Abstract
Grain mold is one of the most devasting diseases in sorghum [Sorghum bicolor L. (Moench)] that affects the endosperm and deteriorates the pericarp tissue, reducing the quality of the grain. Today, sorghum breeding programs have a limited number of sources of resistance for the development of resistant cultivars. Therefore, the USDA-Agriculture Research Service, National Plant Germplasm System Sudan core collection was assessed to identify new sources of grain mold resistance based on seed emergence and deterioration. A total of 246 accessions were evaluated for two years and a subset of 46 accessions with grain mold resistance were subsequently evaluated for two additional years together with 11 breeding resistant lines from the sorghum association panel. The analysis identified 39 grain mold resistance accessions including seven that showed both high seedling emergence (> 82%) and low seed deterioration (< 2.15). Phylogenetic analysis revealed that five accessions (PI 570382, PI 570776, PI 570330, PI 570702, and PI 570348) that clustered distantly from reference sets and showed both high seedling emergence and low seed deterioration can be classified as new resistance sources. Genome-wide association analysis using 147,069 SNPs identified two genomic regions in chromosome 2 and 3 associated with seedling emergence rate and seed deterioration, respectively. The analysis of both genomic regions found two genes of interest associated with phenylpropanoid metabolic process and phosphorylase kinase. These Sudanese grain mold resistance accessions provide new genetically diverse germplasm for breeding programs and insights in the defense resistance responses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The drought-tolerance of sorghum [Sorghum bicolor (L.) Moench] makes its worldwide production possible, especially in the dry regions of multiple countries. In fact, this important dietary staple cereal was domesticated in the arid regions of Northeast Africa, an area that extends from Ethiopia to Sudan (Mann et al. 1983). These two countries enclose most of the genetic diversity of the crop, however, through its distribution to different environmental regions of Africa the crop diversified into five races (Kafir, Durra, Bicolor, Caudatum and Guinea). Each race shows a particular panicle shape and kernel morphology that is associated to the environment where they were re-domesticated (Snowden 1936). Moreover, the distribution of sorghum to south Africa resulted in the selection of temperate adapted genes that make the plant insensitive to day-length (Murphy et al. 2011). Today, sorghum is the fifth most important cereal worldwide (FAOSTAT 2022) and is cultivated in tropical and temperate regions. Most of the genetic diversity of the crop is enclosed in tropical germplasm that flower during short day length (i.e. < 12 h; sensitive to day-length), while temperate adapted germplasm (i.e. insensitive to day-length or day-neutral) includes most of the improved cultivars and commercial seed hybrids. Consequently, public and private sorghum breeding programs in temperate regions have a narrow genetic diversity that increases the vulnerability of its production (Menz et al. 2004).
Ex situ sorghum germplasm collections have been established in different countries to preserve the crop genetic diversity and to make it available for breeding programs. But the large size of many sorghum collections makes it impossible to adequately screen the germplasm and identify the most valuable accessions. Core collections are representative subsets (~ 10–20%) of the germplasm collection that are selected to capture most of the genetic diversity in the large collection (Brown 1989). For instance, the United State Department of Agriculture (USDA) and its National Plant Germplasm System (NPGS) conserve a sorghum collection of > 40,000 accessions from 114 countries. Thus, a core collection of 3001 accessions was established using collection sites to facilitate its screening and preservation (Dahlberg et al. 2004). Likewise, the International Crops Research Institute for the Semi-arid Tropics (ICRISAT) preserves a sorghum collection of > 36,000 accessions and established a core collection of 2247 accessions based on morpho-agronomic traits (Grenier et al. 2001). A proportional sampling of this core collection led to a mini-core collection of 242 accessions for extensive phenotyping and genotyping characterization (Morris et al. 2013; Upadhyaya et al. 2009). Indeed, the introgression of new genetic diversity into sorghum breeding programs is initiate with the screening and identification of most valuable accessions within these core sets and later by the allele mining in these large germplasm collections.
The development of new genotyping techniques based on next-generation sequencing led to the genetic characterization of germplasm collections. In this regard, genotyping-by-sequencing [GBS; Elshire et al. (2011)] is a low cost and effective techniques that identify large number of single nucleotide polymorphism (SNPs) suitable for genome-wide association analysis (GWAS) and genetic diversity study. The GBS analysis of the ICRISAT mini-core collection resulted in the identification of ~ 265,000 SNPs (Morris et al. 2013). Likewise, NPGS core collection from Ethiopia, Sudan, and Yemen, as the whole collections from Nigeria, Senegal and Niger have been subject to GBS analysis providing insight of its genetic diversity (Cuevas et al. 2017; Cuevas and Prom 2020; Faye et al. 2019; Maina et al. 2018; Morales-Salva 2022; Olatoye et al. 2018). Certainly, the Ethiopian and Sudanese collections are the most genetically diverse and both revealed a large number of rare alleles (Cuevas et al. 2017; Cuevas and Prom 2020). The population structure analysis of these collections showed signature of selection for agronomic traits and of adaptation to agro-environmental regions (Cuevas et al. 2017; Faye et al. 2019). Moreover, phylogenetic analyses between these NPGS tropical accessions and temperate adapted breeding lines present in the sorghum association panel [SAP; Morris et al. (2013), Faye et al. (2019)] are being used to identify and select the most valuable accessions for breeding programs. The high throughput screening of these NPGS core collections for abiotic and biotic factors that affect sorghum production might lead to the identification of new important loci and the development of new and improved tropical and temperate adapted germplasm.
