Abstract
Corynebacterium (C.) diphtheriae is one of the two etiological pathogens for human diphtheria with significant morbidity and mortality. Recently, members of its biovar Belfanti have been described as two novel species, C. belfantii and C. rouxii. The most important virulence factor and also the premise to cause diphtheria is the isolate’s capacity to encode and express the diphtheria toxin (DT). In contrast to C. ulcerans, which represents a potentially zoonotic pathogen, C. diphtheriae (incl. the novel deduced species) has almost exclusively been found to comprise a human pathogen. We here report three rare cases of C. rouxii isolation from dogs suffering from disseminated poly-bacterial exsudative to purulent dermatitis and a traumatic labial defect, respectively. The isolates were identified as C. diphtheriae based on commercial biochemistry and matrix-assisted laser desorption/ionisation–time of flight mass spectrometry (MALDI-TOF MS) analysis. However, recently described specific spectral peaks were highly similar to spectra of C. rouxii, which was confirmed by whole genome sequencing. Further investigations of the dog isolates for the presence of DT by tox gene qPCR revealed negative results. The findings from this study point out that skin infections in companion animals can be colonized by uncommon and so believed human specific pathogens, thereby resembling the clinical signs of cutaneous diphtheria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Corynebacterium (C.) diphtheriae is the type species of the genus and the etiological microorganism of human diphtheria. Historically and based on DNA homologies, the species C. diphtheriae, C. ulcerans and C. pseudotuberculosis form the C. diphtheriae group and have the potential to cause human infections (Pascual et al. 1995; Riegel et al. 1995). However, C. pseudotuberculosis is predominantly an animal pathogen, leading to caseous lymphadenitis in small ruminants, horses and camelids (Funke et al. 1997) and C. ulcerans can be isolated from a broad number of animal hosts including wild, zoo and companion animals as well as from livestock (Abbott et al. 2020; Berger et al. 2019, 2011; Eisenberg et al. 2015; Foster et al. 2002; Higgs et al. 1967; Hirai-Yuki et al. 2013; Hommez et al. 1999; Marini et al. 2014; Morris et al. 2005; Olson et al. 1988; Schuhegger et al. 2009; Sykes et al. 2010; Venezia et al. 2012). A close contact with both symptomatically and asymptomatically infected companion animals is the preferred route for human transmission. Interestingly, most human diphtheria cases in Western Europe are caused by C. ulcerans today (Wagner et al. 2010). In the recent past, systematic changes have led to the description of novel species within the C. diphtheriae group. For one of the four biovars of C. diphtheriae, namely biovar (bv.) Belfanti, species status has been proposed as C. belfantii (Dazas et al. 2018). Atypical strains of the same biovar were the subject of another species description, now designated as C. rouxii (Badell et al. 2020). Recently, former C. ulcerans strains isolated from game animals (wild boar and roe deer) (Contzen et al. 2011; Eisenberg et al. 2014; Rau et al. 2012), that were provisionally denominated as “wild boar cluster” of C. ulcerans (Rau et al. 2019), have now been described as C. silvaticum (Dangel et al. 2020). Therefore, the C. diphtheriae group comprises now six species.
However, isolations of former C. diphtheriae bacteria from animals have rarely been found. Although this organism can be readily cultured in murine macrophage cell lines (Weerasekera et al. 2019), reports of natural infections lastly involved a red fox from Germany caused by C. diphtheriae bv. Belfanti (Sing et al. 2016). Furthermore, these authors also give an exhaustive overview of only 11 further similar cases reported during the last century. Besides the single wildlife isolation, infections were otherwise found in companion animals (dogs, cats, horses) and livestock (cattle) with close contact to supposed human shedders.
In contrast to the dermonecrotic exotoxin phospholipase D, a major virulence factor displayed by both C. ulcerans and C. pseudotuberculosis and involved in caseous lymphadenitis, strains of C. diphtheriae, C. ulcerans and C. silvaticum might carry lysogenic β-corynephages which can harbor a tox gene encoded diphtheria toxin (DT), a virulence factor inhibiting protein synthesis (Funke et al. 1997). However, C. belfantii and C. rouxii strains described so far were all non-toxigenic (Badell et al. 2020) or rarely reported as tox gene negative (Dazas et al. 2018). DT contributes to the formation of pseudomembranes in larynx and nasopharynx, which is a hallmark of acute and potentially life-threatening sequelae of diphtheria.
Here we report three cases of C. rouxii isolations from dogs in Germany in order to broaden the knowledge on these rare microorganisms and to include them into the differential of skin lesions in dogs.
Material and methods
Clinical investigation
Case 1
A nearly 6-year old, male, castrated Newfoundlander with a history of disseminated exsudative to purulent dermatitis for few weeks was presented to a local veterinarian. A sample of the external skin of the throat region was submitted for microbiological culture.
