Abstract
Invasive species can facilitate the spread of pathogens by first providing asymptomatic host reservoirs, and then driving disease outbreaks in native populations through pathogen spillover. An example of this are invasive crayfish species in Europe (Faxonius limosus, Pacifastacus leniusculus, Procambarus clarkii), which carry the deadly plague agent (Aphanomyces astaci). Effective disease management requires comprehensive monitoring, however, pathogen detection in carrier populations with low pathogen prevalence and intensities is challenging. We simultaneously collected and analysed crayfish tissue samples of invasive crayfish populations and water samples to compare A. astaci detection in different sample types using quantitative PCR. Combined, the two sampling methods revealed A. astaci presence with DNA concentrations above limit of detection (LOD; the lowest concentration which can be detected with reasonable certainty) in 13 of 23 invasive crayfish populations. In four additional sites, A. astaci DNA concentrations below LOD were found in water. In four populations only were A. astaci concentrations above LOD detected in both sample types and in three populations in concentrations above LOD in tissue but below LOD in water. The likely reason for these discrepancies is the low A. astaci prevalence and concentration in resistant invasive crayfish, which limit detection reliability. Consistency may be improved by timing surveys with seasonal periods of high A. astaci abundance and by increasing water sampling effort. Considering the ease of collecting eDNA samples, compared to crayfish tissue sampling, eDNA methods would facilitate frequent and comprehensive surveys. However, remaining uncertainties in eDNA-based detection reveal the relevance of combining monitoring tools to improve detection of invasive pathogens and their management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive species can disrupt the structure and functioning of communities and ecosystems, threatening the survival of endangered species (Strayer 2010). In addition to direct effects through competition or predation, co-dispersal of parasites with invasive host species often compounds the harmful effects on local biodiversity, especially when the introduced parasite can also infect resident biota (Dunn and Hatcher 2015). The spread of a species carrying parasites into new territory can lead to novel combinations of parasites and hosts, i.e. to a spillover event, where the parasite acquires a new host species in its invasive range (Strauss et al. 2012). In their introduced range, invasive parasites can mediate the competition between species (Price et al. 1988; Dunn and Hatcher 2015). If the new native host is more susceptible to the parasite than its original, non-native host, the non-native host can acquire a competitive advantage, which promotes its spread and its chances of becoming invasive (Strauss et al. 2012). Invasive species and their parasites can therefore become serious threats to highly susceptible native species, since invasive species can act as an asymptomatic carrier and reservoir species for the parasite. Such reservoir species can be crucial for the persistence of an invasive parasite (Reynolds 1988).
The oomycete Aphanomyces astaci is the causative agent of crayfish plague, the most serious disease threatening European native freshwater crayfish species (Holdich et al. 2009). It is therefore listed among the 100 worst invasive species worldwide (Lowe et al. 2000). Native European crayfish are highly susceptible to the disease, which is transmitted by free-swimming zoospores, and local population extinction has been documented in a matter of weeks after contracting the pathogen (Unestam and Weiss 1970; Alderman et al. 1987). Originating from North America, A. astaci has a long history of co-evolution with North American crayfish species, such as the signal crayfish (Pacifastacus leniusculus), which appear to be asymptomatic carriers of the pathogen in their introduced range in Europe (Holdich et al. 2009). Infection in crayfish usually happens through lesions in the epicuticle, the outermost layer of the exoskeleton (Unestam and Weiss 1970). Once infected, growth of A. astaci hyphae is stopped or slowed by melanisation in P. leniusculus, while the melanisation response of the European noble crayfish (Astacus astacus) is too slow to prevent the parasite from spreading (Nyhlén and Unestam 1980; Cerenius et al. 2003). The widespread presence of invasive North American crayfish species in Europe, acting as disease carrier and reservoir species, poses severe infection risks to native crayfish populations (Holdich et al. 2009; Kouba et al. 2014). Crayfish transported in the ballast water of trans-Atlantic ships are suspected as the source of initial A. astaci invasions (Holdich 2003). The first outbreak was recorded in Italy in 1859 and several outbreaks were observed throughout Europe thereafter (Alderman 1996). The intentional release of P. leniusculus in Sweden, to compensate for the dwindling populations of native A. astacus, further promoted the spread of the crayfish plague (Bohman et al. 2006). Today, invasive North American crayfish and A. astaci are found in most European countries (Kouba et al. 2014).
In Switzerland, three North American crayfish species, P. leniusculus, spiny-cheek (Faxonius limosus) and red swamp crayfish (Procambarus clarkii) have successfully colonised large waterways and lakes, while populations of the native species A. astacus, white-clawed (Austropotamobius pallipes) and stone crayfish (Austropotamobius torrentium) only persist in isolated waterbodies or smaller, hard to reach streams (Stucki and Zaugg 2005). To preserve the remaining populations and allow their recovery, management plans have been devised (Stucki and Zaugg 2011; Elmiger et al. 2018). For the effective implementation of such plans, close surveillance of native and invasive crayfish populations and their disease status is crucial. The advancement of molecular methodologies has enabled the development of fast and reliable A. astaci detection using PCR (Oidtmann et al. 2002, 2004, 2006; Hochwimmer et al. 2009) and quantitative real-time PCR (qPCR) assays (Vrålstad et al. 2009) of infected crayfish tissue. Molecular diagnostics on crayfish tissue samples have therefore become the default testing method for crayfish plague (Kozubíková et al. 2009; Vrålstad et al. 2011; Kokko et al. 2012; Schrimpf et al. 2012). Soft cuticle from the abdomen and the tail fans of invasive crayfish species have shown highest A. astaci detection rates and sampling of both cuticle types increases detection success (Oidtmann et al. 2006; Vrålstad et al. 2011). However, acquisition of crayfish tissue for testing is laborious and costly since crayfish need to be captured in high numbers for reliable results of infection status, especially when A. astaci prevalence in the population is low (Schrimpf et al. 2013). Furthermore, searching and trapping activities can unnecessarily disturb the local habitats and species beyond the target crayfish. Therefore, an alternative method involving the detection of A. astaci in water, based on the detection of environmental DNA (eDNA; Bass et al. 2015; Thomsen and Willerslev 2015), has been developed and experimentally tested with water spiked with A. astaci zoospores (Strand et al. 2011), with ambient water of infected P. leniusculus (Strand et al. 2012), and successfully applied in lakes with P. leniusculus populations carrying A. astaci (Strand et al. 2014). In field surveys, methods for detection of A. astaci in water have performed equally well, or better, in detecting infected sites, than crayfish trapping and tissue extraction methods (Strand et al. 2014; Wittwer et al. 2018, 2019). Being less costly and labour-intensive than conventional trapping methods with subsequent examination of single crayfish individuals, the eDNA-based method can greatly facilitate a regularly updated A. astaci monitoring scheme. However, rigorous validation is required to assess the efficacy and reliability of such an eDNA-based detection method in comparison to more established methods.
