Abstract
A cross-sectional study was conducted to identify factors for infections of pigs with key respiratory pathogens: porcine circovirus type 2 (PCV2), porcine reproductive and respiratory syndrome virus (PPRSv), Mycoplasma hyopneumoniae (M. hyo), Actinobacillus pleuropneumoniae (App), and gastrointestinal (GI) parasites in Uganda. A structured questionnaire was used to collect data on management practices associated with infections. Ninety (90) farms and 259 pigs were sampled. Sera were screened against 4 pathogens using commercial ELISA tests. The Baerman’s method was used to identify parasite species in faecal samples. Logistic regression was done to identify risk factors for infections. Results showed individual animal seroprevalence of PCV2 was 6.9% (95% CI 3.7–11.1), PRRSv 13.8% (95% CI 8.8–19.6), M. hyo 6.4% (95% CI 3.5–10.5), and App 30.4% (95% CI 24.8–36.5). The prevalence of Ascaris spp. was 12.7% (95% CI 8.6–16.8), Strongyles spp was 16.2% (95% CI 11.7–20.7), and Eimeria spp. was 56.4% (95% CI 50.3–62.4). Pigs infested with Ascaris spp. were more likely to test positive to PCV2, odds ratio (OR) 1.86 (CI 1.31–2.60; p = 0.0002). For M. hyo, infection with Strongyles spp. was a risk factor (OR 12.9, p < 0.001). Pigs that had Strongyles and Ascaris spp. Infections (ORs 3.5 and 3.4, p < 0.001 respectively) were likely to have co-infections. The model showed that use of cement, elevated floor, and limiting contacts with outside pigs were protective while using mud and helminth infestations increased risks of co-infections. This study provided evidence that improved housing and biosecurity are critical in reducing pathogen incidence in herds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In Uganda, pig production has grown rapidly in recent years from approx. 0.7 million pigs in 1990 to 4.4 million in 2019 (UBOS 2020). This reflects a rise in the demand for pork (Ouma et al. 2014), which offers significant opportunities to pig producers for livelihood improvement. In Uganda’s current production systems, the lack of implementation of biosecurity measures constitute key factors for the spread of swine diseases such as African swine fever (Muhanguzi et al. 2012; Muhangi et al. 2014; Dione et al. 2016). Recent studies reveal that among diseases, respiratory and gastrointestinal (GIT) helminth infections are common in Ugandan pigs, contributing to the disease burden and thus affecting productivity in the sector (Ikwap et al. 2014; Roesel et al. 2017). In Lira district, Uganda, a recent multi-pathogen study revealed occurrence of Mycoplasma hyopneumoniae (M. hyo), Actinobacillus pleuropneumoniae (App), Leptospira spp., porcine reproductive and respiratory syndrome virus (PPRSv), and porcine circovirus (PCV2) type 2 (Dione et al. 2018). Other studies confirmed presence of PCV2 in Ugandan pigs (Jonsson 2013; Ojok et al. 2013; Eneku et al. 2018). Three main production systems are identified: farrow to finish, farrow to wean, and wean to finish. In some farms, pigs are often not segregated by age groups and are fed together. Coupled with low biosecurity, this exposes younger pigs to infectious diseases. In Uganda, no pig vaccination was done against any pig disease during this study.
To date, no information exists on husbandry and management factors associated with important respiratory pathogen infections in Uganda’s smallholder pig systems. This hampers the design of effective interventions at farm level. Evidence from previous studies shows Metastrongylu spp. and Ascaris spp. compromise lung function due to the damage induced by their migratory larvae, thereby excerbating the effect of other viral and bacterial agents (Adedeji et al. 1989; Brewer and Greve 2011). This interaction increases disease duration and/or severity, with associated negative effects on productivity (Thacker et al. 1999). This study was designed to (i) identify risk factors for co-infections with respiratory pathogens, (ii) examine associations between gastrointestinal parasite infestations and key respiratory pathogens (PCV2, PRRSv, M. hyo, and App), and (iii) investigate associations between pathogens occurrence, farm management, and biosecurity practices, with a view to inform control and preventive measures at herd level.
