Abstract
Variants which disrupt splicing are a frequent cause of rare disease that have been under-ascertained clinically. Accurate and efficient methods to predict a variant’s impact on splicing are needed to interpret the growing number of variants of unknown significance (VUS) identified by exome and genome sequencing. Here, we present the results of the CAGI6 Splicing VUS challenge, which invited predictions of the splicing impact of 56 variants ascertained clinically and functionally validated to determine splicing impact. The performance of 12 prediction methods, along with SpliceAI and CADD, was compared on the 56 functionally validated variants. The maximum accuracy achieved was 82% from two different approaches, one weighting SpliceAI scores by minor allele frequency, and one applying the recently published Splicing Prediction Pipeline (SPiP). SPiP performed optimally in terms of sensitivity, while an ensemble method combining multiple prediction tools and information from databases exceeded all others for specificity. Several challenge methods equalled or exceeded the performance of SpliceAI, with ultimate choice of prediction method likely to depend on experimental or clinical aims. One quarter of the variants were incorrectly predicted by at least 50% of the methods, highlighting the need for further improvements to splicing prediction methods for successful clinical application.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The diagnosis of rare disorders has been revolutionised in recent years thanks to the availability and widespread adoption of next generation sequencing technologies capable of detecting disease-causing variants. With the ever-decreasing prices of whole-exome sequencing (WES) and whole-genome sequencing (WGS) comes an increased use of these approaches, leading to the detection of more genetic variants than ever before. This brings with it a major challenge in understanding what these variants do, since our ability to detect them has far outstripped our ability to meaningfully interpret their effects, particularly outside of protein coding regions. As a result, even with WGS, around half of patients with rare disorders do not get a diagnosis (Turro et al. 2020; Stranneheim et al. 2021).
While estimates vary widely (Lord and Baralle 2021), it is thought somewhere between 15 and 60% of disease causing variants affect splicing (Krawczak et al. 1992; López-Bigas et al. 2005). Generally speaking, in diagnostic and research variant prioritisation pipelines, variants which fall within the 2 bp canonical splice acceptor or donor sites will be classed as splice-affecting, while variants outside of those small regions are often not assessed for splicing impact. It is common for intronic and synonymous variants to be filtered out, while missense variants are generally assessed for their impact on protein structure and function without consideration for the role they may play in splicing. All of these variant types, however, can and do impact splicing and cause disease. This approach has led to an under-ascertainment of splice-affecting variants clinically (Lord et al. 2019). What is needed, particularly with the increasing use of WGS over WES enabling the detection of far more intronic variants than before, is a way to effectively triage which variants are splice-affecting and which are not.
Currently, under ACMG/AMP guidelines (Richards et al. 2015), in silico splicing prediction approaches may be used as supporting evidence for genetic diagnosis if multiple independent tools predict an impact on splicing. Experimental validation of splicing effects using RT-PCR, mini-genes or RNAseq is often required to confidently establish a variant’s impact on splicing, but such approaches are time-consuming and expensive to perform at scale. Recent years have seen a plethora of innovative new approaches to splicing prediction, with many new tools being generated, often utilising machine learning. If a high degree of accuracy and reliability can be obtained from in silico approaches, we may be able to move away from requiring experimental confirmations, or at the least, have an efficient method to triage variants most in need of validation. This would require highly accurate algorithms and extensive testing in the clinical setting to give sufficient confidence in these optimal approaches.
The Splicing Variants of Unknown Significance (VUS) challenge in the 6th Critical Assessment of Genome Interpretation (CAGI6) sought to assess splicing prediction accuracy on a set of clinically ascertained, functionally validated variants. This enabled performance comparison of many cutting-edge splicing prediction approaches and gave insights into the types of variants not currently well-captured by these methods.
