Abstract
Emerging evidence suggests that the APOBEC family is implicated in multiple cancers and might be utilized as a new target for cancer detection and treatment. However, the dysregulation and clinical implication of the APOBEC family in clear cell renal cell cancer (ccRCC) remain elusive. TCGA multiomics data facilitated a comprehensive exploration of the APOBEC family across cancers, including ccRCC. Remodeling analysis classified ccRCC patients into two distinct subgroups: APOBEC family pattern cancer subtype 1 (APCS1) and subtype 2 (APCS2). The study investigated differences in clinical parameters, tumor immune microenvironment, therapeutic responsiveness, and genomic mutation landscapes between these subtypes. An APOBEC family-related risk model was developed and validated for predicting ccRCC patient prognosis, demonstrating good sensitivity and specificity. Finally, the overview of APOBEC3B function was investigated in multiple cancers and verified in clinical samples. APCS1 and APCS2 demonstrated considerably distinct clinical features and biological processes in ccRCC. APCS1, an aggressive subtype, has advanced clinical stage and a poor prognosis. APCS1 exhibited an oncogenic and metabolically active phenotype. APCS1 also exhibited a greater tumor mutation load and immunocompromised condition, resulting in immunological dysfunction and immune checkpoint treatment resistance. The genomic copy number variation of APCS1, including arm gain and loss, was much more than that of APCS2, which may help explain the tired immune system. Furthermore, the two subtypes have distinct drug sensitivity patterns in clinical specimens and matching cell lines. Finally, we developed a predictive risk model based on subtype biomarkers that performed well for ccRCC patients and validated the clinical impact of APOBEC3B. Aberrant APOBEC family expression patterns might modify the tumor immune microenvironment by increasing the genome mutation frequency, thus inducing an immune-exhausted phenotype. APOBEC family-based molecular subtypes could strengthen the understanding of ccRCC characterization and guide clinical treatment. Targeting APOBEC3B may be regarded as a new therapeutic target for ccRCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Renal cell carcinoma (RCC) is one of the most common and lethal cancers in the urogenital system and originates from epithelial cells [1]. According to Global Cancer Statistics reports, more than 431,000 new cases and approximately 180,000 deaths are documented from RCC worldwide [2]. Clear cell renal cell carcinoma (ccRCC) is the most common type of RCC, accounting for nearly 80% of RCC cases [3]. Traditional radiotherapy and chemotherapy are invalid for metastatic renal cancer, which leads to limited choices for RCC treatment. Targeted therapy and immunotherapy have brought some hope for RCC patients; however, only some patients respond well, and most of them will develop drug resistance [4]. Notably, somatic mutation is one of the most important characteristics for tumor progression and drug resistance. Considering the chromosome 3p loss directly decide aggressive phenotype in ccRCC, which induced the mutation of tumor drive including VHL, PBRM1 and SET2, a comprehensive understanding of somatic mutations and mutational progression might facilitate the revolutionary treatment of ccRCC. The development and validation of large-scale sequencing data offers the possibility to decipher mutational signatures. Among various mutation features, APOBEC (apolipoprotein B mRNA editing catalytic polypeptide-like) family members play an important role in DNA modification mutations.
The APOBEC family, containing 11 members, was gradually identified as an exogenous mutation factor in human cells [5]. The main biological function of the APOBEC family is related to DNA replication and genomic mutation [6]. Since the APOBEC family plays a role in the transcription process, dysregulation of the APOBEC family could induce mutations in genomic stability, thus promoting tumorigenesis and progression [7]. Recent studies have revealed the role of the APOBEC family in cancers. Serebrenik [8] found that APOBEC3B was overexpressed in clear cell ovarian carcinoma and could induce DNA damage, thus sensitizing tumor cells to additional genotoxic stress by cisplatin. Law [9] indicated that inhibiting APOBEC3B activity could reverse therapy resistance in breast cancer. Faden [10] also found that APOBEC mutagenesis was tightly correlated with tumor immune evasion and the ICI response rate in head and neck squamous cell carcinoma. Robertson [11] found that bladder cancer progression was associated with APOBEC mutations, and APOBEC signature mutations accounted for an important proportion of the total mutation load. Subsequent studies have also shown that APOBEC-associated mutations were significantly associated with invasive bladder cancer and poor patient outcomes [12]. Tan [13] found APOBEC3A functional polymorphisms played an important role in the progression of renal cell carcinoma. Even with the increasing knowledge of the APOBEC family in cancers, a comprehensive understanding is still limited. In addition, the function of the APOBEC family mutations in the progression of ccRCC remains largely elusive.
In this study, we carried a systematic analysis of the APOBEC family to facilitate the risk stratification of ccRCC, which included molecular features, biological function, immune landscape, and agents’ sensitivity. Based on remodeling algorithm, two distinctive ccRCC subgroups were identified and further tested, and a novel APOBEC related risk model was constructed based on each subtypes’ biomarkers. This study also encouraged the combination treatment of anti-angiogenesis and ICI for patients in APCS1.
Materials and methods
Data collection and processing
The pipeline was summarized in Fig. S1. We obtained pancancer datasets at multi omics from The Cancer Genome Atlas database with the use R package TCGAbiolinks [14]. Another ccRCC dataset were collected from the Express-array database with the accession number of E-TAB-1980. Patients enrolled in this work without prognostic information or expression profiles were excluded. All fragments per kilobase per million mapped fragments (FPKM) were transformed into Transcripts per million (TPM) for further analysis (TPM = (FPKM / sum of all FPKM) * 1,000,000).
