Abstract
Fusarium wilt of bananas (Musa spp.), caused by Fusarium oxysporum f. sp. cubense (Foc) causes up to 100% yield loss in bananas. Foc race 1 in particular is very devastating to dessert bananas in Uganda. One of the effective control strategies for the disease is the development of resistant cultivars through breeding. The objectives of this study were to identify suitable banana germplasm for generating a segregating population for resistance to Foc race 1 and understand the mode of inheritance of resistance to Foc race 1. Twenty-two banana accessions sourced from the National Agricultural Research Organisation in Uganda were challenged with Foc race 1 in a screen house experiment. Monyet, resistant to Foc race 1 and Kokopo, susceptible, were selected and crossed to generate 142 F1 genotypes. These F1 genotypes were also challenged with Foc race 1 in a screen house experiment. Data were collected on rhizome discoloration index (RDI), leaf symptom index (LSI) and pseudo-stem splitting (PSS), and analysed for variability. The banana accessions evaluated showed varying degrees of resistance to Foc race 1. Segregation ratios for resistant versus susceptible progenies fitted 13:3 (χ2 = 0.12, P = 0.73) for RDI and 11:5 (χ2 = 3.04, P = 0.08) for PSS. Estimated broad sense heritability was 27.8% for RDI, 13.9% for LSI and 14.7% for PSS. The results suggest that resistance to Foc race 1 in banana is controlled by at least two dominant genes with epistatic interaction and that heritability of resistance to Foc race 1 is low in Musa spp.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Banana (Musa spp.) a heterogeneous, outcrossing and vegetatively propagated crop (Ortiz and Swennen 2014), is cultivated in more than 130 countries in the world (FAOSTAT 2016). Its total production worldwide is estimated at 162 Mio. Metric Tons (MMT), with 21 MMT (14%) deemed for export, earning about US$100 billion (FAOSTAT 2016). Although industrialised nations view banana essentially as a dessert item, many regions of the developing world consider bananas as an essential staple that contributes significantly to the caloric intake of low-income subsistence farmers (Etebu and Young-Harry 2011; Brown et al. 2017). It is a food and cash crop for more than 70 million smallholder farmers in the Great Lakes Region of Africa, with an annual production worth US$ 4.3 billion, which is about 5% of the region’s gross domestic product (EAC 2012).
Uganda is the largest producer of bananas in Africa with an annual total production of 10 MMT (Kilimo-Trust 2012). Most of the bananas grown in the country are the locally evolved clones known as the East African highland bananas (EAHBs, denoted Musa AAA-EA). The EAHBs include cooking ‘Matooke’ and brewing ‘Mbidde’ types, representing 70% and 20% of the total bananas produced, respectively. The rest of the bananas grown are dessert types, that include ‘Gros Michel' (syn. ‘Bogoya', AAA), ‘Pisang Awak’ (syn. ‘Kayinja', ABB) and Ney Poovan (‘Sukali Ndiizi', AAB and ‘Kisubi’, AB), constituting about 9%, and the plantains (AAB), constituting 1% (Karamura and Karamura 1994; Gold et al. 2002). Dessert bananas are widely eaten when ripe and used in local beer breweries (Van Asten et al. 2010; Karangwa et al. 2016).
Fusarium wilt, also known as Panama disease, is the most important lethal disease of dessert bananas (Bidabadi and Sijun 2018). It is a soil-borne fungal disease caused by Fusarium oxysporum f. sp. cubense (Foc) (Ploetz 2015a, b). Fusarium oxysporum f. sp. cubense race 1 is the primary cause of Fusarium wilt disease of dessert bananas in Uganda (Karangwa et al. 2016). Foc race 1 is reported to cause an estimated yield loss of > 60% in dessert bananas (Tushemereirwe et al. 2000). Controlling Fusarium wilt using chemical, biological and cultural control methods has not been very effective (Guo et al. 2013) partly due to long-term survival of the spores in soil and due to the ability to evolve into new strains able to infect resistant cultivars (Su et al. 1986; Mostert et al. 2017). Cultural practices such as pruning symptomatic leaves, culling and burying of diseased plants have been applied for the control of Fusarium wilt, however, these practices lead to further propagation of the disease as the spores can survive in soil for long periods, with or without an alternate host. Chemical control methods are also hazardous to the environment, domestic animals and humans (Ploetz 2000; Pérez-vicente et al. 2014; Ploetz 2015a, b).
