Abstract
Since the 1990s, the cryptic whitefly (Bemisia tabaci) has been linked to severe viral disease pandemics affecting cassava, a crucial staple crop in eastern Africa. This surge in whitefly populations has also been observed in other crops and uncultivated plants. While previous surveys have connected the increase on cassava to two specific populations, SSA1 and SSA2, the dynamics behind the population growth on other plants remain unclear. Additionally, other B. tabaci species, including EA1, IO, MED, SSA9, and SSA10, have been found on cassava in smaller numbers. This study aimed to identify the host plants that support the growth and development of different B. tabaci in Uganda by collecting fourth-instar nymphs from cassava and 20 other common host plants. Host transfer experiments were conducted to test the ability of seven species (EA1, MEAM1, MED-Africa Silver Leafing (ASL), SSA1-subgroup1, SSA1-Hoslundia, SSA6, and SSA12) to develop on cassava. The identities of the nymphs were determined using partial mitochondrial cytochrome oxidase 1 sequences. Twelve B. tabaci species were identified, including two novel species, based on the 3.5% nucleotide sequence divergence. Cassava was colonised by SSA1-SG1, SSA1-SG2, and SSA2. The most prevalent species were SSA1-SG1, MED-ASL, and SSA13, which were also the most polyphagous, colonising multiple plant species. Several whitefly species colonised specific weeds, such as Aspilia africana and Commelina benghalensis. The polyphagous nature of these species supports continuous habitats and virus reservoirs. Effective management of whitefly populations in eastern Africa requires an integrated approach that considers their polyphagy and the environmental factors sustaining host plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bemisia tabaci (Gennadius) is a group of cryptic species, whose members differ in their levels of polyphagy (Dinsdale et al. 2010; Mugerwa et al. 2018; Vyskočilová et al. 2019). B. tabaci Middle East-Asia Minor1 (MEAM1) and Mediterranean (MED), for example, are highly polyphagous (Brown et al. 1995; Shah and Liu 2013; Xia et al. 2017), whereas some species have a limited host range. Examples of monophagous species include the Jatropha race, which colonises Jatropha gossypifolia L. in Puerto Rico (Bird 1957) and sub-Saharan Africa (SSA) 6 that colonises Ocimum gratissimum L. in Uganda (Sseruwagi et al. 2005). Polyphagous insect pests utilise various host plants from different plant families, which allows them to adapt to changes in agroecosystems such as senescence of preferred host plants, harvest, and changing cropping patterns (Kennedy and Storer 2000; Sivakoff et al. 2013). Therefore, knowledge of the host plant range and level of polyphagy of B. tabaci species is important for the development of effective management strategies of these pests. In addition, the choice of host plants determines the pathways of spread of whitefly-transmitted viruses (Polston and Capobianco 2013) and a wider vector host plant range may increase transmission of viruses to crops.
African cassava B. tabaci, namely SSA1 and SSA2, have been reported to be the most frequent species colonising cassava in Uganda (Legg et al. 2002, 2014; Sseruwagi et al. 2006; Mugerwa et al. 2018). Other B. tabaci species such as SSA9, SSA10, MED, Indian Ocean (IO) and East Africa (EA) 1 have also been collected from cassava, although in relatively low numbers (Legg et al. 2014; Mugerwa et al. 2018). Studies showed that the globally invasive whitefly species MEAM1 and MED (De Barro et al. 2011; Wang et al. 2011) occurred mainly on non-cassava host plants in Uganda, with MED being more prevalent than MEAM1 (Sseruwagi et al. 2005; Mugerwa et al. 2018; Namuddu et al. 2023). Adult whitefly surveys showed that the SSA1, particularly the subgroup1 (SSA1-SG1), was the most abundant cassava-feeding B. tabaci and it occurred on more than 30 plant species (Legg et al. 2014; Mugerwa et al. 2018; Mugerwa et al. 2021). However, adults are of limited reliability when studying the actual host plant range of B. tabaci, because their occurrence on a plant may not necessarily signify that it is good for nymphal growth and development (Sseruwagi et al. 2006). The adults, for example, could have been casually on the plant, searching for a suitable host for feeding and oviposition (van Lenteren and Noldus 1990; Walling 2008) at the time of collection.
When fourth-instar nymphs were collected from field host plants, Sseruwagi et al. (2006) revealed that the SSA1 could develop on five additional host plant species commonly found adjacent to cassava fields. These were: (i) Manihot glaziovii (Müll.Arg.) Allem, (ii) J. gossypifolia, (iii) Euphorbia heterophylla L., (iv) Aspilia africana (Pers.) C. D. Adams, and (v) Abelmoschus esculentus L.. In addition to collecting fourth-instar nymphs from alternative host plants in close proximity to cassava, it is also important to collect the fourth-instar nymphs from the non-cultivated host plants, in the surrounding vegetation near cassava. This would help to establish if they are the source of whiteflies that infest cassava when it is planted in a new field, facilitating the continuous high whitefly populations on cassava. It was hypothesised that highly fecund B. tabaci species could displace other whitefly species from cassava. In this study, host transfer experiments for some of the B. tabaci species that have been collected from cassava in low numbers were conducted to determine their ability to colonise cassava. The aim of this work, therefore, was to carry out a comprehensive investigation into the cassava-colonising B. tabaci species in Uganda and the alternative host plant species suitable for their feeding and development.
Materials and methods
Collection of fourth-instar nymphs within and adjacent to cassava fields
Fourth-instar B. tabaci nymphs were collected from cassava and eight cultivated plant species that are commonly intercropped or grown in close proximity to cassava and 13 non-cultivated plant species usually found within or adjacent to cassava fields (Table 1). Plants adjacent to cassava fields were defined as those positioned in a margin of 0–25 m away from the edges of the nearest cassava field. At least 10 nymphs were collected from two to three leaves of a given plant species and preserved in 90% ethanol in 1.5-mL Eppendorf tubes. The unparasitised fourth-instar nymphs were collected with the aid of a Peak 1966 Light Loupe 10 × hand lens from plants without virus symptoms.
Collection of fourth-instar nymphs from plants situated at least 300 m from cassava
Samples of fourth-instar B. tabaci nymphs on A. africana, Phyllanthus niruri L., Hoslundia opposita Vahl, O. gratissimum, Vernonia amygdalina Delile, Lantana camara L., E. heterophylla and Sida acuta Burm. f. in the vegetation situated > 300 m from cassava fields were collected for analysis. It proved difficult, however, to find a wide range of cultivated host plants with fourth-instar nymphs. Therefore, a field experiment was set up where the position of experimental plants could be manipulated. The non-cultivated plants species Bidens pilosa L., Ageratum conyzoides (L.) L., Commelina benghalensis L., M. glaziovii, Pavonia urens Cav. and cultivated ones, V. unguiculata, A. hypogaea, P. vulgaris, Gossypium hirsutum L., Solanum lycopersicum L., A. esculentus and Ipomoea batatas (L.) Lam. were set up in an experimental plot in an open field more than 300 m from cassava at the National Crops Resources Research Institute (NaCRRI), Namulonge, Uganda (N0.521257, E32.631322). Plants were grown in 3-L buckets in an insect-proof screen house until they had three to five fully expanded leaves. They were then transferred to the open field surrounded by a forest, shrubs and land that had been left to fallow. Prior to transferring them to the field, the buckets and plants were fully covered with plastic to avoid any infestation from cassava and/or sweet potato whiteflies during transportation. The area where the buckets were placed had the grass cut back every two weeks. Each plant species had six replicates, and each plant was assigned a random number. The plants were spaced at 1-m intervals in three rows in a 2 m × 2 m grid and allocated randomly to a position. They were ordered 1 to 72 with three rows and each row contained 24 plants. The plants were watered every five days during the dry season. After three weeks and every week thereafter for two months, the plants’ leaves were examined for fourth-instar nymphs. The fourth-instar B. tabaci nymphs were collected and preserved in 90% ethanol.