Methods
Variant selection and validation
As previously described in Wai et al. 2020 (Wai et al. 2020), a total of 64 variants were ascertained through Wessex Regional Genetics Laboratory in Salisbury (52 variants) or the Splicing and Disease research study (12 variants) at the University of Southampton, ethically approved by the Health Research Authority (IRAS Project ID 49685, REC 11/SC/0269) and by the University of Southampton (ERGO ID 23056). Informed consent was provided for all patients for splicing studies to be conducted. All variants had been, or were undergoing RT-PCR analysis to determine their impact on splicing using RNA from whole blood collected in PAXgene tubes, again as previously described (Wai et al. 2020).
Eight variants were excluded from the final analysis (unable to establish splicing impact before analysis period (n = 3), incorrect gene/variant annotations given in the data set distributed (n = 3), variant found to impact gene expression rather than splicing (n = 2)), giving a total of 56 variants in the final assessment set (Supplementary Table 1), which span a wide range of rare disease and cancer predisposition associations, none of which had had their impact on splicing published previously.
The Splicing VUS challenge
Variants were distributed as a tab delimited text file, including the following information: HGNC identifier, chromosome, position, reference allele, alternative allele, gene and strand. Entrants also had access to 256 previously published variants (Wai et al. 2020) obtained and validated by the same approach to aid in method development/testing.
Challenge participants submitted their entries in the form of tab delimited text files, including the variant information, a binary prediction of whether a variant affected splicing or not (1 = yes, 0 = no), along with a score for the probability of the variant affecting splicing and the level of confidence in the prediction given. All assessments were based on the binary splice-affecting prediction alone.
Challenge assessment
The performance of each prediction model was assessed by calculating and comparing a series of metrics: overall accuracy, area under the receiver operating characteristic curve (AUC), sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV). AUC and confidence intervals (2000 stratified bootstrap replicates) were calculated using the pROC package (Robin et al. 2011) in R v3.5.1 (R Core Team 2018), and plots made with ggplot2 (Wickham 2009). Performance of each method was compared across binned splicing locations—Near Acceptor (acceptor ± 10 bp), Near Donor (donor ± 10 bp), Exonic Distant (exonic, 11 bp or more from either splice site), and Intronic Distant (intronic, 11 bp or more from either splice site. For grouped analyses, exonic distant and intronic distant variants were grouped together due to low numbers). These scores were based on the concordance of the binary classification of the variants provided by each team/model (1 = splice-affecting and 0 = not splice-affecting) with the experimental validation of the splicing impact.
SpliceAI (Jaganathan et al. 2019) and CADD v1.6 (Kircher et al. 2014) (which incorporates SpliceAI predictions) were included in the assessment alongside the challenge models as a comparison to emerging industry standards. CADD-phred scores were obtained by uploading a VCF to the CADD webserver (https://cadd.gs.washington.edu/score). SpliceAI scores were obtained from Ensembl’s Variant Effect Predictor (VEP) web interface (McLaren et al. 2016) (44 variants scored) or using the SpliceAI webserver from the Broad Institute (https://spliceailookup.broadinstitute.org/, 11 variants that were not scored by VEP; options: hg38, masked scores, max distance 50 bp). A cutoff of 0.2 was used for SpliceAI scores, and 18 for CADD.
Results
Variant characteristics of challenge set
Of the 56 variants in the final analysis, the majority (n = 49, 87.5%) were SNVs, with 7 indels (12.5%). The variants fell within 42 different genes, broadly representative of clinical genetics referrals in the UK, with the majority of genes having a single variant in the set, and only 7 genes with >1 variant (BRCA1 n = 6, FBN1 n = 4, MYBPC3 n = 3, BRCA2 n = 2, SCN5A n = 2, APC n = 2, USP7 n = 2). 37 variants (66%) were found to affect splicing, while 19 (34%) had no observable impact.