Sorghum is affected by various biotic factors associated with the environmental production regions. The warm and humid environmental conditions in sub-topical and tropical regions are characterized by the presence of multiple fungal diseases such as grain mold, anthracnose (Colletotrichum sublineola), rust (Puccinia purpurea), downy mildew (Peronosclerospora sorghi), and ergot (Claviceps spp.) (Fakrudin et al. 2021). Grain mold is caused by a complex of more than 40 pathogenic and opportunistic fungi including Fusarium thapsinum (Klittich, Leslie, Nelson, & Manasas.), Fusarium semitectum (Berk. & Ravenel), Curvularia lunata (Wakk.) Boedijn, Alternaria alternata (Fr.) Keissler, and Colletotrichum sublineola, and is one of the most devastating diseases worldwide (Thakur et al. 2006). The disease affects the endosperm and embryo tissue reducing its viability and nutritional value (Bandyopadhyay et al. 2000) and deteriorate the pericarp tissue affecting the appearance of the grain (Forbes 1986). Yield losses associated to grain mold diseases can reach 100% in highly susceptible cultivars (Navi et al. 2005). Moreover, mycotoxin contamination associated with grain mold is one of the most important threats to sorghum grain quality and safety (Ackerman et al. 2021). Today, the use of resistant cultivars provides the most effective method to control diseases (Thakur et al. 2007), however, the narrow genetic diversity in breeding programs limits the number of resistance sources. Therefore, the identification of new resistance sources in temperate and tropical germplasm is imperative to sustain an adequate control of the disease.
The inheritance of grain mold resistance response is not well understood. Panicle architecture has an important role in the disease development because the airflow and low moisture in open panicles structures reduce grain mold incidence (Ackerman et al. 2021; Cuevas et al. 2018). However, resistance sources found in the SAP showed different panicle shapes indicating resistant response is also associated with genes related to plant immunity system (Cuevas et al. 2019). Genome-wide association analysis using tropical and temperate sorghum germplasm has delimited several genomic regions associated with the observed resistance response (Cuevas et al. 2019; Nida et al. 2019; Prom et al. 2020). Candidate gene analysis within these genomic regions include genes associated to plant immunity system (e.g., R genes), biochemical pathway and grain pigmentation [e.g. YELLOW SEED1 (Y1)] revealing the complexity of the defense response. These loci explained a modest proportion of the estimated heritability; thus, family inheritance research is still necessary to improve the understanding of the resistance response. It is necessary to identify multiple genetically diverse resistance sources for the development of multiple perennial segregating populations that enable genomic dissection of the grain mold resistance responses.
The NPGS Sudan core collection is genetically diverse germplasm that enclosed agronomically valuable accessions (Cuevas and Prom 2020). Nevertheless, this core collection needs to be extensively phenotyped for multiple biotic and abiotic factors. In the current study, the NPGS Sudan core collection was evaluated for: (1) identification of new grain mold resistant accessions, (2) determination of the genetic relationship between Sudanese resistant accessions and resistance sources present in SAP, and (3) identification of genomic regions that are associated with grain mold resistance through genome-wide association analysis.
Material and methods
Germplasm and field experiment
A total of 246 accessions from the NPGS Sudan core collection (Cuevas and Prom 2020; Dahlberg et al. 2004) were evaluated for grain mold resistance at the USDA-ARS Tropical Agriculture Research Station in Isabela (18.471569, − 67.043472) and Mayaguez (18.211536, − 67.135562) in 2016 and 2017, respectively. The Sudanese core set, three susceptible lines (RTx430, RTx2536, RTx2911 and BTx623), and two resistant lines [Sureño and PI 267548 (Prom and Erpelding 2009)] were planted at both locations using a completely randomized design with a single replication, with plots measuring 1.8 m in length with 0.9 m between rows.
These two-year evaluations were used to select a subset of 46 accessions that showed grain mold resistance based on its average seed deterioration (≤ 2.5 on a 5-point scale) and emergence rate (≥ 75%). This subset, susceptible checks (RTx430, RTx2536 and RTx2911) and the resistant checks from the SAP [‘Summac’, ‘Rox Orange’, ‘Red Amber’, ‘Keller’, ‘Kansas Orange’, ‘SC309’, ‘SC609’, ‘Sureño’, ‘SC15’ ‘SC719’ and ‘SC13’; (Cuevas et al. 2019)] and PI 267548 were planted in 2020 and 2021 in randomized block design consisting of four blocks with plots measuring 1.8 m in length with 0.9 m between rows at Isabela, Puerto Rico.
