Abstract
Linear model trees are regression trees that incorporate linear models in the leaf nodes. This preserves the intuitive interpretation of decision trees and at the same time enables them to better capture linear relationships, which is hard for standard decision trees. But most existing methods for fitting linear model trees are time consuming and therefore not scalable to large data sets. In addition, they are more prone to overfitting and extrapolation issues than standard regression trees. In this paper we introduce PILOT, a new algorithm for linear model trees that is fast, regularized, stable and interpretable. PILOT trains in a greedy fashion like classic regression trees, but incorporates an L2 boosting approach and a model selection rule for fitting linear models in the nodes. The abbreviation PILOT stands for PIecewise Linear Organic Tree, where ‘organic’ refers to the fact that no pruning is carried out. PILOT has the same low time and space complexity as CART without its pruning. An empirical study indicates that PILOT tends to outperform standard decision trees and other linear model trees on a variety of data sets. Moreover, we prove its consistency in an additive model setting under weak assumptions. When the data is generated by a linear model, the convergence rate is polynomial.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Despite their long history, decision trees such as CART (Breiman et al., 1984) and C4.5 (Quinlan, 1993) remain popular machine learning tools. Decision trees can be trained quickly with few parameters to tune, and the final model can be easily interpreted and visualized, which is an appealing advantage in practice. As a result, these methods are widely applied in a variety of disciplines including engineering (Shamshirband et al., 2020), bioinformatics (Shaikhina et al., 2019), agriculture (Tariq et al., 2023), and business analysis (Aydin et al., 2022; Golbayani et al., 2020). In addition to their standalone use, decision trees have seen widespread adoption in ensemble methods, often as the best available “weak learner”. Prime examples are random forests (Breiman, 2001) and gradient boosting methods such as XGBoost (Chen & Guestrin, 2016) and LightGBM (Ke et al., 2017). In this work we assume that the target variable is continuous, so we focus on regression trees rather than on classification.
A limitation of classical regression trees is that their piecewise constant nature makes them ill-suited to capture continuous relationships. They require many splits in order to approximate linear functions, which is undesirable. There are two main approaches to overcome this issue. The first is to use ensembles of decision trees such as random forests or gradient boosted trees. These ensembles smooth the prediction and can therefore model a more continuous relation between predictors and response. A drawback of these ensembles is the loss of interpretability. Combining multiple regression trees no longer allows for a simple visualization of the model, or for an explainable stepwise path from predictors to prediction. The second approach to capture continuous relationships is to use model trees. Model trees have a tree-like structure for the partition of the space, but allow for non-constant fits in the leaf nodes of the tree. Model trees thus retain the intuitive interpretability of classical regression trees while being more flexible. Arguably the most intuitive and common model tree is the linear model tree, which allows for linear models in the leaf nodes.
There are several algorithms for fitting linear model trees. The first linear model tree algorithm to be proposed, which we will abbreviate as FRIED (Friedman, 1979), uses univariate piecewise linear fits to replace the piecewise constant fits in CART. Once a model is fit on a node, its residuals are passed on to its child nodes for further fitting. The final prediction is given by the sum of the linear models along the path. Therefore, the final model can be written as a binary regression tree with linear models in the leaf nodes. The coefficients of these linear models in the leaf nodes are given by the sum of the coefficients of all linear models along the path from the root to the leaf node. Our experiments suggest that this method may suffer from overfitting and extrapolation issues. The FRIED algorithm received much less attention than its more involved successor MARS (Friedman, 1991).
The M5 algorithm (Quinlan, 1992) is by far the most popular linear model tree, and is still commonly used today (Shamshirband et al., 2020; da Silva et al., 2021; Khaledian & Miller, 2020; Pham et al., 2022; Fernández-Delgado et al., 2019). It starts by fitting CART. Once this tree has been built, linear regressions are introduced at the leaf nodes. Pruning and smoothing are then applied to reduce its generalization error. One potential objection against this algorithm is that the tree structure is built completely oblivious of the fact that linear models will be used in the leaf nodes.
The GUIDE algorithm (Loh, 2002) fits multiple linear models to numerical predictors in each node and then applies a \(\chi ^2\) test comparing positive and negative residuals to decide on which predictor to split. The algorithm can also be applied to detect interactions between predictors. However, no theoretical guarantee is provided to justify the splitting procedure. There are also algorithms that use ideas from clustering in their splitting rule. For example, SECRET (Dobra & Gehrke, 2002) uses the EM algorithm to fit two Gaussian clusters to the data, and locally transforms the regression problem into a classification problem based on the closeness to these clusters. Experimental results do not favor this method over GUIDE, and its computational cost is high.
The SMOTI method (Malerba et al., 2004) uses two types of nodes: regression nodes and splitting nodes. In each leaf, the final model is the multiple regression fit to its ‘active’ variables. This is achieved by ‘regressing out’ the variable of a regression node from both the response and the other variables. The resulting fit is quite attractive, but the algorithm has a complexity of \(\mathcal {O}(n^2p^3)\), where n denotes the number of cases and p the number of predictors. This can be prohibitive for large data sets, and makes it less suitable for an extension to random forests or gradient boosting.
Most of the above algorithms need a pruning procedure to ensure their generalization ability, which is typically time consuming. An exception is LLRT (Vogel et al., 2007) which uses stepwise regression and evaluates the models in each node via k-fold cross validation. To alleviate the computational cost, the algorithm uses quantiles of the predictors as the potential splitting points. It also maintains the data matrices for fitting linear models on both child nodes so that they can be updated. Unfortunately, the time complexity of LLRT is quite high at \(\mathcal {O}(knp^3 + n p^5)\) where k is the depth of the tree.
It is also interesting to compare ensembles of linear model trees with ensembles of classical decision trees. Recently, certain piecewise linear trees were incorporated into gradient boosting (Shi et al., 2019). For each tree, they used additive fitting (Friedman, 1979) or half additive fitting where the past prediction was multiplied by a factor. To control the complexity of the tree, they set a tuning parameter for the maximal number of predictors that can be used along each path. Empirical results suggest that this procedure outperformed classical decision trees in XGBoost (Chen & Guestrin, 2016) and in LightGBM (Ke et al., 2017) on a variety of data sets, and required fewer training iterations. This points to a potential strength of linear model trees for ensembles.
To conclude, the main issue with existing linear model trees is their high computational cost. Methods that apply multiple regression fits to leaf and/or internal nodes introduce a factor \(p^2\) or \(p^3\) in their time complexity. And the methods that use simple linear fits, such as FRIED, still require pruning which is also costly on large data sets. In addition, these methods can have large extrapolation errors (Loh et al., 2007). Finally, to the best of our knowledge there is no theoretical support for linear model trees in the literature.
In response to this challenging combination of issues we propose a novel linear model tree algorithm. Its acronym PILOT stands for PIecewise Linear Organic Tree, where ‘organic’ refers to the fact that no pruning is carried out. The main features of PILOT are:
-
Speed: It has the same low time complexity as CART without its pruning.
-
Regularized: In each node, a model selection procedure is applied to the potential linear models. This requires no extra computational complexity.
-
Explainable: Thanks to the simplicity of linear models in the leaf nodes, the final tree remains highly interpretable. Also a measure of feature importance can be computed.
-
Stable extrapolation: Two truncation procedures are applied to avoid extreme fits on the training data and large extrapolation errors on the test data, which are common issues in linear model trees.
-
Theoretically supported: PILOT has proven consistency in additive models. When the data is generated by a linear model, PILOT attains a polynomial convergence rate, in contrast with CART.
The paper is organized as follows. Section 2 describes the PILOT algorithm and discusses its properties. Section 3 presents the two main theoretical results. First, the consistency for general additive models is discussed and proven. Second, when the true underlying function is indeed linear an improved rate of convergence is demonstrated. We refer to the Appendix for proofs of the theorems, propositions and lemmas. In addition to these, we derive the time and space complexity of PILOT. Empirical evaluations are provided in Sect. 4, where PILOT is compared with several alternatives on a variety of benchmark data sets. It outperformed other tree-based methods on data sets where linear models are known to fit well, and outperformed other linear model trees on data sets where CART typically performs well. Section 5 concludes.
2 Methodology
In this section we describe the workings of the PILOT learning algorithm. We begin by explaining how its tree is built and motivate the choices made, and then derive its computational cost.
We will denote the \(n \times p\) design matrix as \(\varvec{X}=({X}_1, \dots ,{X}_n)^\top\) and the \(n \times 1\) response vector as \(Y=(y_1,\dots ,y_n)^\top\). We consider the standard regression model \(y = f(X) + \epsilon\), where \(f: \mathbb R^p\rightarrow \mathbb R\) is the unknown regression function and \(\epsilon\) has mean zero.
As a side remark, the linear models and trees we consider here are equivariant to adding a constant to the response, that is, the predictive models would keep the same fitted parameters except for the intercept terms. Therefore, in the presentation we will assume that the responses are centered around zero, that is, \(Y_{min} = -Y_{max}\) without loss of generality.
2.1 Main structure of PILOT
A typical regression tree has four ingredients: a construction rule, the evaluation of the models/splits, a stopping rule, and the prediction. Most regression tree algorithms are built greedily from top to bottom. That is, they split the original space along a predictor and repeat the procedure on the subsets. This approach has some major advantages. The first is its speed. The second is that it starts by investigating the data from a global point of view, which is helpful when detecting linear relationships.
Algorithm 1 presents a high-level description of PILOT. It starts by sorting each predictor and storing its ranks. At each node, PILOT then selects a predictor and a univariate piecewise linear model via a selection procedure that we will describe in Sects. 2.2 and 2.3. Then it fits the model and passes the residuals on to the child nodes for further fitting. This recursion is applied until the algorithm stops. There are three stopping triggers. In particular, the recursion in a node stops when:
-
the depth reaches the preset maximal depth of the tree \(K_{max}\);
-
the number of cases in the node is below a threshold value \(n_{\text {fit}}\);
-
none of the candidate models do substantially better than a constant prediction, as we will describe in Sect. 2.3.
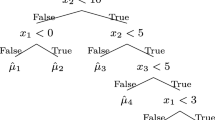
When all nodes have reached a stopping trigger, the algorithm is done. The final prediction is given by aggregating the predictions of the linear models from the root to the leaves, as in the example in Fig. 1.
Note that PILOT retains good interpretability. The tree structure visualizes the decision process, while the simple linear models in each step reveal which predictor was used and how the prediction changes with respect to a small change in this predictor. For example, in Fig. 1, the result from the node X3>0.4 can be interpreted as: given \(X1<2.3\) and the previous prediction \(0.3*X1+0.2\), the best additional predictor is X3 where the split of the space happens. Furthermore, if \(X3>0.4\), \(0.9*X3+0.1\) is added to the prediction and \(0.5*X3-0.4\) otherwise, meaning that there was a correlation between X3 and the residuals after the first prediction. Since the final prediction in each leaf node is linear, it is easy to interpret. Moreover, we will define a measure of feature importance similar to that of CART, based on the variance reduction in each node. This is because PILOT selects only one predictor in each node, which makes the gain fully dependent on it. This differs from methods such as M5, GUIDE, and LLRT that use multiple predictors in each node, making it harder to fairly distribute the gain over several predictors in order to derive an overall measure of each predictor’s importance. We refer to Sect. 4.5 for a more detailed discussion.
2.2 Models used in the nodes
As stated in the high-level summary in Algorithm 1, at each node PILOT selects a fit from a number of linear and piecewise linear models. In particular, PILOT considers the following regression models on each predictor, shown in Fig. 2:
-
pcon: A Piecewise CONstant fit, as in CART.
-
lin: Simple LINear regression.
-
blin: A Broken LINear fit: a continuous function consisting of two linear pieces.
-
plin: A two-Piece LINear fit that need not be continuous.
-
con: A CONstant fit. We stop the recursion in a node after a con model is fitted.
