Abstract
Joubert syndrome (JS) is a recessively inherited congenital ataxia characterized by hypotonia, psychomotor delay, abnormal ocular movements, intellectual disability, and a peculiar cerebellar and brainstem malformation, the “molar tooth sign.” Over 40 causative genes have been reported, all encoding for proteins implicated in the structure or functioning of the primary cilium, a subcellular organelle widely present in embryonic and adult tissues. In this paper, we developed an in vitro neuronal differentiation model using patient-derived induced pluripotent stem cells (iPSCs), to evaluate possible neurodevelopmental defects in JS. To this end, iPSCs from four JS patients harboring mutations in distinct JS genes (AHI1, CPLANE1, TMEM67, and CC2D2A) were differentiated alongside healthy control cells to obtain mid-hindbrain precursors and cerebellar granule cells. Differentiation was monitored over 31 days through the detection of lineage-specific marker expression by qRT-PCR, immunofluorescence, and transcriptomics analysis. All JS patient-derived iPSCs, regardless of the mutant gene, showed a similar impairment to differentiate into mid-hindbrain and cerebellar granule cells when compared to healthy controls. In addition, analysis of primary cilium count and morphology showed notable ciliary defects in all differentiating JS patient-derived iPSCs compared to controls. These results confirm that patient-derived iPSCs are an accessible and relevant in vitro model to analyze cellular phenotypes connected to the presence of JS gene mutations in a neuronal context.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ciliopathies are an expanding group of disorders due to mutations in genes involved in the formation and function of the primary cilium, a subcellular organelle implicated in key biological processes both during embryonic and adult life. Ciliopathies affect several organs such as the central nervous system (CNS), retina, kidneys, liver, and skeleton, reflecting the multiple roles of primary cilia in different tissues (Mitchison and Valente 2017). They are both genetically heterogeneous and pleiotropic, as in the case of Joubert syndrome (JS, MIM213300), a ciliopathy characterized by a unique cerebellar and brainstem malformation termed the “molar tooth sign” (MTS). This consists of cerebellar vermis dysplasia, a deepened interpeduncular fossa, and thickened, elongated, and poorly oriented superior cerebellar peduncles (SCPs), which often fail to decussate (Romani et al. 2013). Clinically, neurological manifestations of JS comprise developmental delay, oculomotor apraxia, hypotonia, ataxia, intellectual deficit, and breathing dysregulation in the neonatal age. Other manifestations may involve eye (Leber congenital amaurosis, retinal dystrophy), kidney (nephronophthisis, polycystic kidney disease), skeleton (mainly polydactyly), liver (congenital fibrosis), and oral-facial and laterality defects, defining heterogeneous phenotypic subgroups sharing the MTS (Mitchison and Valente 2017).
More than 40 genes have been identified to date, overall accounting for about two-thirds of the cases. Despite this extreme genetic heterogeneity, five JS major genes (TMEM67, AHI1, CPLANE1, CEP290, and CC2D2A) were found to account for about half of genetically characterized cases (Bachmann-Gagescu et al. 2015; Mitchison and Valente 2017). Interestingly, while dysfunction of all these genes results in the same specific brain malformation (the MTS), they encode for proteins involved in different aspects of cilia biology: (i) TMEM67 is part of the Tectonic Complex within the transition zone (TZ), a specialized region of the cilium acting as ciliary gate and regulating both ciliogenesis and ciliary membrane composition (Garcia-Gonzalo et al. 2011); moreover, TMEM67 has been implicated in non-canonical Wnt signaling (Abdelhamed et al. 2015; Lee et al. 2017); (ii) AHI1 is located at the basal body and is known to stabilize the canonical Wnt pathway effector beta-catenin (Lancaster et al. 2011); (iii) CPLANE1, part of the CPLANE complex, resides at the basal body and has recently been related to Kif7 and the Shh pathway (Asadollahi et al. 2018); (iv) CEP290 localizes at the TZ where it plays an important regulatory role, preventing inappropriate entry of membrane-associated proteins into cilia and regulating the assembly pathway of the various protein complexes to build a functional TZ (Li et al. 2016); (v) finally, CC2D2A is a TZ protein involved in the organization of the vesicle fusion machinery at the periciliary membrane (Ojeda Naharros et al. 2017). So far, the role of the primary cilium in the developing cerebellum was only partially elucidated. It still remains unclear how malfunction of the proteins which impair cilia biology at different levels may eventually result in such a consistent neurodevelopmental defect as the MTS.
The development of induced pluripotent stem cells (iPSCs) from somatic cells represents a simple and elegant model to study human diseases, providing the means to maintain the specific patient’s genotype in vitro. iPSCs possess a cilium (Barabino et al. 2020) and can differentiate into almost every cell type, representing a powerful model to study the effect of selected mutations on the differentiation process leading to specific phenotypes. Here, we established iPSCs from JS patients carrying mutations in four different major genes (AHI1, TMEM67, CPLANE1, and CC2D2A) and characterized their ciliary and differentiation properties compared to iPSCs derived from healthy controls.
Materials and methods
Generation of iPSCs from control and JS patients
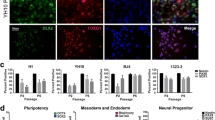
Dermal fibroblasts obtained from 2 healthy donors (CT-iPSCs, both young adult males) and 4 JS patients (JS-iPSCs, two males and two females, age range 10–27 years) carrying mutations in AHI1, CPLANE1, TMEM67, and CC2D2A (Table 1) were isolated and reprogrammed into iPSCs using the CytoTune™-iPS 2.0 Sendai Reprogramming Kit (Thermo Fisher Scientific). iPSC lines used for this study were produced by pooling of cells after reprogramming and characterized to confirm the expression of stemness markers and normal karyotype (Online Resources Figs. 1–8) as described in Online Resources. For all experiments, iPSCs were used at passages from 12 to 17.
