Abstract
Several countries have implemented stocking programmes to enhance abundance and fish production by releases of hatchery-reared fish. However, due to fluctuations in population size, stocking history, and potential indirect effects of straying of hatchery-reared fish, it is often difficult to predict how these factors will affect genetic diversity and differentiation patterns among wild populations. This study characterized the population genetic structure and temporal variability of four Estonian sea trout populations by evaluating the degrees of direct and indirect genetic impacts of stocking over two decades using 14 microsatellite loci. Our results demonstrate considerable temporal change combined with weak genetic structuring among studied sea trout populations. We found a reduction of the overall level of genetic differentiation combined with the tendency for increased genetic diversity, and an effective number of breeders (Nb) over the study period. Furthermore, we found that immigration rates (m) from hatchery stocks were highest in the population subjected to direct stocking and in non-stocked populations that were located geographically closer to the stocked rivers. This work suggests that hatchery releases have influenced the genetic diversity and structuring of studied sea trout populations. However, the impact of hatchery releases on the adaptive variation and fitness-related traits in wild trout populations remains to be revealed by more informative genetic markers. This study illustrates the dynamic nature of the population genetic structure of sea trout and the value of long-term genetic monitoring for management and conservation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the past century, widespread declines and even extirpations of salmonid populations due to increasing human-related activities such as fisheries, pollution, habitat destruction, fragmentation, and alteration have occurred throughout most of salmonid fishes natural ranges (Parrish et al. 1998; Susnik et al. 2004; HELCOM 2011; Perrier et al. 2013). Stocking of captive-reared fish of native, non-native, or mixed origin has become an integral part of the process to support threatened and endangered populations (Aprahamian et al. 2003; Hansen et al. 2009; Laikre et al. 2010). Although stocking is an important tool to achieve management goals, numerous studies have shown that stocking with hatchery-reared fish can have a major impact on the genetic structure of wild salmonid populations (Eldridge et al. 2009; Hansen et al. 2009; Marie et al. 2010; Ozerov et al. 2016; Östergren et al. 2021). Several studies have reported that stocking practices can result in variable admixture rates between donor and source populations (Campos et al. 2008; Sønstebø et al. 2008; Hansen et al. 2009; Perrier et al. 2011; Ozerov et al. 2016). Populations influenced by hatchery releases may show a reduction of genetic differentiation (Susnik et al. 2004; Eldridge and Naish 2007; Hansen et al. 2009, 2010; Ozerov et al. 2016) and loss or increase of genetic variability and potential disruption of local adaptations (Hansen et al. 2001a, b; Borrell et al. 2008; Laikre et al. 2008; Eldridge et al. 2009; Ozerov et al. 2016; Östergren et al. 2021). Furthermore, hatchery-reared fish frequently show less accurate homing behaviour than wild conspecifics (Jensen et al. 2005; Vasemägi et al. 2005; Jonsson and Jonsson 2006; Hansen and Mensberg 2009), and released fish have been recovered in rivers other than those into which they were stocked (Vasemägi et al. 2005; Sønstebø et al. 2008; Degerman et al. 2012).
Genetic analysis of samples taken at two or more time points has become increasingly popular to assess the changes in diversity and population structure of wild fish (Laikre et al. 1998; Ostergaard et al. 2003; Palm et al. 2003; Jensen et al. 2005; Campos et al. 2007; Borrell et al. 2008; Nielsen and Hansen 2008; Hansen et al. 2009; Gudmundsson et al. 2013; Ozerov et al. 2013, 2016; Christensen et al. 2018). This is because population genetic inferences based on a single sample typically are able to provide only a snapshot of evolutionary and demographic processes. In contrast, temporal analyses have led to important insights on the genetic stability of fish populations and changes in effective population size (Jorde and Ryman 1995; Hansen et al. 2002; Laikre et al. 2002; Ostergaard et al. 2003; Palm et al. 2003; Jensen et al. 2005; Campos et al. 2007; Borrell et al. 2008), population responses to pronounced climate changes (Christensen et al. 2018) and habitat fragmentation (Yamamoto et al. 2004; Sandlund et al. 2014). Furthermore, several studies have assessed the short- and long-term genetic effects of stocking and fish farm escapees on wild salmonid populations using samples gathered over time (Tessier and Bernatchez 1999; Hansen et al. 2000, 2006; Finnegan and Stevens 2008; Eldridge et al. 2009; Hansen et al. 2009; Hansen and Mensberg 2009; Hansen et al. 2010; Gudmundsson et al. 2013; Perrier et al. 2013; Valiquette et al. 2014; Ozerov et al. 2016; Pritchard et al. 2016).
Anadromous brown trout (Salmo trutta L.), often called sea trout, reproduces in streams and rivers where juveniles spend one to several years before they undergo smoltification and migrate to the sea, where they reach maturity after one or more years, and subsequently return back to their native rivers to spawn (Klemetsen et al. 2003). In the Baltic Sea, the sea trout is one of the important diadromous fish species (ICES 2020). However, the populations have declined throughout their range in the Baltic Sea basin as a result of a number of anthropogenic stressors, including habitat degradation, migration barriers, poaching, pollution, and overfishing (HELCOM 2011b; Pedersen et al. 2012; HELCOM 2013; ICES 2019, 2020). Currently, ca. 500 sea trout populations reproduce naturally in the Baltic rivers (HELCOM 2011). Earlier studies have demonstrated that trout populations in the Baltic Sea are hierarchically structured according to the geographical regions (Koljonen et al. 2014; Pocwierz-Kotus et al. 2014; Östergren et al. 2016). On a smaller geographical scale, e.g., among rivers of the same region or even among tributaries within large river systems, genetic relationships between populations also tend to reflect their geographical proximity and connectivity (Hansen et al. 2009; Lehtonen et al. 2009; Samuiloviene et al. 2009; Koljonen et al. 2014; Östergren et al. 2016).
In Estonia, sea trout populations are found in about 75 rivers and streams, and more than half of them (39) are descending to the Gulf of Finland area (HELCOM 2011; ICES 2020). Due to the high harvest rate and deterioration of habitat quality in the 1990s, sea trout parr densities in Estonia decreased and stayed at a low level until the middle of the 2010s (HELCOM 2011b; Pedersen et al. 2012). As a result, a stocking programme in the Gulf of Finland was implemented from the 2000s to the end of the 2010s, when the situation in stocked rivers had improved. The rivers were subjected to supportive stocking using hatchery-reared offspring of local wild spawners (2001–2020) or alternatively, F1 offspring of hatchery broodstocks that were created based on local sea trout in the state-owned Põlula Fish Rearing Centre hatchery (2008–2014) (Ministry of the Environment 2020; Põlula Fish Rearing Centre 2021; pers. comm E. Saadre). The earlier population genetic characterization of Estonian sea trout, conducted using microsatellite markers, showed close genetic relationships among Gulf of Finland populations, with pairwise FST estimates ranging from 0.002 to 0.041 (Koljonen et al. 2014). However, due to fluctuations in population size and the recent history of stocking, we currently do not know how these processes may have influenced genetic diversity and differentiation patterns among Estonian sea trout populations. Furthermore, estimates of demographic parameters obtained from temporal samples, such as effective population size (Ne), which determines the extent of random genetic drift and inbreeding, as well as the efficacy of selection (Frankham et al. 2002), can add useful information about the status of populations.
In this study, we characterized the spatial and temporal genetic variability of four Estonian sea trout (Salmo trutta L.) populations over a period of more than 20 years based on 14 microsatellite loci. We compared the genetic structure of these populations prior to, and several generations after, stocking activities which provided an excellent opportunity to explore the long-term effects of hatchery releases. Furthermore, we describe the genetic effects of stocking on sea trout populations which either (i) have directly experienced hatchery releases (i.e. direct effect) or (ii) have not experienced direct hatchery releases but may have been impacted by straying of hatchery fish (i.e. indirect effect). Our main goals were to: (i) assess the temporal changes in genetic diversity and differentiation over time, (ii) quantify changes in effective population size and identify potential genetic bottlenecks, and (iii) evaluate the degree of introgression from hatchery releases.