Whiteflies did not colonise A. esculentus (okra), V. unguiculata (cowpea) and M. glaziovii (tree cassava), in the experimental study area at Namulonge, but whitefly nymphs have been seen developing on these species elsewhere. We widened the search, therefore, to Lira for okra (N2.418294, E32.150063), Kabanyolo for cowpea (N0.466478, E32.610252) and Mukono for tree cassava (N0.435072, E32.768698) (Table 1), where these plant species could be found separated from cassava by distances greater than 300 m. Leaf samples for each host plant species were kept in separate plastic ‘Ziplock’ bags. Geo-coordinates (latitude and longitude) were recorded for each collection site using a geographical positioning system (GPS, Garmin eTrex Vista Cx) and used to generate a map (Fig. 1). All samples were collected during the second rainy season of 2017 (August to December).
Growing plants
Plant species and whitefly colonies (Table 1) were raised in an insect-proof screen house at NaCRRI, Namulonge at 27 ± 5 °C, 70 ± 5% relative humidity (RH) and 12:12 light: dark photoperiod (L:D). Plants were raised on sterilised forest soil mixed with chicken manure at a ratio of 10:3. The soil mixture was sterilised by steaming overnight and allowed to cool before use. At three fully expanded leaf growth stage, plants were individually placed in Lock and Lock (LL) whitefly-proof cages (Wang et al. 2011). The LLs were modified with two additional side openings in the upper container covered by 160-µm nylon mesh for good ventilation. At the Natural Resources Institute, UK, two substrates: Multipurpose Jiffy substrates (Jiffy products, UK) and John Innes No. 2 compost (John Innes manufacturing association, UK), were sterilised by freezing for three days and mixed in a 1:1 ratio. The sterilised mixture was used to grow the different host plants that were propagated by seed (Table 1) and eggplant cv. Black Beauty. They were planted in potting cups (7 × 7 × 8 cm) and thereafter transferred to the growth room. The plants were raised and maintained in the growth room under a controlled environment of 25 ± 2 °C, 65 ± 5% RH, and 14:10 h L: D. Cassava, sweet potato, M. glaziovii and C. benghalensis cuttings were treated with a systemic insecticide and fungicide and planted in potting cups (7 × 7 × 8 cm) in the glasshouse. Plants were kept under quarantine in the glasshouse for two months before use in experiments. Experimental plants with three fully expanded leaves were also enclosed in LL cages.
Rearing whiteflies
From each of the selected plant species, leaf samples with fourth-instar nymphs were collected from plants at least 300 m from cassava and kept in separate plastic ‘Ziplock’ bags. The samples were placed in petri dishes for adults to emerge. The emerged whiteflies were used to establish a live pure colony (one male and one female per plant), on the same host plant species from which they collected, individually enclosed in LL cages. The pure colonies were kept in LL cages for 2–4 months and then transferred to BugDorm whitefly-proof insect-rearing cages (60 cm × 60 cm × 60 cm) containing two or three plants.
The raised whiteflies were transferred to NRI, UK, for molecular analysis and host plant transfer experiments. Briefly, a day before the transfer, five leaves with fourth-instar nymphs were collected from each B. tabaci colony at NaCRRI, Namulonge and these were placed in petri dishes (90 × 15 mm). The petri dishes were sealed with parafilm, wrapped in cloth, placed in perforated polythene bags, and carefully packed in a box for shipping.
Upon arriving at NRI, emerged adults were released onto different egg plants (cv. Black Beauty) in LL cages and monitored in the controlled environment room at 26 ± 3 °C, 65 ± 5% RH and 12:12 h L:D. The whitefly colonies were kept in LL cages until their identity was determined. After 2–4 months, the colonies had built up to about 200 whiteflies per LL cage. The colonies were then transferred to BugDorm whitefly-proof insect-rearing cages containing two or three plants (with at least five fully expanded leaves) of the same plant species, from which each colony had been collected.
Suitability of cassava for feeding and oviposition by B. tabaci species
The suitability of cassava for feeding and development was measured in terms of survival and oviposition by the B. tabaci species that were successfully established at NRI (Table 1). The SSA1-Hoslundia (Namuddu et al. 2023) was also included in the study to investigate whether it was similar to SSA1-SG1 and could develop on cassava. The SSA1-SG1 cassava colony that was collected from Kayingo in Uganda in 2016 and established at NRI, UK (Mugerwa et al. 2018), was used as the control. Cassava plants cv. (NAROCASS1, Magana and Columbian) with three fully expanded leaves were placed in LL cages. NAROCASS1 is an improved variety that is resistant to cassava mosaic disease (CMD) and tolerant to cassava brown streak disease (CBSD), but is susceptible to whiteflies. Magana is a Ugandan landrace that is susceptible to CMD and CBSD, but relatively resistant to whiteflies (Omongo et al. 2012), while Columbian variety is the standard used for rearing cassava whiteflies at NRI, UK. The experimental design involved four replicates and a random number was assigned to each plant. Twenty-five pairs of five-day-old, non-sexed adults were collected from colonies that had been maintained on their respective host plants for at least four generations. The adults were taken from the apical leaves with the aid of an aspirator and introduced onto cassava plants. Surviving adults were counted daily until the eighteenth day after release, when early fourth-instar larvae were observed for SSA1-SG1. All instar stages and any eggs (dead or alive) were counted on each plant, with the aid of a hand lens (× 40), to determine the total B. tabaci eggs oviposited during this period. After counting, plants were kept for a further month to observe any adult emergence of B. tabaci EA1, MEAM1, MED-ASL, SSA1-Hoslundia, SSA6 and SSA12 on cassava. The experiments were carried out under controlled conditions (26 ± 3 °C, 65 ± 5% RH and 12:12 h L:D).
Extraction of DNA from an individual whitefly nymph
Genomic DNA was extracted from two individual fourth-instar nymphs randomly picked from each sample (Eppendorf tube), using a Chelex method (White et al. 2009). Each whitefly in 75 µL of 10% Chelex® (Sigma-Aldrich, St Louis, USA) containing 6 µL of 10 mg/mL proteinase K was crushed in a 1.5-mL Eppendorf tube with zirconium oxide beads using a bullet blender homogeniser (Next Advance, Averill Park, NY). The Eppendorf tubes with the extract were centrifuged for 1 min at 15,871 g. The extract was then incubated for one hour at 37 °C and then for 10 min at 96 °C to denature the enzyme. The DNA extracts were stored at − 20 °C until use.