Changes in gene expression in control and JS-derived iPSCs over the 31-day differentiation treatment. Relative fold change in the expression of ATOH1 (a), NEUROD1 (b), ZIC2 (c), PAX6 (d), and LMX1B (e) measured in each JS line normalized to D0 and then compared to the mean of CT-iPSC values. Only the CPLANE1 JS-iPSC line survived up to the day 31 time-point. n = 3 biological replicates (with 2 technical replicates for each); * is shown when the comparison, at each time point, of each JS-iPSCs vs CT-iPSCs (ctrl) is statistically significant (p < 0.05) using Wilcoxon-Mann–Whitney tests
Differentiation of iPSCs into cerebellar granule cells
Differentiation of iPSCs towards mid-hindbrain precursors and GCs was performed using a previously published protocol (Erceg et al. 2012; Compagnucci et al. 2016) further optimized in the lab. Briefly, proliferating iPSCs were cultured as a monolayer on matrigel-coated dishes in StemFlex medium (Thermo Fisher Scientific RRID:SCR_018018) with Penicillin/Streptomycin until they reached 70% confluence. iPSCs were then switched to a differentiating medium containing specific cocktails of cytokines (BMPs, FGF family members, Shh and Wnt family members) added at specific time points to induce first mid-hindbrain precursors and later cerebellar granule cells after 31 days of culture (Online Resource Fig. 9).
Immunofluorescence
iPSCs treated on coverslips were fixed in paraformaldehyde for 10 min, then permeabilized in PBS with 0.2% Triton X for 5 min, rinsed and blocked in PBS with 10% BSA prior to overnight incubation with the following primary antibodies: anti-ATOH1 (Abcam ab 85,513 RRID:AB_1924798), anti-NEUROD1 (Abnova PAB3857 RRID:AB_1575596), anti-ZIC2 (Millipore AB15392 RRID:AB_1977437), anti-β3-TUBULIN (Merck MAB1637 RRID:AB_2210524), followed by 1 h incubation with anti-rabbit Alexa fluor 555 and anti-mouse Alexa fluor 488 (1:5000) (Thermo Fisher Scientific). Nuclei were stained with Hoechst 33258 (Thermo Fisher Scientific). Images were analyzed with a confocal microscope (Zeiss). Ciliary count and length measurements were performed in iPSCs at the first differentiation time-point (D8). Cells at day 8 of differentiation were fixed in paraformaldehyde for 10 min then permeabilized in PBS with 0.2% Triton X for 5 min, rinsed and blocked in PBS with 10% BSA prior to overnight incubation with anti AcTUBULIN (Thermo Fisher Scientific cat 32-2700 RRID:AB_2533073) and anti-PERICENTRIN (Sigma HPA016820 RRID:AB_1855079), followed by 1 h incubation with rabbit Alexa fluor 555 and Anti mouse Alexa fluor 488 (1:5000). These experiments were performed in triplicate for each iPSC line, and cilia were measured and counted from 10 randomly selected fields on a microscope slide, counting a minimum of 100 cells per sample (n = 3 independent experiments) to calculate the percentage of ciliated cells (as the number of ciliated cells divided by the total number of cells). For ciliary length, the length from base to tip was analyzed with ImageJ (RRID:SCR_003070). The values shown for CT-iPSCs are the mean of the values for the two iPSC lines.
qRT-PCR
Total RNA was extracted using the High Pure RNA Isolation Kit (Roche RRID:SCR_004098) and then reversely transcribed with SuperScript II Reverse Transcriptase (Thermo Fisher Scientific) according to the manufacturer’s instructions. The resulting cDNAs for 3 biological replicates (with 2 technical replicates each) were analyzed by real-time PCR using SYBR green on the Light Cycler 480 (Roche RRID:SCR_020502) using primers listed in Online Resource Table 2–4, with β-ACTIN used as internal control. The relative expression was calculated using the 2ΔΔCT method. Relative fold change in marker gene expression was normalized to D0 for each line.
Statistical analysis
Descriptive statistics for the proportion of cells able to make primary cilia ciliary counts and cilium length are calculated as mean and standard deviation while qRT-PCR data are calculated as median and interquartile range (IQR), given the non-normal distribution of most variables according to the Shapiro-Wilk test. Instead, for categorical variables, absolute and relative frequencies were calculated. To compare impaired ciliation in controls and JS-derived iPSCs, two-tailed t-tests were performed. To assess the differences between qRT-PCR groups, Kruskal-Wallis tests were performed and post-hoc comparisons of each group vs the control group were conducted using the non-parametric Wilcoxon-Mann-Whitney test. A p-value of < 0.05 was considered statistically significant. Bonferroni multiple testing correction was not applied to the post-hoc p-values due to the small sample size and the explorative nature of the analyses.
Bulk transcriptomics
Total RNA was extracted from each cell line at day 8 and day 24 as described above. RNA quality and integrity, checked by RNA Integrity Number (RIN), were evaluated on an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Total RNA was sequenced with the Illumina NovaSeq 6000 technology in service at CeGaT GmbH (Tübingen, Germany) to obtain paired-end reads of 100 bp. At least 3 independent biological replicates were used for each line. Reads were trimmed with Trimmomatic (v0.39) (Bolger et al. 2014) to remove adapters and exclude low-quality reads from the analysis and aligned to the human reference genome using STAR aligner (v2.5.3a) (Dobin et al. 2013). FeatureCounts (v1.6.4) (Liao et al. 2014) was employed to calculate the reads mapping to exons based on the GENCODE human basic gene annotation. The resulting counts matrix represented transcript quantification summarized at the gene level, with the exclusion of reads mapping to multiple genomic locations. The counts matrix underwent a filtering process, retaining genes that exhibited a minimum of 1 CPM (counts per million) in at least 9 samples. Differential gene expression analysis was performed with DESeq2 (Love et al. 2014) bioconductor library (v1.30.1). The rpkm values were computed with edgeR (v3.40.2) bioconductor library (Robinson et al. 2010). Genes with statistically significant expression changes were identified using the following criteria: padj < 0.03, log2 fold change > 1, and log2 fold change < − 0.3. Heatmaps were made with ComplexHeatmap R package (v2.11.1) (Gu et al. 2016). Visualization of differentially expressed genes was achieved by creating a Volcano Plot using the ggplot2 package in R (v3.4.4). Transcriptomics data files are available on GEO (accession number: GSE254556) and analysis scripts are available on Zenodo (https://doi.org/10.5281/zenodo.10355929).