The temperature and relative humidity in the four experiments were recorded using an Onset HOBO U23 Pro V2 located within research plots. Aerial watering was supplied in the absence of rainfall, while plots and plants were maintained using mechanical tillage and hand hoeing for weed control.
Grain mold response
The grain mold response was determined based on visual evaluations of seed mold and weathering degradation (hereafter referred to as seed degradation) and seedling emergence. Three to five panicles with uniform flowering time per plot were covered with mesh bags for exposure to environmental conditions and to avoid damage by birds, harvested at maturity stage (30–40 days after anthesis), dried, and threshed using a single plant thresher (Almaco Single Plant and Head Thresher; Allan Machine Company). Approximately 30–50 g of seeds from each panicle were visually rated for seed degradation using a 5-point visual scale (Thakur et al. 2007; Isakeit et al. 2008) where 1 indicates seed is bright with no mold and no discoloration resulting from weathering; 2 indicates seed is not as bright and has little or no mold but has some discoloration (1–10% molded kernels); 3 indicates seed is not bright, with some mold and some discoloration (11–25% molded kernels); 4 indicates seed is almost entirely covered in mold, and the pericarp is degraded (26–50% molded kernels); and 5 indicates seed is covered entirely with mold, and the pericarp is degraded and looks dead (> 50% molded kernels). Subsequently, thirty seeds selected randomly from each panicle were planted in flats containing Metro Mix 200 potting medium and incubated in a greenhouse at room temperature. The emergence rate was based on the percentage of seedlings observed after 10 days.
Phenotypic analysis
For the first two years of experiments, we implemented a randomized complete block design (RCDB) with two blocks (i.e. years) for analysis of seed degradation and emergence rate. The subset of 46 accessions and 15 reference lines evaluated for four years were analyzed as a RCDB with four blocks. Analysis of variance for seed degradation and emergence rate were completed in SAS (SAS Institute, Cary, NC) using the Proc mixed covtest method type 3 procedure where years were treated as fixed and accessions and blocks as random effects. The repeatability of the experiment for both traits was estimated by using the following formula:
where σ2g and σ2e refer to the accessions and error variances, respectively, while e is the number of blocks (Bernardo 2002). The performance of accessions for seed degradation and emergence rate was compared using the Tukey–Kramer’s post-hoc test at the 5% level of significance. The five populations present in the Sudan core collection (Cuevas and Prom 2020) were also compared for seed degradation and emergence rate using the Tukey–Kramer’s post-hoc test at the 5% level of significance.
Genome-wide association analysis
The genomic characterization of Sudan core collection is a public resource available in the National Center for Biotechnology Information (NCBI) Sequence Read Archive database [Bio Project PRJNA565973; (Cuevas and Prom 2020)]. These 246 Sudanese accessions enclose 147,069 SNPs with minor allele frequencies (MAF) ≥ 0.05 suitable for genome-wide association analysis. The Bayesian-information and Linkage-disequilibrium Interatively Nested Keyway (BLINK) method was implemented for association analysis using the Genome Association and Prediction Integrated Tool (GAPIT) in R (Lipka et al. 2012). Because previous population structure analysis of the Sudanese germplasm collections found that it consisted of five populations (Cuevas and Prom 2020) four components of the ancestry matrix from STRUCTURE (Pritchard et al. 2000) analysis were included as covariates to control population structure and family relatedness. Log quantile–quantile (QQ) p-value plots were visually examined to determine the appropriateness of the association model. Log quantile–quantile (QQ) p-value plots were visually examined to assess the overall quality of the linear model and how effectively it corrected both population structure and family relatedness. Benjamini and Hochberg (1995) false discovery rate as implemented in GAPIT at the 1% level of significance was used to define associated SNPs. Manhattan and QQ plots were visualized using the CMplot package for R software (Yin 2022). The length of the genomic region enclosing the associated significance SNPs were determined based on the linkage disequilibrium (LD) analysis using the block function of PLINK (Purcell et al. 2007). Candidate genes within associated genomic regions were identified based on the annotations of BTx623 v3. sorghum genome reference (www.phytozome.com).
Phylogenetic analysis
The genetic relationship among Sudanese grain mold resistance and 15 reference lines from the SAP was determined based on clustering analysis. A subset of 1,818 unlinked SNPs (r2 < 0.10) with > 95% of the genotype data was produced using PLINK (Purcell et al. 2007). The IQ-TREE 2 software (Minh et al. 2020) was used to create the phylogenetic tree using the maximum-likelihood method and select the tree with ModelFinder (Kalyaanamoorthy et al. 2017). Branch supports were determined using the ultrafast bootstraping (Hoang et al. 2018) method as implemented in IQ-TREE 2 (Minh et al. 2020). The phylogenetic tree was visualized using Interactive Tree of Life (Letunic and Bork 2011).