Polymerase chain reaction (PCR) of mtCO1 DNA, purification and sequencing
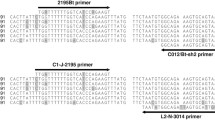
The 3’ region of the mtCO1 gene was amplified using a forward primer, 2195Bt (TGRTTTTTTGGTCATCCRGAAGT), and reverse primer C012/Bt-sh2 (TTTACTGCACTTTCTGCC) with an expected amplicon size of 867 bp (Mugerwa et al. 2018). The total reaction mixture of 30 µL contained 2 µL of the DNA extract, 0.2 µM of each primer pair, 6 µL of 5X MyTaq Buffer HS (Bioline), 1.0 U of MyTaq Polymerase (Bioline) and molecular grade water (Sigma-Aldrich, UK). PCR cycling was performed in a Veriti 96-well thermal cycler (Applied Biosystems, UK) programmed as follows: Initial denaturation at 95 °C/3 min, 10 cycles of 95 °C/30 s, annealing at 45 °C/30 s and 72 °C/1 min; 20 cycles of 95 °C/30 s, annealing at 50 °C/30 s and 72 °C/1 min and final cycle at 72 °C/10 min. Electrophoresis of PCR products was done on 1% (w/v) agarose gels in 0.5 X Tris/borate/EDTA buffer, stained with RedSafe™ (iNtRON Biotechnology, Korea). PCR products on the agarose gel were visualised under UV light at 302 nm on a trans-illuminator GBOX, CHEMI 16 (Syngene, UK), and those with the expected size were purified using the exonuclease 1 and antarctic phosphatase (Exo1-AP) treatment (Werle et al. 1994) with slight modifications. The PCR products (20 µL) were mixed with 1 U Exo1 and 1 U AP (New England Biolabs, UK) and incubated for 20 min at 37 °C. The enzymes were inactivated for 10 min at 80 °C. Purified PCR products were sent for Sanger sequencing in both directions at GATC Biotech (Germany).
Phylogenetic analysis of partial mtCO1 sequences
The partial mtCO1 Sanger sequences generated in this study were first screened using quality checks in Geneious version 10.2.3. Firstly, a consensus sequence was generated using the reverse and forward sequences for each individual whitefly (Kearse et al. 2012). The new consensus sequences of this study were aligned with high-throughput sequencing (HTS)-derived full mitogenome sequences downloaded directly from GenBank, using MAFFT (Katoh and Standley 2013). All Sanger sequences which contained indels were eliminated and not considered for further analysis. The remaining Sanger sequences together with HTS-sequences were then trimmed to 657 bp and translated to amino acid residues from appropriate codon positions using the invertebrate mitochondrial DNA genetic codes to (i) identify potential premature stop codons and (ii) enable amino acid residue alignment against the HTS reference CO1 amino acid data set. Sanger sequences that had premature stop codons and amino acid substitutions in highly conserved regions as identified within the trimmed HTS reference CO1 gene set were eliminated. Then 29 unique haplotypes were extracted from the remaining sequences (n = 179) and aligned with equivalent reference whitefly sequences downloaded directly from the GenBank in accordance with Kunz et al. (2019). MEGA-X was used to determine the best substitution model. Phylogenetic analysis was done using BEAST2, which employs Markov Chain Monte Carlo (MCMC) sampling to approximate posterior probabilities of phylogenies (Green 1995). It also utilises the BEAGLE library, an application programming interface, and library of high-performance statistical phylogenetic inference (Ayres et al. 2012). BEAUti version 2.5.1 was used to generate the xml file for analysis in BEAST version 2.5.1. Tree prior was set to 'Yule' with strict clock model and uniform birth rate. The site model was specified as TN93, base frequencies were estimated, and Gamma plus invariant sites was the site heterogeneity model. The MCMC was run for 10 million generations and sampled every 1000th generation (the first 10% of trees were discarded as burn-in). TreeAnnotator version 2.5.1 was used to generate a final tree and was visualised by FigTree version 1.4.3 to determine the similarities between reference sequences and haplotype sequences. Bemisia afer (accession no. KF734668) was used as the outgroup for the B. tabaci complex.
Results
Host plants utilised by B. tabaci nymphs in Uganda
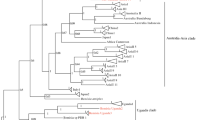
Fourth-instar nymphs collected from cassava and common host plants in Uganda were identified based on phylogenetic analysis of partial mtCO1 sequences (657 bp) (Dinsdale et al. 2010) and are shown in Table 2. The whiteflies clustered with already published B. tabaci species and other whitefly species and they grouped into 14 genetically distinct groups (Fig. 2) that included: SSA1(SSA1-SG1, SSA1-SG2 and SSA1-Hoslundia), SSA2, SSA6, SSA10, SSA12, SSA13, EA1, IO, MEAM1, MED-ASL, B. Uganda1, B. afer and two new whitefly species. The results showed that B. tabaci species, SSA1 (SSA1-SG1 and SSA1-SG2) and SSA2 colonised cassava. However, the SSA1-Hoslundia, a subgroup of SSA1 did not colonise cassava, but utilised six other host plants, H. opposita, A. africana, M. glaziovii, A. conyzoides, S. acuta and I. batatas (Table 3). SSA1-SG1 utilised 11 alternative hosts whereas, SSA1-SG2 colonised alternative hosts, E. heterophylla and M. glaziovii (Table 3). SSA2 was not found on any of the alternative host plants sampled.
BEAST2 phylogenetic tree based on partial mtCOI sequences (657 bp) of individual fourth-instar whitefly nymphs collected from cassava and other host plant species within and adjacent to cassava, and plants at least 300 m away from cassava. Numbers noted at the nodes are posterior probabilities. Numbers in red besides a sample sequence are the number of individuals with that particular haplotype and with an asterisk (*) are reference sequences
The whitefly species SSA6, SSA10, SSA12, SSA13, EA1, IO, MEAM1, MED-ASL, B. Uganda1, B. afer and two unknowns colonised non-cassava host plants. Most of them colonised more than two plant species except SSA6, SSA10, SSA12 and B. afer, which developed only on O. gratissimum, V. amygdalina, C. benghalensis and M. glaziovii, respectively (Table 3). One of the unknowns (Unknown1) that shared a maximum partial mtCO1 nucleotide identity of 77.6% to a known whitefly species, B. afer (accession no. AF418673) colonised A. hypogaea (Tables 2 and 3), while the second one (Unknown2) that shared only a maximum nucleotide identity of 87.6% to a published putative species, SSA2 (accession no. KF425619) utilised A. africana (Tables 2 and 3).
Host preferences were observed for the different cryptic species. SSA1-SG1 was closely associated Euphorbiaceae, Phyllanthacea and Asteraceae plant families, whereas MED-ASL mainly colonised the Malvaceae and Fabaceae families (Table 3). SSA1-SG2 was more common on Euphorbiaceae while SSA13 was more common on the Verbenaceae. MEAM1 was more prevalent on the Solanaceae. IO and SSA12 were more frequent on the Commelinaceae. Other cryptic species colonised particular plant species within a given plant family (Table 3).
Distribution of whitefly species on different plants in the field
Whitefly species within cassava fields
Of the cassava B. tabaci identified in the study, SSA1-SG1 was the most predominant on cassava. It was more prevalent on cassava than on intercropped plant species and non-cultivated plants within cassava fields (Fig. 3). SSA1-SG1 and SSA1-SG2 equally colonised E. heterophylla. SSA1-SG1 colonised most of the host plants found within cassava fields except P. vulgaris, S. acuta, C. benghalensis, V. unguiculata and A. hypogaea (Fig. 3). MED-ASL and SSA12 colonised only V. unguiculata and C. benghalensis, respectively. At least two whitefly species colonised the same plant species within cassava fields, except MED-ASL and Unknown1, which utilised V. unguiculata and A. hypogaea, respectively (Fig. 3).