Materials and methods
Study area
This study was done in Lira district, mid-northern Uganda, where the International Livestock Research Institute (ILRI) previously implemented a smallholder pig value chain development project (SPVCD) in 2011. In this project, a value chain assessment was conducted to select study sites using pig density, poverty levels, and market access (Ouma 2017). Our study used market access to select subcounties based on value chain domains into rural production for urban consumption (R-U) and urban production for urban (U-U) consumption (Ouma 2017). The total pig population in Lira district was estimated to be 30,000 in 2020 (district veterinary officer, personal communication). Pigs are produced under housed, tethered, and free-range systems (Kungu et al. 2019).Under these systems, pigs are housed in permanent or temporal structures made of cement, wood, or papyrus. Tethering is when pigs are tied on a rope (on a pole) to graze around the homestead, while free-range is when pigs are allowed to freely roam in the neighborhood in search of own feeds and water. Routine preventive measures such as anthelmintics are generally not practiced until pigs show visible signs of illness. In these smallholder pig production systems, biosecurity and hygiene practices are generally poor, which partly explains a high disease incidence in herds.
Study design and sampling of subcounties, parishes, and villages
A cross-sectional serologic study was conducted from October to December 2018. We used multistage sampling to select subcounties and villages. In the first stage, four subcounties were selected (from a total of 9): two (central division and railways) representing U-U consumption and two (Adekokwok and Ngetta) representing R-U consumption. In stage two, two (2) villages with the highest pig density were selected for the study.
Sample size determination
To determine the sample size, a formula for simple random sampling was used (Dohoo et al. 2003). A previous study in Lira district found a seroprevalence of M. hyo in pigs of 20.9% (Dione et al. 2018). Adjusting for test sensitivity and specificity, true prevalence was computed to be 24%. The required sample size of pigs was obtained from Eq. (1):
where n = is the required sample size, Zα is the standard z-score from a normal distribution (1.96), p = estimated prevalence of disease (24%), q = 1 − p (76%), and d = allowable error (6%). Using this formula, an unadjusted sample size of 195 pigs was computed. To adjust for within-farm clustering, we sampled 3 pigs per herd, thus the design effect (Deff) was obtained from Eq. 2 below:
where icc is the intra-cluster correlation (0.2) for respiratory disease (Dohoo et al. 2003) and n1 is the number of pigs sampled per herd (3), thus the Deff calculated is 1.4. The adjusted sample size was calculated from the equation: N = n1 × number of pigs sampled per herd (3). From this, adjusted sample size of 273 pigs was derived.
Sampling of farms and pigs
In each selected village, a list of pig keeping households/farms was obtained from the district veterinary office and the area local councils. Random sampling of farms was done until the required sample size was obtained. We sampled farms regardless of health status or anthelmintic treatments to examine differences in farm husbandry practices and how these influence occurrences of specific pathogens in farms. Using a sampling frame of all pig farmers generated with field research assistants, three (3) pigs per herd were sampled until the required sample size was reached. Only pigs ≥ 2.5 months old were selected for sampling, since pigs below that age are reported to retain maternal antibodies to PCV2 and PRRSv post weaning, which could interfere with serologic tests (Opriessnig et al. 2004; Gillespie et al. 2009). App-acquired colostral antibodies were reported to decay within 2 months postpartum (Vigre et al. 2003). We sampled pigs from 2.5 months and above regardless of health status, clinical signs, or feed types given.
Data collection methods
A structured questionnaire with closed questions was designed, pre-tested by the first author in Mukono district, and revised before use. Research assistants were trained in its use before it was administered to each household head or farm manager. To ensure consistency, all questions were translated to a local language spoken in the area (Langi). The questionnaire captured data on potential risk factors for infection with respiratory pathogens.