Results
iPSCs can be differentiated towards mid-hindbrain precursors and cerebellar neurons
CT-iPSCs (Ali et al. 2021) were first used to optimize the differentiation protocol to produce mid-hindbrain precursors and then cerebellar granule cells (CGCs), as previously reported (Erceg et al. 2012; Compagnucci et al. 2016). Differentiation was assessed every 7 days from day 0 (proliferating cells) to day 31 through qRT-PCR analysis of the differentiation markers ATOH1, NEUROD1, and ZIC2 (Fig. 1a–c). As expected, both CT-iPSC lines expressed these markers according to the sequential differentiation steps induced by the protocol (Online Resource Fig. 9). The expression of the early neurogenic marker ATOH1 was already evident at day 8, steadily increased till day 24, and remained high in both lines until the end of the protocol. The expression of NEUROD1, a transcription factor promoting cerebellar differentiation, was induced after 2 weeks and remained highly expressed until day 31. ZIC2, which is expressed in pluripotent cells and is maintained in granule neuron precursors (Luo et al. 2015), was detected throughout the protocol.
To further substantiate these observations, two additional neural genes important for the correct development and organization of mature cerebellum, PAX6, and LMX1B were monitored (Fig. 1d, e). In line with previous observations (Luo et al. 2015), CT-iPSCs showed a progressive increase of PAX6 levels at early time points, followed by a decrease upon lineage maturation. LMX1B, a key gene for midbrain and cerebellum development, was detected by D8 in CT-iPSCs with a strong increase after D24.
Cerebellar differentiation is impaired in iPSCs derived from JS patients
To evaluate how disruption of distinct JS-associated genes may impact on neuronal differentiation, iPSCs were generated from primary fibroblasts of four JS patients carrying pathogenic variants in either AHI1, CPLANE1, TMEM67, or CC2D2A (Table 1).
These iPSC lines (JS-iPSCs) were validated for pluripotency marker expression and karyotype integrity according to standard procedures (Ali et al. 2021). JS-iPSCs were treated according to the differentiation protocol validated in CT-iPSCs, and the expression of differentiation markers was analyzed by qRT-PCR over the 31-day time-course. D31 analysis could only be performed on CPLANE1 JS-iPSCs due to the consistent cell loss experienced by the other JS-iPSCs lines over the last week of differentiation (after D24).
All JS-iPSC lines showed altered expression profiles for the selected differentiation markers compared to control cells (Fig. 1). Expression levels of ATOH1 transcript were consistently lower in all JS lines compared to CT-iPSCs at D8 and D16, and at D24 for CPLANE1, AHI1, and TMEM67 lines. Similarly, NEUROD1 transcript levels were lower in patient lines particularly at D8 and D16, but significatively so at D24 for the AHI1 line. ZIC2 expression was not detected in AHI1 and CC2D2A JS-iPSCs, whereas it was present but lower than controls in CPLANE1 and TMEM67-mutated lines, especially at D24. Moreover, LMX1B mRNA levels were significantly lower in the TMEM67 line compared to controls at D16 and D24, while PAX6 expression increased in JS-iPSCs lines along the differentiation protocol, although with some variability across different mutant lines (Online Resource Table 1). Principal component analysis of D14 expression levels confirmed the clear difference in marker expression profile between differentiated controls and clustered mutant lines (Online Resource Fig. 10).
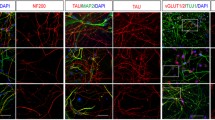
To confirm the altered mRNA expression profile seen in JS-iPSCs, immunofluorescence analysis was performed to investigate the expression of ATOH1, NEUROD1, and ZIC2 proteins in JS patient cells and in healthy controls over the differentiation treatment. In keeping with the qRT-PCR results, ATOH1, NEUROD1, and ZIC2 proteins were either reduced compared to controls (for AHI1 and CPLANE1 lines), or undetectable (for TMEM67 and CC2D2A lines) (Fig. 2 lower panels and Online Resource Fig. 11). Immunostaining for β3-TUBULIN, a microtubule protein of the tubulin family marking neurons, showed a well-defined network of positive cells in CT-iPSCs, particularly at D31 of differentiation (Online Resource Fig. 11). In contrast, JS-iPSCs showed rare branching of β3-TUBULIN-positive neurons (Fig. 2 arrows and Online Resource Fig. 11).
Differentiation of iPSCs towards mid-hindbrain and cerebellar lineages. Immunofluorescence co-staining for β3-TUBULIN (green) with the differentiation markers (red) ATOH1 (a-e, a’-e’), NEUROD1 (a’’–e’’, a’’’–e’’’) and ZIC2 (a’’’’–e’’’’, a’’’’’–e’’’’’) performed in CT-iPSCs (upper panel) and JS-iPSC lines (lower panels) after 8 and 24 days of differentiation (additional time points shown in Online Resource Fig. 11). Nuclei were counterstained with Hoechst (blue). Scale bar = 100 µm
In light of these observations, a differential gene expression analysis was performed on total RNA extracted from D8 and D24 for each differentiated iPSC line to survey a larger number of marker genes. Differentially expressed genes (DEGs) between CT and JS lines were analyzed at both timepoints (Fig. 3). Results show differences in the expression already at D8, with genes relating to the central nervous system development and in particular cerebellum markers (LMX1A, OLIG3) (Consalez et al. 2021; Lowenstein ED et al. 2021) being expressed only in controls cells. At D24, differences in gene expression became more evident, underlining for CT samples the expression of genes specific to the terminal phases of cerebellar differentiation (ASTN2, CNTN1, WNT3, PLXNA2, LMX1B) (Consalez et al. 2021) and the expression of genes encoding proteins of neuronal functionality (GRIA4, SYT8, GRIK1, P2RX2, PCP4, GRIK2) (Hansen et al. 2021; Rubio et al. 2001; Harashima et al. 2011). In contrast, the expression of these genes was significantly lower in JS lines, which showed early differentiation markers still predominantly expressed (ITGB1, CXCL12, HIF1A, EN2, GBX2, BMPR1B, CUL2, RPL37, RPL39, RPL35, PSMD2, PSMD3) (Consalez et al. 2021; Gilthorpe et al. 2002). Taken together, these data support the previous observation that JS-iPSC lines show an impaired progression along the differentiation time course and a reduced ability to reach the maturation state seen in controls.
