Abstract
Benthic phototrophic communities play a crucial role in the functioning of shallow lake ecosystems. The complexity of benthic algal communities poses challenges for exploring their fine structures using traditional methods such as light microscopy. However, confocal laser scanning microscopy (CLSM) offers the potential for microscale-level structural investigation of phototrophic biofilms, thereby contributing to a deeper understanding of the structural and compositional changes within these communities. The primary objective of this study was to develop a relatively rapid, semi-automated, and freely available image processing method to facilitate the tracking of spatio-temporal alterations in intact biofilms. To evaluate our image processing method, we conducted an eight-week long outdoor mesocosm experiment, manipulating temperatures based on two global warming scenarios (assuming intermediate- and high degree of greenhouse gas emission), with increases of 3°C and 5°C, respectively, compared to the reference/unheated mesocosms. Eight limestone cubes were placed on the bottom of each mesocosm to provide 'natural' surfaces for phytobenthos colonization, which was subsequently analysed by confocal microscopy. We hypothesized that increasing water temperature might have a significant impact on the taxonomic composition of algal biofilms, particularly on the abundance of phototrophic prokaryotes and eukaryotes. The designed 'FIJI macro' efficiently analysed the recorded images, including single images, series of Z-stack images, and projection images. Our image processing method effectively discriminated photosynthetic prokaryotes and eukaryotes based on their pigment composition and proved suitable for the high-throughput analysis of structural and compositional changes in algal biofilms. Moreover, this study confirmed that climate-induced warming can lead to an increasing dominance of cyanobacteria in benthic phototrophic communities at the expense of eukaryotic algae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Benthic communities are essential components of healthy aquatic ecosystems as they play a fundamental role in maintaining ecological integrity and biodiversity, in addition to ecosystem services they provide. In benthic habitats, photoautotrophic microorganisms have an important functional role in primary production, food web interactions, benthic-pelagic coupling, and nutrient exchange across the water-sediment interface (Crespo and Pardal 2020). Moreover, benthic diatom communities have been used regularly for monitoring the ecological status and trends of changes in aquatic systems (Serôdio and Lavaud 2020; B-Béres et al. 2023). Benthic phototrophic communities of shallow lakes are commonly dominated by the representatives of cyanobacteria, diatoms, and green algae (Stevenson 1996). These microorganisms are usually attached to solid submerged surfaces and are embedded in a mucilaginous matrix composed of extracellular polymeric substances (EPS) and organic detritus. EPS is a three-dimensional structure consisting primarily of polysaccharides, and has a major role in the development of algal biofilms (Wu 2017a). Biofilm formation is a complex process, generally divided into five main steps: (1) initial attachment; (2) irreversible attachment; (3) colonization; (4) maturation, and (5) dispersion (Sehar and Naz 2016; Osorio et al. 2021; Sauer et al. 2022). Structure of phototrophic benthic communities can change dynamically and is shaped by a wide range of environmental factors, such as nutrient- and light availability, temperature, hydrodynamic conditions, and properties of the substratum they are attached to (Wu 2017b; Fanesi et al. 2019; 2022; Osorio et al. 2021). In addition to these abiotic variables, some biotic factors (e.g., interactions between algal species, microbial succession) can also affect the formation and structure of benthic biofilms (Fanesi et al. 2019; Guo et al. 2021; Krsmanovic et al. 2021; Lamprecht et al. 2022). The complexity and opaqueness of algal biofilms make it difficult to explore their fine structures using ‘traditional’ methods, such as light microscopy (Azeredo et al. 2017).
Since its commercialisation in the late 1980’s, confocal laser scanning microscopy (CLSM) has been applied in various fields of science like applied physics, life sciences, material sciences, and engineering. For now, CLSM has become one of the most widespread fluorescent imaging techniques for three-dimensional (3D) structural analysis of cells as well as tissues (Paddock and Eliceiri 2014; Bayguinov et al. 2018). Among other applications, the development of advanced optical imaging techniques allows the structural investigation of phototrophic biofilms at microscale level (Neu et al. 2010). Studying the 3D architecture of algal biofilms may contribute to a better understanding of their functional and ecological roles, as well as their responses to changing environmental conditions (Zancarini et al. 2017; Lamprecht et al. 2022; Manirakiza et al. 2022).
In practice, CLSM can serve as a non-invasive, non-destructive, label-free, and cost-efficient method for obtaining high-resolution images of intact biofilms (Mueller et al. 2006; Neu et al. 2010). This technique uses a focused laser beam to scan the sample and, by means of using a spatial pinhole, captures fluorescence images at different depths in a sample (Kuehn et al. 1998; Achinas et al. 2020), allowing the reconstruction of 3D images. However, like any other methods used for studying algal biofilms, CLSM also involves some specific challenges and limitations. For example, resolution of CLSM may not be sufficient to resolve fine structural details within dense and heterogeneous samples, particularly in thicker sections where axial resolution degrades. Very thick and highly scattering algal biofilms can limit the penetration depth of CLSM and restrict the ability to effectively visualize the deeper layers. In tissue samples, the effective imaging depth is typically around 100-200 µm (Pawley 2006; Neu and Lawrence 2015). High laser intensities and/or prolonged exposure to the laser light can damage living cells within the biofilms and affect biofilm integrity (Dailey et al. 2006).
Importantly, eukaryotic algae and prokaryotes (cyanobacteria) have different pigment compositions which strongly influence their fluorescence properties; the autofluorescence of eukaryotic algae is dominated by that of chlorophylls, while cyanobacterial autofluorescence is composed of two spectrally distinct components, i.e., of chlorophylls and phycobiliproteins. Hence, although CLSM is typically not suitable for accurate species identification, some algal groups and photosynthetic prokaryotes can be distinguished based on their spectral properties (Carreira et al. 2015; Lamprecht et al. 2022).