Results
Grain mold resistance in NPGS Sudan core collection
The grain mold resistance response varied among NPGS Sudanese accessions. The grain mold evaluation of the core set at Isabela and Mayaguez, Puerto Rico showed that the seedling emergence rate and seed deterioration averaged 60.90% ± 19.44 and 3.51 ± 0.61, respectively (Supplementary Table S1). The analysis found that 46 and 12 accessions had seedling emergence rates of ≥ 80% and seed deterioration ≤ 2.5, respectively, indicating that most of the accessions in the NPGS Sudan core collection are susceptible to grain mold. The low repeatability of seedling emergence (0.55) and seed degradation (0.47) exhibited the large effect of environmental factors on both traits. In fact, the average temperature and relative humidity in Mayaguez (80.56° F and 84.27%) were higher than the observed in Isabela (70.67° F and 74.94%).
The four-year grain mold evaluation for the subset of 46 accessions confirmed multiple resistance sources. The seedling emergence rate of 38 accessions (> 82%) were similar to those observed in eight reference lines (Table 1; Rox Orange, SC309, SC609, Sumac, Sureño, Kansas Orange, Keller and PI 267548). In contrast, just two accessions (PI 568457 and PI 570382) exhibited lowest seed deterioration (< 1.77) similar to those observed in five reference lines (Table 2; Sumac, Red Amber, Roz Orange, SC15 and Keller). But other five accessions (PI 570702, PI 570776, PI 570685, PI 570330, and PI 570348) had also low seed deterioration (< 2.15) that were similar to those observed in the reference lines SC13 and PI 267548. These seven accessions with low seed deterioration (< 2.15) also exhibited high emergence rates (> 86%), thus, might be useful as new grain mold resistant sources in breeding programs.
The population structure of NPGS Sudan core collection and its phylogenetic relationship with other grain mold sources of resistance provided insight into how this new resistance germplasm could be exploited in breeding programs. Our analysis found that NPGS Sudan populations 5 and 1 enclose accessions with high seedling emergence rate (72.98 ± 12.36) and lower seed deterioration (3.11 ± 0.56), respectively (Table 3). In fact, 18 of the 38 accessions with high seedling emergence rate (> 82%) belong to population 5. Nevertheless, seed deterioration was high in the five populations (> 3.11) indicating that NPGS Sudan population structure is not associated with resistance to this trait. The phylogenetic analysis revealed a close genetic relationship between eleven Sudanese resistance accessions belonging to population 2 and reference lines (Fig. 1). Remarkably, the accessions PI 568457, PI 569151, PI 570685 and PI 569957 shared nodes with the reference lines Sumac, Keller, SC1494 and SC309, respectively. Moreover, these eleven Sudanese resistance accessions cluster with Kafir (PI 571108, PI 571109, PI 568457, PI 570944 and PI 571234) and Caudatum/Bicolor (PI 569151, PI 570685, PI 569957, PI 569342, PI 152615 and PI 563145) genetic background present in the SAP. The other 28 Sudanese grain mold resistance accessions belong to population 5 (27 accessions) and 3 (1 accession) and constitute a clearly separate cluster from the reference lines. Remarkable, the seven accessions with low seed deterioration (< 2.15) and high emergence rate (> 86%) were distributed across the phylogenetic tree suggesting the existence of different resistance sources.
Association analysis and candidate genes
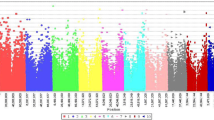
The association analysis for seedling emergence rate and seed deterioration led to the identification of two novel genomic regions associated with grain mold resistance (Table 4; Fig. 2). In chromosome 2, we identified a 38 bp genomic region associated with seedling emergence (Ch.2: 7,998,687; − log(p-value) = 7.56] that explained 14.02% of the phenotypic variance. This genomic region is within a 17 kb intergenic region flanked by the putative genes Sobic.002G076400 (mediator of RNA polymerase II transcription subunit 33A) and Sobic.002G076500 (Serine/threonine protein phosphatase). Moreover, we found that a 1.5 kb sequence with this intergenic region has homology with the Arabidposis gene AT2G48110 (E-value 2e−45) which is associated with the phenylpropanoid metabolic process. The association analysis for seed deterioration detected a 28.96 kb region in chromosome 3 [Ch3: 52,139,909; − log(p-value) = 7.43] that explains 35.2% of the phenotypic variance. This genomic region encloses the four putative genes (Sobic.003G196300, Sobic.003G196400, Sobic.003G196500 and Sobic.003G196600) that encode a nucleoplasmin ATPase (Sobic.003G196300), phosphorylase kinase (Sobic.003G196400) and two unknown functional proteins (Sobic.003G196500 and Sobic.003G196600).
Discussion
Sorghum, a drought tolerant crop is produced primarily in drought prone areas of Sudan (Abdalla and Gamar 2011). This country has been divided into five ecological zones and sorghum is mainly cultivated in three: the desert, semi-desert, and low-rainfall savannah zone (Mahgoub 2014). Nevertheless, the soil and climate are highly diverse within the low-rainfall savannah zone where the annual rainfall ranges from 300 to 800 mm and is highest in the south region. Thus, the low frequency of grain mold resistance observed in NPGS Sudan germplasm (16%) could be associated with the limited sorghum production in the south regions of Sudan. Certainly, seedling emergence is under selection by farmers in rural agriculture systems where a subset of their own seed is retained for the next production cycle. The seed quality and uniform seedling emergence affects the crop establishment, growth, and yield in many crop production systems (Finch-Savage 2020). In Zambia and Ethiopia, most farmers-saved sorghum seeds showed seedling emergency > 75%, suggesting that these farmers use adequate storage system, and their cultivars are adapted to the region (Tripp et al. 1998; Tsega 1994).