Identification of distinct APOBEC family-based subgroups
We summarized 11 APOBEC family member according to prior studies and reviews (Table S1). After filtering ccRCC patients with complete APOBEC family member expression profiles and clinical outcomes, we used the R package ConsensusClusterPlus to perform unsupervised clustering. Two parameters, including principal score and silhouette statistic, were adopted to identify optimal number.
Enrichment analysis between subgroups
We used the R package DEseq2 to calculate differentially expressed genes (DEGs) between APCS1 and APCS2, with thresholds settled at an adjusted p value < 0.01 and an abstract log fold change > 2 [15]. After identifying those DEGs, we analyzed the potential biological processes involving those signatures, including GO, KEGG, GSEA and GEVA [16]. All annotation files for DEGs to explain biological function were extracted from ConsensusPathDB and MSigDB [17, 18].
Differences in immune landscape and ICI sensitivity
We adopted several classic deconvolution algorithms to quantify the immune cell infiltration degree between APCS1 and APCS2 [19,20,21,22]. In addition, ssGSVA was introduced to compare the immune components’ difference from Cibersort [23]. Three classic immune score based on immune and stromal components were quantified by R package ESTIMATE. The immune check point (ICI) therapy sensitivity of each ccRCC patient was evaluated by TIDE website, which consisted of several parameters, including immune exhaustion and exclusion scores [24].
Mutation spectrum characteristics between subgroups
To investigate the impact of APOBEC on genomic mutation of ccRCC, we used R package Maftools to quantify each subtypes mutation profile and frequency [25]. The coexisting and exclusive mutation landscape and subtype’s onco-pathway were investigated via functions of Maftools and visualized by ggplot2. With the use of GISTIC 2.0, the region gain and loss frequency between subtypes were compared [26] based on Euclidean distance [27].
Drug susceptibility prediction
Based on the expression profile of ccRCC patients, we calculated each patients’ sensitivity to molecular and pre-clinical drugs from Genomics of Cancer Drug Sensitivity (GDSC) database [28]. R package pRRophetic was adopted to estimate IC50 and cross validate the sensitivity difference [29]. In addition, two drug therapy sensitivity databases, CellMiner and CCLE, containing IC50 informations of cancer cell lines, were utilized to further detect potential agents [30].
Construction of a risk prediction model based on subtype biomarkers
The genes for risk model construction were retrieved from biomarkers of APCS1 and APCS2. We next applied RSRVH to identify most important biomarkers to induce the variables adopted for model construction, and the details could be seen in our previous works [31,32,33]. Finally, TCGA-KIRC and JAPAN-KIRC were utilized as the training and test cohorts, respectively, to judge the accuracy of the risk model.
Validation of APOBEC3B expression level in the Changhai cohort
All RNA were extracted from tissues of 35 ccRCC patients. RT‒qPCR was adopted to validate APOBEC3B expression in paired tumor and adjacent normal tissues. The Primer sequence information of APOBEC3B was as follows: F: CGCCAGACCTACTTGTGCTAT, R: CATTTGCAGCGCCTCCTTAT (Sangon Biotech).
Statistical analysis
All statistical analysis was finished by R software (Version 4.2.2). The Kruskal‒Wallis and Wilcoxon tests were applied to compare quantitative variables; the Chi-square test was adopted to compare qualitative characteristics. Correlations index of spearman or pearson among variables were calculated by R package corrplot. The R packages survival and pROC were used to depict Kaplan‒Meier and time ROC curves. Only p values < 0.05 (adjust by FDR) in this work were treated as statistically significant different.
Results
Genomic variation landscape of the APOBEC family in ccRCC
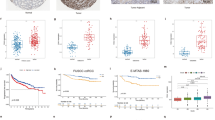
The workflow of this study was illustrated in Fig. S1. Combined with expression and prognostic information of pancancer, we found that most APOBEC family members correlated with inferior prognostic value in ccRCC and LGG and a protective role in BRCA, CESC and HNSC (Fig. 1A). Most APOBEC family genes displayed distinctive expression patterns between normal and cancer tissues at the pancancer level. Figure 1B indicated that APOBEC2 was downregulated, while APOBEC3B was upregulated in LUAD, LUSC, and ESCA et al. We also found that DNA methylation was significantly negatively correlated with APOBEC3G, APOBEC3C and APOBEC3D in most cancer types, which might help explain the aberrant expression pattern of the APOBEC family (Fig. 1C). For copy number variations (CNVs), APOBEC4 had the highest variation rate among all APOBEC family members (Fig. 1D). Regarding the correlation of expression and copy number variation, we concluded that APOBEC3C and APOBEC3F were significantly regulated by genomic variation (Fig. 1E, Fig. S2A). Heterozygous amplifications frequently presented in APOBEC4 and APOBEC1, while heterozygous deletions often appeared in APOBEC3H, 3G, 3F, 3D, 3C and 3A (Fig. S2B). In addition, we investigated the genomic location of the APOBEC family, which indicated that three APOBEC families, APOBEC3A, APOBEC3B and APOBEC3C, are located at chromosome 22 (Fig. 1F). We also analyzed the mutation frequency of APOBEC family and found that those mutated frequently in UCEC, READ, COAD and UCS (Fig. S2C). Finally, we detected the correlation of drug sensitivity and expression level among the APOBEC family, which revealed that most drugs were negatively correlated with the APOBEC family (Fig. S2D). These results reminded us that aberrant APOBEC family expression could induce therapy resistance in cancers.
