Abstract
We aimed to develop and validate a deep learning-based system using pre-therapy computed tomography (CT) images to detect epidermal growth factor receptor (EGFR)-mutant status in patients with non-small cell lung cancer (NSCLC) and predict the prognosis of advanced-stage patients with EGFR mutations treated with EGFR tyrosine kinase inhibitors (TKI). This retrospective, multicenter study included 485 patients with NSCLC from four hospitals. Of them, 339 patients from three centers were included in the training dataset to develop an EfficientNetV2-L-based model (EME) for predicting EGFR-mutant status, and the remaining patients were assigned to an independent test dataset. EME semantic features were extracted to construct an EME-prognostic model to stratify the prognosis of EGFR-mutant NSCLC patients receiving EGFR-TKI. A comparison of EME and radiomics was conducted. Additionally, we included patients from The Cancer Genome Atlas lung adenocarcinoma dataset with both CT images and RNA sequencing data to explore the biological associations between EME score and EGFR-related biological processes. EME obtained an area under the curve (AUC) of 0.907 (95% CI 0.840–0.926) on the test dataset, superior to the radiomics model (P = 0.007). The EME and radiomics fusion model showed better (AUC, 0.941) but not significantly increased performance (P = 0.895) compared with EME. In prognostic stratification, the EME-prognostic model achieved the best performance (C-index, 0.711). Moreover, the EME-prognostic score showed strong associations with biological pathways related to EGFR expression and EGFR-TKI efficacy. EME demonstrated a non-invasive and biologically interpretable approach to predict EGFR status, stratify survival prognosis, and correlate biological pathways in patients with NSCLC.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In terms of morbidity and mortality, lung cancer is one of the most common cancers worldwide [1]. Non-small cell lung cancer (NSCLC) has been shown to account for approximately 80–85% of all lung cancer cases [2]. Identifying activating mutations in the epidermal growth factor receptor (EGFR) in patients with NSCLC and developing targeted therapies with EGFR tyrosine kinase inhibitors (TKI) have drastically improved the clinical diagnosis and treatment of NSCLC. Currently, the detection of EGFR mutations relies mainly on invasive tumor tissue testing. However, approximately 75% of patients with NSCLC have advanced-stage tumors at diagnosis, making surgical resection or pathological puncture impractical [3]. Moreover, EGFR gene sequencing faces significant limitations, such as high genetic heterogeneity of NSCLC and poor DNA quality [4]. On the other hand, previous studies have reported that the response rate to clinical EGFR-TKI therapies in EGFR-mutant NSCLC patients is approximately 70%, whereas the remaining 30% of patients with positive EGFR mutations do not show the expected survival outcomes [5,6,7]. In addition, clinical studies have indicated that stratifying the prognosis of patients based solely on clinical characteristics such as gender and smoking history was not satisfactory [8, 9]. Therefore, developing a non-invasive method to detect the EGFR-mutant status and predict whether patients could benefit from EGFR-TKI is critical.
Tremendous progress has been made using artificial intelligence, especially deep learning, in the clinical diagnosis and treatment of NSCLC [10,11,12,13]. Previous studies have revealed radiogenomic correlations between gray-level texture features on intratumoral computed tomography (CT) and EGFR mutation status in patients with lung cancer [8, 11, 14, 15]. Meanwhile, it has been proven that the morphometric features of medical images, which are imperceptible to human eyes but detectable by deep learning, are associated with the prognosis of patients with NSCLC [13, 16, 17]. Most previous studies have proposed deep learning models that achieve only a single goal, and rarely achieve both EGFR gene mutation status detection and prognostic evaluation of patients receiving EGFR-TKI therapies [15, 16]. In addition, the accuracy of the models proposed in previous studies is limited, and deep learning models with higher accuracy are needed for clinical applications [8, 15]. Moreover, current deep learning models are limited by the intrinsic “black box” attribute, and studies on the validation of the credibility and biological interpretation of these models are limited [18,19,20].
The EfficientNet network proposed by the Google Brain team in 2019 ensures the model’s accuracy while reducing its parameter scale [21]. The EfficientNetV2 network was proposed in 2021 as an update to the EfficientNet network, which has made remarkable achievements in the diagnosis and prognosis of various human diseases [16, 22,23,24].
In this study, we proposed an EfficientNetV2-L-based deep learning system (EME) to non-invasively predict the EGFR-mutant status in patients with NSCLC and stratify the prognosis of EGFR-mutant patients receiving TKI therapies. Additionally, we investigated the EME as a biologically validated approach to detect EGFR mutation and predict EGFR-TKI survival prognosis.
Methods
Study Design and Participants
This was a retrospective, multicenter study conducted in compliance with the Declaration of Helsinki and approved by the ethics committee of each center. The requirement for informed consent was waived because of the study’s retrospective nature. The overall study design is shown in Fig. 1. The inclusion criteria were as follows: (1) patients diagnosed with NSCLC in four centers from north, eastern, and central China between January 2015 and May 2021; (2) patients who underwent CT examination before treatment; (3) patients who underwent an EGFR-mutant status test; and (4) patients with EGFR positive mutation who were treated with the recommended EGFR-TKI therapies. The exclusion criteria were as follows: (1) patients without baseline CT images; (2) patients without records of the EGFR mutation status; and (3) patients without clinical characteristics. To reveal the correlation between EME and biological pathways of EGFR, we used patients from The Cancer Genome Atlas lung adenocarcinoma (TCGA-LUAD) dataset comprising different races from the USA and Canada (inclusion and exclusion criteria of patients from the TCGA-LUAD dataset are presented in Supplementary A). In this study, patients from the first three hospitals were used as the training dataset, and those from the remaining hospitals were the test dataset. The number of patients in the training and test datasets to develop and validate the EME model follows the 70%:30% principle.