Host plant resistance is an effective alternative to chemical, cultural and biological methods for controlling diseases in banana such as Fusarium wilt (Ploetz 2000). It is durable, environmentally safe and user-friendly for small-scale farmers. Natural sources of fungal and other disease resistance exist in wild species and in synthetic diploids of banana developed by breeding programmes (Uma et al. 2011; MusaNet 2016). These diploids have been used in disease resistance in introgressive hybridization programmes (Tushemereirwe et al. 2014; Brown et al. 2017). Conventional banana breeding is highly challenging due to several factors including a long-life cycle, leading to a long breeding cycle (Popova 2011; Brown et al. 2017) and due to the large space requirement, resulting in high costs. The polyploid nature and low female fertility of most popular cultivars of banana (Nyine et al. 2018) and limited knowledge on the genetics of resistance to pests and diseases, have also significantly hindered banana breeding (Heslop-Harrison and Schwarzacher 2007). The success of genetic resistance breeding strategies is affected by the number of genes involved and the nature of inheritance (Boerma and Hussey 1992; Mundt 2014). Therefore, identifying the sources of resistance and studying the genetics underlying resistance to Foc race 1 is pertinent to support banana breeding programmes. The objectives of the present study were: (1) to identify suitable banana germplasm to utilise in generating a segregating banana population for resistance to Foc race 1 and (2) to understand the mode of inheritance of resistance to Foc race 1.
Materials and methods
Plant germplasm used in the development of segregating population for Fusarium wilt resistance.
A total of 22 parental banana germplasm comprised of 18 diploids, one tetraploid and three triploids (Table 1) was used. The germplasm was sourced from the National Agricultural Research Organisation (NARO) and International Institute of Tropical Agriculture (IITA) in Uganda. Triploid bananas were used as controls. Selection of the germplasm was based on good agronomic traits and varying degrees of resistance to several pests and diseases of economic importance in Uganda.
Experimental site
The experiments for challenging the collected parental banana germplasm and F1 population with Foc race 1 inoculum were conducted in a screenhouse at the National Agricultural Research Laboratories (NARL), Kawanda from April 2015 to June 2018. The National Agricultural Research Laboratories are located in Central Uganda at 32°36′E and 0°25′N, 1210 m above sea level. Kawanda is a hotspot of many pathogens and pests, including Mycosphaerella fijiensis ‘Morelet’, Foc race 1, weevils and nematodes.
Preparation of Foc race 1 inoculum
Foc race 1, VCG 0124 inoculum was prepared following a protocol described by Ssali et al. (2013) with some modifications. The Foc fungus was isolated by culturing corm pieces (1cm3) from suckers of symptomatic ‘Sukali Ndiizi’ that was collected from a “hotspot” previously diagnosed with Foc race 1, VCG 0124 infection at NARL, Kawanda, in Uganda. The corm pieces were sterilised by soaking them in 15% Sodium hypochlorite and then in 70% ethanol for 15 min at each soaking stage. The corm pieces were rinsed four times in sterile water and plated onto potato dextrose agar media (PDA) supplemented with streptomycin (300 μg/mL). The Foc cultures were incubated in the dark at 25 °C with routine subculturing until pure cultures with purplish-whitish mycelia were achieved. A PDA plate (90 × 15 mm) fully colonised by pure Foc mycelium, was inoculated into 1 kg of sterile millet grains that had been autoclaved at 121 °C for 30 min then cooled before inoculation. The inoculated millet substrate was incubated for 10 days at 25 °C in the dark with daily agitation to ensure uniform fungal growth.
Foc race 1 disease challenge assay
Three-month-old tissue-cultured banana plants were planted into plastic polythene pots containing 10 kg of sterile loam soil mixed with 100 g of millet grains colonized with Foc race 1. The experiments were set up in a randomised complete block design (RCBD) with six replications. After planting, experimental plants were maintained in a screen house at 28 °C for 12 h of daylight for 60 days. Fusarium wilt disease was assessed on each plant using the severity level of the three key disease symptoms namely, rhizome discoloration index (RDI), leaf symptom index (LSI) and pseudo-stem splitting (PSS) as described by Viljoen et al. (2017) (Table 2). Specifically, LSI data was recorded at 14 days after inoculation by scoring the yellowing of leaves, while data on PSS and RDI were recorded at 60 days after inoculation.
Developing an F1 population segregating for Fusarium wilt resistance
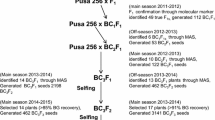
Monyet (Musa acuminata subsp Zebrina), a resistant parental genotype to Foc race 1 VCG 0124 was crossed with Kokopo (Musa acuminata subsp Banksii), a susceptible parental genotype, to generate 142 F1 progenies. Controlled hand pollinations were conducted as described by Ortiz and Vuylsteke (1995). Embryos of the F1 hybrid seeds were extracted for culture as described by Vuylsteke et al. (1990). The ploidy level of the progenies was determined by flow cytometry method as described by Doležel and Bartoš (2005).
