Abstract
Aim
Volatile organic compounds (VOCs) are being studied as potential biomarkers in many infections. Therefore, this study aimed to analyze the volatile profile of three Gram-positive bacteria of clinical relevance to identify potential volatile biomarkers that allow their differentiation.
Methods and results
L. monocytogenes, S. aureus, and E. faecalis clinical isolates were inoculated in a thioglycollate medium until grown. Then, VOCs were extracted by solid-phase microextraction, and the data obtained were subjected to multivariate analysis. According to our results, there was a high production of aldehydes in E. faecalis. In the case of alcohols, they only increased in L. monocytogenes, while ketones were produced significantly in all three bacteria, mainly due to acetoin. Acids were produced significantly in E. faecalis and L. monocytogenes.
Conclusions
Potential biomarkers of L. monocytogenes could be 1-butanol and 2-methylbutanoic acid. In the case of E. faecalis, the VOC most related to its presence was nonanal. Lastly, potential biomarkers of S. aureus could be isoamyl butanoate and methionol, although some pyrazines have also been associated with this bacterium.
Significance and impact of the study
The identification of potential biomarkers of these clinically relevant bacteria could open the way for the diagnosis of these infections through the analysis of volatile compounds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Infections caused by Gram-positive bacteria are a public health problem due to the outbreaks of food poisoning that it can cause and the worrying increase of bacterial resistance to antibiotics that have been taking place in recent years. Among the Gram-positive bacteria involved in outbreaks of food poisoning, the most notable are Listeria monocytogenes, Bacillus cereus, Staphylococcus aureus, Clostridium perfringens, and Clostridium botulinum. L. monocytogenes causes listeriosis, an infection related to eating contaminated food that affects 1600 people and causes 260 deaths each year worldwide (CDC 2024). Listeriosis is the most severe human foodborne disease due to high hospitalization (99%) and mortality (15.6%) rates (Jiménez et al. 2022). In Spain, in 2019, an outbreak of food poisoning by L. monocytogenes was reported with 217 infected people and three deaths associated with the industrially produced larded meat with the CC388 clonal complex consumption (Ministerio de Sanidad, Consumo y Bienestar Social, 2019). In recent years, the upward trend in the number of outbreaks by Listeria spp., as in the number of hospitalized cases, is worrying (Jiménez et al. 2022).
S. aureus also usually causes food poisoning outbreaks by a toxigenic mechanism, giving rise to the sudden appearance of vomiting and diarrhea since preformed enterotoxins are ingested in food (Vila et al. 2009 2008). Between 2019 and 2020, 121 outbreaks of food poisoning by staphylococcal enterotoxins were reported in Europe, affecting almost 1900 people (Jiménez et al. 2022). In addition, methicillin-resistant S. aureus (MRSA), as multiresistant Enterococcus spp., are increasingly causing nosocomial infections due to the appearance of new MRSA clones and mec gene variants that are not detected by the most common molecular systems (Cantón & Ruiz-Garbajosa 2013). The multiresistant bacteria appearance calls for better use of antimicrobials and the application of epidemiological measures, including the detection of carriers, that reduce their transmission.
