Abstract
Arbuscular mycorrhizal fungi (AMF) represent an important group of root symbionts, given the key role they play in the enhancement of plant nutrition, health, and product quality. The services provided by AMF often are facilitated by large and diverse beneficial bacterial communities, closely associated with spores, sporocarps, and extraradical mycelium, showing different functional activities, such as N2 fixation, nutrient mobilization, and plant hormone, antibiotic, and siderophore production and also mycorrhizal establishment promotion, leading to the enhancement of host plant performance. The potential functional complementarity of AMF and associated microbiota poses a key question as to whether members of AMF-associated bacterial communities can colonize the root system after establishment of mycorrhizas, thereby becoming endophytic. Root endophytic bacterial communities are currently studied for the benefits provided to host plants in the form of growth promotion, stress reduction, inhibition of plant pathogens, and plant hormone release. Their quantitative and qualitative composition is influenced by many factors, such as geographical location, soil type, host genotype, and cultivation practices. Recent data suggest that an additional factor affecting bacterial endophyte recruitment could be AMF and their associated bacteria, even though the mechanisms allowing members of AMF-associated bacterial communities to actually establish in the root system, becoming endophytic, remain to be determined. Given the diverse plant growth–promoting properties shown by AMF-associated bacteria, further studies are needed to understand whether AMF may represent suitable tools to introduce beneficial root endophytes in sustainable and organic agriculture where the functioning of such multipartite association may be crucial for crop production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, the rising demand for the production of environmentally safe and high-quality food has caused a major shift in agricultural management, which is growing increasingly sustainable by making use of practices able to maintain and enhance soil fertility and health (FAO 2011). In this context, soil microorganisms play a key role in the modulation of soil biochemical, biological, and nutritional processes (Azcón-Aguilar and Barea 2015). Such microbiota thrives in a privileged niche at the soil–root interface (rhizosphere) that is a rich source of nourishment, represented by sugars, amino acids, and organic acids in the form of root exudates (Philippot et al. 2013). The complex microbial communities establishing in the rhizosphere have profound effects on plant growth, nutrition, and health (Compant et al. 2010; Hayat et al. 2010). Members of the rhizospheric microbiota may establish an intimate relationship with their host plants, colonizing roots and also aboveground plant compartments, becoming endophytes, i.e., microorganisms which can be isolated from, or detected within, surface-sterilized plant organs and do not cause visible harm to the host organism (Hardoim et al. 2008). Bacterial endophytes of plant roots may reach a density of 104–108 bacterial cells per gram of root tissue and may have important roles in plant growth promotion, fitness, and protection against pathogens. In exchange, the plant endosphere provides the endophytic microbiota with a more uniform, protected, and nutrient-rich environment than the nearby soil (Hardoim et al. 2015; Liu et al. 2017b). The use of green fluorescent protein labeling, image analysis, and fluorescence in situ hybridization–confocal laser scanning microscopy has allowed the localization of bacterial endophytes in intercellular spaces, at the base of lateral roots and root tips, in xylem vessels, and inside root hairs (Compant et al. 2010; Gaiero et al. 2013; Liu et al. 2017a).
Arbuscular mycorrhizal fungi (AMF) represent an important group of root endophytes in view of their key role in the enhancement of plant nutrition, health, and product quality. AMF (phylum Glomeromycota) are beneficial soil fungi, establishing mutualistic symbioses with about 80% of land plants including the major food crops, such as cereals, pulses, vegetables, and fruit trees and industrial crops like cotton, sunflower, and oil palm (Smith and Read 2008). They are obligate biotrophic organisms which obtain carbon from the host plant, providing in exchange soil mineral nutrients (such as P, N, S, K, Ca, Cu, and Zn), absorbed and translocated by the extraradical mycelium (ERM) extending from colonized roots into the soil. Therefore, the ERM represents an efficient auxiliary absorbing system because of its high surface-to-volume ratio, hyphal P absorption beyond the P depletion zone around roots, and the occurrence of many nutrient transporters in its hyphae (Smith and Read 2008; Pepe et al. 2017; Kameoka et al. 2019). Moreover, AMF improve plant performance and health by increasing plant tolerance to biotic and abiotic stresses (Sikes et al. 2009; Bitterlich et al. 2018) and induce changes in plant secondary metabolism leading to enhanced biosynthesis of health-promoting phytochemicals (Avio et al. 2018; Agnolucci et al. 2020). Overall, AMF provide multifunctional ecosystem services and are utilized as biofertilizers, biostimulants, and bioenhancers in agriculture (Gianinazzi et al. 2010; Rouphael et al. 2015).
Arbuscular mycorrhizal fungi live closely associated with large and diverse bacterial communities which may colonize spores, sporocarps, and extraradical hyphae, originating a complex and metabolically active environment called the mycorrhizosphere (Rambelli 1973). Such microbiota show different plant growth–promoting properties (plant hormone, antibiotic, and siderophore production; N2 fixation; P solubilization) and mycorrhiza helper activities (spore germination and mycelial growth promotion, mycorrhizal establishment facilitation) also have been observed, leading to enhancement of host plant performance (Bharadwaj et al. 2008a; Battini et al. 2016; 2017; Sharma et al. 2020). The potential functional complementarity and synergistic activity of AMF and their associated microbiota necessitate studies aimed at understanding the complex network of interactions between them and their host plants (Turrini et al. 2018). This is all the more important when implementing AMF inocula in sustainable and organic agriculture where the functioning of such multipartite associations may be crucial for crop production. Notwithstanding, scanty information is available on the relationship between AMF-associated bacteria and the bacterial microbiota colonizing roots after AMF inoculation. Here, we (i) provide an overview of recent developments regarding the recruitment of root endophytic bacteria, (ii) present data on the diversity and functionality of AMF-associated bacterial communities, and (iii) discuss the possible role of AMF in shaping the structure and composition of endophytic bacterial communities recruited by plant roots.
Endophytic bacteria recruited by plant roots
Characteristics and importance of root endophytic bacteria
Bacterial endophytes are able to recognize plant root exudates, adhere to the root surface, form a biofilm, and then enter roots, colonizing their inner tissues. According to current knowledge, passive penetration can take place at wounds, root cracks, secondary root emergence points, and root tips, while active colonization can involve cell wall–degrading hydrolytic enzymes (Compant et al. 2010). The presence of flagella, pili, lipopolysaccharides, exopolysaccharides, some special membrane proteins, quorum-sensing signals, chemotactic abilities, protein secretion systems, and twitching motility also may have importance in the invasion processes of certain endophytic bacteria, as shown by comparative genome analyses (Sessitsch et al. 2012; Hardoim et al. 2015; Pinski et al. 2019). Some of the required endophytic/symbiotic genes may be coded on plasmids or “symbiosis islands,” suggesting the possibility of horizontal transfer of these functional genes among the members of soil bacterial communities (Finan 2002). Nevertheless, vertical ways of endophyte transmission (seed-borne endophytes) also have been confirmed (Truyens et al. 2015) in which cases the transmitted bacteria were able to colonize the rhizospheres of new plantlets (Kaga et al. 2009; Hameed et al. 2015).
