Abstract
In recent years, multidrug-resistant Acinetobacter baumannii has emerged globally as a major threat to the healthcare system. It is now listed by the World Health Organization as a priority one for the need of new therapeutic agents. A. baumannii has the capacity to develop robust biofilms on biotic and abiotic surfaces. Biofilm development allows these bacteria to resist various environmental stressors, including antibiotics and lack of nutrients or water, which in turn allows the persistence of A. baumannii in the hospital environment and further outbreaks. Investigation into therapeutic alternatives that will act on both biofilm formation and antimicrobial resistance (AMR) is sorely needed. The aim of the present review is to critically discuss the various mechanisms by which AMR and biofilm formation may be co-regulated in A. baumannii in an attempt to shed light on paths towards novel therapeutic opportunities. After discussing the clinical importance of A. baumannii, this critical review highlights biofilm-formation genes that may be associated with the co-regulation of AMR. Particularly worthy of consideration are genes regulating the quorum sensing system AbaI/AbaR, AbOmpA (OmpA protein), Bap (biofilm-associated protein), the two-component regulatory system BfmRS, the PER-1 β-lactamase, EpsA, and PTK. Finally, this review discusses ongoing experimental therapeutic strategies to fight A. baumannii infections, namely vaccine development, quorum sensing interference, nanoparticles, metal ions, natural products, antimicrobial peptides, and phage therapy. A better understanding of the mechanisms that co-regulate biofilm formation and AMR will help identify new therapeutic targets, as combined approaches may confer synergistic benefits for effective and safer treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The introduction, misuse, and overuse of antibiotics have facilitated the development of antimicrobial resistance (AMR) far beyond its original, natural, selection-based evolution [1, 2]. AMR occurs when microorganisms no longer respond to medicines they previously were sensitive to. AMR dissemination is facilitated by the horizontal transfer of a broad range of antibiotic resistance genes in bacteria from humans, animals, and in the environment [2,3,4]. Hence, a One-Health-based approach is key to our attempts at better understanding and, ultimately, solving this global threat [2, 5, 6]. As the clinical pipeline of new antimicrobials dries up, increased difficulties in treating bacterial infections due to AMR lay the terrifying foundations for emerging pandemic-size healthcare challenges [2, 7].
Bacteria may live as free-swimming, planktonic organisms. In their natural environments, however, bacteria are mostly sessile, adhered to a substrate, and form complex multispecies communities known as biofilms. Biofilms are the cause of enormous medical challenges and represent a life form that naturally resists exposure to environmental attacks, including that of antibiotics [8]. Biofilm communities secrete their own extracellular matrix, typically consisting of polysaccharides, proteins, and DNA. This matrix reduces the free diffusion of antimicrobials into the mature biofilms and facilitates the development of antibiotic resistance, which makes biofilms significantly more difficult to eradicate than planktonic organisms. Intense ongoing research initiatives investigate quorum sensing molecules, as well as the genes and second messengers, such as c-di-GMP and cAMP, that are implicated in biofilm formation [8,9,10,11,12]. The mechanisms whereby these genetic biofilm regulators may affect the development of AMR remain poorly understood. Research into co-regulatory mechanisms of biofilm and AMR development is sorely needed, as it may help identify new antimicrobial targets.
Acinetobacter baumannii is an opportunistic pathogen responsible for many nosocomial infections that include ventilator-associated pneumonia (VAP) and bloodstream infections, especially in patients hospitalized in intensive care units (ICUs) [13,14,15]. New risks have appeared with the emergence of the COVID-19 pandemic. The COVID-19 pandemic has spawned an overuse of antimicrobials, and co-infection with other respiratory pathogens in COVID-19 patients is an emerging concern. Even though the prevalence of secondary bacterial infections in COVID-19 patients may exceed 50%, reports about secondary infections or co-infections with opportunistic pathogens in COVID-19 patients still remain scarce [16,17,18]. Co-infections in COVID-19 patients may significantly worsen disease outcomes. Recent evidence suggests that co-infection and secondary infection with A. baumannii represent a significant threat to these patients. Indeed, infections with multidrug-resistant (MDR) A. baumannii in COVID-19 patients are being reported with increased frequency, particularly in ICUs [19,20,21,22,23,24,25,26,27,28]. Spread of MDR A. baumannii in hospitals can occur via several routes, including ventilator-associated transmission and air dispersal [23, 29, 30]. Recent findings demonstrate that the lower respiratory tract bacterial microbiome of critically ill COVID-19 patients favors the establishment of carbapenem-resistant A. baumannii [31, 32]. A. baumannii is known to form biofilms in host tissues, as well as on a variety of inert surfaces, including plastic and metals found in medical equipment [33,34,35,36]. This ability further complicates the control of such infections and warrants new prevention measures targeting biofilm-forming antimicrobial-resistant A. baumannii [13, 15].
With a focus on A. baumannii, the aim of this review is to discuss mechanisms that control biofilm formation and AMR development. We discuss the current understanding of genes that may co-regulate both of these processes as a path towards the development of novel therapeutic strategies.
Clinical significance of A. baumannii
A. baumannii is a ubiquitous opportunistic pathogen belonging to the class of Gram-negative Gammaproteobacteria and is a non-fermentative, non-motile, catalase-positive coccobacillus [13, 36]. In addition to respiratory and bloodstream infections, A. baumannii may cause infections in the urinary tract and the skin and may lead to endocarditis and meningitis [13, 14, 30, 36, 37]. Some of these infections are related with the formation of biofilms, such as VAP and catheter-associated infections [37,38,39]. The mortality rate associated with infection by A. baumannii may exceed 50% [13, 30, 36, 40,41,42,43,44]. Recent reports also demonstrate that A. baumannii is responsible for community-acquired infections, such as community-acquired pneumonia with or without bacteremia. Often severe and fatal, these infections are more prevalent in patients with associated risks, such as diabetes mellitus and chronic lung diseases, and mortality can reach 64% [45,46,47,48,49].
A. baumannii expresses key virulence factors including lipopolysaccharides, capsular polysaccharides, proteases, phospholipases, outer membrane porins, outer membrane vesicles, biofilm-associated protein (Bap), iron-chelating systems, surface glycoconjugates, and protein secretion systems [14, 36, 50, 51]. However, the pathogenicity of the Acinetobacter genus, which today contains over 50 species, cannot be explained solely on the basis of phenotypic and chemotaxonomic methods, and virulence appears to be, at least in part, strain-dependent [14, 50, 51]. Indeed, this pathogen harbors a versatile genetic machinery that allows it to not only exhibit variable, strain-dependent virulence aspects but also rapidly generate environment-specific resistance and survival factors [14, 50,51,52]. As a result, it may be found in a broad range of habitats, including water, soil, food, and on the surfaces of medical equipment, and it may persist in environments that are inhospitable to many other bacterial pathogens. The most clinically relevant Acinetobacter spp. are grouped in the Acinetobacter calcoaceticus-baumannii (Acb) complex which includes 5 pathogenic Acinetobacter species, namely A. baumannii, Acinetobacter nosocomialis, Acinetobacter pittii, Acinetobacter seifertii, and Acinetobacter dijkshoorniae. A non-pathogenic one, Acinetobacter calcoaceticus also belongs to this group [14, 50,51,52]. A. baumannii exhibiting MDR profiles are encountered with increasing frequency [50, 52,53,54,55,56]. Resistance to last-resort antibiotics such as carbapenems and colistin has already been reported, allowing such strains to cause pan-drug-resistant infections that are presently impossible to eradicate [54,55,56,57,58,59,60,61]. A novel siderophore cephalosporin antibiotic, cefiderocol, was recently approved as a therapeutic agent for Gram-negative bacterial infections in both Europe and the USA [62]. As observed in P. aeruginosa, one of the innovative aspects of this antibiotic is that cefiderocol can cross the outer membrane, relying on a “Trojan horse” strategy. Cefiderocol creates chelating complexes with extracellular free iron, and thus, these complexes are transported into the bacteria via its iron transporters. Inside the cell, cefiderocol inhibits penicillin-binding protein 3, impairing cell wall synthesis [63,64,65]. Cefiderocol has proven to have potent activity against carbapenem-resistant A. baumannii [65,66,67]. However, cefiderocol-resistant A. baumannii isolates have already been reported [68,69,70,71]. It is estimated that approximately 44% of all A. baumannii isolates are MDR, with the highest incidence found in the Middle East and Latin America, where this rate may exceed 70%. In the European Region, the percentage of MDR A. baumannii can reach 43% [50, 72]. Results from the Central Asian and European Surveillance of Antimicrobial Resistance network and the European Antimicrobial Resistance Surveillance Network demonstrate an alarming increase in the reported cases of Acinetobacter spp., which have doubled (+121%), passing from 5375 cases reported for 2019 to 10885 cases reported for 2021. Regarding the percentage of carbapenem-resistant Acinetobacter spp., this value varied from below 1% to over 50% throughout the region. East and South Europe were the areas that showed the highest percentages of carbapenem-resistant Acinetobacter spp. Moreover, between 2017 and 2021, MDR Acinetobacter spp. increased from 32.1 to 36.8%. [73, 74]. As a result, A. baumannii has become a critical healthcare problem worldwide; this threat has received even greater attention due to the high prevalence of this bacterium in patients with COVID-19 [22, 23, 32].
A. baumannii is on the World Health Organization’s drug-resistant bacteria and antimicrobial resistance research priority list, and the Center for Diseases Control and Prevention has classified carbapenem-resistant Acinetobacter as an urgent threat to public health [75, 76]. The critical concerns around AMR in A. baumannii are compounded by the ability of this pathogen to form biofilms on biotic and abiotic surfaces. Within biofilms, bacteria exhibit limited metabolic activity, and their extracellular polysaccharide matrix shelters them from antibiotics and host immune factors. As a result, therapeutic options to treat MDR biofilm-forming A. baumannii have become ineffective.
A. baumannii biofilms
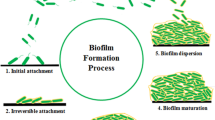
A. baumannii rapidly forms sessile biofilm aggregates upon adherence to its substrate. Biofilm communities produce their own extracellular polysaccharide matrix and, in turn, release planktonic (free-swimming) bacteria that may establish new biofilms elsewhere, a bacterial survival process that exists throughout nature [8, 12, 77, 78] (see Fig. 1). The extracellular matrix contains polysaccharides, proteins, and nucleic acids and confers to the biofilm key viscoelastic, cohesive, and hydrating properties [8, 79, 80]. This dynamic mode of growth allows to retain water, to shelter against environmental stress, and facilitates quorum sensing and horizontal gene transfer [8, 80,81,82,83].
The five stages in biofilm formation (A) and A. baumannii virulence factors associated with biofilm formation (B). (A) Biofilm formation includes five different stages: (1) initial attachment, when planktonic cells reversibly attach to a surface; (2) irreversible attachment, when bacteria irreversibly attach to a surface and start cell-to-cell adhesion; (3) growth and extracellular polymeric substances (EPS) production, with bacteria starting to produce EPS; (4) biofilm maturation and microcolony formation, when biofilms become fully mature and microcolonies start to appear; and (5) detachment and dispersal, when biofilm bacteria start being released from the biofilm to make biofilms elsewhere. (B) A. baumannii has several virulence factors associated with biofilm formation, including the Csu Pili, which is related to the adhesion of the bacteria to abiotic surfaces such as plastic; AbOmpA, a porin involved in the adhesion of the bacteria to biotic surfaces such as epithelial cells; Bap, a protein involved in the maturation of biofilms and also BfmRS, a regulatory system involved in biofilms formation through the regulation of Csu Pili
The formation of biofilms protects bacteria against antimicrobial substances, pH variability, UV radiation, extreme temperature, desiccation, nutrient starvation, and host immunity [81,82,83,84]. On abiotic surfaces, the biofilm mode of growth of A. baumannii protects against severe desiccation, which allows it to persist and may facilitate outbreaks [35, 36, 85,86,87]. Biofilms consume ambient nutrients, electron donors, and acceptors and may incorporate other bacterial species as well as dead and viable host cells [8, 78, 82, 83, 88,89,90]. The biofilm mode of growth may increase AMR by a given bacterial pathogen 100 to 1000-fold [87, 91,92,93]. A broad range of mechanisms cooperate to produce the elevated AMR observed in biofilms, including interactions of the antimicrobials with biofilm matrix elements, reduced bacterial growth rates, quorum sensing, and various drivers of antibiotic resistance that can fuel horizontal genetic transfer inside the biofilm [8, 90, 94,95,96,97,98]. Horizontal gene transfer is a phenomenon that occurs faster within biofilms than in planktonic cells. Inside biofilms, the bacterial evolution and development of drug-resistant bacteria can be achieved by the transfer of mobile genetic elements encoding antibiotic resistance genes, such as plasmids. This gene transfer can be fueled by the exposition to sub-minimum inhibitory concentrations (sub-MICs) of antibiotics inside the biofilm. Thus, biofilms are considered important reservoirs for the dissemination of AMR [95, 98, 99]. Several studies have established a positive correlation between biofilm formation and the degree of AMR in A. baumannii, with extensively drug-resistant (XDR) strains forming more robust biofilms than MDR strains [100,101,102]. Yet, other reports indicate that XDR strains tend to form weaker biofilms than non-MDR and MDR strains [93, 103], highlighting the urgent need to better understand co-regulatory mechanisms of biofilm formation and AMR.