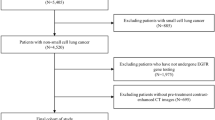
Study flowchart. CT, computed tomography; EGFR, epidermal growth factor receptor; EME, the deep learning model proposed in this study; GSVA, gene set variation analysis; NSCLC, non-small cell lung cancer; TCGA-LUAD, The Cancer Genome Atlas lung adenocarcinoma; TKIs, tyrosine kinase inhibitors; PFS, progression-free survival
According to the EGFR mutation testing (for detailed procedures, see Supplementary B), patients were labeled as EGFR-mutant or EGFR wild-type. Patients could have multiple EGFR mutation types, such as exon 19 deletions and L858R mutations [25]. Patients without any detected EGFR mutations were labeled as EGFR wild-type. For patients with EGFR mutations, follow-up of progression-free survival (PFS), defined as the time from initiation of treatment to the first occurrence of disease progression according to RECIST 1.1 criteria or death from any cause [26], was recorded.
All baseline CT images were reviewed by local radiologists. The ITK-SNAP software [27] (version 3.6.0; http://www.itksnap.org) was used to segment the regions of interest of the primary CT scans by two radiologists with > 5 years of experience. An NSCLC expert with > 10 years of experience, blinded to the above two radiologists, reviewed all the segmentation results. The detailed CT acquisition and reconstruction parameters are described in Supplementary C.
Construction of the EME Model
Previous studies indicated that the peritumoral area on medical images may be highly correlated with disease progression and prognosis in patients with NSCLC [28,29,30]. Therefore, not only the tumor region but also, at most, 10 mm of its peripheral region was included in a patch. The patches were then resized to 256 × 256 pixels with three channels to input into the EME model.
The EME model was based on the cutting-edge EfficientNetV2-L model [31]. The details of the development of the EME model are listed in Supplementary D. The output of the EME is the score predicting the presence or absence of EGFR mutations for each tumor slice. For each patient, the EME score was calculated by averaging the scores for all tumor slices.
Further, to stratify the PFS, we used LASSO Cox regression to build a basic EME-prognostic model based on the deep learning features extracted from the last layer connected to the fully connected layer. A stepwise regression method based on the Akaike information criterion (AIC) was applied to construct the final EME-prognostic model [32]. Once the EME-prognostic model was constructed, a well-validated prognostic tool for signature cut-off selection in the X-tile software (version 3.6.1) [33] was used to select the optimal cut-off score to stratify patients into high- or low-risk subgroups.
Model Comparison and Integration
To further evaluate the performance of the EME model, radiomics features of each patient were extracted using Pyradiomics (version 3.0.1) [34]. A detailed description of the radiomics features is presented in Supplementary E.
Radiomics models were then constructed to predict EGFR-mutant status in patients with NSCLC and to stratify the prognosis after receiving EGFR-TKI. Additionally, fusion models were built by integrating the EME and radiomics models to verify the improvement of EME performance by integrating radiomics.
Feature Visualization
To elucidate the “black box” of the EME model, first, the gradient-weighted class activation maps (Grad-CAM) were used to produce activation maps highlighting the relevant regions activated by the EME model [35]. In addition, we used a nonlinear dimensionality reduction method called t-distributed Stochastic Neighbor Embedding (t-SNE) to reduce the feature dimensions and visualize the relationship between the EME features and EGFR mutation status [36]. Moreover, we used a heat map to describe the relationship between the EME-prognostic features and the prognosis of EGFR-mutant NSCLC patients.
Biological Correlation Analysis
The EME was applied to patients with NSCLC from the TCGA-LUAD dataset. Pathway enrichment of the RNA-seq data of TCGA-LUAD patients was conducted, and the top-ranked key pathways activating the EGFR mutation and regulating the response of EGFR-TKI therapy in patients with NSCLC were identified. Gene set variation analysis (GSVA) was performed to obtain the patient-level GSVA score of each key pathway, and the correlation of the EME signature and the patient-level GSVA scores were evaluated to clarify the relationship of the radiologic features and the underlying biological pathway regulation.
Statistical Analysis
All statistical analyses were conducted using the R software (Version 4.2.0). To compare baseline characteristics, the chi-square test was used for categorical variables, and the Mann–Whitney U test was used for continuous variables. For predicting the EGFR mutation status, we used the area under the curve (AUC), accuracy, precision, recall, and F1 score to evaluate the performance of the models. The Delong test was used to compare the AUCs of different models [37]. The bootstrap method was used to calculate 95% confidence intervals (CI). Multivariate logistic regression analysis was performed using the “glmnet” R package to construct the radiomics model.
For prognosis evaluation, LASSO Cox regression analysis was conducted using the “glmnet” R package, and survival analysis was performed using the “Survival” R package. To reduce the complexity of the prognostic model, we used the AIC stepwise regression method to construct a final effective prognostic model with minimal AIC score. Afterward, Kaplan–Meier analysis and log-rank test were used to visualize the survival curves of the two subgroups using the “survminer” R package. Harrell’s concordance index (C-index) was used to evaluate the performance of the prognostic models. Moreover, we calculated each feature’s contribution for the prognosis task by utilizing the SHapley Additive exPlanation (SHAP) values obtained from the “kernelshap” R package [38, 39].
Additionally, the analysis of t-SNE was conducted using the “Rtsne” R package, and scatter diagrams were drawn using the “ggplot2.” A heat map of the correlation among the prognostic features was drawn using the “ComplexHeatmap” R package. Pathway enrichment was performed using the “clusterProfiler” package by querying the annotated gene set database of Gene Ontology. Biological pathways with a false discovery rate were considered to be statistically significant in the enrichment analysis. Pearson's correlation coefficient was used to determine the relationship between the EME signature and key biological pathways. Statistical significance was set at P < 0.05.