These models extend the piecewise constant pcon fits of CART to linear fits. In particular, plin is a direct extension of pcon while blin can be regarded as a regularized and smoothed version of plin. For variables that are categorical or have but a few unique values, only pcon and con are used.
To guard against unstable fitting, the lin and blin models are only considered when the number of unique values in the predictor is at least 5. Similarly, the plin model is only considered if both potential child nodes have at least 5 unique values of the predictor.
PILOT reduces to CART (without its pruning) if only pcon models are selected, since the least squares constant fit in a child node equals its average response.
Note that a node will not be split when lin is selected. Therefore, when a lin fit is carried out in a node, PILOT does not increase its reported depth. This affects the reported depth of the final tree, which is a tuning parameter of the method.
It is possible that a consecutive series of lin fits is made in the same node, and this did happen in our empirical studies. In a node where this occurs, PILOT is in fact executing \(L^2\) boosting (Bühlmann, 2006), which fits multiple regression using repeated simple linear regressions. It has been shown that \(L^2\) boosting is consistent for high dimensional linear regression and produces results comparable to the Lasso. It was also shown that its convergence rate can be relatively fast under certain assumptions (Freund et al., 2017). PILOT does not increment the depth value of the node for lin models, to avoid interrupting this boosting procedure.
2.3 Model selection rule
In each node, PILOT employs a selection procedure to choose between the five model types con, lin, pcon, blin and plin. It would be inappropriate to select the model based on the largest reduction in the residual sum of squares (\(\textrm{RSS}\)), since this would always choose plin, as that model generalizes all the others. Therefore, we need some kind of regularization to select a simpler model when the extra \(\textrm{RSS}\) gain of going all the way to plin is not substantial enough.
After many experiments, the following regularization scheme was adopted. In each node, PILOT chooses the combination of a predictor and a regression model that leads to the lowest BIC value
which is a function of the residual sum of squares \(\textrm{RSS}\), the number of cases n in the node, and the degrees of freedom v of the model. With this selection rule PILOT mitigates overfitting locally, and applying it throughout the tree leads to a global regularization effect.
It remains to be addressed what the value of v in (1) should be for each of the five models. The degrees of freedom are determined by aggregating the number of model parameters excluding the splits, and the degrees of freedom of a split. The model types con, lin, pcon, blin and plin contain 1, 2, 2, 3, and 4 coefficients apart from any split points. There is empirical and theoretical evidence in support of using 3 degrees of freedom for a discontinuity point (Hall et al., 2017). We follow this recommendation for pcon and plin, which each receive 3 additional degrees of freedom. Also blin contains a split point, but here the fitted function is continuous. To reflect this intermediate complexity we add 2 degrees of freedom to blin. We empirically tested 1 to 3 additional degrees of freedom for blin, and the performance was generally not very sensitive to this choice. Of course, these values could be considered hyperparameters and could thus be tuned by cross-validation. This would come at a substantial increase in computation time though, so we provide default parameters instead. In conclusion, we end up with v = 1, 2, 5, 5, and 7 for the model types con, lin, pcon, blin and plin.
The BIC in (1) is one of several model selection criteria that we could have used. We also tried other measures, such as the Akaike information criterion (AIC) and the adjusted AIC. It turned out that the AIC tended to choose plin too often, which reduced the regularization effect. The adjusted AIC required a pruning procedure to perform comparably to the BIC criterion. As pruning comes at a substantial additional computational cost, we decided in favor of the BIC, which performed well.
Alternatively, one option is to compute the degrees of freedom for the aggregated model in each node, and then compute the adjusted AIC to decide when to stop (Bühlmann, 2006). But in this approach hat matrices have to be maintained for each path, which requires more computation time and memory space. Moreover, as the number of cases in the nodes changes, the evaluations of the hat matrices become more complicated. For these reasons, we preferred to stay with the selection rule (1) in each node.
Algorithm 2 shows the pseudocode of our model selection procedure in a given node. We iterate through each predictor separately. If the predictor is numerical, we find the univariate model with lowest BIC among the 5 candidate models. For con and lin this evaluation is immediate. For the other three models, we need to consider all possible split points. We consider these in increasing order (recall that the ordering of each predictor was already obtained at the start of the algorithm), which allows the Gram and moment matrices for the linear models to be maintained and updated efficiently in each step. On the other hand, if the predictor is categorical, we follow the approach of CART as in Section 9.2.4 of Hastie et al. (2009). That is, we first compute the mean \(m_c\) of the response of each level c and then sort the cases according to \(m_c\). We then fit the model pcon in the usual way.
Once the best model type has been determined for each predictor separately, Algorithm 2 returns the combination of predictor and model with the lowest BIC criterion.
2.4 Truncation of predictions
PILOT relies on two truncation procedures to avoid unstable predictions on the training data and the test data.
The first truncation procedure is motivated as follows. The left panel of Fig. 3 shows the data in a node of PILOT. The selected predictor is on the horizontal axis. The plot illustrates a situation where most of the data points in a node are concentrated in a part of the predictor’s range, causing the linear model to put a lot of weight on these points and little weight on the other(s). This can result in extreme predictions, for instance for the red point on the right. Although such unstable predictions might be corrected by deeper nodes, this could induce unnecessarily complicated models with extra splits. Moreover, if this effect occurs at a leaf node, the prediction cannot be corrected. To avoid this issue we will truncate the prediction function. Note that if we add a constant to the response, PILOT will yield the same estimated parameters except for the intercept terms that change the way they should. Denote \(B=\max (Y)=-\min (Y)\), thus the original responses have the range \([-B,B]\). We then clip the prediction so it belongs to \([-1.5B,1.5B]\). Then we compute the response Y minus the truncated prediction to get the new residuals, and proceed with building the tree for the next depth. The whole process is summarized in Algorithm 3. We found empirically that this works well.
The first truncation method is not only applied when training the model, but also when predicting on new data. The range of the response of the training data is stored, and when a prediction for new data would be outside the proposed range \([-1.5B,1.5B]\) it is truncated in the same way. This procedure works as a basic safeguard for the prediction. However, this is not enough because a new issue can arise: it may happen that values of a predictor in the test data fall outside the range of the same predictor in the training data. This is illustrated in the right panel of Fig. 3. The vertical dashed lines indicate the range of the predictor on the training data. However, the predictor value of the test case (the red point) lies quite far from that range. Predicting its response by the straight line that was fitted during training would be an extrapolation, which could be unrealistic. This is a known problem of linear model trees. For instance, in our experiments in Sect. 4 the out-of-sample predictions of FRIED and M5 became extremely large on some data sets, which turned out to be due to this exact problem.
Therefore we add a second truncation procedure as done in Loh et al. (2007). This second truncation step is only applied to new data because in the training data, all predictor values are by definition inside the desired range. More precisely, during training we record the range \([x_{\hbox {\tiny min}}, x_{\hbox {\tiny max}}]\) of the predictor selected in this node, and store the range of the corresponding predictions \([{\hat{f}}(x_{\hbox {\tiny min}}),{\hat{f}}(x_{\hbox {\tiny max}})]\). When predicting on new data, we truncate the predictions so they stay in the training range. More precisely, we replace the original predictions \({\hat{f}}(x_{\hbox {\tiny test}})\) on this node by \(\max (\min ({\hat{f}}(x_{\hbox {\tiny test}}),{\hat{f}}(x_{\hbox {\tiny max}})),{\hat{f}}(x_{\hbox {\tiny min}}))\).
The two truncation methods complement each other. For instance, the first approach would not suffice in the right panel of Fig. 3 since the linear prediction at the new case would still lie in \([-1.5B,1.5B]\). Also, the second approach would not work in the left panel of Fig. 3 as the unusual case is included in the range of the training data. Our empirical studies indicate that combining both truncation approaches helps eliminate extreme predictions and therefore improve the stability of our model. Moreover, this did not detract from the overall predictive power. Algorithm 4 describes how both truncation methods are applied when PILOT makes predictions on new data.
2.5 Stopping rules versus pruning
It is well known that decision trees have a tendency to overfit the data if the tree is allowed to become very deep. A first step toward avoiding this is to require a minimal number of cases in a node before it can be split, and a minimal number of cases in each child node. In addition, PILOT also stops splitting a node if the BIC model selection criterion selects con.
Several decision trees, including CART, take a different approach. They let the tree grow, and afterward prune it. This indeed helps to achieve a lower generalization error. However, pruning can be very time consuming. One often needs a cross-validation procedure to select pruning parameters. It would be possible for PILOT to also incorporate cost-complexity pruning as in CART. It would work exactly the same as in CART, and additionally consider a series of lin fits as a single model to be kept or removed in the pruning. However, in our empirical study we found that adding this pruning to PILOT did not outperform the existing con stopping rule on a variety of datasets. Also, we will see in the next section that the generalization error vanishes asymptotically. Moreover, not pruning is much more efficient computationally. For these reasons, PILOT does not employ pruning. The computational gain of this choice makes it more feasible to use PILOT in ensemble methods.
3 Theoretical results
3.1 Universal consistency
In this section we prove the consistency of PILOT. We follow the settings in Klusowski (2021) and assume the underlying function \(f\in \mathcal {F}\subset L^2([0,1]^p)\) admits an additive form
where \(f_i\) has bounded variation and \(X^{(j)}\) is the j-th predictor. We define the total variation norm \(||f||_{TV}\) of \(f\in \mathcal {F}\) as the infimum of \(\sum _{i=1}^p||f_i||_{TV}\) over all possible representations of f, and assume that the representation in (2) attains this infimum.
For all \(f, g \in L^2([0,1]^p)\) and a dataset \(X_1, \ldots , X_n\) of size n, we define the empirical norm and the empirical inner product as
For the response vector \(Y = (Y_1,\ldots , Y_n)\) we denote
The general \(L^2\) norm on vectors and functions will be denoted as \(||\cdot ||\) without subscript.
To indicate the norm and the inner product of a function on a node T with t observations we replace the subscript n by t. We denote by \(\mathcal {T}_k\) the set of tree nodes at depth k, plus the leaf nodes of depth lower than k. In particular, \(\mathcal {T}_K\) contains all the leaf nodes of a K-depth tree.
Now we can state the main theorem:
Theorem 1
Let \(f\in \mathcal {F}\) with \(||f||_{TV}\, \leqslant A\) and denote by \({\hat{f}}(\mathcal {T}_K)\) the prediction of a K-depth PILOT tree. Suppose \(X\sim P\) on \([0,1]^{p_n}\), the responses are a.s. bounded in \([-B,B]\), and the depth \(K_n\) and the number of predictors \(p_n\) satisfy \(K_n\rightarrow \infty\) and \(2^{K_n}p_n\log (np_n)/n\rightarrow 0\). Then PILOT is consistent, that is
Note that the conditions on the depth and the dimension can easily be satisfied if we let \(K_n=\log _2(n)/r\) for some \(r>1\) and \(p_n=n^{s}\) such that \(0<s<1-1/r\). The resulting convergence rate is \(\mathcal {O}(1/\log (n))\). This is the same rate as previously obtained for CART under very similar assumptions (Klusowski, 2021).
The key of the proof of Theorem 1 is to establish a recursive formula for the training error \(R_k:=||Y-{\hat{f}}(\mathcal {T}_k)||^2_n-||Y-f||^2_n\) for each depth k. Then we can leverage results from empirical process theory (Györfi et al., 2002) to derive an oracle inequality for PILOT, and finally prove consistency.