Materials and methods
Study area, sample collection and stocking information
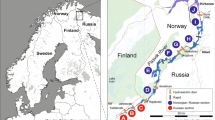
A sampling of sea trout juveniles (0 + and 1 + age; n = 600) was carried out using electrofishing during the annual national salmonid parr density surveys in four northern Estonian rivers flowing into the Gulf of Finland of the Baltic Sea (R. Vasalemma, R. Loobu, R. Selja, and R. Kunda). The surveys were carried out by the Estonian Marine Institute, the University of Tartu and electrofished areas spanned from 100 to 400 m2 depending on location and water level (Fig. 1, red lines). Each river had physical obstacles to the upstream migration of fish, such as waterfalls or dams, and available stretches for sea trout upstream migration varied between rivers (Table S2). Sampling of juveniles was carried out in August-September. Small pieces of fin clips were sampled non-lethally and stored in 96% ethanol for later genetic analysis. The first samples from each river were collected in 1997–2001 and were assumed not to have been affected by stocking activities because during these years large-scale stocking activities had just started, while subsequent samples were collected in 2017–2019 and were assumed to reflect the more recent status of the Estonian sea trout gene pool, which is potentially affected by stocking activities (Table S3). To directly compare the temporal data sets all individuals were divided into cohorts according to their year of birth: earlier period samples (year-classes 1996–2001) and later period samples (year-classes 2017–2019; Table 1 and S3). In addition, to allow evaluation of the degree of introgression from hatchery releases, the samples of juvenile individuals (0 + and 1 + age; n = 436) from four wild populations (Pudisoo, Mustoja, Selja, and Kunda Rivers) and samples of adult individuals (9 + age; n = 159; fin clips) from Põlula Fish Rearing Centre hatchery broodstock were collected in 1997–2014 (Fig. 1; Table S1 and S2). No recent hatchery releases have been carried out in the Rivers Vasalemma, Loobu, and Kunda, while regular releases of hatchery-reared trout into the River Selja occurred during the period 2001–2006 (Table S2). The River Selja was subjected to supportive breeding using hatchery-reared offspring based on River Selja and River Pudisoo wild spawners (Table S2). Since 1994, the density of wild sea trout parr on spawning sites showed significant increases for three out of the four studied rivers (Vasalemma, Loobu, and Selja; simple linear regression’s R2 = 0.49–0.68, P < 0.05), while the density of parr in River Kunda, did not change significantly (R2 = 0.29, P > 0.05) (Kesler et al. 2021). Further information on the hatchery releases to Estonian rivers flowing into the Gulf of Finland can be found in Fig. 1 and Table S2.
Map illustrating the geographical location of the studied sea trout rivers (bold red lines), the rivers in which regular releases of hatchery-origin sea trout juveniles were carried out (black lines), and the rivers, where the broodstocks and the hatchery-reared juveniles of wild spawners originated are additionally marked with an asterisk. Black dots on the red lines indicate sampling points. Numbers correspond to the rivers in Table S2. Inserted line plots indicate the mean density of 0 + sea trout per 100 m2 in the four studied rivers based on monitoring data from 1994 to 2020 (Kesler et al. 2021)
DNA extraction and microsatellite analysis
DNA from fin clips collected in 1997–2001 (a total of 278 individual samples) was isolated according to the simplified method of Laird et al. (1991) and samples collected in 2012–2019 (a total of 784 individual samples) were extracted using NucleoSpin® Tissue kit (Macherey-Nagel GmbH). All samples (n = 1062) were genotyped at 14 microsatellite loci: SsOsl417, SsOsl311 (Slettan et al. 1995), Str60INRA, Str15INRA, Str73INR (Estoup et al. 1993), Ssa407 (Cairney et al. 2000), Bs131 (Estoup et al. 1998), SsOsl438 (Slettan et al. 1996), Strutta58 (Poteaux et al. 1999), OneU9 (Scribner et al. 1996), Ssa85 (McConnell et al. 1995), Sssp1605 (Paterson et al. 2004), Ssa197 (Oreilly et al. 1996) and Str85INRA (Presa and Guyomard 1996). Two multiplex PCR reactions (7 loci per multiplex) were carried out in a total reaction volume of 10 µl which contained 1 × Type-it Multiplex PCR Master Mix (Qiagen), 50–400 nM of each primer (concentration and fluorescent labelling of specific primers are described in Appendix 1 Table A2 in Koljonen et al. 2014), and ca. 10–20 ng of DNA template. Amplifications were performed using the following temperature profile: initial denaturation at 95 °C for 5 min followed by 26 cycles of denaturation at 95 °C for 30 s, annealing at 56 ºC for 60 s, and extension at 72 °C for 30 s, all of which were followed by a final extension at 60 °C for 30 min. The amplification products were separated by capillary electrophoresis on AB3500 (1997–2014 samples) and AB3500XL (2017–2019 samples) Genetic Analyzers (Applied Biosystems, Foster City, CA) using LIZ600 (Applied Biosystems, Foster City, USA) as the internal molecular size. The sizes of the microsatellite alleles were determined using GeneMapper software v. 5.0 (Applied Biosystems, Foster City, CA). Microsatellite genotypes were initially scored automatically and were double-checked manually.
Data analysis
To assess the levels of genetic diversity, the number of unique alleles (private alleles) in a year-class, mean number of alleles (A), allelic richness (AR), observed (HO), and expected heterozygosity (unbiased genetic diversity, HE), were calculated using FSTAT v. 2.9.3.2 (Goudet 1995) and MICROSATELLITE TOOLKIT v. 3.1.1 (Park 2001). The FSTAT v. 2.9.3.2 software was also used to calculate Weir and Cockerham’s (1984) within year-class inbreeding coefficient (FIS) and pairwise FST values. Each year-class was checked for the presence of null alleles and scoring errors due to stuttering or large allele dropouts using the Brookfield 1 estimator (Brookfield 1996), implemented in the program MICRO-CHECKER v. 2.2.3 (Van Oosterhout et al. 2004). The significance of the FIS was estimated by the bootstrap method implemented in GENETIX v. 4.05.2 software (Belkhir et al. 2004). GENEPOP v. 4.7.5 (Rousset 2008) was used to test deviation from the Hardy-Weinberg equilibrium (HWE) (10,000 iterations each) for every locus year-class combination with Fisher’s exact test. All probability tests were based on the Markov chain method (Guo and Thompson 1992; Raymond and Rousset 1995) using 1,000 de-memorization steps, 100 batches, and 1,000 iterations per batch. All tests included Bonferroni corrections (Rice 1989). To identify closely related individuals (full- and half-siblings) the software COLONY v. 2.0.5.0 (Jones and Wang 2010) was used. The model used evaluated the number of full-sib families in all year-classes assuming polygamous reproduction among males and females (Fleming 1996).
Temporal stability of population genetic structure
Statistical significance of differences in the estimates of genetic differentiation (Weir and Cockerham’s (1984) FST) of all studied populations and their year-classes between earlier (1996–2001) and later (2017–2019) time periods was estimated using FSTAT v. 2.9.3.2 (Goudet 1995). FSTAT was also applied for estimating the statistical significance of differences in the estimates of FST between earlier and later time periods for each population (groups of year-classes) separately using a permutation scheme implemented in FSTAT v. 2.9.3.2 (two-sided test with 1,000 permutations; Goudet 1995). Hierarchical analysis of molecular variance (AMOVA) (Excoffier et al. 1992) incorporated in ARLEQUIN v. 3.5.2.2 (Excoffier and Lischer 2010) was used to quantify spatial and temporal genetic variation and its statistical significance. We quantified the amount of spatial and temporal variation for the whole data set (1996–2019) and for the two periods separately (1996–2001 and 2017–2019). The hierarchy levels were set among populations (groups of year-classes of the same population; FCT), among year-classes within populations (FSC), and within year-classes (FST). The variance components were tested statistically by non-parametric randomization tests using 10,000 permutations. The genetic distances between year-classes were estimated according to Nei’s genetic distance (DA) (Nei et al. 1983), and a population tree was constructed with the neighbour-joining (NJ) algorithm (Saitou and Nei 1987) using the POPULATIONS v. 1.2.31 software (Langella 1999). Branch support was estimated with 1,000 bootstrap replications over loci and the resulting tree was visualized in MEGA v. 6.06 software (Tamura et al. 2013). Statistical significance of the differences in the average estimates of the AR and HE between 1996–2001 and 2017–2019 time periods for each population (groups of year-classes) were assessed using a non-parametric Wilcoxon signed-rank test and a permutation scheme implemented in FSTAT v. 2.9.3.2 (two-sided test with 1,000 permutations; Goudet 1995).
