Abstract
Chromosome instability (CIN) is a cancer hallmark that drives tumour heterogeneity, phenotypic adaptation, drug resistance and poor prognosis. High-grade serous ovarian cancer (HGSOC), one of the most chromosomally unstable tumour types, has a 5-year survival rate of only ~30% — largely due to late diagnosis and rapid development of drug resistance, e.g., via CIN-driven ABCB1 translocations. However, CIN is also a cell cycle vulnerability that can be exploited to specifically target tumour cells, illustrated by the success of PARP inhibitors to target homologous recombination deficiency (HRD). However, a lack of appropriate models with ongoing CIN has been a barrier to fully exploiting disease-specific CIN mechanisms. This barrier is now being overcome with the development of patient-derived cell cultures and organoids. In this review, we describe our progress building a Living Biobank of over 120 patient-derived ovarian cancer models (OCMs), predominantly from HGSOC. OCMs are highly purified tumour fractions with extensive proliferative potential that can be analysed at early passage. OCMs have diverse karyotypes, display intra- and inter-patient heterogeneity and mitotic abnormality rates far higher than established cell lines. OCMs encompass a broad-spectrum of HGSOC hallmarks, including a range of p53 alterations and BRCA1/2 mutations, and display drug resistance mechanisms seen in the clinic, e.g., ABCB1 translocations and BRCA2 reversion. OCMs are amenable to functional analysis, drug-sensitivity profiling, and multi-omics, including single-cell next-generation sequencing, and thus represent a platform for delineating HGSOC-specific CIN mechanisms. In turn, our vision is that this understanding will inform the design of new therapeutic strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many human tumours are characterised by extensive copy number variation (CNV), which arises due to an underlying chromosome instability (CIN) phenotype (Ciriello et al. 2013). CIN leads to continuous gain and loss of chromosomes and/or acquisition of structural rearrangements, in turn driving tumour heterogeneity, phenotypic adaptation and drug resistance (Patch et al. 2015; Schwarz et al. 2015; McPherson et al. 2016; Naffar-Abu Amara et al. 2020; Vasudevan et al. 2020; Ippolito et al. 2021; Lukow et al. 2021). Despite an intense focus on the causes of CIN, we still do not understand the full spectrum of molecular drivers, possibly reflecting the presence of multiple mechanisms and/or disease-specific CIN drivers.
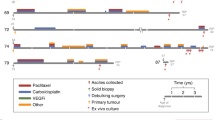
Our focus is on high-grade serous ovarian cancer (HGSOC); one of the most chromosomally unstable cancer types (Ciriello et al. 2013). HGSOC is the commonest histological subtype of ovarian cancer, representing approximately 80% of all cases (Jayson et al. 2014). It is frequently diagnosed at an advanced stage having already undergone metastatic spread beyond the pelvic intraperitoneal tissues. While most cases initially respond to chemotherapy, most women will develop drug-resistant disease (Clamp et al. 2019) (Fig. 1). A known driver of CIN is defective DNA damage repair, and in the case of HGSOC, possibly up to 50% are homologous recombination deficient (HRD), frequently caused by mutations in the BRCA1 and BRCA2 tumour suppressor genes (TCGA 2011; Denkert et al. 2022; Morgan et al. 2023a). Almost 20 years ago, a major advance was the discovery that BRCA1/2-mutant cells are exquisitely sensitive to PARP-1/2 inhibitors (PARPi) (Bryant et al. 2005; Farmer et al. 2005), paving the way for new therapeutic strategies that have had a major beneficial impact in the clinic (Mirza et al. 2016; Coleman et al. 2017; Pujade-Lauraine et al. 2017; Moore et al. 2018; Gonzalez-Martin et al. 2019; Monk et al. 2022). As such, PARPi provide an excellent paradigm illustrating how CIN mechanisms can be exploited to improve patient outcomes. Further exploitation of HGSOC CIN will be important because, in addition to the paucity of actionable oncogenic mutations, at most only 50% are HRD and thus predicted to respond to PARPi. A major research goal therefore is to define the spectrum of CIN mechanisms in HGSOC to identify additional tumour-cell-specific vulnerabilities that can be therapeutically exploited.
Treatment timelines of patients with HGSOC. Line graphs showing CA125 levels, measured via blood sampling, for patients 74 and 110 over time following diagnosis. Graphs are annotated to show surgery (blue up arrows), when ascites were collected (orange down arrows), which ascites generated OCMs (orange stars) and when the patient died (black star). Vertical coloured bars along the top of the plot area show the timing of the indicated chemotherapy treatments
Delineating the full spectrum of CIN mechanisms will require model systems that reflect the diverse CIN phenotypes observed in human tumours. Studies on established human cancer cell lines have been instrumental in dissecting some aspects of CIN mechanisms, and importantly CIN manifests in HGSOC-derived cell lines, with evidence of both mitotic defects and DNA replication stress (Penner-Goeke et al. 2017; Nelson et al. 2020; Tamura et al. 2020). However, while cell lines are experimentally tractable, they have several weaknesses. Often, cell lines were established decades ago in sub-optimal culture conditions that may have selected specific phenotypes (Domcke et al. 2013; Ince et al. 2015; Nelson et al. 2020). Further propagation in vitro likely selects out the fitter subclones best adapted to cell culture conditions, possibly eliminating lesser fit clones that might only survive in vivo. Established cell lines also often lack detailed clinical annotations (e.g., histology, chemotherapy exposure and/or clinical response), and matched pre- and post-treatment lines are rare.
A major advance addressing some of the limitations of established cell lines is the development of Living Biobanks, collections of patient-derived cell cultures or organoid models that are clinically annotated and better capture the heterogeneity observed in human tumours. A key development came from colorectal cancer (CRC), with the discovery of culture techniques that allowed expansion of CRC tumour cells in organoid structures (Sato et al. 2011). This technology has now been extended to other cancers (Gao et al. 2014; Boj et al. 2015; Sachs et al. 2018), including ovarian cancer (Kopper et al. 2019; de Witte et al. 2020). In this review article, we describe our experience developing a Living Biobank of patient-derived ovarian cancer models (OCMs) (Nelson et al. 2020). OCMs are highly purified tumour fractions that have extensive proliferative potential and can be analysed at early passage. They have highly diverse karyotypes, displaying extensive intra- and inter-patient heterogeneity that falls into several subclasses (Fig. 2), and as such provide an attractive starting point for delineating CIN mechanisms.
HGSOC is characterised by extensive chromosome instability. Genome-wide chromosome copy number profiles determined by shallow single-cell whole-genome sequencing (scWGS) of OCMs derived from patients 38, 64, 152 and 246. For each OCM, each row represents a single cell, with chromosomes plotted as columns and the copy number indicated by the colour. The four OCMs shown represent examples whereby genomes are marked by whole-chromosome aneuploidies, rearranged chromosomes, tetrasomies or monosomies. Karyotypes previously shown in Nelson et al. 2020 and Coulson-Gilmer et al. 2021 (Licenses at https://creativecommons.org/licenses/by/4.0/)
Solid sampling versus ascites collection
Standard treatment for ovarian cancer is cytoreductive surgery followed by platinum-based chemotherapy (Jayson et al. 2014), with ~60% of patients in the UK receiving neo-adjuvant chemotherapy. Beyond that, maintenance therapy includes the PARPi olaparib or niraparib, the anti-angiogenic agent bevacizumab, or olaparib plus bevacizumab. Following relapse, a variety of chemotherapeutic strategies can be used, with treatment decision often based on the platinum-free interval (McGee et al. 2017) (Fig. 1). Secondary cytoreductive surgery is less common, but ascites will frequently be removed for symptomatic benefit using therapeutic abdominal paracentesis. Our biopsy pipeline delivers both solid surgical samples and ascitic fluid, and we have developed OCMs from both (Nelson et al. 2020), albeit with a clear bias towards ascites (Fig. 3).
