Abstract
Purpose
This study is to develop a new protocol that combines the use of epigenetic cues and mechanical stimuli to assemble 3D spherical structures, arbitrarily defined “epiBlastoids,” whose phenotype is remarkably similar to natural embryos.
Methods
A 3-step approach is used to generate epiBlastoids. In the first step, adult dermal fibroblasts are converted into trophoblast (TR)-like cells, combining the use of 5-azacytidine, to erase the original phenotype, with an ad hoc induction protocol, to drive cells towards TR lineage. In the second step, epigenetic erasing is applied once again, in combination with mechanosensing-related cues, to generate inner cell mass (ICM)-like organoids. Specifically, erased cells are encapsulated into micro-bioreactors to promote 3D cell rearrangement and boost pluripotency. In the third step, TR-like cells are co-cultured with ICM-like spheroids in the same micro-bioreactors. Subsequently, the newly generated embryoids are transferred to microwells to favor epiBlastoid formation.
Results
Adult dermal fibroblasts are successfully readdressed towards TR lineage. Cells subjected to epigenetic erasing and encapsulated into micro-bioreactors rearrange in 3D ICM-like structures. Co-culture of TR-like cells and ICM-like spheroids into micro-bioreactors and microwells induces the formation of single structures with uniform shape reminiscent in vivo embryos. CDX2+ cells localized in the out layer of the spheroids, while OCT4+ cells in the inner of the structures. TROP2+ cells display YAP nuclear accumulation and actively transcribed for mature TR markers, while TROP2− cells showed YAP cytoplasmic compartmentalization and expressed pluripotency-related genes.
Conclusion
We describe the generation of epiBlastoids that may find useful application in the assisted reproduction field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Early mammalian embryogenesis encompasses several events that lead to the formation of the blastocyst that comprises two distinct structures: the trophectoderm (TE) and inner cell mass (ICM) [1]. The TE consists of trophoblast (TR) cells that contribute to the formation of the placenta, nourishing and supporting the fetus during intrauterine life [2,3,4]. The ICM contains pluripotent cells that are able to differentiate in the three definitive germ layers, giving rise to all tissues of the embryo proper [5].
The process of development during which embryonic cells specialize and differentiate are driven by molecular mechanisms and mechanical cues that have long aroused great interest. However, due to the in vivo microenvironment inaccessibility, the material paucity, and the ethical and legal issues, the study of the peri-implantation period in human remains a daunting task [6].
During the last few years, several efforts have been directed towards the creation of in vitro models that recapitulate embryogenesis in vivo. In this context, the successful and scalable generation of blastocyst-like structures (termed blastoids), using different methods and cell sources, have been reported in both mouse and human [7,8,9,10,11,12,13].
Here, we describe a new 3-step approach that allows the generation of 3D multicellular spherical structures, remarkably similar to natural embryos (Fig. 1). To this purpose, starting from easily accessible terminally differentiated cells, we combine the use of epigenetic cues and mechanical stimuli. In particular, in the first step, we convert adult dermal fibroblasts into TR-like cells, using 5-azacytidine (5-aza-CR) to erase the original cell phenotype, and apply an ad hoc induction protocol to drive cells into the TR lineage [14, 15]. In the second step, we combine epigenetic and mechanosensing-related stimuli to generate ICM-like spheroids. Specifically, we apply the same epigenetic erasing protocol and encapsulate cells into polytetrafluoroethylene (PTFE) micro-bioreactors to promote 3D cell rearrangement and boost pluripotency [16, 17]. In the third and last step, we co-culture chemically induced TR-like cells with ICM-like spheroids in the same micro-bioreactor and, subsequently, into microwells, to further encourage differentiation and favor the formation of 3D spherical structures resemble in vivo blastocysts and that we arbitrarily defined “epiBlastoids.”
The results obtained demonstrate that the approach here described allows for the generation of a blastocyst-like model that is able to mimic the physiological organization of early embryos in vivo and may find useful application in the assisted reproduction field.
Materials and methods
All reagents were purchased from Thermo Fisher Scientific unless otherwise indicated.
Ethical statement
Human cell isolation from healthy adult individuals was approved by the Ethical Committee of the Ospedale Maggiore Policlinico, Milano (CE 479_20071.6). All experiments were performed in accordance with the approved guidelines.
Isolation and culture of human dermal fibroblasts
Human fibroblasts were isolated from fresh skin biopsies of 6 healthy individuals (3 women and 3 men). Tissue fragments of approximately 2 mm3 were transferred onto petri dishes (Sarstedt) previously coated with 0.1% porcine gelatin (Sigma-Aldrich) and cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 20% Fetal Bovine Serum (FBS), 2 mM glutamine (Sigma-Aldrich), and antibiotics (Sigma-Aldrich). After 6 days of culture, fibroblasts grown out of the original skin explants and fragments were carefully removed. Cells were subsequently cultured in fibroblast culture medium (FCM) composed of DMEM, 10% FBS, 2 mM glutamine (Sigma-Aldrich), and antibiotics (Sigma-Aldrich) and maintained in 5% CO2 at 37 °C. Passages were carried out twice a week at 1:3 ratio. From each individual (n = 6) were obtained one primary cell line that was used at least in triplicate in 3 independent experiments.