Quorum sensing
Quorum sensing is a cell-to-cell communication system that regulates bacterial behavior in both Gram-positive and Gram-negative bacteria in response to environmental stress [104,105,106,107]. This process depends on the production, detection, and down-stream signaling of secreted chemical molecules (autoinducers). Using this communication system, bacteria are capable to respond to increased cell density, to control biofilm growth, and to produce extracellular polysaccharides. It also allows bacteria to regulate genes implicated in virulence and drug resistance, an area of research receiving considerable attention in view of its potential to develop new therapeutics [108, 109]. A number of compounds and processes are known to contribute to quorum sensing (summarized in Table 1).
In Gram-negative bacteria, quorum sensing compounds include N-acyl-homoserine lactones (AHL), composed of a homoserine lactone ring and a fatty acid acyl group variable in size, depending upon the bacterial species. Short (4 to 8 carbons) or long (10 to 16 carbons) AHL diffuses through the cell wall and acts as autoinducers [127,128,129,130]. Gram-positive bacteria use secreted oligopeptides as autoinducers and a two-component regulatory system (TCS) to regulate what genes and peptides need to be expressed. These TCS rely on membrane-bound histidine-kinase receptors and intracellular regulators [130, 131]. Furanosyl borate diester or tetrahydroxy furan (also called autoinducer-2, AI-2) is another type of signaling molecule used by both Gram-negative and Gram-positive bacteria and is known to serve as a signal for interspecies communications [132, 133]. Another autoinducer signal molecule involved in quorum sensing communication has been recently discovered in enterohemorrhagic Escherichia coli (EHEC) serotype O157:H7 and is referred to as autoinducer-3 (AI-3) [124]. AI-3 was characterized as 3,6-dimethylpyrazin-2-one [125]. EHEC bacteria sense AI-3 signaling molecules through the sensor kinase QseC belonging to the TCS QseBC (which is also involved in “sensing” of the host-derived hormones epinephrine and norepinephrine), thus enabling the control of expression of virulence genes, including those regulating the production of Shiga toxin [134].
Recent findings have significantly advanced our understanding of quorum sensing in regulating biofilm formation and AMR in both Gram-positive and Gram-negative bacteria, including in A. baumannii [50, 108,109,110, 135,136,137,138]. Recent evidence indicates that quorum sensing deficiency is associated with the formation of thinner biofilms that become susceptible to kanamycin [139]. In P. aeruginosa, quorum sensing regulates the expression of superoxide dismutase and catalase genes that confer resistance to hydrogen peroxide [140]. A three-day exposure of A. baumannii to sub-inhibitory concentrations of antibiotics increases biofilm formation and AMR in association with significant genome alterations linked to these phenotypic changes [141]. Pharmacological interference with the quorum sensing system reduces pathogenicity and facilitates the elimination of a given pathogen by host immunity [137, 142, 143]. These observations add to the ever-increasing number of reports supporting the hypothesis that biofilm formation and development of AMR are, at least in part, genetically co-regulated processes and make the inhibition of bacterial communication an attractive target for new drug development [98, 108, 109, 144,145,146]. Indeed, intensive ongoing research activities aim at developing quorum sensing inhibitors or quorum quenching enzymes to decrease the virulence of bacterial pathogens, including A. baumannii [138, 143, 147, 148].
Quorum sensing in A. baumannii is similar to what is observed in other Gram-negative bacteria, and it regulates biofilm formation, AMR, motility, and virulence [111, 112, 136, 137, 149,150,151]. AHL serves as the autoinducer-1. It is produced via the autoinducer synthase AbaI, which in turn binds to its cognate receptor AbaR (see Fig. 2), a system that is homologous to the canonical LuxI/LuxR system found in other Gram-negative bacteria [111, 149, 150]. AbaI-produced AHL binding to AbaR triggers a cascade of reactions leading to the quorum sensing response. A. baumannii AbaI is 27.5% identical and 47.3% similar to the LasI of a pathogenic and an environmental isolate of P. aeruginosa, respectively, and has a close resemblance to the LuxI family members described for Vibrio fischeri [110, 111, 138, 149]. A. baumannii also appears to express a luxI gene that contributes to AHL production [152]. In this bacterium, acyl side chains range from 10 to 16 carbons, although a number of Acinetobacter strains show various AHL profiles and produce more than one AHL [149, 153,154,155]. The most common AHL in A. baumannii is hydroxy-C12-homoserine lactone [110, 112, 155]. The quorum sensing system AbaI/AbaR is known to contribute to biofilm formation and antibiotic resistance in A. baumannii [110, 112]. Exogenous supplementation of AHL stimulates the formation of biofilms in non-biofilm-forming A. baumannii clinical isolates and enhances biofilm production in weakly adherent bacteria [156]. AHL also plays a role in A. baumannii drug resistance; an abaI deficient A. baumannii mutant was more susceptible to meropenem and piperacillin than the wild-type strain. However, its resistance was restored in the presence of AHL supplementation, promoting the expression of several drug resistance-related genes that include blaOXA-51, ampC, and the efflux pumps adeA and adeB genes [112]. Indeed, the AbaI/AbaR-dependent biofilm formation has been linked to overexpression of antimicrobial resistance genes, including those controlling efflux pumps [113, 114]. Yet, the molecular pathways regulating the complex AbaI/AbaR system remain incompletely understood.
AbaI/AbaR quorum sensing system in A. baumannii. In this system, S-adenosyl methionine (SAM) and acyl-acyl carrier protein substrates (Acyl ACP) are the essential components for the production of AHLs. AbaI-regulated AHLs are released into the extracellular environment. Then, AHLs bind to the receptor AbaR present in the other cells, triggering a cascade of reactions that culminates in the control of expression of several target genes, including genes involved in biofilm formation and also for the production of more AHLs
Biofilm-associated genes in A. baumannii
In A. baumannii, several genes are involved in biofilm formation and development. Some of these determinants and related functions are presented in Table 2.
Adherence is the critical step for the development of a biofilm. Thus, the factors that play a role in this first stage of biofilm formation, including Csu pili, AbOmpA, A. baumannii Bap, and the TCS BfmRS, will be discussed in the following paragraphs. Other genes that may also be involved in the co-regulation of biofilm formation in A. baumannii including blaPER-1, epsA, and ptk will be addressed.
Csu pili
The attachment to a surface by planktonic bacteria is the first step in bacterial biofilm formation [170]. A. baumannii Csu pili contribute to the attachment and biofilm formation on abiotic surfaces [86]. In A. baumannii, Csu pili are formed by the type I chaperone-usher pili assembly system named CsuA/BABCDE. In these bacteria, the csu operon has six open reading frames, namely csu A/B, A, B, C, D, and E [86]. At the structure level, Csu pili are elaborated from four subunit proteins, CsuA/B, CsuA, CsuB, and CsuE, being assembled by CsuC and CsuD proteins [33, 86]. A positive correlation between biofilm formation and AMR, predominantly regulated by csuE, was reported recently. Concurrently, a high prevalence of csuE was observed among MDR A. baumannii clinical isolates, reaching 100% in some reports [56, 171]. At the Csu pili tip, CsuE is made of three hydrophobic finger-like structures responsible for the adherence of the bacteria to abiotic surfaces, such as hydrophobic plastics [33]. As a result, these bacteria exhibit a greater ability to form biofilms on polycarbonate hydrophobic materials than on glass, which is a hydrophilic material. The use of hydrophilic materials for medical devices may help prevent biofilm-associated infections [33, 35]. Sub-inhibitory concentrations of antibiotics like trimethoprim and sulfamethoxazole can inhibit A. baumannii ATCC 17978 biofilm formation. Through the inhibition of folate biosynthesis and then promoting folate stress, these antibiotics repress csuA/B expression, which, in turn, inhibits the development of biofilms [172]. A. baumannii Csu pili are not implicated in adherence to biotic surfaces, such as respiratory tract epithelial cells [173].
A. baumannii outer membrane protein A (AbOmpA)
The OmpA of A. baumannii, also known as OmpAb [174], Omp38 [175, 176], or AbOmpA [177] (hereafter AbOmpA), is a permeability-relevant porin protein present in the outer membrane (OM) [174]. AbOmpA is considered as one of the major proteins in the OM of A. baumannii, and it is involved in specific processes of pathogenesis, such as adherence to and invasion of the host epithelial cells [157, 158, 174], disruption of the mitochondria, and cell death [176, 177]. AbOmpA is also related to the virulence capacity of outer membrane vesicles released from A. baumannii [178, 179]. AbOmpA may also induce dendritic cell death that can lead to a deficient host immune response [177]. In contrast, other studies have shown that AbOmpA stimulates the host immune system, activating and maturing dendritic cells and promoting the differentiation of CD4+ T cells [180]. Overexpression of AbOmpA presents a risk factor for the development of A. baumannii pneumonia and bacteremia, dissemination to organs like the lungs and spleen, and mortality [179, 181].
AbOmpA plays a role in A. baumannii biofilm formation in abiotic surfaces like polystyrene, and it is essential for the adherence of the bacteria to several biotic surfaces, such as human alveolar epithelial cells [158]. AbOmpA is also involved in intrinsic resistance to some antimicrobials. An abompA mutant A. baumannii ATCC 17978 strain showed increased susceptibility to trimethoprim than the wild-type strain. This increase in susceptibility may reflect the interaction of the OmpA-like domain of the AbOmpA with the inner membrane transporters of the resistance–nodulation–division efflux pumps superfamily [182].
A. baumannii biofilm-associated protein (Bap)
Biofilm-associated proteins (Bap) have been identified in both Gram-positive and Gram-negative bacteria. These proteins share common structural and functional characteristics. They are bacterial surface proteins with a high molecular weight, possess a core domain of tandem repeats, and are involved in biofilm development [183].
First identified in S. aureus in 2001 [184], Bap was shown to be a cell wall protein associated with S. aureus primary attachment to abiotic surfaces, such as polystyrene, cell-to-cell adhesion, and, thus, biofilm formation. A S. aureus Bap homologous protein was then discovered in A. baumannii [159].
The A. baumannii Bap protein is involved in biofilm formation and maturation and participates in intercellular adhesion [159]. Bap protein plays an important role in bacterial adherence to human bronchial epithelial cells and human neonatal keratinocytes [160]. Also, Bap expression in A. baumannii is related to mature biofilm formation on abiotic surfaces, such as polypropylene and titanium. This protein is involved in the maintenance of cell surface hydrophobicity [160], a feature that is considered an important factor in adherence ability and biofilm formation in a variety of bacteria [185,186,187].
Many clinical A. baumannii strains have the bap gene [188, 189]. Bacteria that express the bap gene produce stronger biofilms, and the addition of affinity-purified Bap-specific antibodies inhibits biofilm formation [188]. In contrast, Bap-negative isolates recovered from bloodstream infections show a low biofilm formation ability [189]. The bap gene is often present in MDR A. baumannii strains, providing support to the hypothesis that this gene may help co-regulate biofilm formation and AMR in A. baumannii [190].
The two-component regulatory system BfmRS in A. baumannii
TCSs are involved in the sensing and transduction of extracellular stimuli and are prototypically composed of a histidine kinase sensor and a response regulator. Histidine kinase senses a stimulus and sends the information to the response regulator by transferring a phosphoryl group, promoting conformational changes in the regulatory domain. In turn, the response regulator acts as an activator or a repressor for the specific gene’s transcription [191].
TCSs have been found in both Gram-positive and Gram-negative bacteria [161, 191, 192]. They play a role in several bacterial functions, including in the control of genes regulating efflux pumps [162, 193, 194] and biofilm formation [161, 195]. In A. baumannii, the TCS BfmRS regulates biofilm formation [161], capsular polysaccharide biosynthesis, as well as osmotic and oxidative stress responses. BfmRS may also modulate A. baumannii motility by controlling the expression of type IV pili [196, 197].
The BfmRS in A. baumannii is a TCS composed of a cytoplasmic response regulator, the BfmR, encoded by the bfmR gene, and a cytoplasmic membrane sensor kinase, the BfmS, which is encoded by the bfmS gene. The cytoplasmic protein BfmR is involved in biofilm development, cell morphology, and pellicle formation [161, 198]. BfmR is essential for attachment and biofilm formation in abiotic surfaces, such as polystyrene, as shown through a mutation in the bfmR gene. As a transcriptional activator, BfmR is involved in the regulation of Csu pili expression and, thus, adherence and biofilm development [161, 197]. Additionally, another TCS is also involved in the regulation of Csu pili, the TCS GacSC, and thus implicated in biofilm formation [199]. A. baumannii clinical isolates often express bfmS in association with elevated biofilm formation [56].