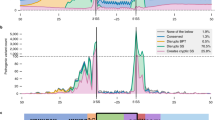
Variants were divided into 5 groups by their positions relative to intron–exon boundaries. There were 16 variants within 10 bp of a splice acceptor site (NearAcc), and 23 within 10 bp of a splice donor site (NearDon). 10 exonic variants > 10 bp from either splice site were classed as Exonic > 10. Intronic variants > 10 bp from their nearest splice site were termed Intronic Distant (six upstream of the acceptor, one downstream of the donor). The locations of all variants relative to the intron–exon boundary and whether the variants were determined to be splice disrupting or not are given in Fig. 1.
Challenge participants
Eight teams submitted predictions for the challenge, with two teams submitting predictions from multiple models, giving 12 models altogether. Table 1 gives a summary of the approach taken by each model, which was provided by the challenge entrants upon submission of their predictions, but blinded to the assessors until after the assessment period.
Model performance across 56 variants
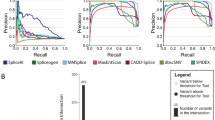
Table 2 summarises the performance metrics of the 12 models, along with CADD and SpliceAI. Full variant information, scores and binary predictions for the 12 models, SpliceAI and CADD and experimental outcome of splicing status are given in Supplementary Table 1. The ROC plots for each model are shown in Fig. 2, and Supplementary Fig. 1 shows the performance of each method on each variant across the splicing region.
Receiver operating characteristic (ROC) curves of model performance based on prediction scores. For Area Under Curve (AUC), see Table 2
No single approach performed optimally on all assessment metrics (Table 2). Overall accuracy was joint highest in Teams 4 and 8 at 0.82, with Team 4 also achieving the highest binary outcome AUC at 0.839 (Fig. 2). Team 8 ranked highest on the related metrics for sensitivity (0.919) and NPV (0.800), indicating its permissive prediction approach (i.e., favouring sensitivity over specificity). Conversely, Team 5’s Model 2 performed the best in terms of specificity (0.947) and PPV (0.947), with the lowest proportion of false positive findings. All three models by Team 1, plus Team 4 and Team 6 achieved over 70% in both sensitivity and specificity, indicating more balanced performance.
Included as comparators were SpliceAI with a cutoff of 0.2 and CADD with a cutoff of 18. SpliceAI was competitive with the challenge entrants, ranking near-top but not top on all metrics, and indeed top in the AUC when measured using prediction score rather than binary prediction outcome. CADD, however, performed poorly on the challenge set with specificity in particular being very low (0.263).
Performance comparison by variant type
To get an overall impression of the performance of the methods on different types of variants, variants were grouped by location relative to their nearest splice site (Fig. 3), as described in Methods. All methods performed better on exonic distant variants than intronic distant variants, with the exception of SpliceAI, which correctly predicted all seven intronic distant variants. Across methods, there was a high degree of consistency in the proportion of variants correctly predicted in the near acceptor region, and a high degree of variance in performance in the intronic distant set. The types of error differed across regions, with the near acceptor region and exonic distant region having very few false positive predictions across all methods, while almost all methods gave false positive predictions in the near donor and intronic distant regions (Supplementary Fig. 2).
We also compared the performance of the approaches on SNVs vs indels, and found all methods except CADD had higher accuracy on SNVs than indels (Supplementary Fig. 3).
Some variants are consistently mispredicted
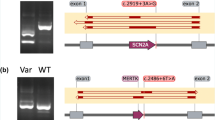
21 of the variants (37.5%) were correctly predicted by all 12 submitted prediction methods. None of the variants were incorrectly predicted by all methods, but 14 variants (25%) were predicted correctly by ≤50% of the methods, with two variants only being correctly predicted by a single method. These were a splice-affecting single nucleotide deletion 4 bp from a splice acceptor site in KANSL1 (correctly predicted by Team 3) and an SNV in the last base of an exon in TRPM6 which despite altering the conserved last G nucleotide did not affect splicing in functional testing (correctly predicted by Team 4; Fig. 4).
Discussion
The CAGI6 Splicing VUS challenge assessed the performance of 14 prediction approaches on a set of 56 clinically relevant variants whose impact on splicing had been functionally tested using RT-PCR. A variety of approaches were adopted, and several methods equalled or exceeded the performance of the emergent field leader, SpliceAI.