Blood sample collection and storage
Each pig was properly restrained as described in the ILRI Standard Operating Procedures (SOPs) manual, Sect. 2, parts (c) and (d) (ILRI 2004). Smaller pigs were restrained by hand while larger ones were restrained with a metallic pig catcher (model BZ002; MG. Livestock, Shandong, China) placed behind the upper incisor teeth and the snout raised upward. Blood was then collected from the cranial vena cava or jugular vein using a 21G, 1.5″ needle into plain 5-mL BD® vacutainer tubes. The tubes were labeled with animal identification details and then placed in an ice box at 4–6 °C. After collection, samples were delivered (within 3 h) to the district veterinary laboratory for temporary storage. Blood samples were left to stand at room temperature (20 °C) overnight and serum harvested the following day into 2 mL cryotubes (Sarstedt®, Germany), labeled, and stored in a fridge at − 20 °C until testing.
Serological analysis
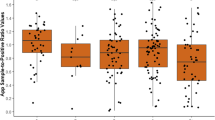
In the lab, sera were screened using ELISA assays according to manufacturers’ instructions for each pathogen: M. hyo and App-ApxIV (IDDEXX, Westbrook, Maine, USA) for PRRSv and PCV2 assays (Krishgen Biosystems, India). Cut-off sample to positive ratios (S/P%) for M. hyo were > 0.40 (positive) and < 0.30 (negative), App was ≥ 50% (positive) and < 40% (negative). PCV2 and PRRSv S/P cut-off ratios for positive and negative samples were ≥ 0.2 and < 0.2 respectively. Suspect samples were re-tested. Test sensitivity (Se) and specificity (Sp) for M. hyo ELISA were 85.6 and 99.6%, respectively; Se and Sp for App-ApxIV Ab ELISA test were 97.8 and 100%, respectively. Se and Sp for PRRSv were 94.0 and 94.0%, while for PCV2 Se and Sp these were 92.0 and 94.0%, respectively. Test Se and Sp were used to calculate true prevalence of respiratory infections at α = 0.05 significance level.
Fecal sample collection and analysis
Fecal samples (~ 3 g) were collected from the rectum of each pig using gloved hands into 5-mL plastic containers, labeled, and placed in ice box at 4 °C. Samples were taken to the district veterinary lab for temporary storage at 4 °C. Samples were transferred to the Central Diagnostic Laboratory, College of Veterinary Medicine, Animal Resources and Biosecurity (CoVAB), Makerere University for analysis 1 week after collection. Helminth species were identified and fecal egg counts per gram (EPGs) were quantified using the Baermann and McMaster methods, respectively (MAFF 1986).
Data analysis and presentation
Data was coded and entered into Excel 16.0, and any errors in entry were corrected by cross-checking with questionnaires. RStudio was used for data analysis and presentation (R Core Team 2019). True prevalence was computed by adjusting for apparent prevalence using prevalence 0.2.0 package in R, taking into account test sensitivities and specificities (Devleesschauwer et al. 2022). Multivariable logistic regression analysis of risk factors for each pathogen was performed. The response variable was the ELISA test result with predictors pig age and husbandry practices (house type, parasites, drug use, pig mixing, hygiene score, and drainage). The model below was fitted to predict respiratory infections as a function of pig characteristics and husbandry practices (house type, parasites, drug use, pig age, pig mixing, hygiene, and drainage):
where \(\mathrm{ln}\frac{\widehat{p}}{(1-\widehat{p)}}\) is the expected log of odds of infection, β0 is the model intercept, β1, β2 are regression coefficients, and x1i, x2i, are the respective explanatory variables. Interaction terms were tested for each model, and confounding was checked by inclusion and exclusion of variables to observe a change in model coefficients. A cumulative link mixed-effects (CLMM) model was fitted to estimate the odds of co-infection (2 or more pathogens) with farm as a random effect. The CLMM model was fitted using R packages “factoextra” and “ordinal” using a backward elimination method such that all variables whose p-value > 0.2 were dropped. Residual plots and R-square statistics were used to assess the fitted models. Table 1 below presents housing and husbandry practice variables used in fitting the models.
