Abstract
Tomatoes are a major global food staple but Phytophthora infestans (an Oomycete) causes late-blight, a devastating disease that precludes commercial tomato production from moist temperate areas such as the United Kingdom and Northern Europe. We dissected the genetic architecture of resistance to late-blight as well as traits that improve yield and fruit quality in a tomato cross between a popular breeding, line NC 2 CELBR, which produces large fruits, and an heirloom cultivar called ‘Koralik’ which produces small, sweet fruits. We used an F2 mapping population to identify quantitative trait loci (QTL) for phenotypes including number of fruits, size of fruits, total crop yield, and soluble solids content in two different environments. Surprisingly, we found very few QTLs shared between the two environments, underscoring the importance of the local environment and genotype-by-environment interactions. We also assayed the virulence of three different isolates of P. infestans to identify QTLs that confer some resistance to the pathogen. We found nine crop-related QTLs and two QTLs for late-blight resistance-related phenotypes. One late-blight resistance QTL was inherited from Koralik (Chromosome 11, 70.2–83.5 cM) and it probably represents an undiscovered source of late-blight resistance. Yield QTLs were also located on chromosome 11 where Koralik alleles increase fruit number and yield, and adjacent regions decrease fruit size. On Chromosome 9, Koralik alleles increase fruit sweetness (Brix) by 25%. These results indicate that Koralik is a valuable donor parent that can be used by tomato breeders in targeted breeding strategies for fresh market tomatoes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tomatoes (Solanum lycopersicum L.) are a major food staple around the world with global annual production at over 177 tonnes in 2016 (www.fao.org/faostat), but due to pathogens present in temperate environments the vast majority are grown in arid regions or protected environments. Two broad groups of tomato cultivars exist; some intended for consumption raw and fresh and others intended for processing. Those intended for direct consumption tend to be smaller, sweeter, and have higher wet mass while tomatoes designed for processing have higher dry mass, contain less seed gel, and are better suited for canning and use in soups, sauces, and other cooked foods (Salunkhe and Kadam 1998). Typically, processing cultivars have fruit that ripens simultaneously while cultivars bred primarily for the fresh market or amateur growers ripen sequentially to allow picking over a longer season (Salunkhe and Kadam 1998).
Tomato crops are susceptible to a range of pathogens that can severely impact yield. One of the most severe pathogens is the water mould Phytophthora infestans (Oomycetes) which causes late-blight and can lead to complete crop failure (Fry 2008). Late-blight primarily affects crops in moist, temperate regions as it requires moisture on the leaves to complete its life cycle (Fry 2008). In arid regions such as California, tomatoes are only vulnerable to P. infestans infection during periods of rainfall at which times they are treated with fungicides (Hartz et al. 2008). In temperate regions, such as Northern Europe, commercial outdoor tomato production is rare due to the threat of infection. Today, all significant commercial tomato production in the U.K. takes place in greenhouses where moisture is tightly regulated to reduce the risk of infection (Heuvelink 2005). Thus there is little demand from commercial growers for blight-resistant tomatoes in the U.K. However, among amateur gardeners there is a large potential market for blight-resistant outdoor cultivars, as tomatoes are one of the most popular garden crops (Staub 2010). Therefore developing blight-resistant cultivars is of great interest to breeders whose target market is amateur gardeners.
A plant that carries innate genetic resistance to pathogens is preferable over one that requires fungicide. Fortunately, a number of loci have been found that confer resistance to late-blight and these can be capitalised upon by an informed breeding program. Major genes from wild relatives that confer race-specific resistance to P. infestans have been used in breeding: Ph-1 is no longer effective against current races (Foolad and Panthee 2012) whereas Ph-2 and Ph-3 (Foolad et al. 2008) remain effective against most races of the pathogen, especially when used together in hybrid cultivars such as Mountain Magic and Crimson Crush. Ph-2 has been mapped to chromosome 10 (Foolad and Panthee 2012) and Ph-3 to chromosome 9 (Chen et al. 2014; Robbins et al. 2010), and the gene underlying Ph-3 has been identified (Zhang et al. 2014). Additional loci have been designated as Ph-4, Ph-5.1 and Ph-5.2 (on chromosomes 2, 1 and 10 respectively) but they have not been widely used by breeders (reviewed by Stroud 2015).