Following structural and compositional changes over time can provide an insight into the dynamics of the formation/ageing of algal biofilms (Papageorgiou 2004). A major goal of our work was to develop a rapid, straightforward and freely available CLSM image analysing method in order to track the spatio-temporal changes in algal biofilms. For a high throughput workflow, we constructed a ‘macro’ set (macro: a program that automates a series of commands, e.g., in an image processing environment). To validate this method and to test the image processing macro we developed, major structural and/or compositional changes/differences in the studied algal biofilms were a prerequisite. Previous studies have demonstrated that climate change-related increase in water temperature can cause substantial alterations in the distribution, composition and diversity of aquatic organisms, as well as in their biomass production and growth rates (Daufresne et al. 2009; Capon et al. 2021; Pörtner et al. 2022). Optimal growth temperature of different algae can vary over a wide range. In the temperate climate zone, this range is generally between 15 and 30°C (Ras et al. 2013; Singh and Singh 2015). As a consequence of global warming, there could be a shift in the taxonomical composition of natural algal communities in the temperate climate zone, manifested in the increasing of dominance cyanobacteria at the expense of diatoms and chlorophytes (DeNicola 1996; Nalley et al. 2018). Building upon this observation, we hypothesized that increasing temperature might exert significant effects on the species composition of algal biofilms under experimental conditions too, leading to an anticipated increase in cyanobacterial abundance along with the decline of eukaryotic photoautotrophs. To simulate the impact of elevated temperature on benthic algal communities, we conducted an outdoor mesocosm experiment, where water temperatures were manipulated based on two global warming scenarios issued by the Intergovernmental Panel on Climate Change, assuming intermediate (SSP2-4.5) and high degree of greenhouse gas emissions (SSP5-8) (Pörtner et al. 2022). The applied heating regimes corresponded to 3°C and 5°C higher average water temperatures compared to the unheated (reference) treatment. Shallow lakes are especially vulnerable to climate change because these water bodies are ‘isolated’ and fragmented in the landscape, lake temperature and water supply are strongly and rapidly affected by ambient warming, many species living in these habitats have limited abilities to range shifting, and most of the shallow lakes are already exposed to the multitude of anthropogenic stressors (Woodward et al. 2010b; Meerhoff et al. 2012).
Materials and methods
Mesocosm study design
An 8-week-long experiment was conducted in the outdoor mesocosm facility (see Fig. 1) of the Balaton Limnological Research Institute (Tihany, Hungary), between 8 July and 2 September 2021. This system comprised 12 cylindrical plastic tanks, each with a maximum volume of approximately 5 m3 and a water depth of 1.6 m.
Two months prior to the experiment's commencement, limestone bricks (a total of 24 pieces, each with a surface area of 100 × 100 mm) were positioned in the littoral zone of Lake Balaton (Hungary) to allow for the growth of natural biofilm. Subsequently, the surfaces of these bricks were scraped into a beaker to obtain a living sample for inoculation; 50 mL of this bulk sample was then added to each tank to initiate biofilm formation. On the same day (8 July), the mesocosm tanks were filled with unfiltered water from Lake Balaton to a depth of 0.8 m resulting in a water volume of 2.5 m3. Additionally, eight limestone cubes, each measuring 10×10×10 mm were placed on the bottom of each mesocosm tank to provide 'natural' surfaces for phytobenthos colonization. One of these cubes was removed weekly from each mesocosm for the analysis of biofilm development.
Water temperatures in the heated tanks were raised by +3°C and +5°C, respectively, compared to the reference/control tanks (maintained at ambient temperature). Each treatment consisted of four replicates, resulting in a total of 12 experimental units. Water temperature was continuously monitored at 10-minute intervals using PONSEL (type: PPHRB-3931; AQUALABO, Champigny-sur-Marne, France) sensors installed in each mesocosm.
Sample preparation for imaging
After immersion into liquid agar in order to fix motile cells, the limestone substrates were placed upside-down onto a Greiner Bio-One 62861 CELLview Cell Culture Dish (Greiner AG, Austria). Five regions of interest were randomly selected on each substrate surface, one at the centre and four around it. During the confocal optical sectioning (confocal imaging) the series of images were taken between the inner (limestone surface) and outer layer of the biofilm. The actual thickness of the biofilm was determined as a distance between the two farthest layers consisting of fluorescence information in the Z sectioning.
CLSM
Benthic biofilms were imaged using a Leica SP8 confocal microscope (Leica Microsystems, Germany) equipped with a ‘HC PL APO CS’ 40× dry long working distance objective with a numerical aperture of 0.85. Images with a size of 1024 × 512 pixels were recorded, covering an area of 384 μm × 192 μm. The 375 nm pixel size was in accordance with the approximate 380 nm resolution of the imaging system considering both the applied objective and the detection wavelengths. Chlorophyll and phycobilisome (PBS) autofluorescence were excited using the 488 and 638 nm laser lines, respectively, and were detected over the 690–790 nm (with artificial red colour on the figures) and 650–680 nm (with artificial green colour on the figures) spectral windows. The sensitivity of the detectors (photomultiplier tubes, PMTs) was constant for all our measurements: PMT1 (Chl) gain: 720 V and PMT2 (PBS) gain: 750 V. Scan speed was specified as 400 Hz. A triple band 488/552/638 main beam splitter was applied for separating the reflected excitation light and the fluorescence. The 552 nm middle band of the dichroic mirror which was not used for a laser line was sufficiently far from the spectral range of the acquired fluorescence as it has not affected the imaging. During the optical sectioning (Z series of the confocal acquisition) the distance of the subsequent focal layers was 5 μm.
