Abstract
Metastatic progression is regulated by metastasis promoter and suppressor genes. NME1, the prototypic and first described metastasis suppressor gene, encodes a nucleoside diphosphate kinase (NDPK) involved in nucleotide metabolism; two related family members, NME2 and NME4, are also reported as metastasis suppressors. These proteins physically interact with members of the GTPase dynamin family, which have key functions in membrane fission and fusion reactions necessary for endocytosis and mitochondrial dynamics. Evidence supports a model in which NDPKs provide GTP to dynamins to maintain a high local GTP concentration for optimal dynamin function. NME1 and NME2 are cytosolic enzymes that provide GTP to dynamins at the plasma membrane, which drive endocytosis, suggesting that these NMEs are necessary to attenuate signaling by receptors on the cell surface. Disruption of NDPK activity in NME-deficient tumors may thus drive metastasis by prolonging signaling. NME4 is a mitochondrial enzyme that interacts with the dynamin OPA1 at the mitochondria inner membrane to drive inner membrane fusion and maintain a fused mitochondrial network. This function is consistent with the current view that mitochondrial fusion inhibits the metastatic potential of tumor cells whereas mitochondrial fission promotes metastasis progression. The roles of NME family members in dynamin-mediated endocytosis and mitochondrial dynamics and the intimate link between these processes and metastasis provide a new framework to understand the metastasis suppressor functions of NME proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 NME genes as suppressors of metastasis
Metastasis is the main cause of death in cancer patients. During metastatic disease, complex factors involving the tumor cells and their microenvironment promote tumor dissemination from the primary site, tumor cell survival in the bloodstream, extravasation and colonization of a secondary site [1]. Breakdown of intercellular adhesion in the tumor epithelium and acquisition of invasive and migratory traits allow epithelial tumor cells to breach the basement membrane and degrade the interstitial – mostly – type I collagen in the stromal extracellular matrix (ECM). These processes, referred collectively to as the epithelial–mesenchymal transition (EMT), are critical events in tumor progression [2, 3]. As in tumorigenesis, metastasis is regulated both positively (by promoters) and negatively (by suppressors). Metastasis suppressor genes – unlike tumor suppressors – inhibit metastasis without affecting growth of the primary tumor [4]. Genetic alterations in these genes – i.e. loss of heterozygosity, spontaneous mutations and polymorphisms – are rare in primary tumors, thus metastasis suppressor genes are probably inactivated by epigenetic mechanisms during metastatic dissemination.
The human genome is estimated to contain ~ 30 genes that encode proteins with metastasis suppressor activity [4,5,6,7,8]. The first example of such a metastasis suppressor gene, NME1 [9], has been characterized extensively and is the subject of numerous reviews [10,11,12,13,14]. The role of the related NME2 gene in the context of metastasis suppression is much less well documented and remains controversial [15,16,17,18]. NME3 has not yet been found to have metastasis suppressor activity, whereas NME4 was described very recently as a new metastasis suppressor gene, the first one in mitochondria [19].
NME genes encode highly conserved multifunctional proteins, some of which are nucleoside diphosphate kinases (NDPKs). NDPKs catalyze the transfer of a γ-phosphate group from nucleoside triphosphates (mainly ATP) to nucleoside diphosphates (mainly GDP) [20,21,22]. These proteins have many important physiological functions in bioenergetics, cytoskeleton and membrane dynamics, signal transduction, metabolism, and development and they are also involved in pathological processes, including metastasis, through mechanisms that are beginning to be deciphered [23,24,25,26]. In mammals, the NME gene family comprises ten members that can be divided into two phylogenetic groups [23, 27]. The group I genes (NME1–NME4) encode proteins that are 58–88% identical to each other and have only one NDPK domain (Table 1). The four NME1 to NME4 proteins are ubiquitous and their catalytic activities have similar kinetics [28]. NME1, NME2 and NME3 are mainly located in the cytosol and at the plasma membrane, whereas NME4 is located exclusively in mitochondria [23, 24, 29, 30]. NME1 and NME2 assemble in vitro and in vivo into stable, catalytically active homohexamers and/or heterohexamers [31, 32]. Formation of such oligomers may contribute to regulating their subcellular localization and/or cellular functions. NME1 and NME2 are the two most abundantly expressed and are thought to be responsible for at least 80% of the cell’s NDPK activity. The phylogenetic group II genes encode more divergent proteins that are only 22–44% identical to each other and to the group I enzymes. Group II NMEs contain one or several full-length or truncated NDPK domains [27]. Despite these domains, none has a demonstrated catalytic activity. In addition, these proteins contain other domains, such as the Dpy-30, DM10, or thioredoxin domains, which may regulate the localization and/or the function of the proteins, possibly by modulating their interactions with various partners. Most group II NME proteins are found only in ciliated cells; the exception is NME6, which is ubiquitous and is located in mitochondria [33].
1.1 NME1, a prototypic metastasis suppressor
Two original discoveries established NME1 as a metastasis suppressor. First, up-regulation of NME1 in several metastatic tumor cell lines from multiple histological types, including melanoma, breast, colon, lung, liver, ovary, prostate, and oral carcinoma cell lines, reduced their metastatic potential both in experimental and spontaneous mouse models of metastasis [34,35,36,37,38,39,40,41,42]. Second, the incidence of lung metastases increased significantly in NME1 knockout mice prone to develop hepatocellular carcinoma [43]. In addition, numerous studies analyzing human patients with melanoma or epithelial tumors of the breast, liver, colon or ovary, found that loss of NME1 expression correlated with a greater risk of metastasis and a poorer clinical prognosis [10, 37, 44,45,46,47,48,49]. By contrast, in neuroblastoma and in hematological malignancies, NME1 overexpression correlated with metastasis dissemination and unfavorable outcome [50,51,52]. These conflicting reports of both positive and negative correlation of NME1 expression with metastasis depending on the tumor type, are highly suggestive of context-specific mechanisms.
Overexpression of NME1 in invasive tumor cell lines that have low levels of endogenous NME1 expression reduces the migratory and invasive potential of the cells. This is observed in cell lines derived from melanoma and from breast, colon, lung, liver, ovarian, prostate and oral carcinoma [42, 53,54,55,56,57,58,59]. By contrast, silencing NME1 in non-invasive tumor cell lines that express high NME1 levels, results in a migratory and invasive phenotype in melanoma, glioblastoma, and carcinoma cells [60,61,62,63,64]. Silencing NME1 in epithelial tumor cells results in an intermediate phenotype with both epithelial and mesenchymal features, refered to as a ‘partial’ EMT [65]. Indeed, NME1-depleted cells express both epithelial and mesenchymal markers, indicating that they do not transition fully to a mesenchymal state. Nevertheless, this ‘partial’ EMT phenotype is more metastatic than the fully mesenchymal state [66, 67], providing an explanation for the high metastatic potential of NME1-deficient tumor cells.
In human cancer samples, there is an inverse correlation between NME1 levels and the expression of well-known EMT markers [65]. Consistent with this, NME1 is downregulated or absent at the invasive front of clinical samples of hepatocellular carcinoma and colon cancers whereas it is robustly present in the center of the tumors [60]. In addition, the ability of NME1 to influence tumor invasion in vivo has been demonstrated in a xenograft model in which human breast tumor cells were injected into the primary duct of the mammary gland in immunodeficient mice [68, 69]. In this model, tumor cells form intraductal tumors that recapitulate the transition from carcinoma in situ to invasive breast carcinoma [68, 70]. Gene-editing inactivation of NME1 in the injected cells accelerated the development of invasive lesions [61].
1.2 NME2, a controversial metastasis suppressor
The anti-metastatic activity of NME2 has been demonstrated mainly in breast, lung and oral squamous carcinoma, and in melanoma [15, 40, 71,72,73]. Moreover, analysis of gene expression datasets from breast, colon, lung and ovarian tumors revealed that there is significantly less NME2 expression in metastatic tumors than in non-metastatic tumors [74].
NME2 has been reported to be an inhibitor of cell motility and invasion [18, 55, 57, 75, 76], suggesting that it might have a similar function to NME1 during tumor progression as an inhibitor of metastasis. In contrast to these earlier studies, however, our work rather suggests that NME2 is not a suppressor of EMT and tumor invasion [60, 61, 65, 77]. Indeed, loss of NME2, unlike NME1, does not impair the invasive capacity of tumor cells or the transition from in situ to invasive breast carcinoma in the intraductal xenograft model [61]. Importantly, inactivation of either NME1 or NME2 expression did not alter the protein level of the other isoform [61], suggesting that there is no compensatory effect when one isoform is deficient.
NME1 and NME2 isoforms are 88% identical in their amino acid sequence, however, sixteen out of eighteen non-identical amino acids are located at the peripheral surface of the hexamer [78]. Thus, NME1 and NME2 homohexamers may interact with different partners and may have distinct cellular functions. Indeed, there is evidence that NME1 and NME2 have more specific partners than partners in common [78]. Furthermore, the relative abundance of NME1 and NME2 proteins and the stoichiometry of NME1–NME2 heterohexamers may provide additional specific regulation and function.
In addition to specific binding partners and regulation of function of NME1 and NME2, the two proteins may function at distinct times in tumorigenesis. We hypothesise that that NME1 and NME2 function as NDPKs at early stages of tumorigenesis, providing nucleoside triphosphates to sustain the high proliferation of tumor cells. At later stages, however, NME1 might become the only isoform controlling EMT, invasion and metastasis. Consistent with this hypothesis, cells induced to proliferate have high levels of NME1 [60, 79,80,81,82,83] and cells overexpressing NME2 proliferate faster than control cells in culture [73]. Accordingly, NME1 and NME2 are overexpressed early in the primary tumor and only NME1 expression is reduced or repressed at later stages in colorectal and hepatic tumors that have undergone metastasis [43, 60, 84]. Additionally, in breast cancer, we found a similar up-regulation of NME1 in ductal carcinoma in situ when compared to the surrounding non-malignant tissues, whereas NME1 levels were significantly reduced in invasive tumor foci and in microinvasive carcinoma buds extending beyond the ruptured basement membrane [61]. By contrast, NME2 was upregulated in ductal carcinoma in situ, but remained highly expressed in invasive tumors supporting the conclusion that NME2 is not a repressor of invasion.