Case 2
The second dog was a nearly 2-year old female, spayed mixed breed dog with a history of a defect in the right upper lip of supposed shooting trauma resulting in the inability of unimpaired feeding. The dog also displayed purulent nasal discharge and was presented to the university clinic by a society for the prevention of cruelty to animals. The dog was fed by hand with small balls formed of canned dog food. On examination the middle third of the right upper lip, approximately 60% of the width and two thirds of the length of the hard palate were missing. Correspondingly, the nasal cavity was visible over approximately two thirds of its length. The mucosa of the nasal cavity was inflamed; hypertrophied and purulent discharge was present. The complete blood count and chemistry profile were within normal limits. Computed tomography (CT) revealed an osseous defect of the right maxilla and the left mandible. Furthermore, multiple injured teeth, retained tooth roots, various bullet fragments in the soft tissue and enlarged retropharyngeal and mandibular lymph nodes were detected on CT. Accordingly, a gunshot injury with secondary chronic rhinitis was diagnosed. Injured teeth, the retained tooth roots and the teeth of the caudal part of the right maxilla were removed in order to plan soft tissue reconstruction. A sample of the nasal cavity was submitted for microbiological culture and sensitivity testing.
Case 3
The third dog was an approximately 2-year-old male castrated mixed breed dog from Romania with a known history of atopic dermatitis and Malassezia (M.) pachydermatis infection. It was presented with otitis externa and a purulent dermatitis accompanied with pruritus, thick and wrinkled skin, lichenification, hyperpigmentation and alopecia on the whole body except the forehead. Samples from the affected skin and the right external ear canal were submitted for microbiological examination.
Microbiological culture and identification
Swabs were directly streaked on Columbia agar with 5% sheep blood and Gassner’s agar and cultivated using aerobic and microaerophilic atmosphere conditions for 48 h at 37 °C. A brain heart infusion broth was inoculated and incubated in the same fashion to enrich sub-lethally damaged bacteria and plated after 24 h on the above mentioned media. Yeast and mould growth was investigated using a Sabouraud glucose agar with gentamicin and chloramphenicol at 30 °C. All culture media were provided by Oxoid, Wesel, Germany. Isolates were further evaluated using Gram’s staining and matrix assisted laser desorption/ionisation—time of flight mass spectrometry (MALDI-TOF MS; microflex LT Mass Spectrometer, MALDI Biotyper™; Bruker Daltonik, Bremen, Germany) using the direct smear method in sample preparation. Primary identification was done with the commercial MALDI-Biotyper database (MBT 8468; Bruker Daltonik). Corynebacterium spp. strains were further identified by biochemical differentiation using the API Coryne system (bioMérieux, Nürtingen, Germany).
Antimicrobial sensitivity testing
Antimicrobial sensitivity testing (AST) was carried out using broth microdilution testing. Briefly, a commercially available panel layout for pet animals (Micronaut/Bruker according to guidelines of the research group antimicrobial resistance of the German Veterinary Society DVG) was used. In this layout, 14 different antimicrobials were employed ([ranges given in µg ml−1]; amoxicillin/clavulanic acid [0.031/0.063–8/16], ampicillin [0.125–8], cephalexin [0.5–16], cefovecin [0.25–4], chloramphenicol [1–16], clindamycin [0.031–2], enrofloxacin [0.016–2], erythromycin [0.125–4], gentamicin [0.063–4], pradofloxacin [0.004–1], oxacillin [0.063–2], penicillin [0.063–4], tetracycline [0.5–8] and trimethoprim/sulfamethoxazole [0.25/4.75–2/38]). Resulting MIC values were interpreted as sensitive, resistant and intermediate resistant by clinical breakpoints according to CLSI M100 29th ed. for broth microdilution testing with special reference to Marosevic et al. (2020).
Molecular characterization of Corynebacterium isolates
The presence of the tox gene was investigated by real-time PCR (Schuhegger et al. 2008a). Nearly full-length sequences of the 16S rRNA and rpoB gene sequences were deduced from the draft genome sequences and were further used for blast (Yoon et al. 2017). For phylogenetic analysis phylogenetic trees based on nearly full-length 16S rRNA and rpoB gene sequences were constructed after alignment with muscle with RAxML (Stamatakis 2014) with the Maximum-Likelihood (ML) method based on General Time Reversible (GTR) model with a discrete Gamma-distribution (+ G) and rapid bootstrap analysis. Both trees are based on 1439–1530 (16S) and 3495–3537 (rpoB) nucleotide positions and each 100 replications (bootstrap analysis) (Felsenstein 1985). Isolates were further subjected to tox gene qPCR as previously described (Schuhegger et al. 2008b). Whole genome sequencing (WGS) and an average nucleotide identity (ANI) analysis of the sequencing data was done using Illumina MiSeq sequencing, spades assembly and pyani v0.2.7 using ANIb and ANIm as described earlier (Dangel et al. 2020).
