Abstract
Species belonging to the bacterial phylum Planctomycetes are ubiquitous members of the microbial communities in aquatic environments and are frequently isolated from various biotic and abiotic surfaces in marine and limnic water bodies. Planctomycetes have large genomes of up to 12.4 Mb, follow complex lifestyles and display an uncommon cell biology; features which motivate the investigation of members of this phylum in greater detail. As a contribution to the current collection of axenic cultures of Planctomycetes, we here describe strain Pla52T isolated from wood particles in the Baltic Sea. Phylogenetic analysis places the strain in the family Pirellulaceae and suggests two species of the recently described genus Stieleria as current closest neighbours. Strain Pla52nT shows typical features of members of the class Planctomycetia, including division by polar budding and the presence of crateriform structures. Colonies of strain Pla52nT have a light orange colour, which is an unusual pigmentation compared to the majority of members in the phylum, which show either a pink to red pigmentation or entirely lack pigmentation. Optimal growth of strain Pla52nT at 33 °C and pH 7.5 indicates a mesophilic (i.e. with optimal growth between 20 and 45 °C) and neutrophilic growth profile. The strain is an aerobic heterotroph with motile daughter cells. Its genome has a size of 9.6 Mb and a G + C content of 56.0%. Polyphasic analyses justify delineation of the strain from described species within the genus Stieleria. Therefore, we conclude that strain Pla52nT = LMG 29463T = VKM B-3447T should be classified as the type strain of a novel species, for which we propose the name Stieleria varia sp. nov.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Planctomycetes is a phylum of mostly aquatic bacteria, which can be found in various limnic and marine water bodies. Together with the medically and biotechnologically relevant phyla Chlamydiae and Verrucomicrobia and other sister phyla, the phylum Planctomycetes constitutes the PVC superphylum (Rivas-Marín and Devos 2018; van Niftrik and Devos 2017; Wagner and Horn 2006). Due to several presumptively eukaryotic characteristics (Fuerst and Sagulenko 2011), Planctomycetes were initially considered to be exceptions to the typical bacterial cell plan. However, with the introduction of novel microscopic techniques and the development of genetic tools for Planctomycetes (Jeske et al. 2016; Jogler et al. 2011; Rivas-Marín et al. 2016), several of their eukaryote-like characteristics have been reassessed. For example, the proposed intracellular compartments turned out to be rather invaginations of the cytoplasmic membrane (Boedeker et al. 2017). The identification of peptidoglycan in Planctomycetes (Jeske et al. 2015; Van Teeseling et al. 2015) led to the reinterpretation of their cell envelope architecture as being similar to that of Gram-negative bacteria (Devos 2014). Nevertheless, certain aspects of their cell biology are still exceptional. All characterised Planctomycetes lack ‘canonical’ divisome proteins including the otherwise universal FtsZ (Jogler et al. 2012; Rivas-Marin et al. 2020). Members of Planctomycetia, the class with the currently highest number of described species within the phylum Planctomycetes, divide by budding, whereas species belonging to the classes Phycisphaerae and Candidatus Brocadiae divide by binary fission (Wiegand et al. 2020).
The lifecycle of most Planctomycetes is complex and involves alternation between sessile cells attached to various abiotic and biotic aquatic surfaces, and flagellated swarmer cells (Faria et al. 2018; Lage et al. 2019). The sessile cells bud to form flagellated swarmer cells, which swim and relocate before settling down to attach and begin reproduction. In this context, Planctomycetes were found to be frequent colonisers of macroalgae (Bengtsson and Øvreås 2010; Faria et al. 2018; Lage and Bondoso 2014) and can even be the dominating phylum in microbial communities on biotic surfaces. For example, as recently shown, Planctomycetes can account for more than 80% of the bacterial community in seagrass meadows in the Mediterranean Sea (Kohn et al. 2020). Such findings appear counterintuitive when taking into account that the growth rates of Planctomycetes are often lower than those of many of their natural bacterial competitors occupying the same ecological niches (Frank et al. 2015). The observation that, despite slower growth, Planctomycetes can be abundant members in marine microbial communities led to the hypothesis that they apply different strategies to compensate for the disadvantage in growth speed, although most of these stategies remain undiscovered. It has been assumed that these strategies may involve the ability to produce bioactive secondary metabolites (Kallscheuer et al. 2019b; Panter et al. 2019), the observed resistance against several antibiotics (Cayrou et al. 2010; Godinho et al. 2019) and/or a metabolism well-adapted to digestion of phototroph-derived compounds, including complex polysaccharides (Wecker et al. 2009; Wegner et al. 2013).