The high frequency of Caudatum race in the Sudanese sorghum germplasm make it one of the most valuable sorghum germplasms for worldwide breeding programs (Aruna and Cheruku 2019). Certainly, the panicle shape and high yielding observed in landraces belonging to this sorghum race appeal to the interest of sorghum breeder. Twenty-six of the grain mold resistance accessions identified herein belongs to Caudatum race and ten are Caudatum intermediate with Kafir (5 accessions), Guinea (4 accessions) and Bicolor (1 accession) races. Therefore, most of these resistant accessions must have valuable agronomic traits. In addition, three accessions with superior seedling emergence (PI 569957, PI 570824, and PI 570825) are resistant to anthracnose (Colletotrichum sublineola) (Cuevas and Prom 2020). These accessions together with the seven accessions with both high seedling emergence and low seed deterioration are the most valuable accessions for sorghum breeding programs.
The conversion of tropical landraces to temperate adapted germplasm is the first step to utilize new resistance sources in breeding programs. Through the Sorghum Conversion Program, [SCP; Stephens et al. (1967)] > 1000 tropical accessions were adapted to temperate regions of which Sudanese accessions were amongst the most selected germplasms. The phylogenetic analysis of both Sudanese and SAP grain mold resistance accessions revealed that some Sudanese resistance sources are likely accessible in temperate adapted germplasm. In fact, the temperate adapted lines SC1494 and SC309 originate from the tropical Sudanese accessions PI 570380 and PI 152594, respectively. Certainly, these two Sudanese accessions must be genetically related to PI 570685 and PI 569957 which cluster near to SC1494 and SC309, respectively. Sumac and Keller are both sweet sorghum improved germplasm whose pedigrees could be traced to Sudanese accessions PI 568457 and PI 569151, respectively. Therefore, sorghum conversion and pre-breeding programs need to be focused on grain mold resistance observed in PI 570382, PI 570776, PI 570330, PI 570702, and PI 570348 which belong to population 5 and could be classified as new sources of resistance.
The introgression of diseases resistance genes into temperate adapted lines is the most adequate strategy to utilize new resistant sources in breeding programs. The complex inheritance of grain mold resistant (Sharma et al. 2010; Rodriguez-Herrera et al. 2000) and its low heritability requires the evaluation of large segregating populations across multiple environments. In this regard, the selected grain mold resistant accessions need to be crossed with a common elite temperate adapted lines to establish multiple segregating populations. Later, the development of recombinant inbred lines (RILs) populations could lead to the identification of new resistant lines with superior agronomic traits and to find associations with other genomic regions with grain mold resistance responses. To accelerate the release of converted germplasm, the reinstated sorghum conversion program (RSC) utilized GBS analysis in early backcross generation (BC1) to select superior lines with genomic regions associated with temperate adaptation (Klein et al. 2013). This backcross system can also be employed with grain mold resistance accessions; however, it will be necessary to increase the population size because 75% of the genome will be derived from the susceptible temperate adapted line. Certainly, the development of pre-breeding germplasm (Sharma et al. 2013) with grain mold resistance and temperate adaptation could be the most adequate strategy to enhance genetic diversity of breeding programs.
The fixation of grain mold resistant and temperate adaptation (i.e. day-neutral) alleles at early generation could be effective for the development of new pre-breeding resistance germplasm. Flowering time and photoperiod sensitivity in sorghum are well understood, and its major genes have been cloned (Yang et al. 2014). But the inheritance of grain mold resistance is not well understood, and its genetic control varies across germplasm. For instance, the SNPs associated to seedling emergence and seed deterioration are not associated to previously identified loci in SAP (Cuevas et al. 2019), NPGS Ethiopian germplasm (Cuevas and Prom 2024) and Ethiopian landraces (Nida et al. 2019). The inconsistency of these results suggests different genetic backgrounds enclose diverse resistance mechanisms that might be combined for a broader resistance response. The genomic regions associated with seedling emergence and seed deterioration explain a limited portion of the phenotypic variance. However, both genomic regions enclose putative genes that could be associated with plant immunity response. For instance, the phenylpropanoid metabolic pathway is activated in plants as a response to pathogen attack to produce many broad-spectrum antimicrobial compounds (Troncoso-Rojas et al. 2013; Dixon et al. 2002). Indeed, the rapid degeneration of non-functional alleles in susceptible lines make not possible its annotation in the BTx623 reference genome. The phosphorylase kinase (Sobic.003G196400), like other kinases could be recognized as microbe elicitors or cell damage that activate the plant defense responses (Gao et al. 2008). We observed that the association of this locus is determined by the allelic variation found in population 5 where PI 570318, PI 570513 and PI 570335 have the resistant allele. The development of high-throughput SNP markers for this locus could be used for the early fixation of resistance allele in segregating population derived from these grain mold resistant accessions.