Results
Patient Characteristics
Between January 2015 and May 2021, 542 patients with NSCLC from four independent hospitals were recruited. As shown in Fig. 2, 485 patients with NSCLC from three centers were enrolled and grouped into a training dataset (339 patients), and the patients from the remaining fourth center were used as the test dataset (146 patients) to develop and validate the EME model. Balanced subgroups were found in our training (165 patients with EGFR positive mutation and 174 patients with EGFR wild-type) and test (72 patients with EGFR positive mutation and 74 patients with EGFR wild-type) dataset, and the detailed characteristics are presented in Table 1. The characteristics and flowchart for patients from TCGA-LUAD are presented in Supplementary Table S1 and Fig. S1, respectively.
Prognostic analysis was performed for all EGFR-mutant patients (237 patients). Among them, 165 patients from 2 hospitals were included in the prognostic training dataset, with a median PFS of 269 days (range, 18–1948 days), and 72 patients from another hospital were included in the prognostic test dataset, with a median PFS of 247.5 days (range, 45–1128 days).
Models for EGFR-Mutant Status Detection
The AUC of the EME model was 0.922 (95% CI 0.878–0.939) in the training dataset and 0.907 (95% CI 0.840–0.926) in the test dataset. The accuracy of the EME model was 0.891 (95% CI 0.866–0.915) for the training dataset and 0.829 (95% CI: 0.800–0.864) for the test dataset. There was no statistically significant difference between the training and test datasets (P = 0.062, Delong test). The detailed results are presented in Table 2. Notably, the EME model showed consistent performance across the three centers in the training datasets (confusion matrices in different centers are shown in Supplementary Fig. S2).
For comparison, we constructed a radiomics model containing 35 features to predict the EGFR mutation status (detailed description is provided in Supplementary F). The contribution and importance of the top 15 radiomics features for predicting EGFR mutation status described by SHAP values are shown in Supplementary Fig. S3. As shown in Table 2, the AUC of the radiomics model reached 0.825 (95% CI 0.755–0.896) in the test dataset, which is significantly lower than that of the EME model (P = 0.007, Delong test). Subsequently, a fusion model containing the EME score and radiomics features was constructed (a detailed description is provided in Supplementary G). The AUC of the fusion model reached 0.941 (95% CI 0.905–0.978) in the test dataset, which was significantly higher than that of the radiomics model (P = 0.001, Delong test). However, there was no statistically significant difference between the EME and the fusion models in the test dataset (P = 0.895, Delong test).
Models for EGFR-TKI Prognostic Evaluation
The final EME-prognostic model with 49 features is presented in Supplementary H. The C-index of the EME-prognostic model was 0.778 (95% CI 0.732–0.825) in the prognosis training dataset and 0.711 (95% CI 0.657–0.775) in the prognosis test dataset. The contribution and importance of the top 15 EME features for survival prognosis described by SHAP values are shown in Supplementary Fig. S4.
X-tile indicated that an optimal cut-off of 2.16 was determined to stratify patients into high- and low-risk subgroups. The results showed that patients with an EME-prognostic score lower than 2.16 showed a significantly better PFS than those with an EME-prognostic score higher than 2.16 (training dataset, hazard ratio (HR) 0.099, 95% CI 0.022–0.453, P < 0.001; test dataset, HR 0.341, 95% CI 0.180–0.647, P < 0.001). The Kaplan–Meier curves of EGFR-mutant NSCLC patients in the prognostic training and test datasets are presented in Fig. 3.
Moreover, a radiomics-prognostic model was constructed for comparison. A total of 15 radiomics features were selected; the C-index of the prognostic model reached 0.726 (95% CI 0.678–0.773) in the prognostic training dataset and 0.551 (95% CI 0.478–0.625) in the test dataset. X-tile indicated an optimal cut-off of 1.09 on the prognosis training dataset for stratifying the risk groups (HR 0.285, 95% CI 0.127–0.643, P < 0.001). However, there was no statistically significant difference between the two groups in the test dataset (HR 0.691, 95% CI 0.412–1.158, P = 0.083). The Kaplan–Meier curves based on the three models mentioned above are shown in Fig. 3.
Visualization
To illustrate the feature maps of EME, the GRAD-CAM activation maps of four patients with NSCLC are shown in Fig. 4, revealing that the EME model accurately targeted the locations of the intratumoral and peritumoral areas. Moreover, tumor activation maps revealed distinct imaging patterns between EGFR-mutant and EGFR wild-type patients. We found that heat maps of the EGFR-mutant patients expressed richer heterogeneity in the intratumoral and peritumoral areas, suggesting a higher probability of EGFR mutation expression than patients without EGFR mutations.
The GRAD-CAM activation maps of patients with NSCLC. (a-b) A 61-year-old male with an EME score of 0.996, biopsy-confirmed EGFR wild-type. (c-d) A 55-year-old male with an EME score of 0.996, biopsy-confirmed EGFR wild-type. (e–f) A 67-year-old male with a PFS of 630 days, biopsy-confirmed EGFR exon 19 deletion mutation (with an EME score of 0.030), and the EME-prognostic model predicted as low risk after receiving EGFR-TKI (with an EME-prognostic score of − 3.771). (g-h) A 63-year-old female with a PFS of 132 days, biopsy-confirmed EGFR exon 20 insertion mutation (with an EME score of 0.369), and the EME-prognostic model predicted as high risk after receiving EGFR-TKI (with an EME-prognostic score of 2.281). EGFR, epidermal growth factor receptor; NSCLC, non-small cell lung cancer; EGFR, epidermal growth factor receptor; PFS, progression-free survival; TKI, tyrosine kinase inhibitors
Further, Fig. 5a and b show the relationship between the EME features and EGFR-mutant status based on t-SNE analysis in the training and test datasets, showing that hidden features in the EME were closely related to EGFR-mutant status.