1.3 NME4, a new metastasis suppressor
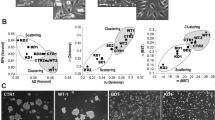
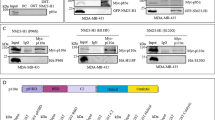
The NME4 gene encodes nucleoside diphosphate kinase D (NDPK-D). Mutations in NME4 that inactivate either the enzymatic activity of NDPK-D or its ability to bind cardiolipin in the mitochondrial inner membrane both induce a strong metastatic phenotype in the cervical carcinoma cell line HeLa and in the breast adenocarcinoma cell line MDA-MB-231 [19], including pronounced cell scattering, loss of intercellular adhesion, increased cell migration in 2D and 3D assays, and increased invasion through a type I collagen matrix. Overexpression of wild-type NDPK-D had the opposite, anti-metastatic effect. Conversely, silencing NME4 in the breast epithelial carcinoma cell line ZR75-1 reduced cell–cell adhesion and increased migration [19]. The metastasis suppressor activity of NME4 was most clearly demonstrated in a metastasis assay in mice in which HeLa cells overexpressing wild-type NDPK-D were injected intravenously [19]. Significantly fewer lung metastases were seen in these mice than in mice injected with HeLa cells overexpressing kinase-dead NDPK-D or expressing a low level of wild-type NDPK-D. These effects were specific to the altered function/expression of mitochondrial NME4 and not due to modified expression of NME1 and NME2 [19].
Low levels of NME4 expression also correlated with high levels of metastasis in various types of cancer in humans. NME4 expression is lower in hepatocarcinoma-derived cell lines with high metastatic potential than it is in those with low metastatic potential [85]. In human cancer, NME4 expression correlates negatively with markers of EMT and tumor aggressiveness [19]. In several cohorts of breast cancer patients, expression of NME4 is negatively associated with markers of mesenchymal cells, the EMT, and tumor invasion, but is positively associated with epithelial markers. In oral cancer, a miRNA that promotes cell migration, invasion and metastasis by inhibiting NME4 expression, miR-196, is highly expressed and correlates with lymph node metastasis [86]. Consistent with the role of NME4 as a metastasis suppressor in human subjects, low expression of NME4 is associated with shorter overall survival (i.e. poor prognosis) in patients with breast tumors or with several other tumor types [19].
1.4 Other NME genes
Among the other members of the NME gene family, there is sparse evidence for their functions as suppressors of metastasis. Human NME3 and NME5 might modulate tumor cell motility depending on the cell context [55, 87], but the mechanism(s) are unclear. Also, in one study, overexpression of NME3 inhibited the metastatic potential of colorectal tumor cells [87]. Further work will be required to pursue the characterization of these and other NME family genes roles in cancer.
2 The dynamin connection
In this section we will review the cellular and molecular mechanisms whereby NME proteins suppress metastasis; in particular, we will detail a mechanism related to NME’s interplay with dynamin GTPases.
Classical dynamins are GTPase motor proteins that are required for endocytosis in all eukaryotic cells [88]. They are responsible for the scission of endocytic vesicles from the plasma membrane during clathrin-mediated endocytosis as well as in some clathrin-independent endocytic pathways [89, 90]. The interaction of NME proteins with dynamins is conserved across species from the nematode Caenorhabditis elegans, to the fruit fly Drosophila melanogaster, mice and humans [26, 91,92,93,94]. NMEs in dynamin-mediated endocytosis might suppress metastasis by inhibiting cell migration and cell invasion, and maintaining cell–cell adhesion, all of which are important for EMT and metastasis processes [3, 95].
2.1 NME/dynamin interplay in cell migration and chemotactism
The first evidence for a functional link between NME proteins and dynamin came from a study in Drosophila [91], that showed that Awd, the counterpart of mammalian NME1 and NME2 facilitated dynamin-mediated neurotransmitter uptake at neuromuscular junctions in the fly. Further studies reported a key implication of Awd function in endocytosis during cell migration [96, 97]. In cooperation with the Drosophila homolog of dynamin, Shibire, Awd inhibits cell migration by promoting endocytosis of chemotactic receptors including the receptors for FGF and PDGF/VEGF, from the surface of migrating tracheal cells during tracheogenesis and of migrating border cells during oogenesis [96,97,98,99,100,101]. Loss of awd in these two cell types decreases endocytosis, leading to up-regulation of the receptors on the cell surface and increasing migration. By contrast, overexpression of awd, increases the endocytosis rate of receptors from the cell surface, so decreasing cell migration. The severity of the awd phenotype is exacerbated in a shibire mutant background whereas overexpression of awd can revert the phenotype associated with a dominant-negative shibire mutation. Likewise in mammalian cells, NME1 mediates endocytosis of the FGF receptor, FGFR1, induced by expression of the von Hippel-Lindau (VHL) protein and prevents cell migration [102]. Interestingly, a loss-of-function mutant in VHL [103] resembles the tracheal phenotype in the awd mutants [96], suggesting that the functional relationship between VHL and NME is evolutionary conserved and is important during development. In addition, mammalian tumor cell lines overexpressing NME1 have increased endocytosis of the EGF receptor and also migrate less than the control cells, and both the increased endocytosis and suppression of migration are blocked by inhibitors of dynamin [57]. In the nematode Caenorhabditis elegans, the NDPK homolog of NME1 and NME2, NDK-1, also influences migration of distal tip cells [104]. Although the underlying mechanism is unknown, the genes encoding C. elegans NDK-1 and dynamin, DYN-1, interact genetically [104]. Additionally, in a genome-wide RNAi screen for genes involved in membrane trafficking, knockdown of NDK-1 caused failure of receptor-mediated endocytosis [105]. Thus, it is possible that NDK-1 regulates the amount of a chemotactic receptor on the surface of the distal tip cells, in the same way as it does in Drosophila and in mammals.
Together, the evidence discussed above indicates that NME proteins facilitate endocytosis of surface receptors and possibly other proteins, altering their availability to transduce migration signals, which, in turn, can suppress cell migration and chemotactism.
2.2 NME/dynamin interplay during cell invasion
Regulation of endocytosis rate by NME proteins may also influence the ability of tumor cells to invade surrounding tissues by clearing membrane-bound proteases from the cell surface. Transmembrane membrane type 1 metalloproteinase (MT1-MMP) is instrumental during cancer progression by mediating proteolytic breaching of tissue barriers, basement membrane and interstitial stromal type I collagen. MT1-MMP is cleared from the cell surface by dynamin-dependent clathrin-mediated endocytosis and is found in clathrin-coated pits associated with dynamin [106,107,108]. Overexpression of NME1 in breast tumor cells was found to increase MT1-MMP endocytosis resulting in removal of the protease from the cell surface, whereas silencing of NME1 decreases MT1-MMP uptake [61]. Collectively, these data indicate that NME1 controls the endocytic clearance and surface exposure of MT1-MMP in human breast cancer. As a consequence, loss of NME1 function enhances both matrix degradation and the invasive potential of breast tumor cells in vitro. At the mechanistic level, MT1-MMP, NME1 and dynamin, interact in clathrin-coated vesicles at the plasma membrane. Loss of NME2, by contrast, has no such effect. Consistent with a role for NME1 in endocytosis of MT1-MMP, in human hepatoma and colorectal tumor cell lines in which NME1 is silenced, expression of a mutant MT1-MMP deleted of its catalytic domain inhibits invasion [60]. Conversely, overexpression of proteolytically active invasion-promoting MT1-MMP increases invasion by cells in which NME1 is silenced. Thus, NME1 enhances endocytosis of MT1-MMP, so suppressing cell invasion.
2.3 NME/dynamin interplay in cell adhesion
Another way by which the function of NME proteins in dynamin-mediated endocytosis may suppress metastasis is by promoting cell adhesion. Evidence for this mechanism, again, came first from studies of the awd mutation in the Drosophila homolog of NME1 and NME2. During oogenesis in mutant awd larvae, adherens junction components, including E-cadherin, β-catenin and α-spectrin, in follicle epithelial cells are mislocalized in the oocyte, disrupting the integrity of the epithelium structure, whereas awd overexpression promotes the turnover of these components by controlling endocytosis [109]. Consistent with this, awd mutations cause developmental defects in the imaginal discs, the sac-like epithelial structures in Drosophila larvae from which legs, antennae and wings develop in the adult fly [110, 111]. A kinase-dead awd mutation failed to rescue the awd mutant phenotype in contrast to the wild-type awd [112], indicating that Awd kinase activity is essential for cell adhesion during Drosophila development.
Consistent with a role for Awd in cell adhesion, dynamin is also necessary to maintain epithelial integrity [113, 114]. In epithelial tissues with a shibire mutation, E-cadherin accumulates in the cytoplasm and adherens junction stability is disrupted, indicating that E-cadherin endocytosis is regulated in epithelial tissues and necessary to maintain epithelium integrity [113,114,115,116]. Similarly, in C. elegans embryos, dynamin-mediated endocytosis is crucial to maintain cell polarity [117]. Moreover, in non-invasive hepatoma and colon tumor epithelial cell lines, silencing NME1 reduces the amount of E-cadherin on the cell surface, correlating with reduced cell–cell adhesion [60]. Also in mammalian epithelial cells, NME1 promotes dynamin-mediated endocytosis of E-cadherin [118].
