Abstract
I-motifs are tetra-helixes that may form in cytosine-rich strands. They are based on cytosine–cytosine+ base pairs that require the N3 hemi-protonation of the nucleobases, and therefore, the stability of these non-canonical DNA arrangements depends on pH. These structures are promising targets for the development of new cancer therapies since they are enriched in the promoters of oncogenes where they can play a role in the regulation of transcription. The proximal promoter of the EGFR oncogene has multiple regions with a significant potential to form such a tetra-helix arrangement. Here, we present the thermodynamic characterization of a C-rich sequence located 37 nucleotides upstream of the transcription starting site of EGFR. We confirmed the ability of this sequence to fold into an I-motif. By applying a global analysis of calorimetric and spectroscopic data, we derived the dependency of the apparent standard Gibbs free energy change associated with the I-motif folding upon temperature and pH. The results showed that, in contrast to in silico prediction, only 4 CC+ base pairs formed while additional GC and TT base pairings were detected in the I-motif. Noteworthy, a single residue mutation at G14 largely shifts the equilibrium toward the formation of multimeric species.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
I-motifs (iMs) and G-quadruplexes (G4s) are tetra-helical nucleic acid secondary structures that may form at sequences containing four runs of repeated cytosines and guanines, respectively [1, 2]. These nucleotide patterns are complementary thus potentially locating iMs and G4s at the same genomic sites. In the human genome, there is an interesting enrichment of potential iM/G4 forming sequences at regions endowed with relevant biological functions, such as telomeres and oncogene promoters [3, 4]. A detailed description of the molecular events that relate the tetra-helices formation to specific functional roles is not fully disclosed but, it has been proven that their conversion from the double-stranded DNA at gene promoters largely impacts the efficiency of gene transcription [5, 6].
iMs and G4s are based on non-canonical hemi-protonated cytosine–cytosine (CC+) and Hoogsteen guanine–guanine base pairs, respectively [7, 8]. Concerning iMs, CC+ base pairs hold together two parallel oriented strands and two of such parallel duplexes intercalate in an antiparallel orientation. Therefore, iMs are highly ordered structures where the adverse entropy change of folding is balanced by a favorable reduction of the enthalpy [9, 10]. Many factors contribute to this enthalpy change. Among them, theoretical calculations indicate that the three hydrogen bonds of the CC+ base pairs and the bonding networks in the minor grooves between the H1’/H4’ and O4’ of two consecutive sugar residues are the most relevant stabilizing components. Indeed, unlike what is observed in the canonical B-DNA double helix, π–π stacking interactions between consecutive CC+ do not significantly contribute to the enthalpy of the folded state [11].
To support the formation of three hydrogen bonds within the CC+ base pair, the N3 of cytosines must be hemi-protonated. This makes iMs pH-dependent structures and the apparent standard Gibbs free energy change of folding (ΔG θapp (T, pH)) is generally minimized at pH 4.5 which corresponds to the pKa of cytosine N3 [12]. For a long time, the acidic conditions required for their folding were considered incompatible with the presence of iM in the intracellular environment. However, by using iM fluorescent-labeled antibodies and in-cell NMR experiments, it has been proven that they exist in the cell nuclei as well [13, 14]. Noteworthy, the intracellular environment of cancer cells is often slightly acidic and this further increases iM formation [15]. This evidence confirmed iMs as promising targets for the development of new anticancer therapies. On these bases, a detailed structural characterization of the genomic regions potentially prone to fold into iMs is the first step toward a better understanding of the finely tuned mechanisms that correlate their formation with the regulation of oncogene transcription.
Many tumors, such as breast or lung cancers and glioblastoma, are related to an overexpression of the Epidermal Grow Factor Receptor oncogene (EGFR) [16, 17]. Currently, targeted treatments are used in therapy. They are represented by monoclonal antibodies and tyrosine kinase inhibitors which specifically target the EGFR protein. However, frequently these approaches result ineffective because of drug resistance development [18, 19]. A new valid therapeutic strategy is to downregulate the oncogene overexpression at its roots, by targeting the gene itself.
The proximal promoter of EGFR has a large guanine–cytosine-rich region with several potential iM/G4 forming sites. Among them, in the region comprising 500 nucleotides upstream of the transcription starting site, we identified two main potential iM/G4 forming sequences that comprise four runs with at least three consecutive cytosines, EGFR-272 and EGFR-37, located at 272 and 37 nucleotides upstream of the transcription starting site, respectively. In previous works, we characterized the conformational features of both the guanine and cytosine-rich strands of EGFR-272 [20, 21]. We observed that, in solution, the guanine-rich strand folds into two different G4s in thermodynamic equilibrium and we showed that the use of G4-targeted small molecules can drive a reduction of the EGFR expression in treated cells which resulted to be particularly cytotoxic for metastatic castration-resistant prostate cancer cells [22]. Conversely, the cytosine-rich strand folds into a single iM structure. This better-defined structure is bound and stabilized by small molecules as well and thus it represents an alternative complementary target. To fully exploit the potential of iM at EGFR promoter as a target, here we report a comprehensive characterization of the folding of the iM assumed in solution by the cytosine-rich EGFR-37 site through calorimetric and spectroscopic analyses.