For the recursive formula we first note that \(R_k=\sum _{T\in \mathcal {T}_K}w(T)R_k(T)\) where \(w(T):=t/n\) is the weight of node T and \(R_k(T):=||Y-{\hat{f}}(\mathcal {T}_k)||^2_t-||Y-f||^2_t\). Then we immediately have \(R_{k+1}=R_k-\sum _{T\in \mathcal {T}_k}w(T)\Delta ^{k+1}(T)\) where
is the impurity gain of the model on T. Here \(t_l\) and \(t_r\) denote the number of cases in the left and right child node. If no split occurs, one of these numbers is set to zero. For PILOT we need to remove con nodes that do not produce an impurity gain from the recursive formula. To do this, we define \(C_k^+=\{T|T=\textsc {con}, T\in \mathcal T_{k-1}, R_k(T)>0\}\), i.e., the set of ‘bad’ nodes on which con is fitted, and similarly \(C_k^-\) for those with \(R_k(T) \leqslant 0\). Then we can define \(R_{C_k^+}:=\sum _{T\in C_k^+}w(T)R_k(T)\), the positive training error corresponding to the cases in these con nodes. We can then consider \(\widetilde{R}_k:=R_k-R_{C_k^+}\) and show that asymptotically both \(R_k\) and \(R_{C_k^+}\) become small.
Now the problem is reduced to relating the impurity gain of the selected model to the training error before this step. Recently, a novel approach was introduced for this kind of estimation in CART (Klusowski, 2021). However, the proof used the fact that in each step the prediction is piecewise constant, which is not the case here. Therefore, we derived a generalized result for pcon in PILOT (Lemma 3 in Appendix 2.1). For controlling the gain of the other models, we can use the following proposition:
Theorem 2
Let \(\Delta _1\), \(\Delta _2\) and \(v_1\), \(v_2\) be the impurity gains and degrees of freedom of two regression models on some node T with t cases. Let \(R_0\) be the initial residual sum of squares in T. We have that
-
If model 1 does better than con, i.e. \(BIC_{\hbox {\tiny {CON}}}> BIC_1\), we have \(t\Delta _1/R_0> C(v_1,t)>0\) for some positive function C depending on \(v_1\) and t.
-
If \(BIC_{\hbox {\tiny {CON}}} > BIC_1\) and \(BIC_2 > BIC_1\) with \(v_2 \geqslant v_1\) we have \(\frac{\Delta _1}{\Delta _2} \geqslant \frac{v_1-1}{v_2-1}.\)
Moreover, if con is chosen at a node, it will also be chosen in subsequent models.
We thus know that the gain of the selected model is always comparable to that of pcon, up to a constant factor.
The proposition also justifies the con stopping rule from a model selection point of view: when con is selected for a node, all the subsequent models for this node will still be con. A con model is selected when the gain of other regression models do not make up for the complexity they introduce. Since con only regresses out the mean, we can stop the algorithm within the node once the first con fit is encountered.
We can show the following recursive formula:
Theorem 3
Under the same assumptions as Theorem 1 we have for any \(K\geqslant 1\) and any \(f\in \mathcal {F}\) that
Moreover, if we let \(K=\log _2(n)/r\) with \(r>1\) we have \(R_{C_K^+}\sim \mathcal {O}(\sqrt{\log (n)/n^{(r-1)/r}})\).
The BIC criterion ensures that the training errors in all con nodes vanish as \(n\rightarrow \infty\). This allows us to prove the main theorem by using the preceding results and Theorems 11.4 and 9.4 and Lemma 13.1 of Györfi et al. (2002). All intermediate lemmas and proofs can be found in the “Appendix 2”.
3.2 Convergence rates on linear models
The convergence rate resulting from Theorem 1 is the same as the previously derived one of CART (Klusowski, 2021). This is because we make no specific assumptions on the properties of the true underlying additive function. Therefore this rate of convergence holds for a wide variety of functions, including poorly behaved ones. In practice however, the convergence could be much faster if the function is somehow well behaved. In particular, given that PILOT incorporates linear models in its nodes, it is likely that it will perform better when the true underlying function is linear, which is a special case of an additive model. We will show that PILOT indeed has an improved rate of convergence in that setting. This result does not hold for CART, and to the best of our knowledge was not proved for any other linear model tree algorithm.
In order to prove the convergence rate of PILOT in linear models, we apply a recent result of Freund et al. (2017). For the convergence rate of the \(L^2\) boosting algorithm, they showed in their Theorem 2.1 that the mean squared error (MSE) decays exponentially with respect to the depth K, for a fixed design matrix. The proof is based on the connection between \(L^2\) boosting and quadratic programming (QP) on the squared loss \(||Y-\varvec{X}\beta ||^2_n\) . In fact, the \(L^\infty\) norm of the gradient of the squared loss is equal to the largest impurity gain among all simple linear models, assuming each predictor has been centered and standardized. Therefore, the gain of the k-th step depends linearly on the mean squared error in step \(k-1\), which results in a faster convergence rate. By the nature of quadratic programming, the rate depends on the smallest eigenvalue \(\lambda _{min}\) of \(\varvec{X}^\top \varvec{X}\).
In our framework we can consider a local QP problem in each node to estimate the gain of lin, which gives a lower bound for the improvement of the selected model (except when con is selected). To ensure that we have a constant ratio for the error decay, we have to make some assumptions on the smallest eigenvalue of the local correlation matrix of the predictors. This eigenvalue can be regarded as a measure of multicollinearity between the predictors, which is a key factor that affects the convergence rate of least squares regression.
To be precise, we require that the smallest eigenvalue of the theoretical correlation matrix (the jl-th entry of which is \(\mathbb {E}[(X-{\bar{X}})(X-{\bar{X}})^\top ]_{jl}/(\sigma _j\sigma _l)\) where \(\sigma _j\) is the standard deviation of the j-th predictor) is lower bounded by some constant \(\lambda _0\) in any cubic region that is sufficiently small. Then we apply two concentration inequalities to show that the smallest eigenvalue of a covariance matrix of data in such a cube is larger than \(\lambda _0\) with high probability. Finally, we can show that the expectation of the error decays exponentially, which leads to a fast convergence rate.
Our conditions are:
-
Condition 1: The PILOT algorithm stops splitting a node whenever
-
the number of cases in the node is less than \(n_{\hbox {\tiny min}}=n^\alpha\), \(0<\alpha <1\).
-
the variance of some predictor is less than \(2\sigma _0^2\) where \(0<\sigma _0<1\).
-
the volume of the node is less than a threshold \(\eta\), \(0<\eta <1\).
-
-
Condition 2: We assume that \(X\sim P\) on \([0,1]^p\) and the error \(\epsilon\) has a finite fourth moment. Moreover, for any cube C with volume smaller than \(\eta\) we assume that\(\lambda _{min}(Cor(X|X\in C)) \geqslant 2\lambda _0>0\), where Cor(X) is the correlation matrix.
The first condition is natural for tree-based methods. For the second one, an immediate example would be a setting when all the predictors are independent. Another example is when they follow a multivariate Gaussian distribution, which becomes a truncated Gaussian when restricted to \([0,1]^p\) or to some other cube C. It can be shown that the Gaussian distribution has relatively small correlation in cubes C of small volume; see, e.g., Muthen (1990) for the two-dimensional case. This is not surprising since on small cubes the density tends to a constant. As a result, the smallest eigenvalue of the correlation matrix on such cubes will be bounded from below.
Under these conditions we obtain the following result, whose proof can be found in “Appendix 3”.
Theorem 4
Assume the data are generated by the linear model \(Y\sim \varvec{X}\beta + \epsilon\) with n cases. Under Conditions 1 and 2, the difference between the training loss \(L^k_n\) of PILOT at depth k and the training loss \(L^*_n\) of least squares linear regression satisfies
where
Combining this result with the techniques in the proof of Theorem 1, we can show polynomial rate convergence of our method on linear functions.
Theorem 5
(Fast convergence on linear models) Assume the conditions of Theorem 4 hold and that |Y| is a.s. bounded by some constant B. Let \(K_n= log_\gamma (n)\). Then we have for any \(0<\alpha <1\) and corresponding \(n_{\hbox {\tiny min}} = n^{\alpha }\) that
The choice of the tuning parameter \(\alpha\) thus determines the convergence rate. When we use a low \(\alpha\) we get smaller nodes hence more nodes, yielding looser bounds in the oracle inequality and for the errors in con nodes, leading to a slower convergence. This is natural, as we have fewer cases for the estimation in each node. If we choose \(\alpha\) relatively high we obtain a faster rate, which is intuitive since for \(\alpha \rightarrow 1\) we would have only a single node.
3.3 Time and space complexity
Finally, we demonstrate that the time complexity and space complexity of PILOT are the same as for CART without its pruning. The main point of the proof is that for a single predictor, each of the five models can be evaluated in a single pass through the predictor values. Evaluating the next split point requires only a rank-one update of the Gram and moment matrices, which can be done in constant time. Naturally, the implicit proportionality factors in front of the \(\mathcal {O}(.)\) complexities are higher for PILOT than for CART, but the algorithm is quite fast in practice.
Theorem 6
PILOT has the same time and space complexities as CART without its pruning.
Proof
For both PILOT and CART we assume that the p predictors have been presorted, which only takes \(\mathcal {O}(np\log (n))\) time once.
We first check the complexity of the model selection in Algorithm 2. It is known that the time complexity of CART for split finding is \(\mathcal {O}(np)\). For the pcon model, PILOT uses the same algorithm as CART. For con we only need to compute one average in \(\mathcal {O}(n)\) time. For lin, the evaluation of \((D^{\textsc {lin}})^\top D^{\textsc {lin}}\) and \((D^{\textsc {lin}})^\top Y\) also takes \(\mathcal {O}(n)\). In the evaluation of the plin model, the Gram matrices always satisfy \(G^{\textsc {plin}}_l+G^{\textsc {plin}}_r=G^{\textsc {lin}}=(D^{\textsc {lin}})^\top D^{\textsc {lin}}\) and the moment matrices satisfy \(M^{\textsc {plin}}_l+M^{\textsc {plin}}_r=(D^{\textsc {lin}})^\top Y\). These matrices have at most \(2^2 = 4\) entries, and can be incrementally updated in \(\mathcal {O}(1)\) time as we evaluate the next split point. For blin the reasoning is analogous, with at most \(3^2 = 9\) matrix entries.
In each model, inverting the Gram matrix only takes \(\mathcal {O}(1)\) because its size is fixed. Therefore, we can evaluate all options on one presorted predictor in one pass through the data, which remains \(\mathcal {O}(n)\), so for all predictors this takes \(\mathcal {O}(np)\) time, the same as for CART. For the space complexity, the Gram and moment matrices only require \(\mathcal {O}(1)\) of storage. Since CART also has to store the average response in each child node, which takes \(\mathcal {O}(1)\) storage as well, the space complexity of both methods is the same.
For the tree building in Algorithm 3, the time complexity of computing the indices \(I_{l,r}\) and the residuals is \(\mathcal {O}(n)\) at each depth \(1 \leqslant k \leqslant K_{max}\). CART also computes the indices \(I_{l,r}\) which requires the same complexity \(\mathcal {O}(n)\) for each step. Therefore, the overall time for both methods is \(\mathcal {O}(K_{max}n)\). Since the PILOT tree only has \(\mathcal {O}(1)\) more attributes in each node than CART, the space complexity of PILOT remains the same as that of CART.
For the initialization and prediction parts, the results are straightforward. \(\square\)
We can estimate the proportionality factor of the \(\mathcal {O}(np)\) time of the split step as follows. We have to update 3 additional pairs of Gram and moment matrices: one pair for blin, where the Gram matrix is \(3 \times 3\) and the moment matrix is \(3 \times 1\), and two pairs for plin where Gram matrices are \(2 \times 2\) and the moment matrices are \(2 \times 1\). The number of additional entries is thus \(1\times (3^2+3)+2\times (2^2+2)=24\). Therefore the time needed for PILOT is roughly 30 times that of CART. Note that this is pretty low compared with other linear model tree methods, many of which have higher asymptotic complexities. Moreover, PILOT does not contain the common pruning step of CART, which is computationally quite demanding.