Living Biobank ascites pipeline. A Bar graph summarising the biopsy pipeline, showing that from June 2016 to March 2023, 715 biobank alerts yielded 454 ascites samples from 209 patients, in turn yielding 127 OCMs. B–D Summary of OCM collection with pie charts showing breakdown of subtypes based on pathology assessment (B); chemo-naïve (CN) vs. post-treatment (C); and longitudinal subsets (D). In (D), while 61 patients are represented by a single OCM (blue), 9 patients have 2 longitudinal OCMs (grey), and 3 patient subsets have 3 longitudinal OCMs (orange), etc.
For histological diagnosis and molecular characterisation, solid specimens (tumour tissue either from a diagnostic biopsy or surgical resection) are often considered the gold standard. Key advantages are the ability to sample both primary and metastatic sites, and — via analysis of spatially resolved tumour material — probe heterogeneity between sites and enable the reconstruction of evolutionary trajectories (Schwarz et al. 2015; McPherson et al. 2016; de Witte et al. 2020; Burdett et al. 2023). Because secondary cytoreductive surgery occurs infrequently in the treatment of HGSOC, solid biopsies have limited potential to deliver temporally resolved samples. Furthermore, single surgical samples may not fully capture disease heterogeneity (Hoogstraat et al. 2014; Schwarz et al. 2015; McPherson et al. 2016; Morgan et al. 2023b). In terms of isolating viable tumour cells, solid biopsies can be challenging when tumour material is limited, e.g., with core diagnostic biopsies, and especially following neo-adjuvant chemotherapy, where 60–70% of patients achieve a response, i.e. little or no primary tumour remaining (Morgan et al. 2021). At this point, specimens can be non-viable and so generating a culture is unlikely. However, if grossly visible tumour material is present and/or tumour-rich regions can be isolated by microdissection, ex vivo cultures can be developed even following neoadjuvant chemotherapy (Hill et al. 2018). Another limitation of surgical biospecimens is that they may come from early-stage disease that is typically cured with surgery plus platinum chemotherapy (Trimbos et al. 2003; Collinson et al. 2014). However, death from advanced HGSOC is commonly associated with chemotherapy-resistant disease, which emerges many months or even years later and is not captured at the time of primary cytoreductive surgery (Fig. 1).
The accumulation of ascites presents an alternative method to sample ovarian cancer cells. The presence of tumour cells in the peritoneal cavity can drive fluid build-up by VEGF-mediated increase in capillary permeability and compromised lymphatic drainage (Kipps et al. 2013; Ford et al. 2020). In turn, cytokines, chemokines, and growth factors present in ascites can promote tumour cell survival and further metastatic spread. Excessive fluid is frequently drained for symptom control and, because ascites contains large numbers of tumour cells, it provides excellent opportunities for translational research. Moreover, because abdominal paracentesis provides a safe method for repeat sampling, it opens the opportunity to collect longitudinal samples, including chemo-naïve and spanning multiple treatments (Fig. 1). Because HGSOC is often diagnosed late, when up to 90% of patients will develop ascites (Huang et al. 2013; Ford et al. 2020), this method can capture a wide spectrum of disease. Ascitic fluid can also capture intra-tumour heterogeneity, with one study demonstrating that >92% of somatic mutations detected across multiple intra- and extraovarian solid lesions were represented in ascites-derived tumour samples (Choi et al. 2017). Moreover, genomics datasets from primary disease (generally solid) and chemo-resistant disease (generally ascitic) are largely consistent (TCGA 2011; Patch et al. 2015). In terms of probing biology and exploring therapeutic strategies, ascites collection permits access to chemotherapy-resistant disease, since resistant tumour cells may be absent or represent only a minor proportion of primary cytoreductive surgical samples.
Optimisation of culture media
The development of better experimental models to study ovarian cancer is a major research focus (Bowtell et al. 2015). Indeed, as reviewed recently (Tomas and Shepherd 2023), extensive effort has been applied to develop patient-derived 2D cell cultures, more complex spheroid, organoid or co-culture models, as well as xenografts (Bertozzi et al. 2006; Shepherd et al. 2006; Latifi et al. 2012; Sueblinvong et al. 2012; Thériault et al. 2013; Davidowitz et al. 2014; Lengyel et al. 2014; Ince et al. 2015; Liu et al. 2017; Thu et al. 2017; Hill et al. 2018; Kopper et al. 2019; Maru et al. 2019; Phan et al. 2019; Fritz et al. 2020; Hoffmann et al. 2020; Maenhoudt et al. 2020; Brodeur et al. 2021; Ito et al. 2023; Vias et al. 2023). Such efforts have been required because establishing primary cell cultures from tumours using traditional cell culture techniques has historically been challenging, with very low success rates due to tumour cell ‘senescence’ and with the emerging cell lines reflecting rare subclones (Ince et al. 2015). A major breakthrough was the development of highly specialised cell culture conditions capable of propagating isolated CRC cells as organoids (Sato et al. 2011), an approach then adapted to breast (Sachs et al. 2018) and epithelial ovarian cancers (Kopper et al. 2019; de Witte et al. 2020). A parallel breakthrough was the development of Ovarian Carcinoma Modified Ince (OCMI) media, by T Ince, with J Brugge, G Mills and colleagues, which allows propagation of epithelial ovarian cancer cells as 2D monolayers (Ince et al. 2015). Prior to adopting OCMI, our attempts to develop proliferative ex vivo HGSOC cultures were unsuccessful; while fibroblasts isolated from ascites proliferated in traditional RPMI-based formulations, the associated tumour cells did not. Adopting OCMI had a transformative effect; as of March 1st, 2023, we have received 454 ascites samples from 209 patients and thus far generated 127 OCMs (Fig. 3). The ‘take-rate’ at first pass is approximately 30% and, in some cases, OCMs have been generated following second and third attempts by fine-tuning initial conditions. Importantly, the vast majority of OCMs can also be revived after cryopreservation; thus far only two OCMs do not revive. Generation of highly purified tumour fractions is usually possible in under five passages, allowing extensive analyses on early passages. If the tumour cells are p53-deficient, this process can be accelerated by selectively killing p53-proficient stromal cells with Nutlin-3 (Nelson et al. 2020). OCMs cultured in OCMI have extensive proliferative potential, with some propagated beyond 50 passages. Prolonged propagation is anticipated to select for the fitter, faster growing subclones that may be more chromosomally stable over time (Nelson et al. 2020). Indeed, in due course OCMs are anticipated to behave like established cell lines.
Ex vivo cultures retain the hallmark characteristics of HGSOC
A key question is whether ascites-derived OCMs reflect the primary tumour. While at first this question seems straightforward, upon closer inspection it is more nuanced. In many cases, OCMs and the corresponding primary tumour are separated by many months if not years (Fig. 1). Considering both the extensive genomic plasticity caused by CIN, and significant selection pressures exerted by multiple rounds of chemotherapy, one might expect the tumour cells sampled in ascites to have diverged considerably from the original primary tumour.
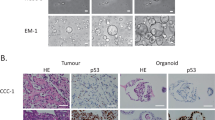
OCMs derived from patient 64 illustrate this incredible plasticity (Nelson et al. 2020). OCMs 64-1 and 64-3 were generated from ascites collected from the same patient 49 days apart, the first and third abdominal drains respectively, almost 2.5 years after surgery. Microscopy revealed that many cells in OCM.64-3 had similar morphology to those in OCM.64-1, with large, atypical nuclei, and negative PAX8 and EpCAM expression (Fig. 4). However, we also identified a second population in OCM.64-3 that had smaller nuclei and were positive for both PAX8 and EpCAM. By exploiting the differential EpCAM status, we physically separated the two sub-populations to create OCM.64-3-Ep+ and OCM.64-3-Ep-. This revealed that the EpCAM-negative population expressed high levels of MYC and had a gene expression profile that more closely resembled OCM.64-1 (Fig. 4).