Results
Bacterial cultures
Case 1
After 24 h of incubation, a poly-bacterial growth was noted in the Newfoundlander including Staphylococcus (St.) schleiferi, Streptococcus (S.) canis, Escherichia coli as well as Acinetobacter baumannii. However, small coryneform colonies were also noted on Columbia sheep blood agar that were preliminarily identified as C. diphtheriae (isolate 191012535 = KL1355).
Case 2
The second dog’s microbiological examination revealed growth of St. aureus, St. schleiferi, Pseudomonas aeruginosa, Neisseria animaloris and C. diphtheriae (isolate 45746 = KL 1306).
Case 3
St. pseudintermedius, S. dysgalactiae, C. amycolatum, M. pachydermatis and C. diphtheriae (isolate 1899/20–5 = KL 1663) were cultured from the skin sample of the third dog. The ear sample revealed growth of St. pseudintermedius, S. dysgalactiae, Proteus mirabilis and M. pachydermatis. C. diphtheriae could not be detected in the ear canal.
Phenotypic characterization
All three corynebacteria isolates were confirmed as Gram-positive coryneform rods and further identified as C. diphtheriae by their biochemical patterns (API Coryne code 0010324) showing positive results for glucose and ribose fermentation and for alpha-glucosidase activity and additionally a negative nitrate reaction (Table 1). Using MALDI-TOF MS in combination with the commercial Bruker database (MBT 8468), respective isolates were again identified as C. diphtheriae with scores above 2.00, which allows an identification to species level in the previous taxonomy (Rau et al. 2019). The commercial database included 8 reference spectra of C. diphtheriae and 280 reference spectra of 79 other Corynebacterium species, whereby the currently described species C. belfantii, C. rouxii and C. silvaticum have not yet been included. Consequently, the commercial database was extended by adding custom-made, quality-controlled main spectra projections (MSPs) of the three novel type strains of C. belfantii, C. rouxii and C. silvaticum to the database (Rau et al. 2016, 2019). Further information regarding the isolates under investigation and an exchange option for the user-made MSPs is provided in the MALDI-UP catalogue at https://maldi-up.ua-bw.bwl.de (Rau et al. 2016). The combined database consisting of the commercial Bruker database and the user-made additions for the type strains of C. belfantii, and C. rouxii was used for the species decision via MALDI-TOF MS. With this database the MALDI-TOF MS spectra of the isolates from the three dogs were assigned to C. rouxii with score values higher than 2.50 for the first hit. Subsequently, a dendrogram was created using MSPs of the three dog isolates from this study as well as from one red fox (Sing et al. 2016) and type strains of the C. diphtheriae group (Fig. 1).
Dendrogram created by cluster analysis of reference main spectra (MSP) obtained by MALDI-TOF mass spectrometry (MALDI Biotyper, Version 3.1, Bruker Daltonik) of three Corynebacterium (C.) rouxii isolates from dogs in comparison to a selection of strains from the C. diphtheriae group including spectra of the type strains (T) of C. belfantii, C. diphtheriae, C. rouxii, C. silvaticum, and C. ulcerans. Non-human isolates were indicated with the animal source of isolation. Details of the isolates and reference spectra were given in MALDI-UP (https:maldi-up@ua-bw.bwl.de)
Antimicrobial susceptibility testing
AST revealed the following minimum inhibitory concentrations (MIC; in µg ml−1 in brackets) for respective antimicrobials: amoxicillin/clavulanic acid: (0.125/0.062–0.25/0.125), ampicillin (≤ 0.125–0.25), cephalexin (≤ 0.5–1.0), cefovecin (0.5–1.0), chloramphenicol (≤ 1.0), clindamycin (0.25), enrofloxacin (0.062), erythromycin (≤ 0.125), gentamicin (1.0) and pradofloxacin (0.008–0.016) were found susceptible; resistance was detected for tetracycline (0.125–0.25), oxacillin (1.0–2.0) and trimethoprim/sulfamethoxazole (≤ 0.25/4.75), penicillin was tested intermediate susceptible in case 1 (0.25) and susceptible in cases 2 and 3 (0.125).
Molecular characterization of C. rouxii isolates
Nearly full-length gene sequences were confirmed by 16S rRNA gene sequencing as members of the C. diphtheriae group using the curated EzBioCloud database (Yoon et al. 2017). The highest sequence similarity (99.64%) was shared with the type strain of C. rouxii FRC0190T (Acc. No. MN535983), followed by C. diphtheriae (strain NCTC 11397 T [Acc. No. LN831026]; 99.05%) and C. belfantii (strain FRC0043T [Acc. No. OANN01000138]; 98.93%). 16S rRNA and rpoB gene sequences were further used for phylogenetic analysis together with other type strains of the C. diphtheriae group. The dog isolates from this study clustered in a distinct branch together with C. rouxii in both phylogenies (Suppl. Figs. S1, S2). All three isolates were non-toxigenic as shown by a negative tox qPCR.