Bacteria colonizing the root endosphere profit from enhanced nutrient availability and environmental homeostasis provided by the plant, while the host plant may receive benefits from the endophytes in the form of direct and indirect growth promotion, stress reduction, or inhibition of plant pathogens (Gaiero et al. 2013; Hardoim et al. 2008; 2015; Compant et al. 2010; Pinski et al. 2019). As to nutrient mobilization, many endophytic bacteria were reported to possess N2-fixing, nitrifying, denitrifying, and P-solubilizing abilities or to be able to produce siderophores (Sessitsch et al. 2012; Hameed et al. 2015). Bacterial root endophytes influence plant hormone levels directly or indirectly, affecting growth, stress, and immune responses of host plants. Many endophytes may be able to release indoleacetic acids (IAA), gibberellins, cytokinins, ethylene, abscisic acid, jasmonates, and volatile compounds, while some of them also act as a sink for 1-aminocyclopropane-1-carboxylate (ACC), a precursor of ethylene production, because of a cytoplasmic ACC deaminase enzyme. The features mentioned suggest that bacterial endophytes may have a fundamental role in “fine-tuning” the hormonal balance of the host plant (Forchetti et al. 2007; Glick 2014; Hardoim et al. 2015). Furthermore, a key feature of endophytic strains is detoxification of reactive oxygen species, reactive nitrogen species, glutathione synthases, and glutathione-S-transferases, ameliorating different effectors of plant stress responses (Sessitsch et al. 2012). In the unique ecological niche where they thrive, some root endophytic bacteria may produce bioactive secondary metabolites, such as antibiotic and antiviral compounds (Strobel et al. 2004; Ryan et al. 2008; Ek-Ramos et al. 2019), while others have shown potential for enhancement of phytoremediation procedures because of their ability to decompose contaminants (Ryan et al. 2008; Mitter et al. 2019). Some authors have reported that the root microbiota may contribute to improved food quality through biofortification or production of health-promoting metabolites (Rehman et al. 2018; Ku et al. 2019) and potentially affect the quality of processed food products. For example, Minervini et al. (2015) demonstrated that endophytic lactic acid bacteria occurred not only in roots and various organs of durum wheat plants (Triticum turgidum ssp. durum) but also in the flour, possibly inducing changes in microbial community structure and properties of sourdough and derived products. Accordingly, the isolation and selection of functional root endophytic bacterial strains may be of particular interest for agriculture, industrial biotechnology, and medicine.
Recruitment of bacterial root endophytes by host plants
The quantitative and qualitative composition of root endophytic bacterial communities is influenced by many factors, mainly by geographical location, soil source, host genotype, and cultivation practice (Edwards et al. 2015). In Table 1, data from recent metagenomic studies of endophytic bacteria are reported. During notable work of defining the core root microbiome of the non-mycorrhizal species Arabidopsis thaliana L., based on 16S rDNA high-throughput sequencing data, two research groups revealed that the composition of root endophytic bacterial communities may be influenced by host genotype (Bulgarelli et al. 2012; Lundberg et al. 2012). Other studies demonstrated that the host plant may modulate the occurrence of root microbiota recruited from the nearby soil environment. Manter et al. (2010), utilizing a pyrosequencing approach, investigated the root endophytic communities of 20 potato cultivars and clones, revealing significant differences among the taxonomic profiles within those different plant genotypes. In that study, the identified bacterial operational taxonomic units (OTUs) affiliated with 238 genera and 15 phyla, demonstrating the remarkable diversity and variability of root endophytes. Plant genotype was found to shape the community composition of bacteria associated with the roots of 10 different rice cultivars (Hardoim et al. 2011), while differences in the bacterial root microbiota in the non-mycotrophic Brassicaceae family were found to be largely quantitative (Schlaeppi et al. 2014). Using the Illumina MiSeq platform, Marasco et al. (2018) found that grapevine (Vitis vinifera L.) rootstock genotypes influenced the taxonomic composition of their endophytic bacterial communities, although plant growth–promoting traits were not significantly different among the cultivars, showing a homeostasis of the plant/bacterial endophyte relationship. Culture-independent techniques, PCR-denaturing gradient gel electrophoresis (DGGE) and Illumina MiSeq sequencing of the 16S rDNA of root endophytic bacterial communities confirmed the selectivity of genotypes in durum wheat, as different cultivars hosted significantly different bacterial communities in their root tissues (Agnolucci et al. 2019b). However, Singer et al. (2019) found conserved community structures across different genotypes of Panicum virgatum and Panicum hallii.
Not only the genotype, but also the phenological stage of the host plant may cause variability in the composition of root endophytic bacterial communities (Van Overbeek and Van Elsas 2008). Such an effect was clearly demonstrated in field-grown durum wheat, where bacterial taxa affiliated with Firmicutes showed fluctuating relative abundance in roots and other plant organs during the growing season (Minervini et al. 2015). Likewise, endophytic bacterial community compositions changed significantly across the growing stages in roots of sweet potato (Marques et al. 2015) and of non-mycorrhizal sugar beet (Shi et al. 2014).
Additional studies have revealed significant shifts in the composition and function of root bacterial microbiota as an effect of environmental variability. Soil type and geographic location contribute variability to potential soilborne colonizers causing significant differences in the quantitative and qualitative composition of root endophytic bacterial communities (Conn and Franco 2004; Lundberg et al. 2012; Schlaeppi et al. 2014; Edwards et al. 2015; Hameed et al. 2015). Other abiotic factors, such as stress (Naylor et al. 2017), flooding (Ferrando and Scavino 2015), suboptimal mineral nutrition (Hameed et al. 2015), seasons (Mocali et al. 2003), or agricultural management practices (Seghers et al. 2004) also have been identified as drivers of root endophytic bacterial community changes.
Regarding the taxonomic position of members of root bacterial microbiota, Liu et al. (2017b) reviewed previously published datasets, revealing that the main phyla were represented by Proteobacteria (ca. 50% in relative abundance), Actinobacteria (ca. 10%), Firmicutes (ca. 10%), and Bacteroidetes (ca. 10%). They also reported that Chloroflexi, Cyanobacteria, Planctomycetes, Verrucomicrobia, Nitrospirae, and Armatimonadetes were common in root tissues, while others, for example, Acidobacteria and Gemmatimonadetes, almost were excluded from the root endosphere. As previously mentioned, however, several studies have confirmed the active roles of host plants in the recruitment of selected bacteria from the nearby soil environment. For example, the genera Enterobacter, Pseudomonas, and Stenotrophomonas (Proteobacteria) represented the core bacterial endophytes in the roots of sweet potato and rice (Marques et al. 2015; Sessitsch et al. 2012). Accordingly, Pseudomonas-like OTUs dominated in the roots of Populus deltoides (34%) (Gottel et al. 2011), while Proteobacteria, Actinobacteria, and Bacteroidetes were the dominating phyla in root bacterial communities of A. thaliana (Bulgarelli et al. 2012; Lundberg et al. 2012; Bodenhausen et al. 2013), wheat, and tomato (Liu et al. 2017a; Lee et al. 2019). In addition to such taxa, Firmicutes occurred in the roots of Aloe vera and Capsicum annuum (Akinsanya et al. 2015; Barraza et al. 2020). Overall, the strong selection by the host plant results in recruitment of an endophytic bacterial community much simpler than that of the rhizosphere and of the nearby soil environment (Novello et al. 2017).