Chromosome instability generates highly divergent subclones. A scWGS-derived karyotypes of EpCAM-positive and EpCAM-negative subpopulations present in the OCM generated from the 3rd ascites sample collected form patient 64. B Table summarising characteristics of OCMs 64-1 and the two 64-3 subpopulations. C Speculative ploidy reversal event to explain how the two 64-3 subpopulations might have been generated. Karyotypes in A adapted from Nelson et al. 2020 (License at https://creativecommons.org/licenses/by/4.0/)
Karyotype analysis revealed that OCM.64-1 was dominated by tetrasomies (Fig. 2) (Nelson et al. 2020). By contrast, OCM.64-3-Ep- harboured disomies and trisomies, while OCM.64-3-Ep+ harboured numerous monosomies (Fig. 4). Importantly, the p53 mutation — p.V216M — was identical, and unique in the collection to date, indicating a clonal origin (Fig. 4). Interestingly, the disomies in OCM.64-3-Ep- were mirrored by monosomies in OCM.64-3-Ep+ (Fig. 4). One possible explanation is that an unequal mitosis resulted in a ploidy reversal event, giving rise to the two cell types found in 64-3 (Fig. 4). Note that ploidy reversal has been described in polyploid hepatocytes as part of a mechanism to generate genomic diversity (Duncan 2013). If ploidy reversal did occur, this would represent an additional mode of punctuated tumour cell evolution, yielding very rapid genomic divergence. Nevertheless, this subset of OCMs illustrates the remarkable plasticity of HGSOC cells in terms of key tumour markers, gene expression profiles and karyotype. In turn, illustrating that perhaps beyond truncal TP53 mutations, we should be cautious in terms of our expectations when comparing primary tumours and ascites-derived cells, especially when separated by extended periods of time and/or chemotherapy regimens.
Despite the complexity outlined above, we have compared OCMs with their corresponding archival tumour blocks using a panel of standard markers used to diagnose HGSOC in the clinic. Analysis of CK7, PAX8, WT1 and p53 expression (Fig. 5), aided by specialist pathology support, was remarkably congruent (Nelson et al. 2020; Coulson-Gilmer et al. 2021). In addition, targeted amplicon sequencing of primary tumour DNA by a clinically accredited diagnostic service, using a multi-gene panel that included TP53, showed excellent congruence with Sanger sequencing of RT-PCR products from matched OCMs.
Primary HGSOC can display very different histopathologies. Representative 20× immunohistochemistry images of the primary tumours from patients 92 and 109, stained to detect p53, PAX8, WT1 and Cytokeratin 7. Patient 92 images adapted from Coulson-Gilmer et al. 2021 (License at https://creativecommons.org/licenses/by/4.0/). Scale bar, 100 μm. Panels are representative images from single experiment
A related, but perhaps more relevant, question is whether the OCM workflow generates models that reflect the tumour sampled by the respective ascites. Importantly, the pioneering study of Ince et al., demonstrated that the OCMI media maintain the genomic and transcriptomic landscape of the original tumour, and that xenograft tumours show morphology typical of human tumours (Ince et al. 2015). Moreover, by generating OCMs within 5–6 passages, our workflow enables analysis before the expansion of subclonal populations. Additional evidence supporting the generation of reflective models comes from the analysis of OCMs prepared from sequential ascites; in many cases the karyotypes and gene expression profiles are similar (Fig. 6).
OCM gene expression analysis. Principal component (PC) analysis of RNAseq-derived global gene expression profiles, distinguishing stromal and tumour clades, and showing the close relationship of longitudinal OCMs samples from patients 64, 66, 74, 110, 118 and 124, with numbers inside the symbol indicating the ascites number. 69* is a stromal culture. Published data collated from Nelson et al. 2020, Barnes et al. 2021, Coulson-Gilmer et al. 2021
With a take rate of ~30%, another key question is selection bias; does the workflow only select for a subset of HGSOC subtypes? OCM gene expression profiles do display substantial heterogeneity (Fig. 6). In addition, analysis of TP53 mutations shows that the proportion of missense mutations versus truncating mutations is similar to that described by the TCGA, as is the nature of the missense mutations (Fig. 7) (TCGA 2011; Cerami et al. 2012; Gao et al. 2013). Interestingly however, of the 42 OCMs sequenced so far, we are yet to identify an R273 missense mutation, despite this mutation being the most frequent in the TCGA analysis.
TP53 mutation profile. A Pie charts showing the number of different TP53 mutation subtypes in the TCGA dataset compared with the subset of OCMs for which TP53 data is currently available. B Comparison of missense TP53 mutations in the TCGA dataset (grey) versus the OCM subset (purple). OCM data collated from Nelson et al. 2020, Coulson-Gilmer et al. 2021; TCGA data from cBioPortal (Cerami et al. 2012; Gao et al. 2013)
It has been suggested that ex vivo culture may select against BRCA1/2-mutant tumours (Hill et al. 2018; Hoffmann et al. 2020; Vias et al. 2023). When we screened a subset of 32 OCMs, 8 were found to be sensitive to PARP inhibition, suggesting an HR-defect (Coulson-Gilmer et al. 2021). Also, in a subset of 20 OCMs derived from patients with known BRCA1/2 status, seven had germline BRCA1/2 mutations (Barnes et al. 2021; Coulson-Gilmer et al. 2021). When we analysed OCMs from four of these seven, three harboured BRCA1/2 mutations (Coulson-Gilmer et al. 2021). Thus, while the number of OCMs fully analysed to date is still relatively small, there is no obvious evidence yet of a selection bias against BRCA1/2-mutant or HRD tumours. Indeed, some OCMs appear to reflect the complex mechanisms responsible for drug resistance in patients. Using Rad51 foci formation in response to ionising radiation as a functional readout of HR status, we established that OCM.109 is HRD and harbours a BRCA1 mutation (Coulson-Gilmer et al. 2021). However, it is PARPi-resistant suggesting a resistance mechanism that bypasses the HR defect. OCM.246 was derived from a patient with a germline BRCA2 mutation who received olaparib maintenance monotherapy prior to biopsy sampling. Interestingly, this OCM harbours a putative intragenic reversion predicted to restore the BRCA2 open reading frame, reflecting reversion mechanisms previously described in patients (Christie et al. 2017; Burdett et al. 2023), and the OCM displays intermediate PARPi resistance (Coulson-Gilmer et al. 2021).
Another mechanism of acquired drug resistance in HGSOC is chromosome translocation events leading to upregulation of the drug efflux pump encoded by ABCB1 (Patch et al. 2015; Christie et al. 2019). A number of OCMs demonstrate upregulated ABCB1 expression and can be re-sensitized to paclitaxel using the efflux inhibitor Elacridar (not shown). Taking all this together, our experience to date is consistent with the notion that the biopsy pipeline and OCM workflow have generated a diverse collection of ovarian cancer models that reflects the disease heterogeneity observed in traditional sample collections. Plus, the biobank reflects various drug resistance mechanisms that have been described previously. As such, the Living Biobank provides a unique opportunity to probe aspects of HGSOC biology and explore novel therapeutic strategies.
OCMs display ongoing CIN
While HGSOC is driven by CIN, mutations in genes directly involved in chromosome replication and segregation are extremely rare in cancer (Matthews et al. 2022). To delineate CIN mechanisms, HGSOC has been studied by whole-genome sequencing (WGS). One landmark study defined two mutational trajectories, the first characterised by HRD, with BRCA1/2 mutations, amplification of MYC and loss of RB1; the second characterised by homologous recombination proficiency (HRP) with foldback inversions (FBI) correlating with CCNE1 amplification and PTEN loss (Wang et al. 2017). While elegant, this dualistic model is likely an oversimplification and indeed, a second key study using shallow WGS identified seven CNV signatures, including two HRD signatures and five HRP signatures (Table 1) (Macintyre et al. 2018).