Image processing
Images were processed using the FIJI (Schindelin et al. 2012) (version 1.52c) open source Java-based image processing program. Due to the high number of bundled plugins, FIJI is capable of importing and processing Leica native microscope image format (LIF). For dealing with the images, we constructed a macro set (attached as a supplement) containing step-by-step tasks of the image processing. Macro element (Set Threshold method) generates a dialog window for determining the minimum intensity thresholds for separate recognition of the prokaryotic and eukaryotic microorganisms as well as to adjust the corresponding channel-intensity parameters. These parameters were determined/tested on some randomly selected images in order to have parameter values for the batch processing. At this point the user can decide whether to project the Z slices onto one layer or perform the calculation layer by layer.
Macro elements deal with one single file (testing purposes), and with all the images of a selected directory, respectively. Both macros call the makeSeparation() and the calcRatio() functions (defined in the same macro set file). The makeSeparation() function performs the Z projection if this option had been selected earlier and evaluates the channels according to the pixel intensity ratios (Strašková et al. 2019). The macro language of ImageJ allows defining complex conditions as well, not simply thresholding of intensity ratios. The output classes (and the corresponding output colours) are the following: a) low intensity / black (background, noise, insufficient fluorescence signal), b) intensive chlorophyll fluorescence without significant PBS contribution / green (eukaryotic organisms) c) both chlorophyll and PBS fluorescence signals are intensive (prokaryotic organisms / red) d) pixel ratios are not classified using the given parameters (“other” / blue). In case of optimal initialization, this “other” / blue category remains empty. Then, the calcRatio() function calculates the “ratio of pixel numbers” falling under the category b) and c) (not ratio of fluorescence intensity) and provides a prokaryotic to eukaryotic ratio on a segmented or Z-projected image.
Statistical analyses
Treatment effects were examined on number of pixels per unit image area and on green/red pixel ratio using one-way ANOVA, followed by Tukey’s post-hoc tests. The number of artificial green and red colour pixels, as well as the green/red pixel ratio followed lognormal distribution; therefore, these variables were logarithmically transformed. In order to avoid losing zero values, +1 were added to each value of the log transformed variables. Even though the dependent variables in the models neither followed a normal distribution according to Shapiro-Wilk tests, nor had homogeneous variance according to Bartlett’s tests, the high sample sizes allowed us to perform robust ANCOVA tests. Dependent variables were number of artificial green colour pixels (log), number of artificial red colour pixels (log) and green/red pixel ratio (log), while treatment (unheated; +3°C; +5°C) was the independent variable. In each model ‘Date’, ‘Chlorophyll-a concentration’ and ‘Layer thickness’ were involved as covariates. Tank number was included in each model as random factor. All statistical analyses were performed in the R environment (Core Team R 2021), by using “car”, “nlme”, and “multcomp” packages (Pinheiro and Bates 2006; Hothorn et al. 2008; Fox and Weisberg 2018).
Results
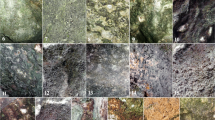
Some characteristic CLSM images (both unprocessed and processed) are shown in Fig. 2. These sample images were recorded at the beginning (15 Jul 2021), in the middle (5 Aug 2021) and at the end (2 Nov 2021) of the experiment.
Maximum intensity projection of Z-stack images from algal biofilm (a-c, g-i, m-o) captured by confocal laser scanning microscopy and the subsequently processed images (d-f, j-l, p-r) using our developed FIJI macro at the beginning (15/7/2021; 1st week), in the middle (5/8/2021; 4th week) and at the end (2/9/2021; 8th week) of the experiment. Autofluorescence of chlorophyll and phycobilisome were excited using the 488 and 638 nm lasers and were detected over 690–790 nm and 650–680 nm spectral windows. On the processed images, artificial green and red colour pixels indicate the photosynthetic prokaryotes and eukaryotes, respectively
During the experiment, the mean water temperature was 23.1±2.7°C (average±standard deviation) in the unheated (control) tanks and it varied between 17.0°C and 27.9°C (Fig. 3, red line). In the heated tanks (+3°C and +5°C constantly warmer relative to the control), water temperature varied between 20.0 and 30.5°C and between 22.0 and 32.5°C, with a mean of 26.1±2.7°C and 28.1±2.7°C, respectively (Fig. 3).
On each CLSM image, prokaryotic and eukaryotic photoautotrophs were discriminated based on their distinct autofluorescent properties, using a custom developed FIJI macro (see section Image processing). On the processed images the numbers of artificial green and red colour pixels indicate the relative amount of prokaryotic and eukaryotic photoautotrophs, respectively. Plotting the results of such quantification over time provides an insight into the dynamics of a forming phototrophic biofilm (Figs. 4 and 5).