Generation of TR-like cells through 5-aza-CR exposure and trophoblast induction
Cells at passages between 6 and 8 were plated into 0.1% gelatin (Sigma-Aldrich) pre-coated 4-well multidishes (Nunc) at concentration of 7.8 × 104 cells/cm2. Based on our previous studies [15,16,17,18,19,20,21,22,23,24], 24 h after seeding, fibroblasts were treated with 1 µM 5-aza-CR (Sigma-Aldrich) for 18 h. At the end of 5-aza-CR exposure, cells were incubated in Embryonic Stem Cell (ESC) medium consisting of DMEM-low glucose: HAM’S F10 (1:1), 5% FBS, 10% K.O. serum, 2 mM glutamine (Sigma-Aldrich), 0.1 mM β-mercaptoethanol (Sigma-Aldrich), nucleoside mix, 1% non-essential amino acids, 1000 IU/ml ES-growth factor (LIF, Chemicon), and 5 ng/ml b-FGF (R&D System) [19, 25, 26] in 5% CO2 at 37 °C for 3 h. TR differentiation was then induced using mouse embryonic fibroblast (MEF) conditioned medium—obtained by culturing 2 × 104 (cells/cm2) inactivated MEF in ESC medium without b-FGF for 24 h—supplemented with 10 ng/ml Bone Morphogenetic protein 4 (BMP4, Sigma-Aldrich), 1 µM activin/nodal signaling inhibitor (A83-01, Sigma-Aldrich), and 0.1 µM basic fibroblast growth factor (FGF2) signaling inhibitor (PD173074, Sigma-Aldrich) [17, 27, 28]. Cells were maintained in low O2 condition (5% O2, 5% CO2, and 90% N2 atmosphere) at 37 °C for 11 days. Culture medium was refreshed every other day.
Creation of ICM-like spheroids through 5-aza-CR exposure and cell encapsulation in PTFE micro-bioreactors
Fibroblasts at passages between 6 and 8 were exposed to 1 μM 5-aza-CR (Sigma-Aldrich) and encapsulated in PTFE (Sigma-Aldrich) micro-bioreactors for 18 h [16, 17]. More in detail, a PTFE powder bed with particle size of 1 μm (Sigma-Aldrich 430,935) was created inside a petri dish (Sarstedt), and 1 × 104 cells/30 μl of 1 μM 5-aza-CR in FCM was dispensed on it. The dish was gently rotated in a circular motion, and the powder particles completely covered the surface of the liquid drop, generating the micro-bioreactors. These latters were then transferred in a new petri dish and incubated in 5% CO2 at 37 °C, using a humidified chamber to avoid dehydration.
Production of epiBlastoids by assembling TR-like cells and ICM-like spheroids
TR-like cells differentiated for 11 days were detached from culture supports, centrifuged at 150 g for 5 min, and resuspended in G1-PLUS medium (Vitrolife) to obtain 3 × 104 cells/30 μL. In parallel, ICM-like spheroids were recovered by puncturing with a needle the PTFE micro-bioreactors and transferred in a drop of G1-PLUS medium (Vitrolife). A 40 μL drop containing 3 × 104 TR-like cells (30 μL) and one single ICM-like spheroid (10 μL) was dispensed onto new PTFE powder bed. The newly obtained PTFE micro-bioreactors were maintained in culture for 2 days at 37 °C in 5% CO2. Subsequently, the generated epiBlastoids were collected, transferred into non-adherent microwells (AggreWell™, Stemcell technologies) and grown in G2-PLUS medium (Vitrolife). After 24 h of culture in non-adherent microwells, 105 PODS Activin A (Cell guidance systems) were resuspended into 150 μl G2-PLUS medium (Vitrolife). EpiBlastoids were maintained in 5% CO2 at 37 °C and kept in culture for additional 4 days.
Morphometric evaluation
EpiBlastoids were observed under an Eclipse TE200 microscope (Nikon), equipped with a digital camera (Nikon). Pictures were acquired with NIS-Elements Software (Version 4.6; Nikon). Spheroid diameters were measured using ImageJ software (ImageJ software version 1.53j).
EpiBlastoid cell separation
EpiBlastoids were dissociated to single cell suspension by a double enzymatic digestion with collagenase IV (300 U/ml, Sigma) for 30 min and trypsin–EDTA solution (Sigma) for 20 min, followed by mechanical dissociation by pipetting. Cell suspension was filtered with a 30 µm nylon mesh (Pre-Separation Filters, 30 μm, # 130–041-407, Miltenyi Biotec) and centrifuged at 300 g for 5 min. Supernatants were removed, and trophoblast cell surface antigen 2 (TROP2)+ cells were isolated using Magnetic-Activated Cell Sorting (MACS, Miltenyi Biotec) protocol, following the manufacture’s instruction. TROP2− cells were isolated by applying the same protocol and collecting the flow-through. The two cell populations obtained were subjected to gene expression and immunostaining analysis.
Gene expression analysis
TaqManGene Expression Cells-to-CT kit was used to extract RNA, following the manufacturer’s instruction, and DNase I was added in lysis solution at 1:100 concentration. Quantitative real-time PCR was performed with CFX96 Real-Time PCR detection system (Bio-Rad Laboratories) using the predesigned gene-specific primers and probe sets from TaqManGene Expression Assays listed in Table 1. GAPDH and ACTB were used as internal reference genes. Target gene quantification was carried out with CFX Manager software (Bio-Rad Laboratories). cDNA obtained from JAR cell line was used as positive controls for TR-like and TROP2+ cells. cDNA obtained from the human ESC line HES7 was used as positive controls for ICM-like spheroids and TROP2− cells.
Western blot analysis
Constitutive proteins were extracted from cell lysates using the ReadyPrep Protein Extraction Kit (Bio-Rad). Cell nuclear extracts were isolated using the NXtract CelLytic NuCLEAR Extraction Kit (Sigma-Aldrich). Protein concentration was measured by Coomassie Blue-G Dye-binding method. Also, 100 μg of proteins were resuspended in sample buffer (1:1) consisting of 4% SDS (Sigma-Aldrich), 10% 2-mercaptoethanol (Sigma-Aldrich), 20% glycerol (Sigma-Aldrich), 0.004% bromophenol blue (Sigma-Aldrich), and 0.125 M Tris–HCl (Sigma-Aldrich) at pH 6.8. Equal amounts of proteins were loaded, electrophoresed on SDS–polyacrylamide gels, transferred onto 0.45 μm pore size nitrocellulose membranes (Hybond-C Extra, GE Healthcare Life Sciences), and probed with YAP primary antibody (1:1000, Cell signaling, 14,074). Protein bands were visualized by the WesternBreeze chemiluminescent kit, and densitometric analysis was performed using the ImageJ software (ImageJ software version 1.53j). GAPDH (1:1000, Abcam, ab8245) was used as loading control for protein normalization.