Matched deep sequencing assigned potential pathways to CNV signatures; one HRD signature was associated with BRCA1/2 mutations and loss of PTEN, while the other was non-BRCA1/2-related with MYC amplification (Macintyre et al. 2018). HRP signatures were associated with various trajectories including oncogenic RAS, inactivation of CDK12, or cell cycle deregulation (Table 1). Multiple signatures were observed to co-exist in the same sample, including HRD and HRP signatures. Also, the composite signature was predictive, e.g., patients with a high degree of signature 1 had poor prognosis. More recently, a study of 7880 tumours from 33 different tissues was used to devise 17 pan-cancer CNV signatures, including three signatures associated with impaired homologous recombination (IHR) alongside varying degrees of replication stress (Drews et al. 2022). One of these IHR signatures correlated with the two HRD signatures identified in ovarian cancers. Another recent study identified 21 pan-cancer CNV signatures, nine of which were present in ovarian cancer, including one that may be unique to ovarian cancer that could not be assigned to a biological process (Steele et al. 2022).
While ground-breaking, these studies expose important new questions. The presence of multiple signatures is complicated by bulk sequencing archival material; single-cell analyses will be required to disentangle whether individual cells exhibit multiple signatures, or whether this reflects intra-tumour heterogeneity and/or specific microenvironments (Shah 2018). Thus, well-defined in vitro models and/or derived subclones amenable to functional experiments will be required to test hypotheses correlating signatures with cell behaviours (Macintyre et al. 2018).
Our vision is that Living Biobanks will provide opportunities to address these issues. Indeed, a key advantage of viable cultures is the ability to analyse highly purified tumour fractions unfettered by contaminating, genetically normal stromal cells, and the microenvironment. Moreover, they are amenable to single-cell analyses, including both shallow WGS and RNA sequencing (Nelson et al. 2020). But most significantly, as viable, proliferating cultures, they can be subjected to functional experiments designed to probe the status of specific signalling and cell cycle pathways, thereby enabling hypotheses that emerge from interrogation of molecular features to be tested more rigorously using phenotypic assays.
As proof-of-principle, to assess CIN functionally in OCMs, we analysed patterns of mitotic chromosome segregation using time-lapse microscopy, facilitated by stable integration of a GFP-tagged histone to visualise the chromatin (Nelson et al. 2020). This revealed highly chaotic and heterogeneous mitoses, with rates of abnormalities far higher than previously observed in established cell lines. Rates of lagging chromosomes, anaphase bridges and cytokinesis/abscission failures were all elevated. The difficulties with chromosome alignment very often resulted in a protracted mitosis, indicating a robust spindle assembly checkpoint. And indeed, when challenged with microtubule toxins, OCMs underwent longer mitotic delays. Interestingly, because of the self-imposed protracted mitosis, we observed several instances of cohesion fatigue (Daum et al. 2011; Stevens et al. 2011); to our knowledge this is the first time this has been seen without experimentally blocking mitosis (Nelson et al. 2020).
While many of the highly abnormal cell divisions did give rise to viable progeny, and certainly sufficient to maintain a proliferative culture, cell fate profiling revealed a number of dead-ends, consistent with the notion some genomes are incompatible with life (Nelson et al. 2020). Nevertheless, the extent of mitotic chaos was surprising and suggests that a key feature of HGSOC is deactivation of post-mitotic and/or apoptotic pathways that would normally eliminate genetic deviants. Moreover, it also supports the notion that OCMs provide interesting alternatives to established cell lines for analysing HGSOC CIN mechanisms. Indeed, although established cell lines exhibit ongoing CIN (Lengauer et al. 1997; Penner-Goeke et al. 2017; Tamura et al. 2020), a limited number of subclones tend to dominate (Wangsa et al. 2018), presumably because they represent the fittest, fastest growing cells (Domcke et al. 2013; Ince et al. 2015; Nelson et al. 2020). Accordingly, one might expect that OCMs that start out highly heterogenous would become less complex over time, as the fitter subclones give rise to more progeny with every passage. Empirical evidence supports this. When we analysed spindle poles as a proxy for CIN, comparing OCMs at early and late passages, complexity reduced over time with bipolar spindles becoming more dominant, presumably because they are both already fitter and more likely to give rise to viable daughters (Nelson et al. 2020). This further highlights the advantages of being able to analyse OCMs at early passage when the population is still complex. Ideally, one would want to be able to isolate and expand different subclones; while this is possible (Naffar-Abu Amara et al. 2020), it can be challenging to expand single cells in vitro. However, advances in bar coding technology mean that it is possible to trace lineages without the need for exerting the stress associated with single-cell cloning (Gutierrez et al. 2021). Such bar-coding technologies open up exciting opportunities to study genome evolution and the emergence of drug resistance in patient-derived tumour material.
The heterogenous mitoses described above were observed when the OCMs were cultured as 2D monolayers (Nelson et al. 2020). Interestingly, it has been found that tissue architecture can impact chromosome segregation fidelity (Knouse et al. 2018). In particular, when mouse epithelial cells were cultured as 3D spheroids the rates of chromosome mis-segregation were very low, but this rate increased to ~7% in 2D culture. This raises the possibility that the mitotic errors observed in OCMs may in part be an artefact of in vitro 2D culture. We suspect that this is not the case. In the OCMs, chromosome mis-segregation rates were often around 50%, far higher than was observed in the primary mouse epithelial cells. Moreover, when we specifically grew the OCMs in a 3D environment, we observed equally high rates of segregation error (Nelson et al. 2020). Moreover, we observed additional classes of abnormal mitoses in 3D, including chromosome ejection at anaphase, possibly reflecting the ability of a 3D environment to better anchor ectopic spindle poles.
Taking together the various WGS studies, plus our OCM-derived observations (shallow scWGS karyotyping, time-lapse microscopy, and more traditional M-FISH-based karyotyping) (Nelson et al. 2020), a very consistent picture emerges — HGSOC genomes are highly dynamic, undergoing persistent and high rates of CIN. A key next step will be to align these different modalities and integrate CIN signatures with mutational profiles derived from bulk WGS data and gene expression signatures from RNA sequencing. This multi-omics data can then be aligned with clinical outcome data, as well as functional phenotypes derived from cell-based analysis and drug-sensitivity profiling to test hypotheses (see below). Thus, the OCMs represent an invaluable resource to delineate mechanisms underlying aberrant mitoses and CIN in HGSOC cells. Of particular value will be matched longitudinal OCMs, especially those that include OCMs from both chemotherapy-naïve and post-treatment disease, to better understand how CIN drives the emergence of drug resistance in patients.
A platform for drug discovery
CIN has the ability to drive the emergence of drug resistance in patients; for example, chromosome translocations within ABCB1 can lead to overexpression of the MDR1/p-glycoprotein drug efflux pump (Patch et al. 2015; Christie et al. 2019). Importantly, we identified ABCB1 translocations in a number of OCMs and have shown that drug sensitivity can be restored by co-exposure with efflux inhibitors (not shown). Interestingly, in OCM.246 we identified three different ABCB1 translocations (Williams et al. 2020), as well as a putative BRCA2 reversion mutation (Coulson-Gilmer et al. 2021), illustrating both the incredible capacity of CIN to alter the genome and the intense selective pressure that chemotherapy exerts. Also, these observations provide further evidence that OCMs provide a window into the drug resistance mechanisms seen in patients (Patch et al. 2015; Christie et al. 2017; Christie et al. 2019; Burdett et al. 2023), and that OCMs provide a potentially interesting platform for drug-sensitivity profiling to complement multi-omics analyses.
To measure drug sensitivities of OCMs, we have optimised a high-throughput assay that uses object counting to measure proliferation (Nelson et al. 2020; Coulson-Gilmer et al. 2021; Golder et al. 2022). In brief, OCMs expressing a GFP-tagged histone are analysed by time-lapse microscopy and changes in green object count over time are used as a proxy for proliferation. The doubling time is then calculated by determining the inverse gradient of the linear portion of a log2 transformation of the fluorescent object count, normalised to t = 0 h (Golder et al. 2022). This approach has advantages over traditional end-point viability assays that infer cell viability by measuring ATP metabolism, which can be confounded by cytostatic effects whereby cells stop proliferating but remain metabolically active (Niepel et al. 2019). Moreover, the approach is very data rich, providing single-cell-level resolution over time. Interrogating time-lapse sequences can provide additional information in terms of cell fate and behaviour simply not apparent in population-based end-point assays. Using this approach, we have measured proliferation rates of numerous OCMs (Pillay et al. 2019; Nelson et al. 2020; Coulson-Gilmer et al. 2021; Golder et al. 2022).