Visualization of the features in the EME system. The t-SNE analysis of the relationship between the EME features and EGFR-mutant status in the a training and b test datasets. c Heat map revealing the relationship between the EME-prognostic features and risk of all EGFR-mutant NSCLC patients receiving EGFR-TKI therapies. EGFR, epidermal growth factor receptor; NSCLC, non-small cell lung cancer; TKI, tyrosine kinase inhibitors; t-SNE, t-distributed Stochastic Neighbor Embedding
Moreover, a heat map (Fig. 5c) containing all EGFR-mutant patients showed the relationship between the EME-prognostic features and risk in patients with NSCLC.
Biological Correlation Analysis
In the TCGA-LUAD cohort, we found significant negative correlations between the EME-prognostic scores and five top-ranked biological pathways (EGFR signaling pathway, regulation of EGFR signaling pathway, cellular response to epidermal growth factor (EGF) stimulus, response to EGF, and protein tyrosine kinase activity, P < 0.05, see Fig. 6 and Supplementary Table S2). As mentioned above, the results indicated that as the EME-prognostic scores increased, patients were more unlikely to benefit from EGFR-TKI. These findings suggested that the prognosis of patients with NSCLC predicted by the EME was positively correlated with the expression of the pathways. Specifically, the results indicated that the survival prognosis predicted by the EME was positively correlated with the expression level of the EGFR signaling pathway and the regulation of the EGFR signaling pathway. As previously reported, patients with EGFR mutations or increased copy number of the EGFR gene are more likely to benefit from EGFR-TKI therapies and potentially show better survival outcomes [40, 41], suggesting that the EME model could potentially reveal the clinical outcome of patients with NSCLC which are regulated by the underlying biological mechanism of EGFR. Moreover, the results showed that the prognosis of patients with NSCLC predicted by the EME was significantly correlated with the pathways of cellular response to EGF stimulus and response to EGF and protein tyrosine kinase activity, which are associated with EGFR binding and kinase activity activation. This finding is consistent with previous reports that EGF binding activates the EGFR tyrosine kinase activity [42, 43].
Discussion
In this study, we constructed and validated a deep learning system called EME to detect EGFR mutation status in patients with NSCLC, predict the prognosis of EGFR-mutant NSCLC patients receiving EGFR-TKI therapies, and reveal the biological correlation of the EME-prognostic score. The EME system showed competitive performance in predicting EGFR-mutant status and evaluating EGFR-TKI therapy prognosis. In addition, this study demonstrated the biological pathway correlation between the EME score and EGFR regulation in patients with NSCLC, indicating that the EME system is an efficient non-invasive approach for advancing individualized clinical decision-making for patients with NSCLC.
Previous studies have indicated that the proportion of patients with NSCLC with EGFR mutations has reached 10% in the USA and 40% in East Asia [44, 45]. Currently, the determination of EGFR mutation status relies on invasive testing of tumor tissue, which entails multiple manual interventions and is accompanied by significant spatial and temporal intratumoral heterogeneity [4]. Therefore, developing non-invasive and assessable radiological assessment for EGFR mutation detection and identifying patients who are likely to benefit from EGFR-TKI are crucial for clinical decision-making.
Compared with results from previous studies, the EME system achieved competitive accuracy in predicting EGFR-mutant status [8, 15]. Wang et al. proposed a deep learning model to predict the EGFR mutation status in lung adenocarcinoma using CT, and their model reached an AUC of 0.810 in the test dataset [15]. Recently, a new CT-based deep learning model was proposed to predict the EGFR mutation status and reached an AUC of 0.813 in the test dataset [8]. According to the results presented here, the EME model achieved an AUC of 0.907 (95% CI 0.840–0.926), which is superior to that of the previous models.
Due to individual differences, it is difficult to accurately identify patients who can benefit from EGFR-TKI in clinical practice. Even patients with detected EGFR mutations may not benefit from EGFR-TKI [5,6,7]. Previous studies have indicated the ability of deep learning models to evaluate the survival prognosis in EGFR-mutant NSCLC patients receiving EGFR-TKI therapies [8, 12, 13]. The EME-prognostic model proposed in this study further demonstrated its value in separating patients receiving EGFR-TKI treatment into high- and low-risk prognostic groups.
Furthermore, the majority of models proposed in previous studies were constrained to accomplish only one objective, namely predicting EGFR mutation status [15] or evaluating the prognosis of patients receiving EGFR-TKI therapies [16]. The EME model proposed in this study can not only detect EGFR mutation status, but it can also predict EGFR-TKI prognosis in patients with NSCLC. This capability facilitates early treatment initiation, thereby enhancing the potential for improved therapeutic outcomes. Moreover, since deep learning is a “black box,” the mechanisms involved are difficult to understand. Whereas previous studies have not further explained the internal mechanism of the reported deep learning models [15, 16], this study interprets it from heat maps and scatter distributions and identifies the relevant biological pathways, which improves the interpretability of the model from a biological point of view.