Distribution of whitefly species that colonised cultivated and non-cultivated host plants within cassava fields. P. vulgaris, I. batatas, V. unguiculata and A. hypogea were intercrops. SSA: sub-Saharan Africa, SG: subgroup, EA1: East Africa1, IO: Indian Ocean, MEAM1: Middle East-Asia Minor1 and MED-ASL: Mediterranean African silver leafing
Whitefly species that colonised host plants adjacent to cassava fields
SSA1-SG1 was the only cassava B. tabaci that colonised alternative host plants adjacent to cassava. It utilised three out of the 14 host plants sampled, all of which were non-cultivated (Fig. 4). MED-ASL was the most polyphagous species and it was found on six host plants, three of which were cultivated. B. Uganda1 colonised four host plants. SSA13, EA1, IO, MEAM1 each colonised two host plants and MEAM1 utilised only cultivated host plants (Fig. 4). SSA1-Hoslundia, SSA6 and B. afer colonised only one host plant (Fig. 4).
Distribution of whitefly species that colonised cultivated and non-cultivated host plants adjacent to cassava fields. P. vulgaris, I. batatas, S. lycopersicum, A. esculentus and V. unguiculata were cultivated plant species. SSA: sub-Saharan Africa, SG: subgroup, EA1: East Africa1, IO: Indian Ocean, MEAM1: Middle East-Asia Minor1 and MED-ASL: Mediterranean African silver leafing
Whitefly species that colonised host plants more than 300 m from cassava fields
SSA1-SG1, SSA1-Hoslundia, and SSA1-SG2 colonised six, four and one host plant species, respectively, more than 300 m from cassava. All host plants colonised by the three whitefly species were non-cultivated (Fig. 5). EA1, MED-ASL and B. Uganda1 utilised both cultivated and non-cultivated plants, whereas MEAM1 colonised only cultivated host plants (Fig. 5). SSA6, SSA10, SSA12, SSA13 and IO developed on non-cultivated plant species (Fig. 5).
Distribution of whitefly species that colonised cultivated and non-cultivated host plants more than 300 m from cassava fields. Nymphs on A. africana, P. niruri, H. opposita, O. gratissimum, V. amygdalina, L. camara, E. heterophylla and S. acuta were collected from the bush; A. esculentus, M. glaziovii and V. unguiculata were collected from farmers’ fields in Lira, Mukono and Kabanyolo, respectively; and the rest were from the experimental plot at NaCRRI. P. vulgaris, G. hirsutum, I. batatas, S. lycopersicum, A. esculentus and V. unguiculata were cultivated plant species. SSA: sub-Saharan Africa, SG: subgroup, EA1: East Africa1, IO: Indian Ocean, MEAM1: Middle East-Asia Minor1 and MED-ASL: Mediterranean African silver leafing
Suitability of cassava for feeding and oviposition by B. tabaci
Adult survival
The suitability for adult feeding was measured in terms of average adult survival on cassava. Cassava cultivar had no significant effect on the adult survival of the seven B. tabaci species: EA1, MEAM1, MED-ASL, SSA1-SG1, SSA1-Hoslundia, SSA6 and SSA12 (F = 0.35, DF = 2, P = 0.70), and there was no significant interaction between cassava cultivar and B. tabaci species (F = 1.14, DF = 12, P = 0.34). The type of B. tabaci species affected adult survival significantly on cassava (F = 101.81, DF = 6, P < 0.0001; Fig. 6). SSA1-SG1 had the longest survival time (10.4 ± 1.8 d), while SSA12 had the shortest survival time (1.1 ± 0.3 d). Only SSA1-SG1 lived longer than 3 days on cassava.
Survival of different B. tabaci species on cassava (data on the three cultivars pooled together), evaluated for 18 days. Error bars represent standard error of the means (N = 4). Different letters above the bars indicate significant differences (P < 0.0001). SSA: sub-Saharan Africa, MEAM1: Middle East-Asia Minor1, MED-ASL: Mediterranean African silver leafing and SG: subgroup
Fecundity
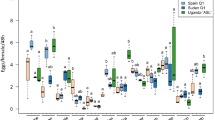
Cassava cultivar had no significant effect on the number of eggs laid by the different B. tabaci species (P = 0.55, GLM analysis of deviance) and no significant interaction occurred between cassava cultivar and B. tabaci species (P = 0.60, GLM analysis of deviance). B. tabaci species had a significant effect on the number of eggs laid on cassava (P < 0.0001, GLM analysis of deviance; Fig. 7). SSA12 laid the least number of eggs (0.3 ± 0.2), while SSA1-SG1 laid the highest number of eggs on cassava (243.7 ± 101.9). The nymphs for EA1, MEAM1, MED-ASL, SSA1-Hoslundia, SSA6 and SSA12 did not develop beyond the first-instar and eggs that did not hatch were shrivelled or dehydrated (Fig. 8).
Number of eggs laid by different B. tabaci species in 18 days on cassava (data on the three cultivars pooled together). Error bars represent standard error of the means (N = 4). Different letters above the bars indicate significant differences (P < 0.0001). SSA: sub-Saharan Africa, MEAM1: Middle East-Asia Minor1, MED-ASL: Mediterranean African silver leafing and SG: subgroup
First-instar nymphs and eggs on cassava A: healthy first-instar, B: dehydrated first-instar, C: shrivelled first-instar, D: healthy egg, E: shrivelled egg, and F: dehydrated egg. The nymphs of non-cassava B. tabaci died at first-instar stage and the eggs that did not hatch were shrivelled or dehydrated. The photographs were taken at × 50 magnification on a stereomicroscope (Nikon SMZ18) with an attached DSLR camera (Nikon D5300) and illuminated with a LED light source (Photonic Optics F3000)
Discussion
This study comprehensively investigated host plants utilised by B. tabaci species especially for growth and development in order to understand the role that alternative host plants play in their population dynamics.
The findings indicate that in Uganda, cassava is only colonised by cassava B. tabaci, SSA1 (-SG1 and -SG2), and SSA2, which was consistent with previous studies on cassava-colonising B. tabaci (Legg et al. 2002, 2014; Sseruwagi et al. 2005, 2006). The very closely related SSA1-Hoslundia (a subgroup of SSA1 at mtCOI level), however, was not detected on cassava but colonised A. africana, M. glaziovii, H. opposita, A. conyzoides, S. acuta and I. batatas. A. africana, H. opposita, A. conyzoides and S. acuta are wild hosts and are considered to be medicinal, except S. acuta. The medicinal plants are often found on field edges that have been left by farmers to fallow or hedge barriers in cases where there no nearby shrubs or forests. Medicinal plants are of great importance to women who pick, process and sell them. A. africana is used to stop bleeding and to treat wounds (Komakech et al. 2019). H. opposita treats skin blisters, worms, diarrhoea and yellow fever (Namukobe et al. 2011). It is also used as a stomach cleanser and to treat postnatal pain and wounds. A. conyzoides is used to treat wounds and provides strength to pregnant women (Namukobe et al. 2011; Tugume et al. 2016). M. glaziovii was introduced into East Africa in the nineteenth century as a source of rubber (Munro 1983) but is currently used as an ornamental and hedge-barrier plant in Uganda. I. batatas is an important food crop in Uganda (Mwanga and Ssemakula 2011) that was introduced in East Africa in the sixteenth century (Grigg 1974). It is commonly intercropped with cassava by small scale farmers. It was hypothesised that the inability of some B. tabaci to colonise cassava could be due to the poisonous hydrogen cyanide (Legg 1996).