Estimation of effective population size, effective number of breeders, and bottlenecks
Three different methods were applied to estimate effective population size (Ne). First, we used the linkage disequilibrium (LD) method (Hill 1981) implemented in NeESTIMATOR v2 (Do et al. 2014) to estimate Ne based on alleles with allele frequencies larger than 5%. Second, Ne was estimated by the sibship assignment (SA) approach implemented in COLONY v. 2.0.5.0 (Jones and Wang 2010). This approach estimates demographic parameters from the multilocus genotypes of a sample allowing calculation of the probabilities that a pair of offspring taken at random from the population are half- or full-sibs. The COLONY runs were performed with the following options: female and male polygamy, random mating, full-likelihood method, and medium-length run. Since both LD and SA methods require only a single sample, the Ne was separately calculated for each year-class resulting in 3–6-point estimates per population. The estimation from a sample of individuals taken at random from a single year-class in a population with overlapping generations gives an estimate of an annual effective number of breeders (Nb) that produced the cohort (Wang 2009), rather than the Ne per generation (Waples et al. 2013). Therefore, for LD and SA approach we will use hereafter the notation as Nb, rather than Ne. Thirdly, Ne was estimated using a maximum-likelihood method implemented in the program MLNe v. 1.0 (Wang and Whitlock 2003). This method estimates Ne considering migration as an additional source of variation in allele frequencies and also quantifies the immigration rate (m) from the assumed source population. For this, we created four putative source populations (potential sources of immigrants) based on spawners of a Põlula hatchery captive broodstock (source 1) and trout samples from the rivers, which were used as spawners for producing hatchery-reared offspring for subsequent stocking purposes: pooled samples of wild-caught fish from the R. Pudisoo and R. Mustoja (source 2), and earlier period samples from R. Selja (source 3) and R. Kunda (source 4) (Table S1 and S2). MLNe requires a user-specified upper limit for Ne, which was set at 200 after checking that similar results were obtained with higher upper limits for Ne. Furthermore, we assumed an average generation time of 3.5 years (Rannak et al. 1983). As the intervals between year-classes were not integers, all estimates were adjusted according to the equations provided by Wang and Whitlock (2003).
To detect recent population genetic bottlenecks, two tests were used: the Wilcoxon sign-rank test which is based on heterozygosity excess, and the mode-shift test which evaluates the allele frequency distribution. Both bottleneck tests were performed with the program BOTTLENECK v. 1.2.02 (Piry et al. 1999) using the stepwise mutation model (SMM) and the two-phase model (TPM) comprising 95% SMM and 5% infinite allele model with the variance for mutation size set to 12 (Piry et al. 1999).
Results
Genetic diversity at microsatellite loci
We found no strong evidence for allele dropouts nor scoring errors due to stuttering, although MICRO-CHECKER suggested a putative null allele at 8 out of 14 microsatellite loci in 13 year-classes (one to two loci per year-class; Table S3). Estimated null allele frequencies ranged from 0.063 to 0.177 (Brookfield 1 estimator), and the highest frequency of null alleles was found in the locus SSsp1605 (Table S3). However, as only 15 out of 252 tests for null alleles were significant (5.9%, i.e., close to the expected Type-I error level), we decided not to exclude any loci from further analysis. Moreover, omitting some loci with putative null alleles (e.g. SSsp1605) had only a negligible effect on the main results (data not shown). Deviations from the Hardy-Weinberg equilibrium (P < 0.05) were detected in 22 of 252 locus-sample combinations. After correcting for multiple tests (Rice 1989), only a single combination remained significant (α = 0.0002; Table S3). The significant deviation was not driven by the deficit nor the excess of the heterozygotes (Table S3).
When sampling juveniles of trout using electrofishing, there is a risk of sampling individuals that share the same parents, i.e. that are full- or half-siblings. Analysis with COLONY v. 2.0.5.0 showed that our samples consisted of only small full-sib families (maximum of two full-sibs). We therefore did not exclude any individuals from further analysis, since small sibling groups are not expected to seriously bias subsequent population genetic inferences, while removing individuals may reduce the precision and statistical power of different genetic analysis approaches (Waples and Anderson 2017).
A total of 175 alleles were observed across the 14 microsatellite loci with an average of 12.5 alleles per locus, ranging from four alleles at Str60INRA to 38 alleles at Ssa407. The number of alleles varied little between the two time periods: a total of 157 and 159 alleles were observed during the earlier and later time periods, respectively. However, we detected 16 alleles during the earlier period samples that were not observed in the later period, while 18 alleles were only observed in more recent samples (data not shown). All microsatellite loci were polymorphic in all year-classes, except Vas-98, for which the locus OneU9 was monomorphic. The average HE of the studied loci was 0.645 and varied from 0.000 (OneU9) to 0.966 (Ssa407) (Table S3). The mean genetic diversity estimates over 14 microsatellite loci did not vary dramatically between populations and temporal replicates; it was the lowest in Lb-99 (AR = 5.07, HE = 0.578) and the highest in Vas-18 (AR = 6.50, HE = 0.697) (Table S3). The number of private alleles was similar during both earlier and later period samples and their frequencies were low (below 3%, except Strutta58 in Kun-19 with a frequency of 10.7%) (data not shown).
Spatio-temporal genetic variation and differentiation
Genetic diversity estimates, expressed as mean allelic richness (AR) and expected heterozygosity (HE), were on an average a little higher in the later period samples (mean AR = 5.8, range 5.1–6.5; mean HE = 0.65, range 0.62–0.70) compared to the earlier period (mean AR = 5.6, range 5.1–5.9; mean HE = 0.64, range 0.58–0.67) (Table 1). However, the change in AR and HE was not significant (Wilcoxon signed-rank test P > 0.05 and two-sided test P > 0.05; Goudet 1995). Genetic differentiation, measured as FST, among year-classes from different populations, was low (global FST = 0.029) with pairwise estimates of FST ranging from 0.004 (between Sel-17 and Kun-98) to 0.075 (between Lb-99 and Sel-19) (Table 2). Among periods, the level of differentiation between year-classes of the studied populations was higher in the earlier period (1996–2001, average pairwise FST = 0.035) than in the later period (2017–2019, average pairwise FST = 0.021) (Table 2). However, the change based on average FST estimates was not significant (two-sided tests P > 0.05; Goudet 1995). Genetic differentiation between year-classes from different populations showed significant estimates in 106 out of 119 pairwise comparisons (89.1%). All but one non-significant comparison (Vas-98 and Sel-19) involved the River Kunda samples. Comparisons between year-classes within the same river revealed significant differentiation in 21 out of 34 tests (61.8%). The largest temporal changes occurred in the Loobu and Selja populations (average pairwise FST = 0.031), while the Kunda population was more stable (average pairwise FST = 0.015) (Table 2). However, the changes between the later and earlier time period for each population (groups of year-classes) were not significant (two-sided tests P > 0.05; Goudet 1995). The hierarchical gene diversity analysis using AMOVA revealed that the highest proportion of variation was present within year-classes (FST = 0.026; P < 0.001), followed by variation among year-classes (FSC = 0.022; P < 0.001) and among populations (FCT = 0.004; P < 0.001) (Table 3). However, when both time periods were analysed separately, the relationships between temporal and spatial variation differed. For example, during the earlier period (1996–2001) temporal variation within populations (FSC = 0.023; P < 0.001) was almost four times higher than spatial differentiation (FCT = 0.006; P < 0.05). On the other hand, based on later samples (2017–2019), both temporal and spatial differentiation estimates were similar to each other (FSC = 0.010; P < 0.001; FCT = 0.012; P > 0.05) (Table 3). The AMOVAs, performed for two time periods separately, also indicated that genetic differentiation among populations was marginally higher during the earlier period (FST = 0.029, P < 0.001) compared to the later period (FST = 0.022, P < 0.001) (Table 3). The pattern of close genetic relationships among populations was further supported by a DA genetic distance-derived NJ dendrogram (Fig. 2). The NJ dendrogram also reflected the geographical relationships between populations (Fig. 2), with nearby Loobu, Selja, and Kunda samples grouping together (Fig. 2). Similarly, to River Vasalemma, all temporal replicates from the River Loobu grouped together, while those of R. Selja and R. Kunda were located on different branches of the tree.
Unrooted neighbour-joining dendrogram based on DA genetic (Nei et al. 1983) distances, demonstrating the genetic relationships between temporally replicated samples of four Estonian sea trout populations in the Gulf of Finland in Northern Estonia, the Baltic Sea. Temporally replicated samples of the same population are represented by the same colour and shape (green-square, yellow-triangle, red-circle, and blue-rhombus). The number on the node indicates branches with bootstrap support > 50% in 1,000 replicates. See Table 1 for the full names of the populations. (Colour figure online)
Effective population size, effective number of breeders, and migration rate estimates
The single sample estimates of the effective number of breeders (Nb) were variable between year-classes and rivers, ranging from 19.3 to 169.8 based on the linkage disequilibrium (LD) method and from 16 to 69 for the sibship assignment (SA) method (Table 1). Despite both methods being based on different assumptions and methodologies, a strong positive correlation between Nb estimates derived from the LD and the SA methods (R2 = 0.752, P < 0.05) indicates the consistency of these estimates. For the Vasalemma, Loobu, and Selja populations, we observed a tendency of increasing Nb estimates over time using both the LD and the SA methods (Table 1). The increase of Nb estimates coincided with the increase of estimated 0 + sea trout parr densities based on national monitoring data (Fig. 1).