To help inform breeding strategy for outdoor cultivars we used an experimental cross between NC 2 CELBR, commonly used as a parent of F1 hybrid cultivars, and the heirloom ‘Koralik’ to identify genomic regions that confer resistance to late-blight as well as regions that control various desirable fruit-related phenotypes. NC 2 CELBR is a tomato breeding line that is homozygous for the late-blight resistant alleles at both Ph-2 and Ph-3 (Gardner and Panthee 2010; Panthee et al. 2015). It grows as a vigorous, determinate bush type and fruits are around 100 grams. The heirloom cultivar Koralik is also a vigorous bush type that originated in Poland (Bralewski et al. 2006). It has a high yield of small, sweet fruits averaging 2.5 g. Koralik is homozygous resistant at the Ph-2 locus but does not carry any resistance alleles at Ph-3 (Stroud 2015). While exact breeding records are unavailable, the unusually vigorous, irregular plant habit and small fruit suggest that Koralik may be derived from a wild parent, Solanum pimpinellifolium or S. lycopersicum var. cerasiforme. High levels of P. infestans resistance exist in wild populations of both of these species, in many cases conferred by genes other than Ph-2 or Ph-3 (Foolad et al. 2014; Arellano Rodríguez et al. 2013) and so Koralik may carry one or more novel resistance genes. A cross between Koralik and NC 2 CELBR should segregate alleles for resistance as well as fruit size, number, and total crop yield; all traits that could be capitalised on when designing a new cultivar. Our goal was to identify any QTLs that explain variation in pathogen resistance and crop traits segregating within this cross.
Materials and methods
Crossing scheme
The crossing and experiments in two environments were carried out at the Henfaes Research Centre, U.K. (53°14′20.4″N 4°01′12.0″W). We bred an F2 mapping population by crossing three Koralik individuals with three NC 2 CELBR individuals and then selfing the F1 hybrids. All plants were housed in a climate-controlled greenhouse with 22 °C daytime temperature and a 7 °C minimum night-time temperature under a 16 h-light to 8 h-dark regimen and potted in Melcourt Silvamix potting compost (Melcourt Industries Limited, Tetbury). Parents were crossed in both directions such that each parental line contributed a male and female parent in different crosses. To avoid self-fertilization, anthers were removed before they began to shed pollen. Flowers were hand-pollinated by rubbing mature anthers on the exposed stigma of the emasculated flower. Once fruit had developed, the F1 seed was extracted from the fruit and cleaned of seed-gel with an 8 g/L sodium carbonate solution, incubated at room temperature (18–22 °C) for 24–48 h, washed, and dried at 40 °C for 24 h. The seed was then sown, and reciprocal F1 plants were grown over the autumn and winter and allowed to self-fertilise in the spring. F2 seed was harvested, cleaned, and sown and after approximately 3 weeks, 90 healthy seedlings were potted and allowed to grow through the winter. In spring, the 90 F2 lines were re-potted and grown on. Clones of each plant were made by collecting 3 to 4 side shoots from each plant which were rooted in tap water and grown on in 7L pots of compost. One clone from each F2 individual was moved into a greenhouse and another into a polytunnel where it was transplanted directly into the soil through a weed-suppressing membrane.
During growth, plants in the greenhouse were treated with Steinernema feltiae nematode (Nemasys, BASF plc, Cheadle Hulme) to combat fungus gnats, and sprayed weekly with SB Plant Invigorator (Fargro, Littlehampton, West Sussex) to treat powdery mildew (Oidium neolycopersici and Leveillula taurica). Plants in the poly-tunnel were allowed to grow unfertilised and not treated with pesticides nor stimulants.
Phenotyping
Ripe tomatoes were harvested fortnightly from both the greenhouse and the polytunnel and the number and total weight of the fruit crop was recorded. Average fruit size was calculated as the total weight of the crop divided by the total number of tomatoes harvested. The ripest tomato from each harvest was analysed for soluble solids content (Brix) using an Atago PAL-1 digital refractometer (Atago Co. Ltd., Tokyo) and these values were averaged across the growing season to find the typical Brix content for each F2 clone.