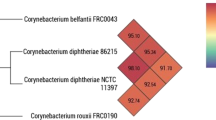
The pairwise ANI comparison of the WGS assemblies from the three dog-derived isolates using PyANI (Pritchard 2019) based on MUMmer (ANIm) and blastn + (ANIb) algorithms for reproducibility revealed sequence identities of 91–92% with NCTC11397T (C. diphtheriae), 92–93% with FRC0043T (C. belfantii), but > 99% with FRC0190T (C. rouxii) and are thus well above the species delineation threshold of 95–96% (Goris et al. 2007; Richter and Rossello-Mora 2009) with highest similarity to C. rouxii as shown in Fig. 2. The ANI analysis with data from Dazas et al. (2018) and Badell et al. (2020) and type strains of further members of the C. diphtheriae group affiliated the three isolates as well with C. rouxii (Suppl. Table S1 A and B).
ANIb comparison of dog derived isolates (KL1306, KL1355, KL1663; marked in bold) with type strains NCTC11397T (C. diphtheriae), FRC0043T (= DSM105776T = KL1687; C. belfantii), FRC0190T (= DSM110354T = KL1688; C. rouxii). Boxes with ANI values ≤ 95% are shown in red, with ≥ 94—< 96% in yellow and > 96% in green. ANI comparison with additional data from further C. belfantii, C. rouxii and additional Corynebacterium spp. isolates is given in Suppl. Table S1
Treatment regimes
Case 1
A topic treatment of the inflamed skin with Pyoderm® shampoo (Virbac Animal Health, Bad Oldesloe, Germany) and Rebohexanid® foam (Alfavet, Neumünster, Germany; twice daily) was administered until AST results were available. The pruritus was treated with oral application of Oclacitinib (Apoquel®; Zoetis, Berlin, Germany; 0.4–0.6 mg kg−1 q 24 h). The antimicrobial treatment involved amoxicillin/clavulanic acid twice daily orally for one week. Following AST results, treatment was changed to clindamycin twice daily (5.5 mg kg−1) orally for another week. The dermatitis resolved completely.
Case 2
Two weeks after healing of the mucosa a surgical attempt was performed to close the defect of upper lip and hard palate. Complete closure of the lip and only partial closure of the hard palate was possible due to an injury related anomaly of the angularis oris artery and vein. The dog was treated with pain medication and amoxicillin/clavulanic acid (12.5 mg kg−1) three-times orally for ten days postoperatively. 20 months postoperatively the dog was able to eat and drink independently, no nasal discharge was present and the cosmetic outcome was good.
Case 3
The infection was treated with cephalexin (28.6 mg kg−1) twice daily orally and ketoconazole (10 mg kg−1) once a day orally over five weeks. Furthermore, the dog was bathed once daily with Maleseb® shampoo (Dechra Veterinary Products A/S, Uldum, Denmark) containing miconazole and chlorhexidine. The pruritus was treated by oral application of Oclacitinib (Apoquel®, Zoetis, Berlin, Germany, 0.4–0.6 mg kg−1) daily. A hypoallergenic diet (Anallergenic®, Royal Canin, Cologne, Germany) was fed for at least eight weeks.
The otitis was treated once a day with Otomax® (MSP Tiergesundheit-Intervet Deutschland GmbH, Unterschleissheim, Germany) containing gentamicin, dexamethason and clotrimazol.
After six weeks the dog was presented with most symptoms in remission but symptoms worsened again later without antibiotic therapy. In two subsequent microbiological examinations (ten and twenty-four weeks after the first microbiological examination) C. diphtheriae could no longer be detected.
With respect to zoonotic potential, responsible medical authorities have been informed after receiving initial results of the microbiological culture and sensitivity testing. We are unaware whether testing of owners was conducted. Some of them could be contacted and instructed to wear gloves during handling and to isolate the dog at home. Because clinical signs improved and some dog owners resided far away from the veterinary clinic, follow-up examinations were not conducted in all the cases.