Recently, a class of N-acylated tyrosine derivatives, designated stieleriacines, has been identified in the Planctomycete Stieleria maiorica Mal15T, which can influence the microbial community composition of biofilms inhabited by this species (Kallscheuer et al. 2020a). Structurally related compounds of the same class were also found in the closely related species Stieleria neptunia (Sandargo et al. 2020). In silico genome analyses indicate that the ability to produce secondary metabolites is widespread in the phylum, in particular in the class Planctomycetia. The hitherto investigated planctomycetal genomes feature sizes of up to 12.4 Mb (Ravin et al. 2018) and between 1 and 13 putative secondary metabolite-associated gene clusters were identified during in silico genome analyses (Wiegand et al. 2020). These clusters are similar to previously investigated clusters, e.g. those found in Actinobacteria, and may be similarly involved in the biosynthesis of non-ribosomal peptides, polyketides, terpenes, bacteriocins and others. Consequently, Planctomycetes are considered to be untapped producers of small molecules with potential therapeutically useful bioactivities (Calisto et al. 2019; Graça et al. 2016; Jeske et al. 2016).
To extend our knowledge on Plantomycetes in general and the genus Stieleria in particular, we herein characterise strain Pla52nT by using physiological, microscopic, genomic, and phylogenetic methods. Based on these analyses, we propose that strain Pla52nT constitutes the third species of the recently described genus Stieleria.
Materials and methods
Isolation of strain Pla52nT and cultivation
Strain Pla52nT was isolated from wood particles placed in an incubator and stored for two weeks (August–September 2014) at a depth of 2 m in the Baltic Sea, below a landing stage at Heiligendamm (‘Seebrücke Heiligendamm’, 54.146 N 11.843 E) (Oberbeckmann et al. 2018). In the laboratory, biofilms formed on the wood particles were removed by incubation with β-galactosidase (2 mg/mL, 30 °C, pH 4.7) for 30 min and subsequent sonication for 10 min at 30 °C. M1 medium buffered with 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) and supplemented with N-acetyl glucosamine (NAG) and artificial seawater (ASW) (designated M1H NAG ASW medium) (Boersma et al. 2019) was used for the cultivation. The medium was solidified with 8 g/L gellan gum and additionally supplemented with 500 mg/L streptomycin, 100 mg/L ampicillin and 20 mg/L cycloheximide. The cell suspension obtained after sonication was streaked on an M1H NAG ASW plate, incubated at 20 °C for six weeks and regularly checked for the presence of colonies. Colonies obtained were then subjected to 16S rRNA gene amplification and sequencing according to a previously published protocol (Rast et al. 2017). This step was included to check whether strains are members of the phylum Planctomycetes. Colonies of strains confirmed as members of the phylum Planctomycetes were re-streaked on M1H NAG ASW plates, which then served as a source for the inoculation of liquid cultures in M1H NAG ASW medium. After several days of cultivation, exponentially growing cells were used for subsequent cultivation experiments. Determination of the pH optimum for growth was performed by cultivation of strain Pla52nT in M1H NAG ASW at 28 °C with 100 mM of the following buffers: 2-(N-morpholino)ethanesulfonic acid (MES) for pH 5.0-6.5, HEPES for pH 7.0-8.0, 3-(4-(2-hydroxyethyl)piperazin-1-yl)propane-1-sulfonic acid (HEPPS) for pH 8.0 and N-cyclohexyl-2-aminoethanesulfonic acid (CHES) for pH 9.0-10.0. Cultivations for determination of the temperature optimum for growth were performed in M1H NAG ASW medium at pH 7.5. Growth of the strain was measured as optical density at 600 nm (OD600). Maximal growth rates µmax were obtained by determination of the slope in the plot of the natural logarithmic function of average OD600 values from biological triplicates against the cultivation time. The slope from at least five data points in the exponential growth phase was used as growth rate µmax (in h−1). The generation time td (in h) was calculated using the equation td = ln(2)/µmax.