Diversity and functionality of AMF-associated bacterial communities
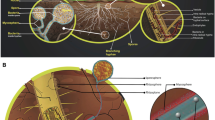
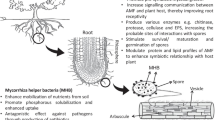
The services provided by AMF often are facilitated by the large and diverse beneficial bacterial communities living closely associated with spores, sporocarps, and extraradical mycelium, frequently embedded in spore wall layers and, in sporocarpic species, in the microniches formed by peridial hyphae (Walley and Germida 1996; Filippi et al. 1998; Iffis et al. 2014) (Table 2). The mycorrhizosphere microbiota show diverse functional activities, ranging from the role of “mycorrhiza helper” (MH) to that of “plant growth promoter” (PGP). MH bacteria (MHB) can promote spore germination, mycelial growth, and mycorrhiza establishment, while PGP bacteria (PGPB) have the ability to enhance plant growth, nutrition, health, and stress resistance (Barea et al. 2002; Frey-Klett et al. 2007). In addition, some components of such beneficial microbiota possess both MH and PGP traits (Xavier and Germida 2003; Battini et al. 2017). Overall, MHB and PGPB show activities promoting and complementing those of AMF (Turrini et al. 2018; Giovannini et al. 2020). The metabolic traits underlying MH functions include growth factor production and detoxification of antagonistic substances, while PGP properties can range from N2 fixation, nutrient mobilization, and nutrient uptake facilitation to plant hormone, antibiotic, and siderophore production, or systemic resistance induction (Frey-Klett et al. 2007; Hayat et al. 2010).
From a taxonomic viewpoint, the composition of AMF-associated bacterial microbiota strongly depends on AMF identity. Indeed, PCR-DGGE analysis of 16S rDNA showed that the bacterial communities associated with AMF spores were more influenced by fungal than host plant species. Overall, PCR-DGGE allowed the detection of bacterial sequences affiliated with the genera Cellvibrio, Chondromyces, Lysobacter, Pseudomonas (Proteobacteria), and Flexibacter (Bacteroidetes). Such bacteria, in particular the genus Flexibacter, well known for their ability to degrade biopolymers, were suspected of feeding on the spore wall, which consists mainly of chitin (Roesti et al. 2005). The same molecular approach revealed differences in the composition of spore-associated bacterial communities of two AMF, Gigaspora margarita and Gigaspora rosea, and showed that most of the bacterial sequences from G. margarita were affiliated with Proteobacteria (Azospirillum, Azovibrio, Polyangium, Ramlibacter, Rubrivivax, Sphingomonas, Rhizobium) and Actinobacteria (Streptomyces, Amycolatopsis, and Pseudonocardia) (Long et al. 2008). Interestingly, by PCR-DGGE and band sequencing, Agnolucci et al. (2015) revealed that the spores of six different AMF harbored unique bacterial communities, which were not correlated with the taxonomic positions of the fungi. The sequences were affiliated with Actinomycetales, Bacillales, Burkholderiales, Pseudomonadales, and Rhizobiales, all orders encompassing taxa known as PGP bacteria. These three mentioned works reached consistent conclusions, suggesting that the differences in the composition of spore walls or spore exudates may affect the recruitment of spore-associated bacterial communities. In contrast, bacterial communities closely associated with AMF spores were reported to be shaped not only by fungal identity, but also by the identity of the host plant (Iffis et al. 2016). Gammaproteobacteria were more abundant in spores collected from Solidago canadensis soil samples than from Populus balsamifera and Lycopus europaeus, whereas spores belonging to the genus Glomus were correlated with Betaproteobacteria, Actinobacteria, Bacilli, and Sphingobacteria. In this case, the authors suggested that the strategy of differential bacteria recruitment by diverse AMF species and isolates also might reflect variations in the composition of spores/hyphae exudates, attracting specific microbial communities (Iffis et al. 2016). The underlying mechanisms of such differential recruitment among different AMF remain to be thoroughly investigated.
A novel study reported a surprisingly high diversity of bacteria associated with AMF vesicles and intraradical spores extracted from microdissected roots of Solidago rugosa. The dominant sequences belonged to the genera Sphingomonas, Pseudomonas, Massilia, and Methylobacterium (Proteobacteria) while Bradyrhizobium, Bosea (Proteobacteria), Bacillus, and Paenibacillus (Firmicutes) were found at lower frequencies (Iffis et al. 2014).
Only a limited number of studies have investigated the bacterial communities closely associated with the surface of AMF hyphae. One of the first works, by using bromodeoxyuridine immunocapture and confocal microscopy, determined the specific attachment of a strain of Bacillus cereus to AMF hyphae (Artursson and Jansson 2003). Toljander et al. (2006) reported that five different strains of gfp-tagged soil bacteria, inoculated into in vitro cultures of two Glomus isolates growing in a controlled artificial system – T-DNA-transformed roots – exhibited different levels of hyphal attachment. In particular, Paenibacillus brasiliensis, Bacillus cereus, Paenibacillus peoriae (Firmicutes), and Pseudomonas fluorescens (Proteobacteria) attached to AMF hyphae, while Arthrobacter chlorophenolicus (Actinobacteria) did not. Such differences can be ascribed to AMF hyphal exudates that have been reported to affect the composition of bacterial communities (Toljander et al. 2007). Similar in vitro experimental systems showed that Streptomyces and members of the Oxalobacteraceae family, i.e., Duganella, Janthinobacterium, and Massilia, were specifically attached to the surface of AMF hyphae (Scheublin et al. 2010). Interestingly, 26 Burkholderia spp. strains and one Rhizobium miluonense strain were able to strongly attach to Rhizophagus irregularis hyphae and to solubilize phosphate (Taktek et al. 2015). Consistent data were obtained by a recent work reporting the isolation of 128 bacterial strains from the hyphae of Rhizoglomus irregulare (syn. Rhizophagus irregularis), of which 12 showed phosphate-solubilizing activity (Sharma et al. 2020). A distinct bacterial community closely associated with extraradical hyphae of Glomus versiforme, and conserved across divergent soils, was mainly represented by Proteobacteria (50% relative abundance), Actinobacteria (10%), Chloroflexi (9%), Acidobacteria (7%), Bacteroidetes (6%), and Fibrobacteres (4%) (Emmett et al. 2021). An in vivo study reported, for the first time, that AMF hyphae may act as “transport agents” or “highways” for bacteria. Indeed, a gfp-tagged nitrogen-fixing rhizobial strain, Bradyrhizobium diazoefficiens, was able to tightly adhere to Glomus formosanum hyphae, facilitating bacterial translocation to their legume host plant and the formation of N-fixing nodules in the root system (de Novais et al. 2020). Recently, Illumina MiSeq metagenome sequencing allowed the identification of 276 bacterial genera, belonging to 165 families, 107 orders, and 23 phyla, mostly represented by Proteobacteria, Bacteroidetes, and Actinobacteria, associated with Rhizoglomus irregulare commercial inoculum. Such richness and diversity are remarkable, given that no bacteria were deliberately added to the AM symbiont. It is interesting to note that the predominant bacterial taxa correspond to those recurrently found in the root endosphere of the majority of plant species investigated so far (Table 1).