When the potential immigration from hatchery releases across the whole study period was taken into account, the Ne estimates were similar for R. Loobu and R. Selja (range Ne = 37.2–62.3) for all four potential sources of immigration. On the other hand, the Ne estimate for R. Kunda and R. Vasalemma varied depending on the source of putative immigration. In three out of four different scenarios of immigration, the Ne estimates for the R. Kunda population were similar to the R. Loobu and R. Selja (range Ne = 63.2–78.8) and the Ne estimates for R. Vasalemma were mostly 2–4 times higher (range Ne = 102.3–145.1) (Table 4). The estimates of immigration rate (m) also varied among the studied rivers and depended on the putative immigration source. Considering the studied populations separately, and taking into account all putative sources of immigration from hatchery releases, the highest estimates of m were observed in the R. Loobu and R. Selja (range m = 0.122–0.458 and m = 0.122–0.251, respectively) (Table 4). Variable m estimates were observed for R. Vasalemma and R. Kunda. For R. Vasalemma, the estimated impact of hatchery immigration was low, or even absent, from source 1 and source 2, while it was higher from sources 3 and 4 (m = 0.171 and m = 0.124, respectively). For R. Kunda, the highest influx of immigrants originated from putative sources 3 and 2 (m = 0.171 and m = 0.224, respectively), and the lowest from putative hatchery sources 1 and 4 (m = 0.087 and m = 0.066, respectively) (Table 4).
Recent bottleneck events
The mode-shift test did not reveal a shift from the L-shaped allele frequency distribution for any year-class of any studied population, suggesting a mutation-drift equilibrium (data not shown). Similarly, Wilcoxon sign-rank tests, conducted using both SMM and TMP models, did not reveal evidence for recent population bottlenecks (all P-values > 0.05).
Discussion
We evaluated the changes in the spatio-temporal population structure among four sea trout populations in northern Estonia over two decades. Our results demonstrate weak genetic structuring among populations, a tendency for reduction of genetic differentiation, and an increased level of genetic diversity (heterozygosity and allelic richness) over time. Furthermore, we found evidence that both direct and indirect (via straying) effects of stocking most likely have influenced the genetic make-up of the studied wild trout populations. Our results demonstrate the dynamic nature of the population genetic structure of sea trout and suggest that hatchery releases have significant but variable effects on the genetic composition of wild populations.
Spatio-temporal variation
Analysis of the microsatellite DNA variation in four Estonian sea trout populations revealed a level of genetic diversity (14 loci; average AR = 5.67, average HE = 0.65; Table 1 and Table S3) comparable with that found in a survey of microsatellite variation in other Baltic Sea trout rivers, e.g. within the Luga River, Gulf of Finland (six common loci with our study; average AR = 3.93, average HE = 0.70; Lehtonen et al. 2009), among Lithuanian (seven common loci; average AR = 4.23, average HE = 0.64; Samuiloviene et al. 2009), Swedish (eight common loci; average AR = 6.22, average HE = 0.69; Östergren et al. 2016) and Polish populations (five common loci; average HE = 0.66; Was and Wenne 2003). Furthermore, the study demonstrated a low level of spatial structuring (global FST = 0.029) of the studied sea trout populations which is in concordance with earlier work on the spatial structuring of seven sea trout populations in the Gulf of Finland (FST = 0.022, Koljonen et al. (2014). Despite the low level of genetic divergence between populations, significant genetic differentiation was observed in 106 out of 119 pairwise comparisons. We also observed considerable temporal changes in allele frequencies over a period of 23 years (i.e., 1996–2019). This was supported by several lines of evidence. First, the hierarchical analysis of molecular variance (AMOVA) showed that temporal variation observed over a period of four to six generations between temporal replicates was high in comparison to spatial differences; temporal variation explained approximately five times more genetic variation (2.20%) than the spatial variation (0.46%). Moreover, the level of temporal variation changed over time. During the earlier period (1996–2001), temporal variation within populations was four times higher than the spatial differentiation, whereas the variance explained by the temporal and spatial components was more similar in the later samples (2017–2019). Secondly, the allele frequency changes over time were evident from significant genetic differentiation between year-classes of the Vasalemma, Loobu, and Selja populations, while only R. Kunda population was genetically more stable. Thirdly, temporal changes were particularly evident in the NJ dendrogram for the Selja and Kunda populations (Fig. 2), where year-classes showed no tendency for clustering together. For Vasalemma and Loobu populations, year-classes of the later period showed more close genetic relationships and were clearly separated from those collected during the earlier period (Fig. 2). Overall, significant temporal variation observed in the studied trout populations is in accordance with several earlier studies of resident and anadromous brown trout (Laikre et al. 2002; Ostergaard et al. 2003; Jensen et al. 2005; Lehtonen et al. 2009). Laikre et al. (2002) and Ostergaard et al. (2003) for example showed that populations inhabiting a highly unstable environment experience considerable temporal genetic changes because of genetic drift and frequent population turnover. Similarly, temporal fluctuations in allele frequencies have been observed in small Danish trout populations (Jensen et al. 2005) and in smaller tributaries of larger river systems (Lehtonen et al. 2009). However, the observed temporal genetic changes in the sea trout populations studied by us cannot be explained by an unstable environment since both Baltic salmon and sea trout breeders successfully reproduce annually in these rivers and thus, the observed changes can be explained by direct or indirect impacts of hatchery releases rather than variable environment (see below).
The pattern of higher temporal variation during the earlier period and a tendency for increased effective population size estimates in the later period for all rivers except R. Kunda indicates that the number of spawners most likely has increased in time and this is also reflected by an increase in juvenile densities in the studied rivers (ICES 2019, 2020). However, despite generally lower effective population size estimates in the earlier period, we didn’t find significant genetic bottleneck signals in any of the analyzed samples. In terms of analysis power, our set of 14 highly variable microsatellite loci is expected to be sufficient for the detection of a drastic decrease in effective population size, since simulations have shown that using 10–20 polymorphic loci and at least 30 individuals should be sufficient to avoid unreasonably high type 1 error rates using mode-shift distortion test for genetic bottlenecks (Piry et al. 1999). Thus, it is likely that the number of spawners during the earlier time period has not been small enough to yield drastic declines in genetic diversity and an increase in genetic divergence between populations because of random genetic drift. Instead, we observed a small decrease in population divergence estimates over time which suggests that the gene flow (immigration of hatchery-reared fish) may have homogenized the among-population divergence over time.
The effective number of breeders
We used two different single-sample approaches to quantify Nb, the linkage disequilibrium (LD) method proposed by Hill (1981) and the sibship assignment (SA) approach developed by Jones and Wang (2010) to better understand the population dynamics of studied trout populations. The Nb estimates for both single-sample approaches used here were rather small (average Nb = 51.2 for LD, and average Nb = 33.6 for the SA), being comparable with the estimated annual number of ascending trout in the nearby river (26 to 125 spawners counted in R. Pirita in 2014–2019; ICES 2020). Similarly, low Nb estimates have been observed among resident and anadromous brown trout, collected in different spawning and nursery grounds in Estonia (Ozerov et al. 2015), in small streams supporting wild brown trout in Norway (Serbezov et al. 2012), as well as in Bornholm (Ostergaard et al. 2003) and Gotland (Laikre et al. 2002). Furthermore, Nb estimates varied substantially from year to year but generally, we observed a tendency of increasing Nb over time. Thus, higher temporal differentiation compared to the spatial component likely reflects the smaller Nb and stronger effect of genetic drift in the earlier time period. In contrast, more stable genetic allele frequencies during the later period likely reflect the reduced level of genetic drift associated with an increase in population size or alternatively, the increased effect of gene flow from hatchery releases.