Hydrogen cyanide (HCN) is stored in an intact form as cyanogenic glucosides in the cell vacuoles of cassava but tissue damage such as insect feeding exposes cyanogenic glucosides to β-glucosidases, leading to the formation of HCN (Calatayud et al. 1994). However, SSA1-Hoslundia colonised M. glaziovii, a wild relative of cassava which also contains cyanogenic glucosides (Joseph et al. 2000), so this is unlikely to be the reason it did not colonise cassava. Studies have shown that wild relatives of cassava contain a higher cyanogenic content than cassava (Wang et al. 2014). SSA1-Hoslundia does not appear to have evolved the ability to colonise cassava. This may be due to the high sugar levels in the cassava phloem sap (Li et al. 2016; Yan et al. 2019), which may create a high osmotic potential that is unsuitable for SSA1-Hoslundia. Transcriptome studies revealed that a cultivated cassava variety (KU50) from South East-Asia had a significantly higher number of genes involved in sucrose transport than in a wild subspecies of cassava (W14) (Wang et al. 2014).
In addition to cassava, SSA1-SG1 colonised 11 plant species from seven families and these included: A. africana, E. heterophylla, P. niruri, M. glaziovii, P. urens, L. camara, H. opposita, O. gratissimum, B. pilosa, V. amygdalina and I. batatas while, SSA1-SG2 colonised E. heterophylla and M. glaziovii. These results complement findings by Sseruwagi et al. (2006) who reported that SSA1 colonised five alternative hosts, M. glaziovii, J. gossypifolia, E. heterophylla, A. africana, and A. esculentus. However, SSA1 species were not detected on A. esculentus in our study area. We also did not identify J. gossypifolia in the area where the study was carried out. MEAM1 and MED-ASL (recently called Uganda ‘ASL’) developed on A. esculentus in the field. MEAM1 has been reported to displace indigenous species in the Americas, Australia and China, particularly in areas where insecticides are used, due to its resistance to several groups of insecticides (Costa and Brown 1991; De Barro et al. 2006; Zang et al. 2006; Hu et al. 2011). Insecticide usage in our study area was limited and thus it could be that MEAM1 and MED-ASL are more specialised in utilising A. esculentus than SSA1. It has been hypothesised that insect herbivores may shift from one hostplant to another as result of competition (Gassmann et al. 2006). Therefore, SSA1-SG1 could have shifted from A. esculentus due to competition from MEAM1 and MED-ASL. SSA2 did not colonise the alternative hosts. This could be due to the replacement by SSA1-SG1 as earlier reported by Legg et al. (2014), leading to the low population that made it hard to be detected on other hosts.
The alternative host plants that were colonised by SSA1-SG1 were mainly wild hosts, of which E. heterophylla and P. urens are common weeds. L. camara, A. africana, P. niruri, H. opposita, O. gratissimum, B. pilosa and V. amygdalina are medicinal plants (Namukobe et al. 2011). Most of them are commonly found in the fallow borders, hedge-barriers, shrubs, or forests except B. pilosa and Phyllanthus niruri, which are common weeds in the gardens. L. camara treats ring worms and tuberculosis (Kirimuhuzya et al. 2009; Tugume et al. 2016). B. pilosa stops bleeding and treats wounds. P. niruri is used to treat measles, wounds, fever and stomach problems (Tugume et al. 2016). O. gratissimum is a remedy for stomach pain (Tugume et al. 2016). V. amygdalina cures malaria, stomach-ache and convulsions (Namukobe et al. 2011; Tugume et al. 2016). Although SSA1-SG1 was polyphagous, it did not colonise A. conyzoides and S. acuta, which were colonised by SSA1-Hoslundia. These clear differences in host plant utilisation between the two populations suggest that these could be separate species. This however, requires further investigation through reciprocal crosses.
SSA1-SG1 was more prevalent on cassava than alternative host plants commonly found in proximity to cassava, suggesting that it has a preference for cassava. Omondi et al. (2005) examined the host preference of cassava whiteflies in Ghana and revealed that they significantly chose cassava for oviposition and development than A. esculentus, S. melongena, S. lycopersicum, S. aethiopicum and V. unguiculata. In addition, SSA1-SG1 colonised 11 alternative host plants found in close proximity to cassava compared to > 300 m from cassava, where it colonised only six hosts. This implies that the high populations on cassava spread to nearby hosts including poor hosts such as L. camara (Namuddu et al., Unpublished data), whereas at a distance of > 300 m, SSA1-SG1 selects the most suitable hosts.
Cassava was introduced in East Africa in eighteenth century (Guthrie 1987). A study by Mugerwa et al. (2018) reported that Africa is the centre of origin of B. tabaci. In this study, 75% (9/12) of the host plants colonised by SSA1-SG1 were wild hosts, implying that these might be the native plants on which this species evolved. The strong preference for cassava may be the result of much more recent evolutionary change due to the intensive and continuous cultivation of cassava. Furthermore, six and one of the host plants colonised by SSA1-SG1 and SSA1-SG2, respectively, were found at least 300 m away from cassava, showing that these populations are capable of surviving on other plant species. These alternative hosts allow these whitefly species to survive in the absence of cassava and colonise, whenever it is planted. The high whitefly populations on cassava in Uganda could be due to the wide host plant range of SSA1-SG1. Studies have shown that the presence of alternative host plants provides a continuous breeding habitat for insect pests leading to increased populations (Joyce 1983; Sétamou et al. 2000; Johannesen and Riedle-Bauer 2014). Sétamou et al. (2000) reported that the abundant wild hosts for a snout moth, Mussidia nigrivenella Ragonot in Benin were responsible for its high numbers on maize during the cropping season. Similarly, Johannesen and Riedle-Bauer (2014) observed that a new host plant, Urtica dioica L. led to increased populations of a plant-hopper, Hyalesthes obsoletus Signoret on grapes in Austria.
The ability of SSA1-SG1 to utilise several alternative hosts increases the risk of transmitting viruses between these hosts and cassava. Findings by Amisse et al. (2019) showed that Cassava Brown Streak Virus (CBSV) infected Zanha africana (Radlk.) Exell and Trichodesma zeylanicum (Burm.f.) R.Br. that are commonly found near cassava fields in Mozambique. They also detected CBSV and Ugandan CBSV in M. glaziovii. In India, Rajinimala et al. (2009) identified the bitter gourd yellow mosaic geminivirus in cassava, A. indica, C. sparsiflorus and M. coromandelianum. The availability of various host plants for SSA1-SG1 may result in the transmission of unknown viruses to cassava that could be epidemiologically important. These host plants may also serve as reservoirs for cassava viruses.
This study found non-cassava B. tabaci species that colonised only non-cultivated plants and those that developed on both cultivated and non-cultivated plants. IO, SSA6, SSA10, SSA12 and SSA13 colonised only non-cultivated plant species. IO colonised P. urens, S. acuta and C. benghalensis. Contrary to our findings, IO (Ug7) was previously reported as a polyphagous species that colonised cultivated plant species namely, P. vulgaris and G. hirsutum and a weed species, C. benghalensis in Uganda (Sseruwagi et al. 2005). SSA6, SSA10 and SSA12 colonised only O. gratissimum, V. amygdalina and C. benghalensis, respectively. This study is in agreement with the findings of Sseruwagi et al. (2005) that SSA6 (Ug3) colonised only O. gratissimum in Uganda. EA1 colonised H. opposita, P. urens, L. camara, B. pilosa and a cultivated plant species, P. vulgaris. EA1 was first identified from a single adult in Tanzania (Legg et al. 2014), and here we report that it is polyphagous, colonising both cultivated and non-cultivated plants. However, its role as a vector requires further investigation. MEAM1 utilised P. vulgaris, A. esculentus, S. lycopersicum and a weed species, S. acuta, while MED-ASL colonised non-cultivated plants including H. opposita, P. urens, S. acuta, A. africana and cultivated plants such as A. esculentus, V. unguiculata, I. batatas and G. hirsutum. Our findings supplement previous studies in Uganda, which reported that MED-ASL (Ug4) was polyphagous and colonised Cucurbita sativus, C. pepo, L. nepetifolia and P. urens, whereas MEAM1 (Ug6) was oligophagous and developed on A. esculentus and C. benghalensis (Sseruwagi et al. 2005). Vyskočilová et al. (2019) also observed that MED-ASL (Uganda ‘‘ASL’’) developed well on G. hirsutum, A. esculentus, I. batatas, S. melongena and C. pepo.