Analysing proliferation in response to drug exposure then enables drug-sensitivity profiling. In brief, we determine the half maximal growth inhibition concentration of drug (GI50) using dose-response curves generated by measuring the area-under-the-curve of fluorescent object count over time for a range of drug concentrations (Golder et al. 2022). The high-throughput nature enables multiple technical replicates and the tractability of the OCMs enables biological replicates to be analysed in quick succession. Various parameters can influence multi-well assay readouts, and recently we explored a number of parameters including cell seeding density and assay duration, as well as analytical approaches to account for variability in cell cycle duration (Golder et al. 2022). While there is heterogeneity due to the complex nature of the OCMs, estimates of doubling times were largely consistent when remeasured 18 months apart, and ex vivo responses to platinum largely reflected patient responses (Nelson et al. 2020; Golder et al. 2022). Thus far, we have evaluated OCM sensitivity to cisplatin, paclitaxel and inhibitors targeting PARP-1/2 and the PAR glycohydrolase (Pillay et al. 2019; Nelson et al. 2020; Coulson-Gilmer et al. 2021). More recently, screening a panel of 16 diverse OCMs, using all possible one-, two-, three- and four-drug combinations of four inhibitors targeting the DNA replication stress response at GI10 concentrations (240 assays in total), demonstrated that the low-dose combination of ATR and CHK1 inhibitors had significant activity against 15 OCMs, identifying a potentially novel therapeutic strategy (Golder et al. 2022).
Due to the extensive proliferative potential of OCMs, drug sensitivity can also be assessed by longer-term colony formation assays, thereby complementing the shorter-term time-lapse-based proliferation assays (Coulson-Gilmer et al. 2021). While OCMs can be cultured in a 3D context, thus far we have measured drug sensitivity of 2D monolayers. Because the microenvironment can influence drug sensitivity, it will be interesting to analyse in vitro chemotherapy responses in more complex 3D and co-culture models (Tomas and Shepherd 2023). However, it is noteworthy that sensitivity and resistance to PARPi, which arguably represent the most significant advancement in recent years for treating patients with HGSOC, manifests very clearly when analysing OCMs as 2D monolayers (Coulson-Gilmer et al. 2021). Indeed, our focus is on exploiting cell cycle vulnerabilities intrinsic to the tumour cells, vulnerabilities that may be less sensitive to the tumour microenvironment. Furthermore, the impact of the microenvironment on response to therapy is highly complex, for example extracellular matrix components have been associated with both chemotherapy sensitivity and resistance (Ahmed et al. 2007; Etemadmoghadam et al. 2009; Helleman et al. 2010; Kozlova et al. 2020; Guo et al. 2021). Clearly, these interactions bring about additional complexity, therefore our approach in the first instance is to focus on dissecting intrinsic tumour cell properties and drug responses. Armed with this knowledge, we will be better placed to explore how the microenvironment modulates tumour cell biology. Importantly, a number of OCMs have been successfully engrafted in immunocompromised mice to form xenograft tumours. These OCM-derived xenograft (ODX) models retain the molecular features of the original OCM, both in vivo and ex vivo following excision and disaggregation (not shown). Such ODX models will provide excellent opportunities to test new therapeutic strategies in vivo that emerge from drug sensitivity profiling of 2D monolayers.
Future perspectives
The Living Biobank currently contains over 120 OCMs from more than 80 patients and is expanding at a rate of 2–3 new OCMs per month. Fifteen ascites-derived OCMs are chemo-naïve and, as the biobank grows, the number of longitudinal cohorts with matched post-treatment OCMs will expand. While focused on HGSOC, we are also collecting smaller cohorts of OCMs derived from other ovarian cancer subtypes as these provide interesting comparators. Although ascites offer various advantages, we have recently received more solid samples (and currently have eight solid-derived OCMs), which increases the potential to broaden the diversity of the biobank. Thus, in summary, the pipeline and workflow we have developed has allowed assembly of a large and diverse collection of ovarian cancer models that reflect the diversity of HGSOC. Importantly, the experimental tractability of OCMs in terms of integrating multi-omics data, including single cell approaches, with functional assays, including high-resolution cell biology approaches and drug-sensitivity profiling, opens up new opportunities to delineate the molecular mechanisms responsible for driving CIN in this particular disease. An important future goal is also to collate the wealth of data associated with the OCMs in a searchable format, so that it is available to researchers in the ovarian cancer community alongside the OCMs.
Abbreviations
- CIN:
-
Chromosome instability
- CN:
-
Chemotherapy-naïve
- CNV:
-
Copy number variation
- CRC:
-
Colorectal cancer
- HGSOC:
-
High-grade serous ovarian cancer
- HR:
-
Homologous recombination
- HRD:
-
Homologous recombination deficient
- HRP:
-
Homologous recombination proficiency
- LGSOC:
-
Low-grade serous ovarian cancer
- IHR:
-
Impaired homologous recombination
- OCM:
-
Ovarian cancer model
- OCMI:
-
Ovarian carcinoma modified Ince
- ODX:
-
OCM-derived xenograft
- PARPi:
-
PARP-1/2 inhibitor
- PC:
-
Principal component
- RNAseq:
-
RNA sequencing
- scWGS:
-
Single-cell whole-genome sequencing
- WGS:
-
Whole-genome sequencing
References
Ahmed AA, Mills AD, Ibrahim AE, Temple J, Blenkiron C, Vias M, Massie CE, Iyer NG, McGeoch A, Crawford R et al (2007) The extracellular matrix protein TGFBI induces microtubule stabilization and sensitizes ovarian cancers to paclitaxel. Cancer Cell 12:514–527
Barnes BM, Nelson L, Tighe A, Burghel GJ, Lin IH, Desai S, McGrail JC, Morgan RD, Taylor SS (2021) Distinct transcriptional programs stratify ovarian cancer cell lines into the five major histological subtypes. Genome Med 13:140
Bertozzi CC, Chang CY, Jairaj S, Shan X, Huang J, Weber BL, Chu CS, Carroll RG (2006) Multiple initial culture conditions enhance the establishment of cell lines from primary ovarian cancer specimens. In Vitro Cell Dev Biol Anim 42:58–62
Boj SF, Hwang CI, Baker LA, Chio II, Engle DD, Corbo V, Jager M, Ponz-Sarvise M, Tiriac H, Spector MS et al (2015) Organoid models of human and mouse ductal pancreatic cancer. Cell 160:324–338
Bowtell DD, Bohm S, Ahmed AA, Aspuria PJ, Bast RC Jr, Beral V, Berek JS, Birrer MJ, Blagden S, Bookman MA et al (2015) Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer 15:668–679