Impact of stocking
Large-scale stocking using hatchery-reared sea trout into rivers flowing into the Gulf of Finland started at the beginning of the 2000s with the aim to support threatened populations in Estonia. Altogether, during this period, more than 700 thousand fry and older fish were stocked into rivers and streams, where the juvenile densities were low (Ministry of the Environment 2020; Põlula Fish Rearing Centre 2021). The impact of stocking of hatchery-reared fish on wild populations has been estimated in a number of studies and frequently associated with the negative consequences on the existing genetic structuring among wild salmonid populations, e.g. stocking often leads to genetic homogenization (Susnik et al. 2004; Eldridge and Naish 2007; Hansen et al. 2009, 2010; Ozerov et al. 2016) and loss of genetic variability and potential disruption of local adaptations (Eldridge et al. 2009; Jasper et al. 2013; Östergren et al. 2021). Based on genetic analyses across more than decades, we found support for both direct and indirect impacts of stocking on the level of genetic divergence among populations. Firstly, we found a tendency, albeit non-significant, for an increase of genetic diversity over time (Table 1 and Table S3) and observed moderate change in the composition of alleles (loss of 16 alleles and addition of new 18 alleles in later period samples compared to earlier period samples) over the 20-year period (data not shown). The most parsimonious explanation for the observed genetic change is the effect of introgression from hatchery-reared trout. Reared fish often possess reduced genetic diversity relative to wild fish (Ryman and Laikre 1991; Blanchet et al. 2008; Araki and Schmid 2010) and as a result, reduce the overall genetic variability of wild populations (Eldridge et al. 2009; Jasper et al. 2013). On the other hand, hatchery fish may also carry unique genetic variation not observed in wild populations (Verspoor 1998) leading to an increase in the genetic variability of wild populations (Marie et al. 2010; Lamaze et al. 2012; Ozerov et al. 2016; Östergren et al. 2021).
Simultaneously with temporal changes in diversity, a temporal decline in genetic divergence was observed between all pairwise population comparisons and also when two sampling periods were compared (average pairwise FST = 0.035 in 1996–2001 vs. FST = 0.021 in 2017–2019), indicating that contemporary wild populations have become genetically more similar to each other over time. This homogenization trend is likely caused by the hatchery releases as evident from the joint estimation of Ne and immigration rate (m) using Wang and Whitlock’s (2003) method. The highest estimated immigration rates from all four putative hatchery sources were observed in R. Selja and R. Loobu (range m = 0.122–0.458 and m = 0.122–0.251, respectively). On the other hand, the effect of hatchery releases was lower in R. Kunda and R. Vasalemma and in some cases, the 95% CI of immigration rate estimates for specific hatchery sources also included zero, suggesting a negligible effect. The high immigration rates to R. Selja are likely explained by direct stocking during the period 2001–2006 (> 63 000 reared juveniles) and relative proximity to other heavily stocked rivers (Fig. 1; Table S2). On the other hand, the high immigration rates of reared fish to R. Loobu are most likely related to the close proximity of heavily stocked rivers of Valgejõgi, Pudisoo, Mustoja, and others (> 370 000 introduced juveniles) since R. Loobu has been not stocked with-hatchery fish (Fig. 1; Table S2). Thus, our results suggest that hatchery releases associated with inaccurate homing and subsequent gene flow have played an important in influencing spatio-temporal genetic structure of studied trout populations. This is not surprising since hatchery-reared salmonids tend to show weaker homing ability than wild conspecifics (Quinn 1993; Schroeder et al. 2001; Jonsson et al. 2003; Jonsson and Jonsson 2006) leading to increased gene flow (Hansen and Mensberg 2009; Hansen et al. 2010; Perrier et al. 2011; Ozerov et al. 2016). Interestingly, we also observed a potential geographical signal on the estimated immigration rates. Among the four studied populations, the lowest immigration rates from putative hatchery sources were observed in R. Vasalemma (range m = 0.000–0.171), which has not been stocked during the last decades and is the most distant from the heavily stocked rivers in the Gulf of Finland. In addition, a larger effective population size of R. Vasalemma compared to other studied populations may act as a buffer against hatchery-origin immigration - populations with smaller effective population sizes are expected to be more prone to admixture with hatchery-reared fish than larger populations (Vasemägi et al. 2005; Östergren et al. 2021). The indirect effects of stocking associated with straying found in this study are similar to earlier findings in other salmonids. For example, Ozerov et al. (2016) found that hatchery introgression via straying has changed the genetic make-up of wild Atlantic salmon (Salmo salar L.) populations of the Gulf of Finland by reducing genetic divergence and increasing genetic diversity. Similar patterns of increase of genetic diversity due to stocking and straying have been observed among wild Atlantic salmon populations of the Gulf of Bothnia in the Baltic Sea (Vasemägi et al. 2005) and also in France (Perrier et al. 2013). Evidence for indirect stocking effects has been also reported for alpine brown trout populations inhabiting Hardangervidda National Park lakes in Norway (Sønstebø et al. 2008); indirect stocking was accounted for in four lakes, and the magnitude of impacts of stocking depended on the distance from the original location of stocking. Alterations of the genetic structure due to gene flow from hatchery-reared fish as a result of direct stocking (Hansen 2002; Eldridge and Naish 2007; Marie et al. 2010; Lamaze et al. 2012) have also been observed frequently in other salmonids.
In conclusion, analysis of samples covering a time span of more than 20 years demonstrated the dynamic nature of the population genetic structure among Baltic Sea trout populations highlighting the important role of random genetic drift and immigration from hatchery releases on trout gene pools. Furthermore, by demonstrating that hatchery releases also affect the genetic composition of untargeted populations via straying, our results have important implications for the conservation and management of wild trout populations. However, further studies using more extensive and informative genomic data sets (e.g. SNP panels, RAD-seq data) are still needed to evaluate the complex and potentially long-term effects of stocking hatchery-reared fish on the adaptive genetic variation and fitness of wild populations. Therefore, regular monitoring and application of genome-wide approaches are necessary to be able to assess the impacts of stocking on wild populations and inform adaptive management strategies. Also, considering that the majority (83%) of peer-reviewed publications (1970–2021) evaluating how hatchery salmonids affected wild salmonids reported adverse effects on diversity, productivity, and abundance of wild salmonids (review by McMillan et al. 2023), implementing limitations on stocking and focusing on other conservation strategies, such as habitat restoration and fisheries management, can help maintain the integrity of wild populations.
Data availability
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.
References
Aprahamian MW, Smith KM, McGinnity P, McKelvey S, Taylor J (2003) Restocking of salmonids - opportunities and limitations. Fish Res 62:211–227. https://doi.org/10.1016/s0165-7836(02)00163-7
Araki H, Schmid C (2010) Is hatchery stocking a help or harm? Evidence, limitations and future directions in ecological and genetic surveys. Aquac 308:S2–S11. https://doi.org/10.1016/j.aquaculture.2010.05.036
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (2004) GENETIX 4.05, logiciel sous windows pour la génétique des populations. Laboratoire Génome, populations, interactions, CNRS UMR 5000. Université de Montpellier II, Montpellier, France