Dysregulation and genome alteration of the APOBEC family in cancers A Impact of the APOBEC family on cancer patient survival. B Difference expression level of the APOBEC family between tumor and normal tissues. C Impact of Correlation of DNA methylation and APOBEC expression level. D–E The CNV frequency and correlation of CNV with gene expression of the APOBEC family. F The genome locations of the APOBEC family on human chromosomes
Identification of two distinctive clusters by APOBEC family expression pattern
Since the APOBEC family displayed a significant prognostic impact across various cancers, we aimed to decode their potential role in ccRCC. We first filtered prognostics related APOBEC family members in ccRCC, which included APOBEC3H, APOBEC3G, APOBEC3D, APOBEC3C and APOBEC3B. Then, we identified the optimal cluster number using ConsensusClusterPlus. Consequently, TCGA ccRCC samples were significantly remodeled into two distinct clusters, termed APOBEC expression pattern cancer subtype 1 (APCS1) and subtype 2 (APCS2) (Fig. 2A–B). The clinical outcomes and clinicopathological features of the two subtypes were compared and shown in Fig. 2C and Table S2. Compared to the APCS2 subgroup, the APCS1 subgroup had a higher TNM stage, shorter overall survival (OS), and progression-free interval (PFI). In addition, we investigated the expression levels of the APOBEC family among the two subtypes and normal tissues. APCS1 was treated as an APOBEC hot or significantly activated subtype, in which APCS1 displayed higher APOBEC family expression levels (Fig. 2D).
Functional enrichment analysis of subtypes
Considering the distinct clinicopathological features between subtypes, we next investigated DEGs between APCS1 and APCS2 (Fig. 3A). GO analysis indicated that DEGs were enriched in monovalent inorganic cation homeostasis, anion transmembrane transport, regulation of pH, cellular monovalent inorganic cation homeostasis and sodium ion transport in the biological process module; apical plasma membrane, vacuolar proton − transporting V − type ATPase complex, and apical part of cell in the cellular component module; and anion transmembrane transporter activity, active ion transmembrane transporter activity, chloride transmembrane transporter activity and inorganic anion transmembrane transporter activity in the molecular function module. (Fig. 3B, Fig. S3A). The KEGG analysis demonstrated that cell adhesion molecules, phagosome, chemokine signaling pathway, natural killer cell-mediated cytotoxicity and cytokine − cytokine receptor interaction were activated in APCS1 (Fig. S3B). GSEA further demonstrated that Adaptive Immune System, Cytokine Signaling in Immune system, Hemostasis, Signaling by Interleukins and Signaling by Rho GTPases were activated in APCS1 (Fig. 3C). In addition, GSVA showed that APCS1 led to an activated fat embolism phenotype, since adipogenesis and fatty acid metabolism were activated compared with APCS2 (Fig. 3D). To further investigate the regulatory mechanism between subgroups, we applied the R package RTN to compare classic transcription factor regulon score. APCS1 displayed higher activity in HNF4A, HNF1A, HNF1B, EPAS1 and ZEB2, while APCS2 contained a higher activity in FOXE1, TBX18 and TP53 (Fig. 3E).
Potential immune escape mechanism between subtypes
In this part, we questioned whether the effect of APOBEC on ccRCC could reshape immune infiltration and induce immune heterogeneity. Interestingly, nearly all APOBEC family members were positively correlated with immune infiltration in ccRCC, especially APOBEC3H, APOBEC3G, APOBEC3D and APOBEC3B (Fig. 4A). We found different immune-related signature expression levels between APCS1 and APCS2. APCS1 contained higher levels of chemokine, chemokine receptor, MHC, immune-inhibitor and immune-stimulator factors (Fig. 4B). In detailed immune cell infiltration, we found that APCS1 displayed an immune hot microenvironment, since most immune cell enrichment scores were higher in APCS1. It should be mentioned that CD4 + T cells, mast cells, NK cells, neutrophils and endothelial cells were more highly infiltrated in APCS2 (Fig. 4C). Similar results were also found in immune-related signature-based GSVA analysis (Fig. 5A, B). The immune checkpoint inhibitor (ICI) therapy response in APCS1 was lower than that in APCS2 (Fig. 5C).
Accordingly, APCS1 displayed an increased stromal, immune and ESTIMATE score compared to APCS2 via the ESTIMATE package (Fig. 6A). Tumor immunity involves several sequential steps, and a successive presentation determines the efficacy of immunity. Thus, we then analyzed the difference in the anticancer immune response between subtypes, which was involved in multiple steps of immune events. Interestingly, the APCS1 subtype was deficient in monocyte recruitment and immune-induced tumor cell killing processes (Fig. 6B). Combined with the clinical outcome and immune infiltration described above, we found a paradox in tumor immunity in APCS1. Nearly all immune inhibitory regulators were higher expressed in APCS1 (Fig. 6C). There was no significant difference in TMB between subtypes (Fig. 6D). The above findings suggested that the overexpression level or aberrant active state of APOBEC signatures might induce an immune exhaustion state in ccRCC.