NME proteins may also regulate cell adhesion to the substratum by modifying the endocytosis of integrin receptors. ICAP-1, an adaptor protein for clathrin-dependent endocytosis of integrins, recruits NME1 and NME2 close to integrins, the clathrin adaptor protein complex, AP2, and dynamin at clathrin-coated pits to ensure integrin turnover at focal adhesions and regulate integrin signaling and cell adhesion [119,120,121].
This abundant evidence that NME proteins are involved in clearing from the cell surface various receptors with key functions in cell–cell adhesion and cell adhesion to the matrix by promoting their dynamin-mediated endocytosis, so inhibiting cell migration and invasion, likely explains the strong metastasis suppressor activity of NME1.
2.4 Control of mitochondrial dynamics by NME family proteins
Additional members of the dynamin superfamily act elsewhere in eukaryotic cells to mediate membrane fission and fusion [122, 123]. Three human mitochondrial dynamin-like proteins, dynamin-related protein 1 (DRP1), optic atrophy protein 1 (OPA1) and mitofusin (MFN), could be implicated in the metastasis suppressor function of mitochondrial NME4 and possibly NME3.
Changes in mitochondria structure and function are potent determinants of EMT and metastasis [124,125,126]. In particular, fragmentation (or fission) of the mitochondrial network facilitates invasion and migration of tumor cells, whereas mitochondrial fusion is rather inhibitory [127]. Generally, metastatic tumor cells express low levels of the fusogenic protein, MFN, as compared to non-metastatic cells, while they express higher levels of the pro-fission protein, DRP1 [128,129,130,131]. In addition, activation of DRP1 [132] or MFN silencing [128] increase the metastatic potential, whereas silencing or pharmacological inhibition of DRP1 or MFN overexpression reduce cell migration and metastasis [128, 129, 133, 134].
The two isoforms of mitofusin, MFN1 and MFN2, are integral membrane proteins of the mitochondrial outer membrane that mediate fusion [122, 123]. NME3 is similarly localized at the outer membrane depending on its N-terminal region [93], and it is known to interact with MFN1 and MFN2 [135]. Silencing of NME3 increases fragmentation of mitochondria [136], suggesting that NME3 enhances outer membrane fusion mediated by mitofusins. Strikingly, expression of NME3 was shown to rescue mitochondrial fusion and elongation in NME3-silenced cells irrespective of its NDPK activity [135]. Further studies will be necessary to elucidate exactly how NME3 functions in mitochondrial fusion and whether this mechanism can contribute to suppress metastasis. In addition, it should be noticed that the NDPK ortholog DYNAMO1 (dynamin-based ring motive-force organizer 1) of mammalian NME3 locally generates GTP for the optimal activity of DRP1 during the division of mitochondria in the red alga, Cyanidioschyson merolae [137]. However, the relevance of these data regarding the role of NME3 in metastasis dissemination is currently unknown.
In addition, the dynamin-like protein, OPA1, mediates the fusion between the inner membranes of mitochondria [122, 123]. NME4, is also located in the intermembrane space and is bound to the mitochondrial inner membrane through cardiolipin, an abundant phospholipid in the inner membrane [138]. Silencing of NME4 alters mitochondrial morphology by producing fragmented, swollen and ‘blebby’ mitochondria reminiscent of those produced upon defective mitochondrial fusion [93]. Depletion of NME4 phenocopies the effect of OPA1 loss-of-function on mitochondria morphology. Moreover, NME4 forms a complex with OPA1 on the mitochondrial inner membrane, facilitated by the binding of both proteins to cardiolipin [138,139,140]. The effect of NME4 binding to OPA1 is not fully understood, however NME4 increases GTP-loading onto OPA1 [93], suggesting that NME4 promotes OPA1-mediated fusion activity. The functions of NME3 and MFN at the outer membrane and NME4 and OPA1 at the inner membrane, respectively, likely contribute to maintaining a fused mitochondrial network important for preventing the EMT and metastasis.
2.5 Molecular mechanisms related to dynamin function displayed by NME family proteins
Members of the dynamin superfamily are evolutionarily conserved membrane-remodeling GTPases involved in both membrane fission and fusion reactions. However, unlike myosin and kinesin motor proteins that use ATP to produce forces, dynamins hydrolyse GTP. In addition, dynamins are very sensitive to intracellular GTP concentration due to their remarkably low affinity for GTP and high intrinsic GTPase activity, resulting in rapid GTP hydrolysis and GDP–GTP exchange, which is enhanced by dynamin oligomerization [90, 93, 141]. Thus, vigourous replenishment of GTP is necessary to sustain cellular activity of dynamin. Unlike for myosins and kinesins, which are fueled by high cytosolic ATP concentration, the lower concentrations of GTP are not sufficient to maintain high rate of GTP loading and GTP hydrolysis by dynamins [142]. Thus, a mechanism of GTP channeling achieved by enzymes that synthesise GTP in close proximity to dynamins is required to secure a high GTP/GDP ratio and favour GTP hydrolysis. The strong NDPK activity of NME proteins [143], together with their high affinity for GDP [144], are ideal to maintain a high local concentration of GTP required for dynamin function. Indeed, several evidence support a model in which NDPKs physically interact with members of the dynamin superfamily to maintain high local GTP concentration for optimal dynamin function in membrane remodeling [94]. NME1 and NME2 fuel cytoplasmic endocytic dynamins at plasma membrane clathrin-coated pits to drive endocytosis (Fig. 1A). In addition, NME1 and NME2 may facilitate the oligomerization of dynamin, which is necessary for membrane fission [57]. Thus, local production and channeling of GTP to endocytic dynamins and stimulation of dynamin oligomerization by NME1 and NME2 contribute to stimulate dynamin’s function in endocytosis. NME4 has the same function on OPA1 by fueling GTP on it at the mitochondria inner membrane to drive inner membrane fusion (Fig. 1B). The molecular mechanism underlying NME3 function on mitofusins must be different to this GTP channeling mechanism as a kinase-dead mutant of NME3 can rescue mitochondrial fusion in NME3-silenced cells. One possibility is that NME3 recruits cytosolic NDPKs, NME1 and/or NME2, to the mitochondrial surface by forming hetero-oligomers. In this scenario, NME1/NME2 recruited by NME3 would channel GTP to mitofusins to promote outer membrane fusion (Fig. 1C). If the molecular mechanisms of action of NME proteins on dynamin superfamily proteins remain to be defined precisely, convergence of subcellular localisations of NME proteins and their dynamin superfamily counterparts, acting as a team is emerging as a new, interesting perspective to explain the antimetastatic activity of NME proteins (Table 1).
Mechanisms of antimetastatic action of group I NME family members. A. The cytosolic NDPKs NME1 and NME2 are recruited to the plasma membrane by their physical interaction with dynamin 2. They generate GTP locally from GDP and ATP and channel GTP to dynamin 2 to optimise dynamin’s activity, which is necessary for fission and endocytosis. The resulting removal of pro-migratory/pro-invasive factors such as MT1-MMP, PDGFR, FGFR and EGFR from the cell surface and turnover of adherens junction proteins such as E-cadherin explain the antimetastatic effects of NME1 and NME2. B. The mitochondrial NDPK NME4 binds the dynamin-related protein OPA1 at the mitochondrial inner membrane to provide GTP for OPA1, which permits mitochondrial inner membrane fusion. This fusion process inhibits metastasis, thus explaining the antimetastatic activity of NME4. C. The localization of NME3 at the mitochondrial surface, where the dynamin-related pro-fusion proteins mitofusins act, suggests that this NDPK might assist mitofusins during mitochondrial outer membrane fusion. NME3 might recruit the cytosolic NDPKs NME1 and NME2 to the mitochondrial surface to produce GTP for mitofusins and promote mitochondrial outer membrane fusion. This fusion process also inhibits metastasis, thus this mechanism may explain the antimetastatic effect of NME3. PDGFR: platelet-derived growth factor receptor, FGFR: fibroblast growth factor receptor, EGFR: epidermal growth factor receptor, MOM: mitochondrial outer membrane, IMS: intermembrane space, MIM: mitochondrial inner membrane, ANT: adenylate translocase, OXPHOS: oxidative phosphorylation, MFN: mitofusins
3 Cytoskeleton regulation by NME family proteins
Additionally mechanisms may explain how NME1 and, in some cases, NME2 can potently suppress cell migration and metastasis. On the one hand, NME proteins inhibit Rho GTPase signaling, by sequestering nucleotide exchange factors necessary for Rho activation in the control of cell cytoskeleton dynamics. Indeed, NME1 sequesters Tiam1 and Dbl-1, the nucleotide exchange factors for Rac1 and cdc42, respectively [145,146,147], whereas NME2 sequesters Lbc, the nucleotide exchange factor for RhoA [148]. The independent identifications of three nucleotide exchange factors as NME-interacting proteins suggest that the Rho GTPase signaling may be biologically relevant. On the other hand, NME1 may suppress metastasis, by reducing transcription of the EDG2 gene, which is involved in metastasis and connected with Rho–ROCK regulation of cell motility [38, 63]. Third, the anti-metastatic function of NME1 may be mediated partially by its ability to inhibit the activity of the actin-severing protein gelsolin [149]. A proteomics study identified NME1 as a binding partner of gelsolin [149]. NME1 inhibited the actin depolymerizing activity of gelsolin, antagonized gelsolin-stimulated tumor cell migration in vitro, and attenuated its pro-metastatic activity in an in vivo model of breast tumor metastasis. Fourth, NME1 is reported to inhibit cell migration by phosphorylating the light chain of the cytoplasmic motor protein myosin [150]. Finally, accumulating evidence suggests that NME1, and often also NME2, interact with and affect the functions of various components and regulators of the cytoskeleton, including acting-binding proteins, intermediate filaments and attachment sites for the cytoskeleton (adherens junctions, desmosomes, and focal adhesions) in cells from a variety of organisms and tissues and during the course of development, suggesting this association is conserved through evolution and may serve an essential function [26].