(a) Temporal pattern of the eukaryotic abundance within forming phototrophic biofilm at different temperatures. Error bars represent standard deviations. (b). Box-plot diagram of the effect of temperature treatments on the eukaryotic abundance within forming phototrophic biofilm. The boxes on panel (b) denote the IQR; whiskers represent the minimum and maximum values, if it is within 1.5 IQR from the boxes. Thick lines through the boxes represent the medians; central rhombus symbols show the means. Asterisk denotes significant difference (*P< 0.05, **P<0.01) according to one-way ANOVA test, followed by Tukey’s post-hoc test. In both panels the number of artificial red colour (eukaryotic) pixels in the processed images are log-transformed (ncontrol = 2004, ncontrol+3°C = 2061, ncontrol+5°C = 2300)
(a) Temporal pattern of the prokaryotic abundance within forming phototrophic biofilm at different temperatures. Error bars represent standard deviations (b). Box-plot diagram of the effect of temperature treatments on the prokaryotic abundance within forming phototrophic biofilm. The boxes on panel (b) denote the IQR; whiskers represent the minimum and maximum values, if it is within 1.5 IQR from the boxes. Thick lines through the boxes represent the medians; central rhombus symbols show the means. Asterisk denotes significant difference (***P<0.001) according to one-way ANOVA test, followed by Tukey’s post-hoc test. In both panels the number of artificial green colour (prokaryotic) pixels in the processed images are log-transformed (ncontrol = 2004, ncontrol+3°C = 2061, ncontrol+5°C = 2300)
Both the eukaryotic and prokaryotic abundance (number of artificial red and green colour pixels per unit image area, respectively, after image processing) changed in a wide range in all temperature treatments (Figs. 4 and 5). The amount of artificial red colour pixels was in the range of 415-55090, 1575-57935 and 147-73740 with an average of 15192±9882, 14719±11127 and 12378±13254 in the control, +3°C and +5°C treatments, respectively (Fig. 4a). Number of artificial green colour pixels changed between 1152-186101 in the control, 2087-96116 in the ‘+3°C’ treatment and 46-309419 in the ‘+5°C’ treatment, with a mean of 21025±22828, 30829±22318 and 75578±63824, respectively (Fig. 5a). Thus, the eukaryotic abundance (number of artificial red colour pixels) decreased with increasing temperatures with concomitant increase of prokaryotic density (number of artificial green colour pixels) with increasing temperatures (Figs. 4 and 5).
Temperature treatment had a significant effect on both the eukaryotic (number of artificial red colour pixels; Df= 2(9), F=4.03, p=0.05) and prokaryotic abundance (number of artificial green colour pixels; Df=2(9), F=15.28, p=0.001). Between the control and the +3°C treatment, neither the prokaryotic nor the eukaryotic abundance (number of artificial green and red colour pixels, respectively) differed significantly (all p> 0.1), whilst the +5°C treatment differed significantly from both other treatments (control – control+5°C: pred=0.003, pgreen<0.001; control+3°C – control+5°C: pred=0.046, pgreen<0.001; Figs. 4b and 5b). Sampling time (Date) also had a significant effect on both the eukaryotic (Df= 7(457), F=16.828, p<0.0001) and prokaryotic abundance (Df=7(457), F=15.908, p<0.0001). Chlorophyll-a concentration (see Supplementary Table S1.) had a significant effect on eukaryotic abundance (Df=1(457), F=30.431, p<0.0001), but not on prokaryotic abundance (Df=1(457), F=0.807, p=0.370), while layer thickness (see Supplementary Table S2.) was a significant factor in explaining the variance in prokaryotic (Df=1(457), F=13.743, p=0.0002) but not in eukaryotic (Df=1(457), F=1.204, p=0.273) abundance. Overall, the regression model explained 27% of the variation in eukaryotic abundance (R2=0.27) and 38% of the prokaryotic abundance (R2=0.38).
The relative prokaryotic to eukaryotic ratio (green-to-red pixel ratio on the processed images) also varied in a broad range from 0.108 to 9.566 in the control, from 0.135 to 22.167 in the +3°C, and from 0.230 to 102.520 in the +5°C treatments with an average of 1.619±1.351, 3.320±3.628 and 13.664±18.000, respectively (Fig. 6a).
(a) Temporal pattern of prokaryote/eukaryote ratio (log-transformed ratio of artificial green and red colour pixels on the processed images) at different temperatures. Error bars represent standard deviations. (b). Box-plot diagram of the effect of temperature treatments on prokaryote/eukaryote ratio. The boxes on panel (b) denote the interquartile range (IQR); whiskers represent the minimum and maximum values, if it is within 1.5 IQR from the boxes; data points beyond this range (outliers) are indicated by circles. Thick lines through the boxes represent the medians; central rhombus symbols show the means. Asterisk denotes significant difference (***P<0.001) according to one-way ANOVA test, followed by Tukey’s post-hoc test. In both panels the green-to-red pixel ratio (relative prokaryotic to eukaryotic ratio) in the processed images are log-transformed (ncontrol = 2004, ncontrol+3°C = 2061, ncontrol+5°C = 2300)
Temperature treatment had a significant effect on prokaryote/eukaryote ratio (Df=2(9), F=24.04, p<0.001), i.e. it increased continuously with increasing temperatures (Fig. 6b). There was a marginally significant difference between the control and +3°C treatment (p=0.053), while the +5°C treatment differed significantly from both other treatments (control – control+5°C: p<0.001, control+3°C – control+5°C: p<00.1; Fig. 6b). Sampling time (Date) (Df=7(457), F=7.513, p<0.0001), chlorophyll-a concentration (Df=1(457), F=14.609, p=0.0002) and layer thickness (Df=1(457), F=5.383, p=0.02) also had a significant effect on the prokaryote/eukaryote ratio. 41% of the variance in prokaryote/eukaryote ratio was explained by the independent variables (R2=0.41).
Discussion
Establishing the current number of CLSM image analysis tools is challenging due to ongoing development and continual release of new tools, making a comprehensive comparison difficult. Nevertheless, these specialized software tools and plugins provide a range of functionalities from basic image processing to advanced quantitative analysis. Due to their robustness and specialized features, such tools are also widely used in studying biofilms.
Mhade & Kaushik (2023) provide a comprehensive review of recently emerged image analysis programs, which can also be used in quantitative analysis of biofilms using confocal micrographs. These image analysis tools vary not only in their scope and relevance to specific biofilm features but also in their compatibility with various operating systems, user interfaces, and raw image requirements.