The BfmRS TCS is also involved in AMR. By mechanisms that are not yet fully understood, the response regulator BfmR is considered an important factor in AMR, including resistance to meropenem and polymyxin E concurrently with biofilm formation. BfmR is also involved in resistance to complement [200, 201]. Moreover, BfmR also participates in the regulation of A. baumannii tolerance to desiccation, explaining its persistence in hospital settings [202]. The sensor kinase BfmS is involved in A. baumannii biofilm formation. Mutation of bfmS also interferes with AbOmpA regulatory pathways and virulence [203, 204].
The A. baumannii BfmRS system is strongly associated with virulence and in the transcriptional regulation of the k locus, which is responsible for exopolysaccharide production in the presence of sub-MICs of antibiotics such as chloramphenicol and erythromycin [205, 206]. Furthermore, BfmRS is involved in resistance against β-lactams, both β-lactamase-dependent and β-lactamase-independent manner. In the presence of β-lactams aggression, BfmRS can regulate the production of β-lactamases and also control the cell division and cell wall degradation processes [207]. The fact that BfmRS is involved in AMR as well as in the regulation of biofilm formation through the control of Csu pili points to a role for this gene in the co-regulation of biofilm formation and AMR in A. baumannii [207].
A. baumannii bla PER-1, epsA, and ptk genes
Other genes that may be involved in the co-regulation of biofilm formation and AMR in A. baumannii are blaPER-1, epsA, and ptk.
The blaPER-1 gene encodes for an extended-spectrum β-lactamase, the PER-1 [166,167,168]. A. baumannii that express blaPER-1 are associated with poor clinical outcome [167]. This gene appears to be essential in the adhesion capacity of MDR A. baumannii clinical isolates to respiratory epithelial cells and was found to contribute to biofilm formation on abiotic surfaces [208]. More research needs to assess whether this property may at least in part contribute to the poor outcome of patients infected with A. baumannii carrying the blaPER-1 gene. The prevalence of this gene in MDR A. baumannii clinical isolates has been extensively evaluated: Studies have reported a prevalence of 2% [56], 44% [189], 53% [102], and 64.2% [209]. Remarkably, in one study, out of 27 MDR A. baumannii clinical isolates, 44% had the blaPER-1 gene. These blaPER-1 positive isolates were recovered from the respiratory tract of patients. In contrast, blaPER-1 negative isolates were recovered from the bloodstream and were low biofilm formers [189], strongly suggesting a relationship between the expression of PER-1 and biofilm formation ability.
The capsule is a well-known virulence factor [210]. epsA and ptk are two genes involved in capsule biosynthesis in A. baumannii. A. baumannii epsA encodes for a putative polysaccharide export outer membrane protein (EpsA), whereas the ptk gene encodes for a putative protein tyrosine kinase (PTK) [165]. Both genes appear to be implicated in the biofilm formation of MDR A. baumannii isolates since a study involving 100 MDR A. baumannii clinical isolates demonstrated a 95% prevalence for both ptk and epsA [56], and all were strong biofilm formers. The presence of the capsule may contribute to a strong biofilm.
Paths towards novel therapeutics
The exponential increase in antibiotic resistance along with the growing awareness of the deleterious impact of biofilms in medicine has led to a growing demand for new therapeutic alternatives. Several alternatives are currently under development, with the objective of combating AMR as well as the biofilms formation in A. baumannii. These include the development of new vaccines, the inhibition of quorum sensing, the use of nanoparticles and metal ions, natural products, antimicrobial peptides, and phage therapy.
Vaccine development
The fight against AMR needs to be multifactorial, including stewardship of antimicrobials use, development of new therapeutics, and infection prevention. Vaccines targeting AMR and biofilm formation are being developed [211, 212]. A number of these aim at mounting immunity against structures of the cell wall that are involved, directly or indirectly, with biofilm formation. For A. baumannii, these include AbOmpA, Bap protein, and subunit proteins of the surface-exposed Csu pili such as CsuA/B [213,214,215,216,217,218]. AbOmpA appears to be the most relevant for future vaccine development against drug-resistant A. baumannii. Along with the fact that AbOmpA-antigen-based immunization demonstrates high protection and survival rates reaching 80% or higher [219, 220], AbOmpA is also highly conserved among A. baumannii clinical isolates, with prevalence reaching 99% [215, 216, 219]. A two-recombinant-pilus proteins vaccine targeting the CsuA/B protein plus a related fimbriae protein, FimA, was found to trigger protective immunity in mice against A. baumannii infection [217]. A Bap protein vaccine is similarly protective in murine models [218].
Quorum sensing interference
Molecules that promote cell-to-cell communication, such as quorum sensing compounds (autoinducers), are attractive targets to reduce/inhibit biofilm formation. Quorum quenching inhibits bacterial cell communication. This can be accomplished by inhibiting the production or bacterial detection of these quorum sensing molecules or by their degradation [221]. Quorum quenching includes quorum quenching enzymes and quorum sensing inhibitors. Quorum quenching enzymes act in the extracellular environment and not intracellularly and hence will not select for resistant bacteria [221, 222].
Strategies of interference in quorum sensing are being developed in an attempt to prevent the formation of biofilms and the development of virulence factors as well as AMR in A. baumannii [136, 138, 223, 224]. As AHL lactonases destabilize quorum sensing in Gram-negative bacteria, these enzymes represent a promising approach to combat biofilm-associated infections and AMR [225, 226]. The utility of these enzymes as a therapeutic strategy against A. baumannii biofilms has been established using the clinical isolate A. baumannii S1 [223]. Similar results were observed with another lactonase enzyme with quorum quenching activity, MomL. This lactonase has the capability to degrade A. baumannii AHL and to dramatically reduce biofilm biomass while concurrently enhancing the susceptibility of the biofilm to antibiotics. Supplementation of antimicrobials with quorum quenching agents may offer synergistic benefits, as recently observed in A. baumannii biofilms exposed to tobramycin supplemented with MomL [224]. More research is warranted using this approach in vivo [224].
Nanoparticles and metal ions
Various metal nanoparticles, particularly silver, are the topic of intense research activity in view of their antimicrobial properties [227]. The antimicrobial properties of metal nanoparticles have been evaluated both in planktonic bacterial cells and biofilms of different Gram-positive and Gram-negative bacterial species, including A. baumannii, demonstrating inhibition of biofilm formation and increased antimicrobial sensitivity [228,229,230,231,232,233].
A number of studies have evaluated the benefits of combining nanoparticles with other compounds to inhibit the formation of susceptible and MDR A. baumannii biofilms, some with promising outcomes. These findings are summarized in Table 3.
Silver nanoparticles inhibit DNA synthesis and induce apoptosis-like in MDR A. baumannii in a concentration-dependent fashion [238]. Silver nanoparticles derived from silver salts can inhibit biofilm formation of MDR A. baumannii strains, at least in part by inhibiting the expression of biofilm-related genes, such as csuA/B, abompA, and bap [230].
Antimicrobial and antibiofilm effects of metal ions solutions have been studied in both Gram-positive and Gram-negative bacteria [239, 240]. Antimicrobial effects have been reported against both planktonic and biofilm A. baumannii upon exposure to single and to combinations of metal ions solutions [239]. The antibacterial properties of metal ions are mediated by a variety of mechanisms including interference with bacterial cell membranes, generation of reactive oxygen species (ROS), protein destabilization, and DNA injury [241]. However, both chromosomal and plasmid-mediated resistance of bacteria to silver have been reported, as well as a co-selection with the use of antibiotics when the resistance genes rely on the same genetic platform, such as a plasmid-mediated silver efflux system, the Sil system [242]. More research is warranted to develop effective metal nanoparticle-based strategies to fight biofilm formation and AMR in MDR A. baumannii.
Natural products
A few natural products under study have also demonstrated a high antibacterial and antibiofilm activity against MDR A. baumannii clinical isolates. These include cinnamaldehyde [243], essential oils [244, 245], and polyphenolic compounds [246]. Cinnamaldehyde has exhibited potent antibacterial and antibiofilm activity against carbapenem-resistant A. baumannii clinical isolates. The minimum bactericidal concentration of cinnamaldehyde on strong biofilm formers was 1.75 mg/mL. At ½ of the MIC, cinnamaldehyde inhibited biofilm formation by approximately 72% [243]. Essential oils can also achieve high antibacterial and antibiofilm activities against MDR A. baumannii clinical isolates. In nanoemulsion or as a pure compound, Thymus daenensis oils have achieved MICs of 45 μg/mL and 87.5 μg/mL, respectively. At ½ of the MIC, emulsions achieved 56.43% of inhibition of MDR A. baumannii clinical isolates biofilms [245]. Polyphenolic compounds have also been studied, presenting significant results, such as lower MICs and biofilm inhibition that can reach 90%, depending on the compound and isolate [246]. More research is warranted to identify the potential use of natural products and their mode of action in the dual control of biofilm formation and AMR in A. baumannii.
Antimicrobial peptides
The study of antimicrobial peptides has enabled the identification of multiple compounds with relevant antibacterial and antibiofilm activity against MDR A. baumannii clinical isolates [247,248,249]. Magainin 2, isolated from the frog Xenopus laevis [247], and octominin, obtained from Octopus minor [248], have significant antibacterial and antibiofilm activity against MDR A. baumannii clinical isolates. At 4 μM, magainin 2 promoted biofilm inhibition in the resistant strains and a biofilm reduction of 66.2% at 256 μM. Octominin at 5 μg/mL promoted a biofilm inhibition of 61.59% and a biofilm reduction of 35.62%. Both magainin 2 and octominin were demonstrated to affect the A. baumannii cell membrane. Magainin 2 destabilizes both A. baumannii inner and outer membranes, whereas octominin permeabilizes the A. baumannii cell membrane, thus leading to bacterial cell death. Research needs to further characterize modes of action as well as identify potential off-target effects in infected patients.
Phage therapy
Bacteriophages (also known as phages) are viruses that infect bacteria and, thus, have a promising potential to be used as an antibacterial therapy [250, 251]. The application of phage therapy as an antibacterial and antibiofilm strategy in MDR A. baumannii clinical isolates has yielded positive results, even in combination with other phages (cocktails) or antibiotics [251, 252].
Studies have assessed the antibacterial and antibiofilm activity of the phages AB7-IBB1 [253] and AB7-IBB2 [254] in MDR A. baumannii clinical isolates. With a multiplicity of infection (MOI) of 0.1 applied in 108 CFU/well, the phage AB7-IBB1 inhibited biofilm formation on both abiotic and biotic surfaces in 40% and 50%, respectively. It also promoted more than 35% biofilm removal with a MOI of 10 in 106 CFU/well. Similarly, phage AB7-IBB2 at a MOI of 0.1 in 108 CFU/well, inhibited biofilm formation and disrupted preformed biofilms on abiotic surfaces in approximately 40%. Moreover, phages AB7-IBB1 and AB7-IBB2 were able to inhibit A. baumannii growth up to 46% and 70% with a MOI of 0.1 in 108 CFU/well, respectively.
Phage therapy has proven to be an effective therapeutic option against MDR A. baumannii clinical isolates. However, A. baumannii can develop phage resistance in a very short period of time. Phage resistance in A. baumannii is related to mutations in k locus genes that culminate in relevant alterations on the capsule; the principal factor in with phage depends on adsorption [255, 256].
Conclusion
The combination of AMR and biofilm formation makes A. baumannii a formidable enemy in healthcare settings, where it can cause a wide range of infections, including pneumonia, bloodstream, wound infections, and urinary tract infections. Resistance to last-resort antibiotics such as carbapenems and colistin has already been reported, allowing such strains to cause pan-drug-resistant infections that are presently impossible to eradicate. To address this global health threat, there is a need for improved surveillance, infection control measures, and the development of new antimicrobial agents and treatment strategies. A. baumannii biofilms provide a protective environment for the bacteria and further enhance their resistance to antimicrobial agents. Biofilms are also associated with the persistence of infections and the development of chronic diseases. A better understanding of the mechanisms that co-regulate biofilm formation and AMR will help identify new therapeutic targets. Key genes that are involved in biofilm formation, for example, the quorum sensing system AbaI/AbaR and the TCS BfmRS, are also implicated in the development of AMR. Many MDR clinical isolates contain genes like abompA and csuE, which are essential for biofilm genesis. New therapeutic strategies concurrently targeting these two phenomena include quorum sensing interference, developments of vaccines mostly targeting AbOmpA, the use of nanoparticles and metal ions, natural products, antimicrobial peptides, and phage therapy. All have shown promising results. However, the use of silver nanoparticles appears to come with a risk of A. baumannii developing plasmid-mediated resistance, highlighting the critical need for more research. Combined approaches that may confer synergistic benefits offer intriguing avenues towards new, effective, and safe therapies.