4 Conclusion
Over the last three decades, extensive analyses of the NME/NDPK family revealed the multifaceted roles of these conserved proteins in cellular pathophysiology and uncovered the underlying molecular mechanisms. The role of the NME/NDPK family in membrane remodeling and nucleotide channeling has become widely recognized as an essential feature of the mechanism of action of several NME family members, including NME1, NME2, NME4, and potentially NME3. Here, the classical NDPK model as a main source of GTP is extended to direct fueling of GTP to GTP-dependent dynamin family proteins through protein/protein interaction. This conclusion is supported by experimental evidence obtained in different species and model systems, including the fruit fly D. melanogaster, nematode C. elegans, mouse and human, indicating an evolutionary conserved mechanism of membrane remodeling controlled by dynamin-NME/NDPK protein interplay. GTP fueling to cytosolic dynamins (through cytosolic NME1 and NME2), which promotes endocytosis of cell surface receptors and cargoes with key function in cell–cell adhesion and cell adhesion to the matrix with, direct impact on cell migration and invasion, likely explains the strong metastasis suppressor potential of NME1 and its alternatively, NME2. GTP fueling to mitochondrial dynamin-related OPA1 (by mitochondrial NME4) promotes mitochondrial inner membrane fusion, a process inhibitory to migration and invasion of tumor cells. The localization of NME3 at the outer mitochondrial membrane, where the fusogenic dynamin-like protein, mitofusin, is recruited to, mediates mitochondrial outer membrane fusion, suggesting that NME3 might likewise participates in outer membrane dynamics, although through a mechanism different of GTP channeling. These emerging roles of NME family members in dynamin-mediated endocytosis and mitochondrial dynamics provide a new framework to explain the antimetastatic activities of NME proteins, which may open new routes to novel therapeutics.
References
Steeg, P. S. (2006). Tumor metastasis: Mechanistic insights and clinical challenges. Nature Medicine, 12(8), 895–904. https://doi.org/10.1038/nm1469
Lu, W., & Kang, Y. (2019). Epithelial-Mesenchymal Plasticity in Cancer Progression and Metastasis. Developmental Cell, 49(3), 361–374. https://doi.org/10.1016/j.devcel.2019.04.010
Brabletz, S., Schuhwerk, H., Brabletz, T., & Stemmler, M. P. (2021). Dynamic EMT: a multi-tool for tumor progression. The EMBO Journal, 40(18), e108647. https://doi.org/10.15252/embj.2021108647
Steeg, P. S., Ouatas, T., Halverson, D., Palmieri, D., & Salerno, M. (2003). Metastasis suppressor genes: Basic biology and potential clinical use. Clinical Breast Cancer, 4(1), 51–62. https://doi.org/10.3816/cbc.2003.n.012
Steeg, P. S. (2003). Metastasis suppressors alter the signal transduction of cancer cells. Nature Reviews Cancer, 3(1), 55–63. https://doi.org/10.1038/nrc967
Rinker-Schaeffer, C. W., O’Keefe, J. P., Welch, D. R., & Theodorescu, D. (2006). Metastasis suppressor proteins: Discovery, molecular mechanisms, and clinical application. Clinical Cancer Research, 12(13), 3882–3889. https://doi.org/10.1158/1078-0432.CCR-06-1014
Smith, S. C., & Theodorescu, D. (2009). Learning therapeutic lessons from metastasis suppressor proteins. Nature Reviews Cancer, 9(4), 253–264. https://doi.org/10.1038/nrc2594
Khan, I., & Steeg, P. S. (2018). Metastasis suppressors: Functional pathways. Laboratory Investigation, 98(2), 198–210. https://doi.org/10.1038/labinvest.2017.104
Steeg, P. S., Bevilacqua, G., Kopper, L., Thorgeirsson, U. P., Talmadge, J. E., Liotta, L. A., et al. (1988). Evidence for a novel gene associated with low tumor metastatic potential. Journal of the National Cancer Institute, 80(3), 200–204. https://doi.org/10.1093/jnci/80.3.200
Hartsough, M. T., & Steeg, P. S. (2000). Nm23/nucleoside diphosphate kinase in human cancers. Journal of Bioenergetics and Biomembranes, 32(3), 301–308. https://doi.org/10.1023/a:1005597231776
Ouatas, T., Salerno, M., Palmieri, D., & Steeg, P. S. (2003). Basic and translational advances in cancer metastasis: Nm23. Journal of Bioenergetics and Biomembranes, 35(1), 73–79. https://doi.org/10.1023/a:1023497924277
Steeg, P. S., Horak, C. E., & Miller, K. D. (2008). Clinical-translational approaches to the Nm23-H1 metastasis suppressor. Clinical Cancer Research, 14(16), 5006–5012. https://doi.org/10.1158/1078-0432.CCR-08-0238
Marshall, J. C., Collins, J., Marino, N., & Steeg, P. (2010). The Nm23-H1 metastasis suppressor as a translational target. European Journal of Cancer, 46(7), 1278–1282. https://doi.org/10.1016/j.ejca.2010.02.042
Matyasi, B., Farkas, Z., Kopper, L., Sebestyen, A., Boissan, M., Mehta, A., et al. (2020). The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis. Pathology Oncology Research, 26(1), 49–61. https://doi.org/10.1007/s12253-020-00797-0
Thakur, R. K., Yadav, V. K., Kumar, A., Singh, A., Pal, K., Hoeppner, L., et al. (2014). Non-metastatic 2 (NME2)-mediated suppression of lung cancer metastasis involves transcriptional regulation of key cell adhesion factor vinculin. Nucleic Acids Research, 42(18), 11589–11600. https://doi.org/10.1093/nar/gku860
Li, Y., Tong, Y., & Wong, Y. H. (2015). Regulatory functions of Nm23-H2 in tumorigenesis: Insights from biochemical to clinical perspectives. Naunyn-Schmiedeberg’s Archives of Pharmacology, 388(2), 243–256. https://doi.org/10.1007/s00210-014-1066-1
Tong, Y., Yung, L. Y., & Wong, Y. H. (2015). Metastasis suppressors Nm23H1 and Nm23H2 differentially regulate neoplastic transformation and tumorigenesis. Cancer Letters, 361(2), 207–217. https://doi.org/10.1016/j.canlet.2015.02.050
Liu, Y. F., Yang, A., Liu, W., Wang, C., Wang, M., Zhang, L., et al. (2015). NME2 reduces proliferation, migration and invasion of gastric cancer cells to limit metastasis. PLoS One, 10(2), e0115968. https://doi.org/10.1371/journal.pone.0115968
Lacombe, M. L., Lamarche, F., De Wever, O., Padilla-Benavides, T., Carlson, A., Khan, I., et al. (2021). The mitochondrially-localized nucleoside diphosphate kinase D (NME4) is a novel metastasis suppressor. BMC Biology, 19(1), 228. https://doi.org/10.1186/s12915-021-01155-5
Cleland, W. W. (1963). The kinetics of enzyme-catalyzed reactions with two or more substrates or products. I. Nomenclature and rate equations. Biochimica et Biophysica Acta, 67, 104–137. https://doi.org/10.1016/0006-3002(63)91800-6
Norman, A. W., Wedding, R. T., & Black, M. K. (1965). Detection of phosphohistidine in nucleoside diphosphokinase isolated from Jerusalem artichoke mitochondria. Biochemical and Biophysical Research Communications, 20(6), 703–709. https://doi.org/10.1016/0006-291x(65)90073-2
Lascu, I., & Gonin, P. (2000). The catalytic mechanism of nucleoside diphosphate kinases. Journal of Bioenergetics and Biomembranes, 32(3), 237–246. https://doi.org/10.1023/a:1005532912212
Lacombe, M. L., Milon, L., Munier, A., Mehus, J. G., & Lambeth, D. O. (2000). The human Nm23/nucleoside diphosphate kinases. Journal of Bioenergetics and Biomembranes, 32(3), 247–258. https://doi.org/10.1023/a:1005584929050
Boissan, M., Dabernat, S., Peuchant, E., Schlattner, U., Lascu, I., & Lacombe, M. L. (2009). The mammalian Nm23/NDPK family: From metastasis control to cilia movement. Molecular and Cellular Biochemistry, 329(1–2), 51–62. https://doi.org/10.1007/s11010-009-0120-7
Boissan, M., & Lacombe, M. L. (2011). Learning about the functions of NME/NM23: Lessons from knockout mice to silencing strategies. Naunyn-Schmiedeberg’s Archives of Pharmacology, 384(4–5), 421–431. https://doi.org/10.1007/s00210-011-0649-3
Boissan, M., Schlattner, U., & Lacombe, M. L. (2018). The NDPK/NME superfamily: State of the art. Laboratory Investigation, 98(2), 164–174. https://doi.org/10.1038/labinvest.2017.137
Desvignes, T., Pontarotti, P., Fauvel, C., & Bobe, J. (2009). Nme protein family evolutionary history, a vertebrate perspective. BMC Evolutionary Biology, 9, 256. https://doi.org/10.1186/1471-2148-9-256
Gonin, P., Xu, Y., Milon, L., Dabernat, S., Morr, M., Kumar, R., et al. (1999). Catalytic mechanism of nucleoside diphosphate kinase investigated using nucleotide analogues, viscosity effects, and X-ray crystallography. Biochemistry, 38(22), 7265–7272. https://doi.org/10.1021/bi982990v
Milon, L., Rousseau-Merck, M. F., Munier, A., Erent, M., Lascu, I., Capeau, J., et al. (1997). nm23-H4, a new member of the family of human nm23/nucleoside diphosphate kinase genes localised on chromosome 16p13. Human Genetics, 99(4), 550–557. https://doi.org/10.1007/s004390050405
Milon, L., Meyer, P., Chiadmi, M., Munier, A., Johansson, M., Karlsson, A., et al. (2000). The human nm23-H4 gene product is a mitochondrial nucleoside diphosphate kinase. Journal of Biological Chemistry, 275(19), 14264–14272. https://doi.org/10.1074/jbc.275.19.14264
Gilles, A. M., Presecan, E., Vonica, A., & Lascu, I. (1991). Nucleoside diphosphate kinase from human erythrocytes. Structural characterization of the two polypeptide chains responsible for heterogeneity of the hexameric enzyme. Journal of Biological Chemistry, 266(14), 8784–8789.