Culture-dependent analyses not only confirmed the high diversity of bacterial communities living in association with AMF, but also showed the PGP activity of strains belonging to Actinomycetales, Bacillales, Enterobacteriales, and Rhizobiales, as IAA and siderophore producers (Agnolucci et al. 2019a). Such activities are important for plant development because the phytohormone IAA is able to modulate the growth and functioning of the root system (Duca et al. 2014), while siderophores, high-affinity iron-chelating compounds, may facilitate plant iron acquisition and control soilborne diseases by means of iron competition (Mimmo et al. 2014). Other culture-based studies reported a high abundance of the mentioned phyla, recording genera such as Micrococcus, Acidovorax, Cellulomonas, Janthinobacterium, Alcaligenes, and Flavobacterium (Xavier and Germida 2003; Bharadwaj et al. 2008b; Cruz et al. 2008). Some bacteria with PGP potentials, isolated from the spores of Rhizophagus intraradices, affiliated with the genera Sinorhizobium/Ensifer, Streptomyces, Bacillus, Arthrobacter, and Fictibacillus, also have shown MH properties (Battini et al. 2016). Indeed, seven of such bacterial isolates significantly increased hyphal length density, while two of them, Streptomyces sp. W77 and Streptomyces sp. W94, additionally were able to promote specific P uptake and translocation in maize plants (Battini et al. 2017).
Overall, the available data show that diverse bacterial taxa are differentially able to attach to AMF structures, i.e., spores, sporocarps, and hyphae, of which exudates might differ in quantity and/or quality, thus promoting or inhibiting the growth and attachment of particular bacterial communities. It is important to note that such specific and close physical relationships may be indicative of complex interactions among AMF, bacteria, and host plants, suggesting that AMF might act as carriers of the endophytic microbiota that establish in roots.
Concluding observations and future prospects
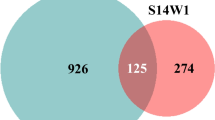
The role of AMF as drivers of the endophytic bacterial communities colonizing plant roots can be revealed by investigating possible overlaps in the taxonomic composition of root endophytic microbiota and AMF-associated bacterial communities. Only a few such comparisons currently are available, but major shifts have been revealed in the composition of the root endophytic bacterial communities of durum wheat after inoculation with the AM symbiont Funneliformis mosseae. The use of two culture-independent approaches, PCR-DGGE analysis of 16S rDNA and high-throughput sequencing of the same gene through Illumina MiSeq, has revealed that AMF inoculation increased the abundance of some genera and species of Actinobacteria and Bacteroidetes in two durum wheat cultivars, Odisseo and Saragolla (Agnolucci et al. 2019b). In particular, Funneliformis mosseae increased the abundance of Actinobacteria, such as Rhodococcus species, in the cv. Saragolla, and that of Streptomyces and Microbacterium spp. in both cultivars. This is an interesting finding, as Actinobacteria, considered promising PGP bacteria (Seipke et al. 2012), have been reported to be closely associated with the spores of six different AMF isolates (Agnolucci et al. 2015). Moreover, in the two durum wheat cultivars, AMF inoculation increased the abundance of the Proteobacteria Pantoea, a PGP genus, including species and strains able to produce siderophores, enhance Zn availability in soil, and induce plant resistance to drought stress (Moreira et al. 2016; Kamran et al. 2017; Chen et al. 2017).
The possible mechanisms operating in the recruitment of different bacterial endophytic communities in roots may be dual. As the physiological status of plants is altered when they are mycorrhizal (e.g., photosynthesis, nutrition and growth, phenology, health) (Smith and Read 2008), certain groups of endophytic bacteria might find specific conditions best suited for their growth and development and thus might be favored in colonization of such a privileged ecological niche. On the other hand, they may gain facilitated access to root tissues during AMF establishment through their close association with AMF hyphae (Toljander et al. 2006; de Novais et al. 2020). These two mechanisms may act simultaneously, and it will be difficult to discriminate between them. Notwithstanding, systematic studies on the differential occurrence of root bacterial endophytes in mycorrhizal and non-mycorrhizal plants still are needed.
In the years to come, a series of key questions remain to be answered. The first question concerns whether members of AMF-associated bacterial communities can actually establish in the root system, becoming endophytic, and in what proportion. If so, would they be able to show the specific PGP properties revealed when isolated and grown in vitro? The results obtained will raise the question as to whether AMF identity and diversity can affect the structure and composition of root endophytic bacterial communities. Moreover, as AMF commercial inocula carry their own associated bacterial microbiota, could they represent suitable tools to introduce beneficial root endophytes in sustainable agriculture? Such questions can only be answered by targeted research on the diversity and functionality of root endophytic communities as affected by the presence of AMF and AMF-associated bacterial communities.
References
Agnolucci M, Battini F, Cristani C, Giovannetti M (2015) Diverse bacterial communities are recruited on spores of different arbuscular mycorrhizal fungal isolates. Biol Fertil Soils 51:379–389. https://doi.org/10.1007/s00374-014-0989-5
Agnolucci M, Avio L, Pepe A, Turrini A, Cristani C, Bonini P, Cirino B, Colosimo F, Ruzzi M Giovannetti M (2019a) Bacteria associated with a commercial mycorrhizal inoculum: community composition and multifunctional activity as assessed by Illumina sequencing and culture-dependent tools Front Plant Sci 9 1956 https://doi.org/10.3389/fpls.2018.01956
Agnolucci M, Palla M, Cristani C, Cavallo N, Giovannetti M, De AM, Gobbetti M, Minervini F (2019b) Beneficial plant microorganisms affect the endophytic bacterial communities of durum wheat roots as detected by different molecular approaches. Front Microbiol 10:2500. https://doi.org/10.3389/fmicb.2019.02500
Agnolucci M, Avio L, Palla M, Sbrana C, Turrini A, Giovannetti M (2020) Health-promoting properties of plant products: the role of mycorrhizal fungi and associated bacteria. Agronomy 10:1864. https://doi.org/10.3390/agronomy10121864
Akinsanya MA, Goh JK, Lim SP, Ting ASY (2015) Metagenomics study of endophytic bacteria in Aloe vera using next-generation technology. Genom Data 6:159–163. https://doi.org/10.1016/j.gdata.2015.09.004
Artursson V, Jansson JK (2003) Use of bromodeoxyuridine immunocapture to identify active bacteria associated with arbuscular mycorrhizal hyphae. Appl Environ Microbiol 69:6208–6215. https://doi.org/10.1128/aem.69.10.6208-6215.2003