References
Davies J, Davies D (2010) Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev 74:417–433. https://doi.org/10.1128/mmbr.00016-10
WHO, FAO, OIE, UNEP (2021) Antimicrobial resistance and the united nations sustainable development cooperation framework. World Health Organization, Geneva, Switzerland
Larsson DGJ, Flach C-F (2021) Antibiotic resistance in the environment. Nat Rev Microbiol 20:257–269. https://doi.org/10.1038/s41579-021-00649-x
Andersson DI, Balaban NQ, Baquero F et al (2021) Antibiotic resistance: turning evolutionary principles into clinical reality. FEMS Microbiol Rev 44:171–188. https://doi.org/10.1093/femsre/fuaa001
Mackenzie JS, Jeggo M (2019) The one health approach—why is it so important? Trop Med Infect Dis 4:5–8. https://doi.org/10.3390/tropicalmed4020088
McCubbin KD, Anholt RM, de Jong E et al (2021) Knowledge gaps in the understanding of antimicrobial resistance in Canada. Front Public Heal 9:1–14. https://doi.org/10.3389/fpubh.2021.726484
World Bank Group (2017) Drug-resistant infections: a threat to our economic future. World Bank Group, Washington, DC
Lockhart JS, Buret AG, Morck DW (2020) Biofilm and biofilm control. In: McDonnel G (ed) Disinfection, Sterilization and Preservation, 6th edn. Lippincott Williams & Wilkins (LWW), Pennsylvania, USA, pp 1320–1336
Liu C, Sun D, Zhu J et al (2020) The regulation of bacterial biofilm formation by cAMP-CRP: a mini-review. Front Microbiol 11:1–7. https://doi.org/10.3389/fmicb.2020.00802
Wolska KI, Grudniak AM, Rudnicka Z, Markowska K (2016) Genetic control of bacterial biofilms. J Appl Genet 57:225–238. https://doi.org/10.1007/s13353-015-0309-2
Ha D-G, O’Toole GA (2015) c-di-GMP and its effects on biofilm formation and dispersion: a Pseudomonas aeruginosa review. Microbiol Spectr 3:1–20. https://doi.org/10.1128/microbiolspec.mb-0003-2014
Wong GCL, Antani JD, Lele PP et al (2021) Roadmap on emerging concepts in the physical biology of bacterial biofilms: from surface sensing to community formation. Phys Biol 18:1–49. https://doi.org/10.1088/1478-3975/abdc0e
Alsan M, Klompas M (2010) Acinetobacter baumannii: an emerging and important pathogen. J Clin Outcomes Manag 17:363–369
McConnell MJ, Actis L, Pachón J (2013) Acinetobacter baumannii: human infections, factors contributing to pathogenesis and animal models. FEMS Microbiol Rev 37:130–155. https://doi.org/10.1111/j.1574-6976.2012.00344.x
Peleg AY, Seifert H, Paterson DL (2008) Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev 21:538–582. https://doi.org/10.1128/cmr.00058-07
Lai C-C, Wang C-Y, Hsueh P-R (2020) Co-infections among patients with COVID-19: the need for combination therapy with non-anti-SARS-CoV-2 agents? Ann Thorac Surg 53:505–512. https://doi.org/10.1016/j.jmii.2020.05.013
Khurana S, Singh P, Sharad N et al (2021) Profile of co-infections & secondary infections in COVID-19 patients at a dedicated COVID-19 facility of a tertiary care Indian hospital: implication on antimicrobial resistance. Indian J Med Microbiol 39:147–153. https://doi.org/10.1016/j.ijmmb.2020.10.014
Sreenath K, Batra P, Vinayaraj EV et al (2021) Coinfections with other respiratory pathogens among patients with COVID-19. Microbiol Spectr 9:1–13. https://doi.org/10.1128/Spectrum.00163-21
Abdollahi A, Aliramezani A, Salehi M et al (2021) Co-infection of ST2IP carbapenem-resistant Acinetobacter baumannii with SARS-CoV-2 in the patients admitted to a Tehran tertiary referral hospital. BMC Infect Dis 21:1–7. https://doi.org/10.1186/s12879-021-06642-2
Apisarnthanarak A, Weber DJ (2021) Strategy to limit multidrug-resistant Acinetobacter baumannii transmission in a cohort coronavirus disease 2019 (COVID-19) critical care unit. Infect Control Hosp Epidemiol 43:1517–1518. https://doi.org/10.1017/ice.2021.289
Gottesman T, Fedorowsky R, Yerushalmi R et al (2021) An outbreak of carbapenem-resistant Acinetobacter baumannii in a COVID-19 dedicated hospital. Infect Prev Pract 3:1–5. https://doi.org/10.1016/j.infpip.2021.100113
Russo A, Gavaruzzi F, Ceccarelli G et al (2022) Multidrug-resistant Acinetobacter baumannii infections in COVID-19 patients hospitalized in intensive care unit. Infection 50:83–92. https://doi.org/10.1007/s15010-021-01643-4
Rangel K, Chagas TPG, De-Simone SG (2021) Acinetobacter baumannii infections in times of COVID-19 pandemic. Pathogens 10:1–13. https://doi.org/10.3390/pathogens10081006
Boral J, Genç Z, Pınarlık F et al (2022) The association between Acinetobacter baumannii infections and the COVID-19 pandemic in an intensive care unit. Sci Rep 12:1–7. https://doi.org/10.1038/s41598-022-25493-8
Ceparano M, Baccolini V, Migliara G et al (2022) Acinetobacter baumannii isolates from COVID-19 patients in a hospital intensive care unit: molecular typing and risk factors. Microorganisms 10:1–13. https://doi.org/10.3390/microorganisms10040722
Mumcuoğlu İ, Çağlar H, Erdem D et al (2022) Secondary bacterial infections of the respiratory tract in COVID-19 patients. J Infect Dev Ctries 16:1131–1137. https://doi.org/10.3855/jidc.16724
Pourajam S, Kalantari E, Talebzadeh H et al (2022) Secondary bacterial infection and clinical characteristics in patients with COVID-19 admitted to two intensive care units of an academic hospital in Iran during the first wave of the pandemic. Front Cell Infect Microbiol 12:1–9. https://doi.org/10.3389/fcimb.2022.784130
Montrucchio G, Corcione S, Lupia T et al (2022) The burden of carbapenem-resistant Acinetobacter baumannii in ICU COVID-19 patients: a regional experience. J Clin Med 11:1–11. https://doi.org/10.3390/jcm11175208
Adelman MW, Bhamidipati DR, Hernandez-Romieu AC et al (2021) Secondary bacterial pneumonias and bloodstream infections in patients hospitalized with COVID-19. Ann Am Thorac Soc 18:1584–1587. https://doi.org/10.1513/AnnalsATS.202009-1093RL
Wong SC, Lam GKM, Chen JHK et al (2021) Air dispersal of multidrug-resistant Acinetobacter baumannii: implications for nosocomial transmission during the COVID-19 pandemic. J Hosp Infect 116:78–86. https://doi.org/10.1016/j.jhin.2021.08.005
Gaibani P, Viciani E, Bartoletti M et al (2021) The lower respiratory tract microbiome of critically ill patients with COVID-19. Sci Rep 11:1–11. https://doi.org/10.1038/s41598-021-89516-6
Kariyawasam RM, Julien DA, Jelinski DC et al (2022) Antimicrobial resistance (AMR) in COVID-19 patients: a systematic review and meta-analysis (November 2019–June 2021). Antimicrob Resist Infect Control 11:1–18. https://doi.org/10.1186/s13756-022-01085-z
Pakharukova N, Tuittila M, Paavilainen S et al (2018) Structural basis for Acinetobacter baumannii biofilm formation. Proc Natl Acad Sci U S A 115:5558–5563. https://doi.org/10.1073/pnas.1800961115
Espinal P, Martí S, Vila J (2012) Effect of biofilm formation on the survival of Acinetobacter baumannii on dry surfaces. J Hosp Infect 80:56–60. https://doi.org/10.1016/j.jhin.2011.08.013
Greene C, Wu J, Rickard AH, Xi C (2016) Evaluation of the ability of Acinetobacter baumannii to form biofilms on six different biomedical relevant surfaces. Lett Appl Microbiol 63:233–239. https://doi.org/10.1111/lam.12627
Gedefie A, Demsis W, Ashagrie M et al (2021) Acinetobacter baumannii biofilm formation and its role in disease pathogenesis: a review. Infect Drug Resist 14:3711–3719. https://doi.org/10.2147/IDR.S332051
Pour NK, Dusane DH, Dhakephalkar PK et al (2011) Biofilm formation by Acinetobacter baumannii strains isolated from urinary tract infection and urinary catheters. FEMS Immunol Med Microbiol 62:328–338. https://doi.org/10.1111/j.1574-695X.2011.00818.x
Dijkshoorn L, Nemec A, Seifert H (2007) An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat Rev Microbiol 5:939–951. https://doi.org/10.1038/nrmicro1789
Gaddy JA, Actis LA (2009) Regulation of A. baumannii biofilm formation. Future Microbiol 4:273–278. https://doi.org/10.2217/fmb.09.5
Yang S, Sun J, Wu X, Zhang L (2018) Determinants of mortality in patients with nosocomial Acinetobacter baumannii bacteremia in Southwest China: a five-year case-control study. Can J Infect Dis Med Microbiol 2018:1–9. https://doi.org/10.1155/2018/3150965
Patel RV, Shah JS, Revathi G et al (2019) Acinetobacter infections: a retrospective study to determine inhospital mortality rate and clinical factors associated with mortality. Infect Prev Pract 1:1–5. https://doi.org/10.1016/j.infpip.2019.100010
Cornejo-Juárez P, Cevallos MA, Castro-Jaimes S et al (2020) High mortality in an outbreak of multidrug resistant Acinetobacter baumannii infection introduced to an oncological hospital by a patient transferred from a general hospital. PLoS One 15:1–14. https://doi.org/10.1371/journal.pone.0234684
Falagas ME, Rafailidis PI (2007) Attributable mortality of Acinetobacter baumannii: no longer a controversial issue. Crit Care 11:11–13. https://doi.org/10.1186/cc5911
Jamulitrat S, Arunpan P, Phainuphong P (2009) Attributable mortality of imipenem-resistant nosocomial Acinetobacter baumannii bloodstream infection. J Med Assoc Thai 92:413–419
Dexter C, Murray GL, Paulsen IT, Peleg AY (2015) Community-acquired Acinetobacter baumannii: clinical characteristics, epidemiology and pathogenesis. Expert Rev Anti Infect Ther 13:567–573. https://doi.org/10.1586/14787210.2015.1025055
Chen MZ, Hsueh PR, Lee LN et al (2001) Severe community-acquired pneumonia due to Acinetobacter baumannii. Chest 120:1072–1077. https://doi.org/10.1378/chest.120.4.1072
Anstey NM, Currie BJ, Withnall KM (1992) Community-acquired Acinetobacter pneumonia in the northern territory of Australia. Clin Infect Dis 14:83–91. https://doi.org/10.1093/clinids/14.1.83
Falagas ME, Karveli EA, Kelesidis I, Kelesidis T (2007) Community-acquired Acinetobacter infections. Eur J Clin Microbiol Infect Dis 26:857–868. https://doi.org/10.1007/s10096-007-0365-6
Wang JT, McDonald LC, Chang SC, Ho M (2002) Community-acquired Acinetobacter baumannii bacteremia in adult patients in Taiwan. J Clin Microbiol 40:1526–1529. https://doi.org/10.1128/JCM.40.4.1526-1529.2002
Harding CM, Hennon SW, Feldman MF (2018) Uncovering the mechanisms of Acinetobacter baumannii virulence. Nat Rev Microbiol 16:91–102. https://doi.org/10.1038/nrmicro.2017.148
Lee C-R, Lee JH, Park M et al (2017) Biology of Acinetobacter baumannii: pathogenesis, antibiotic resistance mechanisms, and prospective treatment options. Front Cell Infect Microbiol 7:1–35. https://doi.org/10.3389/fcimb.2017.00055
Monem S, Furmanek-Blaszk B, Łupkowska A et al (2020) Mechanisms protecting Acinetobacter baumannii against multiple stresses triggered by the host immune response, antibiotics and outside-host environment. Int J Mol Sci 21:1–30. https://doi.org/10.3390/ijms21155498
Consales G, Gramigni E, Zamidei L et al (2011) A multidrug-resistant Acinetobacter baumannii outbreak in intensive care unit: antimicrobial and organizational strategies. J Crit Care 26:453–459. https://doi.org/10.1016/j.jcrc.2010.12.016
Göttig S, Gruber TM, Higgins PG et al (2014) Detection of pan drug-resistant Acinetobacter baumannii in Germany. J Antimicrob Chemother 69:2578–2579. https://doi.org/10.1093/jac/dku170
Chusri S, Chongsuvivatwong V, Silpapojakul K et al (2019) Clinical characteristics and outcomes of community and hospital-acquired Acinetobacter baumannii bacteremia. J Microbiol Immunol Infect 52:796–806. https://doi.org/10.1016/j.jmii.2019.03.004
