Abstract
STAT2 is distinguished from other STAT family members by its exclusive involvement in type I and III interferon (IFN-I/III) signaling pathways, and its unique behavior as both positive and negative regulator of IFN-I signaling. The clinical relevance of these opposing STAT2 functions is exemplified by monogenic diseases of STAT2. Autosomal recessive STAT2 deficiency results in heightened susceptibility to severe and/or recurrent viral disease, whereas homozygous missense substitution of the STAT2-R148 residue is associated with severe type I interferonopathy due to loss of STAT2 negative regulation. Here we review the clinical presentation, pathogenesis, and management of these disorders of STAT2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inborn errors of immunity are important not only in their own right as serious human diseases, but for what they teach us about the action and regulation of pathways of human immunity. Over the last two decades, a range of monogenic disorders have been identified that impact, in opposing ways, the activity of the antiviral type I and III interferon (IFN-I/III) systems. The clinical consequence of these defects (reviewed elsewhere [1,2,3]) includes (i) susceptibility to severe viral disease, including pandemic SARS-CoV-2 [4], due to inadequate IFN-I/III activity; or (ii) a spectrum of autoinflammatory disease phenotypes, collectively termed type I interferonopathies, associated with excessive IFN-I activity. This knowledge has led to a greater appreciation of the protective and pathogenic effects of IFNs in humans. Mendelian disorders of the transcription factor STAT2 have contributed significantly to our understanding of these dual roles of IFN-I in antiviral defense and immunopathology, by underlining the unique function of STAT2 as both positive and negative regulator of IFN-I activity.
STAT2
The principal role of STAT2 is to transduce signals downstream of the receptors for IFN-I and IFN-III. STAT2 was identified by the labs of Darnell and Stark, in elucidating the molecular pathways governing the response to IFN-I. Key discoveries were the identification of an interferon-stimulated response element (ISRE) in DNA [5] that was bound by a polyprotein complex termed interferon-stimulated gene factor 3 (ISGF3) [6]. ISGF3 included a 113-kDa protein, subsequently shown to be the product of the STAT2 gene [7]. Parallel mutagenesis studies in the human fibrosarcoma cell line U6A identified STAT2 to be an essential activator of gene transcription in response to IFN-I but not IFNγ [8]. The relevance of STAT2 to antiviral immunity was subsequently confirmed by generation of STAT2 knockout (Stat2 − / −) mice [9], which showed impaired responses to IFN-I and were susceptible to vesicular stomatitis virus (VSV) infection, similar to Stat1 − / − [10] or Ifnar1 − / − [11] mice.
STAT2 Signaling
These seminal studies led to the canonical model of STAT2 signaling summarized in Fig. 1. In this model, STAT2 is activated by tyrosine phosphorylation through the action of receptor-associated kinases JAK1 [12] and TYK2 [13]. It associates with tyrosine phosphorylated STAT1 and IRF9 in a heterotrimeric transcription factor complex known as interferon-stimulated gene factor 3 (ISGF3). ISGF3 translocates to the nucleus, binding to ISRE in the promoters of hundreds of interferon-stimulated genes (ISGs). A small contribution to the antiviral response is also made by tyrosine phosphorylated homodimers of STAT1, which bind to a separate motif known as the gamma activated sequence (GAS) [14].
While this is a useful conceptual model, findings over the last few years indicate a more complex reality. For instance, STAT2 participates in transcriptionally active complexes other than ISGF3. These “noncanonical” complexes include STAT2:IRF9 [15] and an unphosphorylated form of ISGF3 [16, 17]. The topic of noncanonical STAT2 signaling has recently been reviewed [18]. A recent study also challenges the notion that ISGF3 assembles in the cytosol, as in the canonical model, suggesting instead that uSTAT2:IRF9 is bound to DNA under homeostatic conditions, where it governs basal transcription, and is subsequently displaced by ISGF3 upon IFN-I treatment [15]. Regardless of the precise mode of action of STAT2, its importance to human antiviral immunity has been revealed by the discovery of humans with homozygous STAT2 deficiency, who exhibited a clinical phenotype of increased susceptibility to various viruses [19,20,21,22,23]. Clinical aspects of this disorder will be considered in more detail in a later section.
STAT2 Structure and Interactions
In common with other members of the STAT family, STAT2 has six functional domains—the N terminal domain (NTD), coiled coil domain (CCD), DNA binding domain (DBD), linker domain (LD), Src homology 2 (SH2) domain, and the C-terminal transactivation domain (TAD; Fig. 2).
The activity of STAT2 is regulated by post-translational modification. The most well-known is phosphorylation of Y690 which enhances its transcriptional activity. Mutagenesis studies have identified that phosphorylation of additional residues (S287 [24], T387 [25], S734 [26]) on STAT2 negatively regulates its transcriptional activity. STAT2 T387 is constitutively phosphorylated, negatively regulating ISGF3 binding to DNA; constitutive phosphorylation at T403 maintains STAT2 dimerization with STAT1 [27]. By contrast, IFN-mediated acetylation of STAT2 K390 promotes transcription [28] (Fig. 2).