Discussion
Isolations of bacteria of the C. diphtheriae group and of C. diphtheriae in particular from animals are rarely reported, but need a thorough evaluation under Public Health considerations. This is even more important in the light of recent taxonomical changes that have not yet been incorporated in the standard microbiological tools used in routine diagnostics (e.g. databases in commercial biochemistry and MALDI-TOF MS). Based on phenotypic as well as molecular data, we provide evidence that the isolates from this study and also an isolate from a red fox from a previous report (Sing et al. 2016) were in fact C. rouxii and not C. diphtheriae (Figs. 1, 2, Suppl. Figures 1, 2). This underlines the necessity that all putative C. diphtheriae isolates in animals need a more in depth approach to be identified to species level. Currently, C. rouxii is known from only six isolates that have formerly been assigned to C. diphtheriae bv. Belfanti (Badell et al. 2020). These were isolated from human cutaneous or peritoneum infections, one ascitic fluid and also from one dog with purulent orbital cellulitis. Interestingly, another case report describes the finding of C. diphtheriae bv. Belfanti, isolated from a cow with dermatitis in Switzerland that should be re-evaluated in the light of recent taxonomical changes (Corboz et al. 1996). It remains to be determined whether the negative tox gene is – like in C. belfantii – a constant feature in this species, which might directly influence its role as a human diphtheria pathogen. On the other hand, even severe human infections with non-toxigenic strains must not be underestimated with respect to the pathogenic potential (de Santis et al. 2020; Massmann et al. 2020). With four more animal isolates from three dogs from this study and one fox (Sing et al. 2016) and possibly also from two cat isolates from the USA (Hall et al. 2010) it is tempting to speculate and remains to be determined whether C. rouxii is also a zoonotic pathogen. Unfortunately, animal keepers were not available for bacterial testing in the here presented cases. Close contact between humans and their pets may foster such transmissions principally in both directions, but – unlike to C. ulcerans – the human-to-animal (reverse) zoonosis seems a priori more probable in isolates, formerly known to belong to C. diphtheriae. However, C. rouxii has already been isolated from a woman suffering from osteomyelitis (Hoefer et al. 2021). In accordance with Sing et al. (2016) the characterization of any C. diphtheriae strain isolated from animals – even those sensu lato – is recommended by a reference laboratory as a matter of Public Health concern. For this reason, it is now also important to properly identify isolates of the complete C. diphtheriae species complex (C. diphtheriae [bv. Gravis, Mitis and Intermedius], C. belfantii and C. rouxii). The vast minority of the few animal isolates of former C. diphtheriae has been subjected or is accessible to profound identification.
With respect to phenotypic traits, Badell et al. (2020) point out that the six C. rouxii isolates examined in their study displayed a negative maltose fermentation (utilizing API Coryne) or atypical maltose test results using a Rosco diagnostic method compared to all other C. diphtheriae group species. In contrast the three dog isolates from this study displayed a positive maltose test using Api Coryne.
Optimally and like in other species from the C. diphtheriae group (including former C. diphtheriae, C. ulcerans, C. silvaticum and C. pseudotuberculosis) species differentiation is very well accomplished by MALDI-TOF MS. Presently, the use of commercial databases alone will help to identify the microorganism under study to ‘group-level’, whereas user derived additions to the database have provided great capacity to determine also novel taxa with high accuracy (Dangel et al. 2020; Eisenberg et al. 2014; Rau et al. 2019). Badell et al. 2020 have provided six pairs of C. rouxii-specific biomarkers based on MALDI-TOF MS m/z signals that facilitate the unequivocal differentiation from other C. diphtheriae group species, including C. belfantii. These signal patterns are in good agreement with the signals observed for the isolates from the three dogs and for the isolate from a red fox (Sing et al. 2016) (Suppl. Table. S2). User derived reference spectra have been included in the MALDI user platform (www.maldi-up.ua-bw.de) to propagate non-commercial exchange of well suited, quality approved spectra and to improve clinical and veterinary diagnostics with these rarely recognized microorganisms (Rau et al. 2019).
From a clinical perspective, the skin and dermatitis lesions in the dogs of the present study improved under antimicrobial therapy without further complications. Attempts to re-isolate the corynebacteria failed. A number of different bacterial species were isolated in the depicted cases that might also have caused the presented clinical signs alone or in a poly-bacterial combination. Interestingly, also the red fox suffered from severe subacute phlegmonous inflammation of the subcutaneous tissue and a widespread subacute suppurative inflammation of the mammary gland (Sing et al. 2016). However, from a Public Health perspective, antimicrobial sensitivity patterns for the involved C. rouxiii isolates were assessed using breakpoints for C. diphtheriae in humans. Treatment of Corynebacterium spp. can be challenging since Poor et al. (2017) have documented high MIC levels consistent with supposed antimicrobial resistance in sows, especially for macrolides/lincosamide, tetracyclines and quinolones. Contrarily, the dog isolates from this study were found resistant against trimethoprim/sulfamethoxazole, tetracycline and oxacillin only, which represented a favourable starting point for antimicrobial therapy. Based on therapy attempts in C. ulcerans, Carfora et al. (2018) recommended topical application of erythromycin for two weeks and the systemic administration of cephalexin for three weeks to treat ulcerative lesions in dogs (Carfora et al. 2018).