Detection of pathogens in samples from the environment (mainly foods) and patients usually involves time-consuming growth in selective media, isolation, and biochemical and molecular diagnostic analyses (Reichert-Schwillinsky et al. 2009). Thus, currently, the methodologies used to diagnose diseases have been diversified to a great extent. The search for specific disease biomarkers in non-invasive samples is a trend in several research areas. Specifically, volatile organic compounds (VOCs) are being studied as potential biomarkers in many diseases, such as cancer (Wen et al. 2020) and Alzheimer’s disease (Ubeda et al. 2022). Furthermore, microorganisms can produce species-specific VOCs as a product of their metabolism and could also be helpful for the diagnosis of infection as an odor fingerprinting (Tait et al. 2014). Thus, the VOCs produced by microorganisms have been employed successfully to identify their presence in biological samples with Helicobacter pylori (Ulanowska et al. 2011), Giardia lamblia (Ubeda et al. 2019), and SARS-CoV-2 (Lamote et al. 2020), among several others. Specifically, the volatilomic profile of L. monocytogenes has already been studied in thioglycolate broth (Lepe-Balsalobre et al. 2022), trypticase soy broth (Chen et al. 2017), and milk samples (Tait et al. 2014). Regarding S. aureus, the VOCs produced by this bacterium have also been studied in trypticase soy broth (Chen et al. 2017), blood agar media (Gómez-Mejia et al. 2022), and bronchoalveolar lavage fluid (Nasir et al. 2018). This bacterium, like E. faecalis, has also been studied in urine sample (Storer et al. 2011) and blood culture samples (Dolch et al. 2012).Therefore, this study aimed to analyze the volatile profile of the L. monocytogenes clonal complex responsible for the recent and severe outbreak in Spain (CC388) in comparison with that produced by other Gram-positive bacteria of clinical relevance, such as S. aureus and E. faecalis, to identify potential volatile biomarkers that allow their differentiation.
Materials and methods
This study was not submitted for institutional review board approval because it did not include individual patient data.
Bacterial strains and culture conditions
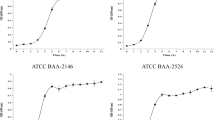
The study included the clinical isolate of Listeria monocytogenes characterized by Multilocus Sequence typing (MLST) belonging to the CC388 clonal complex involved in the epidemic outbreak of 2019 and two clinical isolates of Staphylococcus aureus (ATCC 29213) and Enterococcus faecalis (ATCC 29212) obtained from the American Type Culture Collection (ATCC).
A 0.5 McFarland dilution was prepared in sterile water from each isolate (plate count, 1.2 × 108 CFU/mL). Then, a technical triplicate 100 μl inoculum (1 × 106 CFU/mL) was transferred to three fluid thioglycollate medium 20-mL tubes (Becton, Dickinson and Company, USA) and incubated at 37°C for 24 h. The positivity was evaluated by bacterial count in Columbia Agar with 5% Sheep Blood (Becton, Dickinson and Company, USA). An uninoculated tube was used as a control. Tubes were stored at − 80 °C until analysis.
Extraction of volatile compounds
Solid-phase microextraction (SPME) was employed for the VOC extraction. Tubes were defrosted, and 7.5 mL of the media where microorganisms grew was placed in a 20-mL headspace vial joint to 1.5 g of NaCl and 10 μL of internal standard (4-methyl-2-pentanol, 0.75 mg/L). An MPS Autosampler (Gerstel, USA) incubated the vial for 10 min at 45 °C with agitation at 300 rpm. Then, a 2-cm 50/30 μm Carboxen/DVB/PDMS SPME fiber (Supelco, USA) was exposed to the headspace of the vial for 40 min. Afterward, fiber was desorbed in the injector in a splitless mode for 3 min with the transfer line at a temperature of 250 °C. Analyses were performed in an Agilent 8890 GC system coupled to an Agilent 5977B Inert Plus quadrupole mass spectrometer with a Gerstel autosampler (Müllheim an der Ruhr, Germany). The capillary column and flow rate employed were the same as the analysis. The conditions were as follows: the oven temperature program started at 35 °C held for 4 min, followed by an increase to 220 °C at 2.5 °C/min held for 1 min.
For both analyses, electron ionization mass spectra data were recorded from m/z 29–300 in scan mode with an ionization voltage of 70 eV. All data were recorded using an MS ChemStation (Agilent Technologies, USA).
The VOC identification was done by comparing the mass spectra obtained from each molecule with the reference spectra of the NIST 98 software library and the literature data (Pherobase: www.pherobase.com; NIST Mass Spectrometry Data Center: https://webbook.nist.gov; LRI and Odour database: http://www.odour.org.uk/lriindex.html). When only the software identification was possible, it was treated as tentatively identified. The data showed in this work were expressed as the relative area concerning 4-methyl-2-pentanol (internal standard). The relative concentration was calculated by dividing the peak area of the target ion of each compound by the peak area of the target ion of the internal standard.
