Abstract
Epigenetics encompasses reversible and heritable chemical modifications of non-nuclear DNA sequences, including DNA and RNA methylation, histone modifications, non-coding RNA modifications, and chromatin rearrangements. In addition to well-studied DNA and histone methylation, RNA methylation has emerged as a hot topic in biological sciences over the past decade. N6-methyladenosine (m6A) is the most common and abundant modification in eukaryotic mRNA, affecting all RNA stages, including transcription, translation, and degradation. Advances in high-throughput sequencing technologies made it feasible to identify the chemical basis and biological functions of m6A RNA. Dysregulation of m6A levels and associated modifying proteins can both inhibit and promote cancer, highlighting the importance of the tumor microenvironment in diverse biological processes. Gastrointestinal tract cancers, including gastric, colorectal, and pancreatic cancers, are among the most common and deadly malignancies in humans. Growing evidence suggests a close association between m6A levels and the progression of gastrointestinal tumors. Global m6A modification levels are substantially modified in gastrointestinal tumor tissues and cell lines compared to healthy tissues and cells, possibly influencing various biological behaviors such as tumor cell proliferation, invasion, metastasis, and drug resistance. Exploring the diagnostic and therapeutic potential of m6A-related proteins is critical from a clinical standpoint. Developing more specific and effective m6A modulators offers new options for treating these tumors and deeper insights into gastrointestinal tract cancers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastrointestinal tract cancers are malignant diseases caused by multiple factors, with high incidence and mortality rates worldwide. They mainly include liver, gastric, colorectal, and pancreatic cancers [1,2,3,4]. Early diagnosis and patient prognosis remain formidable challenges due to the subtle symptoms at onset and during tumor invasion and metastasis [5,6,7]. Advances in medical science have led to the widespread use of diagnostic techniques such as endoscopy, tissue biopsy, and imaging technology for the diagnosis and management of gastrointestinal tract cancers [8, 9]. These essential tools allow the identification of the precise location and severity of the specific lesions [10,11,12,13]. Nearly all gastrointestinal tract cancers exhibit genomic and epigenomic DNA alterations, which, along with other microenvironment factors, play a crucial role in initiating and driving the progression of cancers [14,15,16,17,18,19]. Therefore, understanding the molecular mechanisms of tumorigenesis remains essential for improved diagnosis and treatment.
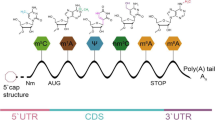
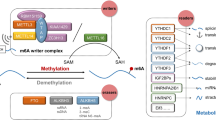
With the advancement of specific antibodies and high-throughput sequencing technologies, researchers can investigate N6-methyladenosine (m6A) sites in greater depth, marking an important milestone in RNA epitranscriptomics [20,21,22,23,24]. This reversible modification is regulated by the balance of "writers" and "erasers" proteins, indicating the potential of these proteins in regulating biological processes [25,26,27,28]. The core members of the highly conserved mRNA methyltransferase complex, known as the m6A "writer" complex, include methyltransferase-like (METTL)3, METTL14, and Wilms’ tumor 1-associating protein (WTAP) [29,30,31]. m6A modification attracts "reader" proteins to exert their biological functions. These proteins can be categorized into three types based on structural domains. The first type features the conserved YTH (YT521-B homology) domain, while the second type includes heterologous nuclear ribonucleoproteins (hnRNP) such as hnRNPC, hnRNPG, and hnRNPA2B1 [32, 33]. The third type of m6A "reader" protein includes insulin-like growth factor-binding protein (IGFBP) family proteins, such as IGFBP1-3 [34,35,36,37,38]. "Reader" proteins regulate almost every aspect of RNA metabolism, including stability, translation, and splicing of m6A transcript products [37, 39, 40]. Finally, m6A modification can be reversed through enzymatic reactions mediated by alpha-ketoglutarate dependent dioxygenase (FTO) and AlkB homolog 5 (ALKBH5), known as m6A "erasers" [41, 42]. m6A modification plays an important role in RNA epitranscriptomics, and its regulation and function are closely related to the interaction of multiple proteins affecting RNA stability, translation, degradation, nuclear localization, and splicing [43,44,45,46,47,48]. Deciphering these processes is crucial for understanding gene expression regulation mechanisms and disease development [49,50,51,52,53,54].