Materials and methods
Materials
All tested oligonucleotides (sequences reported in Table 1) were synthesized and purified by Eurogentec (Liege, Belgium). They were dissolved in Milli-Q water to obtain 1 mM stock solutions (strand concentration) as derived by UV absorbance at 260 nm. Before use, all DNA samples were melted for 5 min at 95 °C and then slowly cooled to room temperature.
Methods
Electrophoretic mobility shift assay
Samples were prepared at 4 µM or 200 µM DNA (strand concentration) in 10 mM Na-cacodylate pH 5.5. They were heated at 95 °C and slowly cooled to room temperature before loading them (250 ng DNA/lane) on a 15% native polyacrylamide gel (acrylamide/bis-acrylamide 19:1) in 1 × TAE (40 mM Tris, 20 mM acetic acid, 1 mM EDTA, pH 5.5). The run was performed at 25 °C for 2 h by applying a voltage of 15 V/cm. Gels were stained by soaking them in a 1 × Sybr Green II in 1 × TBE (89 mM Tris, 89 mM boric acid, 2 mM EDTA) solution. DNA bands were visualized on a Geliance system (Roche).
S1 footprinting
Reaction mixtures were prepared using 3’-FAM-labeled DNA (100 ng/µL) in 10 mM Na-cacodylate, 4.5 mM ZnSO4, pH 5.5. They were heated for 5 min at 95 °C and then slowly cooled to room temperature. After the cooling step, 1 U/µL of S1 endonuclease (Promega) was added. Samples were incubated for different times at 25 °C and the reactions were stopped by adding 2.5 µL of the reaction mixture to 9 µL of stop solution (80% formamide, 60 mM EDTA). Samples were heated for 5 min at 95 °C, chilled on ice, and finally loaded on a 20% polyacrylamide (acrylamide/bis-acrylamide 19:1) denaturing gel in 7 M urea, 1 × TBE (89 mM Tris, 89 mM boric acid, 2 mM EDTA). Reaction products were visualized on a Geliance system (Roche).
NMR spectroscopy
1H NMR spectra were recorded on a Bruker DMX 600 MHz spectrometer, equipped with a 5 mm TXI probe XYZ-Gradient at 25 °C. Samples were prepared at 150 µM DNA in 10 mM Na-phosphate, pH 5.5 with 10% D2O. Before data acquisition, they were melted for 5 min at 95 °C and then slowly cooled down to room temperature. Suppression of water signal was achieved by applying WATERGATE pulse sequence. The NMR spectrum was processed and analyzed by using TOPSPIN software.
Thermal difference spectra
Thermal difference spectra (TDS) were obtained by subtracting the DNA UV–Vis spectrum acquired at 1 °C from the one recorded at 85 °C, thus below and above the oligonucleotide melting temperature, respectively. The experiments were performed in 10 mM Na-cacodylate pH 5.5.
Circular dichroism
CD spectra were acquired using a Jasco J 810 spectropolarimeter equipped with a Peltier as a temperature controller device in a 1 cm length quartz cuvette. Signals were reported as Δε. All experiments were repeated in triplicate. pH-titration experiments were performed by adding HCl to 4 µM DNA samples in 10 mM Na-cacodylate pH 8.5 at 298.15 K. The CD signal recorded at 287 nm was fitted with Eq. (1) according to the Hill formalism:
where n is the Hill coefficient, pHT is the pH of the middle transition, a and b are the Δε of the unfolded and folded species, respectively.
Heating and cooling experiments were performed by setting ± 20 K h−1 temperature slopes and recoding the spectra every 2 K in the 230–330 nm wavelength range. The CD signal recorded at 287 nm during the heating and cooling steps was fitted with Eq. (2) according to van't Hoff’s formalism:
where T is the temperature (K), ΔHθ is the standard enthalpy change of folding (kJ mol−1), Tm is the melting temperature (K), R is the ideal gas constant (8.314 × 10–3 kJ K−1 mol−1) and a, b are the Δε of the unfolded and folded species, respectively.
Differential scanning calorimetry
Differential scanning calorimetry experiments were performed on a Microcal VP-DSC with cells of 502.7 µL in the 1–80 °C temperature range at stated heating–cooling rates. Samples were prepared at 200 µM DNA concentration in the required buffer. Data were reported as molar excess of heat capacity (ΔCp) as a function of the temperature. All experiments were repeated in triplicate.