Zeighami H, Valadkhani F, Shapouri R et al (2019) Virulence characteristics of multidrug resistant biofilm forming Acinetobacter baumannii isolated from intensive care unit patients. BMC Infect Dis 19:1–9. https://doi.org/10.1186/s12879-019-4272-0
Antunes LCS, Visca P, Towner KJ (2014) Acinetobacter baumannii: evolution of a global pathogen. Pathog Dis 71:292–301. https://doi.org/10.1111/2049-632X.12125
Karakonstantis S, Gikas A, Astrinaki E, Kritsotakis EI (2020) Excess mortality due to pandrug-resistant Acinetobacter baumannii infections in hospitalized patients. J Hosp Infect 106:447–453. https://doi.org/10.1016/j.jhin.2020.09.009
Sobouti B, Mirshekar M, Fallah S et al (2020) Pan drug-resistant Acinetobacter baumannii causing nosocomial infections among burnt children. Med J Islam Repub Iran 34:1–4. https://doi.org/10.34171/mjiri.34.24
Valencia R, Arroyo LA, Conde M et al (2009) Nosocomial outbreak of infection with pan–drug-resistant Acinetobacter baumannii in a tertiary care university hospital. Infect Control Hosp Epidemiol 30:257–263. https://doi.org/10.1086/595977
Magiorakos AP, Srinivasan A, Carey RB et al (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18:268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
Takemura M, Wise MG, Hackel MA et al (2023) In vitro activity of cefiderocol against MBL-producing Gram-negative bacteria collected in North America and Europe in five consecutive annual multinational SIDERO-WT surveillance studies (2014–2019). J Antimicrob Chemother 78:2019–2027. https://doi.org/10.1093/jac/dkad200
Ito A, Nishikawa T, Yamano Y et al (2016) Siderophore cephalosporin cefiderocol utilizes ferric iron transporter systems for antibacterial activity against Pseudomonas aeruginosa. Antimicrob Agents Chemother 60:7396–7401. https://doi.org/10.1128/AAC.01405-16
Ito-Horiyama T, Ishii Y, Ito A et al (2016) Stability of novel siderophore cephalosporin S-649266 against clinically relevant carbapenemases. Antimicrob Agents Chemother 60:4384–4386. https://doi.org/10.1128/AAC.03098-15
Ito A, Sato T, Merime Ota MT et al (2018) In vitro antibacterial properties of cefiderocol, a novel siderophore cephalosporin, against Gram-negative bacteria. Antimicrob Agents Chemother 62:1–11. https://doi.org/10.1128/AAC.01454-17
Hackel MA, Tsuji M, Yamano Y et al (2017) In vitro activity of the siderophore cephalosporin, cefiderocol, against a recent collection of clinically relevant Gram-negative bacilli from North America and Europe, Including Carbapenem-Nonsusceptible Isolates (SIDERO-WT-2014 Study). Antimicrob Agents Chemother 61:1–16. https://doi.org/10.1128/AAC.00093-17
Kazmierczak KM, Tsuji M, Wise MG et al (2019) In vitro activity of cefiderocol, a siderophore cephalosporin, against a recent collection of clinically relevant carbapenem-non-susceptible Gram-negative bacilli, including serine carbapenemase- and metallo- β-lactamase-producing isolates (SIDERO-WT-2014 Study). Int J Antimicrob Agents 53:177–184. https://doi.org/10.1016/j.ijantimicag.2018.10.007
Carcione D, Siracusa C, Sulejmani A et al (2021) In vitro antimicrobial activity of the siderophore cephalosporin cefiderocol against Acinetobacter baumannii strains recovered from clinical samples. Antibiotics 10:1–11. https://doi.org/10.3390/antibiotics10111309
Smoke SM, Brophy A, Reveron S et al (2023) Evolution and transmission of cefiderocol-resistant acinetobacter baumannii during an outbreak in the burn intensive care unit. Clin Infect Dis 76:1–3. https://doi.org/10.1093/cid/ciac647
Malik S, Kaminsky M, Landman D, Quale J (2020) Cefiderocol resistance in Acinetobacter baumannii: roles of β-lactamases, siderophore receptors, and penicillin binding protein 3. Antimicrob Agents Chemother 64:1–4. https://doi.org/10.1128/aac.01221-20
Dobias J, Dénervaud-Tendon V, Poirel L, Nordmann P (2017) Activity of the novel siderophore cephalosporin cefiderocol against multidrug-resistant Gram-negative pathogens. Eur J Clin Microbiol Infect Dis 36:2319–2327. https://doi.org/10.1007/s10096-017-3063-z
Giammanco A, Calà C, Fasciana T, Dowzicky MJ (2017) Global assessment of the activity of tigecycline against multidrug-resistant Gram-negative pathogens between 2004 and 2014 as part of the tigecycline evaluation and surveillance trial. Am Soc Microbiol 2:1–10. https://doi.org/10.1128/mSphere.00310-16
European Centre for Disease Prevention and Control, World Health Organization (2023) Antimicrobial resistance surveillance in Europe 2023 - 2021 data. Solna, Sweden
European Centre for Disease Prevention and Control (2023) Antimicrobial resistance surveillance in Europe 2023 - 2021 data. Antimicrob. Resist. Surveill. Eur, In https://www.ecdc.europa.eu/en/publications-data/antimicrobial-resistance-surveillance-europe-2023-2021-data. Accessed 20 Sep 2023
World Health Organization (2017) WHO publishes list of bacteria for which new antibiotics are urgently needed. World Heal. Organ, In https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed. Accessed 8 Jun 2022
US Department of Health and Human Services, CDC (2019) Antibiotic resistance threats in the United States. Centers Dis. Control Prev, In https://www.cdc.gov/drugresistance/biggest-threats.html. Accessed 18 Apr 2022
Costerton JW, Lewandowski Z, Caldwell DE et al (1995) MICROBIAL BIOFILMS. Annu Rev Microbiol 49:711–745. https://doi.org/10.1146/annurev.mi.49.100195.003431
Buret AG, Motta JP, Allain T et al (2019) Pathobiont release from dysbiotic gut microbiota biofilms in intestinal inflammatory diseases: a role for iron? J Biomed Sci 26:1–14. https://doi.org/10.1186/s12929-018-0495-4
Di Martino P (2018) Extracellular polymeric substances, a key element in understanding biofilm phenotype. AIMS Microbiol 4:274–288. https://doi.org/10.3934/microbiol.2018.2.274
Donlan RM, Costerton JW (2002) Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev 15:167–193. https://doi.org/10.1128/cmr.15.2.167-193.2002
Anderl JN, Franklin MJ, Stewart PS (2000) Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob Agents Chemother 44:1818–1824. https://doi.org/10.1128/AAC.44.7.1818-1824.2000
Shiau AL, Wu CL (1998) The inhibitory effect of Staphylococcus epidermidis slime on the phagocytosis of murine peritoneal macrophages is interferon-independent. Microbiol Immunol 42:33–40. https://doi.org/10.1111/j.1348-0421.1998.tb01966.x
Kaya E, Grassi L, Benedetti A et al (2020) In vitro interaction of Pseudomonas aeruginosa biofilms with human peripheral blood mononuclear cells. Front Cell Infect Microbiol 10:1–13. https://doi.org/10.3389/fcimb.2020.00187
Yin W, Wang Y, Liu L, He J (2019) Biofilms: the microbial “protective clothing” in extreme environments. Int J Mol Sci 20:1–18. https://doi.org/10.3390/ijms20143423
Gil-Perotin S, Ramirez P, Marti V et al (2012) Implications of endotracheal tube biofilm in ventilator-associated pneumonia response: a state of concept. Crit Care 16:1–8. https://doi.org/10.1186/cc11357
Tomaras AP, Dorsey CW, Edelmann RE, Actis LA (2003) Attachment to and biofilm formation on abiotic surfaces by Acinetobacter baumannii: involvement of a novel chaperone-usher pili assembly system. Microbiology 149:3473–3484. https://doi.org/10.1099/mic.0.26541-0
Choudhary M, Shrivastava R, Vashistt J (2022) Acinetobacter baumannii biofilm formation: association with antimicrobial resistance and prolonged survival under desiccation. Curr Microbiol 79:1–9. https://doi.org/10.1007/s00284-022-03071-5
Bryers JD (2008) Medical biofilms. Biotechnol Bioeng 100:1–18. https://doi.org/10.1002/bit.21838
Buret A, Ward KH, Olson ME, Costerton JW (1991) An in vivo model to study the pathobiology of infectious biofilms on biomaterial surfaces. J Biomed Mater Res 25:865–874. https://doi.org/10.1002/jbm.820250706
Hathroubi S, Mekni MA, Domenico P et al (2016) Biofilms: microbial shelters against antibiotics. Microb Drug Resist 23:147–156. https://doi.org/10.1089/mdr.2016.0087
Ceri H, Olson ME, Stremick C et al (1999) The calgary biofilm device: new technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J Clin Microbiol 37:1771–1776. https://doi.org/10.1128/jcm.37.6.1771-1776.1999
Wenner M (2009) Quiet bacteria and antibiotic resistance. Sci. Am, In https://www.scientificamerican.com/article/bacteria-antibiotic-resistance/. Accessed 16 Apr 2022
Qi L, Li H, Zhang C et al (2016) Relationship between antibiotic resistance, biofilm formation, and biofilm-specific resistance in Acinetobacter baumannii. Front Microbiol 7:1–10. https://doi.org/10.3389/fmicb.2016.00483
Hall CW, Mah T-FF (2017) Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol Rev 41:276–301. https://doi.org/10.1093/femsre/fux010
Madsen JS, Burmølle M, Hansen LH, Sørensen SJ (2012) The interconnection between biofilm formation and horizontal gene transfer. FEMS Immunol Med Microbiol 65:183–195. https://doi.org/10.1111/j.1574-695X.2012.00960.x
Brito IL (2021) Examining horizontal gene transfer in microbial communities. Nat Rev Microbiol 19:442–453. https://doi.org/10.1038/s41579-021-00534-7
Hennequin C, Aumeran C, Robin F et al (2012) Antibiotic resistance and plasmid transfer capacity in biofilm formed with a CTX-M-15-producing Klebsiella pneumoniae isolate. J Antimicrob Chemother 67:2123–2130. https://doi.org/10.1093/jac/dks169
Bowler P, Murphy C, Wolcott R (2020) Biofilm exacerbates antibiotic resistance: is this a current oversight in antimicrobial stewardship? Antimicrob Resist Infect Control 9:1–5. https://doi.org/10.1186/s13756-020-00830-6
Michaelis C, Grohmann E (2023) Horizontal gene transfer of antibiotic resistance genes in biofilms. Antibiotics 12:1–31. https://doi.org/10.3390/antibiotics12020328
Rao RS, Karthika RU, Singh S et al (2008) Correlation between biofilm production and multiple drug resistance in imipenem resistant clinical isolates of Acinetobacter baumannii. Indian J Med Microbiol 26:333–337. https://doi.org/10.1016/S0255-0857(21)01809-0
Khoshnood S, Sadeghifard N, Mahdian N et al (2023) Antimicrobial resistance and biofilm formation capacity among Acinetobacter baumannii strains isolated from patients with burns and ventilator-associated pneumonia. J Clin Lab Anal 37:1–10. https://doi.org/10.1002/jcla.24814
Bardbari AM, Arabestani MR, Karami M et al (2017) Correlation between the ability of biofilm formation with their responsible genes and MDR patterns in clinical and environmental Acinetobacter baumannii isolates. Microb Pathog 108:122–128. https://doi.org/10.1016/j.micpath.2017.04.039
Rodrigues Perez LR (2015) Acinetobacter baumannii displays inverse relationship between meropenem resistance and biofilm production. J Chemother 27:13–15. https://doi.org/10.1179/1973947813Y.0000000159
Li Y-H, Tang N, Aspiras MB et al (2002) A quorum-sensing signaling system essential for genetic competence in Streptococcus mutans is involved in biofilm formation. J Bacteriol 184:2699–2708. https://doi.org/10.1128/jb.184.10.2699-2708.2002
Pesci EC, Pearson JP, Seed PC, Iglewski BH (1997) Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J Bacteriol 179:3127–3132. https://doi.org/10.1128/jb.179.10.3127-3132.1997
Davies DG, Parsek MR, Pearson JP et al (1998) The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 280:295–298. https://doi.org/10.1126/science.280.5361.295
Huber B, Riedel K, Hentzer M et al (2001) The cep quorum-sensing system of Burkholderia cepacia H111 controls biofilm formation and swarming motility. Microbiology 147:2517–2528. https://doi.org/10.1099/00221287-147-9-2517