Comprehensive and integrated genomic characteristics between subtypes
In addition to transcriptional alterations, we also investigated the disparity in the genome mutation level. We observed that the APCS1 subtype presented a higher mutation rate than APCS2 (87.97% vs 82.89%). The waterfall charts showed the top 20 most frequently mutated genes between the subtypes (Fig. 7A). Specifically, the mutation frequency of VHL, SETD2, BAP1 in APCS1 was higher than APCS2 (56% vs. 44%; 17% vs. 9%; 12% vs. 9%), while PBRM1 was lower than APCS2 (40% vs. 44%). Based on gene mutation, we utilized the DGIdb database and drug interactions in the Maftools package to investigate probable targets for each subtype. The top 3 druggable targets for APCS1 were ADGRV1, ATM, BAP1, FBN2 and HMCN1 (druggable genome); ATM, BAP1, KDM5C, PBRM1 and PTEN (clinically actionable); and ATM, BAP1, KDM5C and SETD2 (histone modification). The top 3 druggable targets for APCS2 were ARID1A, ATM, BAP1, ERBB4 and KDM5C (clinically actionable); ATM, BAP1, ERBB4, KDM5C, and MTOR (druggable genome); and AKAP9, ATM, BAP1, MTOR, and SETD2 (tumor suppressor) (Fig. 7B). The somatic interaction analysis found co-mutation models of SPEN-PTEN, PLXNA4-MUC16, MYCBP2-HMCN1, DNAH10-PBRM1, USH2A-KDM5C, LRP2-SPTA1, ADGRV1-HMCN1, SPTA1-HMCN1 and MUC16-BAP1 in APCS1 and FAT1-DNAH9 and PBRM1-VHL in APCS2 (Fig. S4A). Furthermore, we used the Maftools to explore the somatic mutations in onco-pathways. We found that NRF2 was more affected in APCS1, while RTK − RAS and NOTCH were easily affected in APCS2 (Fig. 7C). Interestingly, several mutation events exerted different prognostic effects on APCS1 and APCS2; for instance, SETD2 and VHL displayed risk roles in APCS2 when compared with APCS1 (Fig. S4B).
Copy number variations were also compared between subtypes. Overall, APCS1 displayed higher amplification and deletion frequencies than APCS2 (Fig. 8D). GISTIC software was used to visualize the CNVs (Fig. 8A and B), and found the gain/loss events showed similar patterns. Of interest, both the APCS1 and APCS2 subgroups contained frequent CNVs in the region of Chr3 and Chr5. The recurrent CNVs of APCS1 included amplification of 5q35.3 (DBN1, DRD1, GPK6, GRM6) and 1q32.1 (KISS1, MDM4, PIK3C2B) and deletion of 9p23 (PTPRD) and 9p21.3 (CKN2B). The repeated CNVs of APCS2 were the amplification of 5q35.1 (STC2) and 5p31.3 (KCTD16) and deletion of 2p37.3 (AGXT, KIF1A, BOK) and 9p21.3 (CDKN2A, CDKN2B). The distinctive CNV landscape might be induced by different APOBEC family expression patterns between subgroups. In detail, APCS1 displayed higher genome amplification at 1p, 1q, 2p, 5p, 5q, 7p, 7q, 8q, 12p, 12q, 19q, 20p, and 22q and higher genome deletion at 3p, 4q, 6p, 6q, 8p, 9p, 9q, 13q, 14q, 18p and 18q, while APCS2 displayed higher genome deletion at 2p and 2q (Fig. 8C). All these findings suggest that aberrant APOBEC family members might cause genomic instability and copy number variation in the genome, as well as changes in oncogenesis pathways, especially RTK − RAS and NOTCH signaling, in ccRCC.
Drug sensitivity analysis and verification of remodeling results
The GDSC database was applied to explore potential therapeutic drugs for APCS1 and APCS2. We observed that APCS1 was sensitive to axitinib, imatinib, lisitinib, and sunitinib, while APCS2 was sensitive to dasatinib and saracatinib (Fig. 9A). In addition, we further evaluated the therapeutic response to 138 small molecular inhibitors between the two subtypes (Table S3). The top ten notable drugs are listed in Figure. The ACS1 subtype was sensitive to CI.1040, AICAR, camptothecin, RO.3306, vinblastine, VX.680, CGP.60474, bosutinib, and LFM. A13 and SL.0101.1, while APCS2 was sensitive to FH535, AS601245, Elesclomol, CCT007093, NSC.87877, Bicalutamide, Vinorelbine, SB590885, AZD.0530 and Bexarotene (Fig. 9B, C). All small molecule drugs provide new insights for the management of different risk stratification subgroups. To confirm the productivity of remodeling system, we used the NTP algorithm and found the heatmap of biomarkers maintained a high inner consistency (Fig. S5A). APCS1 also led to a poorer prognosis than the APCS2 subgroup (Fig. S5B).
Construction and validation of a subtype biomarker-based risk model
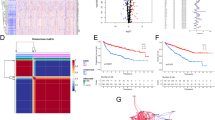
APOBEC family members could affect the prognosis of ccRCC patients, and APCS1 and APCS2 displayed a distinctive phenotype. Thus, we constructed a prognostic risk model based on subgroup biomarkers to aid prognostic prediction. We investigated dysregulated biomarkers between subtypes with the use of univariate Cox regression analysis (Fig. 10A). The random forest supervised classification algorithm helps us to screen the pivotal biomarkers (Fig. 10B). Next, we screened out a risk assessment model composed of five genes (PDIA2, PAEP, FOXI2, CEL and PI3), named ccRCC-AP5 (Fig. 10C). The risk score of each sample was calculated as follows: ccRCC-AP5 = 4.847704* PDIA2 + 6.452028* PAEP -2.850983* FOXI2 + 2.954473* CEL + 5.922656* PI3. To confirm the effectiveness of ccRCC-AP5, we divided patients in TCGA-ccRCC and JAPAN-KIRC into high- and low-risk groups by comparing median cut-off score (Fig. 10D). As expected, the high-risk group patients had worse OS and PFS than the low-risk group patients in both cohorts (Fig. 10E). The promising AUC score were achieved in TCGA-ccRCC cohort, and AUC score were 0.7132, 0.7323, 0.76988, 0.7166 and 0.6967 at 0.5, 1, 2, 3 and 5 year, respectively; the predictive value of ccRCC-AP5 model was also determined in JAPAN-KIRC cohort, and the AUC score were 0.7216, 0.7077, 0.787, 0.7307 and 0.8168 at 0.5, 1, 2, 3 and 5 year, respectively (Fig. 10F). The above results indicated that the ccRCC-AP5 model was reliable and applicable for the prognostic evaluation of ccRCC patients.