Results
In all four subcounties, a total of 259 pigs were sampled from 90 farms. Data of 12 pigs was incomplete (missing values) and as such was excluded from the analysis. The median age category of the respondents was 36–50 years with an age range of 24–70 years. Of the 90 respondents, 41 (45.5%) were males and 49 (54.5%) were females. The median herd size per housed herds was 11 pigs, and for tethered herds it was 4 pigs. Male pigs constituted 53.7% (n = 139) of pigs in the sample while females were 46.3% (n = 120). The median age of sampled pigs was 5 months, and the age range was from 2.5 to 15 months. Table 2 below shows a summary of demographic characteristics.
Prevalence of respiratory pathogens and GIT parasites
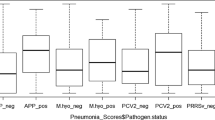
Table 3 below shows true herd and individual level prevalence of selected respiratory pathogens at respective 95% confidence intervals. Results showed that App was the most prevalent pathogen while PCV2 was the least prevalent at both pig and herd levels. Of the GIT parasites, Eimeria spp was the most prevalent while Trichuris spp was the least prevalent parasite found.
Prevalence and proportion of co-infections
Co-infections with two or more pathogens were observed in this study. Among respiratory pathogens, the highest percentage of co-infections was between pathogens and GI parasites. Among pathogens, the highest co-infections occurred between PRRSv and PCV2, followed by M. hyo and PCV2. Only 5 pigs were co-infected with 3 pathogens, and 42.5% (n = 110) of pigs sampled had at least 2 co-infections. Table 4 below shows a summary of percentage of pigs co-infected with pathogens and GI parasites.
Multivariable logistic regression model of risk factors for individual respiratory infections
Results showed that parasite infections, pig confinement (only for App), drug use, and pig age were significant predictors of single respiratory infections (Table 5). Drugs used by farmers were antibiotics, anthelmintics, and multivitamins.
A cumulative link mixed model for mixed respiratory infections with farm as random effect
Table 6 below shows a cumulative link mixed model (CLMM) from R packages (“ordinal,factoextra”) of factors for respiratory co-infections with farm as a random effect. This model was fitted to predict co-infections (regardless of pathogens involved), as these have synergistic effects on the induction of respiratory disease. The model showed that farmer occupation, floor type (cement, elevated floor), and no contacts with outside pigs were protective against co-infections, while pig age, wall type (mud, timber), herd size, and helminth infestations increased risks of respiratory co-infections. A likelihood ratio test (LR stat) statistic (4.02, p < 0.04*) showed that the model below was slightly significant.
Discussion
These results highlight widespread occurrence of selected respiratory pathogens in pigs in the study area. At both individual and herd levels, App was found to be of highest prevalence, followed by PRRSv, PCV2, and, lastly, M. hyo (Table 3). These findings are comparable with those from a recent study (Dione et al. 2018). However, compared to the findings of other studies (Jonsson 2013; Ojok et al. 2013; Eneku et al. 2018), our study found a lower PCV2 seroprevalence. This may be due to differences in the sampling procedures, diagnostic methods used, and the type of production system from which pigs were sampled. In our study, sampling was done in households that confined their pigs in pig sheds or tethered around homesteads, while Jonsson (2013) sampled pigs from a wildlife-livestock interface (near Murchison Falls national park), which probably exposed them to a higher risk of infection from other roaming or wild pigs. Eneku et al. (2018) sampled pigs that presented with clinical signs of PCV2 and therefore had a higher probability of PCV2 detection while another research team (Ojok et al. 2013) sampled pig tissues from a local abattoir.
The PCV2 seroprevalence at individual pig and herd levels found in this study was lower than (54 and 78% respectively) that reported in Mozambique (Laisse et al. 2018). Differences in the sampled population could account for variations in PCV2 prevalence, as the Mozambiqan study was done in slaughter places and was likely on older animals compared to the pigs sampled in this study. The results from this study also show that PCV2 seroprevalence was higher (6.9% vs 1.4%) than that found in Nigeria (Aiki-Raji et al. 2018) and lower than (15.9%) that found in South Africa (Afolabi et al. 2017).