Avio L, Turrini A, Giovannetti M, Sbrana C (2018) Designing the ideotype mycorrhizal symbionts for the production of healthy food. Front Plant Sci 9:1089. https://doi.org/10.3389/fpls.2018.01089
Azcón-Aguilar C, Barea JM (2015) Nutrient cycling in the mycorrhizosphere. J Soil Sci Plant Nutr 15:372–396. https://doi.org/10.4067/S0718-95162015005000035
Barea JM, Azcón R, Azcón-Aguilar C (2002) Mycorrhizosphere interactions to improve plant fitness and soil quality. Antonie Van Leeuwenhoek 81:343–351. https://doi.org/10.1023/A:1020588701325
Barraza A, Castellanos C-C, T, Loera-Muro A, (2020) Bacterial community characterization of the rhizobiome of plants belonging to Solanaceae family cultivated in desert soils. Ann Microbiol 70:1–14. https://doi.org/10.1186/s13213-020-01572-x
Battini F, Cristani C, Giovannetti M, Agnolucci M (2016) Multifunctionality and diversity of culturable bacterial communities strictly associated with spores of the plant beneficial symbiont Rhizophagus intraradices. Microbiol Res 183:68–79. https://doi.org/10.1016/j.micres.2015.11.012
Battini F, Grønlund M, Agnolucci M, Giovannetti M, Jakobsen I (2017) Facilitation of phosphorus uptake in maize plants by mycorrhizosphere bacteria. Sci Rep 7:4686. https://doi.org/10.1038/s41598-017-04959-0
Bharadwaj DP, Lundquist PO, Alström S (2008a) Arbuscular mycorrhizal fungal spore-associated bacteria affect mycorrhizal colonization, plant growth and potato pathogens. Soil Biol Biochem 40:2494–2501. https://doi.org/10.1016/j.soilbio.2008.06.012
Bharadwaj DP, Lundquist PO, Persson P, Alström S (2008b) Evidence for specificity of cultivable bacteria associated with arbuscular mycorrhizal fungal spores. FEMS Microbiol Ecol 65:310–322. https://doi.org/10.1111/j.1574-6941.2008.00515.x
Bidondo LF, Colombo R, Bompadre J, Benavides M, Scorza V, Silvani V, Pérgola M, Godeas A (2016) Cultivable bacteria associated with infective propagules of arbuscular mycorrhizal fungi. Implications for mycorrhizal activity. Appl Soil Ecol 105:86–90. https://doi.org/10.1016/j.apsoil.2016.04.013
Bitterlich M, Franken P, Graefe J (2018) Arbuscular mycorrhiza improves substrate hydraulic conductivity in the plant available moisture range under root growth exclusion. Front Plant Sci 9:301. https://doi.org/10.3389/fpls.2018.00301
Bodenhausen N, Horton MW, Bergelson J (2013) Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PLoS ONE 8(2):e56329. https://doi.org/10.1371/journal.pone.0056329
Bonito G, Benucci GMN, Hameed K, Weighill D, Jones P, Chen KH, Jacobson D, Schadt C, Vilgalys R (2019) Fungal-bacterial networks in the Populus rhizobiome are impacted by soil properties and host genotype. Front Microbiol 10:481. https://doi.org/10.3389/fmicb.2019.00481
Budi SW, Bakhtiar Y, May NL (2012) Bacteria associated with arbuscula mycorrhizal spores Gigaspora margarita and their potential for stimulating root mycorrhizal colonization and neem (Melia azedarach Linn) seedling growth. Microbiol Indones 6:6–6. https://doi.org/10.5454/mi.6.4.6
Bulgarelli D, Rott M, Schlaeppi K, Loren V, van Themaat E, Ahmadinejad N, Assenza F, Rauf P, Huettel B, Reinhardt R, Schmelzer E, Peplies J, Gloeckner FO, Amann R, Eickhorst T, Schulze-Lefert P (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488:91–95. https://doi.org/10.1038/nature11336
Bulgarelli D, Garrido-Oter R, Münch PC, Weiman A, Dröge J, Pan Y, McHardy AC, Schulze-Lefert P (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17:392–403. https://doi.org/10.1016/j.chom.2015.01.011
Chen C, Xin K, Liu H, Cheng J, Shen X, Wang Y, Zhang L (2017) Pantoea alhagi, a novel endophytic bacterium with ability to improve growth and drought tolerance in wheat. Sci Rep 7:41564. https://doi.org/10.1038/srep41564
Coleman-Derr D, Desgarennes D, Fonseca-Garcia C, Gross S, Clingenpeel S, Woyke T, North G, Visel A, Partida-Martinez LP, Tringe SG (2016) Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. New Phytol 209:798–811. https://doi.org/10.1111/nph.13697
Compant S, Clément C, Sessitsch A (2010) Plant growth-promoting bacteria in the rhizo- and endosphere of plants: their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem 42:669–678. https://doi.org/10.1016/j.soilbio.2009.11.024
Conn VM, Franco CMM (2004) Analysis of the endophytic actinobacterial population in the roots of wheat (Triticum aestivum L.) by terminal restriction fragment length polymorphism and sequencing of 16S rRNA clones. Appl Environ Microbiol 70:1787–1794. https://doi.org/10.1128/AEM.70.3.1787-1794.2004
Cruz AF, Ochiai HS, S, Yasuda A, Ishii T, (2008) Isolation and analysis of bacteria associated with spores of Gigaspora margarita. J Appl Microbiol 104:1711–1717. https://doi.org/10.1111/j.1365-2672.2007.03695.x
Cruz AF, Ishii T (2011) Arbuscular mycorrhizal fungal spores host bacteria that affect nutrient biodynamics and biocontrol of soil-borne plant pathogens. Biol Open, 1:52–57. https://doi.org/10.1242/bio.2011014
de Novais CB, Sbrana C, da Conceição JE, Rouws LFM, Giovannetti M, Avio L, Siqueira JO, Saggin OJ Jr, Ribeiro da Silva EM, de Faria SM (2020) Mycorrhizal networks facilitate the colonization of legume roots by a symbiotic nitrogen-fixing bacterium. Mycorrhiza 30:389–396. https://doi.org/10.1007/s00572-020-00948-w
Duca D, Lorv J, Patten CL, Rose D, Glick BR (2014) Indole-3-acetic acid in plant–microbe interactions. Anton Van Leeuw 106:85–125. https://doi.org/10.1007/s10482-013-0095-y
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112:E911–E920. https://doi.org/10.1073/pnas.1414592112
Ek-Ramos MJ, Gomez-Flores R, Orozco-Flores AA, Rodríguez-Padilla C, González-Ochoa G, Tamez-Guerra P (2019) Bioactive products from plant-endophytic Gram-positive bacteria. Front Microbiol 10:463. https://doi.org/10.3389/fmicb.2019.00463
Emmett BD, Lévesque-Tremblay V, Harrison MJ (2021) Conserved and reproducible bacterial communities associate with extraradical hyphae of arbuscular mycorrhizal fungi ISME J 1–13 https://doi.org/10.1038/s41396-021-00920-2
Ferrando L, Fernández Scavino A (2015) Strong shift in the diazotrophic endophytic bacterial community inhabiting rice (Oryza sativa) plants after flooding. FEMS Microbiol Ecol 91:9, fiv104. https://doi.org/10.1093/femsec/fiv104
Filippi C, Bagnoli G, Citernesi AS, Giovannetti M (1998) Ultrastructural spatial distribution of bacteria associated with sporocarps of Glomus mosseae. Symbiosis 24:1–12
Finan TM (2002) Evolving insights: symbiosis islands and horizontal gene transfer. J Bacteriol 184:2855–2856. https://doi.org/10.1128/JB.184.11.2855-2856.2002
Fonseca-García C, Coleman-Derr D, Garrido E, Visel A, Tringe SG, Partida-Martínez LP (2016) The cacti microbiome: interplay between habitat-filtering and host-specificity. Front Microbiol 7:150. https://doi.org/10.3389/fmicb.2016.00150
Food and Agriculture Organization (2011) A policymaker’s guide to the sustainable intensification of smallholder crop production; FAO: Rome, Italy, 2011; Available online: http://www.fao.org/3/a-i2215e.pdf.