Blanchet S, Paez DJ, Bernatchez L, Dodson JJ (2008) An integrated comparison of captive-bred and wild Atlantic salmon (Salmo salar): implications for supportive breeding programs. Biol Conserv 141:1989–1999. https://doi.org/10.1016/j.biocon.2008.05.014
Borrell YJ, Bernardo D, Blanco G, Vazquez E, Sanchez JA (2008) Spatial and temporal variation of genetic diversity and estimation of effective population sizes in Atlantic salmon (Salmo salar, L.) populations from Asturias (Northern Spain) using microsatellites. Conserv Genet 9:807–819. https://doi.org/10.1007/s10592-007-9400-5
Brookfield JFY (1996) A simple new method for estimating null allele frequency from heterozygote deficiency. Mol Ecol 5:453–455. https://doi.org/10.1046/j.1365-294X.1996.00098.x
Cairney M, Taggart JB, Hoyheim B (2000) Characterization of microsatellite and minisatellite loci in Atlantic salmon (Salmo salar L.) and cross-species amplification in other salmonids. Mol Ecol 9:2175–2178. https://doi.org/10.1046/j.1365-294X.2000.105312.x
Campos JL, Posada D, Caballero P, Moran P (2007) Spatio-temporal genetic variability in sea trout (Salmo trutta) populations from north-western Spain. Freshw Biol 52:510–524. https://doi.org/10.1111/j.1365-2427.2006.01721.x
Campos JL, Posada D, Moran P (2008) Introgression and genetic structure in northern Spanish Atlantic salmon (Salmo salar L.) populations according to mtDNA data. Conserv Genet 9:157–169. https://doi.org/10.1007/s10592-007-9318-y
Christensen C, Jacobsen MW, Nygaard R, Hansen MM (2018) Spatiotemporal genetic structure of anadromous Arctic char (Salvelinus alpinus) populations in a region experiencing pronounced climate change. Conserv Genet 19:687–700. https://doi.org/10.1007/s10592-018-1047-x
Degerman E, Leonardsson K, Lundqvist H (2012) Coastal migrations, temporary use of neighbouring rivers, and growth of sea trout (Salmo trutta) from nine northern Baltic Sea rivers. ICES J Mar Sci 69:971–980. https://doi.org/10.1093/icesjms/fss073
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (ne) from genetic data. Mol Ecol Resour 14:209–214. https://doi.org/10.1111/1755-0998.12157
Eldridge WH, Naish KA (2007) Long-term effects of translocation and release numbers on fine-scale population structure among coho salmon (Oncorhynchus kisutch). Mol Ecol 16:2407–2421. https://doi.org/10.1111/j.1365-294X.2007.03271.x
Eldridge WH, Myers JM, Naish KA (2009) Long-term changes in the fine-scale population structure of coho salmon populations (Oncorhynchus kisutch) subject to extensive supportive breeding. Hered 103:299–309. https://doi.org/10.1038/hdy.2009.69
Estoup A, Presa P, Krieg F, Vaiman D, Guyomard R (1993) (CT)n and (GT)n microsatellites: a new class of genetic markers for Salmo trutta L. (brown trout). Hered 71:488–496. https://doi.org/10.1038/hdy.1993.167
Estoup A, Rousset F, Michalakis Y, Cornuet JM, Adriamanga M, Guyomard R (1998) Comparative analysis of microsatellite and allozyme markers: a case study investigating microgeographic differentiation in brown trout (Salmo trutta). Mol Ecol 7:339–353. https://doi.org/10.1046/j.1365-294X.1998.00362.x
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes - application to human mitochondrial - DNA restriction data. Genetics 131:479–491. https://doi.org/10.1093/genetics/131.2.479
Fleming IA (1996) Reproductive strategies of Atlantic salmon: Ecol Evol. Rev Fish Biol Fish 6:379–416. https://doi.org/10.1007/bf00164323
Frankham R, Ballou SEJD, Briscoe DA, Ballou JD (2002) Frantmatter. In: Introduction to conservation genetics. Cambridge: Cambridge University Press
Goudet J (1995) FSTAT (Version 1.2): a computer program to calculate F-Statistics. J Hered 86:485–486. https://doi.org/10.1093/oxfordjournals.jhered.a111627
Gudmundsson LA, Gudjonsson S, Marteinsdottir G, Scarnecchia DL, Danielsdottir AK, Pampoulie C (2013) Spatio-temporal effects of stray hatchery-reared Atlantic salmon Salmo salar on population genetic structure within a 21 km-long Icelandic river system. Conserv Genet 14:1217–1231. https://doi.org/10.1007/s10592-013-0510-y
Guo SW, Thompson EA (1992) Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics 48:361–372. https://doi.org/10.2307/2532296
Hansen MM (2002) Estimating the long-term effects of stocking domesticated trout into wild brown trout (Salmo trutta) populations: an approach using microsatellite DNA analysis of historical and contemporary samples. Molec Ecol 11:1003–1015. https://doi.org/10.1046/j.1365-294X.2002.01495.x
Hansen MM, Mensberg KLD (2009) Admixture analysis of stocked brown trout populations using mapped microsatellite DNA markers: indigenous trout persist in introgressed populations. Biol Lett 5:656–659. https://doi.org/10.1098/rsbl.2009.0214
Hansen MM, Nielsen EE, Ruzzante DE, Bouza C, Mensberg KLD (2000) Genetic monitoring of supportive breeding in brown trout (Salmo trutta L.), using microsatellite DNA markers. Can J Fish Aquat Sci 57:2130–2139. https://doi.org/10.1139/cjfas-57-10-2130
Hansen MM, Nielsen EE, Bekkevold D, Mensberg KLD (2001a) Admixture analysis and stocking impact assessment in brown trout (Salmo trutta), estimated with incomplete baseline data. Can J Fish Aquat Sci 58:1853–1860. https://doi.org/10.1139/cjfas-58-9-1853
Hansen MM, Ruzzante DE, Nielsen EE, Mensberg KLD (2001b) Brown trout (Salmo trutta) stocking impact assessment using microsatellite DNA markers. Ecol Appl 11:148–160. https://doi.org/10.2307/3061063
Hansen MM, Ruzzante DE, Nielsen EE, Bekkevold D, Mensberg KLD (2002) Long-term effective population sizes, temporal stability of genetic composition and potential for local adaptation in anadromous brown trout (Salmo trutta) populations. Mol Ecol 11:2523–2535. https://doi.org/10.1046/j.1365-294X.2002.01634.x
Hansen MM, Bekkevold D, Jensen LF, Mensberg KLD, Nielsen EE (2006) Genetic restoration of a stocked brown trout Salmo trutta population using microsatellite DNA analysis of historical and contemporary samples. J Appl Ecol 43:669–679. https://doi.org/10.1111/j.1365-2664.2006.01185.x
Hansen MM, Fraser DJ, Meier K, Mensberg KLD (2009) Sixty years of anthropogenic pressure: a spatio-temporal genetic analysis of brown trout populations subject to stocking and population declines. Mol Ecol 18:2549–2562. https://doi.org/10.1111/j.1365-294X.2009.04198.x
Hansen MM, Meier K, Mensberg KLD (2010) Identifying footprints of selection in stocked brown trout populations: a spatio-temporal approach. Mol Ecol 19:1787–1800. https://doi.org/10.1111/j.1365-294X.2010.04615.x
HELCOM (2011) Salmon and Sea Trout populations and rivers in the Baltic Sea – HELCOM assessment of salmon (Salmo salar) and sea trout (Salmo trutta) populations and habitats in rivers flowing to the Baltic Sea, vol 126A. Baltic Sea Environment Proceedings, Helsinki Commision, p 79
HELCOM (2013) HELCOM Red List of Baltic Sea species in danger of becoming extinct, vol 140. Baltic Sea Environment Proceedings, Helsinki Commission, p 106
Hill WG (1981) Estimation of effective population-size from data on linkage disequilibrium. Genet Res 38:209–216. https://doi.org/10.1017/s0016672300020553
ICES (2020) Baltic Salmon and Trout Assessment Working Group (WGBAST). ICES Scientific Reports. 2:22. 261 pp. https://doi.org/10.17895/ices.pub.5974
ICES (2019) Baltic Salmon and Trout Assessment Working Group (WGBAST). ICES Scientific Reports. 1:23. 312 pp. https://doi.org/10.17895/ices.pub.4979
Jasper JR, Habicht C, Moffitt S, Brenner R, Marsh J, Lewis B, Fox EC, Grauvogel Z, Olive SDR, Grant WS (2013) Source-Sink estimates of genetic Introgression Show influence of Hatchery Strays on Wild Chum Salmon populations in Prince William Sound, Alaska. PLoS ONE 8. https://doi.org/10.1371/journal.pone.0081916
Jensen LF, Hansen MM, Carlsson J, Loeschcke V, Mensberg KLD (2005) Spatial and temporal genetic differentiation and effective population size of brown trout (Salmo trutta, L.) in small Danish rivers. Conserv Genet 6:615–621. https://doi.org/10.1007/s10592-005-9014-8
Jones OR, Wang JL (2010) COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour 10:551–555. https://doi.org/10.1111/j.1755-0998.2009.02787.x