Transcriptomics analysis of differentially expressed genes in CT and JS iPSC lines. a, b Volcano Plots for T8 (a) and T24 (b) representing the logarithm of fold-change (in x) and the negative logarithm of the p-value (in y) of genes that are significantly regulated (color) determined by predefined thresholds for both fold-change and p-value. c Rpkm expression heatmap (in z-score) of selected genes
iPSCs derived from JS patients show impaired ciliation and reduced length of primary cilia
The proportion of cells able to make primary cilia and the relative ciliary length was compared in control and JS-derived iPSCs at day 8 of differentiation by immunofluorescence staining using acetylated TUBULIN (AcTUBULIN) and PERICENTRIN as markers of the ciliary axoneme and basal body, respectively (Fig. 4a–a’’’’ and Online Resource Fig. 12). The percentage of ciliated cells in all JS-derived iPSCs was significantly lower compared to healthy controls, although to various degrees (control 54.5 ± 3.3%; AHI1 20.1 ± 5% (p < 0.005); CPLANE1 45.5 ± 3.9% (p < 0.05); TMEM67 38.7 ± 2.7% (p < 0.005); CC2D2A 17.1 ± 1.9% (p < 0.005)) (Fig. 4b). Three JS lines (AHI1, TMEM67, CC2D2A) also showed a significant reduction of ciliary length, while no significant change in length was observed in CPLANE1 mutant cells compared to controls (control 1.6 ± 0.05 µm; AHI1 1.3 ± 0.04 µm; CPLANE1 1.6 ± 0.04 µm; TMEM67 1.4 ± 0.03 µm; CC2D2A 1.2 ± 0.07 µm, (p < 0.005 for all comparisons) (Fig. 4c).
Analysis of primary cilia in control and JS-derived iPSCs at day 8 of differentiation. (a–a’’’’) Representative immunofluorescence images showing cilia labelled with anti-AcTUBULIN (green) and anti-PERICENTRIN (red) with Hoechst nuclear counterstain (blue) in each line. Scale bar: 20 µm. Lower magnification views shown in Online Resource Fig. 12. (b) Quantification of the mean proportion of ciliated cells in control and JS-iPSCs. (c) Analysis of ciliary length in control and JS-iPSCs. Results are presented as mean ± standard error of three independent experiments (n = 10 per sample). *p < 0.05; **p < 0.005
Discussion
Joubert syndrome (JS) is characterized by a unique cerebellar and brainstem malformation, the MTS, which is linked to a multiplicity of genetic defects. No therapies or modifying treatments are currently available for this disorder (Rosati et al. 2018). All the > 40 JS-linked genes encode proteins of the primary cilium, a subcellular organelle widely present in embryonic and adult cells. To date, however, it remains unclear how disruption of distinct proteins with diverse roles in the structure or functioning of the cilium may eventually result in such a specific and peculiar defect as the MTS. In this light, iPSCs represent a simple and efficient model to study abnormal neuronal development in ciliopathies such as JS. To date, a few studies have reported the generation of iPSCs carrying mutations in JS-related genes, including AHI1 (Rosati et al. 2018; Altieri et al. 2019; Serpieri et al. 2023), CPLANE1 (Ali et al. 2020), CC2D2A (Ali et al. 2021), CEP290 (Shimada et al. 2017; May-Simera et al. 2018), RPGRIP1L (Wang et al. 2019), and TMEM67 (Barabino et al. 2020); however, these studies did not systematically analyze differentiation markers or compare ciliary phenotypes. The present study employed iPSCs derived from patients carrying pathogenic variants in four of the five common genes causative of JS, which were differentiated towards mid-hindbrain precursors and cerebellar neurons in order to address two main research questions: (1) do JS-derived iPSCs show aberrant differentiation along the cerebellar lineage? (2) Are there any patent differences among cells mutated in distinct genes?
Interestingly, the four iPSC lines revealed shared ciliary and differentiation alterations in vitro. While differentiated CT-iPSCs correctly expressed markers for mid-hindbrain and cerebellar granule cell precursors along the in vitro time-course, JS mutant cells showed a marked alteration in their differentiation capacity. A progressive cell loss was observed in three of the JS-iPSC lines (AHI1, TMEM67, CC2D2A) during the last phase of differentiation treatment (post D24), and all JS-iPSC lines showed a reduced formation of β3-TUBULIN-positive neurons, suggesting a lower differentiation capacity. This in vitro response was reminiscent of observations made for iPSC lines from other ciliopathies (such as Meckel syndrome and Bardet-Biedl syndrome), reporting increased cell loss in culture and differentiation delays (Barabino et al. 2020). The present observation warrants further studies to assess possible defects linked to apoptosis and autophagy pathways, two key mechanisms already linked to ciliogenesis (Attanasio et al. 2007; Tang et al. 2013; Morleo and Franco 2019) and neural differentiation (Mazarakis et al. 1997; Rosiak et al. 2016) and therefore potentially affected in JS-derived cultures. The mRNA expression levels for two genes finely regulated during neural development, LMX1B, and PAX6, were severely misregulated in JS cells. LMX1B is a master gene for the development and maintenance of the isthmic organizer (Mishima et al. 2009; Srivastava et al. 2017), which is thought to secrete planar signals regulating the development of the mid/hindbrain (Shimada et al. 2017). LMX1B expression in all JS-derived cultures was observed to be lower than controls, suggesting a failure of the mutant lines to adequately activate a midbrain/cerebellar differentiation program. In contrast, transcriptional levels of the neuroprogenitor marker PAX6 in JS-iPSCs failed to show the typical differentiation-related reduction seen in CT-iPSCs (Erceg et al. 2010; Ha et al. 2012). The JS-iPSC lines analyzed here also demonstrated an altered expression of the mid-hindbrain markers ATOH1 and NEUROD1, which are both required for the formation of the granule cell population. ATOH1 regulates early neuronal differentiation leading to the formation of CGC precursors, and its deletion results in the elimination of the granule cell population (Brown and Witman 2014; Lange et al. 2021). NEUROD1 acts subsequently to promote the differentiation of CGC precursors towards the mature cerebellar granule cell lineage (Pan et al. 2009); it is also involved in the maintenance of this differentiated population, as evidenced by the increased cell death detected in NeuroD1 knockout models (Warburton-Pitt et al. 2012). In light of this, it is tempting to speculate that the observed reduction of NEUROD1 expression may somehow relate to the massive cell loss experienced by three mutant cell lines. Finally, JS-iPSCs showed reduced ZIC2 expression during the later phase of the differentiation protocol, in particular by D24, as also observed at the protein level. ZIC2 is reportedly expressed in pluripotent cells (Brown and Brown 2009), while in differentiated lineages it is a marker of granule cell precursors, acting together with ZIC1 to control cerebellar development (Schouteden et al. 2015; Bashford and Subramanian 2019). In support of these observations, the transcriptomics analysis underlined the reduced mature marker expression in JS lines. Compared to controls, JS lines showed a profile more typical of early phases of differentiation. Taken together, this altered expression profile in JS-iPSC cultures appears consistent with an inadequate activation of genes controlling neuronal/cerebellar differentiation, a situation not dissimilar to that seen in iPSCs from patients with Bardet-Biedl syndrome mutated in BBS10, which responded to neural crest differentiation with a lower neural crest genes expression and incomplete differentiation (Barrell et al. 2019). Of note, alterations detected at the mRNA level for neuronal markers were largely echoed by immunostaining observations; it will be useful to further confirm these quantitatively, for instance by performing a Western blot analysis, and extend the analysis to glial markers to assess whether additional lineages may contribute to the phenotype. Furthermore, experiments on larger samples of iPSCs from JS patients will address whether distinct JS mutations in the same gene will result in a similar differentiation impairment.