Conclusion
The results of this study demonstrate the existence of phenotypic and genetically valuable sorghum accessions in NPGS Sudan germplasm collection. The grain mold resistance evaluation of this germplasm identified 39 resistant accessions of which seven showed both high seedling emergence (> 82%) and low seed deterioration (< 2.15). Association analysis between grain mold resistance and NPGS Sudanese population structure revealed that most of these accessions belong to populations 2 and 5. Phylogenetic analysis determined that grain mold resistant accessions from population 2 are genetically related with resistant accessions present in the SAP. Thus, the five accessions (PI 570382, PI 570776, PI 570330, PI 570702, and PI 570348) from population five that showed both high seedling emergence and low seed deterioration are the most likely to be classified as new resistance sources for breeding programs. These five tropical grain mold resistant accessions must be first cross with elite germplasm to develop pre-breeding germplasm through pedigree or backcrossing method. The new pre-breeding germplasm should include grain mold resistance, non-photoperiod sensitive (i.e. temperate adaptation) and high yielding plants that could be used to enhance sorghum breeding programs. Genome-wide association analysis identified two genomic regions in chromosome 2 and 3 which enclose candidate genes associated with the plant defense responses. Both genomic regions associated with the resistance response provide insight into the molecular mechanisms although further functional genomics research is required to elucidate its actual role. The development of high throughput molecular markers within both genomic regions could be used to accelerate selection in breeding programs in parallel with phenotypic evaluations.
Data availability
The phenotypic datasets generated during the current study are available in the supplementary table and through the U.S. National Plant Germplasm System; GRIN-Global (https://www.grin-global.org).
References
Abdalla H, Gamar Y (2011) Climate change: selection of sorghum genotype with wide adaptation, AG-17, for rain-fed areas of Sudan. Int J AgriSci 1(3):144–155
Ackerman A, Wenndt A, Boyles R (2021) The sorghum grain mold disease complex: pathogens, host responses, and the bioactive metabolites at play. Front Plant Sci 12:660171. https://doi.org/10.3389/fpls.2021.660171
Aruna C, Cheruku D (2019) Genetic improvement of grain sorghum. In: Aruna C, Visarada KBRS, Venkatesh Bhat B, Tonapi VA (eds) Breeding sorghum for diverse end uses. Elsevier, Amsterdam, pp 157–173
Bandyopadhyay R, Butler DR, Chandrashekar A, Reddy RK, Navi SS (2000) Biology, epidemiology and management of sorghum grain mold. In: Chandrashekar A, Bandyopadhyay R, Hall AH (eds) Technical and institutional options for sorghum grain mold management: proceedings of an international consultation, 18–19 May 2000. ICRISAT, Patancheru, Andhara Pradesh, India, pp 34–71
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Stat Methodol 57(1):289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Bernardo R (2002) Breeding for quantitative traits in plants. Stemma Press, Woodbury
Brown AHD (1989) Core collections: a practical approach to genetic resources management. Genome 31:818–824. https://doi.org/10.1139/g89-144
Cuevas HE, Prom LK (2020) Evaluation of genetic diversity, agronomic traits, and anthracnose resistance in the NPGS Sudan sorghum core collection. BMC Genom 21(1):88. https://doi.org/10.1186/s12864-020-6489-0
Cuevas HE, Prom LK (2024) Association analysis of grain mould resistance in a core collection of NPGS Ethiopian sorghum germplasm. Plant Genet. Res: Characterization and Utilization. Published online 2024:1–10. https://doi.org/10.1017/S1479262124000157
Cuevas HE, Rosa-Valentin G, Hayes CM, Rooney LW, Hoffmann L (2017) Genomic characterization of a core set of the USDA-NPGS Ethiopian sorghum germplasm collection: implications for germplasm conservation, evaluation, and utilization in crop improvement. BMC Genom 18:108. https://doi.org/10.1186/s12864-016-3475-7
Cuevas HE, Prom LK, Rosa-Valentin G (2018) Population structure of the NPGS Senegalese sorghum collection and its evaluation to identify new disease resistant genes. PLoS ONE 13(2):e0191877. https://doi.org/10.3389/fpls.2021.660171
Cuevas HE, Fermin-Perez RA, Prom LK, Cooper EA, Bean S, Rooney WL (2019) Genome-wide association mapping of gain mod resistance in the US sorghum association panel. Plant Genome 12(2):180070. https://doi.org/10.3835/plantgenome2018.09.0070