Data analysis
In order to select two contrasting parents for Fusarium wilt resistance to be used in generating segregating progenies for Fusarium wilt resistance, the data collected from the 22 banana accessions assessed for resistance to Foc race 1 were subjected to analysis of variance (ANOVA) using GenStat (Payne et al. 2011). Means of RDI, PSS, and LSI were separated using least significance differences (LSD) at 5% significance level. The Disease Severity Index (DSI) of each genotype was computed for RDI as described by Mak et al. (2004) and the germplasm were placed into respective resistance groups following the method described by Sutanto et al. (2013) (Table 3). The genetic basis underlying Fusarium wilt resistance in F1 progeny was determined from the observed frequencies using the Chi-square test (Cochran 1952) versus the standard genetic ratios (Laughlin 1918; Mendel 1866). To determine the broad sense heritabilities (H), the genotypic (σ2g), phenotypic (σ2p) and error (σ2e) variances were computed using the formulae of Burton and DeVane (1953) and Kebere et al. (2006) as σ2g = (MSg−MSe)/r; σ2p = σ2g + σ2e and σ2e = MSe, where MSg = genotypic mean square, MSe = environmental variance (error mean square) and r = the number of replications. Heritability was estimated by the formulae of Wricke and Weber (1986): H = σ2g / σ2p.
Results
Variation of the parental banana germplasm for Fusarium wilt
Genotype mean squares determined by ANOVA were highly significant (P < 0.001) for RDI and LSI as the measure of Fusarium wilt resistance and non-significant for PSS (P > 0.05) (Table 4).
Mean performance of the genotypes for rhizome discoloration index and leaf severity index
Genotypes Hutishamba, Mraru, Mshale, Njuru, Nshonowa, Kahuti, Mlelembo which belong to the AA-Mchare group, and Kokopo, which originates from Papua New Guinea, were significantly different from the resistant control (Mbwazirume) according to the standard error mean separations and LSD between means of the genotypes for RDI (Fig. 1). The remaining genotypes were not significantly different from resistant control (Mbwazirume). Long Tavoy and TMB2X8075-7 showed a slightly lower RDI mean score than the resistant control ‘Mbwazirume’. The standard error means separation and LSD between the means of the genotypes for LSI could not significantly differentiate the resistant control ‘Mbwazirume’ from the susceptible control ‘Kayinja’ (Fig. 2). Furthermore, some genotypes such as TMB2X8075-7, Kasaska, Borneo, and Mwitu Pemba showed high LSI values (Fig. 2) although they had lower RDI values (non-significant RDI values compared to resistant control Mbwazirume).
Grouping of parental germplasm into resistance groups using DSI for RDI
Since the standard error means separation and LSD between means of the accessions for the external symptoms/LSI could not significantly differentiate the resistant control ‘Mbwazirume’ from the susceptible control ‘Kayinja’, grouping the genotypes into resistance groups was performed only for values of DSI for RDI. Therefore, based on DSI for RDI values, genotypes were grouped into four resistance classes: highly resistant, resistant, susceptible and highly susceptible. Only 2 genotypes were grouped as highly resistant, 10 as resistant, 8 as susceptible and 2 as highly susceptible (Table 5).
Genetic basis of the banana resistance to Fusarium wilt
A cross combination of Monyet (tetraploid) and Kokopo (diploid) resulted in 142 F1 progenies with a mixture of ploidy levels: 136 triploids (3x), 4 tetraploids (4x) and 2 diploids (2x). The genotype mean squares in the ANOVA for 142 F1 progenies were highly significant (P < 0.001) for RDI, LSI and PSS (Table 6).
Using DSI for RDI, 117 F1 progenies were grouped as resistant (scale of 1.0–3.0) and 25 F1 progenies as susceptible (Scale 3.1–6.0) while using DSI for LSI, 73 F1 progenies were grouped as resistant (Scale 1.0–2.0) and 69 F1 progenies as susceptible (Scale 2.1–5.0). Using DSI for PSS, 88 F1 progenies were grouped as resistant (Scale 1.0) and 54 as susceptible (1.1–3.0). The segregation ratio for RDI fitted the two gene model ratio of 13:3 while PSS fitted the two gene model ratio of 11:5 using a chi square goodness of fit test (Table 7). LSI segregation did not fit either of the one gene model ratios nor the two gene model ratios tested. The 13:3 ratio is described as complete dominance at both gene pairs; however, when either gene is dominant, it overshadows the effects of the other gene, while a ratio of 11:5 indicates complete dominance for both gene pairs only if both kinds of dominant alleles are present; otherwise, the recessive phenotype appears.