Jonsson B, Jonsson N (2006) Cultured Atlantic salmon in nature: a review of their ecology and interaction with wild fish. ICES J Mar Sci 63:1162–1181. https://doi.org/10.1016/j.icesjms.2006.03.004
Jonsson B, Jonsson N, Hansen LP (2003) Atlantic salmon straying from the River Imsa. J Fish Biol 62:641–657. https://doi.org/10.1046/j.1095-8649.2003.00053.x
Jorde PE, Ryman N (1995) Temporal allele frequency change and estimation of effective size in populations with overlapping generations. Genetics 139:1077–1090. https://doi.org/10.1093/genetics/139.2.1077
Kesler M, Svirgsden R, Taal I (2021) Kalanduse riikliku andmekogumise programmi täitmine, teadusvaatlejate paigutamine Eesti lipu all sõitvatele kalalaevadele ning teadussoovituste koostamine kalavarude haldamiseks 2020–2021. aastal. Töövõtulepingu nr 4 – 1/20/3 lõpparuanne 2020.a. kohta. Osa: Lõhe ja meriforell, p 123 (in Estonian). Retrieved from https://envir.ee/elusloodus-looduskaitse/kalandus/uuringud-ja-aruanded. Accessed 3 April 2022
Klemetsen A, Amundsen PA, Dempson JB, Jonsson B, Jonsson N, O’Connell MF, Mortensen E (2003) Atlantic salmon Salmo salar L., brown trout Salmo trutta L. and Arctic Charr Salvelinus alpinus (L.): a review of aspects of their life histories. Ecol Freshw Fish 12:1–59. https://doi.org/10.1034/j.1600-0633.2003.00010.x
Koljonen ML, Gross R, Koskiniemi J (2014) Wild Estonian and Russian sea trout (Salmo trutta) in Finnish coastal sea trout catches: results of genetic mixed-stock analysis. Hereditas 151(6):177–195. https://doi.org/10.1111/hrd2.00070
Laikre L, Jorde PE, Ryman N (1998) Temporal change of mitochondrial DNA haplotype frequencies and female effective size in a brown trout (Salmo trutta) population. Evolution 52:910–915. https://doi.org/10.1111/j.1558-5646.1998.tb03716.x
Laikre L, Jarvi T, Johansson L, Palm S, Rubin JF, Glimsater CE, Landergren P, Ryman N (2002) Spatial and temporal population structure of sea trout at the Island of Gotland, Sweden, delineated from mitochondrial DNA. J Fish Biol 60:49–71. https://doi.org/10.1006/jfbi.2001.1809
Laikre L, Larsson LC, Palme A, Charlier J, Josefsson M, Ryman N (2008) Potentials for monitoring gene level biodiversity: using Sweden as an example. Biodivers Conserv 17:893–910. https://doi.org/10.1007/s10531-008-9335-2
Laikre L, Schwartz MK, Waples RS, Ryman N, Ge MWG (2010) Compromising genetic diversity in the wild: unmonitored large-scale release of plants and animals. Trends Ecol Evol 25:520–529. https://doi.org/10.1016/j.tree.2010.06.013
Laird PW, Zijderveld A, Linders K, Rudnicki MA, Jaenisch R, Berns A (1991) Simplified mammalian DNA isolation procedure. Nucleic Acids Res 19:4293–4293. https://doi.org/10.1093/nar/19.15.4293
Lamaze FC, Sauvage C, Marie A, Garant D, Bernatchez L (2012) Dynamics of introgressive hybridization assessed by SNP population genomics of coding genes in stocked brook charr (Salvelinus fontinalis). Mol Ecol 21:2877–2895. https://doi.org/10.1111/j.1365-294X.2012.05579.x
Langella O (1999) Populations 1.2.32: a population genetic software. Montpellier: Laboratoire Populations, g én étique et evolution. CNRS UPR9034. Available at http//bioinformatics.org/~tryphon/populations/
Lehtonen PK, Tonteri A, Sendek D, Titov S, Primmer CR (2009) Spatio-temporal genetic structuring of brown trout (Salmo trutta L.) populations within the River Luga, northwest Russia. Conserv Genet 10:281–289. https://doi.org/10.1007/s10592-008-9577-2
Marie AD, Bernatchez L, Garant D (2010) Loss of genetic integrity correlates with stocking intensity in brook charr (Salvelinus fontinalis). Mol Ecol 19:2025–2037. https://doi.org/10.1111/j.1365-294X.2010.04628.x
McConnell SK, Oreilly P, Hamilton L, Wright JN, Bentzen P (1995) Polymorphic microsatellite loci from Atlantic salmon (Salmo salar) - genetic differentiation of North-American and European populations. Can J Fish Aquat Sci 52:1863–1872. https://doi.org/10.1139/f95-779
McMillan JR, Morrison B, Chambers N, Ruggerone G, Bernatchez L, Stanford J, Neville H (2023) A global synthesis of peer-reviewed research on the effects of hatchery salmonids on wild salmonids. Fish Manag Ecol 00:1–18. https://doi.org/10.1111/fme.12643
Ministry of the Environment (2020) Ministry of the Environment, Republic of Estonia website. https://envir.ee/elusloodus-looduskaitse/kalandus/kalade-asustamine. Accessed 22 November 2020
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data. 2. Gene frequency data. J Mol Evol 19:153–170. https://doi.org/10.1007/bf02300753
Nielsen EE, Hansen MM (2008) Waking the dead: the value of population genetic analyses of historical samples. Fish Fish 9:450–461. https://doi.org/10.1111/j.1467-2979.2008.00304.x
Oreilly PT, Hamilton LC, McConnell SK, Wright JM (1996) Rapid analysis of genetic variation in Atlantic salmon (Salmo salar) by PCR multiplexing of dinucleotide and tetranucleotide microsatellites. Can J Fish Aquat Sci 53:2292–2298. https://doi.org/10.1139/f96-192
Ostergaard S, Hansen MM, Loeschcke V, Nielsen EE (2003) Long-term temporal changes of genetic composition in brown trout (Salmo trutta L.) populations inhabiting an unstable environment. Mol Ecol 12:3123–3135. https://doi.org/10.1046/j.1365-294X.2003.01976.x
Östergren J, Nilsson J, Lundqvist H, Dannewitz J, Palm S (2016) Genetic baseline for conservation and management of sea trout in the northern Baltic Sea. Conserv Genet 17:177–191. https://doi.org/10.1007/s10592-015-0770-9
Östergren J, Palm S, Gilbey J, Spong G, Dannewitz J, Konigsson H, Persson J, Vasemägi A (2021) A century of genetic homogenization in Baltic salmon-evidence from archival DNA. Proc Royal Soc B 288. https://doi.org/10.1098/rspb.2020.3147
Ozerov MY, Veselov AE, Lumme J, Primmer CR (2013) Temporal variation of genetic composition in Atlantic salmon populations from the Western White Sea Basin: influence of anthropogenic factors? BMC Genet 14(15). https://doi.org/10.1186/1471-2156-14-88
Ozerov M, Jurgenstein T, Aykanat T, Vasemägi A (2015) Use of sibling relationship reconstruction to complement traditional monitoring in fisheries management and conservation of brown trout. Conserv Biol 29:1164–1175. https://doi.org/10.1111/cobi.12480
Ozerov MY, Gross R, Bruneaux M, Vaha JP, Burimski O, Pukk L, Vasemägi A (2016) Genomewide introgressive hybridization patterns in wild Atlantic salmon influenced by inadvertent gene flow from hatchery releases. Mol Ecol 25:1275–1293. https://doi.org/10.1111/mec.13570
Palm S, Laikre L, Jorde PE, Ryman N (2003) Effective population size and temporal genetic change in stream resident brown trout (Salmo trutta, L). Conserv Genet 4:249–264. https://doi.org/10.1023/a:1024064913094
Park SDE (2001) Trypanotolerance in West African cattle and the population structure. PhD dissertation. University of Dublin
Parrish DL, Behnke RJ, Gephard SR, McCormick SD, Reeves GH (1998) Why aren’t there more Atlantic salmon (Salmo salar)? Can J of Fish Aquat Sci 55:281–287. https://doi.org/10.1139/d98-012
Paterson S, Piertney SB, Knox D, Gilbey J, Verspoor E (2004) Characterization and PCR multiplexing of novel highly variable tetranucleotide Atlantic salmon (Salmo salar L.) microsatellites. Mol Ecol Notes 4:160–162. https://doi.org/10.1111/j.1471-8286.2004.00598.x
Pedersen S, Heinimaa P, Pakarinen T (eds) (2012) Workshop on Baltic Sea Trout, Helsinki, Finland, 11–13 October 2011. DTU Aqua. DTU Aqua report No. 248–2012
Perrier C, Guyomard R, Bagliniere JL, Evanno G (2011) Determinants of hierarchical genetic structure in Atlantic salmon populations: environmental factors vs. anthropogenic influences. Mol Ecol 20:4231–4245. https://doi.org/10.1111/j.1365-294X.2011.05266.x
Perrier C, Guyomard R, Bagliniere JL, Nikolic N, Evanno G (2013) Changes in the genetic structure of Atlantic salmon populations over four decades reveal substantial impacts of stocking and potential resiliency. Ecol Evol 3:2334–2349. https://doi.org/10.1002/ece3.629
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90:502–503. https://doi.org/10.1093/jhered/90.4.502
Pocwierz-Kotus A, Bernas R, Debowski P, Kent MP, Lien S, Kesler M, Titov S, Leliuna E, Jespersen H, Drywa A, Wenne R (2014) Genetic differentiation of southeast Baltic populations of sea trout inferred from single nucleotide polymorphisms. Anim Genet 45:96–104. https://doi.org/10.1111/age.12095