Unphosphorylated STAT2 shuttles continuously between the cytosol and the nucleus, but owing to a strong nuclear export signal (NES) is predominantly cytosolic [29]. Here, STAT2 associates with various proteins including IRF9 [30, 31], STAT1 [32, 33], and IFNAR2 [34]. The interactions with STAT1 and IRF9 are mediated by the NTD and CCD of STAT2 respectively. Interestingly, although STAT2:IRF9 or STAT1:STAT2 complexes can be readily identified, there is limited evidence that STAT1 and IRF9 directly interact [15, 31]. The interaction domain with IFNAR2 has not been precisely mapped, but involves the proximal third (1–315) of STAT2, incorporating the NTD and CCD [34]. Upon phosphorylation at Y690, STAT2 rapidly accumulates in the nucleus due to a conformational change that impedes accessibility of the STAT2 NES [29].
Certain aspects of STAT2 structure and function distinguish it from other STATs. For example, STAT2 does not bind directly to DNA, as it lacks a strong DNA binding domain (it relies on STAT1 and/or IRF9 to bind to DNA) [35]. STAT2 makes an essential contribution to ISGF3 function by recruiting transcriptional coactivators through its TAD [35, 36]. It also exhibits the most interspecies divergence of all STAT molecules, particularly in the TAD [37]. Nevertheless, mouse STAT2 TAD can complement human STAT2 activity [38, 39].
Viral Targeting of STAT2
Selection pressure exerted by viruses has been proposed as an explanation for the increased sequence diversity of STAT2 [37]. Reflecting the critical role of STAT2 in the antiviral IFN response [9, 19], certain viruses target STAT2 for degradation as an IFN evasion strategy. Examples are flaviviruses (dengue, Zika, and hepatitis C virus) [40,41,42], paramyxoviruses (parainfluenza virus 2 and respiratory syncytial virus) [43,44,45], and herpesviruses (CMV) [46]. There is evidence that the sequence diversity of STAT2 restricts interspecies viral transmission, in that it impacts the ability of certain viral proteins to bind and degrade STAT2 of other species [40, 41, 47]. The result is that certain viral pathogens only cause disease in humans. Examples include dengue [40] and Zika [41] virus and human metapneumovirus [48].
Regulatory STAT2 Functions
Beyond its importance to antiviral immunity, in recent years STAT2 has been shown to participate in regulation of immune signaling. Functions recently identified include cross-regulation of STAT1 [32] and NF-KB signaling [49], as well as negative regulation of IFNAR signaling [50], via the ubiquitin-specific protease 18 (USP18).
STAT2 was shown to cooperate with the NF-KB pathway to positively regulate the induction of the IL6 gene [49]. IL6 is an important proinflammatory cytokine. Nan and colleagues showed that when STAT2 and IRF9 expression was increased, the uSTAT2:IRF9 complex interacted with p65, bridging the ISRE and KB elements in the IL6 promoter, leading to the enhanced expression of IL6 in response to IL-1B, tumor necrosis factor, or lipopolysaccharide (LPS) [49].
STAT2 also has negative regulatory activity toward cytokine signaling pathways. In resting cells, STAT2 and STAT1 bind strongly [32], with the net effect of retaining STAT1 in the cytosol via the dominant activity of the STAT2 NES (discussed above). Upon exposure to cytokines that activate STAT1 but not STAT2 (e.g., IFNγ, IL27), the interaction inhibits pSTAT1 from forming homodimers and participating in signaling [32]. Thus, in the absence of STAT2, the transcriptional output of these cytokines becomes dominated by STAT1 [32]. Loss of this regulatory function may contribute to certain inflammatory manifestations of STAT2 deficiency [20,21,22], described below.
STAT2 has also been recently shown to participate in negative feedback toward IFN-I signaling [50], where it supports the activity of a key negative regulator, USP18 [51, 52]. USP18 is an essential regulator of IFN-I signaling, as revealed by the profound pathological consequences for USP18-deficient humans and mice [53,54,55]. The precise details of STAT2’s role in supporting USP18-mediated regulation will be discussed in more detail below. The clinical importance of this latter function of STAT2 was recently confirmed by the discovery of children with fatal IFN-I-mediated inflammatory disease attributed to homozygous missense mutations of STAT2 [56, 57].
Inborn Errors of Immunity Caused by Mutations in STAT2
Autosomal Recessive STAT2 Deficiency
Genetics
The human STAT2 gene is found on chromosome 12 and contains 24 exons. Homozygous or compound heterozygous variants in STAT2 leading to complete deficiency of STAT2 protein have been identified in 11 individuals in five kindreds [19,20,21,22, 46]. Five distinct loss of expression variants have been reported, resulting in either frameshift or splicing defects leading to nonsense mediated RNA decay. Heterozygous carriers of these variants appear clinically unaffected. Mutations associated with complete STAT2 deficiency and the associated clinical phenotypes are summarized in Table 1.