To assay for infection resistance, we harvested healthy and mature leaflets from F2 clones in the polytunnel as well as parental controls and a susceptible control. Leaf inoculations were done as described by Day and Shattock (1997), with eighteen leaflets harvested from each clone and exposed, in groups of six, to one of three isolates of P. infestans that occur in Great Britain: 6_A1, 8_A1, or 13_A2 (Cooke et al. 2012). At either 9- or 12- days post-inoculation (depending on the screen) the number of infected leaflets were recorded (expressed as percentage infection efficiency within the parental, control and the F2 population) and the diameter of all lesions was also recorded. For QTL analysis these data were converted into two metrics of late-blight resistance for each F2: infection efficiency (expressed as a binary absence/presence score where any signs of infection counted as presence, and only fully resistant individuals were scored as absent) and average lesion area (mm2), assuming all lesions were circular. Raw data for all F2 phenotypes can be found in Supplemental File 1. We were unable to record reliable infection phenotypes for Koralik and Moneymaker (the susceptible control) inoculated with isolates 6_A1 and 8_A1 because the leaves became contaminated in these tests.
Genotyping
Unexpanded, healthy leaflets were collected from the greenhouse-grown mature F2 plants and freeze-dried using an Edwards Modulyo K4 freeze-dryer and RV5 vacuum pump (Thermo-Fisher Scientific, Renfrew, Renfrewshire). Approximately 20 mg of freeze-dried leaf tissue was ground in a microfuge tube using a Qiagen Tissue Lyser beadmill (Qiagen, Crawley, Sussex). DNA extraction from the ground leaflet samples was carried out using a Qiagen DNEasy Plant Mini Kit (Qiagen, Crawley, Sussex) according to the manufacturer’s instructions. DNA concentration was measured using a NanoDrop 1000 spectrophotometer (NanoDrop Products, Wilmington, DE, USA). The concentration of the extracted DNA was adjusted to 50 ng μL−1 and genotyped by TraitGenetics GmbH (TraitGenetics GmbH, Gatersleben, Germany). Genotyping was carried out using the 7720 locus “SolCAP” SNP array (Sim et al. 2012a, b). This SNP array ties markers back to chromosomes in the SLv2.0 version of the SolCap (Fernandez-Pozo et al. 2014) genome available from https://solgenomics.net/organism/Solanum_lycopersicum/genome.
Genetic map
We used R/qtl (Broman and Sen 2009; Broman 2012) to create a genetic map for the cross. We dropped markers that were genotyped in less than 80 individuals, those with duplicate genotypes, and those with significant segregation distortion. Linkage groups were formed using a recombination frequency of 0.35 and a LOD cutoff of 10. This resulted in 12 linkage groups with between 15 and 56 markers and one which had only 4 markers and was discarded. Each linkage group was ordered with orderMarkers() using the ‘Haldane’ mapping function and the ripple() function was used to test the order. Finally the dropone() function was used to identify and remove internal markers that disproportionately expanded the map. Finally we compared the linkage groups with the physical map to identify chromosomes and verify that marker order was generally preserved between our map and the genome.
QTL mapping
Many phenotypic traits were significantly correlated with greenhouse position and so all non-binary traits were regressed against sample order using a simple linear regression and the residuals were used for QTL mapping. Traits were mapped using the Haley-Knott method (Haley and Knott 1992; Martínez and Curnow 1992) implemented in the scanone() function of R/qtl (Broman and Sen 2009) with the model set to ‘normal’ except for infection efficiencies where the model was ‘binary.’ Significance thresholds were calculated by 1000 permutations with scanone(). Once single QTL were identified, they were used as the start points for stepwiseqtl() which identifies additional additive and interactive (epistatic) QTLs by incrementally building up the model complexity to a set stopping point of 10 QTLs (5 for binary traits) and then incrementally removing levels of complexity back down to the null model of no QTLs. Likelihoods were calculated for each model and more complex models were penalised to avoid over-fitting. Each model is given a penalized LOD score (pLOD) which is the LOD score of the current model minus a penalisation based on the models’ complexity to avoid over-fitting. Due to the penalization correction there is no LOD cutoff. Rather the model with the highest pLOD score best explains the phenotype, thus we have reported all QTL models with a pLOD over 0 and models whose highest pLOD is less than 0 are scored as having no QTLs. Penalties were calculated by permuting scantwo() 1000 times and set so that the false discovery rate alpha is 0.05. Full QTL models from stepwiseqtl() were discarded if the penalised LOD scores increased exponentially at high numbers of QTLs as these are cases where we have too little power to identify true QTLs and over-fitting of the model resulted in extremely high LOD scores. We re-ran these few cases with a stopping point of 5 QTLs and verified that no true QTLs existed. Full models were subjected to one final test where we used fitqtl() to measure the improvement in the model when dropping each term. Individual QTLs whose marginal benefit was insignificant were dropped from the models. For each QTL, we ran an ANOVA on the phenotypes binned by the genotypes of the marker at the peak of the QTL, and then a Tukey HSD test to determine which genotype(s) were significant and whether the relationship between the alleles were overdominant, dominant, additive, recessive, or underdominant.