Our selected tool for analysing CLSM images, the FIJI (ImageJ) software family has being developed for decades. A large, active user community contributes to ongoing development, support, and comprehensive documentation, making it easy to find solutions and troubleshoot issues. It is available for free on different operating systems (Windows, Linux, MacOS), and its software packages include many practical applications. FIJI allows users to create and customize macros and plugins tailored to specific image processing needs, including those unique to algal biofilms. FIJI is compatible with a wide range of native microscope file formats, supporting the reading of metadata and scale information. It also integrates seamlessly with other software tools and workflows commonly utilized in microscopy. In summary, FIJI macros offer a highly customizable and cost-effective solution for processing CLSM images of algal biofilms (Schindelin et al. 2012).
Yet, we must consider that FIJI also has some disadvantages. In fact, it may be challenging to use FIJI without some experience in programming or in scripting languages (such as ImageJ macro language, Python, or Java), which might limit its accessibility to users. FIJI might take a long time to process large datasets, such as those from CLSM analysis of thick biofilms. This slowness can be due to the software's processing algorithms, the size of the dataset, or limitations in the computer's hardware. Handling large image files requires a significant amount of memory (RAM). FIJI may struggle with these demands, leading to problems such as running out of memory or experiencing crashes. This is especially problematic when dealing with high-resolution 3D images, as they consume a large amount of memory during processing and analysis (Schindelin et al. 2012; Hartmann et al. 2021).
Major commercial software products, like Imaris and Velocity, provide advanced visualization tools, user-friendly interfaces, robust 3D and 4D analysis, and excellent support and training resources. However, they are generally expensive, require high-performance computers, and are less customizable than FIJI for specific tasks. Some of the frequently used image analysis tools are based on Matlab (e.g., PHLIP), requiring knowledge of the Matlab environment and, as commercial software, also can entail significant licensing costs. While other Matlab-built image analysis tools aim to be user-friendly and are intended to work independently of Matlab (e.g., COMSTAT as a ported ImageJ plugin and BiofilmQ as a standalone software), their application may still require to go through a learning curve to understand their functionalities and optimal setup for specific analysis tasks (Heydorn et al. 2000; Mueller et al. 2006; Hartmann et al. 2021; Mhade and Kaushik 2023).
In contrast to the aforementioned image analysis tools, the macro we developed is freely accessible, does not require extensive programming knowledge, and is an easy-to-use tool. This custom-developed method provides semi-automated processing for CLSM images and offers high throughput capabilities. It is optimized for analysing the structure and composition of algal biofilms and is well-suited for distinguishing photosynthetic prokaryotes and eukaryotes based on their auto-fluorescence properties. The ‘FIJI macro’ is capable of processing even single images, layers of Z-stack images, or projection images.
While traditional light microscopic examination methods allow for highly accurate species composition analysis, the common practice of scraping samples from surfaces eliminates the spatial information on the biofilm structure (Greenwood et al. 1999). In contrast, scanning electron microscopy (SEM) can offer detailed information about the structure of benthic algal communities. However, due to the fixation method applied, SEM is unsuitable for real-time analysis of living samples. Additionally, SEM is typically better suited for visualizing hard-shelled algae such as diatoms and provides only snapshots from a single vertical or horizontal plane (Larson and Passy 2005). Unlike the aforementioned methods, CLSM enables non-destructive, real-time, three-dimensional visualization of algal biofilms. Due to its inherent properties, it is particularly useful for examining the structural organization of living samples (Elliott 2020). Understanding the spatial organization and patch dynamics of biofilms can reveal important ecological processes such as colonization, population dynamics, interspecific interactions, and succession. Moreover, this microscopic technique can also aid in understanding how benthic algal communities respond to environmental changes (Larson and Passy 2005).
Temperature plays a crucial role in shaping the distribution and abundance of various species by exerting significant effects on biological and biochemical processes such as metabolism (Schulte 2015). At the individual level, increasing temperature can boost photosynthetic- and respiration rates in primary producers. At the population level, since growth is an integral component of metabolism, higher temperatures can also enhance growth rates (DeNicola 1996; Schulte 2015; Zhao et al. 2023). Different species of algae exhibit varying temperature preference for their photosynthesis and growth. Diatoms, for example, have the lowest thermal optima (24.0 ± 0.4°C), followed by chlorophytes (25.7 ± 0.1°C) and cyanobacteria (30.6 ± 2.3°C) (DeNicola 1996; Nalley et al. 2018). In our experiment, the temperature conditions in the control treatment typically favoured the growth of diatoms and green algae, whereas water temperatures in the heated tanks, particularly in the +5°C treatment, promoted the growth of cyanobacteria.
At the ecosystem level, temperature can have a significant impact on biomass and primary production (Daufresne et al. 2009; Woodward et al. 2010b). Typically, periphyton biomass is expected to increase in the temperature range between 0 and 30°C, and tends to decline at higher temperatures. In the temperate climate zone, periphyton generally reaches its peak biomass in spring and late summer / early fall (DeNicola 1996; Liu et al. 2021). However, previous studies have yielded inconsistent results regarding the response of periphyton biomass to global warming. Positive, negative, and neutral effects have all been reported (Mahdy et al. 2015; Kazanjian et al. 2018; Hao et al. 2020; Liu et al. 2021; Pacheco et al. 2022). Furthermore, disparities between different methods used for biomass determination further complicate the comparability of data across studies.