In this study we assessed the performance of an eDNA-based method in detecting A. astaci in water and compared it to crayfish trapping combined with molecular detection from crayfish tissue. In addition, we evaluated the capacity of eDNA-based method to detect the crayfish plague agent associated with invasive, asymptomatic crayfish populations (i.e. with low infection intensity levels). To achieve this, we firstly examined the degree of association between A. astaci detection results using both methods. Secondly, we investigated sources of variation in detection for both methods, including host species identity, crayfish size and sex, A. astaci prevalence and number of infected individuals among captured crayfish and agent levels in crayfish. Third, to confirm the functionality of both methods in a situation with high A. astaci concentrations in crayfish and water, we sampled A. astacus and water samples from an acute crayfish plague outbreak site. We then compared detection rates and DNA concentrations with those from asymptomatic invasive crayfish populations. Lastly, we discussed the implications of our findings for crayfish plague monitoring.
Methods
Site selection and crayfish sampling
Sampling sites were chosen using prior knowledge of invasive crayfish species occurrence in the Canton of Zürich (ZH), Switzerland (n = 21 sites, Table 1). Three additional sites were sampled in the Cantons of Aargau (AG), St. Gallen (SG) and Zug (ZG). Sampling sites comprised different types of waterbodies, ranging from small brooks to large rivers and lakes. Sampling was conducted from May to September 2017. Nine sites were inhabited by P. leniusculus, six by F. limosus, six by P. clarkii and a mixed population of P. leniusculus and F. limosus was found at one site (Table 1). One additional sample was taken in September 2018 from the river Glatt, where an ongoing crayfish plague outbreak was discovered in a population of A. astacus, a native European species.
Depending on accessibility and practicability, crayfish were either captured by hand, trapped, or both (Table 1). Hand-capture was conducted at daytime by searching the crayfish underneath stones and in potential burrows. If less than 20 crayfish were caught by hand, five baited traps (Krebskorb Pirat, Engel-Netze GmbH &Co.KG, Bremerhaven, Germany) were distributed at the sampling site and left overnight. The river Töss was trapped for two nights with 10 baited traps each night and crayfish from Lake Greifensee were bought from a local fisherman. The River Limmat near Neuenhof AG was trapped with 10 traps for one night and crayfish in Lake Zugersee were captured as bycatch in nets. The captured crayfish were anaesthetised with clove oil (Ghanawi et al. 2019), and frozen at − 20 °C until tissue extraction.
eDNA sampling
Water samples were taken before handling crayfish. Crayfish and water samples were collected between mid-May and end of September 2017 (Table 1). Environmental DNA samples were collected on the same date as the crayfish at each sampling site, except for sites “Greifensee”, “Limmat Neuenhof” and “Zugersee”. The water sampling procedure is described in detail and visualised in Fig. 1 of Sieber et al. (2020). In short, a portable peristaltic pump (Alexis peristaltic pump, Proactive Environmental Products LLC, Bradenton FL, USA) was used to pump water through a 47 mm diameter glass fibre filter with 1 µm pore size (Grade GF/B, Whatman, VWR, Dietikon, Switzerland). The tubes containing the eDNA filters were transferred on ice before being stored at − 80 °C until DNA extraction. Three 5 L water samples were collected per site, except when filters clogged early due to suspended particles in the water. In that case, up to six water samples, i.e. filters, were collected. At each sampling site, 5 L of clean MilliQ water were first filtered through the filtration equipment as a negative control to verify cleanliness of the equipment. The filtration equipment was changed between each site. Cleaning of used equipment with bleach (2.5% active chlorine) was done after the fieldwork in the laboratory.
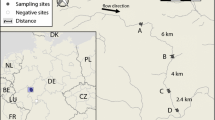
Map of sampling sites and Aphanomyces astaci occurrence (hashed present in crayfish tissue, red present in eDNA, orange present in eDNA below LOD). Labels indicate initials of the crayfish species present at each location (Aa, Astacus astacus; Fl, Faxonius limosus; Pc, Procambarus clarkia; Pl, Pacifastacus leniusculus). Numbers show site identities as listed in Table 1. The grey area of the map indicates borders of Canton Zurich
Crayfish tissue and environmental DNA extraction
Carapace length and sex of each crayfish was determined before tissue sampling. We extracted half of the soft abdominal cuticle and three of the five tail fan tips (uropods) from each crayfish (Fig. S1 in Online Resource 2). The cuticle and uropod samples were stored separately at − 20 °C and DNA was extracted using the DNeasy Blood and Tissue kit (Qiagen AG, Hombrechtikon, Switzerland), following a protocol adapted from Strand et al. (2019) (see Text S1 in Online Resource 1 for a detailed protocol). At the beginning of the project, we also used a CTAB-based and a high salt extraction method on a subset of the sampled crayfish before settling on the DNEasy Blood and Tissue kit. The protocols of all extraction protocols can be found in Online Resource 1 (Texts S1–S3).
A dedicated laboratory used only for processing sensitive samples with low DNA content and for pre-PCR work was used for eDNA extractions. Environmental DNA samples were extracted with the DNEasy Power Water kit (Qiagen AG, Hombrechtikon, Switzerland) as described in Sieber et al. (2020). Extraction runs included a no-template extraction control. The extracted DNA samples were stored at − 20 °C until further analysis.
Real-time quantitative PCR
Both crayfish tissue and eDNA extracts were analysed for A. astaci with real-time quantitative PCR (qPCR) on a LightCycler 480 (Roche, Basel, Switzerland) using the same procedures and protocol. A QIAgility pipetting robot (Qiagen AG, Hombrechtikon, Switzerland) was used for setting up triplicate reactions. For quantification of the samples, a five-fold dilution series consisting of eight dilutions of a double-stranded Gblocks fragment (Integrated DNA Technologies) containing the A. astaci assay target sequence (see Text S4 in Online Resource 1 for sequence information) was included in each qPCR run in triplicate replication. The DNA concentrations ranged from 69,335 to 0.9 copies µl−1. A negative PCR control was included in each qPCR run. The assay developed by Vrålstad et al. (2009) with a modified thermal cycling regime to reduce non-specific amplification according to Strand et al. (2011, 2014) was used. Probe and primer concentrations were optimised for Roche 480 Probes Master Mix in 10 µl reactions. Reactions contained 5 µl of LightCycler 480 Probes Master buffer (Roche, Basel, Switzerland), forward primer AphAstITS-39F at concentration of 50 nM, reverse primer AphAstITS-97R at 900 nM, the MGB probe AphAstITS-60 T (Vrålstad et al. 2009) at 200 nM and 2.5 µl of template DNA. Thermal cycling was initiated by 10 min at 95 °C to activate the DNA polymerase and denature template DNA, followed by 50 cycles of 15 s at 95 °C and 30 s at 62 °C. At the end, a cooling step of 10 s at 40 °C was implemented as suggested by the manufacturer of the thermal cycler. A synthetic template not matching any published sequence data was used as internal positive control to test for PCR inhibition of crayfish tissue and eDNA samples (Carraro et al. 2017). The IPC reactions were setup using methods described in Sieber et al. (2020). The IPC was run separately from the A. astaci assays in triplicate for each crayfish tissue and eDNA sample.