To further clarify the value of CT images in this context, we constructed radiomics models for comparison and fusion. In predicting EGFR mutation status, the radiomics results (Table 2) demonstrated good performance, which confirmed that medical images contain substantial information for detecting EFGR mutation-derived variations [46, 47]. Therefore, a fusion model containing EME and radiomics models was constructed. As listed in Table 2, the fusion model achieved the best performance. However, no statistically significant difference was observed between the EME and fusion models in the test dataset, indicating that the radiomics features were not significant for the fusion model in terms of accuracy improvement, which is consistent with previous works [48,49,50]. Furthermore, in contrast to radiomics models, the EME model does not necessitate the laborious task of manually outlining ROIs, which allows radiologists to be freed from the burdensome workload associated with segmentation, thereby increasing the acceptability of the EME model.
In terms of prognosis evaluation, radiomics features were used to construct a radiomics-prognostic model; however, the results on survival prognosis of the radiomics and fusion model were inferior (C-index < 0.600 on the test dataset), which is consistent with the findings of previous comparative studies [51, 52]. In contrast, the proposed method, in addition to detecting EGFR mutation status, enables the EME model to identify patients who are more likely to derive benefits from EGFR-TKI. This capability facilitates early treatment initiation, thereby enhancing the potential for improved therapeutic outcomes.
Recently, GRAD-CAM and t-SNE have been widely used to reveal hidden features of deep learning models [36, 53]. In this study, the GRAD-CAM activation maps demonstrated that our model primarily directed attention toward the intratumoral and peritumoral regions. This observation aligns with the findings of previous studies, which have concluded that activation maps can assist physicians in identifying high-risk lung areas in patients, thereby facilitating early intervention and enabling adjustments to treatment plans. In addition, the t-SNE analysis revealed a strong correlation between the EME features and EGFR-mutant status, indicating that the EME features correctly detect the distinct features on CT images between patients with EGFR-mutant and EGFR wild-type NSCLC. Moreover, according to the EME-prognostic model features in EGFR-mutant NSCLC patients, the differences between the patient subgroups with different prognostic risks in Fig. 5 revealed the potential feature divergence of patients with distinct responses to EGFR-TKI therapies, which aids in providing better individualized clinical treatment for patients with NSCLC. Moreover, our study demonstrated that the EME-prognostic score was significantly associated with biological pathways linked to the EGFR mutation and efficacy of EGFR-TKI, which could help clinicians better understand the prognostic information on radiological images that are regulated by these underlying biological pathways.
This study had some limitations. First, we only included Asian patients to develop and validate the EME model, and future research should be based on more ethnicities, in order to improve the universality of the deep learning model. Moreover, EGFR mutation subtypes, such as exon 19 deletions and L858R mutations [25], may be associated with patient prognosis; therefore, studies on different subtypes of EGFR mutations should further investigate this. Second, histopathological images may contain more information and should be considered in further studies to comprehensively analyze the image characteristics of EGFR-mutant NSCLC patients. Next, owing to the correlation between the clinical features and survival prognosis of patients with NSCLC [54], a fusion model containing clinical features should be considered. While we have uncovered the correlation between the EME and biological pathways, it is imperative to conduct in vitro or in vivo experiments to provide further evidence and substantiate these biological associations.
In conclusion, our study proposed a non-invasive and biologically interpretable method to detect the EGFR mutation status in patients with NSCLC and predict the prognosis of patients receiving EGFR-TKI based on pre-therapy CT images. The EME deep learning system can facilitate more individualized clinical decision-making for patients with NSCLC.
Data Availability
The corresponding authors confirms that they had full access to all the study data and had the final responsibility for the decision to submit for publication.
Abbreviations
- AIC:
-
Akaike information criterion
- AUC:
-
Area under the curve
- C-index:
-
Harrell’s concordance index
- CI:
-
Confidence interval
- CT:
-
Computed tomography
- EGF:
-
Epidermal growth factor
- EGFR:
-
Epidermal growth factor receptor
- EME:
-
The deep learning model proposed in this study
- Grad-CAM:
-
Gradient-weighted class activation maps
- GSVA:
-
Gene set variation analysis
- HR:
-
Hazard ratio
- LUAD:
-
Lung adenocarcinoma
- NSCLC:
-
Non-small cell lung cancer
- PFS:
-
Progression-free survival
- SHAP:
-
SHapley Additive exPlanation values
- TCGA:
-
The Cancer Genome Atlas
- TKI:
-
Tyrosine kinase inhibitors
- t-SNE:
-
T-distributed Stochastic Neighbor Embedding
References
Siegel RL, Miller KD, Fuchs HE, Jemal A: Cancer statistics, 2022. CA Cancer J Clin 72(1):7-33, 2022.
Herbst RS, Morgensztern D, Boshoff C: The biology and management of non-small cell lung cancer. Nature 553(7689):446-54, 2018.
Cheng I, Le GM, Noone AM, Gali K, Patel M, Haile RW, Wakelee HA, Gomez SL: Lung cancer incidence trends by histology type among Asian American, Native Hawaiian, and Pacific Islander populations in the United States, 1990-2010. Cancer Epidemiol Biomarkers Prev 23(11):2250-65, 2014.
Bi WL, Hosny A, Schabath MB, Giger ML, Birkbak NJ, Mehrtash A, Allison T, Arnaout O, Abbosh C, Dunn IF, Mak RH, Tamimi RM, Tempany CM, Swanton C, Hoffmann U, Schwartz LH, Gillies RJ, Huang RY, Aerts H: Artificial intelligence in cancer imaging: Clinical challenges and applications. CA Cancer J Clin 69(2):127-57, 2019.
Barnet MB, O'Toole S, Horvath LG, Selinger C, Yu B, Ng CC, Boyer M, Cooper WA, Kao S: EGFR-Co-Mutated Advanced NSCLC and Response to EGFR Tyrosine Kinase Inhibitors. J Thorac Oncol 12(3):585-90, 2017.