At the community level, temperature can induce alterations in the species composition and community structure of phytobenthos in aquatic ecosystems. Climate change has been shown to have a decidedly negative impact on species composition, diversity, and functioning of freshwater biotic communities (Woodward et al. 2010a; 2010b). Climate change is reshaping the distribution and interactions of species across various ecosystems, directly influencing population dynamics (Pearce-Higgins et al. 2015). Lengyel et al. (2023) reported that the SSP2-4.5 global warming scenario (+3 °C increase in average temperature compared to the present state) issued by the Intergovernmental Panel on Climate Change moderately affected phytobenthos in terms of biodiversity loss, while the occurrence of SSP5-8.5 scenario (+5 °C) could lead to significant structural and functional changes. These changes include a decline in species richness and loss of functional diversity, along with an increase in the abundance of filamentous life-forms and a decrease in the abundance of Bacillariophyta. Regarding the ecological implications of our findings revealed by analysing CLSM images, we conclude that a 3°C increase in water temperature had a moderate effect on the phytobenthic community, i.e., the ratio of prokaryotes to eukaryotes (green-to-red pixel ratio) showed only marginally significant difference compared to the Control treatment (Fig. 5b). In turn, a 5°C increase in water temperature induced severe changes in the composition of algal biofilms, favouring the dominance of photosynthetic prokaryotes.
These observations are in agreement with previous studies demonstrating that warming has a selective effect on promoting cyanobacterial growth and support the notion that relatively high temperatures provide a distinct advantage to cyanobacteria in the competition with eukaryotic primary producers, such as diatoms, chlorophytes, cryptophytes, and dinoflagellates (Paerl and Paul 2012). Research by He et al. (2018) has also indicated that global warming could exacerbate the adverse effects of eutrophication by encouraging the occurrence of harmful cyanobacterial blooms, thereby impacting ecosystem functioning and services. Consequently, the prevalence of cyanobacterial blooms is expected to become more frequent in the future as global climate change continues (Jeppesen et al. 2012), and the anticipated changes in periphyton communities can trigger alterations in the whole food web through cascading interactions (Shurin et al. 2012; Kazanjian et al. 2018).
Conclusion
In the frame of this study, we developed an image processing macro to streamline the analysis of confocal images of algal biofilms. This macro is freely available in the supplements for download and image analysis. The method we devised enables straightforward examination of the structure and composition of intact algal biofilms, including the differentiation of prokaryotic and eukaryotic photosynthetic organisms based on their spectral properties. By utilizing series of images, our method is also suitable for tracking spatio-temporal changes in phytobenthos, thereby facilitating the monitoring of its responses to environmental changes such as temperature-induced variations. We concluded that a 5°C increase in mean water temperature would lead to significant alterations in the community structure of periphytic organisms in shallow lakes. In essence, if global temperatures were to change in accordance with any of the worst-case scenarios, drastic structural and functional transformations would inevitably occur in shallow lake ecosystems.
Data availability
All data supporting the findings of this study are available from the corresponding author, upon reasonable request.
References
Achinas S, Yska SK, Charalampogiannis N, Krooneman J, Euverink GJW (2020) A technological understanding of biofilm detection techniques: a review. Materials 13:3147
Azeredo J, Azevedo NF, Briandet R, Cerca N, Coenye T, Costa AR, Desvaux M, Di Bonaventura G, Hébraud M, Jaglic Z, Kačániová M, Knøchel S, Lourenço A, Mergulhão F, Meyer RL, Nychas G, Simões M, Tresse O, Sternberg C (2017) Critical review on biofilm methods. Crit Rev Microbiol 43:313–351
Bayguinov PO, Oakley DM, Shih CC, Geanon DJ, Joens MS, Fitzpatrick JA (2018) Modern laser scanning confocal microscopy. Curr Protoc Cytomet 85:e39
B-Béres V, Stenger-Kovács C, Buczkó K, Padisák J, Selmeczy GB, Lengyel E, Tapolczai K (2023) Ecosystem services provided by freshwater and marine diatoms. Hydrobiologia 850:2707–2733
Capon SJ, Stewart-Koster B, Bunn SE (2021) Future of freshwater ecosystems in a 1.5°C warmer world. Front Env Sci 9:784642
Carreira C, Staal M, Middelboe M, Brussaard CP (2015) Autofluorescence imaging system to discriminate and quantify the distribution of benthic cyanobacteria and diatoms. Limnol Oceanogr: Methods 13:169–177
Core Team R (2021) R: a language and environment for statistical computing [computer software]. R Foundation for Statistical Computing, Vienna
Crespo D, Pardal MÂ (2020) Ecological and economic importance of benthic communities. In: Leal Filho W, Azul AM, Brandli L, Lange Salvia A, Wall T (eds) Life below water. Springer, Cham, pp 1–11
Dailey ME, Manders E, Soll DR, Terasaki M (2006) Confocal microscopy of living cells. In: Pawley JB (ed) Handbook Of Biological Confocal Microscopy. Springer, Boston, pp 381–403
Daufresne M, Lengfellner K, Sommer U (2009) Global warming benefits the small in aquatic ecosystems. Proc Nat Acad Sci 106:12788–12793