Data availability
The genetic map and raw genotyping data are available in Supplementary File 1.
Results
Genetic map
The final map is 1084.9 cM long, has 459 makers, and an average spacing of 2.4 cM. No significant segregation distortion was detected. The genetic map and all individual genotypes are found in Supplemental File 1. All linkage groups were anchored to the corresponding chromosome. Oddly, there was a significant amount of linkage between chromosomes 1 and 11 (Supplemental Figure 1). Despite the low recombination frequency and high LOD scores, we found it impossible to properly order the markers if merged into a single linkage group. As the markers originate from different chromosomes in the genome (Sim et al. 2012a) we decided to leave the two chromosomes separate and treat them independently. However this pattern suggests some sort of rearrangement, association, or nondisjunction between these two chromosomes and further karyotype work may prove illuminating.
QTLs for crop yield phenotypes
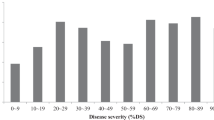
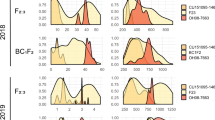
We tracked four metrics of crop productivity: number of tomatoes produced (Fruit Count), average weight of individual tomatoes (Fruit Size), total weight of all fruit (Crop Yield), and soluble solids content (Brix). Each of these were assayed under two different environments: a greenhouse and a polytunnel. The distributions of all phenotypes can be found in Fig. 1. We discovered a QTL for fruit count in each environment (Table 1, Fig. 2a). Surprisingly these were on different chromosomes and had opposite effect directions. In the tunnel, Koralik alleles at a QTL on chromosome 11 increased the fruit count by ~ 6 (ANOVA, F2,87 = 10.931, p = 5.8e − 5) while in the greenhouse Koralik alleles at chromosome 3 decreased the fruit count by ~ 10 (ANOVA, F2,87 = 12.658, p = 1.50e − 5, Fig. 2b). While they have effects in different directions, Tukey tests suggest that in both cases, the Koralik alleles acted dominantly. The best model for fruit size in the tunnel includes four independent QTLs in the tunnel (ANOVA, Chr 2: F2,81 = 18.249, p = 2.9e − 7, Chr 4: F2,81 = 8.035, p = 0.0066, Chr 9: F2,81 = 3.713, p = 0.0286, and Chr 11: F2,81 = 29.619, p = 2.2e − 10) while the best model for fruit size in the greenhouse has only one QTL (ANOVA, Chr 2: F2,87 = 11.005, p = 5.5e − 5, Table 1, Fig. 2a). A Tukey test suggests that for these QTLs, the Koralik alleles at the QTLs on chromosomes 2 and 4 act dominantly and decrease fruit size while the QTL on chromosome 9 may be underdominant (heterozygotes have the lowest fruit size) and the alleles at the QTL on chromosome 11 act additively in the tunnel but dominantly in the greenhouse (Fig. 2c). We found a single QTL for crop yield in the greenhouse on chromosome 11 (ANOVA, F2,87 = 11.047, p = 5.3e − 5, Table 1, Fig. 2a). Koralik alleles at this QTL acted recessively and increased the total crop yield by 75 grams (Fig. 2d). Finally, we found a single QTL for Brix in the tunnel on chromosome 9 (ANOVA, F2,86 = 14.372, p = 4.1e − 6,Table 1, Fig. 2a) where Koralik alleles acted dominantly and increased the Brix by 0.43 (Fig. 2e).