Estimates of broad-sense heritability for the resistance parameters RDI, LSI and PSS as measures of Fusarium wilt were relatively low (Table 8). RDI had a heritability of 27.8%, while LSI and PSS had heritabilities of 13.9% and 14.7%, respectively.
Discussion
Fusarium wilt [Fusarium oxysporum f. sp. cubense (Foc)] is a destructive soil-borne fungal disease that causes heavy yield losses among susceptible bananas worldwide. Foc race 1 was responsible for the destruction of 40,000 hectares of Gros Michel plantations in the Central American/Caribbean region in 1940 (Ploetz and Pegg 2000). In Uganda, Foc race 1 heavily affects the dessert banana cultivars, leading to complete destruction of the fields if not controlled (Tushemereirwe et al. 2004). Fusarium wilt can be appropriately addressed by providing farmers with resistant varieties through breeding. Therefore, identifying sources of resistance to Foc race 1 and understanding genetic mechanisms underlying Foc race 1 resistance are a fundamental step towards breeding resistant banana varieties.
The first part of this study included an assessment of 22 parental banana accessions that had never been utilised for Foc race 1 resistance breeding because their response to Foc race 1 was unknown. However, upon assessment of the parental genotypes, the mean squares in the ANOVA were significantly different for RDI and LSI, implying that they had varying degrees of resistance to Foc race 1. Genotypes that were significantly different from the susceptible controls (Kayinja and Sukali Ndiizi) were subsequently grouped into the highly resistant and the resistant classes by disease severity index (DSI) scores for rhizome discoloration (RDI). Ten out of 18 (~ 55.6%) of the diploids evaluated were classified as either highly resistant or resistant and the tetraploid (Monyet) was grouped as resistant. The results reported in this study are in agreement with those reported by Uma et al. (2011) and Ribeiro et al. (2017) where the authors reported that diploid banana accessions they assessed were resistant to most of the pests and diseases tested. A study by Kumar et al. (2009) also found a majority (4/7 synthetic diploids and 4/6 diploid parents) of the diploid banana cultivars to be resistant to Foc race 1. Genotype Kokopo, a fertile diploid and assumed to be a source of vitamin A because of its orange-fleshed fruit colour, was grouped as susceptible. Kokopo can be utilised for genetic studies for Foc race 1 and also as a source of vitamin A for incorporation into the East African highland banana (EAHBs) breeding programmes. The Mchare varieties (Hutishamba, Mraru, Mshale, Njuru, Nshonowa, Kahuti, Mlelembo), which are the most common edible bananas in Tanzania, were all susceptible to Foc race 1 in the present study. This may pose a threat to the food security in the region and therefore steps to improve the resistance of Mchare bananas against Foc race 1 are highly recommended. Accessions that have been identified as resistant or susceptible in the present study can be used as parents to be integrated into breeding programmes to improve the resistance of dessert, plantain and Mchare bananas to Foc race 1 and for studying the mechanisms underlying Fusarium wilt resistance.
The standard error means separation and the LSD between the means for leaf symptom index (LSI) could not significantly differentiate the resistant control Mbwazirume from the susceptible control Kayinja. Some genotypes, such as TMB2X8075-7, Kasaska, Borneo, Mwitu Pemba that had shown lower RDI (non-significant RDI values compared to resistant control Mbwazirume), showed advanced yellowing symptoms (i.e. high LSI value) and could not be clearly differentiated from the susceptible controls. The advanced yellowing of these genotypes may not be associated with Fusarium wilt infection since they had lower RDI values or no rhizome discoloration, but is possibly associated with mineral deficiency and/ or excessive water. Ribeiro et al. (2017) reported that a plant can show external characteristics such as advanced yellowing (high LSI) due to nutritional deficiency and excess water, but internally may not exhibit rhizome discoloration (high RDI). According to Li et al. (2015), Foc can cause internal corm discoloration without causing any external symptoms such as yellowing of leaves. Ribeiro et al. (2017) and Li et al. (2015) suggested that dissecting the rhizome to verify the absence/presence of discoloration is the most precise evaluation of Fusarium wilt. Therefore, we grouped the germplasm into resistance groups and selected parents contrasting for Foc race 1 resistance, only based on the DSI for RDI.
We selected Monyet (Foc race 1 resistant) and Kokopo (susceptible) as suitable parents for developing a segregating F1 population for assessing the genetic basis of resistance to Foc race 1. The tetraploid Monyet was selected to be used as the female because of its moderate female fertility, while Kokopo, a diploid was used as the source of pollen.