Limit of detection and limit of quantification
The limit of detection (LOD) of a qPCR assay is defined as the lowest DNA concentration, which can still be reliably detected, e.g. with detection rates of 95% or more. The limit of quantification (LOQ) is set at the lowest measured DNA concentration within the linear interval of the standard curve and should indicate the lowest reliably quantifiable DNA concentration (Bustin et al. 2009). To determine the A. astaci assay limit of detection (LOD) and limit of quantification (LOQ) the double-stranded Gblocks fragment (Integrated DNA Technologies) containing the A. astaci assay target sequence was diluted five-fold to create a dilution series with 15 dilutions from concentrations of 54.17 × 107 to 0.09 copies reaction−1. A separate qPCR run with the A. astaci assay was conducted with 30 replicates each of the dilutions 5 to 11, and 40 replicates each of dilutions 12–15 (see Table S1 in Online Resource 2 for detailed results). Detection rates were a 100% in all replicates up to dilution 12. Therefore, the mean cycle value (Cq-value) of positive replicates of dilution 13 with 62.5% detection success was defined as LOD (Cq-value = 38.844; 2.22 copies reaction−1). This is a more permissive LOD than the frequently used 95% detection threshold (Bustin et al. 2009), which we deemed appropriate for low content DNA samples of pathogens. For comparison, we also calculated the LOD of the A. astaci assay at a 95% detection level according to Klymus et al. (2019), which lies at a concentration of 7.76 copies reaction−1. The LOQ was defined as the concentration of the last dilution of the linear range of the standard curve, which was at 11.09 copies reaction−1. The same standard dilution was included in each qPCR run to quantify A. astaci DNA concentrations in samples. The standard curve used to define the LOD and LOQ is visualised in Fig. S2 in Online Resource 2.
Agent levels
To semi-quantitatively categorise A. astaci loads in crayfish tissue, the PCR forming unit (PFU) value of dilution 13 mentioned above was calculated using most probable number (MPN) estimation (Blodgett 2010), i.e. PFU reaction−1 = 2.303 × log10(n × q−1) were n = total number of qPCR replicates and q = number of negative qPCR replicates. Dilution 13 had 15 out of 40 negative replicates, resulting in PFU reaction−1 (dilution 13) = 0.981. This value was used to calculate the PFU values of the remaining dilutions of the series. With the PFU values and mean DNA concentrations for each dilution of the standard curve, the curve equation y = − 1.56 × ln(x) + 39.71 was calculated (x = PFU reaction−1, y = concentration). This equation was used to categorise the crayfish tissue samples into agent levels A0 – A7 according to Vrålstad et al (2009): A0: no detection (n. d.), A1: PFU < 5, A2: 5 ≤ PFU < 50, A3: 50 ≤ PFU < 103, A4: 103 ≤ PFU < 104, A5: 104 ≤ PFU < 105, A6: 105 ≤ PFU < 106, A7: PFU ≥ 106.
Data analysis
The LightCycler 480 Software version 1.5.1 (Roche, Basel, Switzerland) was used to prepare the qPCR raw data as described in Sieber et al. (2020). A crayfish tissue or water sample was considered above LOD if the Cq-value was lower than the LOD in at least one of the three replicate qPCR reactions. Detection of A. astaci was considered successful in crayfish if at least one of the two tissue types were tested positive for A. astaci DNA. The water at a sampling site was considered A. astaci positive if parasite DNA was detected in at least one of the water samples collected at this site. Statistical analyses were conducted in R version 3.6.1. (R Core Team 2019). Parasite detection success was compared between crayfish tissue types and between tissue and water samples using McNemar’s chi-square tests. Main and interaction effects of crayfish species, sex and size (carapace length) on A. astaci detection success in tissue samples were tested with generalised linear mixed effects models (GLMM), and the effects on estimated A. astaci DNA concentrations in tissue samples with linear mixed effects models (LMM), both including sampling site as a random factor. A. astaci DNA concentration estimates from the two tissue types of the same individual (abdominal cuticle and uropod tissue) were compared using a Wilcoxon signed-rank test with continuity correction. The effects of parasite prevalence (number of infected / total number crayfish), maximum parasite agent level and number of infected crayfish on A. astaci detection success in eDNA samples were analysed with GLMMs. Type II Wald chi-square tests were used to test significance of fixed effects. Linear models were used to analyse the correlation between cuticle and uropod tissue types, including species as factor, and to test for effects of parasite prevalence, crayfish agent levels, and mean A. astaci concentrations in crayfish tissue on estimated pathogen eDNA concentrations at the same sampling site. For analyses involving A. astaci prevalence, we only included sites where three or more crayfish were captured. PCR inhibition in DNA samples was quantified using the difference of the IPC’s Cq-values from qPCR reactions containing the DNA extractions and control reactions containing MilliQ water instead of DNA template. We considered ∆ Cq-values ≥ 3 to indicate substantial inhibition. The effect of inhibition on detection success and estimated DNA concentrations was analysed with binomial GLMs and LMs, respectively. Linear models were used to investigate the effect of crayfish size on PCR inhibition. A Kruskal–Wallis rank sum test was used to test for differences of inhibition among invasive crayfish species because the data did not meet parametric assumptions. Normality was tested using Shapiro–Wilk’s test. A Wilcoxon signed rank test with continuity correction was used to compare inhibition between tissue samples of the same crayfish. All tests analysing effects on detection and parasite DNA concentration were conducted once with the complete dataset and once with a subset of samples extracted with the DNEasy Blood and Tissue kit only. This was to ensure that inclusion of samples not extracted with the kit would not drastically change the results. If not stated otherwise, the presented results are from the full dataset.
The R package “eDNAOccupancy” (Dorazio and Erickson 2018) was used for computing hierarchical occupancy models and model selection to test if parasite prevalence in the population, total number of infected crayfish, crayfish agent levels and inhibition scores had an effect on detection probability. Occupancy models were run with 11,000 iterations of the MCMC algorithm. The posterior Predictive Loss (PPLC, Gelfand & Ghosh 1998) and Watanabe-Akaike Information criterions (WAIC, Watanabe 2010) were used for model selection. If adding a covariate or factor did not improve model fit, the covariate was considered not to influence A. astaci detection. The Eq. (1)—(1 − θ)n ≥ 0.95 was used to determine the number of water samples (n) required for successful detection probability of 95%, with θ being the probability of detection of A. astaci DNA in a water sample.
Results
Detection of A. astaci in water and crayfish tissue
Aphanomyces astaci DNA was detected in quantities above LOD in water from five and in tissue samples from twelve out of 23 sites with invasive crayfish (Fig. 1, Table 1). Crayfish tissue sampling was therefore more successful in detecting A. astaci in a population than eDNA sampling (McNemar’s χ2 = 4, df = 1, p = 0.046). A. astaci was detected in both crayfish tissue and water from four sites (Fig. 1). Of the remaining eight sites with detections in tissues, three sites showed a weak signal in water (below LOD) and the other five had no detection in water. In turn, the water samples revealed the presence of A. astaci (above LOD) in one site where it was not detected in crayfish tissue.