QTL and Effect Plots. (a) QTL locations and LOD confidence intervals. Only linkage groups with a QTL are plotted. (b–f) Violin plots are used to show the effect size and direction of QTLs in the tunnel (plots in left column) and greenhouse (right column) for residual Fruit Count (b), residual Fruit Size (c), residual total Crop Yield (d), residual Brix (e) and Infection efficiencies for isolates 6_A1 and 8_A1 (f). For all effect plots NC 2 CELBR homozygotes are on the left, heterozygotes are in the middle and Koralik homozygotes are on the right. The phenotypes for the effect plots are binned by the genotype at the peak of the QTL (see Table 1 for exact locations). Significance thresholds are determined by an ANOVA and a Tukey HSD test, a star and line indicates p < 0.05 for the pairwise comparison underscored by the line. A colour version of this figure is available online
QTLs for infection resistance phenotypes
The three different genotyped isolates (races) of late-blight (P infestans: 6_A1, 8_A1, and 13_A2) showed a marked difference in infection efficiency and large variation in lesion size in the parent strains (Table 2) and F2s (Fig. 1). While infection efficiency of 13_A2 was only 50% in Koralik, it was highly aggressive on the segregating population (only 4 F2 clones were uninfected, Fig. 1). Such high infectivity made it impossible to identify QTLs for resistance to 13_A2 (Fig. 1, Table 2). We found two QTLs for infection efficiency with the other isolates; one each for 6_A1 (ANOVA, F2,84 = 16.663, p = 8.0e − 7) and 8_A1 (ANOVA, F2,84 = 12.846, p = 1.4e − 5, Table 3, Fig. 2a). Both of these QTLs act recessively but explained a high amount of variance in resistance (20–30%). The allele that confers some resistance to 6_A1 comes from NC 2 CELBR while the allele that confers resistance to 8_A1 originates in Koralik (Fig. 2f). While lesion area exhibited a large variance in the F2 population (Fig. 1), we were unable to find any significant QTL models for any of the three isolates of late blight.
Discussion
We developed a tomato linkage map of 1084.9 cM from two inbred salad tomatoes. This map is broadly consistent with the linkage maps for three inter-specific F2 populations published by Sim et al. (2012b) who used the same SNP array. The main differences are some inversions within chromosomes, some short duplications, and segregation distortion on chromosomes 1, 10 and 11 in the previously published maps. Genetic and physical positions generally agree between all four maps, however, we detected a pattern suggesting a novel rearrangement between chromosomes 1 and 11 occurred in the cross. The parents of our map contain only small introgressed regions from S. pimpinellifolium so our map had limited interspecific regions yet the SNP array used for map construction revealed sufficient polymorphic loci (459) for mapping.
This F2 mapping population was used to discover either major genes or QTLs underlying both crop-related and disease-resistance phenotypes segregating in the cross. The parents differ for fruit size and number: Koralik has many, small, sweeter tomatoes while NC 2 CELBR produces fewer, large, and less sugary fruits. Nearly all of the QTLs we identified act in accordance with the parental expectations. For instance, Koralik alleles for fruit number on Chromosome 11 increase the number of fruit produced in the tunnel (Fig. 2b). The one exception is the QTL on Chromosome 3 where Koralik alleles tend to decrease fruit number in the greenhouse (Fig. 2c). Surprisingly, none of the QTL models that we identified for any trait involved epistatic interactions. This may be due to the small size of our mapping population leaving us underpowered to detect epistatic interactions.
Our results underscore the important role of the environment as we found no cases where the same genomic region explained a phenotype in both environments. Indeed in some cases we only found QTLs in one environment (i.e. Crop Yield in the greenhouse and Brix in the tunnel) and for the other phenotypes (fruit number and fruit size) QTLs found in the greenhouse do not appear to play an important role in the tunnel and visa-versa. Most intriguingly, we found four QTLs for fruit size in the tunnel, but only one in the greenhouse which does not overlap with any QTLs from the tunnel. These inconsistencies highlight the importance of the environment and genotype-by-environment interactions. The greenhouse grown plants suffered from insect pests and mildew which did not affect the plants in the tunnel. The greenhouse was warmer than the tunnel, which may have increased the pest prevalence. Plants in the greenhouse received a nematode addition to combat fungus gnats and a weekly spray with SB Plant Invigorator to treat powdery mildew. In addition, the greenhouse plants were grown in pots which required supplementary fertiliser, whereas in the tunnel the plants were grown directly in the ground, unfertilised and with neither pesticides nor stimulants. It is worth noting that the QTL for crop yield in the greenhouse on chromosome 11 overlaps with the QTL for fruit count in the tunnel, suggesting that Koralik alleles in this region continue to act on overall yield under increased pest pressure and therefore this region could be suitable for selection to increase yield stably across both environments.