Conclusion
This is the first case series of canine clinical infections with C. rouxii isolates with focal to disseminated skin diseases. Generally, C. diphtheriae, C. rouxii and C. belfantii are rarely isolated bacteria in animals. Furthermore, the findings from this study point out that such isolates will be missed by using standard diagnostics, and definitely need further evaluation by expert labs. However, purulent dermatitis in companion animals can be colonized by uncommon and so believed human specific pathogens, thereby resembling the clinical signs of cutaneous diphtheria. Because C. rouxii is also a human pathogen, this report broadens the knowledge on this rare microorganism with respect to One Health aspects.
Data availability
All data have been made fully available to the public.
References
Abbott Y, Efstratiou A, Brennan G, Hallanan S, Leggett B, Leonard FC, Markey BK, Tuite C, Fry NK (2020) Toxigenic Corynebacterium ulcerans associated with upper respiratory infections in cats and dogs. J Small Anim Pract 61:554–560. https://doi.org/10.1111/jsap.13185
Badell E, Hennart M, Rodrigues C, Passet V, Dazas M, Panunzi L, Bouchez V, Carmi-Leroy A, Toubiana J, Brisse S (2020) Corynebacterium rouxii sp. nov., a novel member of the diphtheriae species complex. Res Microbiol 171:122–127. https://doi.org/10.1016/j.resmic.2020.02.003
Berger A, Huber I, Merbecks SS, Ehrhard I, Konrad R, Hörmansdorfer S, Hogardt M, Sing A (2011) Toxigenic Corynebacterium ulcerans in woman and cat. Emerg Infect Dis 17:1767–1769. https://doi.org/10.3201/eid1709.110391
Berger A, Dangel A, Peters M, Mühldorfer K, Braune S, Eisenberg T, Szentiks CA, Rau J, Konrad R, Hörmansdorfer S, Ackermann N, Sing A (2019) Tox-positive Corynebacterium ulcerans in hedgehogs, Germany. Emerg Microbes Infect 8:211–217. https://doi.org/10.1080/22221751.2018.1562312
Carfora V, Scarampella F, Iurescia M, Donati V, Stravino F, Lorenzetti S, Menichini E, Franco A, Caprioli A, Battisti A (2018) Non-toxigenic Corynebacterium ulcerans sequence types 325 and 339 isolated from two dogs with ulcerative lesions in Italy. J Veter Diagn Investigat Official Publicat Am Associat Veter Lab Diagn. https://doi.org/10.1177/1040638718764786
Contzen M, Sting R, Blazey B, Rau J (2011) Corynebacterium ulcerans from diseased wild boars. Zoonoses Public Hlth 58:479–488. https://doi.org/10.1111/j.1863-2378.2011.01396.x
Corboz L, Thoma R, Braun U, Zbinden R (1996) Isolation of Corynebacterium diphtheriae subsp. belfanti from a cow with chronic active dermatitis. Schweiz Arch Tierheilkd 138:596–599
Dangel A, Berger A, Rau J, Eisenberg T, Kampfer P, Margos G, Contzen M, Busse HJ, Konrad R, Peters M, Sting R, Sing A (2020) Corynebacterium silvaticum sp. nov., a unique group of NTTB corynebacteria in wild boar and roe deer. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.004195
Dazas M, Badell E, Carmi-Leroy A, Criscuolo A, Brisse S (2018) Taxonomic status of Corynebacterium diphtheriae biovar Belfanti and proposal of Corynebacterium belfantii sp. nov. Int J Syst Evol Microbiol 68:3826–3831. https://doi.org/10.1099/ijsem.0.003069
de Santis A, Siciliano RF, Sampaio RO, Akamine M, Veronese ET, de Almeida Magalhaes FM, Araújo MRE, Rossi F, Magri MMC, Nastri AC, Accorsi TAD, Rosa VEE, Titinger DP, Spina GS, Tarasoutchi F (2020) Non-toxigenic Corynebacterium diphtheriae infective endocarditis with embolic events: a case report. BMC Infect Dis 20:907. https://doi.org/10.1186/s12879-020-05652-w
Eisenberg T, Kutzer P, Peters M, Sing A, Contzen M, Rau J (2014) Nontoxigenic tox-bearing Corynebacterium ulcerans infection among game animals, Germany. Emerg Infect Dis 20:448–452. https://doi.org/10.3201/eid2003.130423