Mitsudomi T, Morita S, Yatabe Y, Negoro S, Okamoto I, Tsurutani J, Seto T, Satouchi M, Tada H, Hirashima T, Asami K, Katakami N, Takada M, Yoshioka H, Shibata K, Kudoh S, Shimizu E, Saito H, Toyooka S, Nakagawa K, Fukuoka M: Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): an open label, randomised phase 3 trial. Lancet Oncol 11(2):121-8, 2010.
Rosell R, Carcereny E, Gervais R, Vergnenegre A, Massuti B, Felip E, Palmero R, Garcia-Gomez R, Pallares C, Sanchez JM, Porta R, Cobo M, Garrido P, Longo F, Moran T, Insa A, De Marinis F, Corre R, Bover I, Illiano A, Dansin E, de Castro J, Milella M, Reguart N, Altavilla G, Jimenez U, Provencio M, Moreno MA, Terrasa J, Muñoz-Langa J, Valdivia J, Isla D, Domine M, Molinier O, Mazieres J, Baize N, Garcia-Campelo R, Robinet G, Rodriguez-Abreu D, Lopez-Vivanco G, Gebbia V, Ferrera-Delgado L, Bombaron P, Bernabe R, Bearz A, Artal A, Cortesi E, Rolfo C, Sanchez-Ronco M, Drozdowskyj A, Queralt C, de Aguirre I, Ramirez JL, Sanchez JJ, Molina MA, Taron M, Paz-Ares L: Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 13(3):239-46, 2012.
Wang S, Yu H, Gan Y, Wu Z, Li E, Li X, Cao J, Zhu Y, Wang L, Deng H, Xie M, Wang Y, Ma X, Liu D, Chen B, Tian P, Qiu Z, Xian J, Ren J, Wang K, Wei W, Xie F, Li Z, Wang Q, Xue X, Liu Z, Shi J, Li W, Tian J: Mining whole-lung information by artificial intelligence for predicting EGFR genotype and targeted therapy response in lung cancer: a multicohort study. Lancet Digit Health 4(5):e309-e19, 2022.
Recondo G, Facchinetti F, Olaussen KA, Besse B, Friboulet L: Making the first move in EGFR-driven or ALK-driven NSCLC: first-generation or next-generation TKI? Nat Rev Clin Oncol 15(11):694-708, 2018.
Ozkan E, West A, Dedelow JA, Chu BF, Zhao W, Yildiz VO, Otterson GA, Shilo K, Ghosh S, King M, White RD, Erdal BS: CT Gray-Level Texture Analysis as a Quantitative Imaging Biomarker of Epidermal Growth Factor Receptor Mutation Status in Adenocarcinoma of the Lung. AJR Am J Roentgenol 205(5):1016-25, 2015.
Li XY, Xiong JF, Jia TY, Shen TL, Hou RP, Zhao J, Fu XL: Detection of epithelial growth factor receptor (EGFR) mutations on CT images of patients with lung adenocarcinoma using radiomics and/or multi-level residual convolutionary neural networks. J Thorac Dis 10(12):6624-35, 2018.
Hosny A, Parmar C, Coroller TP, Grossmann P, Zeleznik R, Kumar A, Bussink J, Gillies RJ, Mak RH, Aerts H: Deep learning for lung cancer prognostication: A retrospective multi-cohort radiomics study. PLoS Med 15(11):e1002711, 2018.
Mu W, Jiang L, Zhang J, Shi Y, Gray JE, Tunali I, Gao C, Sun Y, Tian J, Zhao X, Sun X, Gillies RJ, Schabath MB: Non-invasive decision support for NSCLC treatment using PET/CT radiomics. Nat Commun 11(1):5228, 2020.
Wang C, Xu X, Shao J, Zhou K, Zhao K, He Y, Li J, Guo J, Yi Z, Li W: Deep Learning to Predict EGFR Mutation and PD-L1 Expression Status in Non-Small-Cell Lung Cancer on Computed Tomography Images. J Oncol 2021:5499385, 2021.
Wang S, Shi J, Ye Z, Dong D, Yu D, Zhou M, Liu Y, Gevaert O, Wang K, Zhu Y, Zhou H, Liu Z, Tian J: Predicting EGFR mutation status in lung adenocarcinoma on computed tomography image using deep learning. Eur Respir J 53(3):1800986, 2019.
Deng K, Wang L, Liu Y, Li X, Hou Q, Cao M, Ng NN, Wang H, Chen H, Yeom KW, Zhao M, Wu N, Gao P, Shi J, Liu Z, Li W, Tian J, Song J: A deep learning-based system for survival benefit prediction of tyrosine kinase inhibitors and immune checkpoint inhibitors in stage IV non-small cell lung cancer patients: A multicenter, prognostic study. EClinicalMedicine 51:101541, 2022.
She Y, Jin Z, Wu J, Deng J, Zhang L, Su H, Jiang G, Liu H, Xie D, Cao N, Ren Y, Chen C: Development and Validation of a Deep Learning Model for Non-Small Cell Lung Cancer Survival. JAMA Netw Open 3(6):e205842, 2020.
Nensa F, Demircioglu A, Rischpler C: Artificial Intelligence in Nuclear Medicine. J Nucl Med 60(Suppl 2):29s-37s, 2019.
Yang G, Ye Q, Xia J: Unbox the black-box for the medical explainable AI via multi-modal and multi-centre data fusion: A mini-review, two showcases and beyond. Inf Fusion 77:29-52, 2022.
Madani A, Arnaout R, Mofrad M, Arnaout R: Fast and accurate view classification of echocardiograms using deep learning. NPJ Digit Med 1(1):1-6, 2018.