Gastrointestinal tract cancers are common malignant tumors with complex pathogenesis. Specifically, m6A methylation participates in the occurrence and development of gastrointestinal tract cancers by influencing several biological processes such as tumor cell proliferation, apoptosis, invasion, metabolism, immune response, and metastasis [55,56,57,58]. Furthermore, changes in m6A methylation levels are closely associated with the clinical and pathological features of gastrointestinal tract cancer. Elevated m6A methylation levels are closely correlated with poor clinical prognosis in liver cancer [59,60,61]. In gastric cancer, increased m6A methylation levels are reported to facilitate tumor cell proliferation and invasion while reducing patients' survival [62]. Therefore, m6A methylation has emerged as a potential diagnostic and prognostic marker or a promising therapeutic target [49]. Current studies are developing diagnostic and prognostic methods for different subtypes of gastrointestinal tract cancers, targeting m6A methylation to guide individualized treatment [63,64,65,66]. Combined upregulation of METTL3 and YTHDF1 was validated as a biological marker reflecting the malignancy level of liver cancer and patient prognosis [63]. Additionally, ongoing research aims to develop m6A methylation-targeted treatments to prolong patient survival and relieve cancer symptoms [67]. Further understanding and in-depth research into this modification will expand its application prospects as a novel diagnostic, prognostic indicator, and therapeutic target in gastrointestinal tract cancers [68, 69].
This review aims to explore the correlation between m6A methylation changes, clinical and pathological features, the role of m6A methylation in cancer, and its involvement in cancer processes. Summarizing these findings provides a strong foundation for considering m6A methylation as a new target for developing clinical strategies for the management of gastrointestinal tract cancers.
Molecular mechanisms of m6A modification in gastrointestinal tract cancers
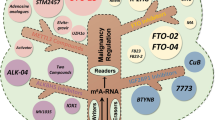
m6A RNA modification is a reversible process involved in the regulation of mRNA stability, splicing, translation, and other processes [70]. Increasing evidence links m6A modification to cancer progression, as it promotes tumor cell growth, survival, and invasion, facilitates the maintenance of stem cell self-renewal and differentiation, and confers resistance to radiotherapy and chemotherapy (Fig. 1) [71,72,73,74].
Overview of the involvement of m6a in several types of common gastrointestinal tract cancers. The dysregulation of m6A regulators plays essential roles in gastrointestinal tract cancers by impacting various biological processes such as cell proliferation, migration, invasion, and metabolic reprogramming
Furthermore, m6A has dual functions in cancer [75]. Its impact on the regulation of target genes depends on three key factors [76, 77]. Firstly, it depends on whether m6A-regulated target genes are oncogenes or tumor suppressor genes. Secondly, it is influenced by the role of the aberrant m6A regulators in cancer as "writers", "erasers", or "readers". Furthermore, the ultimate effect of these "readers" on target genes varies depending on their specific interactions, either promoting or inhibiting RNA expression [45, 78, 79]. Gastrointestinal tract cancers exhibit altered expression levels of m6A regulators, leading to dysregulated m6A modification and aberrant gene expression [80]. Exploring m6A modifications in gastrointestinal tract cancers reveals a new mechanism underlying tumor progression, with promising perspectives for clinical applications [81].
In this section, we focus on m6A methylation level changes in common digestive tract tumors, including liver, gastric, colorectal, and pancreatic cancers (Table 1). Additionally, we elucidate how m6A regulates biological processes and drives tumor progression by influencing target gene expression (Fig. 2).
The main functions and mechanism of m6A in several types of common gastrointestinal tract cancers. Various m6A regulators modulate expression levels of cancer-related genes by influencing RNA stability, post-transcriptional modifications, and translation efficiency, thus participating in the occurrence and development of gastrointestinal tract cancers
Liver cancer
Liver cancer is a common malignancy that is often discovered in advanced stages due to the lack of early symptoms. Hepatocellular carcinoma (HCC) ranks as the fourth leading cause of cancer-related deaths globally [82,83,84,85]. Early-stage HCC benefits significantly from surgical resection, but late-stage diagnosis hampers the feasibility and effectiveness of surgery [86,87,88]. Molecular targeted therapy and immune checkpoint inhibitors are pivotal for treating advanced HCC [89]. Drugs such as sorafenib, lenvatinib, and nivolumab have been widely used in clinical practice [90,91,92]. However, their usage faces several limitations, including high costs [93,94,95,96].