Viral Susceptibility
The primary manifestation of autosomal recessive (AR) STAT2 deficiency is susceptibility to severe and/or recurrent viral disease in individuals without other clinical or laboratory evidence of immunodeficiency. A particularly striking aspect of this phenotype is susceptibility to disease caused by live-attenuated viral (LAV) vaccines, such as measles, mumps, and rubella (MMR) or varicella zoster virus (vVZV). This is in common with other monogenic defects of IFN-I/III immunity (reviewed in [3]). Of the eight STAT2-deficient individuals known to have been exposed to LAV vaccines, all developed viral illness in temporal association. Vaccine-strain viral dissemination was confirmed by PCR in 4/8 cases (in the others, testing was either not done or not reported). One STAT2-deficient individual was identified in adulthood by family screening [19]. The expectation is that she would have been in receipt of measles vaccine in childhood, and had antibodies to measles consistent with exposure to either wild-type virus or vaccine [19]. Thus, susceptibility to LAV vaccines may not be fully penetrant in STAT2-deficient patients. Problems handling other LAV vaccines (such as the yellow fever vaccine) have not been reported in STAT2 deficiency, but would be expected by analogy to homozygous IFNAR1 deficiency [58] or IFN-I autoantibodies [59].
In addition to susceptibility to LAV vaccines, which serves as a “red flag” for defects of IFN-I/III immunity (reviewed in [3]), STAT2-deficient patients also experience increased susceptibility to naturally acquired viral disease. This includes a range of DNA and RNA viruses acquired at the mucosal surface, including influenza virus, enteroviruses, EBV, and adenovirus (Table 1), presumably due to the involvement of STAT2 in both IFN-I and IFN-III signaling pathways [3]. The penetrance of this phenotype is more variable, ranging from death in early infancy or childhood to survival into adulthood with no obvious phenotype [19], recalling the incomplete clinical penetrance of several other inborn errors of innate immunity [60,61,62]. Also in common with such disorders, [63, 64], and despite limited follow-up of STAT2 deficiency to date, there seems to be a reduction in severity of infections over time [19, 22, 23], which might point to the maturation of compensatory adaptive immunity [62]. STAT2-deficient individuals generally have normal laboratory indices of adaptive immunity, and mount appropriate serological responses to vaccination.
STAT2-deficient patient cells demonstrate defects of ISG expression and induction of the antiviral state in response to IFN-I, which can be rescued by STAT2 complementation. This defect can also be overcome in vitro by treatment with IFNγ [21]. Whether IFNγ might offer an option for antiviral therapy in patients with STAT2 deficiency has not been tested. In part, this may be due to concerns that use of IFNγ during acute viral disease might exacerbate the hyperinflammatory state that can accompany viral disease in STAT2 deficiency.
Hyperinflammation
Hyperinflammatory features such as prolonged fevers requiring hospitalization in response to viral infection [20, 21], unprovoked sepsis-like presentations [20, 21], and even HLH [22, 23] have been noted in approximately two-thirds of patients with AR STAT2 deficiency (Table 1). The pathogenesis is unknown, and may be multifactorial. In most cases, hyperinflammation occurred in the context of viral infection or live-attenuated viral vaccination, implying that viral infection is a trigger. However, the occurrence of cases of hyperinflammation without convincing evidence of viral infection [20] raises the possibility of a more complex defect of STAT2-dependent immunoregulation. This is not surprising, considering the emerging evidence for immunoregulatory functions of STAT2.
As in patients, STAT2-deficient mice exhibit inflammatory phenotypes. Stat2 − / − mice are prone to hyperinflammation and macrophage activation following influenza infection [65]. This hyperinflammatory state was able to confer protection against bacterial superinfection. Furthermore, Stat2 − / − mice are more susceptible to endotoxic shock [66], in contrast to Ifnar1 − / − mice which are protected [67]. The mechanism(s) underlying these phenomena have not been elucidated. Deletion of STAT2 in murine macrophages has been shown to alter their cellular response to inflammatory signals. Stat2 − / − macrophages express MHC class II in response to IFN-I, through a mechanism involving IRF1 and STAT1 [68], whereas in WT macrophages MHC class II is typically induced by IFNγ. Thus, loss of STAT2 seemingly alters the transcriptional response to IFN-I, potentially with inflammatory consequences. Whether this is also true in humans has yet to be established. Considering the regulatory functions of STAT2 described later in this review, it is conceivable that STAT2 loss may have more complex effects on immunoregulation. Further studies are warranted to explore the immunological basis of hyperinflammation in STAT2 deficiency.
Diagnosis and Management
Diagnosis of STAT2 deficiency relies on a high index of clinical suspicion and is confirmed by genetic testing. Although not clinically validated, laboratory screening approaches prior to genetic testing may increase the yield. Such assays include analysis of IFN-I signaling activity and/or STAT2 protein expression by immunoblot [19, 23].
Much remains to be learned about the optimal clinical management of STAT2 deficiency. While LAV vaccines (e.g., MMR, varicella, yellow fever) should be avoided, other inactivated and recombinant vaccines are strongly advised. Immunoglobulin supplementation has been proposed as a therapy in STAT2 deficiency, and was associated with a reduction in the frequency of infections and episodes of inflammation in two cases in which it was used [21, 22]. Owing to the range of clinical expressivity noted in STAT2-deficient kindreds, where it is clear that some STAT2-deficient patients live into adulthood with no apparent disease phenotype, there remains a case for individualized therapeutic decision-making.