Forchetti G, Masciarelli O, Alemano S, Alvarez D, Abdala G (2007) Endophytic bacteria in sunflower (Helianthus annuus L.): isolation, characterization, and production of jasmonates and abscisic acid in culture medium. Appl Microbiol Biotech 76:1145–1152. https://doi.org/10.1007/s00253-007-1077-7
Frey-Klett P, Garbaye J, Tarkka M (2007) The mycorrhiza helper bacteria revisited. New Phytol 176:22–36. https://doi.org/10.1111/j.1469-8137.2007.02191.x
Gaiero JR, McCall CA, Thompson KA, Day NJ, Best AS, Dunfield KE (2013) Inside the root microbiome: bacterial root endophytes and plant growth promotion. Am J Bot 100:1738–1750. https://doi.org/10.3732/ajb.1200572
Gianinazzi S, Gollotte A, Binet MN, van Tuinen D, Redecker D, Wipf D (2010) Agroecology: the key role of arbuscular mycorrhizas in ecosystem services. Mycorrhiza 20:519–530. https://doi.org/10.1007/s00572-010-0333-3
Glick BR (2014) Bacteria with ACC deaminase can promote plant growth and help to feed the world. Microbiol Res 169:30–39. https://doi.org/10.1016/j.micres.2013.09.009
Giovannini L, Palla M, Agnolucci M, Avio L, Sbrana C, Turrini A, Giovannetti M (2020) Arbuscular mycorrhizal fungi and associated microbiota as plant biostimulants: research strategies for the selection of the best performing inocula. Agronomy 10:106. https://doi.org/10.3390/agronomy10010108
Gottel NR, Castro HF, Kerley M, Yang Z, Pelletier DA, Podar M, Karpinets T, Uberbacher E, Tuskan GA, Vilgalys R, Doktycz MJ, Schadt CW (2011) Distinct microbial communities within the endosphere and rhizosphere of Populus deltoides roots across contrasting soil types. Appl Environ Microbiol 77:5934–5944. https://aem.asm.org/content/77/17/5934.short
Hameed A, Yeh MW, Hsieh YT, Chung WC, Lo CT, Sen YL (2015) Diversity and functional characterization of bacterial endophytes dwelling in various rice (Oryza sativa L.) tissues, and their seed-borne dissemination into rhizosphere under gnotobiotic P-stress. Plant Soil 394:177–197. https://doi.org/10.1007/s11104-015-2506-5
Hardoim PR, van Overbeek LS, van Elsas JD (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol 16:463–471. https://doi.org/10.1016/j.tim.2008.07.008
Hardoim PR, Andreote FD, Reinhold-Hurek B, Sessitsch A, van Overbeek LS, van Elsas JD (2011) Rice root-associated bacteria: insights into community structures across 10 cultivars. FEMS Microbiol Ecol 77:154–164. https://doi.org/10.1111/j.1574-6941.2011.01092.x
Hardoim PR, van Overbeek LS, Berg G, Pirttilä AM, Compant S, Campisano A, Döring M, Sessitsch A (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79:293–320. https://doi.org/10.1128/mmbr.00050-14
Hayat R, Ali S, Amara U, Khalid R, Ahmed I (2010) Soil beneficial bacteria and their role in plant growth promotion: a review. Ann Microbiol 60:579–598. https://doi.org/10.1007/s13213-010-0117-1
Iffis B, St-Arnaud M, Hijri M (2014) Bacteria associated with arbuscular mycorrhizal fungi within roots of plants growing in a soil highly contaminated with aliphatic and aromatic petroleum hydrocarbons. FEMS Microbiol Lett 358:44–54. https://doi.org/10.1111/1574-6968.12533
Iffis B, St-Arnaud M, Hijri M (2016) Petroleum hydrocarbon contamination, plant identity and arbuscular mycorrhizal fungal (AMF) community determine assemblages of the AMF spore-associated microbes. Environ Microbiol 18:2689–2704. https://doi.org/10.1111/1462-2920.13438
Kaga H, Mano H, Tanaka F, Watanabe A, Kaneko S, Morisaki H (2009) Rice seeds as sources of endophytic bacteria. Microbes Environ 24:154–162. https://doi.org/10.1264/jsme2.ME09113
Kameoka H, Maeda T, Okuma N, Kawaguchi M (2019) Structure-specific regulation of nutrient transport and metabolism in arbuscular mycorrhizal fungi. Plant Cell Physiol 60:2272–2281. https://doi.org/10.1093/pcp/pcz122
Kamran S, Shahid I, Baig DN, Rizwan M, Malik KA, Mehnaz S (2017) Contribution of zinc solubilizing bacteria in growth promotion and zinc content of wheat. Front Microbiol 8:2593. https://doi.org/10.3389/fmicb.2017.02593
Karray F, Gargouri M, Chebaane A, Mhiri N, Mliki A, Sayadi S (2020) Climatic aridity gradient modulates the diversity of the rhizosphere and endosphere bacterial microbiomes of Opuntia ficus-indica. Front Microbiol 11:1622. https://doi.org/10.3389/fmicb.2020.01622
Krishnamoorthy R, Kim K, Subramanian P, Senthilkumar M, Anandham R, Sa T (2016) Arbuscular mycorrhizal fungi and associated bacteria isolated from salt-affected soil enhances the tolerance of maize to salinity in coastal reclamation soil. Agric Ecosyst Environ 231:233–239. https://doi.org/10.1016/j.agee.2016.05.037
Ku YS, Rehman HM, Lam HM (2019) Possible roles of rhizospheric and endophytic microbes to provide a safe and affordable means of crop biofortification. Agronomy 9:764. https://doi.org/10.3390/agronomy9110764
Lasudee K, Tokuyama S, Lumyong S, Pathom-Aree W (2018) Actinobacteria associated with arbuscular mycorrhizal Funneliformis mosseae spores, taxonomic characterization and their beneficial traits to plants: evidence obtained from mung bean (Vigna radiata) and Thai jasmine rice (Oryza sativa). Front Microbiol 9:1247. https://doi.org/10.3389/fmicb.2018.01247
Lecomte J, St-Arnaud M, Hijri M (2011) Isolation and identification of soil bacteria growing at the expense of arbuscular mycorrhizal fungi. FEMS Microbiol Lett 317:43–51. https://doi.org/10.1111/j.1574-6968.2011.02209.x
Lee SA, Kim Y, Kim JM, Chu B, Joa JH, Sang MK, Song J, Weon HY (2019) A preliminary examination of bacterial, archaeal, and fungal communities inhabiting different rhizocompartments of tomato plants under real-world environments. Sci Rep 9:1–15. https://doi.org/10.1038/s41598-019-45660-8
Liu H, Carvalhais LC, Crawford M, Singh E, Dennis PG, Pieterse CMJ, Schenk PM (2017a) Inner plant values: diversity, colonization and benefits from endophytic bacteria. Front Microbiol 8:2552. https://doi.org/10.3389/fmicb.2017.02552
Liu H, Carvalhais LC, Schenk PM, Dennis PG (2017b) Effects of jasmonic acid signalling on the wheat microbiome differ between body sites. Sci Rep 7:41776. https://doi.org/10.1038/srep41766