This study revealed a higher PRRSv seroprevalence compared to the previous findings (Dione et al. 2018). This could suggest either increased herd to herd transmission over the past few years, since PRRSv transmission can occur via several routes (Otake et al. 2010), or because the virus can remain in affected herds as a persistent infection (Murtaugh and Genzow 2011). The finding that the odds of testing seropositive to PPRSv rises with increase in pig age is in consonance with findings from previous studies that showed that neutralizing and anti-PRRSv IgG antibodies can remain persistent for several months (Nelson et al. 1994; Murtaugh et al. 2002). The increased odds of seropositivity to PRRSv due to lack of regular preventive treatments against bacterial and helminth infections were demonstrated in this study. The role of PRRSv in inducing severe disease during co-infections with other pathogens has been previously reported (Halbur et al. 1996; Thacker et al. 1999). This suggests that regular prophylactic treatments are important in reducing the risk of opportunistic co-infections. A similar observation was made for App, in which infection was dose-dependent, accounting for increased incidence in older pigs (Marsteller and Fenwick 1999). The effect of herd size was highlighted in this study. The observation that larger herds (> 20 pigs) increased the odds of PRRSv infection may be related with increased stocking density, as PRRSv is known to be highly infectious. The increase in the odds of PRRSv detection may also be due to its tendency to remain as a persistent infection after entry into a herd (Pileri and Mateu 2016).
Co-infections in this study were lower than in other studies (Gillespie et al. 2009). Co-infections between PRRSv and App were the most prevalent, followed by PCV2 and App. The effect of PCV2 co-infection with other pathogens in increasing the severity and incidence of PCV2-associated disease has been reported in previous studies (Opriessnig et al. 2004; Fablet et al. 2012; Segalés et al. 2013). Other studies reveal that a diversity of pathogens is involved in respiratory disease (Qin et al. 2018). The cumulative link model showed that better floor types (cement or elevated) had protective effects against co-infections (Table 6).While hygiene score was not significantly associated with co-infections, the frequency of cleaning of pens was significantly associated with co-infections: the poorer the level of hygiene and the lower the frequency of cleaning of pens, the higher the risk of co-infections. These findings concur with previous studies which showed pigs raised in clean environments grew significantly faster than those raised on dirty floors (Cargill 2019).
This study revealed associations of particular respiratory pathogens and GIT parasite infections in pigs (Tables 4 and 5). These results are comparable with findings from a study in southwest Uganda which reported a high prevalence of GIT helminth infections (Roesel et al. 2017). The increased odds of Ascaris spp infection in tethered pigs illustrates the importance of biosecurity (e.g., confining pigs) in reducing the risk of infection. Ascaris spp. has been shown to compromise lung function through immunomodulatory mechanisms, thereby excerbating the effect of other viral and bacterial agents, as well as increase disease severity (Adedeji et al. 1989; Brewer and Greve 2011). While pigs infected with Eimeria spp. may show no observable clinical signs, Eimeria spp. has been reported to cause diarrhea in piglets as it damages intestinal mucosa, increasing susceptibility to other pathogens.
In general, results showed that biosecurity variables had a significant influence on pathogen occurrence in farms. These findings agree with previous studies which revealed that good housing, hygiene and reduced stress play a significant role in minimizing the effects of diseases such as PCVAD (Gillespie et al. 2009; Cargill 2019). Poor drainage and hygiene in some farms may have raised the risk of re-infections with parasite eggs from contaminated feeds and water. In addition, the higher parasite infestations observed in some farms (tethering) may also have been due to lack of implementation of routine preventive measures such as deworming. These environmental stressors (poor hygiene, rearing of different age groups in the same pens, overcrowding, poor nutrition, etc.) are known to suppress immunological responses and therefore impede a pig’s ability to fight off infection (Cargill 2019). The role of good ventilation and proper cleaning practices in improving indoor air quality by reducing microbial contamination with respiratory pathogens has been documented (Banhazi et al. 2008; Cargill 2019). Overall, the results of the logistic regression models demonstrated the importance of improved hygiene and biosecurity in reducing mixed respiratory infections.