Dahlberg JA, Burke JJ, Rosenow DT (2004) Development of a sorghum core collection: refinement and evaluation of a subset from Sudan. Econ Bot 58(4):556–567. https://doi.org/10.1663/0013-0001(2004)058[0556:DOASCC]2.0.CO;2
Dixon RA, Achnine L, Kota P, Liu CJ, Reddy MS, Wang L (2002) The phenylpropanoid pathway and plant defence—a genomics perspective. Mol Plant Pathol 3(5):371–390. https://doi.org/10.1046/j.1364-3703.2002.00131.x
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6(5):e19379. https://doi.org/10.1371/journal.pone.0019379
Fakrudin B, Lakshmidevamma T, Ugalat J, Gunnaiah R, Khan J, Gautham Suresh S, Apoorva K, Doddamani M, Kadam S, Rashmi K (2021) Genomic designing for biotic stress resistance in sorghum. In: Kole C (ed) Genomic designing for biotic stress resistant cereal crops. Springer, Cham, pp 213–255
Food and Agriculture Organization (2022) FAOSTAT. Accessed June 1, 2022
Faye JM, Maina F, Hu Z, Fonceka D, Cisse N, Morris GP (2019) Genomic signatures of adaptation to Sahelian and Soudanian climates in sorghum landraces of Senegal. Ecol Evol 9:6038–6051. https://doi.org/10.1002/ece3.5187
Finch-Savage W (2020) Influence of seed quality on crop establishment, growth, and yield. In: Gough RE, Basra AS (eds) seed quality. CRC Press, Boca Raton, pp 361–384
Forbes GA (1986) Characterization of grain mold resistance in sorghum [Sorghum Bicolor (L.) Moench] (histology, fluorescence microscopy). Texas A&M University
Gao M, Liu J, Bi D, Zhang Z, Cheng F, Chen S, Zhang Y (2008) MEKK1, MKK1/MKK2 and MPK4 function together in a mitogen-activated protein kinase cascade to regulate innate immunity in plants. Cell Res 18(12):1190–1198. https://doi.org/10.1038/cr.2008.300
Grenier C, Bramel-Cox PJ, Hamon P (2001) Core collection of sorghum: I. Stratification based on eco-geographical data. Crop Sci 41(1):234–240
Hoang DT, Chernomor O, Von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35(2):518–522. https://doi.org/10.1093/molbev/msx281
Isakeit T, Collins SD, Rooney WL, Prom LK (2008) Reaction of sorghum hybrids to anthracnose, grain mold and grain weathering in Burleson County, Texas, 2007. Plant Dis Manag Rep 2:FC003
Kalyaanamoorthy S, Minh BQ, Wong TK, Von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14(6):587–589. https://doi.org/10.1038/nmeth.4285
Klein RR, Miller FR, Klein PE, Burke JJ (2013) Registration of partially converted germplasm from 44 accessions of the USDA-ARS Ethiopian and Sudanese sorghum collections. J Plant Registrations 7(3):368–372. https://doi.org/10.3198/jpr2012.08.0025crgs
Letunic I, Bork P (2011) Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478. https://doi.org/10.1093/nar/gkab301
Lipka AE, Tian F, Wang QS, Peiffer J, Li M, Bradbury PJ, Gore MA, Buckler ES, Zhang ZW (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28(18):2397–2399. https://doi.org/10.1093/bioinformatics/bts444
Mahgoub F (2014) Current status of agriculture and future challenges in Sudan. Current African issues. Nordiska Afrikainstitutet, Uppsala, Sweden
Maina F, Bouchet S, Marla SR, Hu Z, Wang J, Mamadou A, Abdou M, Saïdou A-A, Morris GP (2018) Population genomics of sorghum (Sorghum bicolor) across diverse agroclimatic zones of Niger. Genome 61(4):223–232. https://doi.org/10.1139/gen-2017-0131
Mann JA, Kimber CT, Miller FR (1983) The origin and early cultivation of sorghum in Africa. Texas Agricultural Experiment Station Bulletin 1454, Texas A&M University, College Staion, TX
Menz MA, Klein RR, Unruh NC, Rooney WL, Klein PE (2004) Genetic diversity of public inbreeds of sorghum determined by mapped AFLP and SSR markers. Crop Sci 44:1236–1244. https://doi.org/10.2135/cropsci2004.1236
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37(5):1530–1534. https://doi.org/10.1093/molbev/msaa015
Morales-Salva A (2022) Genome-wide association analysis of anthracnose resistance in NPGS Yemen sorghum [Sorghum bicolor (L.) Moench] germplasm. University of Puerto Rico-Mayaguez Campus, Mayaguez
Morris GP, Ramu P, Deshpande SP, Hash CT, Shah T, Upadhyaya HD, Riera-Lizarazu O, Brown PJ, Acharya CB, Mitchell SE, Harriman J, Glaubitz JC, Buckler ES, Kresovich S (2013) Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc Natl Acad Sci USA 110(2):453–458. https://doi.org/10.1073/pnas.1215985110
Murphy RL, Klein RR, Morishige DT, Brady JA, Rooney WL, Miller FR, Dugas DV, Klein PE, Mullet JE (2011) Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum. Proc Natl Acad Sci USA 108(39):16469–16474. https://doi.org/10.1073/pnas.1106212108
Navi SS, Bandyopadhyay R, Reddy RK, Thakur RP, Yang XB (2005) Effects of wetness duration and grain development stages on sorghum grain mold infection. Plant Dis 89(8):872–878. https://doi.org/10.1094/PD-89-0872