Microscopy
Microscopic analyses included phase contrast light microscopy and field emission scanning electron microscopy (SEM) and were performed as previously described (Boersma et al. 2019).
Genome information and antiSMASH analysis
Sequencing of the genome of strain Pla52nT was conducted as part of a previous study (Wiegand et al. 2020). Genome and 16S rRNA gene sequence of strain Pla52nT are available from GenBank under accession numbers GCA_007860045 and MK554582, respectively. Analysis of secondary metabolite-associated gene clusters was performed using antiSMASH bacterial version 5.1.2 with relaxed strictness and the following extra features enabled: KnownClusterBlast, ActiveSiteFinder and SubClusterBlast (Blin et al. 2019).
Phylogenetic analysis
Maximum likelihood phylogeny was computed for strain Pla52nT, the type strains of all described planctomycetal species (assessed in May 2020) including strains of the family Pirellulaceae published and described in the recent year (Kallscheuer et al. 2019a, c, 2020a, b, c; Kumar et al. 2020; Peeters et al. 2020a, b; Rensink et al. 2020; Sandargo et al. 2020). Phylogenetic trees based on 16S rRNA gene sequences and multi-locus sequence analysis (MLSA) were calculated as previously described (Boersma et al. 2019). 16S rRNA gene sequences from Opitutus terrae (acc. no. AJ229235), Kiritimatiella glycovorans (acc. no. NR_146840) and Lentisphaera araneosa (acc. no. NR_027571) were used as outgroup in the 16S rRNA gene sequence-based tree. Two members of the family Planctomycetaceae, Planctopirus limnophila and Gimesia maris, were used as outgroup in the MLSA-based tree.
Average nucleotide identities (ANI) were calculated using OrthoANI (Lee et al. 2016), average amino acid identities (AAI) using the aai.rb script of the enveomics collection (Rodriguez-R and Konstantinidis 2016) and percentage of conserved proteins (POCP) as previously described (Qin et al. 2014). The rpoB nucleotide sequences were taken from publicly available genome annotations and the sequence identities for a partial sequence fragment of 1200 bp expected to be amplified with the described primer set were determined according to Bondoso et al. (2013).
Results and discussion
Phylogenetic inference
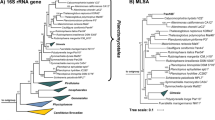
Maximum likelihood phylogenetic trees based on 16S rRNA genes sequences and MLSA place strain Pla52nT in the recently introduced family Pirellulaceae (Dedysh et al. 2019). In this family, strain Pla52nT clusters monophyletically with two recently described species of the genus Stieleria, namely S. maiorica and S. neptunia (Kallscheuer et al. 2020a; Sandargo et al. 2020) (Fig. 1). Five investigated phylogenetic markers also identified these two species as the current closest relatives of strain Pla52nT (Fig. 2). Comparison of the 16S rRNA gene sequence of strain Pla52nT to those of S. maiorica Mal15T and S. neptunia Enr13T yielded similarities of 96.0% and 95.9%, respectively (Fig. 2). Both values are above the proposed genus threshold of 94.5% (Yarza et al. 2014) but below the species threshold of 98.7% (Stackebrandt and Ebers 2006), indicating that strain Pla52nT is a member of the genus Stieleria, but does not belong to either of the two described species. This finding is in accordance with results obtained during an analysis of additional phylogenetic markers, including AAI, POCP and rpoB similarity when applying the proposed genus thresholds of 60%, 50% and 75.5–78%, respectively (Kallscheuer et al. 2019c; Konstantinidis and Tiedje 2005; Qin et al. 2014) (Fig. 2). In addition, none of the values obtained during comparison of strain Pla52nT with its close neighbours were found to be above the species threshold for AAI and ANI of 95% (Kim et al. 2014; Konstantinidis and Tiedje 2005) and for rpoB of 96.3% (Bondoso et al. 2013). The conclusion that strain Pla52nT belongs to a novel species within the genus Stieleria is thus supported by all analysed markers.