Curves were fitted according to Eq. (3):
where T is the temperature (K), ΔHθ is the standard enthalpy change of folding (kJ mol−1), Tm is the melting temperature (K), and R is the ideal gas constant (8.314 × 10–3 kJ K−1 mol−1).
Global fitting analysis
Global fitting analyses were performed on all the datasets derived from CD and DSC experiments according to the simplest model mechanism that successfully described the folding process:
where U is the unfolded species, F is the folded species, n is the number of protons recruited for the folding, T is the temperature (K), ΔGθ(T) is the standard Gibbs free energy change referred to T and R is the ideal gas constant (8.314 × 10–3 kJ K−1 mol−1). Worth noting, for a DNA folding process, it is more appropriate to consider n as the Hill coefficient [23].
Thermodynamic parameters such as ΔGθ(298.15 K), ΔHθ, and n were kept as shared fitting parameters.
The dataset corresponding to CD dependence upon pH, temperature (heating and cooling), and DSC curves (heating and cooling) were simultaneously fitted according to Eqs. 6–7, respectively:
where n is the Hill coefficient, T is the temperature (K), To is 298.15 K, ΔGθ(To) is the standard Gibbs free energy change of folding (kJ mol−1) referred to To, ΔHθ is the standard enthalpy change of folding (kJ mol−1), R is the ideal gas constant (8.314 × 10–3 kJ K−1 mol−1) and a and b are the Δε of the unfolded and folded species, respectively.
ΔHθ was considered a temperature-independent parameter (ΔCp = 0).
Considering the unimolecular iM folding pathway, the apparent standard Gibbs free energy change was pH-dependent:
where U is the unfolded species, F is the folded species, T is the temperature, ΔG θapp (T, pH) is the apparent standard Gibbs free energy change referred to T and pH, and R is the ideal gas constant (8.314 × 10–3 kJ K−1 mol−1).
ΔG θapp (T, pH) was derived at different temperatures ad pHs according to Eq. (11):
where n is the Hill coefficient, T is the temperature (K), To is 298.15 K, ΔGθ(To) is the standard Gibbs free energy change of folding (kJ mol−1) referred to To, ΔHθ is the standard enthalpy change of folding (kJ mol−1) and R is the ideal gas constant (8.314 × 10–3 kJ K−1 mol−1).
Python scripts based on NumPy and SciPy libraries were developed to perform data analysis and global fitting.
Singular value decomposition
Multiple wavelength CD experiments were analyzed by Singular Value Decomposition (SVD). For each experiment, the dataset was converted into a matrix A in which A[i, j] is the Δε at a given i wavelength and j temperature or pH. A matrix was decomposed, into a product of three matrixes: USVT where U is an orthogonal matrix in which column vectors are the eigenvectors of AAT, V is an orthogonal matrix in which column vectors are the eigenvectors of ATA and S is a rectangular diagonal matrix which values are the eigenvalues of both AAT and ATA.
The A matrix can be well approximated according to Eq. (12):
where n is the number of significant optical components contributing to the signal.
Only the species (k) that simultaneously have autocorrelation coefficient of Uk and Vk ≥ 0.75 and the maximum absolute value of UkSkVkT ≥ 10–3 cm−1 were considered.
Python scripts based on NumPy and SciPy libraries were developed to perform this analysis.
Results
Characterization of EGFR-37 folding
The chiroptical properties of DNA change upon its folding into secondary structures, and this can be followed by circular dichroism (CD). In particular, the antiparallel orientation of the strands in the iM structure correlates with a positive CD signal around 290 nm and a negative one around 260 nm. Therefore, the ability of EGFR-37 to assume iM structures was preliminarily evaluated following the CD signal in the 330–230 nm range while moving from basic (pH 8.5) to acidic (pH 4.0) conditions upon progressive additions of HCl to the DNA solution (Fig. 1). The spectrum acquired at pH 5.5 was more intense than the ones recorded at basic conditions and it showed a positive signal at 287 nm and a negative one at 264 nm. This optical fingerprint corresponds to the one expected for an iM. The folding of EGFR-37 into an iM in acidic conditions was further supported by the UV–Vis thermal difference spectrum (TDS) (Fig. S1). Indeed, also this spectroscopic profile varies according to the specific DNA secondary structures [24]. The TDS spectrum of EGFR-37 at pH 5.5 showed a negative signal at 295 nm and a positive one at 240 nm, in line with those reported for iM folded DNA.