Therefore, in terms of future diagnosis and treatment, it is important to deeply understand the molecular mechanisms of m6A regulators in liver cancer development and develop highly effective, low side-effect treatments to mitigate this disease [97]. Research has shown that METTL3 mediates frizzled10 activation, initiating the β-catenin/YAP1 axis in HCC cells. In turn, the frizzled 10-β-catenin/c-Jun axis also transcriptionally activates METTL3 expression, forming a positive feedback loop that promotes self-renewal, expansion of liver cancer stem cells, and metastasis of liver cancer cells [98]. METTL3, as an oncogene, when knocked down, significantly reduces the proliferation, migration, and colony formation of HCC cells due to the YTHDF2-dependent m6A modification on suppressor of cytokine signaling 2 (SOCS2) mRNA. This change increases the expression of SOCS2 [99]. High expression of WTAP was revealed to suppress the expression of ETS proto-oncogene 1 (ETS1) through the m6A-HuR pattern, reducing the expression of tumor suppressors p21 and p27 and promoting the proliferation of HCC cells and tumor growth [100].
Additionally, METTL14 and microprocessor protein DiGeorge Critical Region 8 collaborate to promote the processing of miR-126, suppressing HCC metastasis through an m6A-dependent mechanism [101]. High expression of YTHDF1 enhances the activation of the phosphatidylinositol 3-kinase (PI3K)/protein kinase B (AKT)/mammalian target of rapamycin (mTOR) signaling pathway, promoting proliferation, cell cycle progression, migration, invasion, and epithelial-mesenchymal transition of liver cancer cells [102]. miR-145, a suppressive non-coding RNA in various tumors, downregulates the expression of YTHDF2 and elevates the mRNA m6A level in HCC HepG2 cells, thereby inhibiting the proliferation of HCC cells [103].
Gastric cancer
Gastric cancer is ranked the fifth most common malignancy and the third leading cause of cancer-related mortality in the world [104,105,106,107]. Risk factors encompass environmental and hereditary factors, including genetic mutations, chromosomal abnormalities, differential gene expression, and epigenetic changes [108,109,110,111,112,113]. Environmental factors include Helicobacter pylori infection, age, high salt intake, and low intake of fruits and vegetables [114, 115]. Despite a global decline in the incidence of gastric cancer over the past century, the median survival time for advanced gastric cancer is still less than one year due to late-stage detection [116,117,118,119]. Early detection and treatment remain critical for improving patient survival rates. Recent studies have revealed a significant increase in m6A methylation levels of total RNA in gastric cancer.
Dysregulation of METTL3 contributes to the oncogenesis of gastric cancer as METTL3 regulates the translation of oncogenes [120, 121]. In 415 patients from The Cancer Genome Atlas (TCGA) cohort, METTL3 expression was increased in gastric cancer tissues and associated with poor patient prognosis. Additionally, METTL3 deficiency reduced the expression level of growth factor independence 1 (GFI-1). Knockdown of METTL3 significantly inhibited proliferation, migration, and progression of epithelial-mesenchymal transition in gastric cancer cells [122].
The AKT signaling pathway is an important cellular signaling pathway that promotes the proliferation, migration, and invasion of gastric cancer cells. When METTL3 is downregulated, the AKT signaling pathway cannot be activated properly, inhibiting the proliferation, migration, and invasion ability of gastric cancer cells [123]. Consequently, knocking down METTL14 in vitro activates the Wingless/Integrated (Wnt) and PI3K/Akt signaling pathways, promoting the proliferation and invasion of gastric cancer cells [124].
Demethylase ALKBH5 inhibited the metastatic ability of gastric cancer cells through negative regulation of the expression of protein kinase, membrane-associated tyrosine/threonine 1 (PKMYT1), a member of the serine/threonine protein kinase family. Targeting IGF2BP3 in gastric cancer HGC-27 cells resulted in a significant downregulation of PKMYT1 expression, highlighting the potential role of the ALKBH5/PKMYT1/IGF2BP3 regulatory signaling pathway in gastric cancer metastasis [125].