To illustrate the time complexity of PILOT we ran simulations using different numbers of cases (ranging from 10 to 10,000) and predictors (ranging from 2 to 1000). The functions are piecewise linear and randomly simulated. Here, we tested the running time of the split function, which is the main part of the algorithm, so not including the presorting with time complexity \(\mathcal {O}(np\log (n))\). The other parts of PILOT are not fundamentally different from the corresponding parts in CART, and so the comparison of the split function is of primary interest.
The left panel of Fig. 4 plots the log of the measured runtime versus the log number of cases, in the middle panel versus the log number of predictors and in the right panel versus the log number of predictors times cases. The trends are linear, and the least squares lines have slopes close to 1. This illustrates the \(\mathcal {O}(np)\) time complexity.
4 Empirical evaluation
In this section we evaluate the proposed PILOT algorithm empirically and compare its results with those of some popular competitors. We start by describing the data and methods under comparison.
4.1 Data sets and methods
We analyzed 25 benchmark data sets, with the number of cases ranging from 71 to 21,263 and the number of predictors ranging from 4 to 4,088. From the UCI repository (Dua & Graff, 2017) we used the data sets Abalone, Airfoil, Auto mpg, Bike, Communities, Concrete, Diabetes, Electricity, Energy, Power plant, Real estate, Residential, Skills, slump test, Superconductor, Temperature, Thermography and Wine. From Kaggle we obtained Bodyfat, Boston Housing, Graduate Admission, and Walmart. The California Housing data came from the StatLib repository (http://lib.stat.cmu.edu/datasets/). The Ozone data is in the R-package hdi (Dezeure et al., 2015), and Riboflavin came from the R-package missMDA (Josse & Husson, 2016). Table 1 gives an alphabetical list of the data sets with their sizes.
In the comparative study we ran the following tree-based methods: the proposed PILOT algorithm we implemented in Python with NUMBA acceleration in the split function, FRIED (Friedman, 1979) implemented by us, M5 (Quinlan, 1992) from the R package Rweka (Hornik et al., 2009), and CART. We also ran ridge linear regression from the Scikit-Learn package (Pedregosa et al., 2011) and the lasso (Tibshirani, 1996). When running methods that were not developed to deal with categorical variables, we first applied one-hot encoding to such variables, that is, we replaced each categorical variable with m outcomes by m binary variables.
The settings for hyperparameter tuning are the following. For PILOT we considered the depth parameter \(K_{\text {max}}\) in (3, 6, 9, 12, 15, 18), the minimal number of cases to fit a model \(n_{\text {fit}}\) in (2, 10, 20, 40), and the minimal number of cases in a leaf node \(n_{\text {leaf}}\) in (1, 5, 10, 20). For CART and FRIED we tuned \(K_{\text {max}}\), \(n_{\text {fit}}\) and \(n_{\text {leaf}}\) over the same ranges. For the latter methods, we tuned the data driven parameter \(\alpha _{ccp}\) for cost complexity pruning for each combination of \((K_{\text {max}},n_{\text {fit}},n_{\text {leaf}})\). For M5, \(n_{\text {leaf}}\) is the only parameter since the building, smoothing and pruning procedures are parameter-free. For the Lasso and Ridge models, we tuned the \(\lambda\) of the penalty term.
In order to evaluate the methods, we start by randomly splitting each dataset into 5 folds. For each method, and on each training set (each consisting of 4 folds), we perform 5-fold cross validation to select the best combination of hyperparameters. Finally, we compute the cross-validated mean square error, that is, the average test MSE of each method with hyperparameters tuned on the test folds. The final score of each method is its MSE divided by the lowest MSE on that dataset. A score of 1 thus corresponds with the method that performed best on that dataset, and the scores of the other methods say how much larger their MSE was.
4.2 Results
The results on these datasets are summarized in Table 2. The bold entries correspond to scores of at most 1.05, that is, methods whose MSE was at most 5% higher than the MSE of the best performing method on that dataset. The bottom four lines of the table summarize the performance, and we see that PILOT achieved the best average score with the lowest standard deviation. Moreover, its mean rank and standard deviation of its rank were also the lowest.
Table 2 has some unusual entries that require an explanation. In the last column, the M5 results were partially adjusted because its predictions are not invariant to scaling the predictors. Therefore we ran M5 both with and without predictor scaling, and we present the best result here. Those based on scaling are shown in parentheses. Note that these MSEs are at least 10 times lower than the results of M5 on the unscaled dataset, indicating that M5 is sensitive to the preprocessing of the predictors. Still, the performance of M5 remained poor on the California Housing dataset. On the Superconductor data the hyperparameter tuning time for CART exceeded the preset wall time, despite applying optimized functions (from Scikit-Learn) for cross validation and pruning, so we left this table entry blank. We also noticed that FRIED had extremely high test errors (up to 100 times larger than the best MSE) for several choices of hyperparameters on the Residential dataset. A careful examination revealed that this was caused by extrapolation. At one node, the value of the predictor of a test case was outside the range of the training cases, so lin extrapolated and gave a wild prediction.
Overall, PILOT outperformed FRIED 21 times. It also did better than CART on 22 datasets. As for M5, PILOT outperformed it 15 times. In the comparison with M5 we note that PILOT has a much more stable performance, with a performance standard deviation that is 6 times smaller.
A closer look at the results reveals that PILOT struggled on the Airfoil data. A potential explanation is the fact that the Airfoil data has a rather complicated underlying structure, which is often fit using ensemble methods such as gradient boosting and random forests (Patri & Patnaik, 2015). All tree-based methods including PILOT gave relatively poor predictions on the Riboflavin data, which is high-dimensional with only 71 observations but over 4000 predictors, and whose response has a strong linear relationship with the regressors. This explains why Ridge and Lasso performed so well on these data. The tree-based methods suffered from the relatively small number of cases in the training set, which resulted in a higher test error. Still, PILOT did better than CART, FRIED and M5 on these data, suggesting that it is indeed less prone to overfitting and better at capturing linear relationships.
There are several other datasets on which Ridge and/or Lasso did very well, indicating that these data have linear patterns. The results show that PILOT tended to do quite well on those datasets and to outperform CART on them. This confirms that the linear components of PILOT make it well suited for such data, whereas we saw that it usually did at least as well as the other tree-based methods on the other datasets.
To assess the statistical significance of the performance differences involving PILOT, we conducted Wilcoxon signed rank tests between the PILOT performance and each of the other methods, with the results shown in Table 3. (The M5 result is with the scaled datasets when they did better.) All the p-values are below \(5\%\) and often much lower. To formally test for the superiority of the PILOT results in this study, we can apply the Holm-Bonferroni method (Holm, 1979) for multiple testing, at the level \(\alpha =5\%\). To this end we must sort the five p-values in Table 3 from smallest to largest, and check whether \(p_{(1)} <\alpha /5\), \(p_{(2)} < \alpha /4\), \(p_{(3)} < \alpha /3\), \(p_{(4)} < \alpha /2\), and \(p_{(5)} < \alpha /1\). All of these inequalities hold, so we have significance.
4.3 Results after transforming predictors
It has been observed that linear regression methods tend to perform better if the numerical predictors are not too skewed. In order to address potentially skewed predictors, we reran the empirical experiments after preprocessing all numerical predictors by the Yeo-Johnson (YJ) transformation (Yeo & Johnson, 2000). The parameter of the YJ transformation was fit using maximum likelihood on the training set, after which the fitted transformation parameter was applied to the test set. One way in which such a transformation could help is by mitigating the extrapolation issue, as this is expected to occur mainly on the long tailed side of a skewed predictor, which becomes shorter by the transformation.
Table 4 shows the results after transforming the predictors. The MSE ratios are relative to the lowest MSE of each row in Table 2, so the entries in Table 4 can be compared directly with those in Table 2. We see that the transformation often enhanced the performance of PILOT as well as that of some other methods. On the high dimensional Riboflavin data set the score of PILOT came closer to Ridge and Lasso, whereas the other decision trees kept struggling. Also on the other data sets with linear structure PILOT was comparable with both linear methods. The average and the standard deviation of its score and rank were reduced by the transformation as well. The Wilcoxon tests in Table 5 gave similar results as before, and the Holm-Bonferroni test again yields significance. The overall performance of PILOT remained the best. We conclude that PILOT typically benefits from transforming the data before fitting the tree.
4.4 Depth comparison between the tree-based methods
Here we compare the depth of the trees fitted by CART, PILOT and FRIED, to gain insight into how the inclusion of linear models in the nodes affects the tree structure and tree depth. Figure 5 shows the frequencies of tree depths of these methods, for all folds on all datasets, when the maximal depth was set to 18. We see that the PILOT trees are much more shallow than those built by CART. This is understandable, because CART needs many splits to model approximately linear relations to a reasonable precision. On the other hand, PILOT can avoid many of those splits by fitting linear models. The depth of FRIED trees is more similar to those of PILOT. Moreover, these conclusions remain the same when the experiment is done on the transformed datasets instead of the raw datasets. We did not include the depth of M5 for two reasons: First, not all the reported results of M5 in Table 2 were for the raw datasets, because of its sensitivity to the predictor scales. Second, the maximal depth of M5 cannot be specified as a tuning parameter (Hornik et al., 2009), making it difficult to obtain a fair comparison.
4.5 Feature importance in PILOT
Feature importance is a popular tool for assessing the role of each predictor in the final model (Hastie et al., 2009). Due to the tree structure of PILOT, we can construct a measure of feature importance similar to that of CART. For each predictor, we find the nodes in which it plays a role. We then compute the cumulative variance reduction of the corresponding fitted models. The cumulative variance reductions of all predictors are then normalized so they sum to 1. Note that this construction is exactly like the one used in CART (Hastie et al., 2009).
To illustrate feature importance in PILOT we analyze two examples, the Wine dataset and the Communities dataset. The former is low-dimensional and the latter is high-dimensional, but both datasets have variables that are easy to interpret. We first tuned the hyperparameters of PILOT and then calculated the feature importance. When computing the feature importance in PILOT, repeated lin fits in the same node are counted separately.
The Wine dataset has 11 continuous predictors and 4894 cases. The response is a measure of wine quality, provided as a score between 1 and 10. The feature importances from CART and PILOT are shown in Fig. 6. We see that the main effects are due to variables X2 (volatile acidity), X6 (free sulfur dioxide), and X11 (alcohol percentage), for both CART and PILOT. Figure 7 plots the response versus each of these predictors. We see that these predictors have some explanatory power, which CART can capture by piecewise constants. PILOT also uses these predictors to split on, but for X11 it fits additional linear models to the different regions, as seen in the last panel of Fig. 7. As a result, the feature importance of X11 is higher in PILOT than in CART, leaving less importance in some of the other variables.
We now consider the Communities dataset. Its continuous response measures the per capita violent crimes in communities in the US. The predictors are numerical too, and describe features of the communities such as median income, racial distribution, and average family composition. As the data are relatively high-dimensional with 128 predictors, we will only discuss the predictors with feature importance above 0.01 . Figure 8 shows the feature importances of CART and PILOT. In this example the difference between the two methods is more pronounced. CART identifies X51 (percentage of kids aged 4 or under in family housing with two parents) as the most important predictor, and assigns some additional importance to a few other predictors. On the other hand, PILOT identifies X45 (percentage of males who have never married) as the dominant predictor.
Figure 9 plots the response against these two predictors and superposes the fits on the root nodes of CART and PILOT. For CART this was a piecewise constant model on X51, whereas PILOT selected a blin fit on X45. The former reduces the RSS by 41%, whereas the latter yields a reduction in RSS of 56%. Therefore PILOT captured the underlying relation more accurately in its initial fit. This resulted in a substantially better predictive performance of the final model: from Table 2 we see that the MSE of PILOT was \(1.09/1.31 \approx 83\%\) of the MSE of CART on this dataset.