The segregation ratios of resistant vs susceptible for the F1 progenies fit 13:3 (χ2 = 0.12, P = 0.73) for RDI and 11:5 (χ2 = 3.04, P = 0.08) for PSS. Both 13:3 and 11:5 ratios obtained in current study are a deviation from the expected 9:3:3:1 dihybrid ratio, suggesting that Foc race 1 is under the genetic control of at least two dominant genes with epistatic interactions. Previous studies have reported Fusarium wilt to be under the genetic control of a single dominant and single recessive gene. Single dominant genetic controls of Fusarium wilt reports include Larter (1947) who reported that Foc race1 was controlled by a single dominant gene in a study of tetraploid progenies obtained by the cross of Gros Michel with a diploid accession. Vakili (1965) also reported Foc race1 to be under the control of a single dominant gene in a banana population developed using a homozygous banana parent ‘Pisang Lilin’ as the source of resistance. Fraser-Smith et al. (2016) reported Foc subtropical race 4 (SR4) and TR4 to be under the genetic control of a single dominant gene in an F1 progeny of self-fertilized malaccensis plants. Control of resistance by a recessive gene has also been observed by Ssali et al. (2013) who reported that Foc race 1 inheritance was controlled by a single recessive gene in an F2 population derived from crosses of ‘Sukali Ndizi’ (AAB) and a resistant diploid banana ‘TMB2X8075′ (AA).
The two dominant genes with epistasis obtained in the current study differing from the most reported single dominant gene model, could be because an early F1 generation was used. Ssali et al. (2013) reported that there is more genetic variation in F2 banana individuals, which provides a better platform to study mode of inheritance compared to the low variation in F1 individuals. Another cause of discrepancy in the gene ratio could be due to the low number of F1 progenies used in the current study. Ideal mapping populations should consist of a minimum of 50–250 individuals (Collard et al. 2005). Ortiz and Vuylsteke (1994) attributed the inconclusiveness between the one or two genes model controlling the inheritance of albinism in Musa spp. to the small sample sizes of below 65 genotypes (a problem inherent in the low reproductive fertility of cultivated parthenocarpic Musa). Other factors that could have affected the genetic ratios in the current study are, the use of single environments for evaluations and heterozygosity between parents. Kammili and Raoof (2014) attributed the different inheritance patterns (15:1, 9:7 and 13:3) of Fusarium oxysporum f.sp. ricini observed in castor (Ricinus communis L.) to the use of a single location for the evaluations and high levels of heterozygosity and heterogeneity within parents used in their study. Therefore, another study is recommended to confirm the genes controlling Foc race 1 using an advanced F2 population with a large number of progenies. Also, it will be important to confirm the nature of inheritance of resistance to Foc race 1 using molecular markers. However, there are no available molecular markers associated with Foc race 1 in bananas.
The generation of an F1 population segregating for Foc race 1 resistance, provided an opportunity to determine heritability of resistance to this trait for the first time in Musa spp. We found heritability of 27.8% for RDI, 14.7% for PSS and 13.9% for LSI which can be considered low based on the heritability scale described by Johnson et al. (1955), where heritability of 0–30% is classified as low. Several studies have reported low heritabilities for various pests and diseases in Musa spp. Ssali et al. (2016) reported a low heritability of 24.4% for youngest leaf spotted when studying black Sigatoka in secondary triploid banana ‘Matooke’ (Musa sp., AAA-EA) hybrids. Arinaitwe et al. (2015) reported a low heritability of 24.0% for total corm damage caused by weevils in an F2 diploid population segregating for weevil resistance. Kiggundu (2000) found a low weevil cross section damage heritabilities of 29% for both upper inner and lower outer damage among hybrids of Musa spp. The low heritability values obtained in the current study suggests that environmental factors play a big role in inheritance of resistance to Foc race 1 and therefore, selection based on phenotype is not recommended. Dutta et al. (2013) and Bushra et al. (2017) reported that selection based on phenotype performance is more effective when the heritability estimates are significantly high. It is, therefore, commendable to use molecular markers when selecting for pest and disease resistances in Musa spp. because they are not affected by the environment.
Conclusion
There was high degree of variability among the parental banana germplasm evaluated for Foc race 1, indicating that by hybridizing among them, genetic advance would be achieved for resistance to Foc race 1. Therefore, the genotypes evaluated are recommended for integration in the banana breeding program for Foc race 1 resistance breeding. Resistance to Foc race 1 among F1 progenies evaluated, was controlled by at least two dominant genes with epistatic interaction. Low heritability of resistance to Foc race 1 was observed in the present study, indicating that the expression of this trait was strongly influenced by the environment. Hence, direct phenotypic-based selection for Foc race 1 would probably be ineffective and there is a need for marker assisted selection. There is also a need to develop molecular markers for Foc race 1 in bananas by identifying Foc race 1 quantitative trait loci (QTLs) from the current developed banana population.