Nida H, Girma G, Mekonen M, Lee S, Seyoum A, Dessalegn K, Tadesse T, Ayana G, Senbetay T, Tesso T, Ejeta G, Mengiste T (2019) Identification of sorghum grain mold resistance loci through genome wide association mapping. J Cereal Sci 85:295–304. https://doi.org/10.1016/j.jcs.2018.12.016
Olatoye MO, Hu ZB, Maina F, Morris GP (2018) Genomic signatures of adaptation to a precipitation gradient in nigerian sorghum. G3 Genes Genom Genet 8(10):3269–3281. https://doi.org/10.1534/g3.118.200551
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Prom LK, Erpelding J (2009) New sources of grain mold resistance among sorghum accessions from Sudan. J Trop Subtrop Agroecosyst 10:457–463
Prom LK, Cuevas HE, Ahn E, Isakeit T, Rooney WL, Magill C (2020) Genome-wide association study of grain mold resistance in sorghum association panel as affected by inoculation with Alternaria alternata alone and Alternaria alternata, Fusarium thapsinum, and Curvularia lunata combined. Eur J Plant Pathol 157(4):783–798. https://doi.org/10.1007/s10658-020-02036-3
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575. https://doi.org/10.1086/519795
Rodriguez-Herrera R, Rooney WL, Rosenow DT, Frederiksen RA (2000) Inheritance of grain mold resistance in grain sorghum without a pigmented testa. Crop Sci 40(6):1573–1578. https://doi.org/10.2135/cropsci2000.4061573x
Sharma R, Rao V, Upadhyaya H, Reddy VG, Thakur R (2010) Resistance to grain mold and downy mildew in a mini-core collection of sorghum germplasm. Plant Dis 94(4):439–444. https://doi.org/10.1094/PDIS-94-4-0439
Sharma S, Upadhyaya HD, Varshney RK, Gowda C (2013) Pre-breeding for diversification of primary gene pool and genetic enhancement of grain legumes. Front Plant Sci 4:309. https://doi.org/10.3389/fpls.2013.00309
Snowden JD (1936) The cultivated races of sorghum. Adlard and Son, London
Stephens JC, Miller FR, Rosenow DT (1967) Conversion of alien sorghums to early combine genotypes. Crop Sci 7(4):396. https://doi.org/10.2135/cropsci1967.0011183X000700040036x
Thakur RP, Reddy BVS, Indira S, Rao VP, Navi SS, Yang XB, Ramesh S (2006) Sorghum grain mold. In: Information bulletin No. 72. International crops reserarch institute for the semi-arid tropics, Patancheru, India
Thakur RP, Rao VP, Reddy BVS, Sanjana Reddy P (2007) Grain mold. In: Thakur RP, Reddy BVS, Mathur K (eds) Screening techniques for sorghum diseases, vol Bulletun # 76. International crops research institute for the semi-arid tropics, Patancheru, India
Tripp R, Walker D, Miti F, Mukumbuta S, Zulu M (1998) Seed management by small-scale farmers in Zambia. A study of cowpea, groundnut and sorghum seed in the southern and western provinces (NRI Bulletin 76)
Troncoso-Rojas R, Sánchez-Estrada A, Carvallo T, González-León A, Ojeda-Contreras J, Aguilar-Valenzuela A, Tiznado-Hernández M-E (2013) A fungal elicitor enhances the resistance of tomato fruit to Fusarium oxysporum infection by activating the phenylpropanoid metabolic pathway. Phytoparasitica 41:133–142. https://doi.org/10.1007/s12600-012-0271-z
Tsega M (1994) An inventory and investigation of the optimum local seed storage methods in Wello and Shewa administrative regions. Consultancy Report, Seeds of Survival Program, USC Canada in Ethiopia
Upadhyaya HD, Pundir RPS, Dwivedi SL, Gowda CLL, Reddy VG, Singh S (2009) Developing a mini core collection of sorghum for diversified utilization of germplasm. Crop Sci 49(5):1769–1780. https://doi.org/10.2135/cropsci2009.01.0014
Yang S, Murphy RL, Morishige DT, Klein PE, Rooney WL, Mullet JE (2014) Sorghum phytochrome B inhibits flowering in long days by activating expression of SbPRR37 and SbGHD7, repressors of SbEHD1, SbCN8 and SbCN12. PLoS ONE 9(8):e105352. https://doi.org/10.1371/journal.pone.0105352
Yin L (2022) CMplot: circle manhattan plot. R package version 4.2. 0.
Funding
This research was funded by the USDA-ARS Current Research Information System (Project 6090-21000-053-00-D; HC). Authors thank to USDA, ARS, Plant Genetic Resources Conservation Unit at Griffin, Georgia U.S. for providing the Sudanese sorghum germplasm.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Hugo E. Cuevas and Louis K. Prom. The first draft of the manuscript was written by Hugo E. Cuevas and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cuevas, H.E., Prom, L.K. The NPGS Sudanese sorghum core collection encloses novel grain mold resistant germplasm. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-02039-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-02039-7