The QTL we identified for Brix on chromosome 9 is linked to a marker (solcap_snp_sl_39722) that is positioned on the physical genome only 6.9 kb from a functional SNP within the Lycopersicum Invertase5 (LIN5) gene (Sauvage et al. 2014). LIN5 was identified as the gene underlying the QTL Brix9-2-5 (Fridman et al. 2004) and was found to control soluble solids content (Kühn et al. 2009), so our detection of a QTL for Brix that co-locates with Brix9-2-5 suggests that LIN5 is functioning in Koralik to increase Brix content.
The four QTLs for fruit size identified in the greenhouse (where Koralik alleles reduce fruit size) are all located in regions where QTLs for either fruit weight (fw2.1, fw2.2, fw2.3, fw4.2, fw9 and fw11.1) or fruit size (fs2.1 and fs2.2) have been mapped in at least two other studies (Grandillo et al. 1999). Of these, the regions on chromosomes 2, 9 and 11 are all associated with domestication sweeps (Lin et al. 2014), suggesting that NC 2 CELBR may contain many loci in these that were fixed during domestication and that crossing with Koralik can break some of these linkages and increase allelic diversity.
QTLs for late-blight resistance
We found alleles conferring late-blight resistance donated by both parents. There was much variation in susceptibility to different isolates of P. infestans both within and between parents and F2s (Table 2). We chose traits that may explain both whether the disease will establish and then once it does, how severely it will attack. We were unable to find any QTLs affecting the severity (lesion area), probably because our screen did not provide the resolution for minor QTL detection. However, we did find two QTLs that partially explain whether an individual became infected or not. Neither of these loci conferred absolute protection, but rather they decreased the chance of an infection establishing and are evidence, therefore, that both could be major genes conferring race -specific resistance. The resistance allele detected on chromosome 9 against isolate 6_A1 originated from NC 2 CELBR so is expected to be due to Ph-3 which is known to be segregating in our mapping population. The allele detected on chromosome 11 giving resistance to isolate 8_A1 originated from Koralik. To our knowledge, only one other Ph locus has been mapped on chromosome 11 (Ohlson et al. 2018) but it is not in the same region, so our QTL may thus represent a novel resistance locus.
Our study did not detect Ph-2 (chromosome 10), a finding that supports our previous (Stroud 2015) CAPS marker genotyping data which indicate that Koralik is homozygous for the Ph-2 resistance allele. Since NC 2 CELBR is well known to be homozygous for Ph-2 resistance alleles we can be confident that the Ph-2 locus is not segregating in our mapping population.
Other minor QTLs for late-blight resistance thought to derive from the same wild source as Ph-3 have been identified, including one on chromosome 2 (Chen et al. 2014) and one on chromosome 12 (Panthee et al. 2017). There are a number of reasons that could explain why we did not detect these: our mapping population was smaller, we used UK-derived not US-derived late-blight isolates for infection, or the resistant alleles are not present in either NC 2 CELBR or Koralik.
The great variation in infection status, even among individuals that carry one or both of the resistant alleles, suggests that the best breeding strategy for defence against late-blight may be to select progeny to carry the maximum combination of resistance alleles in the same cultivar (i.e. Ph-3, Ph-2 and the newly identified QTL on chromosome 11). In addition, breeders could combine them all with other recently mapped loci (Merk et al.2012; Ohlson et al. 2018; Arafa et al. 2017). Stacking a diverse range of resistance genes is especially appropriate when developing new cultivars for amateur gardeners given the high genetic variation harboured within the P. infestans population in gardens (Stroud et al. 2016). In addition, we found that the isolate 13_A2 (Cooke et al. 2012) was highly aggressive, supporting emerging reports that Ph-2 and Ph-3 are no longer effective on their own against some recently appearing, more aggressive isolates (Panthee et al. 2017; Merk et al. 2012) but they still contribute to slowing the disease if combined with other resistance loci. Koralik has been identified in this study as a useful parent in this approach because it contributes two late-blight resistance loci (a new QTL and Ph-2) as well as fruit sweetness and some yield component traits for breeding new outdoor salad tomato cultivars.