Eisenberg T, Mauder N, Contzen M, Rau J, Ewers C, Schlez K, Althoff G, Schauerte N, Geiger C, Margos G, Konrad R, Sing A (2015) Outbreak with clonally related isolates of Corynebacterium ulcerans in a group of water rats. BMC Microbiol 15:42. https://doi.org/10.1186/s12866-015-0384-x
Felsenstein J (1985) Confidence limits of phylogenies: an approach using the bootstrap. Evolution 39:783–791
Foster G, Patterson T, Howie F, Simpson V, Davison N, Efstratiou A, Lai S (2002) Corynebacterium ulcerans in free-ranging otters. Vet Rec 150:524
Funke G, von Graevenitz A, Clarridge JE 3rd, Bernard KA (1997) Clinical microbiology of coryneform bacteria. Clin Microbiol Rev 10:125–159
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Hall AJ, Cassiday PK, Bernard KA, Bolt F, Steigerwalt AG, Bixler D, Pawloski LC, Whitney AM, Iwaki M, Baldwin A, Dowson CG, Komiya T, Takahashi M, Hinrikson HP, Tondella ML (2010) Novel Corynebacterium diphtheriae in domestic cats. Emerg Infect Dis 16:688–691. https://doi.org/10.3201/eid1604.091107
Higgs TM, Smith A, Cleverly LM, Neave FK (1967) Corynebacterium ulcerans infections in a dairy herd. Vet Rec 81:34–35
Hirai-Yuki A, Komiya T, Suzaki Y, Ami Y, Katsukawa C, Takahashi M, Yamamoto A, Yamada YK (2013) Isolation and characterization of toxigenic Corynebacterium ulcerans from 2 closed colonies of cynomolgus macaques (Macaca fascicularis) in Japan. Comp Med 63:272–278
Hoefer A, Pampaka D, Herrera-León S, Peiró S, Varona S, López-Perea N, Masa-Calles J, Herrera-León L (2021) Molecular and epidemiological characterization of toxigenic and nontoxigenic Corynebacterium diphtheriae, Corynebacterium belfantii, Corynebacterium rouxii, and Corynebacterium ulcerans isolates identified in Spain from 2014 to 2019. J Clin Microbiol. https://doi.org/10.1128/jcm.02410-20
Hommez J, Devriese LA, Vaneechoutte M, Riegel P, Butaye P, Haesebrouck F (1999) Identification of nonlipophilic corynebacteria isolated from dairy cows with mastitis. J Clin Microbiol 37:954–957
Marini RP, Cassiday PK, Venezia J, Shen Z, Buckley EM, Peters Y, Taylor N, Dewhirst FE, Tondella ML, Fox JG (2014) Corynebacterium ulcerans in ferrets. Emerg Infect Dis 20:159–161. https://doi.org/10.3201/eid2001.130675
Marosevic DV, Berger A, Kahlmeter G, Payer SK, Hormansdorfer S, Sing A (2020) Antimicrobial susceptibility of Corynebacterium diphtheriae and Corynebacterium ulcerans in Germany 2011–17. J Antimicrob Chemother 75:2885–2893. https://doi.org/10.1093/jac/dkaa280
Massmann R, Zavadilová J, Drozenová J, Fiksa D, Smíšková D (2020) Septicemia in an immunocompetent adult in the Czech Republic caused by Corynebacterium diphtheriae nontoxigenic strain biotype mitis: emergence of invasive cases in Western Europe. Braz J Infect Dis 24:89–91. https://doi.org/10.1016/j.bjid.2019.12.003
Morris WE, Uzal FA, Cipolla AL (2005) Pyogranulomatous meningoencephalitis in a goat due to Corynebacterium ulcerans. Vet Rec 156:317–318
Olson ME, Goemans I, Bolingbroke D, Lundberg S (1988) Gangrenous dermatitis caused by Corynebacterium ulcerans in Richardson ground squirrels. J Am Vet Med Assoc 193:367–368
Pascual C, Lawson PA, Farrow JA, Gimenez MN, Collins MD (1995) Phylogenetic analysis of the genus Corynebacterium based on 16S rRNA gene sequences. Int J Syst Bacteriol 45:724–728
Poor AP, Moreno LZ, Matajira CEC, Parra BM, Gomes VTM, Silva APS, Dutra MC, Christ APG, Barbosa MRF, Sato MIZ, Moreno AM (2017) Characterization of Corynebacterium diphtheriae, C. confusum and C. amycolatum isolated from sows with genitourinary infection. Vet Microbiol 207:149–152. https://doi.org/10.1016/j.vetmic.2017.06.008
Pritchard L (2019) PyANI. Copyright The James Hutton Institute 2014–2019. Pyani. Python module for Average Nucleotide Identity analyses; [cited 2021]. https://github.com/widdowquinn/pyani.