Weak, below-LOD signals of A. astaci were obtained from water in additional four sites where none of the crayfish (n = 75) were found infected (Table 1). Six sites were negative for A. astaci in both eDNA and tissue samples. Overall, both eDNA and tissue—based detection revealed 17 A. astaci positive sites, if below LOD detections of A. astaci in water were included. With their inclusion the detection methods agreed on A. astaci presence in seven sites, on its absence in six sites, and five sites each revealed detection in either eDNA or in tissue only.
Hierarchical occupancy models with constant parameters estimated occupancy probability of A. astaci in water per site to be Ψ(·) = 0.314 in invasive crayfish population sites. The estimate of parasite detection per sample was θ(·) = 0.366 and p(·) = 0.726 per qPCR replicate. Thus, to reach detection rates per site of 95% or above, seven water samples would need to be taken. Model fit did not improve much when additional variables were included. If detections of A. astaci concentrations below LOD were considered, the occupancy probabilities increased to Ψ(·) = 0.544, θ(·) = 0.760 and p(·) = 0.940 and only three water samples would need to be taken for detection success to exceed 95%. All tested models are listed in Table S2 in Online Resource 2.
Detection of A. astaci in water samples was more likely with increasing parasite prevalence in the crayfish population (χ2 = 4.042, df = 1, p = 0.044). Furthermore, there was a positive effect of the total number of infected crayfish on eDNA detection success (χ2 = 4.497, df = 1, p = 0.034). Parasite detection rates in water were neither affected by highest parasite agent levels in the crayfish per population (χ2 = 5.417, df = 3, p = 0.144) nor by invasive crayfish species (χ2 = 3.663, df = 2, p = 0.160). Further, we did not observe associations of mean A. astaci concentration in water with parasite prevalence in invasive crayfish (F1,9 = 1.521, p = 0.249), nor with maximum parasite agent levels (F3,8 = 0.753, p = 0.551) or mean A. astaci concentration estimates in crayfish tissues (Fig. 2; abdominal cuticle: F1,6 = 0.985, p = 0.359; uropod: F1,6 = 1.376, p = 0.285).
Mean Aphanomyces astaci concentrations in crayfish tissues (white bars) and/or water (red bars) per sampling site. The bars are labelled with site numbers and names (see Table 1 for more details), followed by abbreviations indicating the crayfish species in brackets (Aa, Astacus astacus; Fl, Faxonius limosus; Pc, Procambarus clarkia; Pl, Pacifastacus leniusculus). Dots show A. astaci concentrations per crayfish and water sample (in crayfish samples black dot = abdominal cuticle, empty = uropod). The active A. astaci outbreak site “Glatt” is distinguishable by the hashed bar pattern
A. astaci detection and concentrations in different types of crayfish tissue
Among the A. astaci positive invasive crayfish (n = 86), the parasite DNA was detected in both tissue types in 44.2% (n = 38) of the crayfish, and in the remaining crayfish, 24 were A. astaci positive in abdominal cuticle tissue and 24 in uropod tissue. The detection success in either tissue type of invasive crayfish was not affected by species (abdominal cuticle: χ2 = 1.331, df = 2, p = 0.214; uropod: χ2 = 1.678, df = 2, p = 0.432), nor sex (abdominal cuticle: χ2 = 0.226, df = 1, p = 0.635; uropod: χ2 = 0.002, df = 1, p = 0.961), nor size (abdominal cuticle: χ2 = 0.021, df = 1, p = 0.886; uropod: χ2 = 0.065, df = 1, p = 0.799) nor any interaction effects. There was a marginally significant interaction effect between sex and size for detection probability in abdominal cuticle samples only (χ2 = 3.917, p = 0.048), which indicated that detection rates were higher for larger females and smaller males.
Estimated A. astaci DNA concentrations in crayfish tissue differed significantly between the two tissue types of the same crayfish, with concentrations in uropod tissue being higher on average (Z = − 2.927, p = 0.003). However, abdominal cuticle and uropod tissue concentrations in the same individual did not correlate significantly (R = 0.126, F1,53 = 0.86, p = 0.358), even when we excluded 42 individuals for which the two types of tissue samples were not extracted with the same method (R = 0.240, F1,44 = 2.695, p = 0.108). Estimated A. astaci DNA concentrations in abdominal cuticle samples of invasive crayfish differed among species (χ2 = 7.656, df = 2, p = 0.022), with P. clarkii showing the highest and P. leniusculus the lowest concentrations on average. The other main effects were not significant (size: χ2 = 0.948, df = 1, p = 0.330; sex: χ2 = 0.989, df = 1, p = 0.320), but there was a significant interaction between species and size (χ2 = 7.942, df = 2, p = 0.019) with F. limosus showing decreasing parasite DNA concentrations with increasing size, while P. leniusculus and P. clarkii demonstrated parasite DNA concentrations slightly increasing with size. Furthermore, a significant interaction between sex and size (χ2 = 5.411, df = 1, p = 0.020) was noted, with parasite DNA concentrations in females slightly increasing, and concentrations in males slightly decreasing with size. Concentrations of A. astaci DNA in uropod tissues of invasive crayfish were not affected by species (χ2 = 2.852, df = 2, p = 0.240), sex (χ2 = 0.072, df = 1, p = 0. 789), or size (χ2 = 0.060, df = 1, p = 0.806) and the variables did not show significant interaction effects.
Native noble crayfish from the single outbreak site showed strong A. astaci signals above LOD and also water samples from this site were clearly positive for the crayfish plague. The diseased A. astacus also showed the highest semi-quantitative A. astaci agent levels we found in this study (A5), while the highest levels in the invasive species were A3 for F. limosus and P. leniusculus and A2 for P. clarkii (Table 1).
PCR inhibition in tissue and eDNA samples
None of the eDNA samples were strongly inhibited (all IPC ∆ Cq < 1) and A. astaci detection success in water samples was not affected by PCR inhibition (χ2 = 1.02, p = 0.313). On the other hand, part of the DNA extractions from crayfish tissue samples (8.3% of cuticle samples, 24.9% of uropod samples) showed substantial signs of inhibition, i.e. IPC ∆Cq > 3, and 9.9% of uropod tissue samples failed to amplify the IPC at all (Fig. 3). Inhibition estimated as ∆ Cq correlated between abdominal cuticle and uropod tissues of the same individuals (F1,389 = 61.410, p < 0.001) and reached higher levels in uropod samples (Z = -2.589, p = 0.01). Furthermore, inhibition in tissue samples of invasive crayfish increased with crayfish size (abdominal cuticle: R = 0.197, F1,426 = 17.213, p < 0.001; uropod: R = 0.369, F1,379 = 42.665, p < 0.001) and differed between species, being highest in P. leniusculus (abdominal cuticle: Kruskal–Wallis χ2 = 106.16, df = 2, p < 0.001; uropod: Kruskal–Wallis χ2 = 103.11, df = 2, p < 0.001).