iPSCs possess a primary cilium which becomes more evident in the course of neuronal differentiation (Rosenbaum and Witman 2002; Wood et al. 2013) and are thus a fitting model to study cellular defects in ciliopathies caused by different genetic mutations. Among the four JS patient lines generated here, a significant reduction in the proportion of mutant iPSCs able to form the primary cilium was consistently observed compared to control cells. Ciliary measurements further indicated a reduced cilium length that was significant in three out of the four JS-iPSC lines (AHI1, TMEM67, CC2D2A), while results for the CPLANE1 line were similar to those of control cells. These results align with previous reports on fibroblasts from patients mutated in either CSPP1, ARL13B, and CEP41, describing cellular defects in cilium morphology, number or defects in the ciliary axoneme (Tuz et al. 2014; Ramsbottom et al. 2018). Barabino et al. also reported abnormal ciliary phenotype in a TMEM67-mutant iPSC line, although in this case more numerous, but shorter and thinner cilia were observed, suggesting that different mutations in the same JS gene can lead to different outcomes. Moreover, iPSCs from RPGRIP1L-mutated JS patients showed a reduction in the number of ciliated neurons formed upon differentiation (Altieri et al. 2019), while JS iPSCs mutated in CEP290, when differentiated towards retinal epithelium lineages, showed smaller cilia associated with incomplete lineage maturation (Rosati et al. 2018). Further in vitro observations made in other ciliopathy models, such as another iPSC line from a CEP290-mutated patient with Leber’s congenital amaurosis and differentiated towards retinal pigment epithelium, also showed alterations in primary cilium formation and incidence (Parfitt et al. 2016). Of note, in our experimental setting, the CPLANE1 line was the only line to survive over the differentiation time-course and showed a normal ciliary length with less evident differences from controls regarding the proportion of ciliated cells. This observation contrasts with previous observations made on CPLANE1 mutant fibroblasts, which showed fewer and shorter primary cilia and diminished response to a SHH agonist compared to controls (Asadollahi et al. 2018). This would call for further analysis in a larger number of CPLANE1 iPSC lines carrying distinct mutations, as we cannot exclude an hypomorphic effect of the CPLANE1 missense variant p. S1290P, potentially resulting in a less detrimental effect on the protein function, as already reported for other variants in distinct JS-related genes (Leitch et al. 2008). Additional investigations including a longitudinal cilium analysis over the cerebellar differentiation time-course, a spatial characterization of the ciliary position for the protein products of the 4 mutated genes (AHI1, CPLANE1, TMEM67, and CC2D2A), and a precise analysis of how different JS mutations impact on the functioning of the corresponding protein at the ciliary level will help to further understand the pathogenic mechanisms underlying the neurodevelopmental defects observed in patients’ cell lines. More generally, since some mutations in the JS-associated genes analyzed here have been associated with other ciliopathy phenotypes such as nephronophthisis and Meckel syndrome (Van De Weghe et al. 2022), it will be interesting to compare iPSCs lines from patients with different ciliopathy phenotypes harbouring mutations within the same gene to explore how these result in defects affecting different organs.
References
Abdelhamed ZA, Natarajan S, Wheway G, Inglehearn CF, Toomes C, Johnson CA, Jagger DJ (2015) The Meckel-Gruber syndrome protein TMEM67 controls basal body positioning and epithelial branching morphogenesis in mice via the non-canonical Wnt pathway. Dis Model Mech 8:527–541. https://doi.org/10.1242/dmm.019083
Ali E, Ferraro RM, Lanzi G, Masneri S, Piovani G, Mazzoldi EL, Serpieri V, Valente EM, Giordano L, Giliani SC (2020) Generation of induced pluripotent stem cell (iPSC) lines from a Joubert syndrome patient with compound heterozygous mutations in C5orf42 gene. Stem Cell Res 49:102007. https://doi.org/10.1016/j.scr.2020.102007
Ali E, Ferraro RM, Guglielmi A, Lanzi G, Masneri S, Piovani G, Mazzoldi EL, Pollara L, Valente EM, Accorsi P, Giordano L, Giliani SC (2021) Establishment of three Joubert syndrome-derived induced pluripotent stem cell (iPSC) lines harbouring compound heterozygous mutations in CC2D2A gene. Stem Cell Research 54:102430. https://doi.org/10.1016/j.scr.2021.102430
Altieri F, D’Anzi A, Martello F, Tardivo S, Spasari I, Ferrari D, Bernardini L, Lamorte G, Mazzoccoli G, Valente EM, Vescovi AL, Rosati J (2019) Production and characterization of human induced pluripotent stem cells (iPSC) CSSi007-A (4383) from Joubert Syndrome. Stem Cell Res 38:101480. https://doi.org/10.1016/j.scr.2019.101480
Asadollahi R, Strauss JE, Zenker M, Beuing O, Edvardson S, Elpeleg O, Strom TM, Joset P, Niedrist D, Otte C, Oneda B, Boonsawat P, Azzarello-Burri S, Bartholdi D, Papik M, Zweier M, Haas C, Ekici AB, Baumer A, Boltshauser E, Steindl K, Nothnagel M, Schinzel A, Stoeckli ET, Rauch A (2018) Clinical and experimental evidence suggest a link between KIF7 and C5orf42-related ciliopathies through Sonic Hedgehog signaling. Eur J Hum Genet 26:197–209. https://doi.org/10.1038/s41431-017-0019-9