CD titration of EGFR-37 with HCl. a CD spectra of 4 µM EGFR-37 in 10 mM Na-cacodylate at 25 °C from pH 8.5 (black solid line) to pH 4.0 (black dashed line) upon addition of HCl. b CD signals recorded at 287 nm (black dots) plotted as a function of pH and fitted according to Eq. 1 (red solid line)
Additionally, native polyacrylamide gel electrophoresis performed in TAE pH 5.5 indicated that EGFR-37 samples in the 4–200 µM concentration range, constantly run as a single band with electrophoretic mobility slightly greater than a 22 residues DNA strand unable to fold into secondary structures (22BM) (Fig. S2). These evidences converged in supporting the folding of EGFR-37 into an intra-molecular iM structure in acidic conditions.
Additionally, the CD spectra acquired at variable pH showed an iso-dichroic point at 277 nm, in line with the presence of only two optical species during the protonation-induced folding process. This was further confirmed by the analysis of the optical components performed on the U, S, and V matrices derived by SVD (Table S1).
The lack of intermediate species indicated that under our experimental conditions, the DNA folding process was highly cooperative. On these bases, the CD data acquired at 287 nm (maximum signal intensity for an iM) were fitted according to Eq. (1), based on the Hill formalism. From this analysis, we derived the pH of middle transition (pHT) and the Hill coefficient (n) which resulted in 6.2 ± 0.1 and 3.8 ± 0.1, respectively (Table 2). The pHT at 298.15 K was significantly higher than the pKa of cytosine N3 which is ≃ 4.5 (referred to 298.15 K) [12]. This shift was expected since, beyond the protonation of cytosines, other energetic contributions (base pairing, hydrogen bonding, etc.) play part in determining the ΔGθ(298.5 K) of the iM folding. As it concerns the n coefficient, its value indicated that the cooperativity of the folding process was markedly positive, a result that fits with the above-described absence of long-living intermediated species. In these conditions, the Hill coefficient can be considered to approach the number of binding sites, that in our model is related to the number of recruited protons responsible for the structural rearrangement [25]. Our analyses cannot discriminate among interactions of protons at distinct DNA domains. However, in the pH range where the structural transition occurred, we can safely rule out the interactions of protons with other DNA acidic functional groups but the N3 of cytosine [12]. Thus, n represented a reliable estimation of the number of protons directly involved in CC + base pairs formation.
To better define the EGFR-37 folding model, we analyzed also the temperature dependency of the iM of EGFR-37 at pH 5.5 by following the heating and cooling processes by CD (Fig. S3). The processes resulted to be fully reversible, and the analysis of the U, S, and V matrices derived by SVD confirmed the presence of only two significant optical components in the solution (Table S1). Thus, it was possible to fit the data corresponding to the CD signal at 287 nm according to Eq. 2 based on van't Hoff's formalism (Fig. 2). Through this analysis, we derived the melting temperature (43.9 ± 0.1) °C and the associated ΔHθ (− 238.1 ± 4.6) kJ mol−1 reported in Table 2.
EGFR-37 heating and cooling in 10 mM Na-cacodylate, pH 5.5 performed at ± 20 K h−1 heating–cooling rate. a CD signals recorded at 287 nm during heating (black dot) and cooling (gray dots) of 4 µM EGFR-37 as a function of pH and fitted according to Eq. 2 (red solid line). b DSC curves acquired during the heating (black solid line) and cooling (gray solid line) scans of 200 µM EGFR-37 and fitting according to Eq. 3 (red solid line)
Due to the monomeric feature of the iM assumed by EGFR-37, it was possible to integrate these data by DSC experiments where we applied the same heating–cooling rate (± 20 °C h−1) to 200 µM DNA samples. The process was confirmed to be fully reversible under these conditions. Consistently, the melting temperatures as well as the enthalpic contributions derived by CD and DSC well overlapped (Table 2).
This result further supported that the EGFR-37 folding process was concentration-independent as expected for an intra-molecular structure and occurred with no formation of other metastable states. Therefore, we used the ensemble of CD (HCl-titration, thermal heating, and cooling) and DSC experiments to run a global fitting according to Eqs. 6–7, respectively (Fig. S4). In the analysis n, ΔGθ(298.15 K), ΔHθ were kept as shared fitting parameters. The derived parameters well compared to those obtained by the individual analyses of every single dataset. This approach allowed us to further test the model mechanism of folding and derive all the associated thermodynamic parameters. The molar fraction profiles of the folded and unfolded species through pH and temperature are reported in Fig. 3.
Noteworthy, the in silico prediction indicated that folded iM EGFR-37 should be able to form up to 6 CC+ base pairs. However, the derived ΔHθ ((−250 ± 1) kJ mol−1, Table 2) was significantly lower than those previously reported for iM models with 6 CC+ base pairs [9], while it was in line with the Hill coefficient that pointed to the recruitment of only 4 protons for the iM folding.