Brodeur MN, Simeone K, Leclerc-Deslauniers K, Fleury H, Carmona E, Provencher DM, Mes-Masson A-M (2021) Carboplatin response in preclinical models for ovarian cancer: comparison of 2D monolayers, spheroids, ex vivo tumors and in vivo models. Sci Rep 11:18183
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, Kyle S, Meuth M, Curtin NJ, Helleday T (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434:913–917
Burdett NL, Willis MO, Alsop K, Hunt AL, Pandey A, Hamilton PT, Abulez T, Liu X, Hoang T, Craig S et al (2023) Multiomic analysis of homologous recombination-deficient end-stage high-grade serous ovarian cancer. Nat Genet 55:437–450
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404
Choi YJ, Rhee JK, Hur SY, Kim MS, Lee SH, Chung YJ, Kim TM, Lee SH (2017) Intraindividual genomic heterogeneity of high-grade serous carcinoma of the ovary and clinical utility of ascitic cancer cells for mutation profiling. J Pathol 241:57–66
Christie EL, Fereday S, Doig K, Pattnaik S, Dawson SJ, Bowtell DDL (2017) Reversion of BRCA1/2 germline mutations detected in circulating tumor DNA from patients with high-grade serous ovarian cancer. J Clin Oncol 35:1274–1280
Christie EL, Pattnaik S, Beach J, Copeland A, Rashoo N, Fereday S, Hendley J, Alsop K, Brady SL, Lamb G et al (2019) Multiple ABCB1 transcriptional fusions in drug resistant high-grade serous ovarian and breast cancer. Nat Commun 10:1295
Ciriello G, Miller ML, Aksoy BA, Senbabaoglu Y, Schultz N, Sander C (2013) Emerging landscape of oncogenic signatures across human cancers. Nat Genet 45:1127–1133
Clamp AR, James EC, McNeish IA, Dean A, Kim JW, O'Donnell DM, Hook J, Coyle C, Blagden S, Brenton JD et al (2019) Weekly dose-dense chemotherapy in first-line epithelial ovarian, fallopian tube, or primary peritoneal carcinoma treatment (ICON8): primary progression free survival analysis results from a GCIG phase 3 randomised controlled trial. Lancet 394:2084–2095
Coleman RL, Oza AM, Lorusso D, Aghajanian C, Oaknin A, Dean A, Colombo N, Weberpals JI, Clamp A, Scambia G et al (2017) Rucaparib maintenance treatment for recurrent ovarian carcinoma after response to platinum therapy (ARIEL3): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 390:1949–1961
Collinson F, Qian W, Fossati R, Lissoni A, Williams C, Parmar M, Ledermann J, Colombo N, Swart A (2014) Optimal treatment of early-stage ovarian cancer. Ann Oncol 25:1165–1171
Coulson-Gilmer C, Morgan RD, Nelson L, Barnes BM, Tighe A, Wardenaar R, Spierings DCJ, Schlecht H, Burghel GJ, Foijer F et al (2021) Replication catastrophe is responsible for intrinsic PAR glycohydrolase inhibitor-sensitivity in patient-derived ovarian cancer models. J Exp Clin Cancer Res 40:323
Daum JR, Potapova TA, Sivakumar S, Daniel JJ, Flynn JN, Rankin S, Gorbsky GJ (2011) Cohesion fatigue induces chromatid separation in cells delayed at metaphase. Curr Biol 21:1018–1024
Davidowitz RA, Selfors LM, Iwanicki MP, Elias KM, Karst A, Piao H, Ince TA, Drage MG, Dering J, Konecny GE et al (2014) Mesenchymal gene program-expressing ovarian cancer spheroids exhibit enhanced mesothelial clearance. J Clin Invest 124:2611–2625
de Witte CJ, Espejo Valle-Inclan J, Hami N, Lõhmussaar K, Kopper O, Vreuls CPH, Jonges GN, van Diest P, Nguyen L, Clevers H et al (2020) Patient-derived ovarian cancer organoids mimic clinical response and exhibit heterogeneous inter- and intrapatient drug responses. Cell Rep 31:107762
Denkert C, Romey M, Swedlund B, Hattesohl A, Teply-Szymanski J, Kommoss S, Kaiser K, Staebler A, du Bois A, Grass A et al (2022) Homologous recombination deficiency as an ovarian cancer biomarker in a real-world cohort: validation of decentralized genomic profiling. J Mol Diagn 24:1254–1263
Domcke S, Sinha R, Levine DA, Sander C, Schultz N (2013) Evaluating cell lines as tumour models by comparison of genomic profiles. Nat Commun 4:2126
Drews RM, Hernando B, Tarabichi M, Haase K, Lesluyes T, Smith PS, Morrill Gavarró L, Couturier DL, Liu L, Schneider M et al (2022) A pan-cancer compendium of chromosomal instability. Nature 606:976–983
Duncan AW (2013) Aneuploidy, polyploidy and ploidy reversal in the liver. Semin Cell Dev Biol 24:347–356
Etemadmoghadam D, deFazio A, Beroukhim R, Mermel C, George J, Getz G, Tothill R, Okamoto A, Raeder MB, Harnett P et al (2009) Integrated genome-wide DNA copy number and expression analysis identifies distinct mechanisms of primary chemoresistance in ovarian carcinomas. Clin Cancer Res 15:1417–1427
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB, Santarosa M, Dillon KJ, Hickson I, Knights C et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434:917–921
Ford CE, Werner B, Hacker NF, Warton K (2020) The untapped potential of ascites in ovarian cancer research and treatment. Br J Cancer 123:9–16
Fritz JL, Collins O, Saxena P, Buensuceso A, Ramos Valdes Y, Francis KE, Brown KR, Larsen B, Colwill K, Gingras AC et al (2020) A novel role for NUAK1 in promoting ovarian cancer metastasis through regulation of fibronectin production in spheroids. Cancers 12:1250
Gao D, Vela I, Sboner A, Iaquinta PJ, Karthaus WR, Gopalan A, Dowling C, Wanjala JN, Undvall EA, Arora VK et al (2014) Organoid cultures derived from patients with advanced prostate cancer. Cell 159:176–187
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:pl1
Golder A, Nelson L, Tighe A, Barnes B, Coulson-Gilmer C, Morgan RD, McGrail JC, Taylor SS (2022) Multiple-low-dose therapy: effective killing of high-grade serous ovarian cancer cells with ATR and CHK1 inhibitors. NAR. Cancer 4:zcac036
Gonzalez-Martin A, Pothuri B, Vergote I, DePont Christensen R, Graybill W, Mirza MR, McCormick C, Lorusso D, Hoskins P, Freyer G et al (2019) Niraparib in patients with newly diagnosed advanced ovarian cancer. N Engl J Med 381:2391–2402
Guo W, He X, Ni J, Ma L, Cheng X, Wang C, Chen X, Wang Y (2021) High serpin family a member 10 expression confers platinum sensitivity and is associated with survival benefit in high-grade serous ovarian cancer: based on quantitative proteomic analysis. Front Oncol 11:761960
Gutierrez C, Al’Khafaji AM, Brenner E, Johnson KE, Gohil SH, Lin Z, Knisbacher BA, Durrett RE, Li S, Parvin S et al (2021) Multifunctional barcoding with ClonMapper enables high-resolution study of clonal dynamics during tumor evolution and treatment. Nat Can 2:758–772
Helleman J, Jansen MP, Burger C, van der Burg ME, Berns EM (2010) Integrated genomics of chemotherapy resistant ovarian cancer: a role for extracellular matrix, TGFbeta and regulating microRNAs. Int J Biochem Cell Biol 42:25–30
Hill SJ, Decker B, Roberts EA, Horowitz NS, Muto MG, Worley MJ Jr, Feltmate CM, Nucci MR, Swisher EM, Nguyen H et al (2018) Prediction of DNA repair inhibitor response in short-term patient-derived ovarian cancer organoids. Cancer Discov 8:1404–1421
Hoffmann K, Berger H, Kulbe H, Thillainadarasan S, Mollenkopf HJ, Zemojtel T, Taube E, Darb-Esfahani S, Mangler M, Sehouli J et al (2020) Stable expansion of high-grade serous ovarian cancer organoids requires a low-Wnt environment. EMBO J 39:e104013