Maximum likelihood phylogenetic analysis of strain Pla52nT. Phylogenetic trees based on 16S rRNA gene sequences and MLSA were computed as described in the Materials and methods section. Bootstrap values after 1000 re-samplings (16S rRNA gene)/500 re-samplings (MLSA) are given at the nodes (in %). The outgroup in the 16S rRNA gene-based tree consists of 16S rRNA genes from three strains outside of the phylum Planctomycetes but part of the PVC superphylum. In the MLSA tree the genomes of Planctopirus limnophila and Gimesia maris (both family Planctomycetaceae) served as outgroup
Analysis of phylogenetic markers used for the delineation of strain Pla52nT from characterised species of the genus Stieleria. Analysed markers included 16S rRNA gene sequence similarity (16S), average amino acid identity (AAI), average nucleotide identity (ANI), identity of a 1200 bp fragment of the gene rpoB and percentage of conserved proteins (POCP)
Morphological and physiological analyses
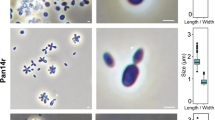
Based on the phylogenetic position of strain Pla52nT, its phenotypic and genomic characteristics were compared to the two Stieleria species (Figs. 3, 4; Table 1). Cells of strain Pla52nT have an average size of 1.8 ± 0.3 × 0.9 ± 0.2 µm (Fig. 3a, c), which is an elongated shape compared to more roundish or pear-shaped cells of S. neptunia Enr13T and S. maiorica Mal15T. The shape of mature Pla52nT cells turned out to vary from ovoid to round grain rice-shaped and is less uniform compared to the other two strains (Fig. 3b, d, e), which is reflected in the proposed name of the novel species represented by the type strain Pla52nT. All three compared strains either occur as single cells or form clusters, however, while S. maiorica and S. neptunia form larger aggregates, strain Pla52nT tends to form rosettes which often assemble to short chains (Fig. 3d). Planktonic cells of all three strains are motile and at least for S. maiorica a clear lifestyle switch with sessile mother cells and swarming motile daughter cells was observed. Cells of the three strains contain crateriform structures and lack a stalk or holdfast structure. In case of strain Pla52nT, matrix or fimbriae originates from one of the poles and forms a characteristic fibre cap. Colonies of the strain have a light orange pigmentation, a quite rare pigmentation amongst characterised strains in the phylum, while the other two species have the more common pink pigmentation. Strain Pla52nT can thus be clearly distinguished from the two described species, even with the naked eye. All three strains are aerobic heterotrophs and can grow up to temperatures of 35–37 °C. The temperature optimum of strain Pla52nT (33 °C) (Fig. 4) falls between the optima of S. maiorica Mal15T (35 °C) and S. neptunia Enr13T (28 °C). Optimal growth is observed at pH 7.5 for all three strains. During laboratory-scale shaking flask cultivations in M1H NAG ASW medium, strain Pla52nT reached a maximal growth rate of 0.061 h−1 (generation time of 11 hours, Fig. 4). Its growth rate is slightly higher than that observed for S. neptunia Enr13T (0.054 h−1, td = 13 h), but considerably lower compared to S. maiorica Mal15T (0.093 h−1, td = 7.5 h).
Microscopy images and cell size plot of strain Pla52nT. The mode of cell division (a) and a general overview of cell morphology (b, d, e) is shown in the micrographs, respectively. For determination of the cell size (c) at least 100 representative cells were counted manually or by using a semi-automated object count tool
Genomic characteristics of strain Pla52nT
The genome of strain Pla52nT has a size of 9.59 Mb and a DNA G + C content of 56%. With such values the strain has currently the smallest genome and lowest G + C content in the genus Stieleria (Table 1). Its genome harbours 7094 genes, of which 6998 are putatively protein-coding. Automated gene annotation yielded 3223 genes coding for hypothetical proteins or proteins of unknown function, accounting for 46% of the total number of annotated proteins in the genome. These values fall within the range of 40–55% hypothetical proteins found to be encoded in most planctomycetal genomes sequenced so far and is comparable to the other two Stieleria species (42–43%). Given the relatively large genomes of the three strains, the presence of giant genes (with open reading frames > 15 kb) is not surprising. Strain Pla52nT harbours 12 such genes, a number comparable to the two strains used for comparison (13–14 giant genes). An in silico analysis of the encoded proteins (> 5000 aa) based on domains detected by InterPro scan points towards a role as adhesion proteins or extracellular proteins with glycosyl hydrolase activity (Table 2). Plasmids were not observed in the genomes within the genus Stieleria. Strain Pla52nT harbours a single copy of the 16S rRNA gene, whereas 3 copies can be found in the other two strains.