Immunostaining
TR-like cells, ICM-like spheroids, and epiBlastoids were fixed in 4% paraformaldehyde for 20 min, washed three times in PBS, permeabilized with 0.5% Triton X-100 (Sigma) for 30 min, and treated with a blocking solution containing 10% goat serum (Sigma) for 30 min. When cells formed spherical structures, these were dissociated and attached to slides, using a cytocentrifuge (Cytospin 4, Thermo Shandon). Primary antibodies for CDX2 (1:50, Santa Cruz Biotechnology, sc-166830), KRT19 (1:500, Abcam, ab76539), and OCT4 (1:50, Chemicon, ab3209) were incubated overnight at + 4 °C. The day after, samples were washed three times in PBS and incubated with the appropriate secondary antibodies (Alexa Fluor) for 45 min at room temperature using a 1:250 dilution. Nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). At the end of the immunostaining procedure, cells were analyzed under an Eclipse E600 microscope (Nikon) equipped with a digital camera (Nikon), and epiBlastoids were transferred and mounted to glass slides and visualized under an Eclipse E600 microscope (Nikon) equipped with a digital camera (Nikon). Pictures were acquired with NIS-Elements Software (Version 4.6; Nikon).
For YAP localization, TROP2+ and TROP2− cells (see above) were attached to slides using the cytocentrifuge Cytospin 4 (Thermo Shandon). Cells were then fixed in 4% paraformaldehyde (Sigma), washed three times in PBS, permeabilized, and blocked with a solution containing 0.3% Triton X-100 (Sigma) and 5% serum for 1 h. YAP primary antibody (1:100, Cell signaling, 14,074) was incubated overnight at + 4 °C. Cells were then incubated with suitable secondary antibody (Alexa Fluor) for 45 min. Nuclei were stained with DAPI (Sigma). Samples were observed under a Nikon Eclipse TE200 (Nikon) equipped with a digital camera (Nikon). Pictures were acquired with NIS-Elements Software (Version 4.6; Nikon).
Cell counting
The number of CDX2, KRT19, and OCT4 immuno-positive cells was counted in 10 randomly selected fields at 200 × total magnification. A minimum of 350 cells were scored in at least three independent replicates. The number of positively stained cells was expressed as a percentage of the total cell counted.
Statistical analysis
Statistical analysis was performed using the Student t-test (SPSS 19.1; IBM). Data were presented as mean ± standard deviation (SD). Differences of p ≤ 0.05 were considered significant and were indicated with different superscripts.
Results
Generation of TR-like cells through 5-aza-CR exposure and trophoblast induction
Fibroblasts obtained from skin biopsies grew out of the original explants within 6 days of culture forming a monolayer with cells displaying the standard elongated morphology (Fig. 2A, Untreated fibroblasts). After 5-aza-CR exposure, cell phenotype changed, and treated fibroblasts acquired an oval or round shape, becoming smaller with larger nuclei, granular and vacuolated cytoplasm (Fig. 2A, Post 5-aza-CR). These changes were accompanied by the onset of the pluripotency-related genes OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, and DPPA2 which were undetectable in untreated fibroblasts (Fig. 2C). In agreement with these observations, epigenetically erased cells significantly downregulated the typical fibroblast markers VIM and THY1 (Fig. 2C). At the end of 11-day TR induction, cells acquired a mature TR morphology, exhibiting round or ellipsoid shape, round nuclei, and well-defined borders (Fig. 2A, trophoblast-like cells). Consistent with these morphological changes, TR-like cells actively transcribed for TR related genes, namely, GCM1, CGA, CGB, HSD17B1, CYP11A1, PGF, ESRRB, GATA2, GATA3, and KRT19 which were originally absent in untreated fibroblasts (Fig. 2C). This was further support by immunocytochemical results, showing cell positivity for the TR markers CDX2 and KRT19 (79.5 ± 2.99% and 78.65 ± 6.01%, respectively; Fig. 2B).
Generation of TR-like cells through 5-aza-CR exposure and trophoblast induction. Fibroblasts exposed to 5-aza-CR lost their typical elongated shape (untreated fibroblasts) and became smaller with larger nuclei and granular cytoplasm (post 5-aza-CR) (scale bars 100 µm; A). At day 11 of trophoblast induction, cells acquired a tight adherent epithelial morphology, exhibiting round or ellipsoid shape, with round nuclei and well-defined borders (trophoblast-like cells; scale bar 100 µm; A). Immunostainings show cell positivity for the TR markers CDX2 (red, left panel) and KRT19 (green, right panel). Nuclei are stained in blue (scale bars 100 μm; B). Transcription levels for pluripotent- (OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, DPPA2), fibroblast- (VIM, THY1), and TR- related genes (GCM1, CGA, CGB, HSD17B1, CYP11A1, PGF, ESRRB, GATA2, GATA3, KRT19) in untreated fibroblasts (white bars), fibroblasts exposed to 5-aza-CR (Post 5-aza-CR, black bars), at day 11 of trophoblast induction (trophoblast-like cells, grey bars) and in JAR cell line (JAR, blue bars). Gene expression values are reported with the highest expression set to 1 and all others relative to this. Different superscripts denote significant differences (P < 0.05; C)
Creation of ICM-like spheroids through 5-aza-CR exposure and cell encapsulation in PTFE micro-bioreactors
After 5-aza-CR exposure and encapsulation in PTFE, fibroblasts became rounded, with large and granulated nuclei, lost their monolayer distribution, and rearranged in 3D spherical structures (Fig. 3A, post 5-aza-CR + PTFE). Consistently with this, the obtained ICM-like spheroids actively transcribed for the main pluripotency-related genes OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, and DPPA2 originally absent in untreated cells (untreated fibroblasts), while significantly decreased the expression of the fibroblast-specific markers VIM and THY1 (Fig. 3C). In addition, immunocytochemical studies demonstrate the presence of 86.31 ± 4.13% OCT4+ cells (Fig. 3B).