4.6 Explainability of PILOT
We illustrate the explainability and interpretability of PILOT on the Graduate Admission dataset, which has 400 cases and 7 predictors. The response variable is the probability of admission, and the predictors include features such as the GPA and GRE scores. Figures 10 and 11 show the CART and PILOT trees of depth 3 on this dataset. The depth of the final fits is higher, but here we focus on depth 3 for illustrative purposes. Note that the first node of the PILOT tree performs two lin fits in a row, which it is allowed to.
Both methods result in a simple cascade of decision rules and found X6 to be an important predictor, which corresponds to the student’s GPA score. CART performs six splits out of seven on X6, whereas PILOT carries out two splits out of three on X6, and has X6 in all the linear models in the leaf nodes.
The left panel of Fig. 12 shows the probability of admission versus the GPA score X6. The GPA is clearly related with the response. Moreover, the relationship looks nearly linear, making a piecewise linear model a much better fit than a piecewise constant model. This is confirmed by the fact that the initial PILOT fit on X6 explains 77.8% of the variance of the response, while the initial CART split on X6 explains 54.1% of the variance. The explained variance of the final trees is 78.3% for PILOT versus 72.4% for CART, while PILOT uses half the number of leaf nodes and less than half the number of splits.
Regarding the other predictors, both methods carry out a split on the GRE score X1 at depth 2. The relation between the admission probability and X1 is shown in the right panel of Fig. 12. Interestingly, PILOT also chooses a piecewise constant split on this variable, and does so at basically the same split point as CART. Therefore, both models suggest that the GRE score is sometimes used as a rough threshold to distinguish between candidates. Moreover, PILOT suggests that students have little to gain by raising the GRE much above this threshold. Figure 11 indicates that PILOT makes slight linear adjustments based on the variables X2 (TOEFL score), X3 (university rating) and X5 (quality of personal statement).
5 Conclusion
This paper presented a new linear model tree algorithm called PILOT. It is computationally efficient, as it has the same time and space complexity as CART. To avoid overfitting, PILOT is regularized by a model selection rule. This regularization adds no computational complexity, and no pruning is required once the tree has been built. This makes PILOT faster than most existing linear model trees. The prediction can easily be interpreted due to its tree structure and its additive piecewise linear nature. To guide variable selection, a measure of feature importance was defined in the same way as in CART. PILOT applies two truncation procedures to the predictions, in order to avoid the extreme extrapolation errors that were sometimes observed with other linear model trees. An empirical study found that PILOT often outperformed the CART, M5, and FRIED decision trees on a variety of datasets. When applied to roughly linear data, PILOT behaved more similarly to high-dimensional linear methods than other tree-based approaches did, indicating a better ability to discover linear structures. We proved a theoretical guarantee for its consistency on a general class of functions. When applied to data generated by a linear model, the convergence rate of PILOT is polynomial. To the best of our knowledge, this is the first linear model tree with proven theoretical guarantees.
We feel that PILOT is particularly well suited for fields that require both performance and explainability, such as healthcare (Ahmad et al., 2018), business analytics (Bohanec et al., 2017; Delen et al., 2013) and public policy (Brennan & Oliver, 2013), where it could support decision making.
A future research direction is to integrate PILOT trees as base learners in ensemble methods, as it is well known that the accuracy and diversity of base learners benefit the performance of the ensemble. On the one hand, we have seen that PILOT gave accurate predictions on a number of datasets. On the other hand, the wide choice of models available at each node allows for greater diversity of the base learner. For these reasons and the fact that PILOT requires little computational cost, we expect it to be a suitable base learner for random forests and gradient boosting.
6 Supplementary information
This consists of Python code for the proposed method, and an example script.
Data availability
The data was downloaded from publicly available repositories.
Code availability
The Python code is submitted as Supplementary Material. It is also available at https://github.com/STAN-UAntwerp.
References
Ahmad, M. A., Eckert, C., & Teredesai, A. (2018). Interpretable machine learning in healthcare. In Proceedings of the 2018 ACM international conference on bioinformatics, computational biology, and health informatics (pp. 559–560).
Aydin, N., Sahin, N., Deveci, M., & Pamucar, D. (2022). Prediction of financial distress of companies with artificial neural networks and decision trees models. Machine Learning with Applications, 10, 100432. https://doi.org/10.1016/j.mlwa.2022.100432
Bohanec, M., Borštnar, M. K., & Robnik-Šikonja, M. (2017). Explaining machine learning models in sales predictions. Expert Systems with Applications, 71, 416–428.
Breiman, L. (2001). Random forests. Machine Learning, 45(1), 5–32.
Breiman, L., Friedman, J. H., Olshen, R. A., & Stone, C. J. (1984). Classification and regression trees. Routledge.
Brennan, T., & Oliver, W. L. (2013). Emergence of machine learning techniques in criminology: Implications of complexity in our data and in research questions. Criminology & Public Policy, 12, 551.
Bühlmann, P. (2006). Boosting for high-dimensional linear models. The Annals of Statistics, 34(2), 559–583.
Chen, T., & Guestrin, C. (2016). Xgboost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining
da Silva, R. G., Ribeiro, M. H. D. M., Moreno, S. R., Mariani, V. C., & Santos Coelho, L. (2021). A novel decomposition-ensemble learning framework for multi-step ahead wind energy forecasting. Energy, 216, 119174. https://doi.org/10.1016/j.energy.2020.119174
Delen, D., Kuzey, C., & Uyar, A. (2013). Measuring firm performance using financial ratios: A decision tree approach. Expert Systems with Applications, 40(10), 3970–3983.
Dezeure, R., Bühlmann, P., Meier, L., & Meinshausen, N. (2015). High-dimensional inference: Confidence intervals, p-values and R-software hdi. Statistical Science, 30(4), 533–558.
Dobra, A., & Gehrke, J. (2002). SECRET: A scalable linear regression tree algorithm. In Proceedings of the 8th ACM SIGKDD international conference on knowledge discovery and data mining (pp. 481–487).
Dua, D., & Graff, C. (2017). UCI machine learning repository. http://archive.ics.uci.edu/ml
Fernández-Delgado, M., Sirsat, M. S., Cernadas, E., Alawadi, S., Barro, S., & Febrero-Bande, M. (2019). An extensive experimental survey of regression methods. Neural Networks, 111, 11–34. https://doi.org/10.1016/j.neunet.2018.12.010
Freund, R. M., Grigas, P., & Mazumder, R. (2017). A new perspective on boosting in linear regression via subgradient optimization and relatives. The Annals of Statistics, 45(6), 2328–2364.
Friedman, J. H. (1979). A tree-structured approach to nonparametric multiple regression. In T. Gasser & M. Rosenblatt (Eds.), Smoothing techniques for curve estimation (pp. 5–22). Springer.
Friedman, J. H. (1991). Multivariate adaptive regression splines. The Annals of Statistics, 19(1), 1–67.
Golbayani, P., Florescu, I., & Chatterjee, R. (2020). A comparative study of forecasting corporate credit ratings using neural networks, support vector machines, and decision trees. The North American Journal of Economics and Finance, 54, 101251. https://doi.org/10.1016/j.najef.2020.101251
Györfi, L., Kohler, M., Krzyzak, A., & Walk, H. (2002). A distribution-free theory of nonparametric regression. Springer.
Hall, A. R., Osborn, D. R., & Sakkas, N. (2017). The asymptotic behaviour of the residual sum of squares in models with multiple break points. Econometric Reviews, 36, 667–698.
Hastie, T., Tibshirani, R., & Friedman, J. H. (2009). The elements of statistical learning: Data mining, inference, and prediction. Springer.
Holm, S. (1979). A simple sequentially rejective multiple test procedure. Scandinavian Journal of Statistics, 6(2), 65–70.
Hornik, K., Buchta, C., & Zeileis, A. (2009). Open-source machine learning: R meets Weka. Computational Statistics, 24(2), 225–232.
Josse, J., & Husson, F. (2016). missMDA: A package for handling missing values in multivariate data analysis. Journal of Statistical Software, 70(1), 1–31.
Ke, G., Meng, Q., Finley, T., Wang, T., Chen, W., Ma, W., Ye, Q., & Liu, T.-Y. (2017). LightGBM: A highly efficient gradient boosting decision tree. Advances in Neural Information Processing Systems, 30
Khaledian, Y., & Miller, B. A. (2020). Selecting appropriate machine learning methods for digital soil mapping. Applied Mathematical Modelling, 81, 401–418. https://doi.org/10.1016/j.apm.2019.12.016
Klusowski, J. M. (2021). Universal consistency of decision trees in high dimensions. arXiv preprint arXiv:2104.13881
Loh, W.-Y. (2002). Regression trees with unbiased variable selection and interaction detection. Statistica Sinica, 12, 361–386.
Loh, W.-Y., Chen, C.-W., & Zheng, W. (2007). Extrapolation errors in linear model trees. ACM Transactions on Knowledge Discovery from Data (TKDD), 1(2), 1–17.
Malerba, D., Esposito, F., Ceci, M., & Appice, A. (2004). Top-down induction of model trees with regression and splitting nodes. IEEE Transactions on Pattern Analysis and Machine Intelligence, 26(5), 612–625.
Maurer, A., & Pontil, M. (2009). Empirical Bernstein bounds and sample variance penalization. arXiv preprint arXiv:0907.3740
Muthen, B. (1990). Moments of the censored and truncated bivariate normal distribution. British Journal of Mathematical and Statistical Psychology, 43(1), 131–143.
Patri, A., & Patnaik, Y. (2015). Random forest and stochastic gradient tree boosting based approach for the prediction of airfoil self-noise. Procedia Computer Science, 46, 109–121.
Pedregosa, F., Varoquaux, G., Gramfort, A., Michel, V., Thirion, B., Grisel, O., Blondel, M., Prettenhofer, P., Weiss, R., Dubourg, V., Vanderplas, J., Passos, A., Cournapeau, D., Brucher, M., Perrot, M., & Duchesnay, E. (2011). Scikit-learn: Machine learning in Python. Journal of Machine Learning Research, 12, 2825–2830.
Pham, Q. B., Kumar, M., Di Nunno, F., Elbeltagi, A., Granata, F., Islam, A. R. M. T., Talukdar, S., Nguyen, X. C., Ahmed, A. N., & Anh, D. T. (2022). Groundwater level prediction using machine learning algorithms in a drought-prone area. Neural Computing and Applications, 13, 10751–10773.
Quinlan, J. R. (1992). Learning with continuous classes. In 5th Australian joint conference on artificial intelligence (Vol. 92, pp. 343–348). World Scientific.
Quinlan, J. R.: C4.5: Programs for machine learning. The Morgan Kaufmann Series in Machine Learning (1993)
Scornet, E., Biau, G., & Vert, J.-P. (2015). Consistency of random forests. The Annals of Statistics, 43(4), 1716–1741.
Shaikhina, T., Lowe, D., Daga, S., Briggs, D., Higgins, R., & Khovanova, N. (2019). Decision tree and random forest models for outcome prediction in antibody incompatible kidney transplantation. Biomedical Signal Processing and Control, 52, 456–462. https://doi.org/10.1016/j.bspc.2017.01.012
Shamshirband, S., Hashemi, S., Salimi, H., Samadianfard, S., Asadi, E., Shadkani, S., Kargar, K., Mosavi, A., Nabipour, N., & Chau, K.-W. (2020). Predicting standardized streamflow index for hydrological drought using machine learning models. Engineering Applications of Computational Fluid Mechanics, 14(1), 339–350.
Shi, Y., Li, J., & Li, Z. (2019). Gradient boosting with piece-wise linear regression trees. In Proceedings of the 28th international joint conference on artificial intelligence. IJCAI’19 (pp. 3432–3438). AAAI Press.
Tariq, A., Yan, J., Gagnon, A. S., Khan, M. R., & Mumtaz, F. (2023). Mapping of cropland, cropping patterns and crop types by combining optical remote sensing images with decision tree classifier and random forest. Geo-spatial Information Science, 26(3), 302–320. https://doi.org/10.1080/10095020.2022.2100287
Tibshirani, R. (1996). Regression shrinkage and selection via the lasso. Journal of the Royal Statistical Society: Series B, 58(1), 267–288.