Management of hyperinflammation in AR STAT2 deficiency is similarly an evolving area, requiring further mechanistic studies to understand its pathophysiology. As mentioned, IVIG may have a role in acute management of hyperinflammatory episodes [21, 22], as it does in other inflammatory syndromes [69]. Its mechanism of action in this context is unclear. There may also be a role for other immunomodulators, by analogy to COVID-19 in adults [70, 71]. However, episodes of hyperinflammation have also been reported to resolve with conservative management [23].
Hematopoietic stem cell transplantation (HSCT) has not been undertaken to date in AR STAT2 deficiency, unlike AR STAT1 deficiency [72]. The latter, in addition to conferring a profound defect of IFN-I/III signaling, critically disables IFNγ signaling between cells of the immune system which are replaced during allogeneic HSCT. In contrast, the viral susceptibility of STAT2 deficiency probably results from impaired IFN-I/III signaling in non-hematopoietic tissues, untouched by HSCT. Furthermore, the transplant process is inevitably accompanied by a temporary loss of adaptive immune protection against viral infection that would be particularly hazardous in the context of impaired innate immunity. This does not necessarily preclude a possible role for HSCT in special circumstances, for example, severe/recurrent treatment-refractory HLH.
Autosomal Recessive STAT2-Associated Type I Interferonopathy (STAT2 Gain of Function)
Genetics
Homozygous missense variants affecting the same arginine 148 residue of STAT2 have been identified in three children in two kindreds with severe early-onset type I interferonopathy [56, 57]. Heterozygous carriers of these variants appear clinically unaffected. The mutations and their associated phenotypes are summarized in Table 2. Type I interferonopathy is a term used to describe a group of Mendelian diseases characterized by neurological and multisystem disease associated with increased IFN-I activity in blood and cerebrospinal fluid (reviewed in [73]). Aicardi-Goutières syndrome [74], which phenocopies congenital viral infection, is a prototypical type I interferonopathy.
Clinical Phenotype
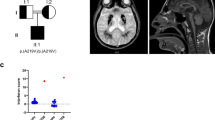
STAT2-associated type I interferonopathy was originally reported in two brothers born of consanguineous parents of Pakistani origin, bearing a very rare homozygous missense substitution of arginine by tryptophan (R148W) in STAT2 [56]. The proband presented with recurrent episodes of sterile systemic inflammation, which met clinical diagnostic criteria for HLH, accompanied by neurological features suggestive of type I interferonopathy [73], such as seizures, intracranial calcifications, hemorrhages, cerebral white matter changes, and developmental regression. Investigations revealed transcriptional evidence of heightened IFN activity in whole blood. There was a clinical response to corticosteroids and the JAK inhibitor ruxolitinib; however, he died due to complications of HSCT. His younger brother was more severely affected, particularly from the neurological perspective, and despite treatment with ruxolitinib did not survive beyond early infancy. In parallel, another individual was reported with a similar clinical phenotype (including seizures and intracranial calcification) associated with a homozygous missense variant affecting the same residue of STAT2—in this case replacing arginine with glutamine (R148Q) [57]. This infant similarly had transcriptional evidence of increased IFN activity in whole blood. Additional features were fistulating adenitis and progressive lung disease which led to his death aged 5 months. There was a family history of infant death in two siblings with a similar clinical syndrome. In all three patients, the phenotype recalled USP18 deficiency [54, 55] as summarized in Table 2.
In cells from patients bearing STAT2 R148 variants, there was evidence of significantly enhanced late transcriptional responses to IFN-I [56, 57]. In the case of R148W cells, there was also a clear phenotype of prolonged phosphorylation of JAK1, STAT1, and STAT2 [56] indicative of a defect of negative regulation of IFNAR signaling upstream of STAT2, reminiscent of USP18 deficiency [54]. This phenotype was not observed in response to IFNγ [56, 57] or other cytokines [56] and thus was specific to IFN-I. Patient fibroblasts from patients [56] or reconstituted U6A cells [57] were insensitive to overexpression of USP18, while knockdown of USP18 had no effect [56], indicating that USP18 function was impaired in the presence of R148W/Q variants. These findings indicated a defect of STAT2’s supportive function toward USP18 [50].
Molecular Pathogenesis
USP18 fulfills its negative feedback function by binding to IFNAR2, displacing JAK1 and altering the conformation of the IFN-IFNAR1-IFNAR2 complex [75]. This impedes JAK1 phosphorylation—an essential step in IFNAR signaling—and consequently blocks tyrosine phosphorylation of STAT1 and STAT2 [50, 52, 75]. USP18 expression is induced by ISGF3 signaling and its regulatory activity continues for the duration of its expression. Thus, USP18 is primarily responsible [52, 76] for the phenomena recognized in IFN biology whereby cells, after IFN-I treatment, become refractory to further restimulation [77]. STAT2 is essential for this function of USP18 [50]. A key question is how the R148W/Q mutations impair this regulatory function of STAT2.