Establishment and verification of the subtype biomarker-based risk model A Volcano plot indicating the univariate Cox coefficient of filtered biomarkers. B Random survival forest analysis identifying the most important signatures. C Optimal combination of signatures. D Risk score distribution in TCGA- and JAPAN-KIRC cohorts. E–F Survival analysis of OS and PFI in the TCGA-ccRCC cohort. G Time-dependent ROC curves for different risk groups in the training and test datasets
The role of APOBEC3B in ccRCC
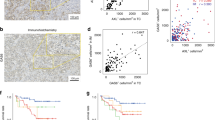
In view of the importance of APOBEC family members in ccRCC, we analyzed which gene led to the greatest proportion of importance. We found that APOBEC3B might be the most survival-related member in ccRCC patients through SVM analysis (Fig. 11A). At the protein level, APOBEC3B was higher expressed in tumor tissues, and its expression level increased with grade and stage system accordingly (Fig. 11B). APOBEC3B was more highly expressed in tumor tissues at the pancancer level (Fig. S6A). Since previous results indicated that APOBEC3B might be involved in ccRCC immune infiltration, we aimed to investigate the detailed mechanism of APOBEC3B in ccRCC. We found that the genomic alteration of APOBEC3B could be significantly related to the increased levels of immune checkpoints, DNA mismatch repair (MMR) regulators and DNA methylation transferases, especially in ccRCC (Fig. 11C–E). Furthermore, the genome mutation state of APOBEC3B could impact the infiltration levels of CD8 + T cells, CD4 + T cells, neutrophils and dendritic cells (Fig. 11F). APOBEC3B was involved in multiple signaling pathways in pancancer, including the MYC-target pathway, G2M-checkpoint and E2F-target (Fig. S6B). We found that high APOBEC3B expression predicted poor survival in TCGA-ccRCC patients (Fig. 11G). In addition, utilizing tissues from Changhai Hospital, we found that APOBEC3B was significantly increased in tumor samples compared with normal tissues (Fig. S7A), and ccRCC patients in our cohort with higher expression levels of APOBEC3B also had a poor prognosis (Fig. S7B). Combining all these results, it might be rational to elevate the immunotherapy response by activating APOBEC3B expression.
The role of APOBEC3B in ccRCC A Number of tree indicating the importance proportion of APOBEC family members. B Protein levels of APOBEC3B in CPTAC-KIRC, classified by tumor grade and histological pathological stage. C–E The association between APOBEC3B and the immune signature, MMR, and DNA methyltransferases at the pancancer level. (DNMT1: red, DNMT2: blue, DNMT3A: green, and DNMT3B: purple). ∗ P < 0:05, ∗ ∗ P < 0:01, and ∗ ∗ ∗ P < 0:001. F The correlation between APOBEC3B expression and clinicopathologic features. G The relationship between APOBEC3B and clinical characteristics in ccRCC
Discussion
The prognosis of ccRCC patients has been prolonged with the advancement of imaging technology and drug development, while intratumor heterogeneity impedes the precise and personalized management of ccRCC patients [26, 34]. In clinical practice, patients with similar clinical stages or pathological grade scores have distinctive clinical outcomes [35, 36]. Considering the characteristics of genomic instability and epigenetic modification of ccRCC, a promising prognostic model considering those traits is extremely urgent. The frequent occurrence of targeted and immune drug resistance is inextricably linked to genomic mutations. The APOBEC family, related to cytidine deaminase and targeting a single DNA strand, mainly induces the generation of C > T mutations, which mainly function to restrict viral infection [5]. Recent studies have redefined the role of the APOBEC family, which might be involved in cancer initiation and progression through its off-target effect [37]. Therefore, exploring the potential biological impact of the APOBEC family ccRCC might facilitate precise management.
This study identified and validated two distinctive APOBEC subgroups in ccRCC. APCS1 displayed high expression level of APOBEC family members combined with immune cells and signatures significantly enriched, which was termed as APOBEChot and immunehot subtypes. APCS2 displayed a lower level of APOBEC family members and turned to an immune cold microenvironment, which was identified as the APOBEClow and immunelow subtype. To verify our remodeling result, we identified subtype signatures and utilized NTP algorithms to perform remodeling analysis in the KIRC-Japan cohort, which reached high reproducibility and accuracy. Furthermore, based on APCS1- and APCS2-related biomarkers, we constructed and verified a prognostic model, AP-RCC5, which facilitated the risk stratification of ccRCC patients and functioned as an independent prognostic factor. These results shed light on a better understanding of APOBEC in cancers and the precise treatment of ccRCC.