Tropp, J. A. (2015). An introduction to matrix concentration inequalities. Foundations and Trends in Machine Learning, 8(1–2), 1–230.
Vogel, D.S., Asparouhov, O., & Scheffer, T. (2007). Scalable look-ahead linear regression trees. In Proceedings of the 13th ACM SIGKDD international conference on knowledge discovery and data mining (pp. 757–764).
Yeo, I.-K., & Johnson, R. A. (2000). A new family of power transformations to improve normality or symmetry. Biometrika, 87(4), 954–959.
Funding
This work was supported by the European Union’s Horizon 2022 research and innovation programme under the MSCA Grant Agreement No. 101103017 (JR), by Grant C16/15/068 of KU Leuven, Belgium (PR), and by the BASF Chair on Robust Predictive Analytics at KU Leuven (JR, PR, TV, RY). This research also received funding from the Flemish Government under the "Onderzoeksprogramma Artificiele Intelligentie (AI) Vlaanderen" programme.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no Conflict of interest.
Additional information
Editor: Annalisa Appice.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendix 1: Preliminary theoretical results
Here we provide notations and preliminary results for the theorems in Sect. 3.
1.1 Notation
We follow the notations in Sects. 2 and 3. The n response values form the vector Y with mean \(\overline{Y}\). The values of the predictors of one case are combined in \(X\in \mathbb {R}^p\). The design matrix of all n cases is denoted as \(\varvec{X}\in \mathbb {R}^{n\times p}\). Given some tree node T with t cases, we denote by \(\varvec{X}_T:=(X_{T_1},\dots ,X_{T_t})^\top \in \mathbb {R}^{t\times p}\) the restricted data matrix and by \(X^{(j)}_T\) its j-th column. The variance of the j-th column is given by \(\hat{\sigma }_{j,T}^2:=\sum _{k=1}^t(X_{T_k}^{(j)}-\overline{X^{(j)}_T})^2/t\). We also let \((\hat{\sigma }_{j,T}^U)^2\) be the classical unbiased variance estimates with denominator \(t-1\). T is omitted in these symbols when T is the root node, or if there is no ambiguity.
We denote by \(\mathcal {T}_k\) the set of nodes of depth k together with the leaf nodes of depth less than k. The PILOT prediction on these nodes is denoted as \({\hat{f}}(\mathcal {T}_k)\in \mathbb R^{n}\), and \({\hat{f}}^k_T\) denotes the prediction of the selected model (e.g. plin) on some \(T\in \mathcal {T}_k\) , obtained by fitting the residuals \(Y-{\hat{f}}(\mathcal {T}_{k-1})\) of the previous level. The impurity gain of a model on a node T with t cases can be written as
where \(T_l\), \(T_r\) are the left and right child nodes, containing \(t_l\) and \(t_r\) cases, and \(P(t_l) = t_l/t\), \(P(t_r) = t_r/t\).
Since \({\hat{f}}(\mathcal {T}_k)\) is additive and we assume that all \(f\in \mathcal {F}\) are too, we can write the functions on the j-th predictor as \({\hat{f}}_j(\mathcal {T}_k)\) and \(f_j\) . The total variation of a function f on T is denoted as \(||f||_{TV(T)}\). For nonzero values \(A_n\) and \(B_n\) which depend on \(n\rightarrow \infty\), we write \(A_n\precsim B_n\) if and only if \(A_n/B_n\le \mathcal {O}(1)\), and \(\succsim\) is analogous. We write \(A_n\asymp B_n\) if \(A_n/B_n= \mathcal {O}(1)\). The difference of the sets \(C_1\) and \(C_2\) is denoted as \(C_1\backslash C_2\).
1.2 Representation of the gain and residuals
In this section we show that the impurity gain of lin and pcon can be written as the square of an inner product. It is also shown that plin can be regarded as a combination of these two models. These formulas form the starting point of the proof of the consistency of PILOT and its convergence rate on linear models.
Proposition 7
Let \(\hat{\beta }\) be the least squares estimator of a linear model fitting \(Y\in \mathbb R^p\) in function of a design matrix \(\varvec{X}\in \mathbb {R}^{n\times p}\). Then we have
Proof
This follows from \(\hat{\beta }= (\varvec{X}^\top \varvec{X})^{-1}\varvec{X}^\top Y\) and \(\langle Y,\varvec{X}\hat{\beta }\rangle _n = Y^\top \varvec{X}\hat{\beta }/n\). \(\square\)
Lemma 1
(Representation of lin) Let T be a node with t cases. For a lin model on the variable \(X^{(j)}\) it holds that:
Moreover, if we normalize the second term in the last expression and denote
where \(w(t) = t/n\), we have
Proof
By Proposition 7 we have
For \(\hat{\beta }\) we don’t need to take the mean of the residuals into account, hence
On the other hand \(\hat{\alpha }= \overline{Y-{\hat{f}}(\mathcal {T}_k)}-\hat{\beta }\overline{X^{(j)}_T}\), therefore
The second result follows from the definition of \(\hat{\alpha }\), \(\hat{\beta }\) and the above formula. \(\square\)
Next we show a similar result for pcon. Note that here our \({\hat{f}}(\mathcal {T}_k)\) is not a constant on each node, so we need to adapt the result for CART in Klusowski (2021) to our case.
Lemma 2
(Representation of pcon) Let T be a node with t cases and \(T_l\), \(T_r\) be its left and right children. We then have
Moreover, if we normalize the second term in the inner product and denote
where \(w(t)=t/n\), we have
Proof
By the definition of pcon, we have
where \(\overline{Y}_l\) is the mean of Y in \(T_l\), \(\overline{{\hat{f}}(\mathcal {T}_k)}_l\) is the mean of \({\hat{f}}(\mathcal {T}_k)\) in \(T_l\), and similarly for the cases in the right child node. In the following, we also denote the constant predictions for the left and right node as \({\hat{f}}^{k+1}_l:={\hat{f}}^{k+1}\mathbbm {1}(X_T\in T_l)\) and \({\hat{f}}^{k+1}_r:={\hat{f}}^{k+1}\mathbbm {1}(X_T\in T_r)\). Thus we have
where (i) follows from the fact that
Therefore,
Finally we have
\(\square\)
Note that plin is equivalent to a combination of the two preceding models (pcon followed by two lin models). As a result, we have a similar expansion for its prediction.
Proposition 8
(Representation of PLIN) For plin, there exists a function \(\tilde{f}^T_{\hbox {\tiny PLIN}}\) depending on X, T, \(T_l\) and \(T_r\) such that
Proof
As plin can be regarded as a pcon followed by two lin fits on the same predictor, we have
where the last equation follows from \(\langle 1,{\widetilde{f}}_{\hbox {\tiny LIN}}\rangle _n=0\) and the fact that the prediction of pcon is constant on two child nodes. Moreover, since \({\widetilde{f}}^{T_r}_{\hbox {\tiny LIN}}\), \({\widetilde{f}}^{T_l}_{\hbox {\tiny LIN}}\), \({\widetilde{f}}^T_{\hbox {\tiny PCON}}\) are orthogonal to each other, we can deduce that
for some A, B and C depending on X and the nodes T, \(T_l\) and \(T_r\). \(\square\)
Appendix 2: Proof of the universal consistency of PILOT
In this section we show the consistency of PILOT for general additive models. As discussed in Sect. 3, we begin with the estimation of the impurity gains of PILOT. Then we derive a recursion formula which allows us to develop an oracle inequality and prove the consistency at the end. For all the intermediate results, we assume the conditions in Theorem 1 holds.
1.1 A lower bound for \(\Delta _{\hbox {\tiny PCON}}\)
The key to the proof of the consistency is to relate the impurity gain \(\Delta ^{k+1}\) to the training error \(R_k:=||Y-{\hat{f}}(\mathcal {T}_k)||^2_n-||Y-f||^2_n\) so that a recursion formula on \(R_k\) can be derived. Recently, Klusowski (2021) developed such a recursion formula for CART. He showed in Lemma 7.1 that for CART, \(\Delta ^{k+1}\) is lower bounded by \(R_k^2\), up to some factors. However, the proof used the fact that \(\langle Y-\overline{Y}_t,\overline{Y}_t\rangle =0\), and this formula no longer holds when \(\overline{Y}_t\) is replaced by \({\hat{f}}(\mathcal {T}_k)\). The reason is that we cannot assume that the predictions in the leaf nodes of a PILOT tree are given by the mean response in that node, due to earlier nodes. To overcome this issue, we provide a generalized result for pcon in the following.
Lemma 3
Assuming \(R_k(T)>0\) in some node T, then the impurity gain of pcon on this node satisfies:
where \({\hat{s}}\), \({\hat{j}}\) is the optimal splitting point and prediction.
Proof
Throughout this proof, the computations are related to a single node T so that we sometimes drop the T in this proof for notational convenience. We first consider the case where the mean of the previous residuals in the node is zero. We will show that
Note that \({\hat{f}}(\mathcal {T}_k)\) is the sum of \(p_n\) linear functions defined on \(p_n\) predictors. Let \(\hat{\beta }_j\) be the slope of the linear function on j-th predictor. Then we define a probability measure \(\Pi (s,j)\) on the space \(\mathbb R\times \{1,\dots ,p_n\}\) by the following Radon-Nikodym derivative:
Here, \((t_l,t_r)\) and \((t'_l,t'_r)\) are the child nodes due to the split point s and predictor j, and \(a_j\) and \(f'_j(x)\) are defined as
where \(X_{(1)}^{(j)},\dots X_{(t)}^{(j)}\) are the ordered data points along the j-th predictor.
The idea is that the optimal impurity gain \(\hat{\Delta }_{\hbox {\tiny PCON}}({\hat{s}},{\hat{j}},t)\) is always larger than the average impurity gain with respect to this probability measure:
where the last inequality follows from Jensen’s inequality, and the representation in Lemma 2 is used. We now focus on the term inside the brackets, for which we obtain an analog of (29) in Klusowski (2021):
For the numerator we have
where (i) again used \(\langle 1, Y-{\hat{f}}(\mathcal {T}_k)\rangle _t=0\).
For the denominator restricted to one predictor, we have
If we sum over all j predictors we have
Putting the bounds on the numerator and denominator together, we obtain:
Now note that
where (i) is the triangle inequality and (ii) follows from the assumption that \(R_k(T)\) is strictly positive.
Finally, we deal with \(\frac{1}{2}||f-{\hat{f}}(\mathcal {T}_k)||_{TV(T)}\) for which we need an upper bound. We have \(\frac{1}{2}||f-{\hat{f}}(\mathcal {T}_k)||_{TV(T)}\, \leqslant \frac{1}{2}||f||_{TV}+\frac{1}{2}||{\hat{f}}(\mathcal {T}_k)||_{TV(T)}\,\). It thus remains to bound the total variation of \({\hat{f}}(\mathcal {T}_k)\) on the node T. We now show that the sum of the total variations along all the predictors is bounded by \((\max {\hat{f}}(\mathcal {T}_k)-\min {\hat{f}}(\mathcal {T}_k))\leqslant 3B<6B\). Since \({\hat{f}}(\mathcal {T}_k)\) is linear, the maximum and minimum values are always attained on vertices of the node (or cube). We can therefore assume without loss of generality that the cube is \([0,1]^{p_n}\) and \({\hat{f}}(\mathcal {T}_k)=\sum _{j=1}^{p_n}\hat{\beta }_jX^{(j)}\) s.t. for \(\forall j<j'\), \(\hat{\beta }_j<\hat{\beta }_{j'}\) and \(\hat{\beta }_j<0\Leftrightarrow j<p_0\). Now we start with the vertex \(v_0\in [0,1]^{p_n}\) such that its j-th variable is 1 if and only if \(j<p_0\) (otherwise 0). Then we move to another vertex \(v_1\) (along the edge) which is identical to \(v_0\) except on the first entry, i.e. its first entry is 0. In addition, we have \(||{\hat{f}}_1(\mathcal {T}_k)||_{TV}\,={\hat{f}}(\mathcal {T}_k)|_{X = v_1}-{\hat{f}}(\mathcal {T}_k)|_{X = v_0}\,\). Similarly, for \(\forall j\), we let the vertex \(v_{j+1}\) be identical to \(v_{j}\) except for the j-th variable (by changing 1 to 0 or 0 to 1) and we have \(||{\hat{f}}_j(\mathcal {T}_k)||_{TV}\,={\hat{f}}(\mathcal {T}_k)|_{X=v_j}-{\hat{f}}(\mathcal {T}_k)|_{X=v_{j-1}}\,\). Furthermore, it holds that \(\sum _{j=1}^{p_n}||{\hat{f}}(\mathcal {T}_k)_j||_{TV}\,=\max {\hat{f}}(\mathcal {T}_k)-\min {\hat{f}}(\mathcal {T}_k)\leqslant 6B\). Therefore, we obtain \(\left( \frac{1}{2}||f-{\hat{f}}(\mathcal {T}_k)||_{TV(T)}\right) ^2 \leqslant \frac{1}{4}(||f||_{TV}+6B)^2\).