Creation of ICM-like spheroids through 5-aza-CR exposure and cell encapsulation in PTFE micro-bioreactors. Fibroblasts (untreated fibroblasts; scale bar 100 μm) treated with 5-aza-CR and encapsulated in PTFE micro-bioreactors form 3D spherical structures (red arrows, Post 5-aza-CR + PTFE; scale bar 50 μm; A). Immunostainings show cell positivity for the pluripotency-related marker OCT4. Nuclei are stained in blue (scale bars 50 μm; B). Transcription levels for pluripotent- (OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, DPPA2) and fibroblast-related genes (VIM, THY1) in untreated fibroblasts (white bars), fibroblasts exposed to 5-aza-CR and encapsulated in PTFE micro-bioreactors (post 5-aza-CR + PTFE, black bars), and human ESC line (ESC, green bars). Gene expression values are reported with the highest expression set to 1 and all others relative to this. Different superscripts denote significant differences (P < 0.05) (C)
Production of epiBlastoids by assembling TR-like cells and ICM-like spheroids
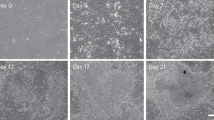
After 2 days of co-culture in PTFE micro-bioreactors, chemically induced TR-like cells and ICM-like spheroids self-assembled in single 3D spherical structures (Fig. 4A). At the end of 7-day culture period, epiBlastoids displayed uniform round shape with sizes ranging from 100 to 200 µm, regardless of the sex of the fibroblasts. More in detail, morphometric analysis indicated that 78.67% of spheroids exhibited a diameter of 150–200 μm, while 21.33% of 100–150 μm (Fig. 4B).
Production of epiBlastoids by assembling TR-like cells and ICM-like spheroids. Representative image of an epiBlastoid (scale bar 50 μm; A). Rates of epiBlastoids displaying diameters ranging from 100 to 150 μm and from 151 to 200 μm *Superscripts denote significant differences (P < 0.05; B). Immunostaining showing CDX2+ cells (red) localize in out layer of the epiBlastoids and OCT4+ cells (green) in the inner compartment. Nuclei are stained in blue (scale bars 50 μm; C). YAP protein is confined in the nuclear compartment of TROP2+ cells, while it is excluded from the nucleus and shifts into the cytoplasm in TROP2− cells. Nuclei are stained in blue (scale bars 100 μm; D). Densitometric analysis of western blots for YAP protein in cytoplasm (C) and nucleus (N) of TROP2+ (black bars) and TROP2− cells (white bars). The values (arbitrary units) are reported as relative optical density of the bands normalized to GAPDH. *Superscripts denote significant differences (P < 0.05; E). Representative YAP and GAPDH western blots for the two cell compartments of each cell type are also shown (F). Transcription levels for pluripotency- (OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, DPPA2) and TR-related genes (GCM1, CGA, CGB, HSD17B1, CYP11A1, PGF, ESRRB, GATA2, GATA3, KRT19) in TROP2+ cells (black bars), TROP2.− cells (white bars), human ESC (ESC, green bars), and JAR cell line (JAR, blue bars). Gene expression values are reported with the highest expression set to 1 and all others relative to this (G)
Immunostaining studies showed CDX2+ cells localized in out layer of the epiBlastoids, while OCT4 was expressed by cells of the inner compartment (Fig. 4C). In addition, immunolocalization and western blot analyses demonstrated that TROP2+ cells were characterized by YAP nuclear accumulation, while TROP2− cells by cytoplasmic compartmentalization of the molecule (Fig. 4D–F).
These morphological observations were also supported by molecular analyses indicating that TROP2+ cells expressed the TR markers GCM1, CGA, CGB, HSD17B1, CYP11A1, PGF, ESRRB, GATA2, GATA3, and KRT19, while TROP2− cells transcribed for the pluripotency-related genes OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, and DPPA2 (Fig. 4G).
Discussion
The possibility to successfully recreate in vitro blastocyst-like structures using embryonic stem cells (ESCs) or induced pluripotent stem cells (iPSCs) has been recently demonstrated both in the mouse [29,30,31,32] and the human [7,8,9,10,11,12]. In the present manuscript, we describe a novel method that combines, for the first time, the use of chemical stimuli and mechanical cues to generate in vitro 3D spherical structures phenotypically similar to natural embryos.
More in detail, in the first step, we induce a high permissivity window in adult dermal fibroblasts isolated from human healthy individuals, using 5-aza-CR which has been previously demonstrated to reactivate pluripotency-related genes [33,34,35,36], to induce a global DNA hypomethylation [16,17,18,19,20,21,22,23,24, 37,38,39,40,41,42,43,44,45,46] and to modulate ten-eleven translocation (TET) gene transcription [23]. In agreement with this, in the present work, adult dermal fibroblasts exposed to 5-aza-CR for 18 h lost their typical elongated morphology and acquired a round or oval shape, with larger nuclei and granular, vacuolated cytoplasm. These morphological changes closely resemble those previously identified in ESCs [47, 48] and iPSCs [49], suggesting the acquisition of features distinctive of a high plasticity phenotype. This was accompanied also by the onset of the main pluripotency-related gene transcription, namely, OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, and DPPA2, which were originally undetectable in untreated fibroblasts (untreated fibroblasts), and by a significant down-regulation of the fibroblast-related markers VIM and THY1, further supporting previously published reports that described 5-aza-CR ability to induce a pluripotent-like phenotype hypomethylation [16,17,18,19,20,21,22,23,24, 37,38,39,40,41,42,43,44,45,46]. Taking advantage of the acquired high permissivity window, cells were readdressed towards the TR lineage, using an induction cocktail containing BMP4 in combination with activin/nodal and FGF2 signaling inhibitors. This differentiation medium has been previously shown to drive cells towards the TR phenotype in both human [27, 50,51,52,53,54,55,56,57,58] and pig [14, 28], with the acquisition of a tight adherent epithelial morphology, round shape, and nuclei, as well as well-defined borders. Consistent with this, immunocytochemical results indicated a high conversion efficiency (~ 80%), which is similar to that scored in human reprogrammed iPSCs differentiated towards TR lineage [27, 56, 59, 60] as well as in TR-like cells obtained from epigenetically converted porcine adult dermal fibroblasts [28]. In addition, the obtained TR-like cells showed active transcription for the TR mature markers GCM1, CGA, CGB, HSD17B1, CYP11A1, PGF, ESRRB, GATA2, GATA3, and KRT19, indicating the activation of the main molecular pathways distinctive of the newly acquired phenotype.