Inhibition score ∆Cq for three invasive crayfish species (white boxplots) and water (red boxplot) samples. Black dots = abdominal cuticle/water, empty dots = uropod tissue samples. A total of 44 uropod tissue samples failed to amplify the IPC and are visualised as a single dot at IPC ∆Cq-values = 30
Even though the IPC indicated PCR inhibition for part of the DNA extractions obtained from crayfish tissue samples, this inhibition did not affect A. astaci detection success significantly (abdominal cuticle: χ2 = 0.724, df = 1, p = 0.395; uropod: χ2 = 0.313, df = 1, p = 0.576). Accordingly, A. astaci DNA concentrations in tissue samples of invasive crayfish were not affected by inhibition either, neither for abdominal cuticle (F1,79 = 0.524, p = 0.471) nor uropod tissue (F1,80 = 2.971, p = 0.089).
Discussion
We investigated the occurrence of crayfish plague agent Aphanomyces astaci in invasive crayfish populations, using molecular detection methods in crayfish tissue and ambient water samples. Considering both methods, 13 of 23 sampled invasive crayfish populations clearly harboured A. astaci, confirming the disease agent reservoir status in around half of the sampled sites. A. astaci was detected in crayfish tissue in twelve and in water in five of the 23 surveyed invasive crayfish populations (Table 1; Fig. 1), making the tissue sampling method more successful in detecting the parasite than the eDNA method. When weak signals (i.e. below LOD) were considered as true positives, the number of sites with A. astaci detections in water or tissue increased to 17, with detection in water in a total of twelve sites, but even then the parasite was detected by both methods in seven sites only. We argue that while we implemented a LOD in this study, the detections below LOD should not be disregarded categorically, since they could likely indicate low levels of parasite DNA in water. As eDNA samples often contain low starting amounts of target DNA, the frequently applied 95% detection threshold, stemming for guidelines for mainly gene expression assays (Bustin et al. 2009), have been challenged for its suitability for eDNA studies (Hunter et al. 2017; Klymus et al. 2019). In this study a modification of the original assay from Vrålstad et al. (2009) was used to improve specificity by increasing the annealing temperature, which entails a trade-off with sensitivity (Strand et al. 2011, 2014). This could have resulted in low amplification of true positive samples. Nevertheless, we cannot exclude the possibility that the below LOD DNA levels measured in water samples are false positive signals, e.g. due to unspecific amplification, as eDNA methodology is prone to false negative and false positive errors (Griffin et al. 2020). Therefore, target DNA levels below LOD should be treated as inconclusive results. We argue that such results warrant further investigation of pathogen presence at the sampling site and should not be discounted categorically, particularly when applied to a deadly pathogen like the crayfish plague.
The infection intensity in most invasive crayfish in Europe is low, making it challenging to confirm disease agent-free status of a population. The highest agent level observed in tissue of an invasive crayfish in this study was A3 which is comparable to levels found by Vrålstad et al. (2009) but lower than in Vrålstad et al. (2011), Filipová et al. (2013) and Wittwer et al. (2018). Therefore, we suggest that variable detection may be a characteristic of asymptomatic carrier populations, and employment of multiple methods will be required to ensure disease-free status of crayfish populations. In the following sections we further discuss how the variation in detection could arise due to several factors.
DNA of A. astaci was not detected in crayfish tissue in 5 sites where DNA was found in water samples, although only one of these eDNA results was above LOD. These results indicate that the tissue sampling method failed to detect A. astaci in some infected populations, unless the eDNA at these sites originated from unsampled infected populations upstream. One possible explanation is that our sample sizes were insufficient for sites with low prevalence of the parasite. Calculations by Schrimpf et al. (2013) showed that, depending on test sensitivity (detection success rate per individual) and population size, 34 to almost a 1000 crayfish need to be sampled for reliable detection (≥ 95% success rate) in populations with low A. astaci prevalence, i.e. 10% or less infected individuals.
Detection rates in samples from the abdominal cuticle and uropods were equally high in our study, but the overlap was only partial. In 48 of the 86 invasive crayfish positive for A. astaci, the parasite DNA was detected in only one of the two tissue types. Sampling two different tissue types thus more than doubled the detection rate, similar to observations by Oidtmann et al. (2006). Detection rates might be improved further if additional parts of the crayfish cuticle are analysed, e.g. the whole soft abdominal cuticle or walking legs (Vrålstad et al. 2011). Individual level variation in site of infection is to be expected as A. astaci infections mostly occur in spots where the epicuticle, the outermost layer of the exoskeleton, is absent or damaged (Unestam and Weiss 1970). Crayfish that tested positive for both tissue types in this study generally contained higher A. astaci DNA concentrations in the uropods than the abdominal cuticle. Vrålstad et al. (2011) discussed the higher degree of exposure to zoospores and larger total exposure area of the uropods compared to the abdominal cuticle as likely reasons for the observed higher parasite concentrations.
Although infection intensity is potentially influenced by host life-history variation, we found little evidence of it affecting detection of A. astaci in analysed crayfish. For example, there was no general increase in detection rate with crayfish size, even though larger individuals yielded larger uropod tissue samples. Sex and species did not influence parasite detection rate, either. However, a significant interaction term indicated that detection rates increased slightly with size in females but not in males. In contrast, Vrålstad et al. (2011) observed higher A. astaci detection rates in females and large crayfish of both sexes. However, comparisons of studies are difficult as we analysed individuals belonging to different species and originating from multiple populations, while Vrålstad et al. (2011) analysed these patterns in a single large lake population of P. leniusculus, where crayfish are all exposed to similar environmental conditions and infection risks. We did indeed observe significant differences among invasive crayfish species in A. astaci concentrations estimated from abdominal cuticle samples. There were also significant interactions between species and size as well as sex and size, which we find difficult to explain biologically. Given that spore release and infection intensity may be influenced by the molting cycle (Svoboda et al. 2013), it may have mattered that we conducted sampling at only one time point, potentially biasing results through seasonal or moult cycle differences. Juvenile crayfish moult often, adult males and females normally moult once, or less frequently, twice a year, females usually after releasing their young in late summer (Westman and Savolainen 2002). Furthermore, crayfish plague might be more prevalent in crayfish populations during periods of highest activity, such as during the mating season. We collected crayfish throughout the warm season (May–September), which could have influenced A. astaci detection variation.
Crayfish plague detection may also be hampered by PCR inhibition. While eDNA samples did not indicate any relevant levels of inhibition, inhibition was observed in some extractions from crayfish tissues (IPC ∆Cq > 3), especially from uropods. However, the magnitude of IPC inhibition was not associated with parasite detection success. These results suggest that although PCR inhibition may occur in some samples, the qPCR assay employed for A. astaci is robust and not significantly influenced by such effects. Nevertheless, it cannot be excluded that inhibition could have influenced A. astaci concentration estimates from crayfish samples, further emphasizing the importance of sampling multiple tissue samples and individuals.