Statistical analysis
The data obtained from the SPME/GC/MS determination were subjected to multivariate analysis performing principal component analysis (PCA) with MetaboAnalyst (Canada).
Results
Volatile profile characterization of L. monocytogenes (CC388), E. faecalis, and S. aureus
The employment of the SPME technique allowed the identification of 78 VOCs present in thioglycollate media after the growth of the bacteria. It was found that the control medium, without bacteria, presented several volatile compounds (Table 1). They were not subtracted from the bacterial culture samples because it is also interesting to observe the decrease in the amount of these compounds in the media since it could be due to volatilization but also to their consumption by these pathogens, which would provide interesting information to understand their metabolism. The metabolic pattern of each microorganism gave rise to different amounts of VOCs of each chemical family found. This first approach shows that the VOC pattern observed in L. monocytogenes is closer to that of E. faecalis than that of S. aureus.
Thus, as can be observed in Fig. 1, in the culture of S. aureus, the aldehydes had decreased slightly concerning the control. In contrast, the other two bacteria showed significant aldehyde production, with E. faecalis being the most productive. In this study, L. monocytogenes was characterized by the production of alcohols, unlike E. faecalis and S. aureus, which did not vary their total amount concerning the control. Our results showed that all bacteria produce ketones, mainly due to acetoin (Table 1). Esters and compounds derived from pyrazine were also determined, detecting most of them in the control medium.
Volatile acids were the only chemical group not present in the control medium determined with SPME–GC–MS. E. faecalis, followed by L. monocytogenes, exhibited a remarkable production of volatile acids (Fig. 1). The production of volatile acids by S. aureus was significantly lower than the amounts detected in the other bacteria studied. The main difference relied on the short-chain fatty acid (SCFA) production. Thus, acetic, propanoic, and 2-methyl butanoic acids were not detected in the media of S. aureus. In addition, butanoic and hexanoic acids were determined in significantly lower amounts than in the E. faecalis and L. monocytogenes media.
Multivariate analysis: principal component analysis
A PCA including all the VOCs and the total sum of each chemical group was performed, as shown in Fig. 2 (84 variables), to interpret the results obtained from the volatile profile comparison of L. monocytogenes, E. faecalis, and S. aureus. The analysis determined eight principal components (PCs) that explained 87.4% of the total variance, with PC1 and PC2 accounting for 48.8% of the accumulated variance (Fig. 3) and permitting a significant separation of the samples. The location in the plane of the samples corroborates that the thioglycollate medium employed for the growth of these bacteria gives rise to similar volatile profiles for L. monocytogenes and E. faecalis, being different to the amounts and types of volatile metabolites resulting for S. aureus growth. Furthermore, it can be observed that acids were clearly correlated with E. faecalis and L. monocytogenes (Fig. 2).
Data scores and loading biplot on the plane of the first two principal components (PC1 against PC2) of the volatiles from L. monocytogenes CC388, E. faecalis, and S. aureus. Numeration of loadings correspond to each compound from the first column of Table 1
Discussion
Although both E. faecalis and L. monocytogenes produce aldehydes, it is interesting in the first bacterium since it has been little studied. Monedeiro et al. (2021) observed that the volatile profile pattern of E. faecalis was the most common, as it had few unique VOCs compared to Escherichia coli, Klebsiella pneumoniae, and Proteus mirabilis. It can be seen in this study that E. faecalis significantly produces benzaldehyde and nonanal, being the only bacterium that generates the latter aldehyde (Table 1), so nonanal could be a potential biomarker of its presence in the media.