The findings from our study revealed that the polyphagy of SSA1-SG1 and other B. tabaci like MED-ASL contributes to the high populations on cassava and other crops, respectively. Ability to utilise various host plants makes the survival of polyphagous B. tabaci more flexible and their control difficult.
García and Zerbini (2019) proposed that uncultivated wild hosts were the original hosts for majority of the causal viruses from which they were transferred to crops by polyphagous whitefly vector populations. The ability of some whitefly species (SSA1-SG1, MED-ASL and B. Uganda1) to colonise both crops and weeds could potentially increase the transmission of viruses between plants resulting in mixed infections. Recombinant begomoviruses have been detected in cassava (Zhou et al. 1997; Berrie et al. 2001) and the viruses often originate from other hosts, suggesting the ability of polyphagous vector populations to facilitate formation of novel viruses.
Our findings also showed that EA1, MEAM1, MED-ASL, SSA1-Hoslundia, SSA6 and SSA12 cannot develop on cassava. Adults of these cryptic species of B. tabaci did not survive on cassava longer than three days. The immatures did not develop beyond first-instar stage and the eggs that did not hatch were shrivelled or dehydrated. In a host transfer experiment, Legg (1996) observed that B. tabaci adults collected from G. hirsutum and I. batatas in the field (mtCO1 species identity was not determined) and established on these host plants, did not develop on cassava and none of the adults lived longer than two days. The first-instar nymphs also died soon after hatching. The shrivelled and dehydrated eggs were probably laid by dying females that could not make a slit in the leaf to insert the pedicel, which absorbs water and solutes from the plant for embryo development and eclosion (Buckner et al. 2002). Mortality of adults and first-instars was probably as result of failure to feed successfully due to the inability to penetrate the leaf cuticle or presence of toxic compounds in cassava phloem sap. Feeding studies conducted by Milenovic et al. (2019) using an electrical penetration graph (EPG) system showed that feeding by non-cassava B. tabaci MED on cassava was characterised by short total phloem ingestion periods (< 1 h) compared to that of the cassava B. tabaci SSA1-SG3, which was > 4 h. There was longer total xylem feeding for MED, suggesting that the whiteflies were not ingesting enough phloem sap and thus became dehydrated. Douglas (2006) reported that phloem-feeding insects encounter a high osmotic pressure created by sucrose in the phloem sap, which can lead to dehydration. The high sucrose levels in cassava (Wang et al. 2014) probably led to dehydration of the non-cassava B. tabaci adults and first-instar nymphs. In addition, cassava is a well-defended plant species that contains cyanogenic glucosides (McMahon et al. 1995) and flavonoids (Prawat et al. 1995) and possibly suitable for only adapted whitefly species.
Some adults of EA1, MEAM1, MED-ASL, SSA1-Hoslundia, SSA6 and SSA12 survived on cassava for at least one day, which increases the possibility of transmission of non-persistent plant viruses that require only insect probing (Polston et al. 2014). When searching for suitable hosts, phloem-feeders make shallow probes to detect the carbohydrate content and presence or absence of secondary metabolites (Walling 2008). Milenovic et al. (2019) observed that during a 12 h EPG recording period, MED whiteflies were able to ingest cassava phloem sap for 41 min.
Conclusions
The data obtained in this study showed that cassava in Uganda is only colonised by cassava B. tabaci i.e. SSA1-SG1, SSA1-SG2, and SSA2. The findings also indicate that the host range of SSA1-SG1 is much wider than that of other cassava-colonising B. tabaci species in Uganda, which plays an important role in its predominance. Its ability to colonise other plant species facilitates its survival during periods when cassava is not available. It could also acquire other whitefly-transmitted viruses, which are not known to infest cassava and may be of epidemiological importance. The alternative hosts may also act as reservoirs for known cassava viruses.
This study provides important information for making decisions on pest management practices. For instance, when making choices of cultural practices, cassava should not be intercropped with I. batatas because it may facilitate the multiplication and spread of SSA1-SG1 in the field. Weed management is a measure that can be used to reduce the role of wild host plants on the population build-up of SSA1-SG1. However, farmers might be reluctant to remove medicinal plants that are of great importance to them. Therefore, whitefly-resistant varieties would be a more cost-effective and sustainable option. Non-cassava B. tabaci, MED-ASL was polyphagous in Uganda, colonising horticultural crops, food crops and weeds, which makes it a threat to agricultural productivity. EA1 was also polyphagous, utilising cultivated and non-cultivated plant species. However, its role as a vector requires further investigation.
Additional information
The 29 unique haplotype sequences of whiteflies identified in this study were deposited in the GenBank sequence database (Accession numbers PP554199–PP554227).
Data availability
Sequence data that support the findings of this study have been deposited in the GenBank sequence database of the National Center for Biotechnology Information (USA), Accession numbers PP554199–PP554227.
References
Amisse JJG, Ndunguru J, Tairo F et al (2019) First report of cassava brown streak viruses on wild plant species in Mozambique. Physiol Mol Plant Pathol 105:88–95. https://doi.org/10.1016/j.pmpp.2018.10.005
Ayres DL, Darling A, Zwickl DJ (2012) BEAGLE: an application programming interface and high-performance computing library for statistical phylogenetics. Syst Biol 61:70–173. https://doi.org/10.1093/sysbio/syr100
Berrie LC, Rybicki EP, Rey ME (2001) Complete nucleotide sequence and host range of South African cassava mosaic virus: further evidence for recombination amongst begomoviruses. J Gen Virol 82:53–58. https://doi.org/10.1099/0022-1317-82-1-53
Bird J (1957) A whitefly-transmitted mosaic of Jatropha gossypifolia, 22nd ed. Agricultural Experiment Station, University of Puerto Rico. https://doi.org/10.5555/19590500526
Brown JK, Frohlich DR, Rosell RC (1995) The sweet potato or silverleaf whiteflies: biotypes of Bemisia tabaci or a species complex? Annu Rev Entomol 40:511–534. https://doi.org/10.1146/annurev.en.40.010195.002455
Buckner JS, Freeman TP, Ruud RL et al (2002) Characterization and functions of the whitefly egg pedicel. Arch Insect Biochem Physiol 49:22–33. https://doi.org/10.1002/arch.10006
Calatayud PA, Rahbé Y, Delobel B (1994) Influence of secondary compounds in the phloem-sap of cassava on expression of antibiosis towards the mealybug Phenacoccus manihoti. Entomol Exp Appl 72:47–57. https://doi.org/10.1111/j.1570-7458.1994.tb01801.x