Tan M, Le QV. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. International conference on machine learning, PMLR May 01, 2019:6105–14, 2019.
Nguyen LH, Pham NT, Do VH, Nguyen LT, Nguyen TT, Nguyen H, Nguyen ND, Nguyen TT, Nguyen SD, Bhatti A, Lim CP: Fruit-CoV: An efficient vision-based framework for speedy detection and diagnosis of SARS-CoV-2 infections through recorded cough sounds. Expert Syst Appl 213:119212, 2023.
Wang H, Liu Y, Xu N, Sun Y, Fu S, Wu Y, Liu C, Cui L, Liu Z, Chang Z, Li S, Deng K, Song J: Development and validation of a deep learning model for survival prognosis of transcatheter arterial chemoembolization in patients with intermediate-stage hepatocellular carcinoma. Eur J Radiol 156:110527, 2022.
Hirokawa M, Niioka H, Suzuki A, Abe M, Arai Y, Nagahara H, Miyauchi A, Akamizu T: Application of deep learning as an ancillary diagnostic tool for thyroid FNA cytology. Cancer Cytopathol 131(4):217-25, 2022.
Robichaux JP, Le X, Vijayan RSK, Hicks JK, Heeke S, Elamin YY, Lin HY, Udagawa H, Skoulidis F, Tran H, Varghese S, He J, Zhang F, Nilsson MB, Hu L, Poteete A, Rinsurongkawong W, Zhang X, Ren C, Liu X, Hong L, Zhang J, Diao L, Madison R, Schrock AB, Saam J, Raymond V, Fang B, Wang J, Ha MJ, Cross JB, Gray JE, Heymach JV: Structure-based classification predicts drug response in EGFR-mutant NSCLC. Nature 597(7878):732-7, 2021.
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, Dancey J, Arbuck S, Gwyther S, Mooney M, Rubinstein L, Shankar L, Dodd L, Kaplan R, Lacombe D, Verweij J: New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45(2):228–47, 2009.
Yushkevich PA, Piven J, Hazlett HC, Smith RG, Ho S, Gee JC, Gerig G: User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31(3):1116-28, 2006.
Dou TH, Coroller TP, van Griethuysen JJM, Mak RH, Aerts H: Peritumoral radiomics features predict distant metastasis in locally advanced NSCLC. PLoS One 13(11):e0206108, 2018.
Chen Q, Shao J, Xue T, Peng H, Li M, Duan S, Feng F: Intratumoral and peritumoral radiomics nomograms for the preoperative prediction of lymphovascular invasion and overall survival in non-small cell lung cancer. Eur Radiol 33(2):947-958, 2023
Wu L, Lou X, Kong N, Xu M, Gao C: Can quantitative peritumoral CT radiomics features predict the prognosis of patients with non-small cell lung cancer? A systematic review. Eur Radiol 33(3):2105-17, 2022.
Tan M, Le Q, editors. Efficientnetv2: Smaller models and faster training. International conference on machine learning PMLR, 2021:10096-10106, 2021
Akaike H, editor. Information theory as an extension of the maximum likelihood principle. In Proc. 2nd int. symp. on information theory: pp 267-281, 1973.
Camp RL, Dolled-Filhart M, Rimm DL: X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res 10(21):7252-9, 2004.
van Griethuysen JJM, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RGH, Fillion-Robin JC, Pieper S, Aerts H: Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Res 77(21):e104-e7, 2017.
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D: Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. International Journal of Computer Vision 128(2):336-59, 2020.
Foersch S, Eckstein M, Wagner DC, Gach F, Woerl AC, Geiger J, Glasner C, Schelbert S, Schulz S, Porubsky S, Kreft A, Hartmann A, Agaimy A, Roth W: Deep learning for diagnosis and survival prediction in soft tissue sarcoma. Ann Oncol 32(9):1178-87, 2021.
DeLong ER, DeLong DM, Clarke-Pearson DL: Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44(3):837-45, 1988.
Tsai SF, Yang CT, Liu WJ, Lee CL: Development and validation of an insulin resistance model for a population without diabetes mellitus and its clinical implication: a prospective cohort study. EClinicalMedicine 58:101934, 2023.
Kim HY, Lampertico P, Nam JY, Lee HC, Kim SU, Sinn DH, Seo YS, Lee HA, Park SY, Lim YS, Jang ES, Yoon EL, Kim HS, Kim SE, Ahn SB, Shim JJ, Jeong SW, Jung YJ, Sohn JH, Cho YK, Jun DW, Dalekos GN, Idilman R, Sypsa V, Berg T, Buti M, Calleja JL, Goulis J, Manolakopoulos S, Janssen HLA, Jang MJ, Lee YB, Kim YJ, Yoon JH, Papatheodoridis GV, Lee JH: An artificial intelligence model to predict hepatocellular carcinoma risk in Korean and Caucasian patients with chronic hepatitis B. J Hepatol 76(2):311-8, 2022.
Cappuzzo F, Hirsch FR, Rossi E, Bartolini S, Ceresoli GL, Bemis L, Haney J, Witta S, Danenberg K, Domenichini I, Ludovini V, Magrini E, Gregorc V, Doglioni C, Sidoni A, Tonato M, Franklin WA, Crino L, Bunn PA, Jr., Varella-Garcia M: Epidermal growth factor receptor gene and protein and gefitinib sensitivity in non-small-cell lung cancer. J Natl Cancer Inst 97(9):643-55, 2005.