Long L, Zhu H, Yao Q, Ai Y (2008) Analysis of bacterial communities associated with spores of Gigaspora margarita and Gigaspora rosea. Plant Soil 310:1–9. https://doi.org/10.1007/s11104-008-9611-7
Long L, Lin Q, Yao Q, Zhu H (2017) Population and function analysis of cultivable bacteria associated with spores of arbuscular mycorrhizal fungus Gigaspora margarita. 3 Biotech, 7:8. https://doi.org/10.1007/s13205-017-0612-1
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, Del RTG, Edgar RC, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90. https://doi.org/10.1038/nature11237
Manter DK, Delgado JA, Holm DG, Stong RA (2010) Pyrosequencing reveals a highly diverse and cultivar-specific bacterial endophyte community in potato roots. Microb Ecol 60:157–166. https://doi.org/10.1007/s00248-010-9658-x
Marasco R, Rolli E, Fusi M, Michoud G, Daffonchio D (2018) Grapevine rootstocks shape underground bacterial microbiome and networking but not potential functionality. Microbiome 6:3. https://doi.org/10.1186/s40168-017-0391-2
Marques JM, da Silva TF, Vollú RE, de Lacerda JRM, Blank AF, Smalla K, Seldin L (2015) Bacterial endophytes of sweet potato tuberous roots affected by the plant genotype and growth stage. Appl Soil Ecol 96:273–281. https://doi.org/10.1016/j.apsoil.2015.08.020
Mayo K, Davis RE, Motta J (1986) Stimulation of germination of spores of Glomus versiforme by spore-associated bacteria. Mycologia 78:426–431. https://www.jstor.org/stable/3793046
Mimmo T, Del Buono D, Terzano R, Tomasi N, Vigani G, Crecchio C, Pinton R, Zocchi G, Cesco S (2014) Rhizospheric organic compounds in the soil microorganism-plant system: their role in iron availability. Eur J Soil Sci 65:629–642. https://doi.org/10.1111/ejss.12158
Minervini F, Celano G, Lattanzi A, Tedone L, de Mastro G, Gobbetti M, de Angelis M (2015) Lactic acid bacteria in durum wheat flour are endophytic components of the plant during its entire life cycle. Appl Environ Microbiol 81:6736–6748. https://doi.org/10.1128/AEM.01852-15
Mitter EK, de Freitas JR, Germida JJ (2017) Bacterial root microbiome of plants growing in oil sands reclamation covers. Front Microbiol 8:849. https://doi.org/10.3389/fmicb.2017.00849
Mitter EK, Kataoka R, de Freitas JR, Germida JJ (2019) Potential use of endophytic root bacteria and host plants to degrade hydrocarbons. Int J Phytoremediation21:928–938. https://doi.org/10.1080/15226514.2019.1583637
Mocali S, Bertelli E, Di Cello F, Mengoni A, Sfalanga A, Viliani F, Caciotti A, Tegli S, Surico G, Fani R (2003) Fluctuation of bacteria isolated from elm tissues during different seasons and from different plant organs. Res Microbiol 154:105–114. https://doi.org/10.1016/S0923-2508(03)00031-7
Moreira FS, da Costa PB, de Souza R, Beneduzi A, Lisboa BB, Vargas LK, Passaglia LMP (2016) Functional abilities of cultivable plant growth promoting bacteria associated with wheat (Triticum aestivum L.) crops. Genet Mol Biol 39:111–121. https://doi.org/10.1590/1678-4685-GMB-2015-0140
Naylor D, Degraaf S, Purdom E, Coleman-Derr D (2017) Drought and host selection influence bacterial community dynamics in the grass root microbiome. ISME J11:2691–2704. https://doi.org/10.1038/ismej.2017.118
Novello G, Gamalero E, Bona E, Boatti L, Mignone F, Massa N, Cesaro P, Lingua G, Berta G (2017). The rhizosphere bacterial microbiota of Vitis vinifera cv. Pinot Noir in an integrated pest management vineyard. Front. Microbiol. 8, 1528. https://doi.org/10.3389/fmicb.2017.01528
Pecundo MH, Chang ACG, Chen T, dela Cruz TEE, Ren H, Li N, (2021) Full-length 16S rRNA and ITS gene sequencing revealed rich microbial flora in roots of Cycas spp. in China. Evol Bioinform 17:1–16. https://doi.org/10.1177/1176934321989713
Pepe A, Sbrana C, Ferrol N, Giovannetti M (2017) An in vivo whole-plant experimental system for the analysis of gene expression in extraradical mycorrhizal mycelium. Mycorrhiza 27:659–668. https://doi.org/10.1007/s00572-017-0779-7
Pei C, Mi C, Sun L, Liu W, Li O, Hu X (2017) Diversity of endophytic bacteria of Dendrobium officinale based on culture-dependent and culture-independent methods. Biotechnol Biotechnol Equip 31:112–119. https://doi.org/10.1080/13102818.2016.1254067
Philippot L, Raaijmakers JM, Lemanceau P, Van Der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol11:789–799. https://doi.org/10.1038/nrmicro3109
Pinski A, Betekhtin A, Hupert-Kocurek K, Mur LAJ, Hasterok R (2019) Defining the genetic basis of plant–endophytic bacteria interactions. Int J Mol Sci 20:1947. https://doi.org/10.3390/ijms20081947
Rambelli A (1973) The Rhizosphere of Mycorrhizae. In: Marks GC, Kozlowski TT (eds) Ectomycorrhizae: their ecology and physiology. Academic Press, pp 299–343. https://doi.org/10.1016/b978-0-12-472850-9.50014-110.1007/s00572-021-01040-7343. https://doi.org/10.1016/b978-0-12-472850-9.50014-1
Rehman A, Farooq M, Naveed M, Nawaz A, Shahzad B (2018) Seed priming of Zn with endophytic bacteria improves the productivity and grain biofortification of bread wheat. Eur J Agron 94:98–107. https://doi.org/10.1016/j.eja.2018.01.017
Roesti D, Ineichen K, Braissant O, Redecker D, Wiemken A, Aragno M (2005) Bacteria associated with spores of the arbuscular mycorrhizal fungi Glomus geosporum and Glomus constrictum. Appl Environ Microbiol 71:6673–6679. https://doi.org/10.1128/AEM.71.11.6673-6679.2005
Rouphael Y, Franken P, Schneider C, Schwarz D, Giovannetti M, Agnolucci M, De Pascale S, Bonini P, Colla G (2015) Arbuscular mycorrhizal fungi act asbiostimulants in horticultural crops. Sci Hortic 196:91–108. https://doi.org/10.1016/j.scienta.2015.09.002
Ryan RP, Germaine K, Franks A, Ryan DJ, Dowling DN (2008) Bacterial endophytes: recent developments and applications. FEMS Microbiol Lett 278:1–9. https://doi.org/10.1111/j.1574-6968.2007.00918.x