Costa HS, Brown JK (1991) Variation in biological characteristics and esterase patterns among populations of Bemisia tabaci, and the association of one population with silverleaf symptom induction. Entomol Exp Appl 61:211–219. https://doi.org/10.1111/j.1570-7458.1991.tb01553.x
De Barro PJ, Bourne A, Khan SA, Brancatini VAL (2006) Host plant and biotype density interactions – their role in the establishment of the invasive B biotype of Bemisia tabaci. Biol Invasions 8:287–294. https://doi.org/10.1007/s10530-005-1261-6
De Barro PJ, Liu S, Boykin LM, Dinsdale AB (2011) Bemisia tabaci : a statement of species status. Annu Rev Entomol 56:1–19. https://doi.org/10.1146/annurev-ento-112408-085504
Dinsdale A, Cook L, Riginos C et al (2010) Refined global analysis of Bemisia tabaci (Hemiptera: Sternorrhyncha: Aleyrodoidea: Aleyrodidae) mitochondrial cytochrome oxidase 1 to identify species level genetic boundaries. Ann Entomol Soc Am 103:196–208. https://doi.org/10.1603/AN09061
Douglas AE (2006) Phloem-sap feeding by animals: problems and solutions. J Exp Bot 57:747–754. https://doi.org/10.1093/jxb/erj067
García-Arenal F, Zerbini FM (2019) Life on the edge: geminiviruses at the interface between crops and wild plant hosts. Annu Rev Virol 6:411–433. https://doi.org/10.1146/annurev-virology-092818-015536
Gassmann AJ, Levy A, Tran T, Futuyma DJ (2006) Adaptations of an insect to a novel host plant: a phylogenetic approach. Funct Ecol 20:478–485. https://doi.org/10.1111/j.1365-2435.2006.01118.x
Ghosh S, Bouvaine S, Maruthi MN (2015) Prevalence and genetic diversity of endosymbiotic bacteria infecting cassava whiteflies in Africa. BMC Microbiol 15:1–17. https://doi.org/10.1186/s12866-015-0425-5
Green PJ (1995) Reversible jump Markov chain Monte Carlo computation and Bayesian model determination. Biometrika 82:711–732. https://doi.org/10.1093/biomet/82.4.711
Grigg DB (1974) The diffusion of Crops and livestock. In: The agricultural systems of the world: an evolutionary approach. Cambridge University Press, London, pp 24–44. https://doi.org/10.1017/S0022050700099769
Guthrie EJ (1987) African cassava mosaic disease and its control. In: International seminar on African cassava mosaic disease and its control. Yamoussoukro, Ivory coast, pp 1–9. https://horizon.documentation.ird.fr/exl-doc/pleins_textes/divers17-09/010006388.pdf
Hu J, De Barro P, Zhao H et al (2011) An extensive field survey combined with a phylogenetic analysis reveals rapid and widespread invasion of two alien whiteflies in China. PLoS ONE 6:e16061. https://doi.org/10.1371/journal.pone.0016061
Johannesen J, Riedle-Bauer M (2014) Origin of a sudden mass occurrence of the stolbur phytoplasma vector Hyalesthes obsoletus (Cixiidae) in Austria. Ann Appl Biol 165:488–495. https://doi.org/10.1111/aab.12158
Joseph T, Yeoh HH, Loh CS (2000) Somatic embryogenesis, plant regeneration and cyanogenesis in Manihot glaziovii Muell. Arg. (ceara rubber). Plant Cell Rep 19:535–538. https://doi.org/10.1007/s002990050769
Joyce RJV (1983) Aerial transport of pests and pest outbreaks. EPPO Bull 13:111–119. https://doi.org/10.1111/j.1365-2338.1983.tb01585.x
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kennedy GG, Storer NP (2000) Life systems of polyphagous arthropod pests in temporally unstable cropping systems. Annu Rev Entomol 45:467–493. https://doi.org/10.1146/annurev.ento.45.1.467
Kirimuhuzya C, Waako P, Joloba M, Odyek O (2009) The anti-mycobacterial activity of Lantana camara a plant traditionally used to treat symptoms of tuberculosis in South-western Uganda. Afri Health Sci 9:40–45. PMCID: PMC2932521
Komakech R, Matsabisa MG, Kang Y (2019) The wound healing potential of Aspilia africana (Pers.) C. D. Adams (Asteraceae). J Evid Based Complement Altern Med 2019:1–12. https://doi.org/10.1155/2019/7957860
Kunz D, Tay WT, Elfekih S et al (2019) Take out the rubbish–removing NUMTs and pseudogenes from the Bemisia tabaci cryptic species mtCOI database. bioRxiv. https://doi.org/10.1101/724765
Legg JP (1996) Host-associated strains within Ugandan populations of the whitefly Bemisia tabaci (Genn.),(Hom., Aleyrodidae). J Appl Entomol 120:523–527. https://doi.org/10.1111/j.1439-0418.1996.tb01646.x
Legg JP, French R, Rogan D et al (2002) A distinct Bemisia tabaci (Gennadius) (Hemiptera: Sternorrhyncha: Aleyrodidae) genotype cluster is associated with the epidemic of severe cassava mosaic virus disease in Uganda. Mol Ecol 11:1219–1229. https://doi.org/10.1046/j.1365-294x.2002.01514.x
Legg JP, Sseruwagi P, Boniface S et al (2014) Spatio-temporal patterns of genetic change amongst populations of cassava Bemisia tabaci whiteflies driving virus pandemics in East and Central Africa. Virus Res 186:61–75. https://doi.org/10.1016/j.virusres.2013.11.018
Li YZ, Zhao JY, Wu SM et al (2016) Characters related to higher starch accumulation in cassava storage roots. Sci Rep 6:19823. https://doi.org/10.1038/srep19823
Maruthi MN, Colvin J, Thwaites RM et al (2004) Reproductive incompatibility and cytochrome oxidase I gene sequence variability amongst host-adapted and geographically separate Bemesia tabaci populations (Hemiptera: Aleyrodidae). Syst Entomol 29:560–568. https://doi.org/10.1111/j.0307-6970.2004.00272.x
McMohan JM, White WLB, Sayre RT (1995) Review article: cyanogenesis in cassava (Manihot esculenta crantz). J Exp Bot 46:731–741. https://doi.org/10.1093/jxb/46.7.731
Milenovic M, Wosula EN, Rapisarda C, Legg JP (2019) Impact of host plant species and whitefly species on feeding behavior of Bemisia tabaci. Front Plant Sci 10:1. https://doi.org/10.3389/fpls.2019.00001
Mugerwa H, Seal S, Wang HL et al (2018) African ancestry of New World, Bemisia tabaci-whitefly species. Sci Rep 8:2734. https://doi.org/10.1038/s41598-018-20956-3
Mugerwa H, Colvin J, Alicai T et al (2021) Genetic diversity of whitefly (Bemisia spp.) on crop and uncultivated plants in Uganda: implications for the control of this devastating pest species complex in Africa. J Pest Sci. https://doi.org/10.1007/s10340-021-01355-6
Munro JF (1983) ‘British rubber companies in East Africa before the First World War.’ J Afr Hist 24:269–379. https://doi.org/10.1017/S0021853700022064
Mwanga ROM, Ssemakula G (2011) Orange-fleshed sweet potatoes for food, health and wealth in Uganda. Int J Agric Sustain 9:42–49. https://doi.org/10.3763/ijas.2010.0546
Namuddu A, Seal S, van Brunschot S et al (2023) Distribution of Bemisia tabaci in different agro-ecological regions in Uganda and the threat of vector-borne pandemics into new cassava growing areas. Front Sustain Food Syst 7:1068109. https://doi.org/10.3389/fsufs.2023.1068109
Namukobe J, Kasenene JM, Kiremire BT et al (2011) Traditional plants used for medicinal purposes by local communities around the Northern sector of Kibale national park, Uganda. J Ethnopharmacol 136:236–245. https://doi.org/10.1016/j.jep.2011.04.044