Putting everything together yields
Finally, we treat the general case where the mean of the residuals in the node is not necessarily zero (which can happen after fitting a blin model). Note that \(R_k(T)=\{||Y-{\hat{f}}(\mathcal {T}_{k})-(\overline{Y}-\overline{{\hat{f}}(\mathcal {T}_k)})||^2_t-||Y-f||^2_t\}+(\overline{Y}-\overline{{\hat{f}}(\mathcal {T}_k)})^2:=R_{k(1)}+R_{k(2)}\), where \(R_{k(1)}\) is the squared training error after regressing out the mean and \(R_{k(2)}\) is the squared mean. Since the first truncation procedure ensures that \(R_k(T)\leqslant 16B^2\), we have \((||f||_{TV}+6B)^2\geqslant 36B^2\geqslant 2R_k(T)=2(R_{k(1)}+R_{k(2)})\), so that
Thus, by the preceding results and the fact that pcon makes the mean of the residuals zero, we have
\(\square\)
1.2 Proof of Proposition 2
As plin generalizes pcon, we always have \(\hat{\Delta }^k_{\hbox {\tiny PLIN}}\geqslant \hat{\Delta }^k_{\hbox {\tiny PCON}}\). For other models, however, it is not possible to develop a lower bound by similarly constructing a carefully designed probability measure as that in Lemma 3. This is because we do not have an indicator function which naturally generates f when associated with \(f'\) in the integral (see the estimation of the numerator). Fortunately, we can rely on the BIC criterion to obtain bounds on the relative impurity gains between the different models. More precisely, Proposition 2 shows that if blin or lin are chosen, their gain has to be comparable (i.e., differ by at most a constant factor) to that of pcon. This ensures no underfitting occurs. Moreover, if con is chosen at one node, all the subsequent models on that node would also be con which justifies the use of the con stopping rule. The proof is given in the following.
Proof
Without loss of generality, we may assume no model fits perfectly. Let us start with the first claim. By the assumption that \(BIC_1>BIC_2\) we have for any \(v_1,v_2\) that
Therefore, for any model i that is selected over con by the BIC criterion, we have
Since \(t\geqslant 2\), \(\log (t)/t\) is always positive, so is the lower bound in the preceding equations.
We now proceed with the second claim. If (12) holds for model 1, we have by (11),
On the other hand, if (12) does not hold for model 1, we still get \({\Delta _2}> C^*{\Delta _1}\) by using (11) for model 2 and con, its inverse inequality for model 1 and con, and the fact that function \(f(x)=(1+ax)/(1+bx)\) is monotonically increasing for \(x\in [-1,0]\) and \(1>a>b>0\).
Next we find the minimum of \(C^*\). We compute the derivative of the numerator and denominator of \(C^*\) with respect to \((\log t)/t\) to get \((v_2-1)\exp ((1-v_2)(\log t)/t)\) and \((v_1-1)\exp ((1-v_1)(\log t)/t)\). Therefore, for any \((\log t)/t:= s\in (0,1/2]\) we have
due to \(v_1\geqslant v_2\). Therefore we conclude that \(C^*\geqslant (v_2-1)/(v_1-1)\) for any \(t\geqslant 2\).
By previous lemmas we know that the impurity gain of the models can be divided into two parts. The first is from regressing out the mean and the second is from the model assumption, which does not depend on the mean of the response variable. Therefore, we can let \(\Delta _2=\Delta _1+\Delta _2'\) in (11) when model 1 is con. Here, \(\Delta _2'\) is the gain after regressing out the mean. Now, if con is better, we have by the inverse inequality of (11) that
which means that the subsequent node still prefers con, even if its gain is 0. \(\square\)
As an example, for our choice of degrees of freedom, \(\Delta _{\hbox {\tiny LIN}}\geqslant \Delta _{\hbox {\tiny PCON}}/4\) if lin is chosen. Similarly, if blin is the preferred model, we must have \(\Delta _{\hbox {\tiny BLIN}}\geqslant \Delta _{\hbox {\tiny PCON}}\) since their degrees of freedom are the same.
1.3 Proof of Theorem 3
The remaining issue is that the RSS in con nodes does not get improved as the depth increases, since subsequent nodes will select con again. Therefore, we first construct a recursion formula which excludes the terms corresponding to the RSS in the con nodes. Then we will show that the training error in these con nodes vanishes asymptotically, which justifies the con stopping rule.
Recall the definitions of \(C_k^+=\{T|T=\textsc {con}, T\in \mathcal {T}_{k-1}, R_{k-1}(T)>0\}\), the set of nodes on which con is fitted before the k-th step, and \(C_k^-\) for those with \(R_k(T) \leqslant 0\). Further define \(C_k^*:=C_k^+\backslash C_{k-1}^+\) and \(C_k^{\#}:=C_k^-\backslash C_{k-1}^-\). Finally, let \(A_k^+:=\{T|T\in \mathcal {T}_{k},T\ne \textsc {con},R_k(T)>0\}\) and \(A_k^-:=\{T|T\in \mathcal {T}_{k},T\ne \textsc {con},R_k(T)\leqslant 0\}\). Note that with these disjoint sets of nodes we now have
Next we can calculate the errors in these sets of nodes. Let \(R_{C_k^+}:=\sum _{T\in C_k^+}w(T)R(T)\) and define \(R_{C_k^-}\), \(R_{C_k^*}\), \(R_{C_k^{\#}}\), \(R_{A_{k-1}^+}\), \(R_{A_{k-1}^-}\) in similar fashion. This yields
Finally, we let \(\widetilde{R}_k:=R_k-R_{C_k^+}\).
Throughout the proof we assume without loss of generality that the gain of the con node fitted at depth k is already included into \(R_k\), therefore \(R_{C^+_k}\) can be regarded as the remaining error after regressing out the mean for con nodes before depth k.
Proof
Without loss of generality we may assume that \(R_{C_k^+} \leqslant R_k\) so that \(R_k>0\). Otherwise the claim follows immediately. We have
where the second inequality follows from the preceding Lemma and Proposition, and (i) is Jensen’s inequality. Since \(R_{k-1}-R_{C_{k-1}^+}\) is positive and the other three terms in \(R^*_{k-1}\) are negative, we may replace \(R^*_{k-1}\) by \(R_{k-1}-R_{C_{k-1}^+}={\widetilde{R}}_{k-1}>0\) on the right hand side of the above inequality. Furthermore, we have by definition,
In fact, by assuming that the first regression model is not con, we already have \({\widetilde{R}}_1\leqslant F/4\) for any initial \(R_0\) by either inequality in (14). Therefore, all the subsequent estimations follow the second inequality and we have for any \(k\geqslant 2\),
By a similar induction argument to Lemma 4.1 of Klusowski (2021), we get the estimation of \(R_K\) after moving \(R_{C^+_K}\) to the right hand side.
It remains to control the errors in the con nodes. We can assume that the con models have regressed out the mean in each \(C_k^+\). The first step is to use Proposition 2 and Lemma 3 to get the following upper bound for any con node T:
where we used Lemma 3 for the first inequality and (13) for the second. Since we assume the response is almost surely bounded by \(\pm B\), we have
Therefore the weighted sum of R(T) on \(C_K^+\) is asymptotically
By the Cauchy-Schwarz inequality we obtain
Therefore if we let \(K=\log _2(n)/r\) with \(r>1\), \(R_{C_K^+}\) is of order \(\mathcal {O}(\sqrt{\log (n)/n^{(r-1)/r}})\), which tends to zero as the number of cases goes to infinity. \(\square\)
1.4 Proof of Theorem 1
Now we can apply Theorem 3 to derive an oracle inequality for our method and finally prove its consistency. We will use Theorem 11.4 together with Theorem 9.4 and Lemma 13.1 from Györfi et al. (2002), since these results do not require the estimator to be an empirical risk minimizer.
Proof
Let f be the true underlying function in \(\mathcal {F}\), and \(\mathcal {F}_n\) be the class of the linear model tree. We first write the \(L^2\) error as \(||f-{\hat{f}}(\mathcal {T}_K)||^2=E_1+E_2\) to apply Theorem 11.4, where
and
By our assumption, we can define \(B_0=\max \{1.5B,1\}\) so that the class of functions and Y are a.s. bounded by \([-B_0,B_0]\), which fulfills the condition of Theorem 11.4. Thus by Theorem 11.4 with \(\epsilon =1/2\) in their notation, it holds that
where \(\alpha ,\beta \rightarrow 0\) as \(n\rightarrow \infty\).
Theorem 9.4 gives the estimation for the covering numbers of \(\mathcal {G}_n\) which denotes the class of functions in the leaf nodes. We note that the condition of the theorem \(\beta /40B_0<B_0/2\) is automatically satisfied for sufficiently large n if \(\beta \rightarrow 0\) is well-defined.
Combining Theorem 9.4 and Lemma 13.1, we get the estimation for the covering number of \(\mathcal {F}_n\). Specifically we have,
Here \(\Lambda _n\) is the set of all possible binary trees on the training set of n cases, and \(\Gamma _n(\Lambda _n)\) is the upper bound for the number of different partitions on that set that can be induced by binary trees. The upper bound on \(\Gamma _n(\Lambda _n)\) follows from Klusowski (2021) and Scornet et al. (2015). The exponent \(V(\mathcal {G}_n)\) is the VC dimension of \(\mathcal {G}_n\), and since we have multivariate linear predictions \(V(\mathcal {G}_n) = p+1\).
If we further let \(\beta \asymp \frac{B_0^2}{n}\) and \(\alpha \asymp 3B_0^4\log (np\log n)2^{K+\log (p+1)}/n\) we have
and
Moreover, \(E_1\leqslant C_1\) for some constant \(C_1\) by the fact that \(||f||_{TV}\;\leqslant A\) and both Y and \(f(\mathcal {T}_k)\) are bounded in \([-B_0,B_0]\) for any k. Thus \(\mathbb {E}[E_1]\leqslant C_1/n\) for some \(C_1>0\). By Theorem 3 we also have for any \(f\in \mathcal {F}\) that
and therefore summing up everything we have,
where \(C_3\) and \(C_4\) only depend on A and B.
By our assumption, the expected error tends to zero. Moreover, if we pick \(K_n=\log (n)/r\) for some \(r>1\) and \(p_n=n^{s}\) such that \(0<s<1-1/r\), the first term in (16) is \(\mathcal {O}(1/\log (n))\) and the second and third terms turn out to be \(o(1/\log (n))\). Therefore, the overall convergence rate becomes \(\mathcal {O}(1/\log (n))\). \(\square\)
Appendix 3: Convergence rate for linear model data
1.1 Preliminaries and Ideas
In the previous section we derived the consistency of PILOT in case the underlying is a general additive function. The convergence rate obtained in that setting is \(\mathcal {O}(1/\log (n))\), the same as CART but rather slow. Of course this is merely an upper bound, and it is not unlikely that we can obtain much faster convergence if the true underlying function is somehow well-behaved.