USP18 is known to interact independently with both STAT2 and IFNAR2 via adjacent domains of USP18 [50]. In the current paradigm, STAT2 recruits USP18 to IFNAR2 (Fig. 3). The USP18:IFNAR2 interaction is substantially impaired (although not completely abolished) in the absence of STAT2 [50]. Consistent with this, deletion of the STAT2 binding site on IFNAR2 reduced the binding of USP18 to IFNAR2 [50]. The R148 residue is located in the CCD of STAT2, a region previously implicated in the interactions with both USP18 [50] and IFNAR2 [34]. Our studies demonstrated impaired interaction between STAT2-R148W and USP18, as measured by coimmunoprecipitation in U6A cells stably expressing WT or STAT2-R148W and treated with IFNα to induce USP18 expression [56] (Fig. 3). This was consistent with prior findings [50] and implied a defect of STAT2-dependent recruitment of USP18 to IFNAR2. Gruber and colleagues confirmed this in coimmunoprecipitation experiments in transiently transfected U6A cells overexpressing WT or STAT2-R148Q and USP18, showing a reduced interaction between USP18 and IFNAR2 in the presence of STAT2-R148Q (Fig. 3) [57]. However, the interactions between STAT2-R148Q and IFNAR2, as well as STAT2-R148Q and USP18, were preserved in coimmunoprecipitation experiments conducted in HEK293 and U6A cells [57]. Further work may resolve the precise molecular mechanism(s) underlying the impairment of USP18 activity in the context of STAT2 R148W/Q variants. Nevertheless, it is clear that the immunological and clinical impact of the loss of this regulatory activity of STAT2 is profound.
Models of STAT2-associated type I interferonopathy (STAT2 gain of function) pathogenesis. USP18 binds STAT2 and IFNAR2, displacing JAK1 from the cytoplasmic domain of IFNAR2 and inducing a conformational change in the IFN-IFNAR1-IFNAR2 complex, leading to impaired signal transduction (left). The R148W variant of STAT2 impairs interaction with USP18 (middle), whereas the R148Q variant demonstrates preserved interaction with USP18 but a defect of recruitment to IFNAR2 (right), leading to a defect of USP18-mediated negative feedback
Diagnosis and Management
Clinical recognition of this very rare disease relies on awareness of the phenotype and is aided by identification of transcriptional evidence of elevated IFN activity in blood, as for other type I interferonopathies [73]. A suitable assay to rapidly screen for STAT2-R148W is to examine peripheral blood by phosflow for prolonged IFNα-induced phosphorylation of STAT1 and/or STAT2 [56]. Based on limited experience in STAT2-R148W [56] and USP18 deficiency [55], rapid initiation of JAK inhibitor treatment is a priority and can be life-saving. Interestingly, the clinical response to ruxolitinib was dose-dependent in USP18 deficiency, requiring doses above 5 mg b.d. to achieve clinical remission [55], although such doses are justified by the grave prognosis.
Concluding Remarks
Molecular defects of STAT2 have already taught us much about the biological function of STAT2 in humans. One of the most interesting aspects is the relatively “mild” clinical phenotype of AR STAT2 deficiency, despite the extent of compromise to IFN-I and IFN-III systems. This presumably reflects both the capacity of viral pathogens to evade innate IFN restriction, and the residual ability of other facets of antiviral immunity to compensate. The variable clinical penetrance of AR STAT2 deficiency, even within the same kindred, is notable. Clearly, given the contribution of viral infection to the disease phenotype, pathogen exposure, infectious dose, and other virological factors are likely to be important determinants of penetrance. Indeed, penetrance is seemingly less variable in circumstances where exposure is more controlled, for example, in the context of administration of live-attenuated viral vaccine(s) to STAT2-deficient patients. In assessing individual patients, serological testing for prior exposure may be helpful to inform the extent of the vulnerability to naturally acquired viruses. A question for future studies is whether incomplete penetrance in STAT2 deficiency may also be governed by the effectiveness of adaptive immune compensation.
Disease associated with STAT2-R148Q has been termed STAT2 GOF. This terminology is convenient as it (i) helps to distinguish it from AR STAT2 deficiency and (ii) conforms to a well-recognized paradigm for other STATs (e.g., STAT1 or STAT3), where both LOF and GOF variants are recognized. However, mutations which lead to a gain of protein function that is pathogenic only in the homozygous state are exceedingly rare in nature [78, 79]. Indeed, GOF variants in STAT1/3 [80,81,82,83] or JAK1 [84, 85] manifest in the heterozygous state. In the case of STAT2, the GOF nomenclature is potentially misleading as it fails to account for the specific molecular defect of R148 STAT2, distinct from its role in ISGF3 [56]. From a protein-centric viewpoint, it is clear that STAT2-R148W/Q mutations impair the regulatory function of STAT2 toward USP18 [56, 57]. In other words, they cause a pathological loss of this particular STAT2 function.
The fascinating aspect of this disorder is that it underscores the unique role of STAT2 as both positive and negative regulator of IFN-I signaling pathway (Fig. 4). A key feature of STAT2 GOF (alongside similar defects such as USP18 deficiency) is its very early onset, apparently without an overt infectious precipitant, suggesting that physiological levels of IFNs produced during homeostasis are sufficient to initiate disease. It is conceivable that additional pathogenic variants in STAT2 might be identified in future that provide further insight into STAT2 biology. By analogy with STAT1 or STAT3 [80,81,82,83], we might predict autosomal dominant variants that confer a GOF of STAT2’s positive transcriptional activity, for example, by interfering with dephosphorylation of Y690, or via the loss of a regulatory residue such as T387. It is debatable whether partial LOF of STAT2, through hypomorphic AR or dominant negative AD inheritance, would be clinically manifest. The wider application of genomic sequencing methods, alongside in vitro and ex vivo measures of interferon signaling and viral susceptibility, will no doubt present opportunities to test these hypotheses in the coming years.