While Teams 4 and 8 had joint highest overall accuracy, there was no single optimal method for the Splicing VUS challenge, since several different models performed optimally on different metrics. Choice of approach may, therefore, be dependent on the specific nature of the predictions required. Seeking a molecular diagnosis for a particular family may favour sensitivity over specificity, since overlooking a causal variant would prevent this aim, so Team 8’s approach with almost 92% sensitivity may be preferred. Seeking confident splice disrupting candidates for functional validation or mechanistic research may call for greater specificity than sensitivity to avoid wasting resources on false positive variants that do not have an impact, in which case Team 5’s model 2 with almost 95% specificity may be the strategy of choice.
SpliceAI and CADDv1.6 were chosen as comparators for the entrants to the Splicing VUS challenge and were run by the assessors on the 56 challenge variants. SpliceAI has been emerging as a field leader in recent years, with accuracies >90% attained in several studies (Wai et al. 2020; Ha et al. 2021; Strauch et al. 2022), although variable performance reported by some (Riepe et al. 2020) which is more consistent with our observed 80.4% overall accuracy in this study.
CADD did not perform well on the challenge variants, achieving an overall accuracy of 62.5%. However, this was predominantly driven by a very low specificity, which is to be expected from CADD, since it is not only the impact on splicing being assessed by the tool, but overall deleteriousness. For example, missense variants which were not found to affect splicing in the challenge set may still have been pathogenic through impact on protein structure and/or function. For such variants, CADD would accurately classify them as deleterious in general, but in our assessment solely of splicing impact, this would appear as a false positive, lowering CADD’s specificity. Notably, the version of CADD included in the assessment (v1.6) includes SpliceAI and additional splicing prediction tools in its underlying model (Rentzsch et al. 2021). Scoring the challenge variants with CADD v1.5 which did not include these splicing metrics resulted in an overall accuracy around 44.6% (data not shown). From these values it is clear that the explicit inclusion of splicing prediction methods within CADD’s underlying model has improved its ability to predict variants that impact splicing. CADD’s broad approach makes it a versatile tool for prediction of deleteriousness for many different variant types. At present, however, if predicting a variant’s splicing impact is the sole aim, the use of more specialised splicing tools is more appropriate.
Of note, SpliceAI featured heavily across the predictive strategies, being the sole predictive method for Team 6 and contributing heavily to the predictions of Team 4, which were weighted by MAF, as well as being run as a comparator by the assessors. Differences in the performance of these approaches highlight that even with the same nominal method, there can be variance in predictions depending on how the scores are obtained, and the thresholds that are used to determine positive predictions. There were just three approaches that did not include SpliceAI as part of their predictions, two utilising instead recent machine learning-based prediction tools SQUIRLS (Danis et al. 2021) and SPiP (Leman et al. 2022), and one based on the splicing prediction tools available within the Alamut software, which has been widely used in clinical practice. Of the three, SPiP was the only method to achieve greater accuracy than SpliceAI.
A major strength of the challenge in terms of providing a real-world assessment of the performance of these tools is the ascertainment of the challenge variants from genuine clinical practice, where potential splice altering variants in genes relevant to the patient’s presentation were referred for validation. This is precisely the type of variant splicing prediction models should be tested on when assessing their suitability for clinical application in rare disorders. It highlights that even in exceptionally well-studied genes, such as the BRCA genes, challenges in variant interpretation remain, since 3 of 8 variants across BRCA1 and BRCA2 were incorrectly predicted by over half of challenge methods, and only two of these were accurately predicted by all methods. However, the relatively small sample size makes it difficult to draw any major inferences and is a significant limitation of the study. Apparent variance in performance may be stochastic at such a sample size, and may not be fully reflective of overall performance in a wider context. It also made drawing firm conclusions about performance in subsets of the data, e.g., split by location, variant type, or disease group challenging. However, ascertaining a large body of clinical variants, validating the splicing impact and keeping that private, as is needed for a blinded challenge, such as the CAGI6 Splicing VUS challenge, raises ethical concerns. Accurate and timely variant interpretation is reliant on sharing of data, and withholding a large body of functionally validated variants from resources, such as ClinVar (Landrum et al. 2018), which are heavily used in clinical assessment of variants does not represent good practice.