Zhao X, Yu Z, Ding T (2020) Quorum-sensing regulation of antimicrobial resistance in bacteria. Microorganisms 8:1–21. https://doi.org/10.3390/microorganisms8030425
Tonkin M, Khan S, Wani YM, Ahmad A (2021) Quorum sensing – a stratagem for conquering multi-drug resistant pathogens. Curr Pharm Des 27:2835–2847. https://doi.org/10.2174/1381612826666201210105638
Niu C, Clemmer KM, Bonomo RA, Rather PN (2008) Isolation and characterization of an autoinducer synthase from Acinetobacter baumannii. J Bacteriol 190:3386–3392. https://doi.org/10.1128/JB.01929-07
Subhadra B, Hwan Oh M, Hee Choi C (2016) Quorum sensing in Acinetobacter: with special emphasis on antibiotic resistance, biofilm formation and quorum quenching. AIMS Microbiol 2:27–41. https://doi.org/10.3934/microbiol.2016.1.27
Dou Y, Song F, Guo F et al (2017) Acinetobacter baumannii quorum-sensing signalling molecule induces the expression of drug-resistance genes. Mol Med Rep 15:4061–4068. https://doi.org/10.3892/mmr.2017.6528
Tang J, Chen Y, Wang X et al (2020) Contribution of the AbaI/AbaR quorum sensing system to resistance and virulence of Acinetobacter baumannii clinical strains. Infect Drug Resist 13:4273–4281. https://doi.org/10.2147/IDR.S276970
He X, Lu F, Yuan F et al (2015) Biofilm formation caused by clinical Acinetobacter baumannii isolates is associated with overexpression of the AdeFGH efflux pump. Antimicrob Agents Chemother 59:4817–4825. https://doi.org/10.1128/aac.00877-15
Oh MH, Han K (2020) AbaR is a LuxR type regulator essential for motility and the formation of biofilm and pellicle in Acinetobacter baumannii. Genes and Genomics 42:1339–1346. https://doi.org/10.1007/s13258-020-01005-8
Gambello MJ, Iglewski BH (1991) Cloning and characterization of the Pseudomonas aeruginosa lasR gene, a transcriptional activator of elastase expression. J Bacteriol 173:3000–3009. https://doi.org/10.1128/jb.173.9.3000-3009.1991
Passador L, Cook JM, Gambello MJ et al (1993) Expression of Pseudomonas aeruginosa virulence genes requires cell-to-cell communication. Science 260:1127–1130. https://doi.org/10.1126/science.8493556
Pearson JP, Gray KM, Passador L et al (1994) Structure of the autoinducer required for expression of Pseudomonas aeruginosa virulence genes. Proc Natl Acad Sci U S A 91:197–201. https://doi.org/10.1073/pnas.91.1.197
Erickson DL, Endersby R, Kirkham A et al (2002) Pseudomonas aeruginosa quorum-sensing systems may control virulence factor expression in the lungs of patients with cystic fibrosis. Infect Immun 70:1783–1790. https://doi.org/10.1128/iai.70.4.1783-1790.2002
Novick RP, Geisinger E (2008) Quorum sensing in staphylococci. Annu Rev Genet 42:541–564. https://doi.org/10.1146/annurev.genet.42.110807.091640
Rutherford ST, Bassler BL (2012) Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harb Perspect Med 2:1–26. https://doi.org/10.1101/cshperspect.a012427
Chen X, Schauder S, Potier N et al (2002) Structural identification of a bacterial quorum-sensing signal containing boron. Nature 415:545–549. https://doi.org/10.1038/415545a
Xavier KB, Bassler BL (2005) Interference with AI-2-mediated bacterial cell-cell communication. Nature 437:750–753. https://doi.org/10.1038/nature03960
Sperandio V, Torres AG, Jarvis B et al (2003) Bacteria-host communication: the language of hormones. Proc Natl Acad Sci 100:8951–8956. https://doi.org/10.1073/pnas.1537100100
Kim CS, Gatsios A, Cuesta S et al (2020) Characterization of autoinducer-3 structure and biosynthesis in E. coli. ACS Cent Sci 6:197–206. https://doi.org/10.1021/acscentsci.9b01076
Hernandez DE, Sintim HO (2020) Quorum sensing autoinducer-3 finally yields to structural elucidation. ACS Cent Sci 6:93–96. https://doi.org/10.1021/acscentsci.0c00033
Newton JA, Fray RG (2004) Integration of environmental and host-derived signals with quorum sensing during plant-microbe interactions. Cell Microbiol 6:213–224. https://doi.org/10.1111/j.1462-5822.2004.00362.x
Parsek MR, Val DL, Hanzelka BL et al (1999) Acyl homoserine-lactone quorum-sensing signal generation. Proc Natl Acad Sci U S A 96:4360–4365. https://doi.org/10.1073/pnas.96.8.4360
Whitehead NA, Barnard AML, Slater H et al (2001) Quorum-sensing in Gram-negative bacteria. FEMS Microbiol Rev 25:365–404. https://doi.org/10.1111/j.1574-6976.2001.tb00583.x
Miller MB, Bassler BL (2001) Quorum sensing in bacteria. Annu Rev Microbiol 55:165–199. https://doi.org/10.1146/annurev.micro.55.1.165
Abee T, Kovács ÁT, Kuipers OP, van der Veen S (2011) Biofilm formation and dispersal in Gram-positive bacteria. Curr Opin Biotechnol 22:172–179. https://doi.org/10.1016/j.copbio.2010.10.016
Sun J, Daniel R, Wagner-Döbler I, Zeng AP (2004) Is autoinducer-2 a universal signal for interspecies communication: a comparative genomic and phylogenetic analysis of the synthesis and signal transduction pathways. BMC Evol Biol 4:1–11. https://doi.org/10.1186/1471-2148-4-36
Lowery CA, Dickerson TJ, Janda KD (2008) Interspecies and interkingdom communication mediated by bacterial quorum sensing. Chem Soc Rev 37:1337–1346. https://doi.org/10.1039/b702781h
Kendall MM, Sperandio V (2016) What a dinner party! Mechanisms and functions of interkingdom signaling in host-pathogen associations. MBio 7:1–14. https://doi.org/10.1128/mBio.01748-15
Anbazhagan D, Mansor M, Yan GOS et al (2012) Detection of quorum sensing signal molecules and identification of an autoinducer synthase gene among biofilm forming clinical isolates of Acinetobacter spp. PLoS One 7:1–12. https://doi.org/10.1371/journal.pone.0036696
Saipriya K, Swathi CH, Ratnakar KS, Sritharan V (2020) Quorum-sensing system in Acinetobacter baumannii: a potential target for new drug development. J Appl Microbiol 128:15–27. https://doi.org/10.1111/jam.14330
López-Martín M, Dubern JF, Alexander MR, Williams P (2021) AbaM regulates quorum sensing, biofilm formation, and virulence in Acinetobacter baumannii. J Bacteriol 203:1–13. https://doi.org/10.1128/JB.00635-20
Zhong S, He S (2021) Quorum sensing inhibition or quenching in Acinetobacter baumannii: the novel therapeutic strategies for new drug development. Front Microbiol 12:1–7. https://doi.org/10.3389/fmicb.2021.558003
Shih PC, Huang CT (2002) Effects of quorum-sensing deficiency on Pseudomonas aeruginosa biofilm formation and antibiotic resistance. J Antimicrob Chemother 49:309–314. https://doi.org/10.1093/jac/49.2.309
Hassett DJ, Ma JF, Elkins JG et al (1999) Quorum sensing in Pseudomonas aeruginosa controls expression of catalase and superoxide dismutase genes and mediates biofilm susceptibility to hydrogen peroxide. Mol Microbiol 34:1082–1093. https://doi.org/10.1046/j.1365-2958.1999.01672.x
Penesyan A, Nagy SS, Kjelleberg S et al (2019) Rapid microevolution of biofilm cells in response to antibiotics. npj Biofilms Microbiomes 5:1–14. https://doi.org/10.1038/s41522-019-0108-3
Chen J, Wang B, Lu Y et al (2019) Quorum sensing inhibitors from marine microorganisms and their synthetic derivatives. Mar Drugs 17:1–27. https://doi.org/10.3390/md17020080
Maeda T, García-Contreras R, Pu M et al (2012) Quorum quenching quandary: resistance to antivirulence compounds. ISME J 6:493–501. https://doi.org/10.1038/ismej.2011.122
Cepas V, López Y, Muñoz E et al (2019) Relationship between biofilm formation and antimicrobial resistance in Gram-negative bacteria. Microb Drug Resist 25:72–79. https://doi.org/10.1089/mdr.2018.0027
Dieltjens L, Appermans K, Lissens M et al (2020) Inhibiting bacterial cooperation is an evolutionarily robust anti-biofilm strategy. Nat Commun 11:1–11. https://doi.org/10.1038/s41467-019-13660-x
Flores-Vargas G, Bergsveinson J, Lawrence JR, Korber DR (2021) Environmental biofilms as reservoirs for antimicrobial resistance. Front Microbiol 12:1–13. https://doi.org/10.3389/fmicb.2021.766242
Mitchell A (2001) Quorum quenching. Nat Rev Mol Cell Biol 2:488–488. https://doi.org/10.1038/35080021
Fong J, Zhang C, Yang R et al (2018) Combination therapy strategy of quorum quenching enzyme and quorum sensing inhibitor in suppressing multiple quorum sensing pathways of P. aeruginosa. Sci Rep 8:1–11. https://doi.org/10.1038/s41598-018-19504-w
Bhargava N, Sharma P, Capalash N (2010) Quorum sensing in Acinetobacter: an emerging pathogen. Crit Rev Microbiol 36:349–360. https://doi.org/10.3109/1040841X.2010.512269
Sun X, Ni Z, Tang J et al (2021) The abaI/abaR quorum sensing system effects on pathogenicity in Acinetobacter baumannii. Front Microbiol 12:1–19. https://doi.org/10.3389/fmicb.2021.679241
Xiong L, Yi F, Yu Q et al (2022) Transcriptomic analysis reveals the regulatory role of quorum sensing in the Acinetobacter baumannii ATCC 19606 via RNA-seq. BMC Microbiol 22:1–12. https://doi.org/10.1186/s12866-022-02612-z
Modarresi F, Azizi O, Shakibaie MR et al (2016) Cloning and expression of quorum sensing N-acyl-homoserine synthase (LuxI) gene detected in Acinetobacter baumannii. Iran J Microbiol 8:139–146
Boşgelmez-Tinaz G, Ulusoy S, Aridoǧan B et al (2005) N-butanoyl-L-homoserine lactone (BHL) deficient Pseudomonas aeruginosa isolates from an intensive care unit. Microbiol Res 160:399–403. https://doi.org/10.1016/j.micres.2005.03.005
Bitrian M, Solari CM, González RH, Nudel CB (2012) Identification of virulence markers in clinically relevant strains of Acinetobacter genospecies. Int Microbiol 15:79–88. https://doi.org/10.2436/20.1501.01.161
González RH, Dijkshoorn L, Van Den Barselaar M, Nudel C (2009) Quorum sensing signal profile of Acinetobacter strains from nosocomial and environmental sources. Rev Argent Microbiol 41:73–78
Prashanth K, Vasanth T, Saranathan R et al (2012) Antibiotic resistance, biofilms and quorum sensing in Acinetobacter species. InTech:179–214. https://doi.org/10.5772/28813
Choi CH, Lee JS, Lee YC et al (2008) Acinetobacter baumannii invades epithelial cells and outer membrane protein A mediates interactions with epithelial cells. BMC Microbiol 8:1–11. https://doi.org/10.1186/1471-2180-8-216
Gaddy JA, Tomaras AP, Actis LA (2009) The Acinetobacter baumannii 19606 OmpA protein plays a role in biofilm formation on abiotic surfaces and in the interaction of this pathogen with eukaryotic cells. Infect Immun 77:3150–3160. https://doi.org/10.1128/IAI.00096-09
Loehfelm TW, Luke NR, Campagnari AA (2008) Identification and characterization of an Acinetobacter baumannii biofilm-associated protein. J Bacteriol 190:1036–1044. https://doi.org/10.1128/jb.01416-07
Brossard KA, Campagnari AA (2012) The Acinetobacter baumannii biofilm-associated protein plays a role in adherence to human epithelial cells. Infect Immun 80:228–233. https://doi.org/10.1128/iai.05913-11
Tomaras AP, Flagler MJ, Dorsey CW et al (2008) Characterization of a two-component regulatory system from Acinetobacter baumannii that controls biofilm formation and cellular morphology. Microbiology 154:3398–3409. https://doi.org/10.1099/mic.0.2008/019471-0
Richmond GE, Evans LP, Anderson MJ et al (2016) The Acinetobacter baumannii two-component system AdeRS regulates genes required for multidrug efflux, biofilm formation, and virulence in a strain-specific manner. MBio 7:1–11. https://doi.org/10.1128/mBio.00430-16
Marchand I, Damier-piolle L, Courvalin P, Lambert T (2004) Expression of the RND-type efflux pump AdeABC in Acinetobacter baumannii is regulated by the AdeRS two-component system. Society 48:3298–3304. https://doi.org/10.1128/aac.48.9.3298-3304.2004