Data Availability
Not applicable
References
Uggenti C, et al. Self-awareness: nucleic acid-driven inflammation and the type I interferonopathies. Annu Rev Immunol. 2019;37:247–67.
Moens L, Meyts I. Recent human genetic errors of innate immunity leading to increased susceptibility to infection. Curr Opin Immunol. 2020;62:79–90.
Duncan CJA, et al. Genetic lesions of type I interferon signalling in human antiviral immunity. Trends Genet. 2021;37(1):46–58.
Zhang Q et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science. 2020;370(6515):eabd4570
Reich N, et al. Interferon-induced transcription of a gene encoding a 15-kDa protein depends on an upstream enhancer element. Proc Natl Acad Sci U S A. 1987;84(18):6394–8.
Levy DE, et al. Interferon-induced nuclear factors that bind a shared promoter element correlate with positive and negative transcriptional control. Genes Dev. 1988;2(4):383–93.
Fu XY, et al. The proteins of ISGF-3, the interferon alpha-induced transcriptional activator, define a gene family involved in signal transduction. Proc Natl Acad Sci U S A. 1992;89(16):7840–3.
Leung S, et al. Role of STAT2 in the alpha interferon signaling pathway. Mol Cell Biol. 1995;15(3):1312–7.
Park C, et al. Immune response in Stat2 knockout mice. Immunity. 2000;13(6):795–804.
Meraz MA, et al. Targeted disruption of the Stat1 gene in mice reveals unexpected physiologic specificity in the JAK-STAT signaling pathway. Cell. 1996;84(3):431–42.
Muller U, et al. Functional role of type I and type II interferons in antiviral defense. Science. 1994;264(5167):1918–21.
Muller M, et al. The protein tyrosine kinase JAK1 complements defects in interferon-alpha/beta and -gamma signal transduction. Nature. 1993;366(6451):129–35.
Velazquez L, et al. A protein tyrosine kinase in the interferon alpha/beta signaling pathway. Cell. 1992;70(2):313–22.
Decker T, et al. Cytoplasmic activation of GAF, an IFN-gamma-regulated DNA-binding factor. EMBO J. 1991;10(4):927–32.
Platanitis E, et al. A molecular switch from STAT2-IRF9 to ISGF3 underlies interferon-induced gene transcription. Nat Commun. 2019;10(1):2921.
Cheon H, et al. IFNbeta-dependent increases in STAT1, STAT2, and IRF9 mediate resistance to viruses and DNA damage. EMBO J. 2013;32(20):2751–63.
Wang W, et al. Unphosphorylated ISGF3 drives constitutive expression of interferon-stimulated genes to protect against viral infections. Sci Signal. 2017;10(476):eaah4248.
Blaszczyk K, et al. The unique role of STAT2 in constitutive and IFN-induced transcription and antiviral responses. Cytokine Growth Factor Rev. 2016;29:71–81.
Hambleton S, et al. STAT2 deficiency and susceptibility to viral illness in humans. Proc Natl Acad Sci U S A. 2013;110(8):3053–8.
Shahni R, et al. Signal transducer and activator of transcription 2 deficiency is a novel disorder of mitochondrial fission. Brain. 2015;138(Pt 10):2834–46.
Moens L, et al. A novel kindred with inherited STAT2 deficiency and severe viral illness. J Allergy Clin Immunol. 2017;139(6):1995-1997 e9.
Alosaimi MF, et al. A novel variant in STAT2 presenting with hemophagocytic lymphohistiocytosis. J Allergy Clin Immunol. 2019;144(2):611-613 e3.
Freij B et al. Life-threatening influenza, haemophagocytic lymphohistiocytosis and probable vaccine strain varicella in a novel case of homozygous STAT2 deficiency. Front Immunol. 2021;11:624415.
Steen HC, et al. Identification of STAT2 serine 287 as a novel regulatory phosphorylation site in type I interferon-induced cellular responses. J Biol Chem. 2013;288(1):747–58.
Wang Y, et al. Negative regulation of type I IFN signaling by phosphorylation of STAT2 on T387. EMBO J. 2017;36(2):202–12.
Steen HC, et al. Phosphorylation of STAT2 on serine-734 negatively regulates the IFN-alpha-induced antiviral response. J Cell Sci. 2016;129(22):4190–9.
Wang Y, et al. A virus-induced conformational switch of STAT1-STAT2 dimers boosts antiviral defenses. Cell Res. 2021;31(2):206–18.
Tang X, et al. Acetylation-dependent signal transduction for type I interferon receptor. Cell. 2007;131(1):93–105.
Frahm T, et al. IFN-type-I-mediated signaling is regulated by modulation of STAT2 nuclear export. J Cell Sci. 2006;119(Pt 6):1092–104.
Rengachari S, et al. Structural basis of STAT2 recognition by IRF9 reveals molecular insights into ISGF3 function. Proc Natl Acad Sci U S A. 2018;115(4):E601–9.