The APOBEC family plays a critical role in regulating the progression and metastasis of cancer by inducing mutations. In addition, these mutations are associated with the dysregulation of various signaling pathways and the immune microenvironment. Several studies have proven that overexpression of APOBEC3A could trigger cell cycle arrest, DNA chain breakage and DNA damage response effects in tumors [38]. A prime example is APOBEC3B, which is responsible for causing base substitutions in the host genome, thereby inducing carcinogenic effects across multiple cancer types, including lung cancer, breast cancer, multiple myeloma, and urothelial cancer [39,40,41]. Based on the expression and mutation characteristics of the APOBEC family, patients were divided into two subtypes. In this work, compared with APCS2, the APCS1 subtype exhibited more aggressiveness and poorer prognosis of patients. We discovered that APCS1 subtype was enriched in phagosomal and chemokine signaling pathways. Jun Nishida found that inflammatory chemokines drove lung metastasis and initiated a metastatic cascade in advanced renal cancer [42]. The APCS1 subtype was characterized by a higher immune component and an immune infiltrate signature, while also possessing immunosuppressive regulators. This paradox of tumor immunity was inherent in the unique immune microenvironment of RCC. Previously, it was believed that high immune infiltration predicted poor prognosis of RCC patients, and the presence of immunosuppressive regulators further impeded the effectiveness of RCC immunotherapy. Mullane [43] found that increased APOBEC3A, APOBEC3D and APOBEC3H expression in urothelial carcinoma was correlated with PD-L1-positive tumor infiltrating mononuclear cell expression. Kobe C Yuen's study revealed that a high level of inflammatory immune microenvironment was associated with atezolizumab insensitivity in metastatic RCC, even for CD8 + T cells [44]. APCS1 subtypes exhibited higher activity of oncogenic transcription factors, which also provided a molecular basis for the high oncogenicity of APCS1 subtypes. Thus, immune disturbances induced by the tumor microenvironment led to poorer responses of APCS1 subtypes to immune checkpoint inhibitors.
Previous works also reminded us that the activity or expression level of APOBEC family could impact sensitivity to ICI therapy [7]. Wang found that APOBEC family members displayed a protective role in non-small cell lung cancer (NSCLC) and predicted durable clinical benefit for advanced NSCLC patients [39]. Faden [10] also observed that APOBEC members could induce IFNγ release thus enhance tumor immunity. In this work, we found APCS1, the APOBEChot subtype, led a low response to ICI therapy even with high immune scores. One reason for the adverse effect of APOBEC turns to that over expression APOBEC family could generate a library of constant mutated antigen, which induced immune exhaustion via a long term of chronic inflammation state [7]. McGral and colleagues [45] also observed that the accumulation of tumor mutation burden caused by APOBEC family failed to predict immunotherapy response. A work from Xia [46] also reminded us that APOBEC3B indicated an inferior prognosis of gastric cancer, and conversely related with CD8+ T cell infiltration. Based on the higher enrichment score of angiogenesis and immune infiltration in APCS1, we hypothesized that the combination of anti-angiogenesis and ICI might reach a better clinical benefit for such group. And our hypothesis is consistent with the standard of care for some advanced ccRCC patients, which suggested that the combination of VEGFR-TKI + ICI work well in progressed ccRCC. The expression of APOBEC family might be one of important characteristics for such group.
Differences in immunotherapy were also associated with gene mutations in APOBEC family-induced subtypes. In our study, the APCS1 subtype flagged more frequent mutations in genes such as VHL, SETD2, and BAP1. VEGFR-targeted inhibitors developed relying on the VHL-HIF pathway are one of the most important treatments for RCC. Xie [47] reported that loss of SETD2 created a suitable epigenetic environment for the amplified transcriptional output of oncogenic drivers. Censoring of BAP1 promoted the biological process of mesenchymal-epithelial transition in renal cancer [48]. Coordinated targeting of the APOBEC family may yield more substantial benefits for therapies that target these genes. Regarding copy number variation, APCS1 showed a higher frequency of amplifications and deletions than APCS2. Fernandes [49] discovered that the most significant somatic CNVs were losses of 3p, 14p, and 6q, as well as gains of 5q and 16q, which aligned with our findings. Therefore, abnormal APOBEC family members might lead to genome instability and copy number variation to promote tumor progression in ccRCC.
Growing evidence indicated that members of the APOBEC family affected tumor effectiveness to various drugs. Different APOBEC family-related subtypes showed varying sensitivities to drugs. Our exploration into molecular inhibitors targeting both subtypes revealed that the aggressive subtype APCS1 demonstrated heightened sensitivity toward the inhibitors CI.1040, AICAR, and camptothecin. CI.1040, which is a small-molecule inhibitor targeting the dual-specificity kinases MEK1 and MEK2, has been proven to suppress pancreatic cancer growth and has significant antitumour activity [50]. AICAR is an AMPK agonist, and Su found that AICAR could induce cancer cell apoptosis, attenuate TGF-β-induced tumor cell invasion, and promote chemosensitivity to docetaxel in prostate cancer cells via the AMPK/mTOR pathway [51, 52]. Luo and colleagues [53] reported a novel drug based on photosensitizer-conjugated CPT, which could cause cancer cell apoptosis through accumulating reactive oxygen species. We proposed that the combination of those drugs with target or immunotherapy agents could achieve synergistic effects in ccRCC patients.
Finally, based on distinctive clinical outcomes and biological heterogeneity between APCS1 and APCS2, we constructed an APOBEC family-related prognostic signature for ccRCC. Five new biomarkers, including three risk factors and two protective factors (PDIA2, PAEP, FOXI2, CEL, PI3), were identified and tested. In addition to showing better predictive performance, several signatures have been reported in previous studies. Zou [54] found that the PIDA2 expression level was positively correlated with tumor progression and the clinical behavior of gallbladder carcinoma. FOXI2, Forkhead Box I2, can enhance DNA-binding transcription factor activity. The aberrant DNA methylation state of FOXI2 was proven to be associated with the prognosis of multiple cancers, including oral cancer, pancreatic ductal adenocarcinoma, colorectal cancer, and esophageal adenocarcinoma [55,56,57]. All five biomarkers were not previously reported in ccRCC, and we found that those biomarkers could affect ccRCC patient prognosis through APOBEC family members via bioinformatics approaches. Wang and colleagues proved that upregulated APOBEC3B in non-small cell lung cancer could predict poor prognosis, while was positively correlated with immunotherapy response [39]. Law’s [58] study found that overexpression of APOBEC3B promoted tamoxifen resistance in ER-breast cancer. Xu [59] reported that high APOBEC3B expression was a predictor of recurrence in patients with low-risk ccRCC. Therefore, targeting APOBEC3B may enhance the efficacy of immunotherapy, though further experiments are required to confirm those findings. Nevertheless, this research also had several limitations. First, owing to the lack of clinical outcome information, our study only enrolled public ccRCC datasets and did not verify the results in our in-house dataset. Second, although we applied some algorithms to predict sensitivity of APCS1 and APCS2 to different molecular drugs, prospective cohort studies and clinical data are still needed.