Hoogstraat M, de Pagter MS, Cirkel GA, van Roosmalen MJ, Harkins TT, Duran K, Kreeftmeijer J, Renkens I, Witteveen PO, Lee CC et al (2014) Genomic and transcriptomic plasticity in treatment-naive ovarian cancer. Genome Res 24:200–211
Huang H, Li YJ, Lan CY, Huang QD, Feng YL, Huang YW, Liu JH (2013) Clinical significance of ascites in epithelial ovarian cancer. Neoplasma 60:546–552
Ince TA, Sousa AD, Jones MA, Harrell JC, Agoston ES, Krohn M, Selfors LM, Liu W, Chen K, Yong M et al (2015) Characterization of twenty-five ovarian tumour cell lines that phenocopy primary tumours. Nat Commun 6:7419
Ippolito MR, Martis V, Martin S, Tijhuis AE, Hong C, Wardenaar R, Dumont M, Zerbib J, Spierings DCJ, Fachinetti D et al (2021) Gene copy-number changes and chromosomal instability induced by aneuploidy confer resistance to chemotherapy. Dev Cell 56:2440–2454.e2446
Ito Y, Kondo J, Masuda M, Matsuzaki S, Onuma K, Kanda M, Watanabe Y, Sakaguchi H, Yoshino K, Ueda Y et al (2023) Ex vivo chemosensitivity assay using primary ovarian cancer organoids for predicting clinical response and screening effective drugs. Hum Cell 36:752–761
Jayson GC, Kohn EC, Kitchener HC, Ledermann JA (2014) Ovarian cancer. Lancet 384:1376–1388
Kipps E, Tan DS, Kaye SB (2013) Meeting the challenge of ascites in ovarian cancer: new avenues for therapy and research. Nat Rev Cancer 13:273–282
Knouse KA, Lopez KE, Bachofner M, Amon A (2018) Chromosome segregation fidelity in epithelia requires tissue architecture. Cell 175(200-211):e213
Kopper O, de Witte CJ, Lohmussaar K, Valle-Inclan JE, Hami N, Kester L, Balgobind AV, Korving J, Proost N, Begthel H et al (2019) An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nat Med 25:838–849
Kozlova N, Grossman JE, Iwanicki MP, Muranen T (2020) The interplay of the extracellular matrix and stromal cells as a drug target in stroma-rich cancers. Trends Pharmacol Sci 41:183–198
Latifi A, Luwor RB, Bilandzic M, Nazaretian S, Stenvers K, Pyman J, Zhu H, Thompson EW, Quinn MA, Findlay JK, Ahmed N (2012) Isolation and characterization of tumor cells from the ascites of ovarian cancer patients: molecular phenotype of chemoresistant ovarian tumors. PLoS One 7:e46858
Lengauer C, Kinzler KW, Vogelstein B (1997) Genetic instability in colorectal cancers. Nature 386:623–627
Lengyel E, Burdette JE, Kenny HA, Matei D, Pilrose J, Haluska P, Nephew KP, Hales DB, Stack MS (2014) Epithelial ovarian cancer experimental models. Oncogene 33:3619–3633
Liu JF, Palakurthi S, Zeng Q, Zhou S, Ivanova E, Huang W, Zervantonakis IK, Selfors LM, Shen Y, Pritchard CC et al (2017) Establishment of patient-derived tumor xenograft models of epithelial ovarian cancer for preclinical evaluation of novel therapeutics. Clin Cancer Res 23:1263–1273
Lukow DA, Sausville EL, Suri P, Chunduri NK, Wieland A, Leu J, Smith JC, Girish V, Kumar AA, Kendall J et al (2021) Chromosomal instability accelerates the evolution of resistance to anti-cancer therapies. Dev Cell 56:2427–2439.e2424
Macintyre G, Goranova TE, De Silva D, Ennis D, Piskorz AM, Eldridge M, Sie D, Lewsley LA, Hanif A, Wilson C et al (2018) Copy number signatures and mutational processes in ovarian carcinoma. Nat Genet 50:1262–1270
Maenhoudt N, Defraye C, Boretto M, Jan Z, Heremans R, Boeckx B, Hermans F, Arijs I, Cox B, Van Nieuwenhuysen E et al (2020) Developing organoids from ovarian cancer as experimental and preclinical models. Stem Cell Reports 14:717–729
Maru Y, Tanaka N, Itami M, Hippo Y (2019) Efficient use of patient-derived organoids as a preclinical model for gynecologic tumors. Gynecol Oncol 154:189–198
Matthews HK, Bertoli C, de Bruin RAM (2022) Cell cycle control in cancer. Nat Rev Mol Cell Biol 23:74–88
McGee J, Bookman M, Harter P, Marth C, McNeish I, Moore KN, Poveda A, Hilpert F, Hasegawa K, Bacon M et al (2017) Fifth Ovarian Cancer Consensus Conference: individualized therapy and patient factors. Ann Oncol 28:702–710
McPherson A, Roth A, Laks E, Masud T, Bashashati A, Zhang AW, Ha G, Biele J, Yap D, Wan A et al (2016) Divergent modes of clonal spread and intraperitoneal mixing in high-grade serous ovarian cancer. Nat Genet 48:758–767
Mirza MR, Monk BJ, Herrstedt J, Oza AM, Mahner S, Redondo A, Fabbro M, Ledermann JA, Lorusso D, Vergote I et al (2016) Niraparib maintenance therapy in platinum-sensitive, recurrent ovarian cancer. N Engl J Med 375:2154–2164
Monk BJ, Parkinson C, Lim MC, O'Malley DM, Oaknin A, Wilson MK, Coleman RL, Lorusso D, Bessette P, Ghamande S et al (2022) A randomized, phase III trial to evaluate rucaparib monotherapy as maintenance treatment in patients with newly diagnosed ovarian cancer (ATHENA-MONO/GOG-3020/ENGOT-ov45). J Clin Oncol 40:3952–3964
Moore K, Colombo N, Scambia G, Kim BG, Oaknin A, Friedlander M, Lisyanskaya A, Floquet A, Leary A, Sonke GS et al (2018) Maintenance olaparib in patients with newly diagnosed advanced ovarian cancer. N Engl J Med 379:2495–2505
Morgan RD, Clamp AR, Barnes BM, Timms K, Schlecht H, Yarram-Smith L, Wallis Y, Valganon-Petrizan M, MacMahon S, White R et al (2023a) Homologous recombination deficiency in newly diagnosed FIGO stage III/IV high-grade epithelial ovarian cancer: a multi-national observational study. Int J Gynecol Cancer 33:1253–1259
Morgan RD, Clamp AR, White DJ, Price M, Burghel GJ, Ryder WDJ, Mahmood RD, Murphy AD, Hasan J, Mitchell CL et al (2023b) Multi-maintenance Olaparib therapy in relapsed, germline BRCA1/2-mutant high-grade serous ovarian cancer (MOLTO): a phase II trial. Clin Cancer Res 29:2602–2611
Morgan RD, McNeish IA, Cook AD, James EC, Lord R, Dark G, Glasspool RM, Krell J, Parkinson C, Poole CJ et al (2021) Objective responses to first-line neoadjuvant carboplatin-paclitaxel regimens for ovarian, fallopian tube, or primary peritoneal carcinoma (ICON8): post-hoc exploratory analysis of a randomised, phase 3 trial. Lancet Oncol 22:277–288
Naffar-Abu Amara S, Kuiken HJ, Selfors LM, Butler T, Leung ML, Leung CT, Kuhn EP, Kolarova T, Hage C, Ganesh K et al (2020) Transient commensal clonal interactions can drive tumor metastasis. Nat Commun 11:5799
Nelson L, Tighe A, Golder A, Littler S, Bakker B, Moralli D, Murtuza Baker S, Donaldson IJ, Spierings DCJ, Wardenaar R et al (2020) A living biobank of ovarian cancer ex vivo models reveals profound mitotic heterogeneity. Nat Commun 11:822
Niepel M, Hafner M, Mills CE, Subramanian K, Williams EH, Chung M, Gaudio B, Barrette AM, Stern AD, Hu B et al (2019) A multi-center study on the reproducibility of drug-response assays in mammalian cell lines. Cell Syst 9:35–48.e35
Patch AM, Christie EL, Etemadmoghadam D, Garsed DW, George J, Fereday S, Nones K, Cowin P, Alsop K, Bailey PJ et al (2015) Whole-genome characterization of chemoresistant ovarian cancer. Nature 521:489–494
Penner-Goeke S, Lichtensztejn Z, Neufeld M, Ali JL, Altman AD, Nachtigal MW, McManus KJ (2017) The temporal dynamics of chromosome instability in ovarian cancer cell lines and primary patient samples. PLoS Genet 13:e1006707