Attanasio M, Uhlenhaut NH, Sousa VH, O’Toole JF, Otto E, Anlag K, Klugmann C, Treier A-C, Helou J, Sayer JA, Seelow D, Nürnberg G, Becker C, Chudley AE, Nürnberg P, Hildebrandt F, Treier M (2007) Loss of GLIS2 causes nephronophthisis in humans and mice by increased apoptosis and fibrosis. Nat Genet 39:1018–1024. https://doi.org/10.1038/ng2072
Bachmann-Gagescu R, Dempsey JC, Phelps IG, O’Roak BJ, Knutzen DM, Rue TC, Ishak GE, Isabella CR, Gorden N, Adkins J, Boyle EA, de Lacy N, O’Day D, Alswaid A, Ramadevi A R, Lingappa L, Lourenço C, Martorell L, Garcia-Cazorla À, Ozyürek H, Haliloğlu G, Tuysuz B, Topçu M, University of Washington Center for Mendelian Genomics, Chance P, Parisi MA, Glass IA, Shendure J, Doherty D (2015) Joubert syndrome: a model for untangling recessive disorders with extreme genetic heterogeneity. J Med Genet 52:514–522. https://doi.org/10.1136/jmedgenet-2015-103087
Barabino A, Flamier A, Hanna R, Héon E, Freedman BS, Bernier G (2020) Deregulation of neuro-developmental genes and primary cilium cytoskeleton anomalies in iPSC retinal sheets from human syndromic ciliopathies. Stem Cell Reports 14:357–373. https://doi.org/10.1016/j.stemcr.2020.02.005
Barrell WB, Griffin JN, Harvey J-L, HipSci Consortium, Danovi D, Beales P, Grigoriadis AE, Liu KJ (2019) Induction of neural crest stem cells from Bardet-Biedl syndrome patient derived hiPSCs. Front Mol Neurosci 12:139. https://doi.org/10.3389/fnmol.2019.00139
Bashford AL, Subramanian V (2019) Mice with a conditional deletion of Talpid3 (KIAA0586) - a model for Joubert syndrome. J Pathol 248:396–408. https://doi.org/10.1002/path.5271
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Brown L, Brown S (2009) Zic2 is expressed in pluripotent cells in the blastocyst and adult brain expression overlaps with makers of neurogenesis. Gene Expr Patterns 9:43–49. https://doi.org/10.1016/j.gep.2008.08.002
Brown JM, Witman GB (2014) Cilia and diseases. Bioscience 64:1126–1137. https://doi.org/10.1093/biosci/biu174
Compagnucci C, Barresi S, Petrini S, Billuart P, Piccini G, Chiurazzi P, Alfieri P, Bertini E, Zanni G (2016) Rho kinase inhibition is essential during in vitro neurogenesis and promotes phenotypic rescue of human induced pluripotent stem cell-derived neurons with Oligophrenin-1 loss of function. Stem Cells Transl Med 5:860–869. https://doi.org/10.5966/sctm.2015-0303
Consalez GG, Goldowitz D, Casoni F, Hawkes R (2021) Origins, development, and compartmentation of the granule cells of the cerebellum. Front Neural Circuits 14:611841. https://doi.org/10.3389/fncir.2020.611841
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21. https://doi.org/10.1093/bioinformatics/bts635
Erceg S, Ronaghi M, Zipancic I, Lainez S, Roselló MG, Xiong C, Moreno-Manzano V, Rodríguez-Jiménez FJ, Planells R, Alvarez-Dolado M, Bhattacharya SS, Stojkovic M (2010) Efficient differentiation of human embryonic stem cells into functional cerebellar-like cells. Stem Cells Dev 19:1745–1756. https://doi.org/10.1089/scd.2009.0498
Erceg S, Lukovic D, Moreno-Manzano V, Stojkovic M, Bhattacharya SS (2012) Derivation of cerebellar neurons from human pluripotent stem cells. Curr Protoc Stem Cell Biol 20:1H.5.1-1H.5.10. https://doi.org/10.1002/9780470151808.sc01h05s20
Garcia-Gonzalo FR, Corbit KC, Sirerol-Piquer MS, Ramaswami G, Otto EA, Noriega TR, Seol AD, Robinson JF, Bennett CL, Josifova DJ, García-Verdugo JM, Katsanis N, Hildebrandt F, Reiter JF (2011) A transition zone complex regulates mammalian ciliogenesis and ciliary membrane composition. Nat Genet 43:776–784. https://doi.org/10.1038/ng.891
Gilthorpe JD, Papantoniou EK, Chédotal A, Lumsden A, Wingate RJT (2002) The migration of cerebellar rhombic lip derivatives. Development 129:4719–4728. https://doi.org/10.1242/dev.129.20.4719
Gu Z, Eils R, Schlesner M (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32:2847–2849. https://doi.org/10.1093/bioinformatics/btw313
Ha TJ, Swanson DJ, Kirova R, Yeung J, Choi K, Tong Y, Chesler EJ, Goldowitz D (2012) Genome-wide microarray comparison reveals downstream genes of Pax6 in the developing mouse cerebellum. Eur J Neurosci 36:2888–2898. https://doi.org/10.1111/j.1460-9568.2012.08221.x
Hansen KB, Wollmuth LP, Bowie D, Furukawa H, Menniti FS, Sobolevsky AI, Swanson GT, Swanger SA, Greger IH, Nakagawa T, McBain CJ, Jayaraman V, Low CM, Dell’Acqua ML, Diamond JS, Camp CR, Perszyk RE, Yuan H, Traynelis SF (2021) Structure function and pharmacology of glutamate receptor ion channels. Pharmacol Rev 73(4):1469–1658. https://doi.org/10.1124/pharmrev.120.000131
Harashima S, Wang Y, Horiuchi T, Seino Y, Inagaki N (2011) Purkinje cell protein 4 positively regulates neurite outgrowth and neurotransmitter release. J Neurosci Res 89(10):1519–1530. https://doi.org/10.1002/jnr.22688
Lancaster MA, Schroth J, Gleeson JG (2011) Subcellular spatial regulation of canonical Wnt signalling at the primary cilium. Nat Cell Biol 13:700–707. https://doi.org/10.1038/ncb2259