Omondi AB, Obeng-Ofori D, Kyerematen RA, Danquah EY (2005) Host preference and suitability of some selected crops for two biotypes of Bemisia tabaci in Ghana. Entomol Exp Appl 115:393–400. https://doi.org/10.1111/j.1570-7458.2005.00296.x
Omongo CA, Kawuki R, Bellotti AC et al (2012) African cassava whitefly, Bemisia tabaci, resistance in African and South American cassava genotypes. J Integr Agric 11:327–336. https://doi.org/10.1016/S2095-3119(12)60017-3
Polston JE, Capobianco H (2013) Transmitting plant viruses using whiteflies. J Vis Exp 81:e4332. https://doi.org/10.3791/4332
Polston JE, De Barro P, Boykin LM (2014) Transmission specificities of plant viruses with the newly identified species of the Bemisia tabaci species complex. Pest Manag Sci 70:1547–1552. https://doi.org/10.1002/ps.3738
Prawat H, Mahidol C, Ruchirawat S et al (1995) Cyanogenic and non-cyanogenic glycosides from Manihot esculenta. Phytochemistry 4:1167–1173. https://doi.org/10.1016/0031-9422(95)00398-q
Rajinimala N, Rabindran R, Ramaiah M (2009) Host range and purification of bitter gourd yellow mosaic virus (BGYMV) in bitter gourd (Momordica charantia). Arch Phytopathol Plant Prot 42:499–507. https://doi.org/10.1080/03235400701191606
Sétamou M, Schulthess F, Gounou S et al (2000) Host plants and population dynamics of the ear borer Mussidia nigrivenella (Lepidoptera: Pyralidae) in Benin. Environ Entomol 29:516–524. https://doi.org/10.1603/0046-225X-29.3.516
Shah MMR, Liu TX (2013) Feeding experience of Bemisia tabaci (Hemiptera: Aleyrodidae) affects their performance on different host plants. PLoS ONE 8:e77368. https://doi.org/10.1371/journal.pone.0077368
Sivakoff FS, Rosenheim JA, Dutilleul P, Carrière Y (2013) Influence of the surrounding landscape on crop colonization by a polyphagous insect pest. Entomol Exp Appl 149:11–21. https://doi.org/10.1111/eea.12101
Sseruwagi P, Legg JP, Maruthi MN et al (2005) Genetic diversity of Bemisia tabaci (Gennadius) (Hemiptera: Aleyrodidae) populations and presence of the B biotype and a non-B biotype that can induce silver-leaf symptoms in squash, in Uganda. Ann Appl Biol 147:253–265. https://doi.org/10.1111/j.1744-7348.2005.00026.x
Sseruwagi P, Maruthi MN, Colvin J et al (2006) Colonization of non-cassava plant species by cassava whiteflies (Bemisia tabaci) in Uganda. Entomol Exp Appl 119:145–153. https://doi.org/10.1111/j.1570-7458.2006.00402.x
Tugume P, Kakudidi EK, Buyinza M et al (2016) Ethnobotanical survey of medicinal plant species used by communities around Mabira central forest reserve, Uganda. J Ethnobiol Ethnomed 12:5. https://doi.org/10.1186/s13002-015-0077-4
van Lenteren JC, Noldus LPJJ (1990) Whitefly-plant relationships: behavioural and ecological aspects. In: Gerling D (ed) Whiteflies: their bionomics, pest status and management. Intercept, Andover, pp 47–89
Vyskočilová S, Tay WT, van Brunschot S et al (2018) An integrative approach to discovering cryptic species within the Bemisia tabaci whitefly species complex. Sci Rep 8:10886. https://doi.org/10.1038/s41598-018-29305-w
Vyskočilová S, Seal S, Colvin J (2019) Relative polyphagy of “Mediterranean” cryptic Bemisia tabaci whitefly species and global pest status implications. J Pest Sci 92:1071–1088. https://doi.org/10.1007/s10340-019-01113-9
Walling LL (2008) Avoiding effective defenses: Strategies employed by phloem-feeding insects. Plant Physiol 146:859–866. https://doi.org/10.1104/pp.107.113142
Wang X, Luan J, Li J et al (2011) Transcriptome analysis and comparison reveal divergence between two invasive whitefly cryptic species. BMC Genom 12:458. https://doi.org/10.1186/1471-2164-12-458
Wang W, Feng B, Xiao J et al (2014) Cassava genome from a wild ancestor to cultivated varieties. Nat Commun 5:5110. https://doi.org/10.1038/ncomms6110
Werle E, Schneider C, Renner M et al (1994) Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Research 22:4354–4355. https://doi.org/10.1093/nar/22.20.4354
White JA, Kelly SE, Perlman SJ, Hunter MS (2009) Cytoplasmic incompatibility in the parasitic wasp Encarsia inaron: Disentangling the roles of Cardinium and Wolbachia symbionts. Heredity 102:483–489. https://doi.org/10.1038/hdy.2009.5
Xia WQ, Wang XR, Liang Y et al (2017) Transcriptome analyses suggest a novel hypothesis for whitefly adaptation to tobacco. Sci Rep 7:12102. https://doi.org/10.1038/s41598-017-12387-3
Yan W, Wu X, Li Y et al (2019) Cell wall invertase 3 affects cassava productivity via regulating sugar allocation from source to sink. Front Plant Sci 10:541. https://doi.org/10.3389/fpls.2019.00541
Zang LS, Chen WQ, Liu SS (2006) Comparison of performance on different host plants between the B biotype and a non-B biotype of Bemisia tabaci from Zhejiang, China. Entomol Exp Appl 121:221–227. https://doi.org/10.1111/j.1570-8703.2006.00482.x
Zein H, Mohamed A, Paredes JR et al (2018) Metabolic resistance, target-site insensitivity of ace1-type acetylcholinesterase, and biotype diversity of whiteflies field populations from Egypt. https://www.ncbi.nlm.nih.gov/nucleotide/MG788326.1
Zhou X, Liu Y, Calvert L et al (1997) Evidence that DNA-A of a geminivirus associated with severe cassava mosaic disease in Uganda has arisen by interspecific recombination. J Gen Virol 78:2101–2111. https://doi.org/10.1099/0022-1317-78-8-2101
Acknowledgements
The authors appreciate the funding from Bill and Melinda Gates Foundation to the Natural Resources Institute (UK), NaCRRI (Uganda) and the Hebrew University of Jerusalem. We thank the farmers who gave us access to collect whitefly nymphs from their fields. We are also grateful to Dr. Stephen Young for his assistance in data analysis. We appreciate Dr. Tadeo Kaweesi for critiquing the manuscript.
Funding
This work was funded by the Bill and Melinda Gates Foundation through the African Cassava Whitefly Project [Grant Agreement OPP1058938]. Under the grant conditions of the Foundation, a Creative Commons Attribution 4.0 Generic License has already been assigned to the Author Accepted Manuscript version that might arise from this submission.
Author information
Authors and Affiliations
Contributions
SM and JC conceptualised the work and acquired funding. AN and RK took part in data collection. AN, SB, and JC participated in data analysis. CO, OM, SM, SS, and JC were involved in supervision. AN drafted the original manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
This study does not contain any studies with human participants or vertebrate performed by any of the authors. Whiteflies are invertebrate insects and, according to the IUCN criteria, are not considered as endangered or protected species. Whitefly samples were collected from farmer’s fields after obtaining their permission.
Additional information
Communicated by Antonio Biondi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Namuddu, A., Malka, O., Seal, S. et al. Is polyphagy of a specific cryptic Bemisia tabaci species driving the high whitefly populations on cassava in eastern Africa?. J Pest Sci (2024). https://doi.org/10.1007/s10340-024-01832-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10340-024-01832-8