References
Arinaitwe IK, Barekye A, Kubiriba J, Sadik K, Karamura E, Edema R (2015) Genetic analysis of weevil (Cosmopolites sordidus) resistance in an F2 diploid banana population. Plant Breed Genet 01:34–42
Van Asten PJ, Florent D, Apio MS (2010) Opportunities and constraints for dried dessert banana (Musa spp.) export in Uganda. Acta Hortic 879:105–112. https://doi.org/10.17660/ActaHortic.2010.879.8
Bidabadi SS, Sijun Z (2018) Banana Fusarium wilt (Fusarium oxysporum f. sp. cubense) control and resistance, in the context of developing wilt-resistant bananas within sustainable production systems. Hortic Plant J. https://doi.org/10.1016/j.hpj.2018.08.001
Boerma HR, Hussey RS (1992) Breeding plants for resistance to nematodes. J Nematol 24:242–252
Brown A, Tumuhimbise R, Amah D, Uwimana B, Nyine M, Mduma H, Swennen R et al (2017) Bananas and plantains (Musa spp.). In: Campos H, Caligari PD (eds) Genetic improvement of tropical crops. Springer, Basel, pp 219–240
Burton GW, DeVane EH (1953) Estimating heritability in Tall fescue (Festuca arundinacea) from replicated clonal material. Agron J 45:478. https://doi.org/10.2134/agronj1953.00021962004500100005x
Bushra R, Muhammad T, Aleena K, Faiza S, Qurban A, Fareeha A, Tayyab H et al (2017) Crop improvement: new approaches and modern techniques. Plant Gene Trait 8:18–30. https://doi.org/10.5376/pgt.2017.08.0003
Cochran WG (1952) The Chi-square test of goodness of fit. Ann Math Stat 23:315–345. https://doi.org/10.1214/aoms/1177729380
Collard B, Jahufer M, Brouwer J, Pang ECK (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196. https://doi.org/10.1007/s10681-005-1681-5
Doležel J, Bartoš J (2005) Plant DNA flow cytometry and estimation of nuclear genome size. Ann Bot 95:99–110. https://doi.org/10.1093/aob/mci005
Dutta P, Dutta PN, Borua PK (2013) Morphological traits as selection indices in rice: a statistical view. Univ J Agric Res 1:85–96. https://doi.org/10.13189/ujar.2013.010308
EAC (2012) East African community facts and figures. EAC Secretariat, Arusha
Etebu E, Young-Harry W (2011) Control of black Sigatoka disease: challenges and prospects. Afr J Agric Res 6:508–514. https://doi.org/10.5897/AJAR10.223
FAOSTAT (2016) Crops and products domain. FAO. https://www.fao.org/faostat/en/#data/QC. Accessed 18 Sept 2018
Fraser-Smith S, Czislowski E, Daly A, Meldrum R, Hamill S, Smith M, Aitken EA (2016) Single gene resistance to Fusarium oxysporum f. sp. cubense Race 4 in the wild banana Musa acuminata subsp. malaccensis. Acta Hortic 1114:95–100. https://doi.org/10.17660/ActaHortic.2016.1114.13
Gold CS, Kiggundu A, Abera MK, Karamura D (2002) Diversity, distribution and farmer preference of Musa cultivars in Uganda. Exp Agric 38:39–50. https://doi.org/10.1017/S0014479702000145
Guo G, Wang B, Ma W, Li X, Yang-Zhu C, Zeng H (2013) Biocontrol of Fusarium wilt of banana: key influence factors and strategies. Afr J Microbiol Res 7:4835–4843. https://doi.org/10.5897/AJMR2012.2392
Heslop-Harrison JS, Schwarzacher T (2007) Domestication, genomics and the future for banana. Ann Bot 100:1073–1084. https://doi.org/10.1093/aob/mcm191
Johnson HW, Robinson HF, Comstock RE (1955) Estimates of genetic and environmental variability in soybeans. Agron J 47:314–318. https://doi.org/10.2134/agronj1955.00021962004700070009x
Kammili A, Raoof MA (2014) Analysis of mode of inheritance of Fusarium wilt resistance in castor (Ricinus communis L.). Plant Breed 133:101–107. https://doi.org/10.1111/pbr.12126
Karamura D, Karamura E (1994) A provisional checklist of banana cultivars in Uganda. NARO-Uganda, Kampala
Karangwa P, Blomme G, Beed F, Niyongere C, Viljoen A (2016) The distribution and incidence of banana Fusarium wilt in subsistence farming systems in east and central Africa. Crop Prot 84:132–140. https://doi.org/10.1016/j.cropro.2016.03.003