Conclusions
This study highlights widespread occurrence of economically important respiratory pathogens in pigs in the study area. This may likely reflect the situation in swine herds in eastern and northern Uganda, where production systems are largely similar. The negative correlations between biosecurity variables and mixed pathogens signify the role of improved biosecurity in reducing the risks of co-infections. In addition, the associations between biosecurity and housing and pathogens provides further support to the above evidence. Further studies to identify PCV2 and PRRSv genotypes that circulate in pigs in this region, as well as quantify their economic impacts on swine productivity, are warranted to guide the design of effective interventions.
Limitations of the study
Statistical analyses were limited to only estimates of prevalence and odds of infections. Potential bias may have been introduced by inaccurate responses to some questions in the questionnaire and misclassification of pig errors due to imperfect assay sensitivities and specificities. Also, the time lapse between fecal sample collection and lab analysis may have affected the sensitivity of the Baerman’s test, as ability of larvae to hatch was reduced. The detection of antibodies may not reflect actual or current infection. This may have led to over- or underestimation of the observed relationships.
Data availability
The original contributions presented in the study are publicly available. The dataset analyzed for this study can be found here: https://data.ilri.org/portal/dataset/multipathogen-survey-and-risk-factors.
References
Adedeji SO, Ogunba EO, Dipeolu OO (1989) Synergistic effect of migrating Ascaris larvae and Escherichia coli in piglets. J Helminthol 63:19–24. https://doi.org/10.1017/S0022149X00008671
Afolabi K, Iweriebor BC, Obi LC, Okoh AI (2017) Molecular detection of porcine circovirus type 2 in swine herds of Eastern Cape province South Africa. BMC Microbiol 17:1–10. https://doi.org/10.1186/s12866-017-1121-4
Aiki-Raji CO, Adebiyi AI, Oluwayelu DO (2018) A slaughterhouse survey for Porcine circovirus type 2 in commercial pigs in Ibadan, southwest Nigeria. Folia Vet 62:30–34. https://doi.org/10.2478/fv-2018-0014
Banhazi TM, Seedorf J, Rutley DL, Pitchford WS (2008) Identification of risk factors for sub-optimal housing conditions in Australian piggeries: part 2 Airbone pollutants. J Agric Saf Health 14:21–39. https://doi.org/10.13031/2013.24122
Brewer MT, Greve JH (2011) Diseases of swine, 11th edn. John Wiley & Sons Inc, New Jersey, USA
Cargill C (2019) The impact of environment on production and health. In: Regional symposium on research into smallholder pig production, health, and pork safety. ILRI, Hanoi, Vietnam
Devleesschauwer B, Torgerson P, Charlier J, et al (2022) Prevalence: tools for prevalence assessment studies.. R package version 0.4.1, https://cran.r-project.org/package=prevalence.
Dione M, Masembe C, Akol J et al (2018) The importance of on-farm biosecurity: sero-prevalence and risk factors of bacterial and viral pathogens in smallholder pig systems in Uganda. Acta Trop 187:214–221. https://doi.org/10.1016/j.actatropica.2018.06.025
Dione M, Ouma E, Opio F et al (2016) Qualitative analysis of the risks and practices associated with the spread of African swine fever within the smallholder pig value chains in Uganda. Prev Vet Med 135:102–112. https://doi.org/10.1016/j.prevetmed.2016.11.001
Dohoo I, Martin W, Stryhn H (2003) Veterinary epidemiologic research, 2nd edn. VER, Incorporated, Charlottetown, Prince Edward Island, Canada
Eneku W, Mutebi F, Mwiine F et al (2018) Porcine circovirus type 2 – systemic disease on pig farms and associated knowledge of key players in the pig industry in central Uganda. Int J Vet Sci Med 6:178–185. https://doi.org/10.1016/j.ijvsm.2018.08.004
Fablet C, Marois-Créhan C, Simon G et al (2012) Infectious agents associated with respiratory diseases in 125 farrow-to-finish pig herds: a cross-sectional study. Vet Microbiol 157:152–163. https://doi.org/10.1016/j.vetmic.2011.12.015