Lange KI, Tsiropoulou S, Kucharska K, Blacque OE (2021) Interpreting the pathogenicity of Joubert syndrome missense variants in Caenorhabditis elegans. Dis Model Mech 14:dmm046631. https://doi.org/10.1242/dmm.046631
Lee S-H, Nam T-S, Li W, Kim JH, Yoon W, Choi Y-D, Kim K-H, Cai H, Kim MJ, Kim C, Choy HE, Kim N, Chay KO, Kim M-K, Choi S-Y (2017) Functional validation of novel MKS3/TMEM67 mutations in COACH syndrome. Sci Rep 7:10222. https://doi.org/10.1038/s41598-017-10652-z
Leitch CC, Zaghloul NA, Davis EE, Stoetzel C, Diaz-Font A, Rix S, Alfadhel M, Al-Fadhel M, Lewis RA, Eyaid W, Banin E, Dollfus H, Beales PL, Badano JL, Katsanis N (2008) Hypomorphic mutations in syndromic encephalocele genes are associated with Bardet-Biedl syndrome. Nat Genet 40:443–448. https://doi.org/10.1038/ng.97
Li C, Jensen VL, Park K, Kennedy J, Garcia-Gonzalo FR, Romani M, De Mori R, Bruel A-L, Gaillard D, Doray B, Lopez E, Rivière J-B, Faivre L, Thauvin-Robinet C, Reiter JF, Blacque OE, Valente EM, Leroux MR (2016) MKS5 and CEP290 dependent assembly pathway of the ciliary transition zone. PLoS Biol 14:e1002416. https://doi.org/10.1371/journal.pbio.1002416
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930. https://doi.org/10.1093/bioinformatics/btt656
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA- seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lowenstein ED, Rusanova A, Stelzer J, Hernaiz-Llorens M, Schroer AE, Epifanova E, Bladt F, Isik EG, Buchert S, Jia S, Tarabykin V, Hernandez-Miranda LR (2021) Olig3 regulates early cerebellar development. Elife 10:e64684. https://doi.org/10.7554/eLife.64684
Luo Z, Gao X, Lin C, Smith ER, Marshall SA, Swanson SK, Florens L, Washburn MP, Shilatifard A (2015) Zic2 is an enhancer-binding factor required for embryonic stem cell specification. Mol Cell 57:685–694. https://doi.org/10.1016/j.molcel.2015.01.007
May-Simera HL, Wan Q, Jha BS, Hartford J, Khristov V, Dejene R, Chang J, Patnaik S, Lu Q, Banerjee P, Silver J, Insinna-Kettenhofen C, Patel D, Lotfi M, Malicdan M, Hotaling N, Maminishkis A, Sridharan R, Brooks B, Miyagishima K, Gunay-Aygun M, Pal R, Westlake C, Miller S, Sharma R, Bharti K (2018) Primary cilium-mediated retinal pigment epithelium maturation is disrupted in ciliopathy patient cells. Cell Rep 22:189–205. https://doi.org/10.1016/j.celrep.2017.12.038
Mazarakis ND, Edwards AD, Mehmet H (1997) Apoptosis in neural development and disease. Arch Dis Child Fetal Neonatal Ed 77:F165–170. https://doi.org/10.1136/fn.77.3.f165
Mishima Y, Lindgren AG, Chizhikov VV, Johnson RL, Millen KJ (2009) Overlapping function of Lmx1a and Lmx1b in anterior hindbrain roof plate formation and cerebellar growth. J Neurosci 29:11377–11384. https://doi.org/10.1523/JNEUROSCI.0969-09.2009
Mitchison HM, Valente EM (2017) Motile and non-motile cilia in human pathology: from function to phenotypes. J Pathol 241:294–309. https://doi.org/10.1002/path.4843
Morleo M, Franco B (2019) The autophagy-cilia axis: an intricate relationship. Cells 8:E905. https://doi.org/10.3390/cells8080905
Ojeda Naharros I, Gesemann M, Mateos JM, Barmettler G, Forbes A, Ziegler U, Neuhauss SCF, Bachmann-Gagescu R (2017) Loss-of-function of the ciliopathy protein Cc2d2a disorganizes the vesicle fusion machinery at the periciliary membrane and indirectly affects Rab8-trafficking in zebrafish photoreceptors. PLoS Genet 13:e1007150. https://doi.org/10.1371/journal.pgen.1007150
Pan N, Jahan I, Lee JE, Fritzsch B (2009) Defects in the cerebella of conditional Neurod1 null mice correlate with effective Tg(Atoh1-cre) recombination and granule cell requirements for Neurod1 for differentiation. Cell Tissue Res 337:407–428. https://doi.org/10.1007/s00441-009-0826-6
Parfitt DA, Lane A, Ramsden CM, Carr A-JF, Munro PM, Jovanovic K, Schwarz N, Kanuga N, Muthiah MN, Hull S, Gallo J-M, da Cruz L, Moore AT, Hardcastle AJ, Coffey PJ, Cheetham ME (2016) Identification and correction of mechanisms underlying inherited blindness in human iPSC-derived optic cups. Cell Stem Cell 18:769–781. https://doi.org/10.1016/j.stem.2016.03.021
Ramsbottom SA, Molinari E, Srivastava S, Silberman F, Henry C, Alkanderi S, Devlin LA, White K, Steel DH, Saunier S, Miles CG, Sayer JA (2018) Targeted exon skipping of a CEP290 mutation rescues Joubert syndrome phenotypes in vitro and in a murine model. Proc Natl Acad Sci U S A 115:12489–12494. https://doi.org/10.1073/pnas.1809432115
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Romani M, Micalizzi A, Valente EM (2013) Joubert syndrome: congenital cerebellar ataxia with the molar tooth. Lancet Neurol 12:894–905. https://doi.org/10.1016/S1474-4422(13)70136-4
Rosati J, Altieri F, Tardivo S, Turco EM, Goldoni M, Spasari I, Ferrari D, Bernardini L, Lamorte G, Valente EM, Vescovi AL (2018) Production and characterization of human induced pluripotent stem cells (iPSCs) from Joubert syndrome: CSSi001-A (2850). Stem Cell Res 27:74–77. https://doi.org/10.1016/j.scr.2018.01.012
Rosenbaum JL, Witman GB (2002) Intraflagellar transport. Nat Rev Mol Cell Biol 3:813–825. https://doi.org/10.1038/nrm952
Rosiak K, Smolarz M, Stec WJ, Peciak J, Grzela D, Winiecka-Klimek M, Stoczynska-Fidelus E, Krynska B, Piaskowski S, Rieske P (2016) IDH1R132H in neural stem cells: differentiation impaired by increased apoptosis. PLoS ONE 11:e0154726. https://doi.org/10.1371/journal.pone.0154726