The highest production of alcohols in L. monocytogenes was mainly due to 1-butanol (Table 1), a short-chain alcohol that was significantly increased in all clonal complexes of L. monocytogenes (Lepe-Balsalobre et al. 2022). This result contrasts with the findings of Yu et al. (2015) who did not observe 1-butanol among the VOCs produced by L. monocytogenes cultured in TSB. This fact might be due to the culture medium, described as one of the main factors defining the VOCs produced by a microorganism (Tait et al. 2014; Sharma et al. 2017). Isoamyl alcohols (2-methyl-1-butanol and 3-methyl-1-butanol) were only produced by L. monocytogenes and E. faecalis, again reflecting the proximity between their metabolic systems. This fact agrees with D’Angelo et al. (2020) on another bacterium of the same genus, Enterococcus faecium, where they observed that leucine was converted to the corresponding keto acid before being decarboxylated to 3-methylbutanal, which could later be reduced to 3-methyl-1-butanol or oxidized to 3-methylbutanoic acid.
The amount of methionol found in the culture medium of S. aureus was significantly higher compared to the other microorganisms. This increase, together with the disappearance of 3-(methylthio)propionaldehyde (methional) in the case of S. aureus, points to a reduction of 3-(methylthio)propionaldehyde to methionol by the alcohol dehydrogenase activity of this bacteria to metabolize the amino acid methionine. This enzymatic activity has been previously described in other bacteria, such as Oenococcus oeni (Vallet et al. 2009).
As mentioned above, the increase of ketones was mainly due to acetoin. The production of this compound by some microorganisms has been widely described through the acid-mixed fermentation pathway of pyruvate. Chen et al. (2017) pointed out that this ketone production indicated the presence of L. monocytogenes and S. aureus compared to Gram-negative bacteria in the TSB medium. Specifically, Yu et al. (2015) stated that acetoin was the key-compound discriminating samples inoculated with L. monocytogenes with an electronic nose. In our study, the highest amount of acetoin in the culture medium was reached by E. faecalis, followed by L. monocytogenes and S. aureus. This fact does not agree with that published by Filipiak et al. (2012), who reported that acetoin meets all the requirements to be a perfect biomarker of S. aureus and shows the great need for more research in this field due to the strong dependence on media and comparison with other bacteria. The acetoin formation causes these bacteria to test positive in the Voges Proskauer test, determining their ability to produce this compound from glucose by butanediol fermentation. This biochemical test is important for the identification of L. monocytogenes and useful as an indicator of its aerobic growth since acetoin is not produced under anaerobic conditions (Romick et al. 1996).
The amounts of pyrazine and compounds derived from it were very similar in L. monocytogenes and the control medium, unlike the results obtained by extracting with dichloromethane (Lepe-Balsalobre et al. 2022). This fact could be due to the different extraction methods since, in the study of the Listeria clonal complexes, the extraction was carried out with a polar solvent, which is a more aggressive technique and extracts volatile compounds, but also a lot of non-volatile molecules. Also, the extraction performed in the present study was at 45°C for 50 min, while in Lepe et al. (2022) it was done at room temperature, and maybe the extraction and/or formation of these compounds could be favored by high temperatures. In contrast, these compounds were higher in the case of S. aureus, mainly due to the increase of 2,5-dimethyl-3-isoamylpyrazine. This compound has been previously described in food matrices contaminated with S. aureus (Fang et al. 2021).
Regarding the esters, due to their lability to acid hydrolysis and evaporation, the apparent consumption by L. monocytogenes and E. faecalis was probably not due to their metabolism (Fig. 1). However, the data point to a production of esters by S. aureus, especially isoamyl butyrate and methyl decanoate (Table 1). The production of esters by S. aureus has already been described (Filipiak et al. 2012), but not that of these VOCs in particular, which seem to be biomarkers of the presence of this pathogen in this medium.
The excretion of volatile acids by E. faecalis has already been reported in milk (Delgado et al. 2002). However, the variety of acids determined in this study has not been described for L. monocytogenes. The volatile acid profile revealed that 2-methylbutanoic acid could be a potential biomarker of L. monocytogenes presence (Table 1). SCFAs such as acetic, butanoic, and hexanoic acids are commonly associated with anaerobic metabolism. Thus, L. monocytogenes shows a fermentative metabolism due to its beta-d-glucosidase activity and generates acids and diacetyl groups from sugars such as glucose. A large number of SCFAs have been described in various intestinal diseases, such as that caused by L. monocytogenes (Ishiguro et al. 2017; Di Cagno et al. 2009). However, it has also been reported that fatty acids in Listeria spp. have an important role in resistance against peptidoglycan hydrolases and regulation of virulence (Sun et al. 2012), so they could be involved in the strong pathogenicity and virulence that the CC388 clonal complex presented in the Spanish outbreak in 2019.