Gillespie J, Opriessnig T, Meng X et al (2009) Porcine circovirus type 2 and porcine circovirus-associated disease. J Vet Intern Med 23:1151–1163. https://doi.org/10.1111/j.1939-1676.2009.0389.x
Halbur PG, Paul PS, Meng XJ et al (1996) Comparative pathogenicity of nine US porcine reproductive and respiratory syndrome virus (PRRSV) isolates in a five-week-old cesarean-derived, colostrum-deprived pig model. J Vet Diagnostic Investig 8:11–20. https://doi.org/10.1177/104063879600800103
Ikwap K, Erume J, Owiny DO et al (2014) Salmonella species in piglets and weaners from Uganda: prevalence, antimicrobial resistance and herd-level risk factors. Prev Vet Med 115:39–47. https://doi.org/10.1016/j.prevetmed.2014.03.009
ILRI (2004) Standard operating procedures (SOPs) for animal experimentation. ILRI, Nairobi
Jonsson L (2013) Emerging infectious diseases: using PCV2 as a model of disease transmission dynamics at the livestock-wildlife interface in Uganda
Kungu JM, Masembe C, Apamaku M et al (2019) Pig farming systems and cysticercosis in northern Uganda. Rev d’élevage médecine vétérinaire des pays Trop 72:115. https://doi.org/10.19182/remvt.31254
Laisse CJ, Souza CK, Pereira PR et al (2018) Detection and phylogenetic characterization of porcine circovirus 2 from pigs in Mozambique. J Vet Diagnostic Investig 30:342–347. https://doi.org/10.1177/1040638718769266
MAFF (1986) Manual of veterinary parasitological laboratory techniques. Great Britain
Marsteller T, Fenwick B (1999) Actinobacillus pleuropneumoniae disease and serology. Swine Heal Prod 7:161–165
Muhangi D, Masembe C, Berg M, et al (2014) Practices in the pig value chain in Uganda; implications to African swine fever transmission. Livest Res Rural Dev 26:18. http://www.lrrd.org/lrrd26/5/muha26094.htm
Muhanguzi D, Lutwama V, Mwiine FN (2012) Factors that influence pig production in central Uganda - case study of Nangabo sub-county, Wakiso district. Vet World 5:346–351. https://doi.org/10.5455/vetworld.2012.346-351
Murtaugh M, Genzow M (2011) Immunological solutions for treatment and prevention of porcine reproductive and respiratory syndrome (PRRS). Vaccine 29:8192–8204. https://doi.org/10.1016/j.vaccine.2011.09.013
Murtaugh MP, Xiao Z, Zuckermann F (2002) Immunological responses of swine to porcine reproductive and respiratory syndrome virus infection. Viral Immunol 15:533–547. https://doi.org/10.1089/088282402320914485
Nelson EA, Christopher-Hennings J, Benfield DA (1994) Serum immune responses to the proteins of porcine reproductive and respiratory syndrome (PRRS) virus. J Vet Diagnostic Investig 6:410–415. https://doi.org/10.1177/104063879400600402
Ojok L, Okuni J, Hohloch C, et al (2013) Detection and characterisation of porcine circovirus 2 from Ugandan pigs. Indian J Vet Pathol 37:77–80. https://www.researchgate.net/publication/256093421
Opriessnig T, Thacker E, Yu S et al (2004) Experimental reproduction of postweaning multisystemic wasting syndrome in pigs by dual infection with Mycoplasma hyopneumoniae and Porcine circovirus type 2. Vet Pathol 41:624–640. https://doi.org/10.1354/vp.41-6-624
Otake S, Dee S, Corzo C et al (2010) Long-distance airborne transport of infectious PRRSV and Mycoplasma hyopneumoniae from a swine population infected with multiple viral variants. Vet Microbiol 145:198–208. https://doi.org/10.1016/j.vetmic.2010.03.028
Ouma E (2017) Overview of ILRI’s smallholder pig value chain efforts in Lira district. Hoima, Uganda
Ouma E, Dione M, Lule P, et al (2014) Characterization of smallholder pig production systems in Uganda: constraints and opportunities for engaging with market systems. Livest Res Rural Dev 26:. https://doi.org/10.22004/ag.econ.160677
Pileri E, Mateu E (2016) Review on the transmission porcine reproductive and respiratory syndrome virus between pigs and farms and impact on vaccination. Vet Res 47:1–13. https://doi.org/10.1186/s13567-016-0391-4