Põlula Fish Rearing Centre (2021) State Forest Management Centre website, Estonia. https://www.rmk.ee/organisation/polula-fish-rearing-centre/restocking-juvenile-fish. Accessed 03 December 2021
Poteaux C, Bonhomme F, Berrebi P (1999) Microsatellite polymorphism and genetic impact of restocking in Mediterranean brown trout (Salmo trutta L). Hered 82:645–653. https://doi.org/10.1046/j.1365-2540.1999.00519.x
Presa P, Guyomard R (1996) Conservation of microsatellites in three species of salmonids. J Fish Biol 49:1326–1329. https://doi.org/10.1111/j.1095-8649.1996.tb01800.x
Pritchard VL, Erkinaro J, Kent MP, Niemela E, Orell P, Lien S, Primmer CR (2016) Single nucleotide polymorphisms to discriminate different classes of hybrid between wild Atlantic salmon and aquaculture escapees. Evol Appl 9:1017–1031. https://doi.org/10.1111/eva.12407
Quinn TP (1993) A review of homing and straying of wild and hatchery-produced salmon. Fish Res 18:29–44. https://doi.org/10.1016/0165-7836(93)90038-9
Rannak L, Arman J, Kangur M (1983) Lõhe ja meriforell. Valgus, Tallinn, p 152 (in Estonian)
Raymond M, Rousset F (1995) GENEPOP (Version-1.2) - Population-Genetics Software for exact tests and ecumenicism. J Hered 86:248–249. https://doi.org/10.1093/oxfordjournals.jhered.a111573
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225. https://doi.org/10.2307/2409177
Rousset F (2008) GENEPOP ' 007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8:103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Ryman N, Laikre L (1991) Efects of supportive breeding on the genetically effective population size. Conserv Biol 5:325–329. https://doi.org/10.1111/j.1523-1739.1991.tb00144.x
Saitou N, Nei M (1987) The neighbor-joining method - a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Samuiloviene A, Kontautas A, Gross R (2009) Genetic diversity and differentiation of sea trout (Salmo trutta) populations in Lithuanian rivers assessed by microsatellite DNA variation. Fish Physiol Biochem 35:649–659. https://doi.org/10.1007/s10695-009-9310-1
Sandlund OT, Karlsson S, Thorstad EB, Berg OK, Kent MP, Norum ICJ, Hindar K (2014) Spatial and temporal genetic structure of a river- resident Atlantic salmon (Salmo salar) after millennia of isolation. Ecol Evol 4:1538–1554. https://doi.org/10.1002/ece3.1040
Schroeder RK, Lindsay RB, Kenaston KR (2001) Origin and straying of hatchery winter steelhead in Oregon coastal rivers. Trans Am Fish Soc 130:431–441. https://doi.org/10.1577/1548-8659(2001)130<0431:oasohw>2.0.co;2
Scribner KT, Gust JR, Fields RL (1996) Isolation and characterization of novel salmon microsatellite loci: cross-species amplification and population genetic applications. Can J Fish Aquat Sci 53:833–841. https://doi.org/10.1139/cjfas-53-4-833
Serbezov D, Jorde PE, Bernatchez L, Olsen EM, Vollestad LA (2012) Short-term genetic changes: evaluating effective population size estimates in a comprehensively described Brown Trout (Salmo trutta) population. Genetics 191:579–592. https://doi.org/10.1534/genetics.111.136580
Slettan A, Olsaker I, Lie O (1995) Atlantic salmon, Salmo salar, microsatellites at the SSOSL25, SSOSL85, SSOSL311, SSOSL417 loci. Anim Genet 26:281–282. https://doi.org/10.1111/j.1365-2052.1995.tb03262.x
Slettan A, Olsaker I, Lie O (1996) Polymorphic Atlantic salmon, Salmo salar L., microsatellites at the SSOSL438, SSOSL439 and SSOSL444 loci. Anim Genet 27:57–58. https://doi.org/10.1111/j.1365-2052.1996.tb01180.x
Sønstebø JH, Borgstrom R, Heun M (2008) Genetic structure in alpine brown trout Salmo trutta L. shows that indirect stocking affects native lake populations. J Fish Biol 72:1990–2001. https://doi.org/10.1111/j.1095-8649.2008.01815.x
Susnik S, Berrebi P, Dovc P, Hansen MM, Snoj A (2004) Genetic introgression between wild and stocked salmonids and the prospects for using molecular markers in population rehabilitation: the case of the Adriatic grayling (Thymallus thymallus L. 1785). Hered 93:273–282. https://doi.org/10.1038/sj.hdy.6800500
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tessier N, Bernatchez L (1999) Stability of population structure and genetic diversity across generations assessed by microsatellites among sympatric populations of landlocked Atlantic salmon (Salmo salar L). Mol Ecol 8:169–179. https://doi.org/10.1046/j.1365-294X.1999.00547.x
Valiquette E, Perrier C, Thibault I, Bernatchez L (2014) Loss of genetic integrity in wild lake trout populations following stocking: insights from an exhaustive study of 72 lakes from Quebec, Canada. Evol Appl 7:625–644. https://doi.org/10.1111/eva.12160
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Vasemägi A, Gross R, Paaver T, Koljonen ML, Nilsson J (2005) Extensive immigration from compensatory hatchery releases into wild Atlantic salmon population in the Baltic Sea: spatio-temporal analysis over 18 years. Heredity 95:76–83. https://doi.org/10.1038/sj.hdy.6800693
Verspoor E (1998) Genetic impacts on wild Atlantic salmon (Salmo salar L.) stocks from escaped farm conspecifics: An assessment risk. In: Canadian Stock Assessment Secretariat Research Document 98/156 20p., Canadian Stock Assessment Secretariat, Halifax, NS
Wang JL (2009) A new method for estimating effective population sizes from a single sample of multilocus genotypes. Mol Ecol 18:2148–2164. https://doi.org/10.1111/j.1365-294X.2009.04175.x
Wang JL, Whitlock MC (2003) Estimating effective population size and migration rates from genetic samples over space and time. Genetics 163:429–446. https://doi.org/10.1093/genetics/163.1.429
Waples RS, Anderson EC (2017) Purging putative siblings from population genetic data sets: a cautionary view. Mol Ecol 26:1211–1224. https://doi.org/10.1111/mec.14022
Waples RS, Luikart G, Faulkner JR, Tallmon DA (2013) Simple life-history traits explain key effective population size ratios across diverse taxa. Proc Royal Soc B 280(9). https://doi.org/10.1098/rspb.2013.1339
Was A, Wenne R (2003) Microsatellite DNA polymorphism in intensely enhanced populations of sea trout (Salmo trutta) in the Southern Baltic. Mar Biotechnol 5:234–243. https://doi.org/10.1007/s10126-002-0068-z
Weir BS, Cockerham CC (1984) Estimating F-Statistics for the analysis of Population structure. Evolution 38:1358–1370. https://doi.org/10.2307/2408641
Yamamoto S, Morita K, Koizumi I, Maekawa K (2004) Genetic differentiation of white-spotted charr (Salvelinus leucomaenis) populations after habitat fragmentation: spatial-temporal changes in gene frequencies. Conserv Genet 5:529–538. https://doi.org/10.1023/B:COGE.0000041029.38961.a0
Acknowledgements
We thank Martin Kesler, the late Mart Kangur, Roland Svirgsten, and Imre Taal from the Estonian Marine Institute, University of Tartu, and Marje Aid, the late Tiit Paaver, Kerli Haugjärv, Magnus Lauringson from the Institute of Veterinary Medicine and Animal Sciences, Estonian University of Life Sciences for assistance with field sampling of Estonian sea trout populations, to Ene Saadre and others from the Põlula Fish Rearing Centre (State Forest Management Centre, Estonia) for providing samples of hatchery broodstock of sea trout. We gratefully acknowledge Ene Saadre for helpful information on Estonian sea trout stocking history. We thank David Arney for language corrections.
Funding
This study was funded by the Estonian Ministry of Education and Research (institutional research funding project IUT8-2), Estonian Research Council (grant PRG852), and Estonian Ministry of Environment (annual contracts for monitoring genetic diversity of salmonid populations in the Gulf of Finland).
Author information
Authors and Affiliations
Contributions
The study was planned and designed by R.G., O.B. and A.V. Samples were collected by R.G. and O.B. Laboratory and statistical analysis of data were performed by O.B. O.B. wrote the main manuscript and all authors reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Burimski, O., Vasemägi, A. & Gross, R. Changes in the spatio-temporal genetic structure of Baltic sea trout (Salmo trutta L.) over two decades: direct and indirect effects of stocking. Conserv Genet 25, 481–497 (2024). https://doi.org/10.1007/s10592-023-01582-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-023-01582-7