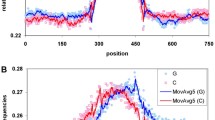
To clarify these discrepancies, we mapped the cytosines paired within the iM structure by performing S1 cleavage footprinting. This enzyme is a nuclease that cuts only single-stranded residues and well performs under acidic conditions. Thus, it can be efficiently exploited to identify unpaired nucleotides within a folded iM structure. The enzymatic cleavage pattern of EGFR-37 at pH 5.5 is reported in Fig. 4. It showed strong cleavage sites at T4, T7-C8, and G19–C20 along with the neighboring residues T18 and C21 that were cleaved although with lower efficiency. This pattern allowed us to locate all these residues in the loops of the iM. Additionally, most of the cytosines at 3’ and 5’ terminals (C1, C2, C25, and C27) were significantly cleaved thus ruling out their involvement in stable CC+ base pairings. Therefore, it resulted that the four runs of cytosines involved in the iM core should be C5–C6, C11–C12–C13, C15–C16–C17, and C23–C24. However, it must be reminded that iM structures have a conserved folding topology corresponding to two parallel duplexes intercalated in antiparallel orientation, from which it derives that cytosines of the first and second runs must pair with those of the third and fourth runs, respectively. Consequently, C5–C6 must pair with two cytosines among C15–C16–C17 and, similarly, C23–C24 with two among C11–C12–C13. As a result, within the iM core, only 4 CC+ base pairs can be present.
iM folding of EGFR-37. a S1 footprinting of 100 ng/µL EGFR-37-FAM in 10 mM Na-cacodylate, 4.5 mM ZnSO4, pH 5.5.. C refers to the untreated EGFR-37-FAM, PM to the purine marker. In the S1 lanes, the time of incubation of EGFR-37-FAM with the enzyme is reported. b Schematic representation of the iM folding of EGFR-37
Notably, T10 and T22 were fully protected from S1 cleavage. This suggested their involvement in the formation of a non-canonical TT base pair, a frequently observed capping module for iM cores [7, 26, 27]. In our sequence, this element would extend the CC+ pairing occurring between the second and fourth cytosine runs. This structural feature implied that C11 and C12 pair to C23 and C24, respectively. According to this folding model, the second loop should comprise C13, G14, and C15, although it was not efficiently cleaved by the enzyme.
However, C3 was protected from S1 whereas C2 and T4 were cleaved and, based on this evidence, we attributed C3–G14 to the formation of an additional GC base pair.
To verify the proposed base-pairing bonding network we performed 1D 1H NMR (Fig. 5). The signals at 15.54 ppm and 14.84 ppm were safely attributed to H3 of cytosines in CC+ pairings although the overlapping contributions prevented us from clearly deriving the effective number of CC+. As far as it concerns the presence of additional base pairings, the imino region of the spectrum showed two well-solved signals at 11.18 and 11.10 ppm deriving from the H3 of thymines involved in the formation of a TT base pair. Conversely, we did not detect any signal belonging to H1 of the guanine involved in GC interaction. It is worth to underly that distinctly from the T10–T22 cupping element, the G14–C3 base pair was not expected to form on the top of a CC+. As a consequence, this base pair is highly exposed to the solvent. Such a condition favors a fast exchange rate of the G14 imino proton with the water thus preventing its detection by NMR.
Mutated sequences of EGFR-37
To validate the presence of the GC base pair in the iM of EGFR-37, we designed a sequence, EGFR-37MUT, in which G14 was substituted with an adenine, and we performed the S1 footprinting on the mutated sequence (Fig. 6).
As can be observed in Fig. 6, cleavage sites were detected at C3 and A14 of EGFR-37MUT, thus confirming they were more accessible to the S1 enzymatic cleavage.
Interestingly, HCl-titration, heating, and cooling experiments followed by CD indicated that the EGFR-37MUT folds into a comparable iM to EGFR-37. Indeed, the recorded CD signals of the two folded sequences almost overlapped as well as the derived Tm and pHT (Fig. S5). Also, the TDS of EGFR-37MUT showed the same shape and intensity as the one acquired for EGFR-37 with a positive pick at 240 nm and a negative one at 290 nm (Fig. S1). The only difference rested in a shoulder at 260 nm which slightly decreased in intensity for the mutated sequence and notably, this is the wavelength range in which a GC base pair formation mainly contributes to the hyperchromic effect [24]. Again, the analysis performed on the U, S, and V matrices derived by SVD, sustained the presence of only two significant optical components in all the CD experiments (Table S2).
However, distinctly from the behavior above reported with EGFR-37, DSC experiments performed at 200 µM EGFR-37MUT, showed no overlapping heating and cooling curves. In particular, multiple transitions were recorded along heating scans (Fig. 7). By increasing the buffer concentration (50 mM Na-cacodylate, pH 5.5), the distribution of the species was further shifted toward those with higher temperature transitions. This profile was conserved even by lowering the scanning rate to 5 K h−1 (Fig. S6).