Some studies have reported the intolerance of S. aureus to SCFAs because they delay and even suppress their growth (Fletcher et al. 2022), and, therefore, this could explain the low contents found in their media. However, 3-methylbutanoic acid that has been previously reported as a unique volatile compound produced by S. aureus grown in TSB (Chen et al. 2017) and Brain Heart Infusion (BHI) broth (Tait et al. 2014) was not found in our experiments.
Multiple growth experiments in defined minimal media have shown that most L. monocytogenes strains require supplementation of the sulfur amino acids methionine and cysteine for growth. This need is explained by the fact that most L. monocytogenes genomes do not express the genes responsible for the sulfate reduction to sulfide, which is subsequently condensed with O-acetylserine to form cysteine (Sauer et al. 2019). The thioglycollate medium contains l-cystine, a dimer of two cysteines linked at their thiol functional groups through a disulfide bond. Some nitrogen and sulfur compounds, such as pyrrole and 2,4-dimethylthiazole, are probably consumed by bacteria. Others, such as 3 methyl-2-thiophenecarboxaldehyde, could be produced from sulfur sources through cysteine consumption (Walker and Schmitt-Kopplin 2021) or formed from the substrate through the Maillard reaction (cysteine + glucose) due to the heating process for 50 min at 45°C during extraction (Umano et al. 1995).
In conclusion, the volatile pattern in the thioglycollate medium observed in L. monocytogenes is more similar to that of E. faecalis than that of S. aureus. Potential biomarkers of L. monocytogenes in this medium could be 1-butanol and 2-methylbutanoic acid. In contrast, in the case of E. faecalis, the VOC most related to its presence and, therefore, a potential volatile biomarker could be nonanal. Lastly, potential biomarkers of S. aureus are isoamyl butanoate and methionol, although some pyrazines have also been associated with this bacterium.
The detection of these volatile biomarkers reveals the possible presence of a specific microorganism and opens the path for future research in biological samples. However, this is a first approach because biological samples contain, in most cases, multiple bacteria that interact with each other, being able to modify the emission of VOCs that they produce when isolated. Therefore, the next research will be focused on the confirmation using biological samples.
Data availability
All data can be provided by the corresponding author upon request.
References
Vila J, Álvarez-Martínez MJ, Buesa J, Castillo J (2009) Diagnóstico microbiológico de las infecciones gastrointestinales. Enfermedades infecciosas y microbiología clínica 27(7):406–411
Cantón R, Ruiz-Garbajosa P (2013) Infecciones causadas por bacterias grampositivas multirresistentes (Staphylococcus aureus y Enterococcus spp.). Enferm Infecc Microbiol Clin 31:543–551
CDC (Centers for Disease Control and Prevention) (2024) Listeria (Listeriosis). Available from: https://www.cdc.gov/listeria/index.html. Accessed 25 Sept 2023
Chen J, Tang J, Shi H, Tang C, Zhang R (2017) Characteristics of volatile organic compounds produced from five pathogenic bacteria by headspace-solid phase micro-extraction/gas chromatography-mass spectrometry. J Basic Microbiol 57:228–237
D’Angelo M, Martino GP, Blancato VS, Espariz M, Hartke A, Sauvageot N et al (2020) Diversity of volatile organic compound production from leucine and citrate in Enterococcus faecium. Appl Microbiol Biotechnol 104:1175–1186
Delgado S, Delgado T, Mayo B (2002) Technological performance of several Lactococcus and Enterococcus strains of dairy origin in milk. J Food Prot 65:1590–1596