Choi AHK, Slamti L, Avci FY et al (2009) The pgaABCD locus of Acinetobacter baumannii encodes the production of poly-β-1-6-N-acetylglucosamine, which is critical for biofilm formation. J Bacteriol 191:5953–5963. https://doi.org/10.1128/JB.00647-09
Russo TA, Luke NR, Beanan JM et al (2010) The K1 capsular polysaccharide of Acinetobacter baumannii strain 307-0294 is a major virulence factor. Infect Immun 78:3993–4000. https://doi.org/10.1128/IAI.00366-10
Vahaboglu H, Öztürk R, Aygün G et al (1997) Widespread detection of PER-1-type extended-spectrum β-lactamases among nosocomial Acinetobacter and Pseudomonas aeruginosa isolates in Turkey: a nationwide multicenter study. Antimicrob Agents Chemother 41:2265–2269. https://doi.org/10.1128/aac.41.10.2265
Vahaboglu H, Coskunkan F, Tansel O et al (2001) Clinical importance of extended-spectrum β-lactamase (PER-1-type)-producing Acinetobacter spp. and Pseudomonas aeruginosa strains. J Med Microbiol 50:642–645. https://doi.org/10.1099/0022-1317-50-7-642
Poirel L, Cabanne L, Vahaboglu H, Nordmann P (2005) Genetic environment and expression of the extended-spectrum β-lactamase blaPER-1 gene in Gram-negative bacteria. Antimicrob Agents Chemother 49:1708–1713. https://doi.org/10.1128/AAC.49.5.1708-1713.2005
Lee HW, Koh YM, Kim J et al (2008) Capacity of multidrug-resistant clinical isolates of Acinetobacter baumannii to form biofilm and adhere to epithelial cell surfaces. Clin Microbiol Infect 14:49–54. https://doi.org/10.1111/j.1469-0691.2007.01842.x
Petrova OE, Sauer K (2012) Sticky situations: key components that control bacterial surface attachment. J Bacteriol 194:2413–2425. https://doi.org/10.1128/JB.00003-12
Yang CH, Su PW, Moi SH, Chuang LY (2019) Biofilm formation in Acinetobacter baumannii: genotype-phenotype correlation. Molecules 24:1–12. https://doi.org/10.3390/molecules24101849
Moon KH, Weber BS, Feldmana MF (2017) Subinhibitory concentrations of trimethoprim and sulfamethoxazole prevent biofilm formation by Acinetobacter baumannii through inhibition of Csu pilus expression. Antimicrob Agents Chemother 61:1–18. https://doi.org/10.1128/AAC.00778-17
de Breij A, Gaddy J, van der Meer J et al (2009) CsuA/BABCDE-dependent pili are not involved in the adherence of Acinetobacter baumannii ATCC19606 to human airway epithelial cells and their inflammatory response. Res Microbiol 160:213–218. https://doi.org/10.1016/j.resmic.2009.01.002
Jyothisri K, Deepak V, Rajeswari MR (1999) Purification and characterization of a major 40 kDa outer membrane protein of Acinetobacter baumannii. FEBS Lett 443:57–60. https://doi.org/10.1016/s0014-5793(98)01679-2
Choi CH, Hyun SH, Lee JY et al (2008) Acinetobacter baumannii outer membrane protein A targets the nucleus and induces cytotoxicity. Cell Microbiol 10:309–319. https://doi.org/10.1111/j.1462-5822.2007.01041.x
Choi CH, Lee EY, Lee YC et al (2005) Outer membrane protein 38 of Acinetobacter baumannii localizes to the mitochondria and induces apoptosis of epithelial cells. Cell Microbiol 7:1127–1138. https://doi.org/10.1111/j.1462-5822.2005.00538.x
Lee JS, Choi CH, Kim JW, Lee JC (2010) Acinetobacter baumannii outer membrane protein A induces dendritic cell death through mitochondrial targeting. J Microbiol 48:387–392. https://doi.org/10.1007/s12275-010-0155-1
Jin JS, Kwon S-O, Moon DC et al (2011) Acinetobacter baumannii secretes cytotoxic outer membrane protein A via outer membrane vesicles. PLoS One 6:1–9. https://doi.org/10.1371/journal.pone.0017027
Tiku V, Kofoed EM, Yan D et al (2021) Outer membrane vesicles containing OmpA induce mitochondrial fragmentation to promote pathogenesis of Acinetobacter baumannii. Sci Rep 11:1–16. https://doi.org/10.1038/s41598-020-79966-9
Lee JS, Lee JC, Lee C-M et al (2007) Outer membrane protein A of Acinetobacter baumannii induces differentiation of CD4+ T cells toward a Th1 polarizing phenotype through the activation of dendritic cells. Biochem Pharmacol 74:86–97. https://doi.org/10.1016/j.bcp.2007.02.012
Sánchez-Encinales V, Álvarez-Marín R, Pachón-Ibáñez ME et al (2017) Overproduction of outer membrane protein A by Acinetobacter baumannii as a risk factor for nosocomial pneumonia, bacteremia, and mortality rate increase. J Infect Dis 215:966–974. https://doi.org/10.1093/infdis/jix010
Il KH, Kim S, Oh MH et al (2017) Outer membrane protein A contributes to antimicrobial resistance of Acinetobacter baumannii through the OmpA-like domain. J Antimicrob Chemother 72:3012–3015. https://doi.org/10.1093/jac/dkx257
Lasa I, Penadés JR (2006) Bap: a family of surface proteins involved in biofilm formation. Res Microbiol 157:99–107. https://doi.org/10.1016/j.resmic.2005.11.003
Cucarella C, Solano C, Valle J et al (2001) Bap, a Staphylococcus aureus surface protein involved in biofilm formation. J Bacteriol 183:2888–2896. https://doi.org/10.1128/jb.183.9.2888-2896.2001
Pompilio A, Piccolomini R, Picciani C et al (2008) Factors associated with adherence to and biofilm formation on polystyrene by Stenotrophomonas maltophilia: the role of cell surface hydrophobicity and motility. FEMS Microbiol Lett 287:41–47. https://doi.org/10.1111/j.1574-6968.2008.01292.x
Kochkodan V, Tsarenko S, Potapchenko N et al (2008) Adhesion of microorganisms to polymer membranes: a photobactericidal effect of surface treatment with TiO2. Desalination 220:380–385. https://doi.org/10.1016/j.desal.2007.01.042
Mirani ZA, Fatima A, Urooj S et al (2018) Relationship of cell surface hydrophobicity with biofilm formation and growth rate: a study on Pseudomonas aeruginosa, Staphylococcus aureus, and Escherichia coli. Iran J Basic Med Sci 21:760–769. https://doi.org/10.22038/IJBMS.2018.28525.6917
Sharon Goh HM, Beatson SA, Totsika M et al (2013) Molecular analysis of the Acinetobacter baumannii biofilm-associated protein. Appl Environ Microbiol 79:6535–6543. https://doi.org/10.1128/AEM.01402-13
Badmasti F, Siadat SD, Bouzari S et al (2015) Molecular detection of genes related to biofilm formation in multidrug-resistant Acinetobacter baumannii isolated from clinical settings. J Med Microbiol 64:559–564. https://doi.org/10.1099/jmm.0.000058
Fallah A, Ahangarzadeh Rezaee M, Hasani A et al (2017) Frequency of bap and cpaA virulence genes in drug resistant clinical isolates of Acinetobacter baumannii and their role in biofilm formation. Iran J Basic Med Sci 20:849–855. https://doi.org/10.22038/IJBMS.2017.9105
Stock AM, Robinson VL, Goudreau PN (2000) Two-component signal transduction. Annu Rev Biochem 69:183–215. https://doi.org/10.1146/annurev.biochem.69.1.183
Matsuo M, Kato F, Oogai Y et al (2010) Distinct two-component systems in methicillin-resistant Staphylococcus aureus can change the susceptibility to antimicrobial agents. J Antimicrob Chemother 65:1536–1537. https://doi.org/10.1093/jac/dkq141
Hu WS, Li PC, Cheng CY (2005) Correlation between ceftriaxone resistance of Salmonella enterica serovar typhimurium and expression of outer membrane proteins OmpW and Ail/OmpX-like protein, which are regulated by BaeR of a two-component system. Antimicrob Agents Chemother 49:3955–3958. https://doi.org/10.1128/AAC.49.9.3955-3958.2005
Huang H, Sun Y, Yuan L et al (2016) Regulation of the two-component regulator CpxR on aminoglycosides and β-lactams resistance in Salmonella enterica serovar typhimurium. Front Microbiol 7:1–10. https://doi.org/10.3389/fmicb.2016.00604
Teschler JK, Cheng AT, Yildiz FH (2017) The two-component signal transduction system VxrAB positively regulates Vibrio cholerae biofilm formation. J Bacteriol 199:1–16. https://doi.org/10.1128/JB.00139-17
Palethorpe S, Farrow JM, Wells G et al (2022) Acinetobacter baumannii regulates its stress responses via the BfmRS two-component regulatory system. J Bacteriol 204:1–20. https://doi.org/10.1128/jb.00494-21
Kim H-J, Kim N-Y, Ko S-Y et al (2022) Complementary regulation of BfmRS two-component and AbaIR quorum sensing systems to express virulence-associated genes in Acinetobacter baumannii. Int J Mol Sci 23:1–14. https://doi.org/10.3390/ijms232113136
Krasauskas R, Skerniškytė J, Armalytė J, Sužiedėlienė E (2019) The role of Acinetobacter baumannii response regulator BfmR in pellicle formation and competitiveness via contact-dependent inhibition system. BMC Microbiol 19:1–12. https://doi.org/10.1186/s12866-019-1621-5
Cerqueira GM, Kostoulias X, Khoo C et al (2014) A global virulence regulator in Acinetobacter baumannii and its control of the phenylacetic acid catabolic pathway. J Infect Dis 210:46–55. https://doi.org/10.1093/infdis/jiu024
Russo TA, Manohar A, Beanan JM et al (2016) The response regulator BfmR is a potential drug target for Acinetobacter baumannii. msphere 1:1–19. https://doi.org/10.1128/msphere.00082-16
Marr CM, MacDonald U, Trivedi G et al (2020) An evaluation of BfmR-regulated antimicrobial resistance in the extensively drug resistant (XDR) Acinetobacter baumannii Strain HUMC1. Front Microbiol 11:1–15. https://doi.org/10.3389/fmicb.2020.595798
Farrow JM, Wells G, Pesci EC (2018) Desiccation tolerance in Acinetobacter baumannii is mediated by the two-component response regulator BfmR. PLoS One 13:1–25. https://doi.org/10.1371/journal.pone.0205638
Liou M-L, Soo P-C, Ling S-R et al (2014) The sensor kinase BfmS mediates virulence in Acinetobacter baumannii. J Microbiol Immunol Infect 47:275–281. https://doi.org/10.1016/j.jmii.2012.12.004
Kim SY, Kim MH, Kim S Il, et al (2019) The sensor kinase BfmS controls production of outer membrane vesicles in Acinetobacter baumannii. BMC Microbiol 19:1–13. https://doi.org/10.1186/s12866-019-1679-0
Geisinger E, Isberg RR (2015) Antibiotic modulation of capsular exopolysaccharide and virulence in Acinetobacter baumannii. PLoS Pathog 11:1–27. https://doi.org/10.1371/journal.ppat.1004691
Wang N, Ozer EA, Mandel MJ, Hauser AR (2014) Genome-wide identification of Acinetobacter baumannii genes necessary for persistence in the lung. MBio 5:1–14. https://doi.org/10.1128/mbio.01163-14
Geisinger E, Mortman NJ, Vargas-Cuebas G et al (2018) A global regulatory system links virulence and antibiotic resistance to envelope homeostasis in Acinetobacter baumannii. PLoS Pathog 14:1–28. https://doi.org/10.1371/journal.ppat.1007030
Lee H-W, Koh YM, Kim J et al (2008) Capacity of multidrug-resistant clinical isolates of Acinetobacter baumannii to form biofilm and adhere to epithelial cell surfaces. Clin Microbiol Infect 14:49–54. https://doi.org/10.1111/j.1469-0691.2007.01842.x
Naeimi Mazraeh F, Hasani A, Sadeghi J et al (2021) High frequency of blaPER-1 gene in clinical strains of Acinetobacter baumannii and its association with quorum sensing and virulence factors. Gene Reports 24:1–7. https://doi.org/10.1016/j.genrep.2021.101232
Morris FC, Dexter C, Kostoulias X et al (2019) The mechanisms of disease caused by Acinetobacter baumannii. Front Microbiol 10:1–20. https://doi.org/10.3389/fmicb.2019.01601
Jansen KU, Knirsch C, Anderson AS (2018) The role of vaccines in preventing bacterial antimicrobial resistance. Nat Med 24:10–20. https://doi.org/10.1038/nm.4465
Ramezanalizadeh F, Rasooli I, Owlia P et al (2022) Vaccination with a combination of planktonic and biofilm virulence factors confers protection against carbapenem-resistant Acinetobacter baumannii strains. Sci Rep 12:1–12. https://doi.org/10.1038/s41598-022-24163-z