EGFR-37MUT heating and cooling in 10 mM Na-cacodylate, pH 5.5 performed at ± 20 K h−1 heating–cooling rate. a CD recorded at 287 nm during heating (black dot) and cooling (gray dots) of 4 µM EGFR-37MUT and fitted according to Eq. 2 (red solid line). b DSC curves acquired during the heating (black solid line) and cooling (black dashed line) scans of 200 µM EGFR-37MUT
In line with this evidence, when EGFR-37MUT samples were resolved by gel electrophoresis, the formation of a slow migrating band was observed by increasing the oligonucleotide concentration (Fig. S2). This allowed us to associate the higher temperature transitions to the slow formation of inter-molecular structures occurring at high concentrations of EGFR-37MUT. This represented an interesting difference between EGFR-37 and EGFR-37MUT, which can only be attributed to the substitution of G14 with an adenine.
This behavior of the mutated sequence prevented us to derive the thermodynamic parameters from DSC data, thus, we limited the global fitting to the CD experimental data sets which were performed at 4 µM DNA concentration, where only the intra-molecular iM was detectable (Fig. S7). Under this condition, the molar fraction distribution of the folded and unfolded species and, consistently, the thermodynamic parameters of the iM of EGFR-37 and EGFR-37MUT well overlapped (Table 3, Fig. S8). This result confirmed that the G14–C3 pairing in EGFR-37 is a weak interaction.
In addition, since the enthalpy of denaturation is model-independent, we calculated it from the data acquired along the cooling processes as area under the DSC curves. The average of three measurements resulted in ΔHθ (182 ± 1) kJ mol−1 and (212 ± 1) kJ mol−1 at 10 and 50 mM Na-cacodylate, respectively. These values appear lower when compared to the ΔHθ derived from the global fitting analysis of the spectroscopic data (Table 3). This reduced system-surrounding heat exchange further corroborates the hypothesis that, at 200 µM DNA concentration, the system reflects an out-of-equilibrium process involving intra- and inter-molecular species in which the intra-molecular one is kinetically favored.
Increasing ionic strength induces the formation of intermolecular species
It was surprising to find that the two tested sequences folded into comparable intra-molecular iM structures from a thermodynamic point of view, but with a divergent attitude toward multimerization. Thus, whether the GC interaction seemed to play a negligible effect on the iM features, still it appeared that the substitution of G14 with an adenine largely drove multimeric species formation. Since the pairing of multiple strands can be promoted by increasing the ionic strength, we decided to test the behavior of EGFR-37 in 50 mM Na-cacodylate, focusing on the heating and cooling profiles. At this ionic strength the CD and DSC experiments, recorded at different DNA concentrations, provided different outputs (Fig. 8).
EGFR-37 heating and cooling in 50 mM Na-cacodylate pH 5.5 performed at ± 20 K h−1 heating–cooling rate. a CD signals recorded at 287 nm during heating (black dot) and cooling (gray dots) of 4 µM EGFR-37 as a function of pH and fitted according to Eq. (2) (red solid line). b DSC curves acquired during the heating (black solid line) and cooling (gray solid line) scans of 200 µM EGFR-37
CD data, acquired at 4 µM DNA concentration, showed a single and fully reversible transition with a melting temperature of (44.1 ± 0.1) °C, a profile that was not significantly different from the one acquired at lower ionic strength. Conversely, when we increased the DNA concertation up to 200 µM, as required by DSC, the reversibility of the process was lost and additional transitions occurring at higher temperatures appeared along the heating scan.
During the cooling scan, a single peak was always recorded with a maximum at 44.5 °C. This value was not significantly different from the one recorded in 10 mM Na-cacodylate, where only the intra-molecular iM occurred. Remarkably, it nicely matched the lower temperature transition in the heating scan. Thus, we attributed it to the intra-molecular iM structure of EGFR-37. As a consequence, the higher temperature transitions can be associated with inter-molecular foldings which resulted to be significantly stabilized by the higher ionic strength also on the wild-type sequence.
Conclusions
Here, we performed an integrated thermodynamic characterization of an iM structure that was expected to form at the site located 37 nucleotides upstream of the transcription starting site of the EGFR oncogene. The sequence was selected to potentially form 6 CC+ base pairs which should grant significant stability even under conditions approaching the physiological ones. However, our results are interestingly in contrast with the in silico prediction. The thermodynamic parameters we derived through a global analysis of calorimetric and spectroscopic data indicated that the iM core is held by 4 CC+, a conclusion supported also by footprinting experiments. This surprising discrepancy prompts us to reconsider that having 4 runs of 3 cytosines is not the only requirement to fold into an iM with 6 CC+ base pairs.