In the second step, we combined epigenetic and mechanosensing-related stimuli to generate ICM-like spheroids. To this purpose, erased adult dermal fibroblasts were encapsulated into PTFE micro-bioreactors to promote 3D cell rearrangement and boost pluripotency. Beside the morphological changes induced by exposure to 5-aza-CR, previously shown to be related to high plasticity, the use of the PTFE micro-bioreactors allowed cells to self-assemble and form multicellular spheroids, displaying a uniform size geometry. This well fits with previous observations indicating that the use of micro-bioreactors efficiently encourages 3D cell aggregation [61, 62], which, in turn, has the ability to support the induction of a pluripotent state and to boost its maintenance [16]. In agreement with this and consistently with our previous study [23], 86.31 ± 4.13% of cells were immune-positive for OCT4 and the generated ICM-like spheroids actively and steadily transcribed for the pluripotency-related genes OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, and DPPA2, and down-regulated VIM and THY1 genes.
In the third and last step of the protocol, we co-cultured TR-like cells with ICM-like spheroids in the PTFE micro-bioreactor for 2 days, to favor the formation of a single 3D spherical structure, composed by the two cell components. We subsequently transferred the newly generated aggregates into microwells and cultured them in a commercially available medium for further 5 days, to encourage epiBlastoid formation. Encapsulation of two different cell types has been previously reported by Jara et al. who applied this approach for the production of pancreatic islet-like structures in vitro [63]. These authors described encapsulation ability to stabilize 3D cell aggregation, maintain differentiation, and support functional activities. In the present manuscript, upon these specific culture conditions, TR-like cells and ICM-like spheroids were able to organize into structures with uniform round shape displaying a diameter ranging from 100 to 200 µm. In particular, morphometric analysis demonstrated that 78.67% of the spheroids exhibited a diameter ranging from 150 to 200 μm, with only 21.33% from 100 to150 μm. These results well fit with the average reported values of natural blastocysts that measure 175–211 μm and with the parameters and criteria reported in Table 2 and currently used to define human blastoid models. In addition, immunostaining studies demonstrated CDX2+ cells externally localized, to surround the epiblastoid, and OCT4+ cells closely assembled within them, showing that our protocol induced cells to spontaneously organize into spheroid complexes displaying TR-like cells homogenously distributed in the outer layer of the structures and ICM-like aggregates confined to the internal compartment. Consistent with this, sorting of epiBlastoid-derived cell suspensions, using the surface TR marker TROP2, led to the obtainment of two distinct cell populations: one consisting of TROP2+ cells and another containing TROP2− cells. The first cell type actively transcribed for GCM1, CGA, CGB, HSD17B1, CYP11A1, PGF, ESRRB, GATA2, GATA3, and KRT19, genes, indicating that TROP2+ cells maintained a transcription pattern typical of TR cells. In addition, TROP2+ cells displayed a distinct compartmentalization of YAP which was mainly accumulated in the nucleus. In agreement with this, several studies reported the direct involvement of the Hippo pathway in the blastocyst formation, highlighting the key role played by the TEAD4/WWTR1/YAP1 complex in promoting CDX2 expression during outside cell maturation to TR [64,65,66,67,68,69,70,71,72,73,74,75,76,77,78]. On the other hand, and in line with this, TROP2− cells showed cytoplasmic retention of YAP and expressed the pluripotency-related genes OCT4, NANOG, REX1, SOX2, KLF17, PRDM14, and DPPA2.
Altogether, the procedure here described allows for an efficient in vitro generation of human epiBlastoids, starting from easily accessible adult dermal fibroblasts and avoiding the use of techniques that require retroviral gene transfection. Considering that the access to human embryo is subjected to tight regulation around the globe due to ethical concerns, the production of epiBlastoid models may overcome some and could find useful application in the assisted reproduction field, for the identification of the most adequate culture conditions and/or for peri- and early post-implantation investigations. In addition, the use of 3D micro-bioreactors may also represent a notable breakthrough in culture system technologies applied to reproduction and may constitute an advantageous micro-environment for long-term culture of blastoids, gastruloids, or organoids.
Data Availability
The data presented in this study are available on request from the corresponding author.