This small but highly clinically relevant challenge assessed the performance of 12 prediction methods plus SpliceAI and CADD on 56 clinically ascertained variants and found SpliceAI weighted by allele frequency and SPiP to be the most accurate overall, while other methods had particular strengths in their sensitivity or specificity. A quarter of variants were incorrectly predicted by half or more of the methods, showing there is still improvement to be made. Furthermore, this challenge was limited to a binary outcome—whether or not splicing was disrupted, but did not address the nature of that disruption. Disruption to splicing is often complex (e.g., multiple different splicing events induced), incomplete (e.g., aberrant and wild-type splicing observed), and can be further complicated by nonsense mediated decay. This will present an even greater challenge for accurate prediction than the binary outcome assessed here. A larger assessment set that would enable further investigation of the types of variants that are consistently incorrectly predicted may help direct efforts for refinement of models moving forwards.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Cheng J et al (2019) MMSplice: modular modeling improves the predictions of genetic variant effects on splicing. Genome Biol 20(1):48
Danis D, Jacobsen JOB, Carmody LC, Gargano MA, McMurry JA, Hegde A, Haendel MA, Valentini G, Smedley D, Robinson PN (2021) Interpretable prioritization of splice variants in diagnostic next-generation sequencing. Am J Hum Genet 108(9):1564–1577
Ha C, Kim JW, Jang JH (2021) Performance evaluation of SpliceAI for the prediction of splicing of NF1 variants. Genes (basel) 12:1308
Jagadeesh KA et al (2019) S-CAP extends pathogenicity prediction to genetic variants that affect RNA splicing. Nat Genet 51(4):755–763
Jaganathan K, Kyriazopoulou Panagiotopoulou S, McRae JF, Darbandi SF, Knowles D, Li YI, Kosmicki JA, Arbelaez J, Cui W, Schwartz GB et al (2019) Predicting splicing from primary sequence with deep learning. Cell 176(3):535–548
Jian X, Boerwinkle E, Liu X (2014) In silico prediction of splice-altering single nucleotide variants in the human genome. Nucleic Acids Res 42(22):13534–13544
Karczewski KJ et al (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581(7809):434–443
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J (2014) A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet 46(3):310–315
Krawczak M, Reiss J, Cooper DN (1992) The mutational spectrum of single base-pair substitutions in mRNA splice junctions of human genes: causes and consequences. Hum Genet 90:41–54
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, Gu B, Hart J, Hoffman D, Jang W et al (2018) ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46(D1):D1062–D1067
Leman R, Parfait B, Vidaud D, Girodon E, Pacot L, Le Gac G, Ka C, Ferec C, Fichou Y, Quesnelle C et al (2022) SPiP: Splicing Prediction Pipeline, a machine learning tool for massive detection of exonic and intronic variant effects on mRNA splicing. Hum Mutat 43(12):2308–2323
López-Bigas N, Audit B, Ouzounis C, Parra G, Guigó R (2005) Are splicing mutations the most frequent cause of hereditary disease? FEBS Lett 579:1900–1903
Lord J, Baralle D (2021) Splicing in the diagnosis of rare disease: advances and challenges. Front Genet 12:689892
Lord J, Gallone G, Short PJ, McRae JF, Ironfield H, Wynn EH, Gerety SS, He L, Kerr B, Johnson DS et al (2019) Pathogenicity and selective constraint on variation near splice sites. Genome Res 29:159–170
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GR, Thormann A, Flicek P, Cunningham F (2016) The ensembl variant effect predictor. Genome Biol 17(1):122
R Core Team (2018) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Rentzsch P, Schubach M, Shendure J, Kircher M (2021) CADD-Splice-improving genome-wide variant effect prediction using deep learning-derived splice scores. Genome Med 13(1):31
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17:405–424