Qin S, Ruan W, Yue H et al (2018) Viral communities associated with porcine respiratory disease complex in intensive commercial farms in Sichuan province, China. Sci Rep 8:1–9. https://doi.org/10.1038/s41598-018-31554-8
R Core Team (2019) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.r-project.org/
Roesel K, Dohoo I, Baumann M et al (2017) Prevalence and risk factors for gastrointestinal parasites in small-scale pig enterprises in central and eastern Uganda. Paras Res 116:335–345. https://doi.org/10.1007/s00436-016-5296-7
Segalés J, Kekarainen T, Cortey M (2013) The natural history of porcine circovirus type 2: from an inoffensive virus to a devastating swine disease? Vet Microbiol 165:13–20. https://doi.org/10.1016/j.vetmic.2012.12.033
Thacker E, Halbur P, Ross R et al (1999) Mycoplasma hyopneumoniae potentiation of Porcine reproductive and respiratory syndrome virus-induced pneumonia. J Clin Microbiol 37:620–627
UBOS (2020) Annual Statistical Abstract. Entebbe
Vigre H, Ersbøll A, Sørensen V (2003) Decay of acquired colostral antibodies to Actinobacillus pleuropneumoniae in pigs. J Vet Med 50:430–435. https://doi.org/10.1046/j.0931-1793.2003.00700.x
Acknowledgements
We thank Jane Poole and Nicholas Ndiwa (Research Methods Group, ILRI Nairobi) for their guidance in statistical analyses. The assistance received from the district veterinary officer of Lira district, Dr Ogwal Anthony, and extension officers Cecil Podpodo and Benard Okello during data collection is highly appreciated. We thank staff of Central Diagnostic Laboratory of CoVAB, Makerere University (Dickson, Edrine, Jacky, and Kanyike) for their support during laboratory analyses of samples. We thank and appreciate the cooperation of pig farmers in Lira district for their voluntary acceptance in this study.
Funding
This work was supported by the German Academic Exchange Service (DAAD) as Peter Oba’s PhD research fellowship through International Livestock Research Institute (ILRI). We acknowledge funding support from the OneCGIAR initiative “Sustainable Animal Productivity for Livelihoods, Nutrition and Gender inclusion (SAPLING)”. This research was conducted as part of the CGIAR Research Program on Livestock, and we thank donors who support its work through their contributions to the CGIAR Trust Fund. The funding agency had no role in the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Michel M. Dione, Barbara Wieland, and Peter Oba conceived the study; Michel M. Dione, Frank N. Mwiine, Joseph Erume, Barbara Wieland, and Peter Oba designed the study; Peter Oba collected the data; Peter Oba and Michel M. Dione analyzed and interpreted the data; Peter Oba wrote the first draft of the manuscript; all authors reviewed subsequent drafts and approved the final version of the manuscript for submission.
Corresponding author
Ethics declarations
Ethics approvals
This study involved a questionnaire-based survey of farmers as well as blood and fecal sampling of pigs. The study protocol was assessed and approved by the institutional review board (IRB no. SBLS/REC/18/008) of Makerere University’s College of Veterinary Medicine, Animal Resources and Biosecurity (CoVAB), Uganda National Council of Science and Technology (UNCST reg no. A590), ILRI’s Institutional Research Ethics Committee (IREC no. IREC2018-23), and ILRI’s Institutional Animal Care and Use Committee (IACUC2018-22). Pig sampling was done by trained veterinarians who adhered to ethical standards of animal welfare.
Consent to participate
Informed (written) consent was obtained from all research participants included in the study by signing consent forms (translated into a local language) given to them.
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Robin Flynn
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oba, P., Wieland, B., Mwiine, F.N. et al. Co-infections of respiratory pathogens and gastrointestinal parasites in smallholder pig production systems in Uganda. Parasitol Res 122, 953–962 (2023). https://doi.org/10.1007/s00436-023-07797-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-023-07797-4