umbilical cord miRNAs to predict neonatal early onset sepsis. PLoS ONE 16(5):e0249548
Acknowledgements
Not applicable.
Funding
This research was supported by the Medical Science and Technology Project of Henan Province (LHGJ20210665).
Author information
Authors and Affiliations
Contributions
ZGH designed the study. ZN and JYJ downloaded and analyzed the data. ZZH completed the qRT-PCR experiment. BY and DG wrote the manuscript. All authors critically reviewed the manuscript. The final manuscript has been approved by all authors.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was supported by the Ethics Committees of Zhengzhou Central Hospital Affiliated to Zhengzhou University. Each child had an informed consent form signed by his or her parents.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Daniele De Luca
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bai, Y., Zhao, N., Zhang, Z. et al. Identification and validation of a novel four-gene diagnostic model for neonatal early-onset sepsis with bacterial infection. Eur J Pediatr 182, 977–985 (2023). https://doi.org/10.1007/s00431-022-04753-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00431-022-04753-9