Rubio ME, Soto F (2001) Distinct Localization of P2X receptors at excitatory postsynaptic specializations. J Neurosci 21(2):641–653. https://doi.org/10.1523/JNEUROSCI.21-02-00641.2001
Schouteden C, Serwas D, Palfy M, Dammermann A (2015) The ciliary transition zone functions in cell adhesion but is dispensable for axoneme assembly in C. elegans. J Cell Biol 210:35–44. https://doi.org/10.1083/jcb.201501013
Serpieri V, Orsi A, Mazzotta C, Cavan S, Rossi E, Scelsa B, Valente EM (2023) Generation of an iPSC line from skin fibroblasts of a patient with Joubert syndrome carrying the homozygous loss of function variant c.787dupC in the AHI1 gene. Stem Cell Res 66:103002. https://doi.org/10.1016/j.scr.2022.103002
Shimada H, Lu Q, Insinna-Kettenhofen C, Nagashima K, English MA, Semler EM, Mahgerefteh J, Cideciyan AV, Li T, Brooks BP, Gunay-Aygun M, Jacobson SG, Cogliati T, Westlake CJ, Swaroop A (2017) In vitro modeling using ciliopathy-patient-derived cells reveals distinct cilia dysfunctions caused by CEP290 mutations. Cell Rep 20:384–396. https://doi.org/10.1016/j.celrep.2017.06.045
Srivastava S, Ramsbottom SA, Molinari E, Alkanderi S, Filby A, White K, Henry C, Saunier S, Miles CG, Sayer JA (2017) A human patient-derived cellular model of Joubert syndrome reveals ciliary defects which can be rescued with targeted therapies. Hum Mol Genet 26:4657–4667. https://doi.org/10.1093/hmg/ddx347
Tang Z, Lin MG, Stowe TR, Chen S, Zhu M, Stearns T, Franco B, Zhong Q (2013) Autophagy promotes primary ciliogenesis by removing OFD1 from centriolar satellites. Nature 502:254–257. https://doi.org/10.1038/nature12606
Tuz K, Bachmann-Gagescu R, O’Day DR, Hua K, Isabella CR, Phelps IG, Stolarski AE, O’Roak BJ, Dempsey JC, Lourenco C, Alswaid A, Bönnemann CG, Medne L, Nampoothiri S, Stark Z, Leventer RJ, Topçu M, Cansu A, Jagadeesh S, Done S, Ishak GE, Glass IA, Shendure J, Neuhauss SCF, Haldeman-Englert CR, Doherty D, Ferland RJ (2014) Mutations in CSPP1 cause primary cilia abnormalities and Joubert syndrome with or without Jeune asphyxiating thoracic dystrophy. Am J Hum Genet 94:62–72. https://doi.org/10.1016/j.ajhg.2013.11.019
Van De Weghe JC, Gomez A, Doherty D (2022) The Joubert-Meckel-nephronophthisis spectrum of ciliopathies. Annu Rev Genomics Hum Genet 23:301–329. https://doi.org/10.1146/annurev-genom-121321-093528
Wang L, De Solis AJ, Goffer Y, Birkenbach KE, Engle SE, Tanis R, Levenson JM, Li X, Rausch R, Purohit M, Lee J-Y, Tan J, De Rosa MC, Doege CA, Aaron HL, Martins GJ, Brüning JC, Egli D, Costa R, Berbari N, Leibel RL, Stratigopoulos G (2019) Ciliary gene RPGRIP1L is required for hypothalamic arcuate neuron development. JCI Insight 4:123337. https://doi.org/10.1172/jci.insight.123337
Warburton-Pitt SRF, Jauregui AR, Li C, Wang J, Leroux MR, Barr MM (2012) Ciliogenesis in Caenorhabditis elegans requires genetic interactions between ciliary middle segment localized NPHP-2 (inversin) and transition zone-associated proteins. J Cell Sci 125:2592–2603. https://doi.org/10.1242/jcs.095539
Wood CR, Huang K, Diener DR, Rosenbaum JL (2013) The cilium secretes bioactive ectosomes. Curr Biol 23:906–911. https://doi.org/10.1016/j.cub.2013.04.019
Acknowledgements
The authors are grateful to Dr. Rosalba Monica Ferraro for help in developing iPSC lines, to Tiziana Stellato and Lorenzo Bianca for assistance with qPCR and immunofluorescence, and to Dr. Amanda Oldani and Dr. Patrizia Vaghi (PASS-Bio Med, core facility Centro Grandi Strumenti, University of Pavia) for their expert input in microscopy and image analysis.
Funding
Open access funding provided by Università degli Studi di Pavia within the CRUI-CARE Agreement. This research was partly supported by Telethon Foundation [grant number GGP20070 to EMV], the Italian Ministry of Health [ERA-NET Neuron NDCil, grant number EUR002 to EMV] and the Italian Ministry for University and Research [grant number PRIN 2022JT5PWC_002 to VS].
Author information
Authors and Affiliations
Contributions
Conceptualization: RDM and EMV; methodology: RDM, ST, SCG, EA, LP; formal analysis: RDM, ST, and LP; patient recruitment and collection of biological material: VL, LG, RF, and BG; investigation: RDM and ST; data curation: RDM and ST; data processing: RDM, SD, MM, LP, MCM, VS; original draft preparation: RDM, LP, and VS; manuscript review and editing: all co-authors; project supervision and funding acquisition: EMV. All authors have read and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Ethics Committee of the University of Pavia (protocol code 20180077857, date 12–09-2018).
Consent to participate
Informed consent was obtained from all subjects involved in the study. Data available from the authors upon reasonable request.
Conflict of interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
De Mori, R., Tardivo, S., Pollara, L. et al. Joubert syndrome-derived induced pluripotent stem cells show altered neuronal differentiation in vitro. Cell Tissue Res 396, 255–267 (2024). https://doi.org/10.1007/s00441-024-03876-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-024-03876-9