Kebere B, Prapa S, Wasana W, Vipa H (2006) Genetic variation, heritability and path-analysis in Ethiopian finger millet [Eleusine coracana (L.) Gaertn] landraces. Kasetsart J Nat Sci 40:322–334
Kiggundu A (2000) Host-plant interactions and resistance mechanisms to Banana weevil cosmopolites sordidus (Germar) in Ungandan musa germplasm. Dissertation, University of the Orange Free State
Kilimo-Trust (2012) Banana value chain(s) in East Africa: consumption, productivity and challenges. Kampala, Uganda
Kumar N (2006) Problems and prospects of banana breeding in India. J Hortic Sci 1:77–94
Kumar N, Damodaran T, Krishnamoorthy V (2009) Breeding banana for combined resistance to Fusarium wilt and nematodes in Tamil Nadu, India. Acta Hortic 828:323–332. https://doi.org/10.17660/ActaHortic.2009.828.33
Larter LMN (1947) Report on banana breeding. Department of Agriculture of Jamaica, Kingstone
Laughlin HH (1918) Modifications of the 9:3:3:1 ratio. Am Nat 52:353–364. https://doi.org/10.2307/2456019
Li WM, Dita M, Wu W, Hu GB, Xie JH, Ge XJ (2015) Resistance sources to Fusarium oxysporum f. sp. cubense tropical race 4 in banana wild relatives. Plant Pathol 64:1061–1067. https://doi.org/10.1111/ppa.12340
Mak C, Mohammed AA, Liew KW, Ho WY (2004) Early screening technique for Fusarium wilt resistance in banana micropropagated plants. Banana Improv 18:219–227
Mendel G (1866) Experiments in plant hybridization. Read at the February 8th, and March 8th, 1865, Meetings of the Brünn Natural History Society, pp 3–47
Mostert D, Molina AB, Daniells J, Fourie G, Hermanto C, Chao C, Viljoen A (2017) The distribution and host range of the banana Fusarium wilt fungus, Fusarium oxysporum f. sp. cubense, in Asia. PLoS ONE 12:e0181630. https://doi.org/10.1371/journal.pone.0181630
Mundt CC (2014) Durable resistance: a key to sustainable management of pathogens and pests. Inf Genet Evol 27:446–455. https://doi.org/10.1016/j.meegid.2014.01.011
MusaNet (2016) Global strategy for the conservation and use of Musa (Banana) genetic resources. Biodiversity International, Montpellier
Nyine M, Uwimana B, Blavet N, Hřibová E, Vanrespaille H, Batte M, Doležel J et al (2018) Genomic prediction in a multiploid crop: genotype by environment interaction and allele dosage effects on predictive ability in Banana. Plant Genome. https://doi.org/10.3835/plantgenome2017.10.0090
Ortiz R, Swennen R (2014) From crossbreeding to biotechnology-facilitated improvement of banana and plantain. Biotechnol Adv 32:158–169. https://doi.org/10.1016/j.biotechadv.2013.09.010
Ortiz R, Vuylsteke DR (1994) Inheritance of albinism in banana and plantain (Musa spp.) and its significance in breeding. Hortic Sci 29:903–905
Ortiz R, Vuylsteke DR (1995) Factors influencing seed set in triploid Musa spp. L. and production of euploid hybrids. Ann Bot 75:151–155. https://doi.org/10.1006/anbo.1995.1006
Payne R, Murray D, Harding S, Baird D (2011) An introduction to GENSTAT for Windows, 14th edn. VSN International, Hemel Hempstead
Ploetz RC (2000) Panama disease: a classic and destructive disease of banana. Plant Health Prog 10:1–7. https://doi.org/10.1094/PHP-2000-1204-01-HM
Ploetz RC (2015a) Fusarium wilt of banana. Phytopathology 105:1512–1521. https://doi.org/10.1094/PHYTO-04-15-0101-RVW
Ploetz RC (2015b) Management of Fusarium wilt of banana: a review with special reference to tropical race 4. Crop Prot 73:7–15. https://doi.org/10.1016/j.cropro.2015.01.007
Ploetz RC, Pegg KG (2000) Fungal diseases of the root, corms and pseudostem: Fusarium wilt. In: Jones DR (ed) Diseases of banana. CABI Publishing, Wallingford, pp 143–159
Popova M (2011) The life cycle of a Banana-the Atlantic. https://www.theatlantic.com/health/archive/2011/12/the-life-cycle-of-a-banana/249551/. Accessed 7 June 2018