Our data indicated that the maintenance of a significantly stable iM is supported by the presence of additional structural elements, in detail a TT and a GC base pair. The presence of a TT base pair that works as a cupping element was not unexpected since several reported high-resolution structures included it at the 3’ or 5’ terminal of the iM core [7, 26, 27]. The same role could be played by the GC which, in principle, should provide a more prominent effect. However, in our sequence, this base pair is spaced out from the iM core thus preventing efficient stacking. This was in line with the fast exchange in the NMR and confirmed by the overlapping thermodynamic profiles of the EGFR-37 and EGFR-37MUT as well. This addresses the C3–G14 as a weak interaction. Remarkably, despite the lack of any iM stabilization roles by this motif, it turned out that the single residue mutation of the guanine at position 14 with an adenine greatly stabilizes inter-molecular species. It would be possible to argue that this is a direct consequence of the presence of adenine. However, we proved that an increment of the ionic strength can induce the same process also on the wild-type sequence while it does not significantly affect the stability of the intra-molecular iM.
These data indicate that a lot of information concerning the structural properties of iM is lacking. Here, we showed how the global analysis of calorimetric and spectroscopic data is a highly valuable strategy to obtain thermodynamic and structural information on iM foldings. This “low-resolution” approach is flexible and reliable and can thus be easily extended to a wide panel of C-rich sequences. The output would help in better addressing the sequence requirements behind iM formation paving the way toward a more precise prediction of iM forming sites and, consequently, in their screening as a potential pharmacological target or in the set up of solid pH-sensible nanodevices.
References
Rigo R, Palumbo M, Sissi C. G-quadruplexes in human promoters: A challenge for therapeutic applications. Biochimica et Biophysica Acta (BBA)—General Subjects. 2017;1861:1399–413. https://doi.org/10.1016/j.bbagen.2016.12.024.
Abou Assi H, Garavís M, González C, Damha MJ. i-Motif DNA: structural features and significance to cell biology. Nucleic Acids Res. 2018;46:8038–56. https://doi.org/10.1093/nar/gky735.
Huppert JL. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 2005;33:2908–16. https://doi.org/10.1093/nar/gki609.
Belmonte-Reche E, Morales JC. G4-iM Grinder: when size and frequency matter. G-Quadruplex, i-Motif and higher order structure search and analysis tool. NAR Genomics and Bioinformatics. 2020;2:lqz005. https://doi.org/10.1093/nargab/lqz005.
Siddiqui-Jain A, Grand CL, Bearss DJ, Hurley LH. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc Natl Acad Sci. 2002;99:11593–8. https://doi.org/10.1073/pnas.182256799.
Kendrick S, Kang H-J, Alam MP, Madathil MM, Agrawal P, Gokhale V, et al. The dynamic character of the BCL2 promoter i-Motif provides a mechanism for modulation of gene expression by compounds that bind selectively to the alternative DNA Hairpin structure. J Am Chem Soc. 2014;136:4161–71. https://doi.org/10.1021/ja410934b.
Gehring K, Leroy J-L, Guéron M. A tetrameric DNA structure with protonated cytosine-cytosine base pairs. Nature. 1993;363:561–5. https://doi.org/10.1038/363561a0.
Sen D, Gilbert W. Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature. 1988;334:364–6. https://doi.org/10.1038/334364a0.
Mergny J-L, Lacroix L, Han X, Leroy J-L, Helene C. Intramolecular folding of pyrimidine oligodeoxynucleotides into an i-DNA Motif. J Am Chem Soc. 1995;117:8887–98. https://doi.org/10.1021/ja00140a001.
Amato J, D’Aria F, Marzano S, Iaccarino N, Randazzo A, Giancola C, et al. On the thermodynamics of folding of an i-motif DNA in solution under favorable conditions. Phys Chem Chem Phys. 2021;23:15030–7. https://doi.org/10.1039/D1CP01779A.
Berger I, Egli M, Rich A. Inter-strand C-H...O hydrogen bonds stabilizing four-stranded intercalated molecules: stereoelectronic effects of O4’ in cytosine-rich DNA. Proceedings of the National Academy of Sciences. 1996;93:12116–21. https://doi.org/10.1073/pnas.93.22.12116.
Ts’o POP. Bases, nucleosides, and nucleotides. In: Ts’o POP, editor. Basic Principles in Nucleic Acid Chemistry. 1974. p. 453–584.
Dzatko S, Krafcikova M, Hänsel-Hertsch R, Fessl T, Fiala R, Loja T, et al. Evaluation of the Stability of DNA i-Motifs in the Nuclei of Living Mammalian Cells. Angew Chem Int Ed. 2018;57:2165–9. https://doi.org/10.1002/anie.201712284.