Change history
12 September 2023
A Correction to this paper has been published: https://doi.org/10.1007/s10815-023-02928-3
References
Aitken RJ, Beaconsfield R, Ginsburg J. Origin and formation of the placenta. 1979. https://doi.org/10.1016/B978-0-08-024435-8.50025-0
Schulz L, Ezashi T, Das P, Westfall S, Livingston K, Roberts R. Human embryonic stem cells as models for trophoblast differentiation Introduction: the trophoblast lineage and its emergence. 2009. https://doi.org/10.1016/j.placenta.2007.10.009
Sheridan MA, Fernando RC, Gardner L, Hollinshead MS, Burton GJ, Moffett A, et al. Establishment and differentiation of long-term trophoblast organoid cultures from the human placenta. 2020. https://doi.org/10.1038/s41596-020-0381-x
Rossant J, Cross JC. Placental development: lessons from mouse mutants. 2001. 538–48. https://doi.org/10.1038/35080570
Romito A, Cobellis G. Pluripotent stem cells: current understanding and future directions. 2016. https://doi.org/10.1155/2016/9451492
Daoud AMP, Popovic M, Dondorp WJ, Bustos MT, Bredenoord AL, de Sousa Lopes SMC, et al. Modelling human embryogenesis: embryo-like structures spark ethical and policy debate. 2020. https://doi.org/10.1093/HUMUPD/DMAA027
Fan Y, Min Z, Alsolami S, Ma Z, Zhang E, Chen W, et al. Generation of human blastocyst-like structures from pluripotent stem cells. 2021. https://doi.org/10.1038/s41421-021-00316-8
Liu X, Tan JP, Schröder J, Aberkane A, Ouyang JF, Mohenska M, et al. Modelling human blastocysts by reprogramming fibroblasts into iBlastoids. 2021. https://doi.org/10.1038/s41586-021-03372-y
Yanagida A, Spindlow D, Nichols J, Dattani A, Smith A, Guo G. Naive stem cell blastocyst model captures human embryo lineage segregation. 2021. https://doi.org/10.1016/j.stem.2021.04.031
Sozen B, Jorgensen V, Weatherbee BAT, Chen S, Zhu M, Zernicka-Goetz M. Reconstructing aspects of human embryogenesis with pluripotent stem cells. 2021. https://doi.org/10.1038/s41467-021-25853-4
Yu L, Wei Y, Duan J, Schmitz DA, Sakurai M, Wang L, et al. Blastocyst-like structures generated from human pluripotent stem cells. 2021. https://doi.org/10.1038/s41586-021-03356-y
Kagawa H, Javali A, Khoei HH, Sommer TM, Sestini G, Novatchkova M, et al. Human blastoids model blastocyst development and implantation. 2021. https://doi.org/10.1038/s41586-021-04267-8
Kagawa H, Javali A, Khoei HH, Sommer TM, Sestini G, Novatchkova M, et al. Protocol for human blastoids modeling blastocyst development and implantation. 2022. https://doi.org/10.3791/63388
Arcuri S, Gandolfi F, Somigliana E, Brevini TAL. A two-step protocol to erase human skin fibroblasts and convert them into trophoblast-like cells. 2021. https://doi.org/10.1007/978-1-0716-1246-0_10
Gandolfi F, Arcuri S, Pennarossa G, Brevini TAL. New tools for cell reprogramming and conversion: Possible applications to livestock. 2019. https://doi.org/10.21451/1984-3143-AR2019-0043
Pennarossa G, Manzoni EFM, Ledda S, deEguileor M, Gandolfi F, Brevini TAL. Use of a PTFE micro-bioreactor to promote 3D cell rearrangement and maintain high plasticity in epigenetically erased fibroblasts. 2019. https://doi.org/10.1007/S12015-018-9862-5
Pennarossa G, Ledda S, Arcuri S, Gandolfi F, Brevini TAL. A two-step strategy that combines epigenetic modification and biomechanical cues to generate mammalian pluripotent cells. 2020. https://doi.org/10.3791/61655
Pennarossa G, Maffei S, Campagnol M, Tarantini L, Gandolfi F, Brevini TALALTAL. Brief demethylation step allows the conversion of adult human skin fibroblasts into insulin-secreting cells. 2013. https://doi.org/10.1073/pnas.1220637110
Pennarossa G, Maffei S, Campagnol M, Rahman MM, Brevini TAL, Gandolfi F. Reprogramming of pig dermal fibroblast into insulin secreting cells by a brief exposure to 5-aza-cytidine. 2014. https://doi.org/10.1007/s12015-013-9477-9
Brevini TAL, Pennarossa G, Rahman MM, Paffoni A, Antonini S, Ragni G, et al. Morphological and molecular changes of human granulosa cells exposed to 5-azacytidine and addressed toward muscular differentiation. 2014. https://doi.org/10.1007/s12015-014-9521-4
Brevini TAL, Pennarossa G, Acocella F, Brizzola S, Zenobi A, Gandolfi F. Epigenetic conversion of adult dog skin fibroblasts into insulin-secreting cells. 2016. https://doi.org/10.1016/j.tvjl.2016.02.014
Pennarossa G, Santoro R, Manzoni EFM, Pesce M, Gandolfi F, Brevini TAL. Epigenetic erasing and pancreatic differentiation of dermal fibroblasts into insulin-producing cells are boosted by the use of low-stiffness substrate. 2018. https://doi.org/10.1007/s12015-017-9799-0
Manzoni EFM, Pennarossa G, DeEguileor M, Tettamanti G, Gandolfi F, Brevini TAL. 5-azacytidine affects TET2 and histone transcription and reshapes morphology of human skin fibroblasts. 2016. https://doi.org/10.1038/srep37017
Brevini TAL, Pennarossa G, Maffei S, Zenobi A, Gandolfi F. Epigenetic conversion as a safe and simple method to obtain insulinsecreting cells from adult skin fibroblasts. 2016. https://doi.org/10.1007/s12015-013-9477-9
Brevini TAL, Pennarossa G, Attanasio L, Vanelli A, Gasparrini B, Gandolfi F. Culture conditions and signalling networks promoting the establishment of cell lines from parthenogenetic and biparental pig embryos. 2010. https://doi.org/10.1007/s12015-010-9153-2
Brevini TAL, Pennarossa G, Maffei S, Gandolfi F. Pluripotency network in porcine embryos and derived cell lines. 2012. https://doi.org/10.1111/j.1439-0531.2012.02060.x
Amita M, Adachi K, Alexenko AP, Sinha S, Schust DJ, Schulz LC, et al. Complete and unidirectional conversion of human embryonic stem cells to trophoblast by BMP4. 2013. https://doi.org/10.1073/pnas.1303094110
Arcuri S, Pennarossa G, Gandolfi F, Brevini TAL. Generation of trophoblast-like cells from hypomethylated porcine adult dermal fibroblasts. 2021. https://doi.org/10.3389/fvets.2021.706106
Rivron NC, Frias-Aldeguer J, Vrij EJ, Boisset J-C, Korving J, Vivié J, et al. Blastocyst-like structures generated solely from stem cells. 2018. https://doi.org/10.1038/s41586-018-0051-0
Sozen B, Cox AL, de Jonghe J, Bao M, Hollfelder F, Glover DM, et al. Self-organization of mouse stem cells into an extended potential blastoid. 2019. https://doi.org/10.1016/j.devcel.2019.11.014
Kime C, Kiyonari H, Ohtsuka S, Kohbayashi E, Asahi M, Yamanaka S, et al. Induced 2C expression and implantation-competent blastocyst-like cysts from primed pluripotent stem cells. 2019. https://doi.org/10.1016/J.STEMCR.2019.07.011
Li R, Zhong C, Yu Y, Liu H, Sakurai M, Yu L, et al. Generation of blastocyst-like structures from mouse embryonic and adult cell cultures. 2019. https://doi.org/10.1016/j.cell.2019.09.029
Taylor SM, Jones PA. Multiple new phenotypes induced in 10 T1/2 and 3 T3 cells treated with 5-azacytidine. 1979; 0092-8674(79)90317-9
Jones PA. Effects of 5-azacytidine and its 2’-deoxyderivative on cell differentiation and DNA methylation. 1985. https://doi.org/10.1016/0163-7258(85)90080-4
Jones PA. Altering gene expression with 5-azacytidine. 1985; 0092–8674(85)90192–8
Glover TW, Coyle-Morris J, Pearce-Birge L, Berger C, Gemmill RM. DNA demethylation induced by 5-azacytidine does not affect fragile X expression. 1986; 38(3):309–18.