Di Cagno R, Rizzello CG, Gagliardi F, Ricciuti P, Ndagijimana M, Francavilla R et al (2009) Different fecal microbiotas and volatile organic compounds in treated and untreated children with celiac disease. Appl Environ Microbiol 75:3963–3971
Dolch ME, Hornuss C, Klocke C, Praun S, Villinger J, Denzer W et al (2012) Volatile organic compound analysis by ion molecule reaction mass spectrometry for Gram-positive bacteria differentiation. Eur J Clin Microbiol Infect Dis 31:3007–3013
Fang S, Liu S, Song J, Huang Q, Xiang Z (2021) Recognition of pathogens in food matrixes based on the untargeted in vivo microbial metabolite profiling via a novel SPME/GC × GC-QTOFMS approach. Food Res Int 142:110213
Filipiak W, Sponring A, Baur MM, Filipiak A, Ager C, Wiesenhofer H et al (2012) Molecular analysis of volatile metabolites released specifically by Staphyloloccus aureus and Pseudomonas aeruginosa. BMC Microbiol 12:113
Fletcher JR, Villareal AR, Penningroth MR, Hunter RC (2022) Staphylococcus aureus overcomes anaerobe-derived short-chain fatty acid stress via FadX and the CodY regulon. J Bacteriol 204:e0006422
Gómez-Mejia A, Arnold K, Bär J, Singh KD, Scheier TC, Brugger SD et al (2022) Rapid detection of Staphylococcus aureus and Streptococcus pneumoniae by real-time analysis of volatile metabolites. iScience 25:105080
Ishiguro R, Tanaka N, Abe K, Nakajima M, Maeda T, Miyanaga A et al (2017) Function and structure relationships of a β-1,2-glucooligosaccharide-degrading β-glucosidase. FEBS Lett 591:3926–3936
Jiménez Manso A, Babich J, Sánchez Moreno MP, Fernández Valenti ML (2022) Ensayos microbiológicos en alimentos en brotes de transmisión alimentaria. Procedimiento de Microbiología Clínica. SEIMC. Available from: https://seimc.org/contenidos/documentoscientificos/procedimientosmicrobiologia/seimc-procedimiento78.pdf. Accessed 12 Sept 2023
Lamote K, Janssens E, Schillebeeckx E, Lapperre TS, De Winter BY, van Meerbeeck JP (2020) The scent of COVID-19: viral (semi-)volatiles as fast diagnostic biomarkers? J Breath Res 14:042001
Lepe-Balsalobre E, Rubio-Sánchez R, Ubeda C, Lepe JA (2022) Volatile compounds from in vitro metabolism of seven Listeria monocytogenes isolates belonging to different clonal complexes. J Med Microbiol 71(6):001553
Ministerio de Sanidad, Consumo y Bienestar Social, Gobierno de España (2019) Informe de fin de seguimiento del brote de listeriosis. Available from: 2023 https://www.sanidad.gob.es/profesionales/saludPublica/ccayes/alertasActual/listeriosis/docs/Informe_cierre_Listeriosis_20190927.pdf. Accessed 21 Sept
Monedeiro F, Railean-Plugaru V, Monedeiro-Milanowski M, Pomastowski P, Buszewski B (2021) Metabolic profiling of VOCs emitted by bacteria isolated from pressure ulcers and treated with different concentrations of Bio-AgNPs. Int J Mol Sci 22:4696
Nasir M, Bean HD, Smolinska A, Rees CA, Zemanick ET, Hill JE (2018) Volatile molecules from bronchoalveolar lavage fluid can ‘rule-in’ Pseudomonas aeruginosa and ‘rule-out’ Staphylococcus aureus infections in cystic fibrosis patients. Sci Rep 8:826
Reichert-Schwillinsky F, Pin C, Dzieciol M, Wagner M, Hein I (2009) Stress- and growth rate-related differences between plate count and real-time PCR data during growth of Listeria monocytogenes. Appl Environ Microbiol 75:2132–2138
Romick TL, Fleming HP, McFeeters RF (1996) Aerobic and anaerobic metabolism of Listeria monocytogenes in defined glucose medium. Appl Environ Microbiol 62:304–307
Sauer JD, Herskovits AA, O´Riordan MXD (2019) Metabolism of the Gram-positive bacterial pathogen Listeria monocytogenes. Microbiol Spectr 7:1–12