In conclusion, this study systematically decoded the aberrance and implication of the APOBEC family at the pancancer level, especially in ccRCC. Two distinctive APOBEC expression pattern ccRCC subtypes, APCS1 and APCS2, were identified and verified. Subtypes displayed different clinical characteristics, immune infiltration landscapes, mutation models and drug sensitivities. A potential biomarker, APOBEC3B, was illustrated in ccRCC progression. All the findings in this study might assist urologists in managing ccRCC patients and shed light on the tumorigenesis and progression of renal cancer.
Data availability
No datasets were generated or analyzed during the current study.
References
Capitanio U, Montorsi F. Renal cancer. Lancet. 2016;387(10021):894–906.
Siegel RL, et al. Cancer statistics, 2022. CA: Cancer J Clin. 2022;72(1):7–33.
Aw SJ, Chang KTE. Clear Cell Sarcoma of the Kidney. Arch Pathol Lab Med. 2019;143(8):1022–6.
Saad AM, et al. Trends in renal-cell carcinoma incidence and mortality in the United States in the last 2 decades: a SEER-based study. Clin Genitourin Cancer. 2019;17(1):46–57.
Harris RS, Dudley JP. APOBECs and virus restriction. Virology. 2015;479–480:131–45.
Yang B, et al. APOBEC: From mutator to editor. J Genet Genomics. 2017;44(9):423–37.
Vile RG, et al. APOBEC and cancer viroimmunotherapy: thinking the unthinkable. Clin Cancer Res. 2021;27(12):3280–90.
Serebrenik AA, et al. The DNA cytosine deaminase APOBEC3B is a molecular determinant of platinum responsiveness in clear cell ovarian cancer. Clin Cancer Res. 2020;26(13):3397–407.
Ek L, et al. The DNA cytosine deaminase APOBEC3B promotes tamoxifen resistance in ER-positive breast cancer. Sci Adv. 2016. https://doi.org/10.1126/sciadv.1601737.
Faden DL, et al. APOBEC mutagenesis is tightly linked to the immune landscape and immunotherapy biomarkers in head and neck squamous cell carcinoma. Oral Oncol. 2019;96:140–7.
Robertson AG, et al. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell. 2017;171(3):540–56.
Lindskrog SV, et al. An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer. Nat Commun. 2021;12(1):2301.
Tan X, et al. Effect of APOBEC3A functional polymorphism on renal cell carcinoma is influenced by tumor necrosis factor-α and transcriptional repressor ETS1. Am J Cancer Res. 2021;11(9):4347.
Tomczak K, Czerwińska P, Wiznerowicz M. The cancer genome atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol. 2015;19(1A):A68-77.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
Yu G, et al. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics: J Integr Biol. 2012;16(5):284–7.
Liberzon A, et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics. 2011;27(12):1739–40.
Kamburov A, et al. ConsensusPathDB—a database for integrating human functional interaction networks. Nucleic Acids Res. 2009;37:D623–8.
Li T, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Can Res. 2017;77(21):e108–10.
Chen B, et al. Profiling tumor infiltrating immune cells with CIBERSORT. In: von Stechow L, editor., et al., Cancer systems biology: methods in molecular biology. New York: Humana Press; 2018. p. 243–59.
Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017;18(1):220.
Racle J, Gfeller D. EPIC: a tool to estimate the proportions of different cell types from bulk gene expression data. In: Boegel S, editor. Bioinformatics for cancer immunotherapy: methods in molecular biology. New York: Humana; 2020. p. 233–48.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7.
Jiang P, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–8.
Mayakonda A, et al. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018;28(11):1747–56.
Jiang A, et al. Establishment of a prognosis prediction model based on pyroptosis-related signatures associated with the immune microenvironment and molecular heterogeneity in clear cell renal cell carcinoma. Front Oncol. 2021;11:4486.
Mermel CH, et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011. https://doi.org/10.1186/gb-2011-12-4-r41.
Cokelaer T, et al. GDSCTools for mining pharmacogenomic interactions in cancer. Bioinformatics. 2018;34(7):1226–8.
Geeleher P, Cox N, Huang RS. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE. 2014;9(9):e107468.
Luna A, et al. Cell miner cross-database (CellMinerCDB) version 1.2: exploration of patient-derived cancer cell line pharmacogenomics. Nucleic Acids Res. 2020;49:D1083–93.
Jiang A, et al. Copper death inducer, FDX1, as a prognostic biomarker reshaping tumor immunity in clear cell renal cell carcinoma. Cells. 2023;12(3):349.
Jiang A, et al. A new thinking: deciphering the aberrance and clinical implication of copper-death signatures in clear cell renal cell carcinoma. Cell Biosci. 2022;12(1):209.
Jiang A, et al. A new thinking: deciphering the aberrance and clinical implication of IGF axis regulation pattern in clear cell renal cell carcinoma. Front Immunol. 2022. https://doi.org/10.3389/fimmu.2022.935595.
Sánchez-Gastaldo A, et al. Systemic treatment of renal cell cancer: a comprehensive review. Cancer Treat Rev. 2017;60:77–89.