Colorectal cancer
In recent years, the heterogeneity of colorectal cancer has attracted increasing attention [126,127,128,129]. Although substantial progress has been made in anti-tumor strategies for colorectal cancer, including multidisciplinary treatment, microbe-based therapies, molecular targeted therapy, and immunotherapy, many patients are still diagnosed at advanced stages [130,131,132]. Early diagnosis and personalized treatment remain crucial for addressing this disease. Therefore, identifying sensitive biomarkers for prognosis, recurrence monitoring, and individualized treatment management of colorectal cancer is of utmost importance [133,134,135]. METTL3 has been shown to play a complex role by participating in various regulatory pathways. It is highly expressed and mediates methylation modification of SRY (sex determining region Y)-box 2 (SOX2), preventing SOX2 mRNA degradation via IGF2BP2 recognition and contributing to cellular self-renewal, migration, and metastasis [136]. However, METTL3 can also activate the p38/ extracellular signal-regulated kinase (ERK) pathway, inhibiting the proliferation and migration of colorectal cancer cells [137].
YTHDF1 is highly expressed in various tumors and is closely associated with oncogenesis. Silencing the expression of YTHDF1 significantly suppresses the activity of the Wnt/β-catenin pathway, inhibiting tumor formation and stem-like activity in colorectal cancer cells [138]. Extensive research has revealed that m6A regulators play a crucial role in regulating abnormal metabolism in tumor cells. In colorectal cancer under high-fat conditions, ALKBH5 and FTO downregulation increases m6A methylation on HK2 mRNA through IGF2BP2. These regulators further activate the class O of forkhead box transcription factors (FOXO) signaling pathway, accelerating glycolysis, proliferation, and tumor progression [139]. Molecular investigations indicate that ALKBH5 collaborates with METTL14/IGF2BPs to suppress tumor cell glycolysis by negatively regulating the Jumonji domain-containing protein 8 (JMJD8)/pyruvate kinase M2 (PKM2) signaling axis, slowing down colorectal cancer progression [140]. m6A modification of circRNAs is also critical for regulating the progression of gastric cancer. Knockdown of METTL14 reduces the circORC5 m6A levels, increasing circORC5 expression and inhibiting tumor progression through the miR-30c-2-3p/AKT1 substrate 1 (AKT1S1) axis [141].
Pancreatic cancer
Pancreatic cancer has a high global incidence and mortality rate and is often diagnosed at an advanced stage, due to nonspecific early symptoms [142,143,144]. Consequently, its prognosis is unfavorable, and it has limited treatment options. Surgery is most effective for localized cases but is often inapplicable due to late diagnosis [145, 146]. Despite therapeutic advancements such as molecular targeted therapy and immunotherapy, pancreatic cancer still has a low five-year survival rate of approximately 10% [147, 148]. Ongoing research focuses on understanding in-depth molecular mechanisms for early detection, innovative therapies, and personalized management [149,150,151,152]. Recent studies have highlighted the significant role of m6A regulators in the tumorigenesis and progression of pancreatic cancer [153, 154]. Elevated METTL14 expression significantly enhanced pancreatic cancer cell proliferation and migration by directly targeting downstream p53 effector related to PMP-22 (PERP) mRNA in an m6A-dependent way [155, 156].
The overexpressed YTHDF2 binds to m6A-modified yes-associated protein 1 (YAP) mRNA, inhibiting YAP expression and promoting the migration-proliferation dichotomy and epithelial-mesenchymal transition of pancreatic cancer cells [157]. ALKBH5 activates Period1 (PER1) gene through m6A demethylation and in an m6A-YTHDF2-dependent manner, reactivating the Ataxia telangiectasia mutated/checkpoint kinase 2/tumor protein 53/cell division cycle 25C signaling pathway and inhibiting the proliferation, migration, and invasiveness of pancreatic cancer cells [158]. ALKBH5 also promotes Wnt inhibitory factor 1 (WIF-1) transcription, suppressing Wnt signaling in pancreatic cancer cells and inhibiting cancer progression [159].
Under hypoxia, the significantly increased transcription and protein levels of ALKBH5 in pancreatic cancer enhance glycolysis and cell migration. ALKBH5 controls m6A modification of histone deacetylase type 4 (HDAC4) recognized by YTHDF2, thereby upregulating HDAC4 expression. Upregulated HDAC4 stabilizes hypoxia-inducible factor-1α (HIF1α) in hypoxic pancreatic cancer cells, creating a positive feedback loop that increases ALKBH5 expression [160]. Additionally, under glucose deprivation, miR-5586-5p induces the overexpression of YTHDF3, leading to m6A modification of DICER1 antisense RNA1 and promoting cell glycolysis, tumor growth, and metastasis in pancreatic cancer [161].