Urano, T., Takamiya, K., Furukawa, K., & Shiku, H. (1992). Molecular cloning and functional expression of the second mouse nm23/NDP kinase gene, nm23-M2. FEBS Letters, 309(3), 358–362. https://doi.org/10.1016/0014-5793(92)80807-s
Proust, B., Radic, M., Vidacek, N. S., Cottet, C., Attia, S., Lamarche, F., et al. (2021). NME6 is a phosphotransfer-inactive, monomeric NME/NDPK family member and functions in complexes at the interface of mitochondrial inner membrane and matrix. Cell & Bioscience, 11(1), 195. https://doi.org/10.1186/s13578-021-00707-0
Leone, A., Flatow, U., King, C. R., Sandeen, M. A., Margulies, I. M., Liotta, L. A., et al. (1991). Reduced tumor incidence, metastatic potential, and cytokine responsiveness of nm23-transfected melanoma cells. Cell, 65(1), 25–35. https://doi.org/10.1016/0092-8674(91)90404-m
Leone, A., Flatow, U., VanHoutte, K., & Steeg, P. S. (1993). Transfection of human nm23-H1 into the human MDA-MB-435 breast carcinoma cell line: Effects on tumor metastatic potential, colonization and enzymatic activity. Oncogene, 8(9), 2325–2333.
Palmieri, D., Horak, C. E., Lee, J. H., Halverson, D. O., & Steeg, P. S. (2006). Translational approaches using metastasis suppressor genes. Journal of Bioenergetics and Biomembranes, 38(3–4), 151–161. https://doi.org/10.1007/s10863-006-9039-9
Boissan, M., & Lacombe, M. L. (2006). Nm23/NDP kinases in hepatocellular carcinoma. Journal of Bioenergetics and Biomembranes, 38(3–4), 169–175. https://doi.org/10.1007/s10863-006-9031-4
Horak, C. E., Mendoza, A., Vega-Valle, E., Albaugh, M., Graff-Cherry, C., McDermott, W. G., et al. (2007). Nm23-H1 suppresses metastasis by inhibiting expression of the lysophosphatidic acid receptor EDG2. Cancer Research, 67(24), 11751–11759. https://doi.org/10.1158/0008-5472.CAN-07-3175
Zhang, Q., McCorkle, J. R., Novak, M., Yang, M., & Kaetzel, D. M. (2011). Metastasis suppressor function of NM23-H1 requires its 3’-5’ exonuclease activity. International Journal of Cancer, 128(1), 40–50. https://doi.org/10.1002/ijc.25307
Baba, H., Urano, T., Okada, K., Furukawa, K., Nakayama, E., Tanaka, H., et al. (1995). Two isotypes of murine nm23/nucleoside diphosphate kinase, nm23-M1 and nm23-M2, are involved in metastatic suppression of a murine melanoma line. Cancer Research, 55(9), 1977–1981.
Parhar, R. S., Shi, Y., Zou, M., Farid, N. R., Ernst, P., & Al-Sedairy, S. T. (1995). Effects of cytokine-mediated modulation of nm23 expression on the invasion and metastatic behavior of B16F10 melanoma cells. International Journal of Cancer, 60(2), 204–210. https://doi.org/10.1002/ijc.2910600213
Fan, Y., Yao, Y., Li, L., Wu, Z., Xu, F., Hou, M., et al. (2013). nm23-H1 gene driven by hTERT promoter induces inhibition of invasive phenotype and metastasis of lung cancer xenograft in mice. Thorac Cancer, 4(1), 41–52. https://doi.org/10.1111/j.1759-7714.2012.00140.x
Boissan, M., Wendum, D., Arnaud-Dabernat, S., Munier, A., Debray, M., Lascu, I., et al. (2005). Increased lung metastasis in transgenic NM23-Null/SV40 mice with hepatocellular carcinoma. Journal of the National Cancer Institute, 97(11), 836–845. https://doi.org/10.1093/jnci/dji143
Bevilacqua, G., Sobel, M. E., Liotta, L. A., & Steeg, P. S. (1989). Association of low nm23 RNA levels in human primary infiltrating ductal breast carcinomas with lymph node involvement and other histopathological indicators of high metastatic potential. Cancer Research, 49(18), 5185–5190.
Florenes, V. A., Aamdal, S., Myklebost, O., Maelandsmo, G. M., Bruland, O. S., & Fodstad, O. (1992). Levels of nm23 messenger RNA in metastatic malignant melanomas: Inverse correlation to disease progression. Cancer Research, 52(21), 6088–6091.
Xerri, L., Grob, J. J., Battyani, Z., Gouvernet, J., Hassoun, J., & Bonerandi, J. J. (1994). NM23 expression in metastasis of malignant melanoma is a predictive prognostic parameter correlated with survival. British Journal of Cancer, 70(6), 1224–1228. https://doi.org/10.1038/bjc.1994.477
An, R., Meng, J., Shi, Q., Dai, X. X., Chen, J. H., Lei, Y. J., et al. (2010). Expressions of nucleoside diphosphate kinase (nm23) in tumor tissues are related with metastasis and length of survival of patients with hepatocellular carcinoma. Biomedical and Environmental Sciences, 23(4), 267–272. https://doi.org/10.1016/S0895-3988(10)60062-1
Liu, L., Li, M., Zhang, C., Zhang, J., Li, G., Zhang, Z., et al. (2018). Prognostic value and clinicopathologic significance of nm23 in various cancers: A systematic review and meta-analysis. International Journal of Surgery, 60, 257–265. https://doi.org/10.1016/j.ijsu.2018.10.035
Leonard, M. K., McCorkle, J. R., Snyder, D. E., Novak, M., Zhang, Q., Shetty, A. C., et al. (2018). Identification of a gene expression signature associated with the metastasis suppressor function of NME1: Prognostic value in human melanoma. Laboratory Investigation, 98(3), 327–338. https://doi.org/10.1038/labinvest.2017.108
Chang, C. L., Zhu, X. X., Thoraval, D. H., Ungar, D., Rawwas, J., Hora, N., et al. (1994). Nm23-H1 mutation in neuroblastoma. Nature, 370(6488), 335–336. https://doi.org/10.1038/370335a0
Niitsu, N., Nakamine, H., & Okamoto, M. (2011). Expression of nm23-H1 is associated with poor prognosis in peripheral T-cell lymphoma, not otherwise specified. Clinical Cancer Research, 17(9), 2893–2899. https://doi.org/10.1158/1078-0432.CCR-10-2999
Almgren, M. A., Henriksson, K. C., Fujimoto, J., & Chang, C. L. (2004). Nucleoside diphosphate kinase A/nm23-H1 promotes metastasis of NB69-derived human neuroblastoma. Molecular Cancer Research, 2(7), 387–394.
Khan, M. H., Yasuda, M., Higashino, F., Haque, S., Kohgo, T., Nakamura, M., et al. (2001). nm23-H1 suppresses invasion of oral squamous cell carcinoma-derived cell lines without modifying matrix metalloproteinase-2 and matrix metalloproteinase-9 expression. American Journal of Pathology, 158(5), 1785–1791. https://doi.org/10.1016/S0002-9440(10)64134-X
Pennino, F. P., Murakami, M., Zollo, M., & Robertson, E. S. (2021). The metastasis suppressor protein NM23-H1 modulates the PI3K-AKT axis through interaction with the p110alpha catalytic subunit. Oncogenesis, 10(4), 34. https://doi.org/10.1038/s41389-021-00326-x
McDermott, W. G., Boissan, M., Lacombe, M. L., Steeg, P. S., & Horak, C. E. (2008). Nm23-H1 homologs suppress tumor cell motility and anchorage independent growth. Clinical & Experimental Metastasis, 25(2), 131–138. https://doi.org/10.1007/s10585-007-9128-0
Leonard, M. K., Novak, M., Snyder, D., Snow, G., Pamidimukkala, N., McCorkle, J. R., et al. (2019). The metastasis suppressor NME1 inhibits melanoma cell motility via direct transcriptional induction of the integrin beta-3 gene. Experimental Cell Research, 374(1), 85–93. https://doi.org/10.1016/j.yexcr.2018.11.010
Khan, I., Gril, B., & Steeg, P. S. (2019). Metastasis Suppressors NME1 and NME2 Promote Dynamin 2 Oligomerization and Regulate Tumor Cell Endocytosis, Motility, and Metastasis. Cancer Research, 79(18), 4689–4702. https://doi.org/10.1158/0008-5472.CAN-19-0492
She, S., Xu, B., He, M., Lan, X., & Wang, Q. (2010). Nm23-H1 suppresses hepatocarcinoma cell adhesion and migration on fibronectin by modulating glycosylation of integrin beta1. Journal of Experimental & Clinical Cancer Research, 29, 93. https://doi.org/10.1186/1756-9966-29-93
Khan, I., & Steeg, P. S. (2018). The relationship of NM23 (NME) metastasis suppressor histidine phosphorylation to its nucleoside diphosphate kinase, histidine protein kinase and motility suppression activities. Oncotarget, 9(12), 10185–10202. https://doi.org/10.18632/oncotarget.23796
Boissan, M., De Wever, O., Lizarraga, F., Wendum, D., Poincloux, R., Chignard, N., et al. (2010). Implication of metastasis suppressor NM23-H1 in maintaining adherens junctions and limiting the invasive potential of human cancer cells. Cancer Research, 70(19), 7710–7722. https://doi.org/10.1158/0008-5472.CAN-10-1887
Lodillinsky, C., Fuhrmann, L., Irondelle, M., Pylypenko, O., Li, X. Y., Bonsang-Kitzis, H., et al. (2021). Metastasis-suppressor NME1 controls the invasive switch of breast cancer by regulating MT1-MMP surface clearance. Oncogene, 40(23), 4019–4032. https://doi.org/10.1038/s41388-021-01826-1
Zhao, R., Gong, L., Li, L., Guo, L., Zhu, D., Wu, Z., et al. (2013). nm23-H1 is a negative regulator of TGF-beta1-dependent induction of epithelial-mesenchymal transition. Experimental Cell Research, 319(5), 740–749. https://doi.org/10.1016/j.yexcr.2012.10.013
Horak, C. E., Lee, J. H., Elkahloun, A. G., Boissan, M., Dumont, S., Maga, T. K., et al. (2007). Nm23-H1 suppresses tumor cell motility by down-regulating the lysophosphatidic acid receptor EDG2. Cancer Research, 67(15), 7238–7246. https://doi.org/10.1158/0008-5472.CAN-07-0962
Tanaka, M., Kuriyama, S., & Aiba, N. (2012). Nm23-H1 regulates contact inhibition of locomotion, which is affected by ephrin-B1. Journal of Cell Science, 125(Pt 18), 4343–4353. https://doi.org/10.1242/jcs.104083
Huna, A., Nawrocki-Raby, B., Padilla-Benavides, T., Gavard, J., Coscoy, S., Bernard, D., et al. (2021). Loss of the Metastasis Suppressor NME1, But Not of Its Highly Related Isoform NME2, Induces a Hybrid Epithelial-Mesenchymal State in Cancer Cells. International Journal of Molecular Sciences, 22(7). https://doi.org/10.3390/ijms22073718.