Tsao MS, Sakurada A, Cutz JC, Zhu CQ, Kamel-Reid S, Squire J, Lorimer I, Zhang T, Liu N, Daneshmand M, Marrano P, da Cunha Santos G, Lagarde A, Richardson F, Seymour L, Whitehead M, Ding K, Pater J, Shepherd FA: Erlotinib in lung cancer - molecular and clinical predictors of outcome. N Engl J Med 353(2):133-44, 2005.
Ferguson KM: Structure-based view of epidermal growth factor receptor regulation. Annu Rev Biophys 37:353-73, 2008.
Voldborg BR, Damstrup L, Spang-Thomsen M, Poulsen HS: Epidermal growth factor receptor (EGFR) and EGFR mutations, function and possible role in clinical trials. Ann Oncol 8(12):1197-206, 1997.
Paez JG, Jänne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, Eck MJ, Sellers WR, Johnson BE, Meyerson M: EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304(5676):1497-500, 2004.
Pao W, Miller V, Zakowski M, Doherty J, Politi K, Sarkaria I, Singh B, Heelan R, Rusch V, Fulton L, Mardis E, Kupfer D, Wilson R, Kris M, Varmus H: EGF receptor gene mutations are common in lung cancers from "never smokers" and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci U S A 101(36):13306-11, 2004.
Yang L, Xu P, Li M, Wang M, Peng M, Zhang Y, Wu T, Chu W, Wang K, Meng H, Zhang L: PET/CT Radiomic Features: A Potential Biomarker for EGFR Mutation Status and Survival Outcome Prediction in NSCLC Patients Treated With TKIs. Front Oncol 12:894323, 2022.
Feng Y, Song F, Zhang P, Fan G, Zhang T, Zhao X, Ma C, Sun Y, Song X, Pu H, Liu F, Zhang G: Prediction of EGFR Mutation Status in Non-Small Cell Lung Cancer Based on Ensemble Learning. Front Pharmacol 13:897597, 2022.
Astaraki M, Yang G, Zakko Y, Toma-Dasu I, Smedby Ö, Wang C: A Comparative Study of Radiomics and Deep-Learning Based Methods for Pulmonary Nodule Malignancy Prediction in Low Dose CT Images. Front Oncol 11:737368, 2021.
Huang W, Wang J, Wang H, Zhang Y, Zhao F, Li K, Su L, Kang F, Cao X: PET/CT Based EGFR Mutation Status Classification of NSCLC Using Deep Learning Features and Radiomics Features. Front Pharmacol 13:898529, 2022.
Whitney HM, Li H, Ji Y, Liu P, Giger ML: Comparison of Breast MRI Tumor Classification Using Human-Engineered Radiomics, Transfer Learning From Deep Convolutional Neural Networks, and Fusion Methods. Proc IEEE Inst Electr Electron Eng 108(1):163-77, 2020.
Song J, Wang L, Ng NN, Zhao M, Shi J, Wu N, Li W, Liu Z, Yeom KW, Tian J: Development and Validation of a Machine Learning Model to Explore Tyrosine Kinase Inhibitor Response in Patients With Stage IV EGFR Variant-Positive Non-Small Cell Lung Cancer. JAMA Netw Open 3(12):e2030442, 2020.
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, van der Laak J, van Ginneken B, Sánchez CI: A survey on deep learning in medical image analysis. Med Image Anal 42:60-88, 2017.
Panwar H, Gupta PK, Siddiqui MK, Morales-Menendez R, Bhardwaj P, Singh V: A deep learning and grad-CAM based color visualization approach for fast detection of COVID-19 cases using chest X-ray and CT-Scan images. Chaos Solitons Fractals 140:110190, 2020.
Xu Y, Hosny A, Zeleznik R, Parmar C, Coroller T, Franco I, Mak RH, Aerts H: Deep Learning Predicts Lung Cancer Treatment Response from Serial Medical Imaging. Clin Cancer Res 25(11):3266-75, 2019.
Funding
This study was funded by the National Natural Science Foundation of China (grant number 92259104).
Author information
Authors and Affiliations
Contributions
Conception and design of the study were done by JS, KD, SL, and NX; development and validation of the model were done by NX, JW, and TL; analysis and interpretation of data were done by NX, JW, and GD; initial draft of the manuscript was done by NX, JW, and GD; all authors contributed to the design, data analysis, and result interpretations. The co-corresponding authors had full access to the data in the study, and took responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of the First Hospital of China Medical University (No. AF-SOP-07–1).
Consent to Participate
Written informed consent was waived by the institutional review board.
Consent for Publication
All authors have consented to the submission of the manuscript and the authors have obtained consent for publication for the content covered in the manuscript.
Conflict of Interest
The authors declare no competing interests.
Disclaimer
The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Critical Relevance Statement
The model proposed in this study may facilitate individualized clinical decision-making for patients with NSCLC, especially for the detection of EGFR-mutant status, the prognosis of EGFR-mutant NSCLC patients receiving EGFR-TKI, and biological interpretation of CT semantic features in patients with advanced NSCLC.
Key Points
• It is critical to propose a biologically interpretable approach for the clinical diagnosis and treatment of patients with non-small cell lung cancer (NSCLC).
• The proposed model can detect epidermal growth factor receptor (EGFR) mutant status in patients with NSCLC and stratify the prognosis of patients receiving EGFR tyrosine kinase inhibitors (TKI) therapies.
• The proposed model strongly correlated with the biological pathways regulating the EGFR mutation and EGFR-TKI efficacy of NSCLC.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, N., Wang, J., Dai, G. et al. EfficientNet-Based System for Detecting EGFR-Mutant Status and Predicting Prognosis of Tyrosine Kinase Inhibitors in Patients with NSCLC. J Digit Imaging. Inform. med. 37, 1086–1099 (2024). https://doi.org/10.1007/s10278-024-01022-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-024-01022-z