Lalani A-KA, et al. Systemic treatment of metastatic clear cell renal cell carcinoma in 2018: current paradigms, use of immunotherapy, and future directions. Eur Urol. 2019;75(1):100–10.
Jiang A, et al. Establishment of a prognostic prediction and drug selection model for patients with clear cell renal cell carcinoma by multiomics data analysis. Oxid Med Cell Longev. 2022;2022:e3617775.
Knisbacher BA, Gerber D, Levanon EY. DNA editing by APOBECs: a genomic preserver and transformer. Trends Genet. 2016;32(1):16–28.
S, Revathi, et al. APOBEC: a molecular driver in cervical cancer pathogenesis. Cancer Lett. 2021. https://doi.org/10.1016/j.canlet.2020.10.004.
Wang S, et al. APOBEC3B and APOBEC mutational signature as potential predictive markers for immunotherapy response in non-small cell lung cancer. Oncogene. 2018;37(29):3924–36.
Natesan D, et al. APOBEC mutational signature and tumor mutational burden as predictors of clinical outcomes and treatment response in patients with advanced urothelial cancer. Front Oncol. 2022;12:816706.
Swanton C, et al. APOBEC enzymes: mutagenic fuel for cancer evolution and heterogeneity. Cancer Discov. 2015;5(7):704–12.
Nishida J, et al. Epigenetic remodelling shapes inflammatory renal cancer and neutrophil-dependent metastasis. Nat Cell Biol. 2020;22(4):465–75.
Mullane SA, et al. Correlation of Apobec Mrna expression with overall survival and pd-l1 expression in urothelial carcinoma. Sci Rep. 2016;6:27702.
Yuen KC, et al. High systemic and tumor-associated IL-8 correlates with reduced clinical benefit of PD-L1 blockade. Nat Med. 2020;26(5):693–8.
McGrail DJ, et al. High tumor mutation burden fails to predict immune checkpoint blockade response across all cancer types. Ann Oncol. 2021;32(5):661–72.
Xia S, et al. Immune inactivation by APOBEC3B enrichment predicts response to chemotherapy and survival in gastric cancer. Oncoimmunology. 2021. https://doi.org/10.1080/2162402X.2021.1975386.
Xie Y, et al. SETD2 loss perturbs the kidney cancer epigenetic landscape to promote metastasis and engenders actionable dependencies on histone chaperone complexes. Nature cancer. 2022;3(2):188–202.
Chen P, et al. Loss of BAP1 results in growth inhibition and enhances mesenchymal-epithelial transition in kidney tumor cells*[S]. Mol Cell Proteom. 2019;18(7):1320–9.
Fernandes FG, et al. Somatic copy number alterations and associated genes in clear-cell renal-cell carcinoma in Brazilian patients. Int J Mol Sci. 2021;22(5):2265.
Lorusso PM, et al. Phase I and pharmacodynamic study of the oral MEK inhibitor CI-1040 in patients with advanced malignancies. J Clin Oncol. 2005;23(23):5281–93.
Guo F, et al. AICAR induces AMPK-independent programmed necrosis in prostate cancer cells. Biochem Biophys Res Commun. 2016;474(2):277–83.
Su C-C, et al. AICAR induces apoptosis and inhibits migration and invasion in prostate cancer cells through an AMPK/mTOR-dependent pathway. Int J Mol Sci. 2019;20(7):E1647.
Luo X, et al. A camptothecin prodrug induces mitochondria-mediated apoptosis in cancer cells with cascade activations. Chem Commun. 2021;57(84):11033–6.
Zou Q, et al. Clinicopathological features and CCT2 and PDIA2 expression in gallbladder squamous/adenosquamous carcinoma and gallbladder adenocarcinoma. World J Surg Oncol. 2013;11:143.
Zhang Z, et al. Analysis of methylation-driven genes in pancreatic ductal adenocarcinoma for predicting prognosis. J Cancer. 2021;12(21):6507–18.
Chen Y, et al. Identification of prognostic risk model based on DNA methylation-driven genes in esophageal adenocarcinoma. Biomed Res Int. 2021;2021:6628391.
Foy JP, et al. New DNA methylation markers and global DNA hypomethylation are associated with oral cancer development. Cancer Prev Res. 2015;8(11):1027–35.
Law EK, et al. The DNA cytosine deaminase APOBEC3B promotes tamoxifen resistance in ER-positive breast cancer. Sci Adv. 2016;2(10):e1601737.
Xu L, et al. High APOBEC3B expression is a predictor of recurrence in patients with low-risk clear cell renal cell carcinoma. Urol Oncol: Semin Orig Invest. 2015. https://doi.org/10.1016/j.urolonc.2015.05.009.
Acknowledgements
We thank Sangerbox for generously sharing their bioinformatic related experience and codes.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Nos. 81902560, 81730073, 81872074), and the China National Key Research and Development Program Stem Cell and Translational Research Key
Author information
Authors and Affiliations
Contributions
Guiying Huang, Xinyou Xie and Jun Zhang designed the research. Guiying Huang and Xianlin Zhan contributed equally to this work. Xinyou Xie, Lihong Shen, Luping Lou, and Yuehong Dai assisted with data acquisition and data analysis. Guiying Huang, Aiming Jiang, Yuzhen Gao, and Yanzhong Wang wrote the manuscript. All authors contributed to the article and approved the final version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, G., Zhan, X., Shen, L. et al. APOBEC family reshapes the immune microenvironment and therapy sensitivity in clear cell renal cell carcinoma. Clin Exp Med 24, 212 (2024). https://doi.org/10.1007/s10238-024-01465-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10238-024-01465-2