Palii SS, van Emburgh BO, Sankpal UT, Brown KD, Robertson KD. DNA methylation inhibitor 5-Aza-2’-deoxycytidine induces reversible genome-wide DNA damage that is distinctly influenced by DNA methyltransferases 1 and 3B. 2008. https://doi.org/10.1128/MCB.01799-07
Brevini TAL, Pennarossa G, Maffei S, Gandolfi F. Phenotype switching through epigenetic conversion. 2015. https://doi.org/10.1071/RD14246
Pennarossa G, Zenobi A, Gandolfi CEE, Manzoni EFM, Gandolfi F, Brevini TAL. Erase and rewind: epigenetic conversion of cell fate. 2015. https://doi.org/10.1007/s12015-015-9637-1
Brevini T, Manzoni E, Gandolfi F. Methylation mechanisms and biomechanical effectors controlling cell fate. 2018. https://doi.org/10.1071/RD17348
Brevini TAL, Pennarossa G, Manzoni EFM, Gandolfi F. Safety and efficacy of epigenetically converted human fibroblasts into insulin-secreting cells: a preclinical study. 2018. https://doi.org/10.1007/5584_2018_172
Tan SJ, Fang JY, Wu Y, Yang Z, Liang G, Han B. Muscle tissue engineering and regeneration through epigenetic reprogramming and scaffold manipulation. 2015. https://doi.org/10.1038/srep16333
Mirakhori F, Zeynali B, Kiani S, Baharvand H. Brief azacytidine step allows the conversion of suspension human fibroblasts into neural progenitor-like cells. 2015. https://doi.org/10.22074/cellj.2015.522
Mirakhori F, Zeynali B, Rassouli H, Salekdeh GH, Baharvand H. Direct conversion of human fibroblasts into dopaminergic neural progenitor-like cells using TAT-mediated protein transduction of recombinant factors. 2015. https://doi.org/10.1016/j.bbrc.2015.02.166
Chandrakanthan V, Yeola A, Kwan JC, Oliver RA, Qiao Q, Kang YC, et al. PDGF-AB and 5-Azacytidine induce conversion of somatic cells into tissue-regenerative multipotent stem cells. 2016. https://doi.org/10.1073/pnas.1518244113
Diomede F, Zini N, Pizzicannella J, Merciaro I, Pizzicannella G, D’Orazio M, et al. 5-Aza exposure improves reprogramming process through embryoid body formation in human gingival stem cells. 2018. https://doi.org/10.3389/fgene.2018.00419
Niwa H. How is pluripotency determined and maintained? 2007. https://doi.org/10.1242/dev.02787
Lai D, Wang Y, Sun J, Chen Y, Li T, Wu Y, et al. Derivation and characterization of human embryonic stem cells on human amnion epithelial cells. 2015. https://doi.org/10.1038/srep10014
Courtot AM, Magniez A, Oudrhiri N, Feraud O, Bacci J, Gobbo E, et al. Morphological analysis of human induced pluripotent stem cells during induced differentiation and reverse programming. 2014. https://doi.org/10.1089/biores.2014.0028
Sudheer S, Bhushan R, Fauler B, Lehrach H, Adjaye J. FGF inhibition directs BMP4-mediated differentiation of human embryonic stem cells to syncytiotrophoblast. 2012. https://doi.org/10.1089/scd.2012.0099
Lee YL, Fong SW, Chen ACH, Li T, Yue C, Lee CL, et al. Establishment of a novel human embryonic stem cell-derived trophoblastic spheroid implantation model. 2015. https://doi.org/10.1093/humrep/dev223
Yang Y, Adachi K, Sheridan MA, Alexenko AP, Schust DJ, Schulz LC, et al. Heightened potency of human pluripotent stem cell lines created by transient BMP4 exposure. 2015. https://doi.org/10.1073/pnas.1504778112
Yabe S, Alexenko AP, Amita M, Yang Y, Schust DJ, Sadovsky Y, et al. Comparison of syncytiotrophoblast generated from human embryonic stem cells and from term placentas. 2016. https://doi.org/10.1073/pnas.1601630113
Jain A, Ezashi T, Roberts RM, Tuteja G. Deciphering transcriptional regulation in human embryonic stem cells specified towards a trophoblast fate. 2017. https://doi.org/10.1038/s41598-017-17614-5
Koel M, Võsa U, Krjutškov K, Einarsdottir E, Kere J, Tapanainen J, et al. Optimizing bone morphogenic protein 4-mediated human embryonic stem cell differentiation into trophoblast-like cells using fibroblast growth factor 2 and transforming growth factor-β/activin/nodal signalling inhibition. 2017. https://doi.org/10.1016/j.rbmo.2017.06.003
Roberts RM, Ezashi T, Sheridan MA, Yang Y. Specification of trophoblast from embryonic stem cells exposed to BMP4†. 2018. https://doi.org/10.1093/biolre/ioy070
Sheridan MA, Yang Y, Jain A, Lyons AS, Yang P, Brahmasani SR, et al. Early onset preeclampsia in a model for human placental trophoblast. 2019. https://doi.org/10.1073/pnas.1816150116
Karvas RM, McInturf S, Zhou J, Ezashi T, Schust DJ, Roberts RM, et al. Use of a human embryonic stem cell model to discover GABRP, WFDC2, VTCN1 and ACTC1 as markers of early first trimester human trophoblast. 2020. https://doi.org/10.1093/molehr/gaaa029
Li Y, Moretto-Zita M, Soncin F, Wakeland A, Wolfe L, Leon-Garcia S, et al. BMP4-directed trophoblast differentiation of human embryonic stem cells is mediated through a ΔNp63+ cytotrophoblast stem cell state. 2013. https://doi.org/10.1242/dev.092155