A. astaci was clearly detected in ambient water in only four of twelve infected invasive crayfish populations (in seven populations if signals below LOD are considered). This contrasts with previous studies which report up to 100% success of eDNA sampling in infected crayfish populations (Strand et al. 2014; Wittwer et al. 2018, 2019). The difference could be related to pathogen prevalence and/or sampling effort. Wittwer et al. (2018, 2019) investigated streams harbouring invasive crayfish populations with infection prevalences of ≥ 60% and took up to 32 eDNA samples per stream, while Strand et al. (2014) collected ten 15 L water samples from lakes containing P. leniusculus with ≥ 50% infection prevalence. A. astaci prevalence in in this study was as low as 4% (Rhein Tössegg), with less than half the populations showing prevalences above 50% (Table 1), and we collected three 5 L water samples per site. The volume of 5 L per water sample was chosen to balance maximal capture of pathogen spores with reasonable filtration time. Indeed, our A. astaci detection success in water increased with higher A. astaci prevalence and the absolute number of infected crayfish and A. astaci was detected in water from three of the four sites harbouring crayfish populations with > 50% infection prevalence. Clearly, the amount of pathogen spores in the water not only depends on prevalence but also on host population density, and it may further be influenced by seasonal variation or by abiotic variables such as flow rate and habitat variability. We were not able to quantify host population density accurately due to the different types of waterbodies surveyed (small streams, large rivers, lakes and ponds) and the different methods of crayfish collection. This may explain the lack of a quantitative association between estimated A. astaci DNA concentrations in water and estimated concentrations/agent levels in crayfish tissues. We did pay attention to collecting water samples when conditions were favorable by avoiding periods of high flow and turbid water after rainfall, but it is of course possible that we did not always sample under optimal conditions. Also, we did not explore any potential effects of habitat variability, which may affect the abundance of A. astaci spores either directly or indirectly via effects on local crayfish population density. The three replicate water samples per site were always taken from the same location. Overall, our results imply that an increased eDNA sampling effort may improve reliability of A. astaci detection in invasive crayfish populations with a low infection prevalence. According to the occupancy modeling results, seven water samples need to be taken per site for a 95% and higher chance of detecting A. astaci concentrations above the LOD in water. However, only three water samples are needed if concentrations below the LOD are considered sufficient for positive detection. Since sampling effort should be reasonable to keep large-scale surveys achievable and cost-effective, it is worth also considering other aspects that could improve reliability. The timing of a survey is a crucial factor for A. astaci detection success in water. In this study, eDNA samples were collected from May to September, which could have contributed to detection variation due to seasonal differences. A. astaci concentrations in water have shown increased levels during crayfish moulting stages in aquaria experiments (Svoboda et al. 2013). Adult crayfish usually moult only once or twice a year, mostly when water temperatures are high during summer (Westin and Gydemo 1986), which indicates our time of sampling was appropriate for increasing A. astaci detection rates. However, Wittwer et al. (2018) took monthly eDNA samples from several sites throughout a year and measured highest A. astaci concentrations in October, coinciding with the mating season when crayfish show increased aggressive behaviour towards each other, which frequently leads to injuries. Due to geographical vicinity of the study system (Germany), we can expect similar seasonal dynamics of A. astaci concentrations in water, which indicate A. astaci eDNA surveys should be conducted later in the year than in this study, i.e. from September to October, to maximise detection success and therefore, reliability of the results. In addition to changes in sampling design, the choice of the extraction method might have an impact on detection reliability as well. The DNEasy Power Water kit used in this study to extract DNA from water samples has been chosen based on its effective inhibitor removal properties but may result in higher loss of DNA than other extraction kits (Eichmiller et al. 2016). Therefore, if the presence of inhibitory compounds is expected to be low, extraction kits with minimal DNA loss could be considered for water samples. Samples that are positively tested for inhibition afterwards could be further diluted or treated with a post-extraction inhibitor removal kit.
All tissue samples of the eight A. astacus indiviudals and the water samples collected at an active crayfish plague outbreak site (Fig. 1) were found positive for A. astaci. The tissue samples had highest parasite concentrations and, therefore, agent levels (A5), of all the collected crayfish in this study, and parasite eDNA concentrations were second highest of all surveyed sites (highest in Riedbach; Table 1; Fig. 2). The same river was sampled downstream from the outbreak site for another survey a month before the outbreak was noticed, and A. astaci eDNA was already found at around a third of the concentrations measured during the outbreak (N. Sieber pers. obs.). While this was one site only, these results suggest that the eDNA method works reliably when parasite loads in a population and therefore in water, are high. The challenges arise from the low amount of parasite spores released by the highly resistant invasive crayfish populations (Strand et al. 2014).
Conclusion
This survey of the crayfish plague agent in asymptomatic invasive crayfish populations showed that two different monitoring methods convey a different picture of A. astaci occurrence. In many cases when crayfish plague infection intensities and prevalence are low, concluding the absence of the plague from a negative result of either method would be misleading. Avenues for optimization of both detection methods are identified. For eDNA-based detection, higher sampling effort would increase detection reliability of asymptomatic crayfish populations. For improved detection of the parasite in crayfish tissue, we suggest analysis of larger numbers of crayfish and more parts of the crayfish cuticle, e.g. the whole abdominal cuticle or leg joints. Both methods would benefit from aligning the time of sampling to seasonal dynamics of the parasite, determined by both host and parasite ecology. Repeated sampling of the same sites during the appropriate season could further improve reliability of the detection result. Decisions on monitoring methods not only depend on reliability of the method, but also on cost and effort, and the ultimate aim of the monitoring and surveillance activity. The effort and cost required for the crayfish tissue sampling method and its suggested improvements is substantially higher than for the eDNA water sampling method. Regular monitoring with the crayfish tissue sampling method alone might therefore not be feasible. Thus, a combination of the two methods would deliver more accurate knowledge of occurrence and spread of the crayfish plague for the implementation of effective management strategies.