Pastushenko, I., Brisebarre, A., Sifrim, A., Fioramonti, M., Revenco, T., Boumahdi, S., et al. (2018). Identification of the tumour transition states occurring during EMT. Nature, 556(7702), 463–468. https://doi.org/10.1038/s41586-018-0040-3
Pastushenko, I., & Blanpain, C. (2019). EMT Transition States during Tumor Progression and Metastasis. Trends in Cell Biology, 29(3), 212–226. https://doi.org/10.1016/j.tcb.2018.12.001
Behbod, F., Kittrell, F. S., LaMarca, H., Edwards, D., Kerbawy, S., Heestand, J. C., et al. (2009). An intraductal human-in-mouse transplantation model mimics the subtypes of ductal carcinoma in situ. Breast Cancer Research, 11(5), R66. https://doi.org/10.1186/bcr2358
Lodillinsky, C., Infante, E., Guichard, A., Chaligne, R., Fuhrmann, L., Cyrta, J., et al. (2016). p63/MT1-MMP axis is required for in situ to invasive transition in basal-like breast cancer. Oncogene, 35(3), 344–357. https://doi.org/10.1038/onc.2015.87
Miller, F. R., Santner, S. J., Tait, L., & Dawson, P. J. (2000). MCF10DCIS.com xenograft model of human comedo ductal carcinoma in situ. JNCI: Journal of the National Cancer Institute, 92(14), 1185–1186. https://doi.org/10.1093/jnci/92.14.1185a
Bhujwalla, Z. M., Aboagye, E. O., Gillies, R. J., Chacko, V. P., Mendola, C. E., & Backer, J. M. (1999). Nm23-transfected MDA-MB-435 human breast carcinoma cells form tumors with altered phospholipid metabolism and pH: A 31P nuclear magnetic resonance study in vivo and in vitro. Magnetic Resonance in Medicine, 41(5), 897–903. https://doi.org/10.1002/(sici)1522-2594(199905)41:5<897::aid-mrm7>3.0.co;2-t
Fukuda, M., Ishii, A., Yasutomo, Y., Shimada, N., Ishikawa, N., Hanai, N., et al. (1996). Decreased expression of nucleoside diphosphate kinase alpha isoform, an nm23-H2 gene homolog, is associated with metastatic potential of rat mammary-adenocarcinoma cells. International Journal of Cancer, 65(4), 531–537. https://doi.org/10.1002/(SICI)1097-0215(19960208)65:4%3c531::AID-IJC23%3e3.0.CO;2-B
Miyazaki, H., Fukuda, M., Ishijima, Y., Takagi, Y., Iimura, T., Negishi, A., et al. (1999). Overexpression of nm23-H2/NDP kinase B in a human oral squamous cell carcinoma cell line results in reduced metastasis, differentiated phenotype in the metastatic site, and growth factor-independent proliferative activity in culture. Clinical Cancer Research, 5(12), 4301–4307.
Thakur, R. K., Yadav, V. K., Kumar, P., & Chowdhury, S. (2011). Mechanisms of non-metastatic 2 (NME2)-mediated control of metastasis across tumor types. Naunyn-Schmiedeberg’s Archives of Pharmacology, 384(4–5), 397–406. https://doi.org/10.1007/s00210-011-0631-0
Jiao, G. J., Zhang, S. J., Li, Y., Wu, W. L., & Liu, H. C. (2018). MicroRNA-645 promotes metastasis of osteosarcoma via targeting tumor suppressor NM23 nucleoside diphosphate kinase 2. Clinical and Experimental Pharmacology and Physiology, 45(12), 1317–1324. https://doi.org/10.1111/1440-1681.13006
Polanski, R., Maguire, M., Nield, P. C., Jenkins, R. E., Park, B. K., Krawczynska, K., et al. (2011). MDM2 interacts with NME2 (non-metastatic cells 2, protein) and suppresses the ability of NME2 to negatively regulate cell motility. Carcinogenesis, 32(8), 1133–1142. https://doi.org/10.1093/carcin/bgr070
Nagle, I., Richert, A., Quinteros, M., Janel, S., Buysschaert, E., Luciani, N., et al. (2022). Surface tension of model tissues during malignant transformation and epithelial-mesenchymal transition. Frontiers in Cell and Developmental Biology, 10, 926322. https://doi.org/10.3389/fcell.2022.926322
Marino, N., Marshall, J. C., & Steeg, P. S. (2011). Protein-protein interactions: A mechanism regulating the anti-metastatic properties of Nm23-H1. Naunyn-Schmiedeberg’s Archives of Pharmacology, 384(4–5), 351–362. https://doi.org/10.1007/s00210-011-0646-6
Caligo, M. A., Cipollini, G., Fiore, L., Calvo, S., Basolo, F., Collecchi, P., et al. (1995). NM23 gene expression correlates with cell growth rate and S-phase. International Journal of Cancer, 60(6), 837–842. https://doi.org/10.1002/ijc.2910600619
Keim, D., Hailat, N., Melhem, R., Zhu, X. X., Lascu, I., Veron, M., et al. (1992). Proliferation-related expression of p19/nm23 nucleoside diphosphate kinase. The Journal of Clinical Investigation, 89(3), 919–924. https://doi.org/10.1172/JCI115672
Sorscher, S. M., Steeg, P., Feramisco, J. R., Buckmaster, C., Boss, G. R., & Meinkoth, J. (1993). Microinjection of an nm23 specific antibody inhibits cell division in rat embryo fibroblasts. Biochemical and Biophysical Research Communications, 195(1), 336–345. https://doi.org/10.1006/bbrc.1993.2049
Cipollini, G., Berti, A., Fiore, L., Rainaldi, G., Basolo, F., & Merlo, G., et al. (1997). Down-regulation of the nm23.h1 gene inhibits cell proliferation. International Journal of Cancer, 73(2), 297–302. https://doi.org/10.1002/(sici)1097-0215(19971009)73:2<297::aid-ijc22>3.0.co;2-b.
Lee, H., Okada, K., Baba, H., Furukawa, K., Chang, S., & Shiku, H. (1997). Up-regulation of nm23/NDP kinase expression in regenerating hepatocytes. International Journal of Oncology, 11(5), 965–970. https://doi.org/10.3892/ijo.11.5.965
Martinez, J. A., Prevot, S., Nordlinger, B., Nguyen, T. M., Lacarriere, Y., Munier, A., et al. (1995). Overexpression of nm23-H1 and nm23-H2 genes in colorectal carcinomas and loss of nm23-H1 expression in advanced tumour stages. Gut, 37(5), 712–720. https://doi.org/10.1136/gut.37.5.712
Xu, Z. Y., Chen, J. S., & Shu, Y. Q. (2010). Gene expression profile towards the prediction of patient survival of gastric cancer. Biomedicine & Pharmacotherapy, 64(2), 133–139. https://doi.org/10.1016/j.biopha.2009.06.021
Lu, Y. C., Chang, J. T., Liao, C. T., Kang, C. J., Huang, S. F., Chen, I. H., et al. (2014). OncomiR-196 promotes an invasive phenotype in oral cancer through the NME4-JNK-TIMP1-MMP signaling pathway. Molecular Cancer, 13, 218. https://doi.org/10.1186/1476-4598-13-218
Qu, L., Liang, L., Su, J., & Yang, Z. (2013). Inhibitory effect of upregulated DR-nm23 expression on invasion and metastasis in colorectal cancer. European Journal of Cancer Prevention, 22(6), 512–522. https://doi.org/10.1097/CEJ.0b013e328361625d
Ferguson, S. M., & De Camilli, P. (2012). Dynamin, a membrane-remodelling GTPase. Nature Reviews Molecular Cell Biology, 13(2), 75–88. https://doi.org/10.1038/nrm3266
Roux, A. (2014). Reaching a consensus on the mechanism of dynamin? F1000Prime Reports, 6, 86. https://doi.org/10.12703/P6-86.