Zeraati M, Langley DB, Schofield P, Moye AL, Rouet R, Hughes WE, et al. I-motif DNA structures are formed in the nuclei of human cells. Nature Chem. 2018;10:631–7. https://doi.org/10.1038/s41557-018-0046-3.
Shirmanova MV, Druzhkova IN, Lukina MM, Matlashov ME, Belousov VV, Snopova LB, et al. Intracellular pH imaging in cancer cells in vitro and tumors in vivo using the new genetically encoded sensor SypHer2. Biochimica et Biophysica Acta (BBA) - General Subjects. 2015;1850:1905–11. https://doi.org/10.1016/j.bbagen.2015.05.001.
Park HS, Jang MH, Kim EJ, Kim HJ, Lee HJ, Kim YJ, et al. High EGFR gene copy number predicts poor outcome in triple-negative breast cancer. Mod Pathol. 2014;27:1212–22. https://doi.org/10.1038/modpathol.2013.251.
Sharma SV, Bell DW, Settleman J, Haber DA. Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer. 2007;7:169–81. https://doi.org/10.1038/nrc2088.
Wykosky J, Fenton T, Furnari F, Cavenee WK. Therapeutic targeting of epidermal growth factor receptor in human cancer: successes and limitations. Chin J Cancer. 2011;30:5–12. https://doi.org/10.5732/cjc.010.10542.
Cooper AJ, Sequist LV, Lin JJ. Third-generation EGFR and ALK inhibitors: mechanisms of resistance and management. Nat Rev Clin Oncol. 2022;1–16. https://doi.org/10.1038/s41571-022-00639-9.
Greco ML, Kotar A, Rigo R, Cristofari C, Plavec J, Sissi C. Coexistence of two main folded G-quadruplexes within a single G-rich domain in the EGFR promoter. Nucleic Acids Res. 2017;45:10132–42. https://doi.org/10.1093/nar/gkx678.
Cristofari C, Rigo R, Greco ML, Ghezzo M, Sissi C. pH-driven conformational switch between non-canonical DNA structures in a C-rich domain of EGFR promoter. Sci Rep. 2019;9:1210. https://doi.org/10.1038/s41598-018-37968-8.
Recagni M, Greco ML, Milelli A, Minarini A, Zaffaroni N, Folini M, et al. Distinct biological responses of metastatic castration resistant prostate cancer cells upon exposure to G-quadruplex interacting naphthalenediimide derivatives. Eur J Med Chem. 2019;177:401–13. https://doi.org/10.1016/j.ejmech.2019.05.066.
Gray RD, Chaires JB. Kinetics and mechanism of K+- and Na+-induced folding of models of human telomeric DNA into G-quadruplex structures. Nucleic Acids Res. 2008;36:4191–203. https://doi.org/10.1093/nar/gkn379.
Mergny J-L, Li J, Lacroix L, Amrane S, Chaires JB. Thermal difference spectra: a specific signature for nucleic acid structures. Nucleic Acids Res. 2005;33:e138. https://doi.org/10.1093/nar/gni134.
Weiss JN. The Hill equation revisited: uses and misuses. Faseb J. 1997;11–835. https://doi.org/10.1096/fasebj.11.11.9285481
Phan AT, Guéron M, Leroy J-L. The solution structure and internal motions of a fragment of the cytidine-rich strand of the human telomere. J Mol Biol. 2000;299:123–44. https://doi.org/10.1006/jmbi.2000.3613.
Han X, Leroy J-L, Guéron M. An intramolecular i-motif: the solution structure and base-pair opening kinetics of d(5mCCT3CCT3ACCT3CC). J Mol Biol. 1998;278:949–65. https://doi.org/10.1006/jmbi.1998.1740.
Funding
The research leading to these results has received funding from AIRC under IG 2021-1D. 26474 project—P.I. Sissi Claudia Sissi and European Union-Next GenerationEU (PNRR M4C2-Investimento 1.4-CN00000041). The Ph.D. fellowship of MG was founded by Cariparo. Open access funding provided by Università degli Studi di Padova within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
MG and CS contributed to conception and design of the study. MG performed all experimental settings and data analyses, CS contributed to sequences selection and design and supervised experimental results and data analyses, MG wrote the first draft of the manuscript, CS cured manuscript writing, review and editing. All authors contributed to manuscript revision, read, and approved the submitted version.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghezzo, M., Sissi, C. Structural characterization of a cytosine-rich potential quadruplex forming sequence in the EGFR promoter. J Therm Anal Calorim 148, 5375–5385 (2023). https://doi.org/10.1007/s10973-023-12060-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10973-023-12060-0