Availability of data and materials
The datasets generated and/or analysed during the current study are available on dryad: https://datadryad.org/stash/share/hxp76PQxR7k_dlw6ifW4CRRu8K3Nwfzc_k0sCMLMjWk
References
Alderman DJ (1996) Geographical spread of bacterial and fungal diseases of crustaceans. OIE Rev Sci Tech 15:603–632. https://doi.org/10.20506/rst.15.2.943
Alderman DJ, Polglase JL, Frayling M (1987) Aphanomyces astaci pathogenicity under laboratory and field conditions. J Fish Dis 10:385–393. https://doi.org/10.1111/j.1365-2761.1987.tb01086.x
Bass D, Stentiford GD, Littlewood DTJ, Hartikainen H (2015) Diverse applications of environmental DNA methods in parasitology. Trends Parasitol 31:499–513. https://doi.org/10.1016/j.pt.2015.06.013
Blodgett R (2010) Bacteriological analytical manual (BAM) appendix 2: most probable number from serial dilutions. Food and Drug Administration. https://www.fda.gov/food/laboratory-methods-food/bam-appendix-2-most-probable-number-serial-dilutions. Accessed 24 Jul 2020
Bohman P, Nordwall F, Edsman L (2006) The effect of the large-scale introduction of signal crayfish on the spread of crayfish plague in Sweden. Bull Fr La Pêche La Piscic 380–381:1291–1302. https://doi.org/10.1051/kmae:2006026
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Carraro L, Bertuzzo E, Mari L, Fontes I, Hartikainen H, Strepparava N, Schmidt-Posthaus H, Wahli T, Jokela J, Gatto M, Rinaldo A (2017) Integrated field, laboratory, and theoretical study of PKD spread in a Swiss prealpine river. Proc Natl Acad Sci USA 114:11992–11997. https://doi.org/10.1073/pnas.1713691114
Cerenius L, Bangyeekhun E, Keyser P, Söderhäll I, Söderhäll K (2003) Host prophenoloxidase expression in freshwater crayfish is linked to increased resistance to the crayfish plague fungus, Aphanomyces astaci. Cell Microbiol 5:353–357. https://doi.org/10.1046/j.1462-5822.2003.00282.x
Dorazio RM, Erickson RA (2018) ednaoccupancy: an r package for multiscale occupancy modelling of environmental DNA data. Mol Ecol Resour 18:368–380. https://doi.org/10.1111/1755-0998.12735
Dunn AM, Hatcher MJ (2015) Parasites and biological invasions: parallels, interactions, and control. Trends Parasitol 31:189–199. https://doi.org/10.1016/j.pt.2014.12.003
Eichmiller JJ, Miller LM, Sorensen PW (2016) Optimizing techniques to capture and extract environmental DNA for detection and quantification of fish. Mol Ecol Resour 16:56–68. https://doi.org/10.1111/1755-0998.12421
Elmiger C, Gouskov A, Philipp U, Hertig A (2018) Flusskrebs-Managementplan Kanton Zürich. Pachtperiode 2018–2026, Zürich
Filipová L, Petrusek A, Matasová K, Delaunay C, Grandjean F (2013) Prevalence of the crayfish plague pathogen Aphanomyces astaci in populations of the signal crayfish Pacifastacus leniusculus in France: evaluating the threat to native crayfish. PLoS ONE 8:e70157. https://doi.org/10.1371/journal.pone.0070157
Gelfand AE, Ghosh SK (1998) Model choice: a minimum posterior predictive loss approach. Biometrika 85:1–11. https://doi.org/10.1093/biomet/85.1.1
Ghanawi J, Saoud G, Zakher C, Monzer S, Saoud IP (2019) Clove oil as an anaesthetic for Australian redclaw crayfish Cherax quadricarinatus. Aquac Res 50:3628–3632. https://doi.org/10.1111/are.14319
Griffin JE, Matechou E, Buxton AS, Bormpoudakis D, Griffiths RA (2020) Modelling environmental DNA data; Bayesian variable selection accounting for false positive and false negative errors. J R Stat Soc Ser C Appl Stat 69:377–392. https://doi.org/10.1111/rssc.12390
Hochwimmer G, Tober R, Bibars-Reiter R, Licek E, Steinborn R (2009) Identification of two GH18 chitinase family genes and their use as targets for detection of the crayfish-plague oomycete Aphanomyces astaci. BMC Microbiol 9:1–17. https://doi.org/10.1186/1471-2180-9-184
Holdich DM (2003) Crayfish in Europe—an overview of taxonomy, legislation, distribution, and crayfish plague outbreaks. In: Holdich DM, Sibley PJ (eds) Proceedings of a conference held on 7th November, 2002. Management and Conservation of Crayfish. Bristol, pp 15–34
Holdich DM, Reynolds JD, Souty-Grosset C, Sibley PJ (2009) A review of the ever increasing threat to European crayfish from non-indigenous crayfish species. Knowl Manag Aquat Ecosyst 2009:394–395. https://doi.org/10.1051/kmae/2009025
Hunter ME, Dorazio RM, Butterfield JSS, Meigs-Friend G, Nico LG, Ferrante JA (2017) Detection limits of quantitative and digital PCR assays and their influence in presence—absence surveys of environmental DNA. Mol Ecol Resour 17:221–229. https://doi.org/10.1111/1755-0998.12619
Klymus KE, Merkes CM, Allison MJ, Jackson CA, Goldberg CS, Helbing CC, Hunter ME, Lance RF, Mangan AM, Monroe EM, Piaggio AJ, Stokdyk JP, Wilson CC, Richter CA (2019) Reporting the limits of detection and quantification for environmental DNA assays. Environ DNA 00:1–12. https://doi.org/10.1002/edn3.29
Kokko H, Koistinen L, Harlioǧlu MM, Makkonen J, Aydin H, Jussila J (2012) Recovering Turkish narrow clawed crayfish (Astacus leptodactylus) populations carry Aphanomyces astaci. Knowl Manag Aquat Ecosyst 404:12p1-12p7. https://doi.org/10.1051/kmae/2012006
Kouba A, Petrusek A, Kozák P (2014) Continental-wide distribution of crayfish species in Europe: update and maps. Knowl Manag Aquat Ecosyst 413:05p1-05p31. https://doi.org/10.1051/kmae/2014007
Kozubíková E, Filipová L, Kozák P, Ďuriŝ Z, Martín MP, Diéguez-Uribeondo J, Oidtmann B, Petrusek A (2009) Prevalence of the crayfish plague pathogen Aphanomyces astaci in invasive American crayfishes in the Czech republic. Conserv Biol 23:1204–1213. https://doi.org/10.1111/j.1523-1739.2009.01240.x
Lowe S, Browne M, Boudjelas S, De Poorter M (2000) 100 of the world’s worst invasive alien species. A selection from the global invasive species database. The Invasive Species Specialist Group (ISSG), Auckland
Nyhlén L, Unestam T (1980) Wound reactions and Aphanomyces astaci growth in crayfish cuticle. J Invertebr Pathol 36:187–197. https://doi.org/10.1016/0022-2011(80)90023-3
Oidtmann B, Bausewein S, Hölzle L, Hoffmann R, Wittenbrink M (2002) Identification of the crayfish plague fungus Aphanomyces astaci by polymerase chain reaction and restriction enzyme analysis. Vet Microbiol 85:183–194. https://doi.org/10.1016/S0378-1135(01)00505-3
Oidtmann B, Geiger S, Steinbauer P, Culas A, Hoffmann RW (2006) Detection of Aphanomyces astaci in North American crayfish by polymerase chain reaction. Dis Aquat Org 72:53–64. https://doi.org/10.3354/dao072053
Oidtmann B, Schaefers N, Cerenius L, Söderhäll K, Hoffmann RW (2004) Detection of genomic DNA of the crayfish plague fungus Aphanomyces astaci (Oomycete) in clinical samples by PCR. Vet Microbiol 100:269–282. https://doi.org/10.1016/j.vetmic.2004.01.019