Antonny, B., Burd, C., De Camilli, P., Chen, E., Daumke, O., & Faelber, K., et al. (2016). Membrane fission by dynamin: what we know and what we need to know. The EMBO Journal, 35(21), 2270–2284. https://doi.org/10.15252/embj.201694613.
Krishnan, K. S., Rikhy, R., Rao, S., Shivalkar, M., Mosko, M., Narayanan, R., et al. (2001). Nucleoside diphosphate kinase, a source of GTP, is required for dynamin-dependent synaptic vesicle recycling. Neuron, 30(1), 197–210. https://doi.org/10.1016/s0896-6273(01)00273-2
Narayanan, R., & Ramaswami, M. (2003). Regulation of dynamin by nucleoside diphosphate kinase. Journal of Bioenergetics and Biomembranes, 35(1), 49–55. https://doi.org/10.1023/a:1023441806530
Boissan, M., Montagnac, G., Shen, Q., Griparic, L., Guitton, J., Romao, M., et al. (2014). Membrane trafficking. Nucleoside diphosphate kinases fuel dynamin superfamily proteins with GTP for membrane remodeling. Science, 344(6191), 1510–1515. https://doi.org/10.1126/science.1253768
Zala, D., Schlattner, U., Desvignes, T., Bobe, J., Roux, A., & Chavrier, P., et al. (2017). The advantage of channeling nucleotides for very processive functions. F1000Res, 6, 724. https://doi.org/10.12688/f1000research.11561.2.
Lamouille, S., Xu, J., & Derynck, R. (2014). Molecular mechanisms of epithelial-mesenchymal transition. Nature Reviews Molecular Cell Biology, 15(3), 178–196. https://doi.org/10.1038/nrm3758
Dammai, V., Adryan, B., Lavenburg, K. R., & Hsu, T. (2003). Drosophila awd, the homolog of human nm23, regulates FGF receptor levels and functions synergistically with shi/dynamin during tracheal development. Genes & Development, 17(22), 2812–2824. https://doi.org/10.1101/gad.1096903
Nallamothu, G., Woolworth, J. A., Dammai, V., & Hsu, T. (2008). Awd, the homolog of metastasis suppressor gene Nm23, regulates Drosophila epithelial cell invasion. Molecular and Cellular Biology, 28(6), 1964–1973. https://doi.org/10.1128/MCB.01743-07
Lee, T., Hacohen, N., Krasnow, M., & Montell, D. J. (1996). Regulated Breathless receptor tyrosine kinase activity required to pattern cell migration and branching in the Drosophila tracheal system. Genes & Development, 10(22), 2912–2921. https://doi.org/10.1101/gad.10.22.2912
Sutherland, D., Samakovlis, C., & Krasnow, M. A. (1996). branchless encodes a Drosophila FGF homolog that controls tracheal cell migration and the pattern of branching. Cell, 87(6), 1091–1101. https://doi.org/10.1016/s0092-8674(00)81803-6
Duchek, P., Somogyi, K., Jekely, G., Beccari, S., & Rorth, P. (2001). Guidance of cell migration by the Drosophila PDGF/VEGF receptor. Cell, 107(1), 17–26. https://doi.org/10.1016/s0092-8674(01)00502-5
McDonald, J. A., Pinheiro, E. M., & Montell, D. J. (2003). PVF1, a PDGF/VEGF homolog, is sufficient to guide border cells and interacts genetically with Taiman. Development, 130(15), 3469–3478. https://doi.org/10.1242/dev.00574
Hsu, T., Adereth, Y., Kose, N., & Dammai, V. (2006). Endocytic function of von Hippel-Lindau tumor suppressor protein regulates surface localization of fibroblast growth factor receptor 1 and cell motility. Journal of Biological Chemistry, 281(17), 12069–12080. https://doi.org/10.1074/jbc.M511621200
Adryan, B., Decker, H. J., Papas, T. S., & Hsu, T. (2000). Tracheal development and the von Hippel-Lindau tumor suppressor homolog in Drosophila. Oncogene, 19(24), 2803–2811. https://doi.org/10.1038/sj.onc.1203611
Fancsalszky, L., Monostori, E., Farkas, Z., Pourkarimi, E., Masoudi, N., Hargitai, B., et al. (2014). NDK-1, the homolog of NM23-H1/H2 regulates cell migration and apoptotic engulfment in C. elegans. PLoS One, 9(3), e92687. https://doi.org/10.1371/journal.pone.0092687
Balklava, Z., Pant, S., Fares, H., & Grant, B. D. (2007). Genome-wide analysis identifies a general requirement for polarity proteins in endocytic traffic. Nature Cell Biology, 9(9), 1066–1073. https://doi.org/10.1038/ncb1627
Uekita, T., Itoh, Y., Yana, I., Ohno, H., & Seiki, M. (2001). Cytoplasmic tail-dependent internalization of membrane-type 1 matrix metalloproteinase is important for its invasion-promoting activity. Journal of Cell Biology, 155(7), 1345–1356. https://doi.org/10.1083/jcb.200108112
Jiang, A., Lehti, K., Wang, X., Weiss, S. J., Keski-Oja, J., & Pei, D. (2001). Regulation of membrane-type matrix metalloproteinase 1 activity by dynamin-mediated endocytosis. Proceedings of the National Academy of Sciences U S A, 98(24), 13693–13698. https://doi.org/10.1073/pnas.241293698
Li, X. Y., Ota, I., Yana, I., Sabeh, F., & Weiss, S. J. (2008). Molecular dissection of the structural machinery underlying the tissue-invasive activity of membrane type-1 matrix metalloproteinase. Molecular Biology of the Cell, 19(8), 3221–3233. https://doi.org/10.1091/mbc.E08-01-0016
Woolworth, J. A., Nallamothu, G., & Hsu, T. (2009). The Drosophila metastasis suppressor gene Nm23 homolog, awd, regulates epithelial integrity during oogenesis. Molecular and Cellular Biology, 29(17), 4679–4690. https://doi.org/10.1128/MCB.00297-09
Dearolf, C. R., Hersperger, E., & Shearn, A. (1988). Developmental consequences of awdb3, a cell-autonomous lethal mutation of Drosophila induced by hybrid dysgenesis. Developmental Biology, 129(1), 159–168. https://doi.org/10.1016/0012-1606(88)90170-4
Dearolf, C. R., Tripoulas, N., Biggs, J., & Shearn, A. (1988). Molecular consequences of awdb3, a cell-autonomous lethal mutation of Drosophila induced by hybrid dysgenesis. Developmental Biology, 129(1), 169–178. https://doi.org/10.1016/0012-1606(88)90171-6
Xu, J., Liu, L. Z., Deng, X. F., Timmons, L., Hersperger, E., Steeg, P. S., et al. (1996). The Enzymatic Activity of Drosophila AWD/NDP Kinase Is Necessary but Not Sufficient for Its Biological Function. Developmental Biology, 177(2), 544–557.
Leibfried, A., Fricke, R., Morgan, M. J., Bogdan, S., & Bellaiche, Y. (2008). Drosophila Cip4 and WASp define a branch of the Cdc42-Par6-aPKC pathway regulating E-cadherin endocytosis. Current Biology, 18(21), 1639–1648. https://doi.org/10.1016/j.cub.2008.09.063
Georgiou, M., Marinari, E., Burden, J., & Baum, B. (2008). Cdc42, Par6, and aPKC regulate Arp2/3-mediated endocytosis to control local adherens junction stability. Current Biology, 18(21), 1631–1638. https://doi.org/10.1016/j.cub.2008.09.029
Classen, A. K., Anderson, K. I., Marois, E., & Eaton, S. (2005). Hexagonal packing of Drosophila wing epithelial cells by the planar cell polarity pathway. Developmental Cell, 9(6), 805–817. https://doi.org/10.1016/j.devcel.2005.10.016
Sasaki, N., Sasamura, T., Ishikawa, H. O., Kanai, M., Ueda, R., Saigo, K., et al. (2007). Polarized exocytosis and transcytosis of Notch during its apical localization in Drosophila epithelial cells. Genes to Cells, 12(1), 89–103. https://doi.org/10.1111/j.1365-2443.2007.01037.x
Nakayama, Y., Shivas, J. M., Poole, D. S., Squirrell, J. M., Kulkoski, J. M., Schleede, J. B., et al. (2009). Dynamin participates in the maintenance of anterior polarity in the Caenorhabditis elegans embryo. Developmental Cell, 16(6), 889–900. https://doi.org/10.1016/j.devcel.2009.04.009
Palacios, F., Schweitzer, J. K., Boshans, R. L., & D’Souza-Schorey, C. (2002). ARF6-GTP recruits Nm23-H1 to facilitate dynamin-mediated endocytosis during adherens junctions disassembly. Nature Cell Biology, 4(12), 929–936. https://doi.org/10.1038/ncb881
Fournier, H. N., Dupe-Manet, S., Bouvard, D., Lacombe, M. L., Marie, C., Block, M. R., et al. (2002). Integrin cytoplasmic domain-associated protein 1alpha (ICAP-1alpha ) interacts directly with the metastasis suppressor nm23-H2, and both proteins are targeted to newly formed cell adhesion sites upon integrin engagement. Journal of Biological Chemistry, 277(23), 20895–20902. https://doi.org/10.1074/jbc.M200200200
Fournier, H. N., Albiges-Rizo, C., & Block, M. R. (2003). New insights into Nm23 control of cell adhesion and migration. Journal of Bioenergetics and Biomembranes, 35(1), 81–87. https://doi.org/10.1023/a:1023450008347
Kyumurkov, A., Bouin, A. P., Boissan, M., Manet, S., Baschieri, F., & Proponnet-Guerault, M., et al. (2023). Force tuning through regulation of clathrin-dependent integrin endocytosis. Journal of Cell Biology, 222(1), https://doi.org/10.1083/jcb.202004025.