Analysis of gene clusters putatively involved in the production of secondary metabolites
Given the relatively large genomes of 9.6–11.0 Mb observed for the hitherto characterised species in the genus Stieleria, they are probably amongst the talented producers of secondary metabolites in the phylum Planctomycetes as confirmed by recently published studies (Kallscheuer et al. 2020a; Sandargo et al. 2020). We thus analysed the three genomes using antiSMASH to check for gene clusters potentially related to secondary metabolite production (Table 3). The analysis indicated that the three strains Pla52nT, S. neptunia Enr13T and S. maiorica Mal15T harbour a total number of 9-11 of such clusters. These numbers are in the upper range when considering the range of 1-13 clusters identified by antiSMASH in strains belonging to the phylum Planctomycetes (Wiegand et al. 2020). S. neptunia Enr13T and S. maiorica Mal15T harbour N-acyl amino acid synthase-encoding genes, which are most likely involved in the biosynthesis of stieleriacines, a class of N-acylated tyrosine derivatives found to be produced by these two strains (Kallscheuer et al. 2020a; Sandargo et al. 2020) (Table 3). However, in the still non-closed genome of strain Pla52nT we could not identify genes coding for putative N-acyl amino acid synthases, which might indicate that strain Pla52nT does not produce stieleriacines. This in not entirely unlikely given that strain Pla52nT is more distantly related than the type strains of the other two Stieleria species (Fig. 2). Stieleriacine production is thus not necessarily a conserved feature within the genus Stieleria.
Other putative clusters associated with secondary metabolite production are related to polyketide and non-ribosomal peptide biosynthesis (Table 3). Two genes or clusters relevant to terpenoid production are likely involved in the production of carotenoids, as indicated by the orange or pink pigmentation of the Stieleria species.
Collectively, the polyphasic analysis justifies delineation of strain Pla52nT from characterised species in the genus Stieleria. Thus, we propose to assign the novel isolate to a novel species, for which we propose the name Stieleria varia sp. nov.
Stieleria varia sp. nov
Stieleria varia (va’ri.a. L. fem. adj. varia varied; corresponding to the varying size of the cells).
Cells are ovoid to round grain rice-shaped (average size: 1.8 ± 0.3 µm × 0.9 ± 0.2 µm; cell size and shape are not uniform), occur as single cells or rosettes, which tend to form short chains. Crateriform structures are formed on one cell pole, while cells lack a stalk or holdfast structure. Matrix or fimbriae are formed at the budding pole. The species is heterotrophic, aerobic, mesophilic and neutrophilic. Daughter cells are motile. Optimal growth is observed at 33 °C and pH 7.5. Colonies have a light orange pigmentation. The type strain genome has a DNA G + C content of 56.0%.
The type strain is Pla52nT (= LMG 29463T = VKM B-3447T, synonym: Pla52neu), isolated from wood particles in the Baltic Sea in September 2014. The type strain has a genome size of 9,586,696 bp (GenBank Accession Number GCA_007860045).