Rau J, Blazey B, Contzen M, Sting R (2012) Corynebacterium ulcerans infection in roe deer (Capreolus capreolus) [in German]. Berl Munch Tierarztl Wochenschr 125:159–162
Rau J, Eisenberg T, Männig A, Wind C, Lasch P, Sting R (2016) MALDI-UP – An internet platform for the exchange of MALDI-TOF mass spectra. User guide for http://maldi-up.ua-bw.de/. Aspects of food control and animal health (eJournal) 2016:1–17
Rau J, Eisenberg T, Peters M, Berger A, Kutzer P, Lassnig H, Hotzel H, Sing A, Sting R, Contzen M (2019) Reliable differentiation of a non-toxigenic tox gene-bearing Corynebacterium ulcerans variant frequently isolated from game animals using MALDI-TOF MS. Vet Microbiol 237:108399. https://doi.org/10.1016/j.vetmic.2019.108399
Richter M, Rossello-Mora R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Riegel P, Ruimy R, de Briel D, Prevost G, Jehl F, Christen R, Monteil H (1995) Taxonomy of Corynebacterium diphtheriae and related taxa, with recognition of Corynebacterium ulcerans sp. nov. nom. rev. FEMS Microbiol Lett 126:271–276
Schuhegger R, Kugler R, Sing A (2008a) Pitfalls with diphtheria-like illness due to toxigenic Corynebacterium ulcerans. Clin Infect Dis. https://doi.org/10.1086/589575
Schuhegger R, Lindermayer M, Kugler R, Heesemann J, Busch U, Sing A (2008b) Detection of toxigenic Corynebacterium diphtheriae and Corynebacterium ulcerans strains by a novel real-time PCR. J Clin Microbiol 46:2822–2823. https://doi.org/10.1128/jcm.01010-08
Schuhegger R, Schoerner C, Dlugaiczyk J, Lichtenfeld I, Trouillier A, Zeller-Peronnet V, Busch U, Berger A, Kugler R, Hormansdorfer S, Sing A (2009) Pigs as source for toxigenic Corynebacterium ulcerans. Emerg Infect Dis 15:1314–1315. https://doi.org/10.3201/eid1508.081568
Sing A, Konrad R, Meinel DM, Mauder N, Schwabe I, Sting R (2016) Corynebacterium diphtheriae in a free-roaming red fox: case report and historical review on diphtheria in animals. Infection 44:441–445. https://doi.org/10.1007/s15010-015-0846-y
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Sykes JE, Mapes S, Lindsay LL, Samitz E, Byrne BA (2010) Corynebacterium ulcerans bronchopneumonia in a dog. J Vet Intern Med 24:973–976. https://doi.org/10.1111/j.1939-1676.2010.0491.x
Venezia J, Cassiday PK, Marini RP, Shen Z, Buckley EM, Peters Y, Taylor N, Dewhirst FE, Tondella ML, Fox JG (2012) Characterization of Corynebacterium species in macaques. J Med Microbiol 61:1401–1408. https://doi.org/10.1099/jmm.0.045377-0
Wagner KS, White JM, Crowcroft NS, De Martin S, Mann G, Efstratiou A (2010) Diphtheria in the United Kingdom, 1986–2008: the increasing role of Corynebacterium ulcerans. Epidemiol Infect 138:1519–1530. https://doi.org/10.1017/s0950268810001895
Weerasekera D, Fastner T, Lang R, Burkovski A, Ott L (2019) Of mice and men: Interaction of Corynebacterium diphtheriae strains with murine and human phagocytes. Virulence 10:414–428. https://doi.org/10.1080/21505594.2019.1614384
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Acknowledgements
For their excellent assistance and support, we thank Mersiha Curić, Martin Dyk, Anna Eckert, Katharina Engel and Jens Heinbächer. The Hessian State Laboratory is supported by Hessian Ministry for the Environment, Climate Change, Agriculture and Consumer Protection (HMUKLV).
Funding
Open Access funding enabled and organized by Projekt DEAL. This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
KS and TE conceived the study. SD and MK were responsible for dog treatments and sampling. KS, TE, JR, GW, AB, AD, CH, CE and AS performed the microbiological and molecular analyses. KS and TE wrote the first draft and all authors revised and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
There is no ethical issue associated with this manuscript, because the presented data was obtained during routine diagnostics in the dogs. They were not involved in any kind of animal experiment. According to competent authorities this kind of research does not require ethics approval or general approval with respect to German law.
Consent to participate
All authors gave their consent to participate in this study.
Consent for publication
All authors gave their consent to publish results from this study and to be listed as a co-author.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schlez, K., Eisenberg, T., Rau, J. et al. Corynebacterium rouxii, a recently described member of the C. diphtheriae group isolated from three dogs with ulcerative skin lesions. Antonie van Leeuwenhoek 114, 1361–1371 (2021). https://doi.org/10.1007/s10482-021-01605-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01605-8