References
Arafa RA, Rakha MT, Soliman NEK, Moussa OM, Kamel SM, Shirasawa K (2017) Rapid identification of candidate genes for resistance to tomato late-blight disease using next-generation sequencing technologies. PLoS ONE 12(12):e0189951
Arellano Rodríguez LJ, Rodríguez Guzmán E, Ron Parra J, Martínez Ramírez JL, Lozoya Saldaña H, Sánchez Martínez J, Lépizi Idefonso R (2013) Evaluación de resistencia a Phytophthora infestans en poblaciones silvestres de Solanum lycopersicum var. cerasiforme. Revista mexicana de ciencias agrícolas 4(5):753–766
Bralewski TW, Holubowicz R, Dorota D, Jin HU (2006) Effect of the biostimulator “Asahi SL” on seed germination of a few cultivars of tomato (Lycopersicon Esculentum Mill.). Not Bot Hort Agrobot. https://doi.org/10.15835/nbha341265
Broman KW (2012) Genetic map construction with R/qtl. Springer, New York
Broman KW, Sen S (2009) A guide to QTL mapping with R/qtl. Springer, New York
Chen A-L, Liu CY, Chen CH, Wang JF, Liao YC, Chang CH, Tsai MH, Hwu KK, Chen KY (2014) Reassessment of QTLs for late-blight resistance in the tomato accession L3708 using a restriction site associated DNA (RAD) linkage map and highly aggressive isolates of phytophthora infestans. PLoS One 9(5):e96417
Cooke DE, Cano LM, Raffaele S, Bain RA, Cooke LR, Etherington GJ, Deahl KL, Farrer RA, Gilroy EM, Goss EM, Grünwald NJ (2012) Genome analyses of an aggressive and invasive lineage of the Irish potato famine pathogen. PLoS Pathog 8(10):e1002940
Day JP, Shattock RC (1997) Aggressiveness and other factors relating to displacement of populations of Phytophthora infestans in England and Wales. Eur J Plant Pathol 103(4):379–391
FAO (2019) FAOSTAT Crops, http://www.fao.org/faostat/en/#data/QC. Accessed 04 Mar 2019
Fernandez-Pozo N, Menda N, Edwards JD, Saha S, Tecle IY, Strickler SR, Bombarely A, Fisher-York T, Pujar A, Foerster H, Yan A (2014) The Sol Genomics Network (SGN)—from genotype to phenotype to breeding. Nucleic Acids Res 43:D1036–D1041
Foolad MR, Panthee DR (2012) Marker-assisted selection in tomato breeding. Crit Rev Plant Sci 31(2):93–123
Foolad MR, Merk HL, Ashrafi H (2008) Genetics, genomics and breeding of late-blight and early blight resistance in tomato. Crit Rev Plant Sci 27(2):75–107
Foolad MR, Sullenberger MT, Ohlson EW, Gugino BK (2014) Response of accessions within tomato wild species, Solanum pimpinellifoliumto late-blight. Plant Breed 133(3):401–411
Fridman E, Carrari F, Liu YS, Fernie AR, Zamir D (2004) Zooming in on a quantitative trait for tomato yield using interspecific introgressions. Science 305(5691):1786–1789
Fry W (2008) Phytophthora infestans: the plant (and R gene) destroyer. Mol Plant Pathol 9(3):385–402
Gardner RG, Panthee DR (2010) NC 1 CELBR and NC 2 CELBR: early blight and late blight-resistant fresh market tomato breeding lines. HortScience 45(6):975–976
Grandillo S, Ku HM, Tanksley SD (1999) Identifying the loci responsible for natural variation in fruit size and shape in tomato. Theor Appl Genet 99(6):978–987
Haley CS, Knott SA (1992) A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69(4):315–324
Hartz T, Miyao G, Mickler J, Lestrange M, Stoddard S, Nunez J, Aegerter B (2008) Processing tomato production in California. University of California, Agriculture and Natural Resources, Publication, California, p 7228
Heuvelink E (ed) (2005) Tomatoes. CABI, Wallingford
Kühn C, Bleiss W, Giavalisco P, Willmitzer L, Sulpice R, Zhou YH, Fernie AR (2009) RNA Interference of LIN5 in tomato confirms its role in controlling brix content, uncovers the influence of sugars on the levels of fruit hormones, and demonstrates the importance of sucrose cleavage for normal fruit development and fertility1. Plant Physiol 150(3):1204–1218
Lin T, Zhu G, Zhang J, Xu X, Yu Q, Zheng Z, Zhang Z, Lun Y, Li S, Wang X, Huang Z (2014) Genomic analyses provide insights into the history of tomato breeding. Nat Genet 46(11):1220