References
Bengtsson MM, Øvreås L (2010) Planctomycetes dominate biofilms on surfaces of the kelp Laminaria hyperborea. BMC Microbiol 10:261
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Boedeker C, Schüler M, Reintjes G, Jeske O, van Teeseling MC, Jogler M, Rast P, Borchert D, Devos DP, Kucklick M (2017) Determining the bacterial cell biology of Planctomycetes. Nat Commun 8:1–14
Boersma AS, Kallscheuer N, Wiegand S, Rast P, Peeters SH, Mesman RJ, Heuer A, Boedeker C, Jetten MS, Rohde M, Jogler M, Jogler C (2019) Alienimonas californiensis gen. nov. sp. nov., a novel Planctomycete isolated from the kelp forest in Monterey Bay. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01367-4
Bondoso J, Harder J, Lage OM (2013) rpoB gene as a novel molecular marker to infer phylogeny in Planctomycetales. Antonie Van Leeuwenhoek 104:477–488
Calisto R, Sæbø EF, Storesund JE, Øvreås L, Herfindal L, Lage OM (2019) Anticancer activity in planctomycetes. Front Mar Sci 5:499
Cayrou C, Raoult D, Drancourt M (2010) Broad-spectrum antibiotic resistance of Planctomycetes organisms determined by Etest. J Antimicrob Chemother 65:2119–2122
Dedysh SN, Kulichevskaya IS, Beletsky AV, Ivanova AA, Rijpstra WIC, Damsté JSS, Mardanov AV, Ravin NV (2019) Lacipirellula parvula gen. nov., sp. nov., representing a lineage of planctomycetes widespread in low-oxygen habitats, description of the family Lacipirellulaceae fam. nov. and proposal of the orders Pirellulales ord. nov., Gemmatales ord. nov. and Isosphaerales ord. nov. Syst Appl Microbiol 43:126050
Devos DP (2014) PVC bacteria: variation of, but not exception to, the Gram-negative cell plan. Trends Microbiol 22:14–20
Faria M, Bordin N, Kizina J, Harder J, Devos D, Lage OM (2018) Planctomycetes attached to algal surfaces: insight into their genomes. Genomics 110:231–238
Frank O, Michael V, Päuker O, Boedeker C, Jogler C, Rohde M, Petersen J (2015) Plasmid curing and the loss of grip–the 65-kb replicon of Phaeobacter inhibens DSM 17395 is required for biofilm formation, motility and the colonization of marine algae. Syst Appl Microbiol 38:120–127
Fuerst JA, Sagulenko E (2011) Beyond the bacterium: planctomycetes challenge our concepts of microbial structure and function. Nat Rev Microbiol 9:403–413
Godinho O, Calisto R, Øvreås L, Quinteira S, Lage OM (2019) Antibiotic susceptibility of marine Planctomycetes. Antonie Van Leeuwenhoek 112:1273–1280
Graça AP, Calisto R, Lage OM (2016) Planctomycetes as novel source of bioactive molecules. Front Microbiol 7:1241
Jeske O, Schüler M, Schumann P, Schneider A, Boedeker C, Jogler M, Bollschweiler D, Rohde M, Mayer C, Engelhardt H (2015) Planctomycetes do possess a peptidoglycan cell wall. Nat Commun 6:1–7
Jeske O, Surup F, Ketteniß M, Rast P, Förster B, Jogler M, Wink J, Jogler C (2016) Developing techniques for the utilization of Planctomycetes as producers of bioactive molecules. Front Microbiol 7:1242
Jogler C, Glöckner FO, Kolter R (2011) Characterization of Planctomyces limnophilus and development of genetic tools for its manipulation establish it as a model species for the phylum Planctomycetes. Appl Environ Microbiol 77:5826–5829
Jogler C, Waldmann J, Huang X, Jogler M, Glöckner FO, Mascher T, Kolter R (2012) Planctomycetes comparative genomics: identification of proteins likely involved in morphogenesis, cell division and signal transduction. J Bacteriol JB. 01325-12
Kallscheuer N, Jogler M, Wiegand S, Peeters SH, Heuer A, Boedeker C, Jetten MS, Rohde M, Jogler C (2019a) Three novel Rubripirellula species isolated from plastic particles submerged in the Baltic Sea and the estuary of the river Warnow in northern Germany. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01368-3
Kallscheuer N, Moreira C, Airs R, Llewellyn CA, Wiegand S, Jogler C, Lage OM (2019b) Pink-and orange-pigmented Planctomycetes produce saproxanthin-type carotenoids including a rare C45 carotenoid. Environ Microbiol Rep 11:741–748