Martinez-Moczygemba M, et al. Distinct STAT structure promotes interaction of STAT2 with the p48 subunit of the interferon-alpha-stimulated transcription factor ISGF3. J Biol Chem. 1997;272(32):20070–6.
Ho J, et al. STAT2 Is a pervasive cytokine regulator due to its inhibition of STAT1 in multiple signaling pathways. PLoS Biol. 2016;14(10):e2000117.
Stancato LF, et al. Preassociation of STAT1 with STAT2 and STAT3 in separate signalling complexes prior to cytokine stimulation. J Biol Chem. 1996;271(8):4134–7.
Li X, et al. Functional subdomains of STAT2 required for preassociation with the alpha interferon receptor and for signaling. Mol Cell Biol. 1997;17(4):2048–56.
Bluyssen HA, Levy DE. Stat2 is a transcriptional activator that requires sequence-specific contacts provided by stat1 and p48 for stable interaction with DNA. J Biol Chem. 1997;272(7):4600–5.
Qureshi SA, et al. Function of Stat2 protein in transcriptional activation by alpha interferon. Mol Cell Biol. 1996;16(1):288–93.
Chowdhury FZ, Farrar JD. STAT2: a shape-shifting anti-viral super STAT. JAKSTAT. 2013;2(1):e123633.
Park C, et al. Murine STAT2 is uncharacteristically divergent. Nucleic Acids Res. 1999;27(21):4191–9.
Paulson M, et al. Stat protein transactivation domains recruit p300/CBP through widely divergent sequences. J Biol Chem. 1999;274(36):25343–9.
Ashour J, et al. Mouse STAT2 restricts early dengue virus replication. Cell Host Microbe. 2010;8(5):410–21.
Grant A, et al. Zika virus targets human STAT2 to inhibit type I interferon signaling. Cell Host Microbe. 2016;19(6):882–90.
Joyce MA, et al. HCV and flaviviruses hijack cellular mechanisms for nuclear STAT2 degradation: up-regulation of PDLIM2 suppresses the innate immune response. PLoS Pathog. 2019;15(8):e1007949.
Ulane CM, Horvath CM. Paramyxoviruses SV5 and HPIV2 assemble STAT protein ubiquitin ligase complexes from cellular components. Virology. 2002;304(2):160–6.
Precious B, et al. In vitro and in vivo specificity of ubiquitination and degradation of STAT1 and STAT2 by the V proteins of the paramyxoviruses simian virus 5 and human parainfluenza virus type 2. J Gen Virol. 2005;86(Pt 1):151–8.
Xu X, et al. Respiratory syncytial virus NS1 protein degrades STAT2 by inducing SOCS1 expression. Intervirology. 2014;57(2):65–73.
Le VTK, et al. Human cytomegalovirus interferes with signal transducer and activator of transcription (STAT) 2 protein stability and tyrosine phosphorylation. J Gen Virol. 2008;89(Pt 10):2416–26.
Wang B, et al. Structural basis for STAT2 suppression by flavivirus NS5. Nat Struct Mol Biol. 2020;27(10):875–85.
Rogers MC, et al. STAT2 limits host species specificity of human metapneumovirus. Viruses. 2020;12(7):724.
Nan J, et al. IRF9 and unphosphorylated STAT2 cooperate with NF-kappaB to drive IL6 expression. Proc Natl Acad Sci U S A. 2018;115(15):3906–11.
Arimoto KI, et al. STAT2 is an essential adaptor in USP18-mediated suppression of type I interferon signaling. Nat Struct Mol Biol. 2017;24(3):279–89.
Malakhova OA, et al. UBP43 is a novel regulator of interferon signaling independent of its ISG15 isopeptidase activity. EMBO J. 2006;25(11):2358–67.
Francois-Newton V, et al. USP18-based negative feedback control is induced by type I and type III interferons and specifically inactivates interferon alpha response. PLoS One. 2011;6(7):e122200.
Goldmann T, et al. USP18 lack in microglia causes destructive interferonopathy of the mouse brain. EMBO J. 2015;34(12):1612–29.
Meuwissen ME, et al. Human USP18 deficiency underlies type 1 interferonopathy leading to severe pseudo-TORCH syndrome. J Exp Med. 2016;213(7):1163–74.
Alsohime F, et al. JAK inhibitor therapy in a child with inherited USP18 deficiency. N Engl J Med. 2020;382(3):256–65.
Duncan CJA, et al. Severe type I interferonopathy and unrestrained interferon signaling due to a homozygous germline mutation in STAT2. Sci Immunol. 2019;4(42):eaav7501.
Gruber C, et al. Homozygous STAT2 gain-of-function mutation by loss of USP18 activity in a patient with type I interferonopathy. J Exp Med. 2020;217(5):e20192319.
Hernandez N, et al. Inherited IFNAR1 deficiency in otherwise healthy patients with adverse reaction to measles and yellow fever live vaccines. J Exp Med. 2019;216(9):2057–70.
Bastard P, et al. Auto-antibodies to type I IFNs can underlie adverse reactions to yellow fever live attenuated vaccine. J Exp Med. 2021;218(4):e20202486.