In this section, we consider such a scenario. In particular, we aim to show a faster convergence rate of PILOT for linear functions. In order to tackle this problem, we cannot use the same approach as for the general consistency result. In particular, a counterpart of Lemma 3 for the gain of lin would still result in a squared error in the left hand side which would restrain the convergence rate. Therefore, we take a different approach and turn to a recent tool proposed by Freund et al. (2017) in the context of boosting models. In our case, we consider a linear regression of \(Y_T\) on \(\varvec{X}_T\) in each node T and estimate the gradient of a related quadratic programming (QP) problem, thereby estimating the impurity gain.
Throughout the proof we use the same notation as that in Freund et al. (2017) to indicate a subtle difference in the excess error compared to Klusowski (2021). We first define the notation of the relevant quantities related to the loss. Let \(L^*_n(T):=\min _{\hat{\beta }}||Y-\varvec{X}_T\hat{\beta }||^2_T\) be the least squares loss on the node T and \(L^*_n:=\min _{\hat{\beta }}||Y-\varvec{X}\hat{\beta }||^2_n\) be the least squares loss on the full data. We further denote the loss of a k depth PILOT by \(L^k_n:=||Y-{\hat{f}}(\mathcal {T}_k)||^2_n\). Finally, \(L^k_n(T):=||Y_T-\varvec{X}_T\beta _T||^2_t\) is its loss in node T for some \(\beta _T\) (we can write the loss like this, since on T the prediction function is linear). When the depth k is not important, we omit the superscript.
In the following we use the notation \(\varvec{{\tilde{X}}}_T\) for the standardized predictor matrix obtained by dividing each (non-intercept) column j in \(\varvec{X}\) by \(\sqrt{n}\hat{\sigma }^U_j\) where \(\hat{\sigma }^U_j\) is the usual estimate of the standard deviation of column j. We then consider \(L_n(T):=||Y_T-\varvec{{\tilde{X}}}_T\tilde{\beta }_T||^2_t\) and write its gradient as \(\nabla L_n(T)|_{\tilde{\beta }= \tilde{\beta }_T}\,\).
Theorem 2.1 of Freund et al. (2017) used a fixed design matrix so that a global eigenvalue condition can be imposed on it. As we are fitting linear models locally, we need a more flexible result. To this end we will impose a distributional condition that makes use of the following matrix concentration inequality:
Theorem 9
(Theorem 1.6.2 of Tropp (2015)) Let \(\varvec{S}_1,\dots ,\varvec{S}_n\) be independent, centered random matrices with common dimension \(d_1\times d_2\), and assume that \(\mathbb {E}[\varvec{S}_i]=0\) and \(||\varvec{S}_i||\leqslant L\). Let \(\varvec{Z} = \sum _{k=1}^n\varvec{S}_i\) and define
Then
1.2 A probabilistic lower bound for \(\Delta _{\hbox {\tiny LIN}}\)
Our first step is to show a probabilistic bound for the impurity gain of \(\Delta _{\hbox {\tiny LIN}}\) based on the ideas in the preceding section.
Lemma 4
Let T be a node with n cases. We define \(L_n(T)\) as the training loss of PILOT and \(L^*_n(T)\) as that of least squares regression. In addition to conditions 1 and 2, we assume that the residuals of the response on T have zero mean. Then there exists a constant \(C_{}\) such that the lin model satisfies
Proof
By the definition of the gradient we have,
where r denotes the residuals \(Y-\tilde{\varvec{X}}\tilde{\beta }_T\) and the last equality follows from (10) and the assumption that the residuals of the response on T have zero mean.
By letting \(h(\cdot )=L_n(\cdot )\), \(Q=\frac{2}{n}\tilde{\varvec{X}}^\top \tilde{\varvec{X}}\) in Proposition A.1 of Freund et al. (2017) and (C9), we have (assuming the difference in the loss is positive),
All that is left to show is that
In order to do so, we compute the minimum eigenvalue of \(\tilde{\varvec{X}}^\top \tilde{\varvec{X}}\). We define the vector \(S_i\in \mathbb R^p\) by its coordinates \(S_i^{(j)}=X_i^{(j)}/\sigma _j\) where \(\sigma _j\) is the true standard deviation of the j-th predictor. By the properties of eigenvalues and singular values, we can split the estimation into three parts
Therefore, we may lower bound the eigenvalues of the matrix \(\tilde{\varvec{X}}^\top \tilde{\varvec{X}}\) using the singular values of 3 other matrices. The left hand side is already bounded by \(2\lambda _0\) by our assumption. For \(\sigma _{max}({\tilde{A}}_n)\), we note that the two matrices inside differ only in the estimation of the standard deviation. If we can show that the difference in the entries are uniformly bounded by some constant, then we could also upper bound its largest singular value. To do this, we use Theorem 10 in Maurer and Pontil (2009), which states that for i.i.d. random variables \(X_1,\dots ,X_n\) in [0,1] and \(\delta >0\), the classical unbiased variance estimates \((\hat{\sigma }^U)^2\) and the true variance \(\sigma ^2\) satisfy
Due to
we only need to control the difference in the variance term, as each \(X_i^{(k)}\) is bounded by 1. If we put \(\sqrt{2\log (1/\delta )/(n-1)}=\lambda _0\sigma ^4_0/2p\) and let \(E_k\) be the event that \(|\sigma _k-\hat{\sigma }^U_k|\leqslant \frac{\lambda _0\sigma ^4_0}{2p}\), we have
due to \(\sigma _0<1\), \(\lambda _0<2\lambda _0\leqslant tr(\mathbb {E}[(S-\overline{S})(S-\overline{S})^\top ])\leqslant p\) and our assumption that the estimated variance should be lower bounded by \(2\sigma _0^2\). On \(E:=\bigcup _k E_k\) we have
where the last inequality is due to \(\sigma _k, \sigma _l^U\leqslant 1\). Thus \(P[\max _{k,l}(|{\tilde{A}}_{kl}|)\leqslant \lambda _0/(2p)]\geqslant 1-p^2\exp (-C_{\lambda _0,\sigma _0,p}n)\) by the union bound. As \(\max _{||x||_2=1}||{\tilde{A}}x||_2\,\leqslant \max _{k,l}(|{\tilde{A}}_{kl}|)p\) we have
For \(\lambda _{max}({\tilde{B}}_n)\) we can apply Theorem 9 to obtain that it is lower bounded by \(\lambda _0/2\) with high probability. In fact, on E the squared \(L^2\) norm of \(S_i\) is bounded by \(p/\sigma _0^2\), and therefore \(||S_iS_i^\top ||=\lambda _{max}(S_iS_i^\top )\leqslant p/\sigma _0^2\) so that \(||\mathbb {E}[SS^\top ]||\leqslant \mathbb {E}[||S_i||^2]\leqslant p/\sigma _0^2\) by Jensen’s inequality. Moreover, for \(\varvec{S}_i:=\mathbb {E}[SS^\top ]-S_iS_i^\top\) we have \(\mathbb {E}[\varvec{S}_i]=0\) and
Using the same argument in Section 1.6.3 of Tropp (2015) we find that \(v(\tilde{B}_n)\leqslant p^2/n\sigma _0^4\) so that on E,
At the end we note that
Summing up all the estimations, we know that by a union bound there exists a constant \(C_{\lambda _0,\sigma _0,p}>0\) such that \(P[\lambda _{min}(\tilde{\varvec{X}}^\top \tilde{\varvec{X}})>\lambda _0]\geqslant 1-\exp (-C_{\lambda _0,\sigma _0,p}n)\). \(\square\)
1.3 Proof of Theorem 4
We can now prove the recursion formula for the expectation of the excess error.
Proof
We first deal with the case where con is not included. We know that whatever non-con model is chosen on T, its impurity gain is always larger than that of lin (because the degrees of freedom of lin is the smallest). Moreover, the first truncation procedure does not deteriorate the gain of the model, since it only eliminates bad predictions. Therefore when \(L^k_n-L^*_n\) is given and positive, by Lemma 4, with probability at least \(\sum _{T\in \mathcal {T}_k}\exp (-C_{\lambda _0,\sigma _0,p}t)\), it holds that
where \(r^k_T:=Y-X_T\beta _T\) are the residuals on T. Here, (i) follows from equation (10). The \(\tilde{\Delta }_{\hbox {\tiny LIN}}(T)\) in (ii) means the gain of lin after regressing out the mean. The last equation in (iii) follows from Lemma 4. (iv) is due to \(||r^k_T-{\bar{r}}^k_T||^2_t+||{\bar{r}}^k_T||^2_t=||r^k_T||^2_t\) and the fact that the coefficient \(\frac{\lambda _0}{4p}\) is less than 1. (v) follows from the fact that the weighted sum of the optimal loss in each leaf node \(\sum _{T\in \mathcal {T}_k}w(T)L^*_{n}(T)\) is less than \(L^*_n\) for each k. As argued in the previous lemma, the factor \(1-\frac{\lambda _0}{4p}\) is always positive.
It remains to control the ‘bad’ probability, i.e. the case where \(\lambda _{min}(T)<\lambda _0\). We have that
By condition 1 it holds that
as long as \(\log K\precsim n^\alpha\) which is satisfied by choosing K of order \(\log n\). To sum up everything, if we let \(G:=\{\forall T\in \mathcal {T}_K, \lambda _ {pmin}(T)>\lambda _0\}\) and assume \(\mathbb {E}[\epsilon ^4]<\infty\), we have for sufficiently large n,
Since we have an exponential decay in the probability, the error can be bounded by \(\mathcal O(1/n)\) if we let \(K\asymp \log n\).
When con is included, we can derive an analog of (15) by using Lemma 4 and Proposition 2 to get for all T with con:
Moreover, since the error term has a finite fourth moment and the true underlying f is linear, we have with probability at least \(1-\exp (C_{\lambda _0,\sigma _0,p}n)\) that
Let us denote by \(\mathcal {T}_K^{\textsc {con}}\) all the con nodes in the tree up to depth K. Then the expectation of the weighted sum of the errors on all of these nodes satisfies
Noticing that in our previous estimation for non-con nodes, the estimation of \(L^{k+1}_n(T)-L^*_n(T)\) is always positive if \(L^k_n(T)-L^*_n(T)>0\) due to \(\lambda _0\leqslant p\) (see the proof of Lemma 4), we were able to first ignore the con nodes to get the recursion formula, and at the end to add back the errors in all con nodes to get the final results. \(\square\)
1.4 Proof of Theorem 5
Finally, by applying the preceding results and the proof of the consistency theorem, we get the convergence rate of PILOT on linear model data.
Proof
We know that \(\mathbb {E}[L^*_n-||\varvec{X}\beta ||^2_n]=\mathcal O(1/n)\) for the true parameter \(\beta\). So if we choose \(K_n= log_\gamma (n)\), by the preceding theorem we have \(\mathbb {E}[L^{K_n}_n-||\varvec{X}\beta ||^2_n]=\mathcal O(N_{leaves}\log (n)/n)\). Finally we can follow the proof of Theorem 1 in which we replace \(2^K\) by \(N_{leaves}\) and use Theorem 4 to derive a similar oracle inequality as (16):
\(\square\)
As a side remark, in the situation where some predictors are correlated with each other, in practice plin and pcon can lead to even faster convergence than pure \(L^2\) boosting.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raymaekers, J., Rousseeuw, P.J., Verdonck, T. et al. Fast linear model trees by PILOT. Mach Learn 113, 6561–6610 (2024). https://doi.org/10.1007/s10994-024-06590-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10994-024-06590-3