Pérez-vicente L, Dita MA, Martínez-de-La-parte E (2014) Technical manual: prevention and diagnostic of fusarium wilt (Panama disease) of banana caused by technical manual prevention and diagnostic of fusarium wilt (Panama disease) of banana caused by Fusarium oxysporum f . sp . cubense Tropical Race 4 (TR4). In worshop on diagnosis of Fusarium Wil. FAO, St Augustine, pp 1–74
Ribeiro RL, De Oliveira S, Santos SA, Amorim EP, Almeida J, Serejo S, Haddad F (2017) Sources of resistance to Fusarium oxysporum f. sp. cubense in banana germplasm. Rev Bras Fruticult 40:1–8. https://doi.org/10.1590/0100-29452018202
Ssali RT, Barekye A, Buregeya H, Erima R, Namanya P, Kubiriba J (2016) Genotypic variability estimates of agronomic traits in secondary triploid banana Matooke (Musa spp., AAA-EA) hybrids. Afr J Plant Sci 10:84–88. https://doi.org/10.5897/AJPS2015.1343
Ssali RT, Kiggundu A, Lorenzen J, Karamura E, Tushemereirwe W, Viljoen A (2013) Inheritance of resistance to Fusarium oxysporum f. sp. cubense race 1 in bananas. Euphytica 194:425–430. https://doi.org/10.1007/s10681-013-0971-6
Su HJ, Hwang S, Ko W (1986) Fusarial wilt of Cavendish bananas in Taiwan. Plant Dis 70:814–818. https://doi.org/10.1094/PD-70-814
Sutanto A, Sukma D, Hermanto C (2013) The study and early evaluation of resistance of banana accessions for wilt disease caused by Fusarium oxyporum f.sp. cubense VCG 01213/16 (TR4). In: Improving food, energy and environment with better crops. 7th Asian crop science association conference, IPB International Convention Center 27–30 September 2011. Research Center for Bioresources and Biotechnology Bogor Agricultural University, Bogor, pp 291–295
Tushemereirwe W, Kangire A, Kubiriba J, Nowakunda K (2000) Fusarium wilt resistant banana considered appropriate replacements for cultivars susceptible to the disease in Uganda. Uganda J Agric Sci 5:62–64
Tushemereirwe WK, Kangire A, Kubiriba J, Nakyanzi M, Gold CS (2004) Diseases threatening banana biodiversity in Uganda. Afr Crop Sci J 12:19–26. https://doi.org/10.4314/acsj.v12i1.27658
Tushemereirwe W, Batte M, Nyine M, Tumuhimbise R, Barekye A, Talengera D, Kubiriba J, Lorenzen J, Swennen R et al (2014) Performance of Narita banana hybrids in the preliminary yield trial, Uganda. NARO-Uganda, Kampala
Uma S, Mustaffa MM, Saraswathi MS, Durai P (2011) Exploitation of diploids in Indian breeding programmes. Acta Hortic 897:215–223. https://doi.org/10.17660/ActaHortic.2011.897.23
Vakili NG (1965) Fusarium wilt resistance in seedlings and mature plants of Musa species. Phytopathology 55:135
Viljoen A, Mahuku G, Massawe C, Ssali RT, Kimunye J, Mostert G, Coyne DL (2017) Banana diseases and pests: field guide for diagnostics and data collection. International Institute of Tropical Agriculture (IITA), Ibadan
Vuylsteke D, Swennen R, De Langhe E (1990) Tissue culture technology for the improvement of African plantains. In: Sigatoka leaf spot diseases of Bananas: Proceedings of an international workshop INIBAP, Montpellier, pp 316–337
Wricke G, Weber WE (1986) Quantitative genetics and selection in plant breeding. de Gruyter, Berlin
Acknowledgements
The authors thank all donors who supported this work through their contributions to the CGIAR Fund (http://www.cgiar.org/who-we-are/cgiar-fund/fund-donors-2/), and in particular to the CGIAR Research Program Roots, Tubers and Bananas (RTB-CRP) and the University of Malaya Collaborative Project IF015-2015.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Arinaitwe, I.K., Teo, C.H., Kayat, F. et al. Evaluation of banana germplasm and genetic analysis of an F1 population for resistance to Fusarium oxysporum f. sp. cubense race 1. Euphytica 215, 175 (2019). https://doi.org/10.1007/s10681-019-2493-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2493-3