Hoppins, S., Lackner, L., & Nunnari, J. (2007). The machines that divide and fuse mitochondria. Annual Review of Biochemistry, 76, 751–780. https://doi.org/10.1146/annurev.biochem.76.071905.090048
van der Bliek, A. M., Shen, Q., & Kawajiri, S. (2013). Mechanisms of mitochondrial fission and fusion. Cold Spring Harb Perspect Biol, 5(6), https://doi.org/10.1101/cshperspect.a011072.
Vyas, S., Zaganjor, E., & Haigis, M. C. (2016). Mitochondria and Cancer. Cell, 166(3), 555–566. https://doi.org/10.1016/j.cell.2016.07.002
Guerra, F., Guaragnella, N., Arbini, A. A., Bucci, C., Giannattasio, S., & Moro, L. (2017). Mitochondrial Dysfunction: A Novel Potential Driver of Epithelial-to-Mesenchymal Transition in Cancer. Frontiers in Oncology, 7, 295. https://doi.org/10.3389/fonc.2017.00295
Altieri, D. C. (2019). Mitochondrial dynamics and metastasis. Cellular and Molecular Life Sciences, 76(5), 827–835. https://doi.org/10.1007/s00018-018-2961-2
Senft, D., & Ronai, Z. A. (2016). Regulators of mitochondrial dynamics in cancer. Current Opinion in Cell Biology, 39, 43–52. https://doi.org/10.1016/j.ceb.2016.02.001
Zhao, J., Zhang, J., Yu, M., Xie, Y., Huang, Y., Wolff, D. W., et al. (2013). Mitochondrial dynamics regulates migration and invasion of breast cancer cells. Oncogene, 32(40), 4814–4824. https://doi.org/10.1038/onc.2012.494
Ferreira-da-Silva, A., Valacca, C., Rios, E., Populo, H., Soares, P., Sobrinho-Simoes, M., et al. (2015). Mitochondrial dynamics protein Drp1 is overexpressed in oncocytic thyroid tumors and regulates cancer cell migration. PLoS One, 10(3), e0122308. https://doi.org/10.1371/journal.pone.0122308
Zhang, Z., Li, T. E., Chen, M., Xu, D., Zhu, Y., Hu, B. Y., et al. (2020). MFN1-dependent alteration of mitochondrial dynamics drives hepatocellular carcinoma metastasis by glucose metabolic reprogramming. British Journal of Cancer, 122(2), 209–220. https://doi.org/10.1038/s41416-019-0658-4
Liang, J., Yang, Y., Bai, L., Li, F., & Li, E. (2020). DRP1 upregulation promotes pancreatic cancer growth and metastasis through increased aerobic glycolysis. Journal of Gastroenterology and Hepatology, 35(5), 885–895. https://doi.org/10.1111/jgh.14912
Jung, J. U., Ravi, S., Lee, D. W., McFadden, K., Kamradt, M. L., Toussaint, L. G., et al. (2016). NIK/MAP3K14 Regulates Mitochondrial Dynamics and Trafficking to Promote Cell Invasion. Current Biology, 26(24), 3288–3302. https://doi.org/10.1016/j.cub.2016.10.009
Wan, Y. Y., Zhang, J. F., Yang, Z. J., Jiang, L. P., Wei, Y. F., Lai, Q. N., et al. (2014). Involvement of Drp1 in hypoxia-induced migration of human glioblastoma U251 cells. Oncology Reports, 32(2), 619–626. https://doi.org/10.3892/or.2014.3235
Peiris-Pages, M., Bonuccelli, G., Sotgia, F., & Lisanti, M. P. (2018). Mitochondrial fission as a driver of stemness in tumor cells: mDIVI1 inhibits mitochondrial function, cell migration and cancer stem cell (CSC) signalling. Oncotarget, 9(17), 13254–13275. https://doi.org/10.18632/oncotarget.24285.
Chen, C. W., Wang, H. L., Huang, C. W., Huang, C. Y., Lim, W. K., Tu, I. C., et al. (2019). Two separate functions of NME3 critical for cell survival underlie a neurodegenerative disorder. Proceedings of the National Academy of Sciences U S A, 116(2), 566–574. https://doi.org/10.1073/pnas.1818629116
Chen, C. W., Tsao, N., Zhang, W., & Chang, Z. F. (2020). NME3 Regulates Mitochondria to Reduce ROS-Mediated Genome Instability. International Journal of Molecular Sciences, 21(14), https://doi.org/10.3390/ijms21145048.
Imoto, Y., Abe, Y., Honsho, M., Okumoto, K., Ohnuma, M., Kuroiwa, H., et al. (2018). Onsite GTP fuelling via DYNAMO1 drives division of mitochondria and peroxisomes. Nature Communications, 9(1), 4634. https://doi.org/10.1038/s41467-018-07009-z
Tokarska-Schlattner, M., Boissan, M., Munier, A., Borot, C., Mailleau, C., Speer, O., et al. (2008). The nucleoside diphosphate kinase D (NM23-H4) binds the inner mitochondrial membrane with high affinity to cardiolipin and couples nucleotide transfer with respiration. Journal of Biological Chemistry, 283(38), 26198–26207. https://doi.org/10.1074/jbc.M803132200
Schlattner, U., Tokarska-Schlattner, M., Ramirez, S., Tyurina, Y. Y., Amoscato, A. A., Mohammadyani, D., et al. (2013). Dual function of mitochondrial Nm23-H4 protein in phosphotransfer and intermembrane lipid transfer: A cardiolipin-dependent switch. Journal of Biological Chemistry, 288(1), 111–121. https://doi.org/10.1074/jbc.M112.408633
Kagan, V. E., Jiang, J., Huang, Z., Tyurina, Y. Y., Desbourdes, C., Cottet-Rousselle, C., et al. (2016). NDPK-D (NM23-H4)-mediated externalization of cardiolipin enables elimination of depolarized mitochondria by mitophagy. Cell Death and Differentiation, 23(7), 1140–1151. https://doi.org/10.1038/cdd.2015.160
Morlot, S., & Roux, A. (2013). Mechanics of dynamin-mediated membrane fission. Annual Review of Biophysics, 42, 629–649. https://doi.org/10.1146/annurev-biophys-050511-102247
Traut, T. W. (1994). Physiological concentrations of purines and pyrimidines. Molecular and Cellular Biochemistry, 140(1), 1–22. https://doi.org/10.1007/BF00928361
Lascu, I., Schaertl, S., Wang, C., Sarger, C., Giartosio, A., Briand, G., et al. (1997). A point mutation of human nucleoside diphosphate kinase A found in aggressive neuroblastoma affects protein folding. Journal of Biological Chemistry, 272(25), 15599–15602. https://doi.org/10.1074/jbc.272.25.15599
Cervoni, L., Lascu, I., Xu, Y., Gonin, P., Morr, M., Merouani, M., et al. (2001). Binding of nucleotides to nucleoside diphosphate kinase: A calorimetric study. Biochemistry, 40(15), 4583–4589. https://doi.org/10.1021/bi002432s
Otsuki, Y., Tanaka, M., Yoshii, S., Kawazoe, N., Nakaya, K., & Sugimura, H. (2001). Tumor metastasis suppressor nm23H1 regulates Rac1 GTPase by interaction with Tiam1. Proceedings of the National Academy of Sciences U S A, 98(8), 4385–4390. https://doi.org/10.1073/pnas.071411598
Murakami, M., Meneses, P. I., Knight, J. S., Lan, K., Kaul, R., Verma, S. C., et al. (2008). Nm23-H1 modulates the activity of the guanine exchange factor Dbl-1. International Journal of Cancer, 123(3), 500–510. https://doi.org/10.1002/ijc.23568
Murakami, M., Meneses, P. I., Lan, K., & Robertson, E. S. (2008). The suppressor of metastasis Nm23-H1 interacts with the Cdc42 Rho family member and the pleckstrin homology domain of oncoprotein Dbl-1 to suppress cell migration. Cancer Biology & Therapy, 7(5), 677–688. https://doi.org/10.4161/cbt.7.5.5665
Iwashita, S., Fujii, M., Mukai, H., Ono, Y., & Miyamoto, M. (2004). Lbc proto-oncogene product binds to and could be negatively regulated by metastasis suppressor nm23-H2. Biochemical and Biophysical Research Communications, 320(4), 1063–1068. https://doi.org/10.1016/j.bbrc.2004.06.067
Marino, N., Marshall, J. C., Collins, J. W., Zhou, M., Qian, Y., Veenstra, T., et al. (2013). Nm23-h1 binds to gelsolin and inactivates its actin-severing capacity to promote tumor cell motility and metastasis. Cancer Research, 73(19), 5949–5962. https://doi.org/10.1158/0008-5472.CAN-13-0368
Suzuki, E., Ota, T., Tsukuda, K., Okita, A., Matsuoka, K., Murakami, M., et al. (2004). nm23-H1 reduces in vitro cell migration and the liver metastatic potential of colon cancer cells by regulating myosin light chain phosphorylation. International Journal of Cancer, 108(2), 207–211. https://doi.org/10.1002/ijc.11546
Acknowledgements
We thank Marie-Lise Lacombe for her continuing interest in our work and Carol Featherstone of Plume Scientific Communication Services for editorial assistance during the preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Prunier, C., Chavrier, P. & Boissan, M. Mechanisms of action of NME metastasis suppressors – a family affair. Cancer Metastasis Rev 42, 1155–1167 (2023). https://doi.org/10.1007/s10555-023-10118-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-023-10118-x