Kallscheuer N, Wiegand S, Peeters SH, Jogler M, Boedeker C, Heuer A, Rast P, Jetten MS, Rohde M, Jogler C (2019c) Description of three bacterial strains belonging to the new genus Novipirellula gen. nov., reclassificiation of Rhodopirellula rosea and Rhodopirellula caenicola and readjustment of the genus threshold of the phylogenetic marker rpoB for Planctomycetaceae. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01374-5
Kallscheuer N, Jeske O, Sandargo B, Boedeker C, Wiegand S, Bartling P, Jogler M, Rohde M, Petersen J, Medema M, Surup F, Jogler C (2020a) The planctomycete Stieleria maiorica Mal15T employs stieleriacines to alter the species composition in marine biofilms. Commun Biol 3:303
Kallscheuer N, Wiegand S, Boedeker C, Peeters SH, Jogler M, Rast P, Heuer A, Jetten MS, Rohde M, Jogler C (2020b) Aureliella helgolandensis gen. nov., sp. nov., a novel Planctomycete isolated from a jellyfish at the shore of the island Helgoland. Antonie van Leeuwenhoek, https://doi.org/10.1007/s10482-020-01403-8
Kallscheuer N, Wiegand S, Heuer A, Rensink S, Boersma AS, Jogler M, Boedeker C, Peeters SH, Rast P, Jetten MS, Rohde M, Jogler C (2020c) Blastopirellula retiformator sp. nov. isolated from the shallow-sea hydrothermal vent system close to Panarea Island. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01377-2
Kim M, Oh H-S, Park S-C, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kohn T, Rast P, Wiegand S, Jetten MS, Kallscheuer N, Jeske O, Boedeker C, Rohde M, Jogler M, Jogler C (2020) The microbiome of Posidonia oceanica seagrass leaves can be dominated by Planctomycetes. Front Microbiol 11:1458
Konstantinidis KT, Tiedje JM (2005) Towards a genome-based taxonomy for prokaryotes. J Bacteriol 187:6258–6264
Kumar D, Gaurav K, Jagadeeshwari U, Ramana ChV (2020) Roseimaritima sediminicola sp. nov., a new member of Planctomycetaceae isolated from Chilika lagoon. Int J Syst Evol Microbiol 70:2616–2623
Lage OM, Bondoso J (2014) Planctomycetes and macroalgae, a striking association. Front Microbiol 5:267
Lage OM, van Niftrik L, Jogler C, Devos D (2019) Planctomycetes. In: Schmidt Thomas M (ed) Reference module in life sciences, 4th edn. Elsevier, Amsterdam, pp 614–626
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Oberbeckmann S, Kreikemeyer B, Labrenz M (2018) Environmental factors support the formation of specific bacterial assemblages on microplastics. Front Microbiol 8:2709
Panter F, Garcia R, Thewes A, Zaburannyi N, Bunk B, Overmann J, Gutierrez MV, Krug D, Müller R (2019) Production of a Dibrominated aromatic secondary metabolite by a planctomycete implies complex interaction with a macroalgal host. ACS Chem Biol 14:2713–2719
Peeters SH, Wiegand S, Kallscheuer N, Jogler M, Heuer A, Jetten MS, Boedeker C, Rohde M, Jogler C (2020a) Lignipirellula cremea gen. nov., sp. nov., a planctomycete isolated from wood particles in a brackish river estuary. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-020-01407-4
Peeters SH, Wiegand S, Kallscheuer N, Jogler M, Heuer A, Jetten MS, Rast P, Boedeker C, Rohde M, Jogler C (2020b) Three marine strains constitute the novel genus and species Crateriforma conspicua in the phylum Planctomycetes. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01375-4
Qin Q-L, Xie B-B, Zhang X-Y, Chen X-L, Zhou B-C, Zhou J, Oren A, Zhang Y-Z (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215
Rast P, Glockner I, Boedeker C, Jeske O, Wiegand S, Reinhardt R, Schumann P, Rohde M, Spring S, Glockner FO, Jogler C, Jogler M (2017) Three novel species with peptidoglycan cell walls form the new genus Lacunisphaera gen. nov. in the family Opitutaceae of the Verrucomicrobial Subdivision 4. Front Microbiol 8: 202
Ravin NV, Rakitin AL, Ivanova AA, Beletsky AV, Kulichevskaya IS, Mardanov AV, Dedysh SN (2018) Genome analysis of Fimbriiglobus ruber SP5T, a planctomycete with confirmed chitinolytic capability. Appl Environ Microbiol 84:e02645-17