Martínez O, Curnow RN (1992) Estimating the locations and the sizes of the effects of quantitative trait loci using flanking markers. Theor Appl Genet 85(4):480–488
Merk HL, Ashrafi H, Foolad MR (2012) Selective genotyping to identify late blight resistance genes in an accession of the tomato wild species Solanum pimpinellifolium. Euphytica 187(1):63–75
Ohlson EW, Ashrafi H, Foolad MR (2018) Identification and mapping of late-blight resistance quantitative trait loci in tomato accession PI 163245. The Plant Genome, 11(3)
Panthee DR, Gardner RG, Ibrahem R, Anderson C (2015) Molecular markers associated with Ph-3 gene conferring late blight resistance in tomato. Am J Plant Sci 6(13):2144
Panthee DR, Piotrowski A, Ibrahem R (2017) Mapping quantitative trait loci (QTL) for resistance to late-blight in tomato. Int J Mol Sci 18(7):1589
Robbins MD, Masud MA, Panthee DR, Gardner RG, Francis DM, Stevens MR (2010) Marker-assisted selection for coupling phase resistance to tomato spotted wilt virus and phytophthora infestans (late-blight) in tomato. HortScience 45(10):1424–1428
Salunkhe DK, Kadam SS (eds) (1998) Handbook of vegetable science and technology: production, composition, storage and processing. Marcel Dekker Inc., New York
Sauvage C, Segura V, Bauchet G, Stevens R, Do PT, Nikoloski Z, Fernie AR, Causse M (2014) Genome-wide association in tomato reveals 44 candidate loci for fruit metabolic traits. Plant Physiol 165(3):1120–1132
Sim S-C, Durstewitz G, Plieske J, Wieseke R, Ganal MW, Van Deynze A, Hamilton JP, Buell CR, Causse M, Wijeratne S, Francis DM (2012a) Development of a large SNP genotyping array and generation of high-density genetic maps in tomato. PLoS One 7(7):e40563
Sim S-C, Van Deynze A, Stoffel K, Douches DS, Zarka D, Ganal MW, Chetelat RT, Hutton SF, Scott JW, Gardner RG, Panthee DR (2012b) High-density SNP genotyping of tomato (Solanum lycopersicum L.) reveals patterns of genetic variation due to breeding. PLoS One 7(9):45520
Staub JE (2010) Alluring lettuces: and other seductive vegetables for your garden. Gibbs Smith, Utah
Stroud JA (2015) Late-blight (Phytophthora infestans) on tomato: evaluation of pathogen population structure in Britain and development of resistant tomato cultivars for growing outdoors (Doctoral dissertation, Bangor University)
Stroud JA, Shaw DS, Hale MD, Steele KA (2016) SSR assessment of Phytophthora infestans populations on tomato and potato in British gardens demonstrates high diversity but no evidence for host specialization. Plant Pathol 65(2):334–341
Zhang C, Liu L, Wang X, Vossen J, Li G, Li T, Zheng Z, Gao J, Guo Y, Visser RG, Li J (2014) The Ph-3 gene from Solanum pimpinellifolium encodes CC-NBS-LRR protein conferring resistance to Phytophthora infestans. Theor Appl Genet 127(6):1353–1364
Acknowledgements
The authors would like to thank Dilip R. Panthee (Mountain Horticultural Crops Research and Extension Center - North Carolina State University, USA) for providing seed of NC 2 CELBR, John Burrows (Pro Veg Seeds, UK) for providing seed of Koralik and David Cooke (James Hutton Institute, Invergowrie, Dundee, UK) for providing late blight isolates. TDB would like to thank Alex Papadopulos and Kristen Crandell (both Bangor University) for helpful discussion about the project. JAS was funded through a Knowledge Economy Skills Scholarship, part funded by the Welsh Government’s European Social Fund with additional support provided by the Sárvári Research Trust and Burpee Europe Ltd. TDB was funded by the Leverhulme Trust research project grant to John F. Mulley and KAS (RPG-2015-450).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Brekke, T.D., Stroud, J.A., Shaw, D.S. et al. QTL mapping in salad tomatoes. Euphytica 215, 115 (2019). https://doi.org/10.1007/s10681-019-2440-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2440-3