Scheublin TR, Sanders IR, Keel C, Van Der Meer JR (2010) Characterisation of microbial communities colonising the hyphal surfaces of arbuscular mycorrhizal fungi. ISME J 4:752–763. https://doi.org/10.1038/ismej.2010.5
Schlaeppi K, Dombrowski N, Oter RG, Themaat VLVE, Schulze-Lefert P (2014) Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc Natl Acad Sci USA 111:585–592. https://doi.org/10.1073/pnas.1321597111
Seghers D, Wittebolle L, Top EM, Verstraete W, Siciliano SD (2004) Impact of agricultural practices on the Zea mays L. endophytic community. Appl Environ Microbiol 70:1475–1482. https://doi.org/10.1128/AEM.70.3.1475-1482.2004
Seipke RF, Kaltenpoth M, Hutchings MI (2012) Streptomyces as symbionts: an emerging and widespread theme? FEMS Microbiol Rev 36:862–876. https://doi.org/10.1111/j.1574-6976.2011.00313.x
Selvakumar G, Krishnamoorthy R, Kim K, Sa TM (2016) Genetic diversity and association characters of bacteria isolated from arbuscular mycorrhizal fungal spore walls. PLoS One, 11(8):e0160356. https://doi.org/10.1371/journal.pone.0160356
Sessitsch A, Hardoim P, Döring J, Weilharter A, Krause A, Woyke T, Mitter B, Hauberg-Lotte L, Friedrich F, Rahalkar M, Hurek T, Sarkar A, Bodrossy L, Van Overbeek L, Brar D, Van Elsas JD, Reinhold-Hurek B (2012) Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. MPMI 25:28–36. https://doi.org/10.1094/MPMI-08-11-0204
Sharma S, Compant S, Ballhausen MB, Ruppel S, Franken P (2020) The interaction between Rhizoglomus irregulare and hyphae attached phosphate solubilizing bacteria increases plant biomass of Solanum lycopersicum. Microbiol Res 240:126556. https://doi.org/10.1016/j.micres.2020.126556
Shi Y, Yang H, Zhang T, Sun J, Lou K (2014) Illumina-based analysis of endophytic bacterial diversity and space-time dynamics in sugar beet on the north slope of Tianshan mountain. Appl Microbiol Biotech 98:6375–6385. https://doi.org/10.1007/s00253-014-5720-9
Sikes BA, Kottenie K, Klironomos JN (2009) Plant and fungal identity determines pathogen protection of plant roots by arbuscular mycorrhizas. J Ecol 97:1274–1280. https://doi.org/10.1111/j.1365-2745.2009.01557.x
Singer E, Bonnette J, Woyke T, Juenger TE (2019) Conservation of endophyte bacterial community structure across two Panicum grass species. Front Microbiol 10:2181. https://doi.org/10.3389/fmicb.2019.02181
Smith SE, Read DJ (2008) Mycorrhizal symbiosis. Academic Press, London
Strobel G, Daisy B, Castillo U, Harper J (2004) Natural products from endophytic microorganisms. J Nat Prod 67:257–268. https://doi.org/10.1021/np030397v
Szymańska S, Borruso L, Brusetti L, Hulisz P, Furtado B, Hrynkiewicz K (2018) Bacterial microbiome of root-associated endophytes of Salicornia europaea in correspondence to different levels of salinity. Environ Sci Pollut Res 25:25420–25431. https://doi.org/10.1007/s11356-018-2530-0
Taktek S, Trépanier M, Servin PM, St-Arnaud M, Piché Y, Fortin JA, Antoun H (2015) Trapping of phosphate solubilizing bacteria on hyphae of the arbuscularmycorrhizal fungus Rhizophagus irregularis DAOM 197198. Soil Biol Biochem 90:1–9. https://doi.org/10.1016/j.soilbio.2015.07.016
Toljander JF, Artursson V, Paul LR, Jansson JK, Finlay RD (2006) Attachment of different soil bacteria to arbuscular mycorrhizal fungal extraradical hyphae is determined by hyphal vitality and fungal species. FEMS Microbiol Lett 254:34–40. https://doi.org/10.1111/j.1574-6968.2005.00003.x
Toljander JF, Lindahl BD, Paul LR, Elfstrand M, Finlay RD (2007) Influence of arbuscular mycorrhizal mycelial exudates on soil bacterial growth and community structure. FEMS Microbiol Ecol 61:295–304. https://doi.org/10.1111/j.1574-6941.2007.00337.x
Truyens S, Weyens N, Cuypers A, Vangronsveld J (2015) Bacterial seed endophytes: genera, vertical transmission and interaction with plants. Environ Microbiol Rep7:40–50. https://doi.org/10.1111/1758-2229.12181
Turrini A, Avio L, Giovannetti M, Agnolucci M (2018) Functional complementarity of arbuscular mycorrhizal fungi and associated microbiota: the challenge of translational research. Front Plant Sci 9:1407. https://doi.org/10.3389/fpls.2018.01407
Van Overbeek L, Van Elsas JD (2008) Effects of plant genotype and growth stage on the structure of bacterial communities associated with potato (Solanum tuberosum L.). FEMS Microbiol Ecol 64:283–296. https://doi.org/10.1111/j.1574-6941.2008.00469.x
Walley FL, Germida JJ (1996) Failure to decontaminate Glomus clarum NT4 spores is due to spore wall-associated bacteria. Mycorrhiza 6:43–49
Wang Y, Zhang W, Ding C, Zhang B, Huang Q, Huang R, Su X (2019) Endophytic communities of transgenic poplar were determined by the environment and niche rather than by transgenic events. Front Microbiol 10:588. https://doi.org/10.3389/fmicb.2019.00588
Xavier LJC, Germida JJ (2003) Bacteria associated with Glomus clarum spores influence mycorrhizal activity. Soil Biol Biochem 35:471–478. https://doi.org/10.1016/S0038-0717(03)00003-8
Zheng Y, Gong X (2019) Niche differentiation rather than biogeography shapes the diversity and composition of microbiome of Cycas panzhihuaensis. Microbiome 7:152. https://doi.org/10.1186/s40168-019-0770-y
Acknowledgements
GU was supported by the University of Pisa PhD Programme (Agriculture, Food and Environment).
Funding
Open access funding provided by Università di Pisa within the CRUI-CARE Agreement. This work was financially supported by the University of Pisa (Fondi di Ateneo).
Author information
Authors and Affiliations
Contributions
MA and AT conceived the study, GU and LA compiled and analyzed data, GU wrote the first draft, and all authors contributed to the manuscript. This paper was part of GU’s doctoral thesis work at the University of Pisa.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ujvári, G., Turrini, A., Avio, L. et al. Possible role of arbuscular mycorrhizal fungi and associated bacteria in the recruitment of endophytic bacterial communities by plant roots. Mycorrhiza 31, 527–544 (2021). https://doi.org/10.1007/s00572-021-01040-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-021-01040-7