Rensink S, Wiegand S, Kallscheuer N, Rast P, Peeters SH, Heuer A, Boedeker C, Jetten MS, Rohde M, Jogler M, Jogler C (2020) Description of the novel planctomycetal genus Bremerella, containing Bremerella volcania sp. nov., isolated from an active volcanic site, and reclassification of Blastopirellula cremea as Bremerella cremea comb. nov. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01378-1
Rivas-Marín E, Devos DP (2018) The paradigms they are a-Changin’: past, present and future of PVC bacteria research. Antonie Van Leeuwenhoek 111:785–799
Rivas-Marin E, Peeters SH, Fernández LC, Jogler C, van Niftrik L, Wiegand S, Devos DP (2020) Non-essentiality of canonical cell division genes in the planctomycete Planctopirus limnophila. Sci Rep 10:1–8
Rivas-Marín E, Canosa I, Santero E, Devos DP (2016) Development of genetic tools for the manipulation of the planctomycetes. Front Microbiol 7:914
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Preprints e1900v1
Sandargo B, Jeske O, Boedeker C, Wiegand S, Wennrich J-P, Kallscheuer N, Jogler M, Rohde M, Jogler C, Surup F (2020) Stieleriacines, N-Acyl Dehydrotyrosines from the marine planctomycete Stieleria neptunia sp. nov. Front Microbiol 11:1408
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
van Niftrik L, Devos DP (2017) Planctomycetes-Verrucomicrobia-Chlamydiae bacterial superphylum: new model organisms for evolutionary cell biology. Front Microbiol 8:1458
Van Teeseling MC, Mesman RJ, Kuru E, Espaillat A, Cava F, Brun YV, VanNieuwenhze MS, Kartal B, Van Niftrik L (2015) Anammox Planctomycetes have a peptidoglycan cell wall. Nat Commun 6:6878
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol 17:241–249
Wecker P, Klockow C, Ellrott A, Quast C, Langhammer P, Harder J, Glöckner FO (2009) Transcriptional response of the model planctomycete Rhodopirellula baltica SH1T to changing environmental conditions. BMC Genom 10:410
Wegner C-E, Richter-Heitmann T, Klindworth A, Klockow C, Richter M, Achstetter T, Glöckner FO, Harder J (2013) Expression of sulfatases in Rhodopirellula baltica and the diversity of sulfatases in the genus Rhodopirellula. Mar Genomics 9:51–61
Wiegand S, Jogler M, Boedeker C, Pinto D, Vollmers J, Rivas-Marín E, Kohn T, Peeters SH, Heuer A, Rast P (2020) Cultivation and functional characterization of 79 Planctomycetes uncovers their unique biology. Nat Microbiol 5:126–140
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, Whitman WB, Euzeby J, Amann R, Rossello-Mora R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645
Acknowledgements
Open Access funding provided by Projekt DEAL. Part of this research was funded by the Deutsche Forschungsgemeinschaft Grants SU 936/1-1, KA 4967/1-1 and JO 893/4-1, Grant ALWOP.308 of the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO), SIAM (Soehngen Institute for Anaerobic Microbiology) Grant No. 024002002 and the Radboud Excellence fellowship. We thank Ina Schleicher for skillful technical assistance and Sonja Oberbeckmann and Matthias Labrenz (IOW Warnemünde, Germany) for sampling support. The staff of the BCCM/LMG Bacteria collection and the All-Russian Collection of Microorganisms (VKM) we thank for support during strain deposition. We also thank Marc Stadler for his generous support.
Author information
Authors and Affiliations
Contributions
F.S and N.K. wrote the manuscript and analysed the cultivation data, S.W. performed the genomic and phylogenetic analysis, A.H. and M.J. isolated the strain and performed the initial cultivation and strain deposition, S.H.P. and C.B. performed the light microscopic analysis and prepared the LM pictures, M.S.M.J. contributed to text preparation and revised the manuscript, M.R. performed the electron microscopic analysis and prepared the SEM pictures, C.J. took the samples, supervised A.H. and the study. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Surup, F., Wiegand, S., Boedeker, C. et al. Stieleria varia sp. nov., isolated from wood particles in the Baltic Sea, constitutes a novel species in the family Pirellulaceae within the phylum Planctomycetes. Antonie van Leeuwenhoek 113, 1953–1963 (2020). https://doi.org/10.1007/s10482-020-01456-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-020-01456-9