DeNicola DM (1996) Periphyton responses to temperature at different ecological levels. In: Stevenson RJ, Bothwell ML, Lowe RL (eds) Algal Ecology. Academic Press, San Diego, pp 149–181
Elliott AD (2020) Confocal microscopy: principles and modern practices. Curr Protoc Cytom 92:e68
Fanesi A, Paule A, Bernard O, Briandet R, Lopes F (2019) The architecture of monospecific microalgae biofilms. Microorganisms 7:352
Fanesi A, Martin T, Breton C, Bernard O, Briandet R, Lopes F (2022) The architecture and metabolic traits of monospecific photosynthetic biofilms studied in a custom flow-through system. Biotechnol Bioeng 119:2459–2470
Fox J, Weisberg S (2018) An R companion to applied regression. Sage Publications, Thousand Oaks
Greenwood J, Clason T, Lowe R, Belanger S (1999) Examination of endopelic and epilithic algal community structure employing scanning electron microscopy. Freshw Biol 41:821–828
Guo K, Wu N, Li W, Baattrup-Pedersen A, Riis T (2021) Microbial biofilm community dynamics in five lowland streams. Sci Total Environ 798:149169
Hao B, Wu H, Zhen W, Jo H, Cai Y, Jeppesen E, Li W (2020) Warming effects on periphyton community and abundance in different seasons are influenced by nutrient state and plant type: a shallow lake mesocosm study. Front Plant Sci 11:523547
Hartmann R, Jeckel H, Jelli E, Singh PK, Vaidya S, Bayer M, Rode DK, Vidakovic L, Díaz-Pascual F, Fong JC (2021) Quantitative image analysis of microbial communities with BiofilmQ. Nature Microbiol 6:151–156
He H, Jin H, Jeppesen E, Li K, Liu Z, Zhang Y (2018) Fish-mediated plankton responses to increased temperature in subtropical aquatic mesocosm ecosystems: implications for lake management. Water Res 144:304–311
Heydorn A, Nielsen AT, Hentzer M, Sternberg C, Givskov M, Ersbøll BK, Molin S (2000) Quantification of biofilm structures by the novel computer program COMSTAT. Microbiology 146:2395–2407
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biomet J 50:346–363
Jeppesen E, Kronvang B, Jørgensen TB, Larsen SE, Andersen HE, Søndergaard M, Liboriussen L, Bjerring R, Johansson LS, Trolle D, Lauridsen TL (2012) Recent climate‐induced changes in freshwaters in Denmark. In: Goldman CR, Kumagai M, Robarts RD (eds) Climatic change and global warming of inland waters: impacts and mitigation for ecosystems and societies. Wiley-Blackwell, Chichester, pp 155–171
Kazanjian G, Velthuis M, Aben R, Stephan S, Peeters ET, Frenken T, Touwen J, Xue F, Kosten S, Van de Waal DB (2018) Impacts of warming on top-down and bottom-up controls of periphyton production. Sci Rep 8:9901
Krsmanovic M, Biswas D, Ali H, Kumar A, Ghosh R, Dickerson AK (2021) Hydrodynamics and surface properties influence biofilm proliferation. Adv Colloid Interface Sci 288:102336
Kuehn M, Hausner M, Bungartz H-J, Wagner M, Wilderer PA, Wuertz S (1998) Automated confocal laser scanning microscopy and semiautomated image processing for analysis of biofilms. Appl Environ Microbiol 64:4115–4127
Lamprecht O, Wagner B, Derlon N, Tlili A (2022) Synthetic periphyton as a model system to understand species dynamics in complex microbial freshwater communities. NPJ Biofilms Microbiomes 8:61
Larson C, Passy SI (2005) Spectral fingerprinting of algal communities: a novel approach to biofilm analysis and biomonitoring. J Phycol 41:439–446
Lengyel E, Stenger-Kovács C, Boros G, Al-Imari TJK, Novák Z, Bernát G (2023) Anticipated impacts of climate change on the structure and function of phytobenthos in freshwater lakes. Environ Res 238:117283
Liu Y, Aznarez C, Jeppesen E, He H, Li W, Levi EE, Pacheco JP, Cao Y (2021) Responses of submerged macrophytes and periphyton to warming under two nitrogen scenarios: a microcosm study. Hydrobiologia 848:1333–1346
Mahdy A, Hilt S, Filiz N, Beklioğlu M, Hejzlar J, Özkundakci D, Papastergiadou E, Scharfenberger U, Šorf M, Stefanidis K (2015) Effects of water temperature on summer periphyton biomass in shallow lakes: a pan-European mesocosm experiment. Aquat Sci 77:499–510
Manirakiza B, Zhang S, Addo FG, Isabwe A, Nsabimana A (2022) Exploring microbial diversity and ecological function of epiphytic and surface sediment biofilm communities in a shallow tropical lake. Sci Total Environ 808:151821
Meerhoff M, Teixeira-de Mello F, Kruk C, Alonso C, González-Bergonzoni I, Pacheco JP, Lacerot G, Arim M, Beklioğlu M, Brucet S, Goyenola G, Iglesias C, Mazzeo N, Kosten S, Jeppesen E (2012) Environmental warming in shallow lakes: a review of potential changes in community structure as evidenced from space-for-time substitution approaches. Adv Ecol Res 46:259–349
Mhade S, Kaushik KS (2023) Tools of the trade: image analysis programs for confocal laser-scanning microscopy studies of biofilms and considerations for their use by experimental researchers. ACS Omega 8:20163–20177
Mueller NL, de Brouwer JFC, Almeida JS, Stal LJ, Xavier JB (2006) Analysis of a marine phototrophic biofilm by confocal laser scanning microscopy using the new image quantification software PHLIP. BMC Ecol 6:1
Nalley JO, O’Donnell DR, Litchman E (2018) Temperature effects on growth rates and fatty acid content in freshwater algae and cyanobacteria. Algal Res 35:500–507
Neu TR, Lawrence JR (2015) Innovative techniques, sensors, and approaches for imaging biofilms at different scales. Trends Microbiol 23:233–242
Neu TR, Manz B, Volke F, Dynes JJ, Hitchcock AP, Lawrence JR (2010) Advanced imaging techniques for assessment of structure, composition and function in biofilm systems. FEMS Microbiol Ecol 72:1–21