Sharma NK, Keerqin C, Wu SB, Choct M, Swick RA (2017) Emissions of volatile odorous metabolites by Clostridium perfringens - in vitro study using two broth cultures. Poult Sci 96:3291–3297
Storer MK, Hibbard-Melles K, Davis B, Scotter J (2011) Detection of volatile compounds produced by microbial growth in urine by selected ion flow tube mass spectrometry (SIFT-MS). J Microbiol Methods 87:111–113
Sun Y, Wilkinson BJ, Standiford TJ, Akinbi HT, O’Riordan MX (2012) Fatty acids regulate stress resistance and virulence factor production for Listeria monocytogenes. J Bacteriol 194:5274–5284
Tait E, Perry JD, Stanforth SP, Dean JR (2014) Bacteria detection based on the evolution of enzyme-generated volatile organic compounds: determination of Listeria monocytogenes in milk samples. Anal Chim Acta 848:80–87
Ubeda C, Lepe-Balsalobre E, Ariza-Astolfi C, Úbeda-Ontiveros JM (2019) Identification of volatile biomarkers of Giardia duodenalis infection in children with persistent diarrhoea. Parasitol Res 118:3139–3147
Ubeda C, Vázquez-Carretero MD, Luque-Tirado A, Ríos-Reina R, Rubio-Sánchez R, Franco-Macías E et al (2022) Fecal volatile organic compounds and microbiota associated with the progression of cognitive impairment in Alzheimer’s disease. Int J Mol Sci 24:707
Ulanowska A, Kowalkowski T, Hrynkiewicz K, Jackowski M, Buszewski B (2011) Determination of volatile organic compounds in human breath for Helicobacter pylori detection by SPME-GC/MS. Biomed Chromatogr 25:391–397
Umano K, Hagi Y, Nakahara K, Shyoji A, Shibamoto T (1995) Volatile chemicals formed in the headspace of a heated D-glucose/L-cysteine Maillard model system. J Agric Food Chem 43:2212–2218
Vallet A, Santarelli X, Lonvaud-Funel A, de Revel G, Cabanne C (2009) Purification of an alcohol dehydrogenase involved in the conversion of methional to methionol in Oenococcus oeni IOEB 8406. Appl Microbiol Biotechnol 82:87–94
Walker A, Schmitt-Kopplin P (2021) The role of fecal sulfur metabolome in inflammatory bowel diseases. Int J Med Microbiol 311:151513
Wen Q, Boshier P, Myridakis A, Belluomo I, Hanna HG (2020) Urinary volatile organic compound analysis for the diagnosis of cancer: A systematic literature review and quality assessment. Metabolites 11:17
Yu Y, Sun X, Liu Y, Pan Y, Zhao Y (2015) Odor fingerprinting of Listeria monocytogenes recognized by SPME-GC-MS and E-nose. Can J Microbiol 61:367–372
Acknowledgements
The authors would like to thank the Virgen del Rocío University Hospital Microbiology Laboratory for supplying the samples used in this study, as the “VI Plan Propio de Investigación y Transferencia” of the University of Seville for the current contract of Dr. Cristina Ubeda (USE-18644-Z).
Funding
Funding for open access publishing: Universidad de Sevilla/CBUA.
Author information
Authors and Affiliations
Contributions
Ricardo Rubio Sánchez and Esperanza Lepe Balsalobre contributed to sample analysis and manuscript writing. Cristina Ubeda contributed to conception and design, data analysis and, interpretation and manuscript writing. José Antonio Lepe contributed to conception and design and sample collection.
Corresponding author
Ethics declarations
Consent for publication
All authors have read and approved the manuscript for publication.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rubio-Sánchez, R., Lepe-Balsalobre, E., Ubeda, C. et al. Volatile biomarkers of Gram-positive bacteria of clinical relevance as a tool for infection diagnosis. Int Microbiol (2024). https://doi.org/10.1007/s10123-024-00511-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10123-024-00511-z