Zhang SY, et al. TLR3 deficiency in patients with herpes simplex encephalitis. Science. 2007;317(5844):1522–7.
Andersen LL, et al. Functional IRF3 deficiency in a patient with herpes simplex encephalitis. J Exp Med. 2015;212(9):1371–9.
Israel L, et al. Human adaptive immunity rescues an inborn error of innate immunity. Cell. 2017;168(5):789-800 e10.
Picard C, et al. Pyogenic bacterial infections in humans with IRAK-4 deficiency. Science. 2003;299(5615):2076–9.
von Bernuth H, et al. Pyogenic bacterial infections in humans with MyD88 deficiency. Science. 2008;321(5889):691–6.
Gopal R, et al. STAT2 Signaling regulates macrophage phenotype during influenza and bacterial super-infection. Front Immunol. 2018;9:2151.
Alazawi W, et al. Stat2 loss leads to cytokine-independent, cell-mediated lethality in LPS-induced sepsis. Proc Natl Acad Sci U S A. 2013;110(21):8656–61.
Karaghiosoff M, et al. Central role for type I interferons and Tyk2 in lipopolysaccharide-induced endotoxin shock. Nat Immunol. 2003;4(5):471–7.
Zhao W, et al. Stat2-dependent regulation of MHC class II expression. J Immunol. 2007;179(1):463–71.
Katz U, et al. Update on intravenous immunoglobulins (IVIg) mechanisms of action and off- label use in autoimmune diseases. Curr Pharm Des. 2011;17(29):3166–75.
Group, R.C. , et al. Dexamethasone in hospitalized patients with COVID-19. N Engl J Med. 2021;384(8):693–704.
Investigators R-C, et al. Interleukin-6 receptor antagonists in critically ill patients with COVID-19. N Engl J Med. 2021;384(16):1491–502.
Naviglio S, et al. Long-term survival after hematopoietic stem cell transplantation for complete STAT1 deficiency. J Clin Immunol. 2017;37(7):701–6.
Crow YJ, Manel N. Aicardi-Goutieres syndrome and the type I interferonopathies. Nat Rev Immunol. 2015;15(7):429–40.
Aicardi J, Goutieres F. A progressive familial encephalopathy in infancy with calcifications of the basal ganglia and chronic cerebrospinal fluid lymphocytosis. Ann Neurol. 1984;15(1):49–54.
Lochte S, et al. Live cell micropatterning reveals the dynamics of signaling complexes at the plasma membrane. J Cell Biol. 2014;207(3):407–18.
Sarasin-Filipowicz M, et al. Alpha interferon induces long-lasting refractoriness of JAK-STAT signaling in the mouse liver through induction of USP18/UBP43. Mol Cell Biol. 2009;29(17):4841–51.
Larner AC, et al. Transcriptional induction by interferon. New protein(s) determine the extent and length of the induction. J Biol Chem. 1986;261(1):453–9.
Cavaco BM, et al. Homozygous calcium-sensing receptor polymorphism R544Q presents as hypocalcemic hypoparathyroidism. J Clin Endocrinol Metab. 2018;103(8):2879–88.
Drutman SB, et al. Homozygous NLRP1 gain-of-function mutation in siblings with a syndromic form of recurrent respiratory papillomatosis. Proc Natl Acad Sci U S A. 2019;116(38):19055–63.
Liu L, et al. Gain-of-function human STAT1 mutations impair IL-17 immunity and underlie chronic mucocutaneous candidiasis. J Exp Med. 2011;208(8):1635–48.
van de Veerdonk FL, et al. STAT1 mutations in autosomal dominant chronic mucocutaneous candidiasis. N Engl J Med. 2011;365(1):54–61.
Flanagan SE, et al. Activating germline mutations in STAT3 cause early-onset multi-organ autoimmune disease. Nat Genet. 2014;46(8):812–4.
Milner JD, et al. Early-onset lymphoproliferation and autoimmunity caused by germline STAT3 gain-of-function mutations. Blood. 2015;125(4):591–9.
Del Bel KL, et al. JAK1 gain-of-function causes an autosomal dominant immune dysregulatory and hypereosinophilic syndrome. J Allergy Clin Immunol. 2017;139(6):2016-2020 e5.
Gruber CN, et al. Complex autoinflammatory syndrome unveils fundamental principles of JAK1 kinase transcriptional and biochemical function. Immunity. 2020;53(3):672-684 e11.
Acknowledgements
We are grateful to the current and past members of the Duncan and Hambleton labs for useful discussions.
Funding
Provided by the Wellcome Trust (211153/Z/18/Z [C.J.A.D.], 207556/Z/17/Z [S.H.], Sir Jules Thorn Trust (12/JTA [S.H.]), and the British Medical Association (C.J.A.D).
Author information
Authors and Affiliations
Contributions
CJAD wrote the first draft; CJAD and SH revised for content.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable
Consent to Participate
Not applicable
Consent for Publication
Not applicable
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Duncan, C.J.A., Hambleton, S. Human Disease Phenotypes Associated with Loss and Gain of Function Mutations in STAT2: Viral Susceptibility and Type I Interferonopathy. J Clin Immunol 41, 1446–1456 (2021). https://doi.org/10.1007/s10875-021-01118-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-021-01118-z