Osorio JHM, Pollio A, Frunzo L, Lens PNL, Esposito G (2021) A review of microalgal biofilm technologies: definition, applications, settings and analysis. Front Chem Eng 3:737710
Pacheco JP, Calvo C, Aznarez C, Barrios M, Meerhoff M, Jeppesen E, Baattrup-Pedersen A (2022) Periphyton biomass and life-form responses to a gradient of discharge in contrasting light and nutrients scenarios in experimental lowland streams. Sci Total Environ 806:150505
Paddock SW, Eliceiri KW (2014) Laser Scanning Confocal Microscopy: History, applications, and related optical sectioning techniques. In: Paddock SW (ed) Confocal Microscopy: Methods and Protocols. Springer, New York, pp 9–47
Paerl HW, Paul VJ (2012) Climate change: links to global expansion of harmful cyanobacteria. Water Res 46:1349–1363
Papageorgiou GC (2004) Fluorescence of photosynthetic pigments in vitro and in vivo. In: Papageorgiou G, Govondjee (eds) Chlorophyll a fluorescence: a signature of photosynthesis. Springer, Dordrecht, pp 43–63
Pawley JB (2006) Fundamental limits in confocal microscopy. In: Pawley JB (ed) Handbook Of Biological Confocal Microscopy. Springer, Boston, pp 20–42
Pearce-Higgins JW, Ockendon N, Baker DJ, Carr J, White EC, Almond RE, Amano T, Bertram E, Bradbury RB, Bradley C (2015) Geographical variation in species’ population responses to changes in temperature and precipitation. Proc Roy Soc B 282:20151561
Pinheiro J, Bates D (2006) Mixed-effects models in S and S-PLUS. Springer, NY
Pörtner HO, Roberts DC, Adams H, Adler C, Aldunce P, Ali E, Begum RA, Betts R, Kerr RB, Biesbroek R (2022) Climate change 2022: Impacts, adaptation and vulnerability. IPCC, Durban
Ras M, Steyer J-P, Bernard O (2013) Temperature effect on microalgae: a crucial factor for outdoor production. Rev Env Sci Bio/Technol 12:153–164
Sauer K, Stoodley P, Goeres DM, Hall-Stoodley L, Burmølle M, Stewart PS, Bjarnsholt T (2022) The biofilm life cycle: expanding the conceptual model of biofilm formation. Nature Rev Microbiol 20:608–620
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B (2012) Fiji: an open-source platform for biological-image analysis. Nature Meth 9:676–682
Schulte PM (2015) The effects of temperature on aerobic metabolism: towards a mechanistic understanding of the responses of ectotherms to a changing environment. J Exp Biol 218:1856–1866
Sehar S, Naz I (2016) Role of the biofilms in wastewater treatment. In: Dhanasekaran D, Thajuddin N (eds) Microbial biofilms-importance and applications. InTech, Riejeka, pp 121–144
Serôdio J, Lavaud J (2020) Diatoms and their ecological importance. In: Leal Filho W, Azul AM, Brandli L, Lange Salvia A, Wall T (eds) Life Below Water. Springer, Cham, pp 1–9
Shurin JB, Clasen JL, Greig HS, Kratina P, Thompson PL (2012) Warming shifts top-down and bottom-up control of pond food web structure and function. Philos Trans Roy Soc B 367:3008–3017
Singh SP, Singh P (2015) Effect of temperature and light on the growth of algae species: a review. Renew Sust Energy Rev 50:431–444
Stevenson RJ (1996) An introduction to algal ecology in freshwater benthic habitats. In: Stevenson RJ, Bothwell ML, Lowe RL (eds) Algal Ecology. Academic Press, San Diego, pp 3–30
Strašková A, Steinbach G, Konert G, Kotabová E, Komenda J, Tichý M, Kaňa R (2019) Pigment-protein complexes are organized into stable microdomains in cyanobacterial thylakoids. Biochim Biophys Acta - Bioenerg 1860:148053
Woodward G, Benstead JP, Beveridge OS, Blanchard J, Brey T, Brown LE, Cross WF, Friberg N, Ings TC, Jacob U (2010a) Ecological networks in a changing climate. Adv Ecol Res 42:71–138
Woodward G, Perkins DM, Brown LE (2010b) Climate change and freshwater ecosystems: impacts across multiple levels of organization. Philos Trans Roy Soc B 365:2093–2106
Wu Y (2017a) Periphyton and its study methods. In: Wu Y (ed) Periphyton. Elsevier, Boston, pp 1–33
Wu Y (2017b) The living environment of periphyton. In: Wu Y (ed) Periphyton. Elsevier, Boston, pp 35–70
Zancarini A, Echenique-Subiabre I, Debroas D, Taïb N, Quiblier C, Humbert J-F (2017) Deciphering biodiversity and interactions between bacteria and microeukaryotes within epilithic biofilms from the Loue River. France. Sci Rep 7:4344
Zhao Y, Zhang Y, Guo J, Wang J, Li Y (2023) Shifts in periphyton research themes over the past three decades. Environ Sci Pollut Res 30:5281–5295
Acknowledgements
We are grateful to Máté Burányi and Réka Horváth for their technical assistance in sample collection and processing.
Funding
Open access funding provided by HUN-REN Balaton Limnological Research Institute. The study was supported by the National Research, Development and Innovation Office (RRF-2.3.1-21-2022-00014), National Brain Project (NAP2022-I-10/2022) and Hungarian Scientific Research Fund (138039).
Author information
Authors and Affiliations
Contributions
Z. N.: Conceptualization, Methodology, Investigation, Visualization, Writing - Original Draft G. Be.: Conceptualization, Methodology, Writing - Review & Editing, Funding acquisition G. S.: Methodology, Software, Investigation, Writing - Review & Editing N. B.: Formal analysis, Visualization, Writing - Review & Editing Zs. P.: Methodology, Resources, Writing - Review & Editing, Funding acquisition G. Bo.: Conceptualization, Investigation, Resources, Writing - Review & Editing, Supervision
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Novák, Z., Bernát, G., Steinbach, G. et al. A semi-automated image processing method for analysing the taxonomic composition of algal biofilms. J Appl Phycol (2024). https://doi.org/10.1007/s10811-024-03339-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10811-024-03339-2