Phan N, Hong JJ, Tofig B, Mapua M, Elashoff D, Moatamed NA, Huang J, Memarzadeh S, Damoiseaux R, Soragni A (2019) A simple high-throughput approach identifies actionable drug sensitivities in patient-derived tumor organoids. Commun Biol 2:78
Pillay N, Tighe A, Nelson L, Littler S, Coulson-Gilmer C, Bah N, Golder A, Bakker B, Spierings DCJ, James DI et al (2019) DNA replication vulnerabilities render ovarian cancer cells sensitive to poly(ADP-Ribose) glycohydrolase inhibitors. Cancer Cell 35(519-533):e518
Pujade-Lauraine E, Ledermann JA, Selle F, Gebski V, Penson RT, Oza AM, Korach J, Huzarski T, Poveda A, Pignata S et al (2017) Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 mutation (SOLO2/ENGOT-Ov21): a double-blind, randomised, placebo-controlled, phase 3 trial. Lancet Oncol 18:1274–1284
Sachs N, de Ligt J, Kopper O, Gogola E, Bounova G, Weeber F, Balgobind AV, Wind K, Gracanin A, Begthel H et al (2018) A living biobank of breast cancer organoids captures disease heterogeneity. Cell 172(373-386):e310
Sato T, Stange DE, Ferrante M, Vries RG, Van Es JH, Van den Brink S, Van Houdt WJ, Pronk A, Van Gorp J, Siersema PD, Clevers H (2011) Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett's epithelium. Gastroenterology 141:1762–1772
Schwarz RF, Ng CK, Cooke SL, Newman S, Temple J, Piskorz AM, Gale D, Sayal K, Murtaza M, Baldwin PJ et al (2015) Spatial and temporal heterogeneity in high-grade serous ovarian cancer: a phylogenetic analysis. PLoS Med 12:e1001789
Shah SP (2018) Copy number signatures in ovarian cancer. Nat Genet 50:1208–1209
Shepherd TG, Thériault BL, Campbell EJ, Nachtigal MW (2006) Primary culture of ovarian surface epithelial cells and ascites-derived ovarian cancer cells from patients. Nat Protoc 1:2643–2649
Steele CD, Abbasi A, Islam SMA, Bowes AL, Khandekar A, Haase K, Hames-Fathi S, Ajayi D, Verfaillie A, Dhami P et al (2022) Signatures of copy number alterations in human cancer. Nature 606:984–991
Stevens D, Gassmann R, Oegema K, Desai A (2011) Uncoordinated loss of chromatid cohesion is a common outcome of extended metaphase arrest. PLoS One 6:e22969
Sueblinvong T, Ghebre R, Iizuka Y, Pambuccian SE, Isaksson Vogel R, Skubitz AP, Bazzaro M (2012) Establishment, characterization and downstream application of primary ovarian cancer cells derived from solid tumors. PLoS One 7:e50519
Tamura N, Shaikh N, Muliaditan D, Soliman TN, McGuinness JR, Maniati E, Moralli D, Durin MA, Green CM, Balkwill FR et al (2020) Specific mechanisms of chromosomal instability indicate therapeutic sensitivities in high-grade serous ovarian carcinoma. Cancer Res 80:4946–4959
TCGA (2011) Integrated genomic analyses of ovarian carcinoma. Nature 474:609–615
Thériault BL, Portelance L, Mes-Masson AM, Nachtigal MW (2013) Establishment of primary cultures from ovarian tumor tissue and ascites fluid. Methods Mol Biol 1049:323–336
Thu KL, Papari-Zareei M, Stastny V, Song K, Peyton M, Martinez VD, Zhang YA, Castro IB, Varella-Garcia M, Liang H et al (2017) A comprehensively characterized cell line panel highly representative of clinical ovarian high-grade serous carcinomas. Oncotarget 8:50489–50499
Tomas E, Shepherd TG (2023) Insights into high-grade serous carcinoma pathobiology using three-dimensional culture model systems. J Ovarian Res 16:70
Trimbos JB, Vergote I, Bolis G, Vermorken JB, Mangioni C, Madronal C, Franchi M, Tateo S, Zanetta G, Scarfone G et al (2003) Impact of adjuvant chemotherapy and surgical staging in early-stage ovarian carcinoma: European organisation for research and treatment of cancer–adjuvant chemotherapy in ovarian neoplasm trial. J Natl Cancer Inst 95:113–125
Vasudevan A, Baruah PS, Smith JC, Wang Z, Sayles NM, Andrews P, Kendall J, Leu J, Chunduri NK, Levy D et al (2020) Single-chromosomal gains can function as metastasis suppressors and promoters in colon cancer. Dev Cell 52:413–428.e416
Vias M, Morrill Gavarró L, Sauer CM, Sanders DA, Piskorz AM, Couturier D-L, Ballereau SP, Hernando B, Schneider MP, Hall J et al (2023) High-grade serous ovarian carcinoma organoids as models of chromosomal instability. Elife 12:e83867
Wang YK, Bashashati A, Anglesio MS, Cochrane DR, Grewal DS, Ha G, McPherson A, Horlings HM, Senz J, Prentice LM et al (2017) Genomic consequences of aberrant DNA repair mechanisms stratify ovarian cancer histotypes. Nat Genet 49:856–865
Wangsa D, Braun R, Schiefer M, Gertz EM, Bronder D, Quintanilla I, Padilla-Nash HM, Torres I, Hunn C, Warner L et al (2018) The evolution of single cell-derived colorectal cancer cell lines is dominated by the continued selection of tumor-specific genomic imbalances, despite random chromosomal instability. Carcinogenesis 39:993–1005
Williams MS, Basma NJ, Amaral FMR, Williams G, Weightman JP, Breitwieser W, Nelson L, Taylor SS, Wiseman DH, Somervaille TCP (2020) Targeted nanopore sequencing for the identification of ABCB1 promoter translocations in cancer. BMC Cancer 20:1075
Acknowledgements
We thank the patients for their commitment to research; the MCRC Biobank for the sample collection; members of the Taylor lab for advice and comments on the manuscript.
Funding
Construction and characterisation of the living biobank is funded by a Cancer Research UK Programme Grant awarded to S Taylor (C1422/A31334).
Author information
Authors and Affiliations
Contributions
L.N. and S.S.T contributed to conceptualisation and design. Methodology development, investigation, and data curation were performed by L.N; A,T; S.L; C.C.G; A.G; S.D; R.D.M. Formal analysis and visualisation was performed by L.N; B.M.B; R.D.M and S.S.T. Project administration was performed by J.C.M. J.C.M and S.S.T wrote the original draft. Supervision and funding acquisition was by S.S.T. All authors reviewed and edited drafts and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
OCMs were established and characteristics published previously (Nelson et al. 2020; Barnes et al. 2021; Coulson-Gilmer et al. 2021; Golder et al. 2022). The research samples were obtained with informed patient consent from the Manchester Cancer Research Centre (MCRC) Biobank. The MCRC Biobank is licensed by the Human Tissue Authority (license number: 30004) and is ethically approved as a research tissue bank by the South Manchester Research Ethics Committee (Ref: 22/NW/0237). The role of the MCRC Biobank is to distribute samples and does not endorse studies performed or the interpretation of results. For more information, see https://www.mcrc.manchester.ac.uk/research/mcrc-biobank.
Consent to participate
Research samples were obtained with informed patient consent from the MCRC Biobank (see above).
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Stefano Santaguida
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nelson, L., Barnes, B.M., Tighe, A. et al. Exploiting a living biobank to delineate mechanisms underlying disease-specific chromosome instability. Chromosome Res 31, 21 (2023). https://doi.org/10.1007/s10577-023-09731-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10577-023-09731-x