Moriel DG, Beatson SA, Wurpel DJ et al (2013) Identification of novel vaccine candidates against multidrug-resistant Acinetobacter baumannii. PLoS One 8:1–9. https://doi.org/10.1371/journal.pone.0077631
Ansari H, Tahmasebi-Birgani M, Bijanzadeh M et al (2019) Study of the immunogenicity of outer membrane protein A (ompA) gene from Acinetobacter baumannii as DNA vaccine candidate in vivo. Iran J Basic Med Sci 22:669–675. https://doi.org/10.22038/ijbms.2019.30799.7427
Lei L, Yang F, Zou J et al (2019) DNA vaccine encoding OmpA and Pal from Acinetobacter baumannii efficiently protects mice against pulmonary infection. Mol Biol Rep 46:5397–5408. https://doi.org/10.1007/s11033-019-04994-2
Luo G, Lin L, Ibrahim AS et al (2012) Active and passive immunization protects against lethal, extreme drug resistant-Acinetobacter baumannii infection. PLoS One 7:1–11. https://doi.org/10.1371/journal.pone.0029446
Ramezanalizadeh F, Owlia P, Rasooli I (2020) Type I pili, CsuA/B and FimA induce a protective immune response against Acinetobacter baumannii. Vaccine 38:5436–5446. https://doi.org/10.1016/j.vaccine.2020.06.052
Fattahian Y, Rasooli I, Mousavi Gargari SL et al (2011) Protection against Acinetobacter baumannii infection via its functional deprivation of biofilm associated protein (Bap). Microb Pathog 51:402–406. https://doi.org/10.1016/j.micpath.2011.09.004
Fereshteh S, Ajdary S, Sepehr A et al (2023) Immunization with recombinant DcaP-like protein and AbOmpA revealed protections against sepsis infection of multi-drug resistant Acinetobacter baumannii ST2Pas in a C57BL/6 mouse model. Microb Pathog 174:1–10. https://doi.org/10.1016/j.micpath.2022.105882
Zhang X, Yang T, Cao J et al (2016) Mucosal immunization with purified OmpA elicited protective immunity against infections caused by multidrug-resistant Acinetobacter baumannii. Microb Pathog 96:20–25. https://doi.org/10.1016/j.micpath.2016.04.019
Bzdrenga J, Daudé D, Rémy B et al (2017) Biotechnological applications of quorum quenching enzymes. Chem Biol Interact 267:104–115. https://doi.org/10.1016/j.cbi.2016.05.028
Uroz S, Dessaux Y, Oger P (2009) Quorum sensing and quorum quenching: the Yin and Yang of bacterial communication. ChemBioChem 10:205–216. https://doi.org/10.1002/cbic.200800521
Chow JY, Yang Y, Tay SB et al (2014) Disruption of biofilm formation by the human pathogen Acinetobacter baumannii using engineered quorum-quenching lactonases. Antimicrob Agents Chemother 58:1802–1805. https://doi.org/10.1128/aac.02410-13
Zhang Y, Brackman G, Coenye T (2017) Pitfalls associated with evaluating enzymatic quorum quenching activity: the case of MomL and its effect on Pseudomonas aeruginosa and Acinetobacter baumannii biofilms. PeerJ 27:1–20. https://doi.org/10.7717/peerj.3251
Hraiech S, Hiblot J, Lafleur J et al (2014) Inhaled lactonase reduces Pseudomonas aeruginosa quorum sensing and mortality in rat pneumonia. PLoS One 9:1–8. https://doi.org/10.1371/journal.pone.0107125
Kiran S, Sharma P, Harjai K, Capalash N (2011) Enzymatic quorum quenching increases antibiotic susceptibility of multidrug resistant Pseudomonas aeruginosa. Iran J Microbiol 3:1–12
Khan I, Saeed K, Khan I (2019) Nanoparticles: properties, applications and toxicities. Arab J Chem 12:908–931. https://doi.org/10.1016/j.arabjc.2017.05.011
Siddique MH, Aslam B, Imran M et al (2020) Effect of silver nanoparticles on biofilm formation and EPS production of multidrug-resistant Klebsiella pneumoniae. Biomed Res Int 2020:1–9. https://doi.org/10.1155/2020/6398165
Verma P, Maheshwari SK (2017) Minimum biofilm eradication concentration (MBEC) assay of silver and selenium nanoparticles against biofilm forming Staphylococcus aureus. J Med Sci Clin Res 05:20213–20222. https://doi.org/10.18535/jmscr/v5i4.77
Hetta HF, Al-Kadmy IMS, Khazaal SS et al (2021) Antibiofilm and antivirulence potential of silver nanoparticles against multidrug-resistant Acinetobacter baumannii. Sci Rep 11:1–11. https://doi.org/10.1038/s41598-021-90208-4
Kalishwaralal K, BarathManiKanth S, Pandian SRK et al (2010) Silver nanoparticles impede the biofilm formation by Pseudomonas aeruginosa and Staphylococcus epidermidis. Colloids Surfaces B Biointerfaces 79:340–344. https://doi.org/10.1016/j.colsurfb.2010.04.014
Applerot G, Lellouche J, Perkas N et al (2012) ZnO nanoparticle-coated surfaces inhibit bacterial biofilm formation and increase antibiotic susceptibility. RSC Adv 2:2314–2321. https://doi.org/10.1039/C2RA00602B
Singh R, Vora J, Nadhe SB et al (2018) Antibacterial activities of bacteriagenic silver nanoparticles against nosocomial Acinetobacter baumannii. J Nanosci Nanotechnol 18:3806–3815. https://doi.org/10.1166/jnn.2018.15013
Scutera S, Argenziano M, Sparti R et al (2021) Enhanced antimicrobial and antibiofilm effect of new colistin-loaded human albumin nanoparticles. Antibiotics 10:1–13. https://doi.org/10.3390/antibiotics10010057
Li X, Gui R, Li J et al (2021) Novel multifunctional silver nanocomposite serves as a resistance-reversal agent to synergistically combat carbapenem-resistant Acinetobacter baumannii. ACS Appl Mater Interfaces 13:30434–30457. https://doi.org/10.1021/acsami.1c10309
Alzahrani RR, Alkhulaifi MM, Alenazi NM et al (2020) Characterization and biological investigation of silver nanoparticles biosynthesized from Galaxaura rugosa against multidrug-resistant bacteria. J Taibah Univ Sci 14:1651–1659. https://doi.org/10.1080/16583655.2020.1854495
Al-mousawi HTM, Al-Janabi NH (2021) Anti-biofilm effects and substantively properties of magnetic iron oxide nanoparticles synthesized against clinical isolates for MDR Acinetobacter baumannii and related with expression of gene. J Genet Environ Resour Conserv 9:122–133
Chen M, Yu X, Huo Q et al (2019) Biomedical potentialities of silver nanoparticles for clinical multiple drug-resistant Acinetobacter baumannii. J Nanomater 2019:1–7. https://doi.org/10.1155/2019/3754018
Vaidya MY, McBain AJ, Butler JA et al (2017) Antimicrobial efficacy and synergy of metal ions against Enterococcus faecium, Klebsiella pneumoniae and Acinetobacter baumannii in planktonic and biofilm phenotypes. Sci Rep 7:1–9. https://doi.org/10.1038/s41598-017-05976-9
Vaidya M, McBain AJ, Banks CE, Whitehead KA (2019) Single and combined antimicrobial efficacies for nine metal ion solutions against Klebsiella pneumoniae, Acinetobacter baumannii and Enterococcus faecium. Int Biodeterior Biodegrad 141:39–43. https://doi.org/10.1016/j.ibiod.2018.06.017
Godoy-Gallardo M, Eckhard U, Delgado LM et al (2021) Antibacterial approaches in tissue engineering using metal ions and nanoparticles: from mechanisms to applications. Bioact Mater 6:4470–4490. https://doi.org/10.1016/j.bioactmat.2021.04.033
McNeilly O, Mann R, Hamidian M, Gunawan C (2021) Emerging concern for silver nanoparticle resistance in Acinetobacter baumannii and other bacteria. Front Microbiol 12:1–20. https://doi.org/10.3389/fmicb.2021.652863
Mohamed SH, Salem D, Azmy M, Fam NS (2018) Antibacterial and antibiofilm activity of cinnamaldehyde against carbapenem-resistant Acinetobacter baumannii in Egypt: In vitro study. J Appl Pharm Sci 8:151–156. https://doi.org/10.7324/JAPS.2018.81121
Ouslimani S, Bendahou M, Abedelmounaim K et al (2023) Antibacterial and anti-biofilm efficiency of twenty Algerian plants essential oils against resistant Acinetobacter baumannii. J Essent Oil-Bearing Plants 26:206–231. https://doi.org/10.1080/0972060X.2023.2179424
Moghimi R, Aliahmadi A, Rafati H et al (2018) Antibacterial and anti-biofilm activity of nanoemulsion of Thymus daenensis oil against multi-drug resistant Acinetobacter baumannii. J Mol Liq 265:765–770. https://doi.org/10.1016/j.molliq.2018.07.023
Mumtaz L, Farid A, Yousef Alomar S et al (2023) Assessment of polyphenolic compounds against biofilms produced by clinical Acinetobacter baumannii strains using in silico and in vitro models. Saudi J Biol Sci 30:1–10. https://doi.org/10.1016/j.sjbs.2023.103743
Kim MK, Kang N, Ko SJ et al (2018) Antibacterial and antibiofilm activity and mode of action of magainin 2 against drug-resistant Acinetobacter baumannii. Int J Mol Sci 19:1–14. https://doi.org/10.3390/ijms19103041
Thulshan Jayathilaka EHT, Rajapaksha DC, Nikapitiya C et al (2021) Antimicrobial and anti-biofilm peptide octominin for controlling multidrug-resistant Acinetobacter baumannii. Int J Mol Sci 22. https://doi.org/10.3390/ijms22105353
Alsaab FM, Dean SN, Bobde S et al (2023) Computationally designed AMPs with antibacterial and antibiofilm activity against MDR Acinetobacter baumannii. Antibiotics 12:1–22. https://doi.org/10.3390/antibiotics12091396
Sharma S, Chatterjee S, Datta S et al (2017) Bacteriophages and its applications: an overview. Folia Microbiol (Praha) 62:17–55. https://doi.org/10.1007/s12223-016-0471-x
North OI, Brown ED (2021) Phage-antibiotic combinations: a promising approach to constrain resistance evolution in bacteria. Ann N Y Acad Sci 1496:23–34. https://doi.org/10.1111/nyas.14533
Luo J, Xie L, Yang M et al (2023) Synergistic antibacterial effect of phage pB3074 in combination with antibiotics targeting cell wall against multidrug-resistant Acinetobacter baumannii in vitro and ex vivo. Microbiol Spectr 11:1–11. https://doi.org/10.1128/spectrum.00341-23
Yele AB, Thawal ND, Sahu PK, Chopade BA (2012) Novel lytic bacteriophage AB7-IBB1 of Acinetobacter baumannii: isolation, characterization and its effect on biofilm. Arch Virol 157:1441–1450. https://doi.org/10.1007/s00705-012-1320-0
Thawal ND, Yele AB, Sahu PK, Chopade BA (2012) Effect of a novel podophage AB7-IBB2 on Acinetobacter baumannii biofilm. Curr Microbiol 65:66–72. https://doi.org/10.1007/s00284-012-0127-2
Wang X, Loh B, Gordillo Altamirano F et al (2021) Colistin-phage combinations decrease antibiotic resistance in Acinetobacter baumannii via changes in envelope architecture. Emerg Microbes Infect 10:2205–2219. https://doi.org/10.1080/22221751.2021.2002671
Gordillo Altamirano F, Forsyth JH, Patwa R et al (2021) Bacteriophage-resistant Acinetobacter baumannii are resensitized to antimicrobials. Nat Microbiol 6:157–161. https://doi.org/10.1038/s41564-020-00830-7
Acknowledgements
Sérgio G. Mendes is supported by a Ph.D. Research Grant with the reference 2021.06289.BD, given by the Portuguese Foundation for Science and Technology I.P. The authors acknowledge Antibe Therapeutics Inc. (Toronto, On, Canada) for financial support through a discovery research grant. Project support was also provided by the Faculty of Pharmacy of the University of Coimbra, Portugal.
Code availability
Not applicable.
Funding
Open access funding provided by FCT|FCCN (b-on).
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the design of this publication. Gabriela J. da Silva and A. G. Buret had the idea for the article. Sérgio G. Mendes and Sofia I. Combo performed the literature search. Sérgio G. Mendes wrote the first draft, and all authors critically reviewed the manuscript.
Corresponding author
Ethics declarations
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mendes, S.G., Combo, S.I., Allain, T. et al. Co-regulation of biofilm formation and antimicrobial resistance in Acinetobacter baumannii: from mechanisms to therapeutic strategies. Eur J Clin Microbiol Infect Dis 42, 1405–1423 (2023). https://doi.org/10.1007/s10096-023-04677-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-023-04677-8