Riepe TV, Khan M, Roosing S, Cremers FPM, ‘t Hoen PAC (2020) Benchmarking deep learning splice prediction tools using functional splice assays. Authorea 42:799–810. https://doi.org/10.22541/au.160081230.07101269
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, Muller M (2011) pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12:77
Stranneheim H, Lagerstedt-Robinson K, Magnusson M, Kvarnung M, Nilsson D, Lesko N, Engvall M, Anderlid BM, Arnell H, Johansson CB et al (2021) Integration of whole genome sequencing into a healthcare setting: high diagnostic rates across multiple clinical entities in 3219 rare disease patients. Genome Med 13:40
Strauch Y, Lord J, Niranjan M, Baralle D (2022) CI-SpliceAI-Improving machine learning predictions of disease causing splicing variants using curated alternative splice sites. PLoS ONE 17:e0269159
Turro E, Astle WJ, Megy K, Graf S, Greene D, Shamardina O, Allen HL, Sanchis-Juan A, Frontini M, Thys C et al (2020) Whole-genome sequencing of patients with rare diseases in a national health system. Nature 583:96–102
Wai HA, Lord J, Lyon M, Gunning A, Kelly H, Cibin P, Seaby EG, Spiers-Fitzgerald K, Lye J, Ellard S et al (2020) Blood RNA analysis can increase clinical diagnostic rate and resolve variants of uncertain significance. Genet Med 22:1005–1014
Wickham H (2009) ggplot2 Elegant graphics for data analysis introduction. Use R. Springer, New York. https://doi.org/10.1007/978-0-387-98141-3_1
Yeo G, Burge CB (2004) Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Comput Biol 11(2–3):377–394
Acknowledgements
We thank the CAGI organisers for their commitment to improving variant interpretation and for making this challenge happen. The CAGI experiment is supported by NIH U24 HG007346. We acknowledge the NIHR Clinical Research Network (CRN) in recruiting the participants and the Musketeers Memorandum, as well as support from the NIHR UK Rare Genetic Disease Consortium. The authors acknowledge the use of the IRIDIS High Performance Computing Facility, and associated support services at the University of Southampton, in the completion of this work. For the purpose of open access, the author has applied a CC BY public copyright licence to any Author Accepted Manuscript version arising from this submission.
Funding
The Baralle Lab is supported by the NIHR Research Professorship awarded to DB (RP-2016-07-011). JL is supported by an Anniversary Fellowship from the University of Southampton. Some of the functional validations of variants were funded by a Wessex Medical Research Innovation Grant awarded to JL. RDB is supported by a New South Wales Health Cardiovascular Disease Senior Scientist Grant.
Author information
Authors and Affiliations
Contributions
DB and JL conceived of the challenge. AGLD, DJB and JL selected variants to include in the set, which had been functionally validated by HAW and DJB. JL assessed challenge entrants and conducted data analysis. CJO conducted additional analyses and presented the findings at the CAGI6 conference. All further authors submitted prediction methods in response to the challenge. JL drafted the manuscript, with revision suggestions and final approval from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose. On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethics approval
Informed consent was provided for all patients for splicing studies to be conducted. Patients were recruited from Wessex Regional Genetics Laboratory in Salisbury (52 variants) or the Splicing and Disease research study (12 variants) at the University of Southampton, ethically approved by the Health Research Authority (IRAS Project ID 49685, REC 11/SC/0269) and by the University of Southampton (ERGO ID 23056).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lord, J., Oquendo, C.J., Wai, H.A. et al. Predicting the impact of rare variants on RNA splicing in CAGI6. Hum. Genet. (2024). https://doi.org/10.1007/s00439-023-02624-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00439-023-02624-3