Horii M, Li Y, Wakeland AK, Pizzo DP, Nelson KK, Sabatini K, et al. Human pluripotent stem cells as a model of trophoblast differentiation in both normal development and disease. 2016. https://doi.org/10.1073/pnas.1604747113
Sarvi F, Arbatan T, Chan PPY, Shen WA. A novel technique for the formation of embryoid bodies inside liquid marbles. 2013. https://doi.org/10.1039/C3RA40364E
Vadivelu RK, Kamble H, Munaz A, Nguyen NT. Liquid marbles as bioreactors for the study of three-dimensional cell interactions. 2017. https://doi.org/10.1007/s10544-017-0171-6
Jara C, Oyarzun-Ampuero F, Carrión F, González-Echeverría E, Cappelli C, Caviedes P. Microencapsulation of cellular aggregates composed of differentiated insulin and glucagon-producing cells from human mesenchymal stem cells derived from adipose tissue. 2020. https://doi.org/10.1186/S13098-020-00573-9
Rayon T, Menchero S, Nieto A, Xenopoulos P, Crespo M, Cockburn K, et al. Notch and hippo converge on Cdx2 to specify the trophectoderm lineage in the mouse blastocyst. 2014. https://doi.org/10.1016/j.devcel.2014.06.019
Leung CY, Zernicka-Goetz M. Angiomotin prevents pluripotent lineage differentiation in mouse embryos via Hippo pathway-dependent and-independent mechanisms. 2013. https://doi.org/10.1038/ncomms3251
Yagi R, Kohn MJ, Karavanova I, Kaneko KJ, Vullhorst D, DePamphilis ML, et al. Transcription factor TEAD4 specifies the trophectoderm lineage at the beginning of mammalian development. 2007. https://doi.org/10.1242/dev.010223
Lorthongpanich C, Messerschmidt DM, Chan SW, Hong W, Knowles BB, Solter D. Temporal reduction of LATS kinases in the early preimplantation embryo prevents ICM lineage differentiation. 2013. https://doi.org/10.1101/gad.219618.113
Yu C, Ji SY, Dang YJ, Sha QQ, Yuan YF, Zhou JJ, et al. Oocyte-expressed yes-associated protein is a key activator of the early zygotic genome in mouse. 2016. https://doi.org/10.1038/cr.2016.20
Menchero S, Rollan I, Lopez-Izquierdo A, Andreu MJ, de Aja JS, Kang M, et al. Transitions in cell potency during early mouse development are driven by notch. 2019. https://doi.org/10.7554/eLife.42930
Nishioka N, Yamamoto S, Kiyonari H, Sato H, Sawada A, Ota M, et al. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. 2008. https://doi.org/10.1016/j.mod.2007.11.002
Hirate Y, Hirahara S, Inoue KI, Suzuki A, Alarcon VB, Akimoto K, et al. Polarity-dependent distribution of angiomotin localizes hippo signaling in preimplantation embryos. 2013. https://doi.org/10.1016/j.cub.2013.05.014
Posfai E, Petropoulos S, de Barros FRO, Schell JP, Jurisica I, Sandberg R, et al. Position- and hippo signaling-dependent plasticity during lineage segregation in the early mouse embryo. 2017. https://doi.org/10.7554/eLife.22906
Kono K, Tamashiro DAA, Alarcon VB. Inhibition of RHO-ROCK signaling enhances ICM and suppresses TE characteristics through activation of Hippo signaling in the mouse blastocyst. 2014. https://doi.org/10.1016/j.ydbio.2014.06.023
Shi X, Yin Z, Ling B, Wang L, Liu C, Ruan X, et al. Rho differentially regulates the hippo pathway by modulating the interaction between amot and Nf2 in the blastocyst. 2017. https://doi.org/10.1242/dev.157917
Cockburn K, Biechele S, Garner J, Rossant J. The hippo pathway member nf2 is required for inner cell mass specification. 2013. https://doi.org/10.1016/j.cub.2013.05.044
Anani S, Bhat S, Honma-Yamanaka N, Krawchuk D, Yamanaka Y. Initiation of Hippo signaling is linked to polarity rather than to cell position in the pre-implantation mouse embryo. 2014. https://doi.org/10.1242/dev.107276
Cao Z, Carey TS, Ganguly A, Wilson CA, Paul S, Knott JG. Transcription factor AP-2γ induces early CdX2 expression and represses HIPPO signaling to specify the trophectoderm lineage. 2015. https://doi.org/10.1242/dev.120238
Frum T, Watts JL, Ralston A. TEAD4, YAP1 and WWTR1 prevent the premature onset of pluripotency prior to the 16-cell stage. 2019. https://doi.org/10.1242/DEV.179861
Acknowledgements
The authors are grateful to Dr. Elena Franca Maria Manzoni for the help in experimental setup.
Funding
Open access funding provided by Università degli Studi di Milano within the CRUI-CARE Agreement. This research was funded by Carraresi Foundation, PSR2021 and HORIZON-WIDERA-2021 project n#101079349 (OH-Boost). The authors are members of the Trans-COST Actions Task-Force on Covid-19.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Fig. 4 in the original version of this article has been replaced
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pennarossa, G., Arcuri, S., De Iorio, T. et al. Combination of epigenetic erasing and mechanical cues to generate human epiBlastoids from adult dermal fibroblasts. J Assist Reprod Genet 40, 1015–1027 (2023). https://doi.org/10.1007/s10815-023-02773-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-023-02773-4