Price PW, Westoby M, Rice B (1988) Parasite-mediated competition: some predictions and tests. Am Nat 131:544–555
R Core Team (2019) R: a language and environment for statistical computing. In: Lang RA (ed) Environ. Stat. Comput. R Found. Stat. Comput. Vienna. https://www.r-project.org/
Reynolds JD (1988) Crayfish extinctions and crayfish plague in central Ireland. Biol Conserv 45:279–285. https://doi.org/10.1016/0006-3207(88)90059-6
Schrimpf A, Maiwald T, Vrålstad T, Schulz HK, Śmietana P, Schulz R (2013) Absence of the crayfish plague pathogen (Aphanomyces astaci) facilitates coexistence of European and American crayfish in central Europe. Freshw Biol 58:1116–1125. https://doi.org/10.1111/fwb.12112
Schrimpf A, Pârvulescu L, Copilaş-Ciocianu D, Petrusek A, Schulz R (2012) Crayfish plague pathogen detected in the Danube Delta—a potential threat to freshwater biodiversity in southeastern Europe. Aquat Invasions 7:503–510. https://doi.org/10.3391/ai.2012.7.4.007
Sieber N, Hartikainen H, Vorburger C (2020) Validation of an eDNA-based method for the detection of wildlife pathogens in water. Dis Aquat Org 141:171–184. https://doi.org/10.3354/dao03524
Strand DA, Holst-Jensen A, Viljugrein H, Edvardsen B, Klaveness D, Jussila J, Vrålstad T (2011) Detection and quantification of the crayfish plague agent in natural waters: direct monitoring approach for aquatic environments. Dis Aquat Org 95:9–17. https://doi.org/10.3354/dao02334
Strand DA, Jussila J, Viljamaa-Dirks S, Kokko H, Makkonen J, Holst-Jensen A, Viljugrein H, Vrålstad T (2012) Monitoring the spore dynamics of Aphanomyces astaci in the ambient water of latent carrier crayfish. Vet Microbiol 160:99–107. https://doi.org/10.1016/j.vetmic.2012.05.008
Strand DA, Jussila J, Johnsen SI, Viljamaa-Dirks S, Edsman L, Wiik-Nielsen J, Viljugrein H, Engdahl F, Vrålstad T (2014) Detection of crayfish plague spores in large freshwater systems. J Appl Ecol 51:544–553. https://doi.org/10.1111/1365-2664.12218
Strand DA, Johnsen SI, Rusch, JC, Agersnap S, Larsein WB, Knudsen SW, M øller PR, Vr ålstad T (2019) Monitoring a Norwegian freshwater crayfish tragedy: eDNA snapshots of invasion, infection and extinction. J Appl Ecol 56:1661–1673. https://doi.org/10.1111/1365-2664.13404
Strauss A, White A, Boots M (2012) Invading with biological weapons: the importance of disease-mediated invasions. Funct Ecol 26:1249–1261. https://doi.org/10.1111/1365-2435.12011
Strayer DL (2010) Alien species in fresh waters: ecological effects, interactions with other stressors, and prospects for the future. Freshw Biol 55:152–174. https://doi.org/10.1111/j.1365-2427.2009.02380.x
Stucki P, Zaugg B (2005) Decapoda. Fauna Helvetica 15. Centre Suisse de Cartographie de la Faune, Neuchâtel
Stucki P, Zaugg B (2011) Aktionsplan Flusskrebse Schweiz. Artenförderung von Edelkrebs, Dohlenkrebs und Steinkrebs, Bern
Svoboda J, Kozubíková-Balcarová E, Kouba A, Buřič M, Kozák P, Diéguez-Uribeondo J, Petrusek A (2013) Temporal dynamics of spore release of the crayfish plague pathogen from its natural host, American spiny-cheek crayfish (Orconectes limosus), evaluated by transmission experiments. Parasitology 140:792–801. https://doi.org/10.1017/S0031182012002223
Thomsen PF, Willerslev E (2015) Environmental DNA—an emerging tool in conservation for monitoring past and present biodiversity. Biol Conserv 183:4–18. https://doi.org/10.1016/j.biocon.2014.11.019
Unestam T, Weiss DW (1970) The host-parasite relationship between freshwater crayfish and the crayfish disease fungus Aphanomyces astaci: Responses to infection by a susceptible and a resistant species. J Gen Microbiol 60:77–90. https://doi.org/10.1099/00221287-60-1-77
Vrålstad T, Johnsen SI, Fristad RF, Edsman L, Strand D (2011) Potent infection reservoir of crayfish plague now permanently established in Norway. Dis Aquat Org 97:75–83. https://doi.org/10.3354/dao02386
Vrålstad T, Knutsen AK, Tengs T, Holst-Jensen A (2009) A quantitative TaqMan® MGB real-time polymerase chain reaction based assay for detection of the causative agent of crayfish plague Aphanomyces astaci. Vet Microbiol 137:146–155. https://doi.org/10.1016/j.vetmic.2008.12.022
Watanabe S (2010) Asymptotic equivalence of Bayes cross validation and widely applicable information criterion in singular learning theory. J Mach Learn Res 11:3571–3594
Westin L, Gydemo R (1986) Influence of light and temperature on reproduction and moulting frequency of the crayfish, Astacus astacus l. Aquac 52:43–50. https://doi.org/10.1016/0044-8486(86)90106-7
Westman K, Savolainen R (2002) Growth of the signal crayfish, Pacifastacus leniusculus, in a small forest lake in Finland. Boreal Environ Res 7:53–61
Wittwer C, Stoll S, Strand D, Vrålstad T, Nowak C, Thines M (2018) eDNA-based crayfish plague monitoring is superior to conventional trap-based assessments in year-round detection probability. Hydrobiologia 807:87–97. https://doi.org/10.1007/s10750-017-3408-8
Wittwer C, Stoll S, Thines M, Nowak C (2019) eDNA-based crayfish plague detection as practical tool for biomonitoring and risk assessment of A. astaci-positive crayfish populations. Biol Invasions 21:1075–1088. https://doi.org/10.1007/s10530-018-1886-x
Acknowledgements
We thank Pascal Reichlin and Alexandre Gouskov for providing crayfish for the study and Andreas Hertig of the cantonal fishing and hunting administrations (Fischerei- und Jagdverwaltung) for the permit for capturing crayfish. We thank the Swiss Federal Institute of Aquatic Science and Technology and the Swiss Federal Office for the Environment for funding. We further thank Bettina Dubach, Pravin Ganesanandamoorthy, Marta Reyes, Dr. Robert Dünner and Jana Jucker for their assistance in the field. Furthermore, we thank Prof. Dr. Jukka Jokela who funded a significant part of the last year of Natalie Sieber’s PhD. Finally, we thank the reviewers for their helpful comments to improve the manuscript.
Funding
Open Access funding provided by Lib4RI – Library for the Research Institutes within the ETH Domain: Eawag, Empa, PSI. This study was funded by Eawag, the Swiss Federal Institute of Technology Zürich (ETH Zürich) and the Swiss Federal Office for the Environment (FOEN).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Natalie Sieber, Hanna Hartikainen and Christoph Vorburger. The first draft of the manuscript was written by Natalie Sieber and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethics approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sieber, N., Hartikainen, H., Krieg, R. et al. Parasite DNA detection in water samples enhances crayfish plague monitoring in asymptomatic invasive populations. Biol Invasions 24, 281–297 (2022). https://doi.org/10.1007/s10530-021-02644-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-021-02644-y