Clinical applications of m6A modification in gastrointestinal tract cancers
Many studies have elaborated on the significance of m6A regulators in gastrointestinal tract cancers, particularly focusing on their roles as diagnostic, prognostic indicators, and potential therapeutic targets. (Fig. 3) [162,163,164,165]. Increasing evidence links changes in m6A modification levels to clinical characteristics in various digestive tract tumors, providing new possibilities for non-invasive early diagnosis of malignant gastrointestinal tract cancers (Table 2) [61, 166, 167].
m6A modification holds promise in gastrointestinal tract cancers for clinical diagnosis, prognosis prediction, and treatment guidance. Kaplan–Meier survival and receiver operating characteristic curves confirmed the reliability of abnormal m6A levels as diagnostic indicators. Univariate and multivariate regression models support m6A as an independent prognostic indicator. Research on m6A regulation mechanisms in the progression of digestive tract tumors suggests that targeting abnormal m6A levels improves chemotherapy and radiotherapy resistance. Modulating m6A levels enhances the efficacy of immunotherapy, improving patient survival rates. Furthermore, targeted drug delivery systems achieve significant anticancer effects
m6A modification as diagnostic and prognostic indicators in gastrointestinal tract cancers
Circulating tumor cells (CTCs), which are cells shed from tumors into the bloodstream, offer a non-invasive method for diagnosing and monitoring cancer [168]. Elevated levels of m6A modification in CTCs compared to whole blood samples suggest its potential as a diagnostic marker for cancer, particularly in gastrointestinal tract cancers. For example, the level of m6A in peripheral blood RNA was significantly higher in patients with gastric cancer than in patients with benign gastric diseases and healthy controls. The study also assessed m6A levels in peripheral blood RNA as a non-invasive diagnostic biomarker for patients with gastric cancer using a receiver operating characteristic (ROC) curve analysis. ROC analysis revealed an area under the curve of 0.929, outperforming commonly used biomarkers such as carcinoembryonic antigen and carbohydrate antigen 199. This indicated that the m6A levels in peripheral blood RNA have high accuracy in diagnosing gastric cancer [169, 170]. In pancreatic cancer, m6A levels showed promise as a predictive and prognostic marker. Reduced ALKBH5 levels in pancreatic cancer tissues were identified through various technologies, and survival analysis indicated a significant association between ALKBH5 expression and poor prognosis in patients with pancreatic cancer [158, 171].
Furthermore, multiple m6A regulators, including METTL3 and ALKBH5, have been identified as independent prognostic indicators for gastrointestinal tract cancers through multivariate Cox regression analysis and Kaplan–Meier survival curves [172,173,174]. High expression of METTL3 in gastric cancer is associated with lymph node metastasis and advanced TNM stage. It serves as an adverse prognostic indicator for patient survival and recurrence. Multivariate Cox regression analysis has further demonstrated that METTL3 is an independent prognosis predictor in patients with gastric cancer. When combined with clinical risk score (TNM stage), METTL3 detection significantly improves prognostic accuracy, underlining its clinical significance in gastric cancer [175]. High expression of METTL3 may serve as an adverse prognostic indicator for patient survival and recurrence. Numerous studies have also analyzed large TCGA datasets to investigate the relationship between abnormal expression of m6A regulators, patient prognosis, and tumor staging in gastrointestinal tract cancers [176, 177]. Robust m6A-related prognostic models and risk-scoring systems have been developed using clinical follow-up data and public databases like TCGA, aiding in predicting disease progression and survival status. Moreover, downregulation of ALKBH5 is correlated with poor prognosis in colorectal cancer patients. Chen Rui's team detected a significant downregulation of ALKBH5 in 1,078 patients with colorectal cancer tissues compared to healthy controls in a ten-year follow-up cohort and the TCGA dataset. This downregulation was associated with poor prognosis in patients with colorectal cancer [140]. In addition, researchers have also developed robust m6A-related prognostic models and risk-scoring systems using clinical follow-up data and public databases such as TCGA. These models and scoring systems can individually evaluate the prognosis and survival of patients with gastrointestinal tract cancers based on their m6A regulator expression and other clinical characteristics, better-predicting disease progression and survival status and providing important references for clinical decision-making [172, 178,179,180,181].
Therapeutic potential of m6A modification in gastrointestinal tract cancers
Various m6A regulators participate in carcinogenic processes in gastrointestinal tract cancers, such as cell proliferation, migration, invasion, stem cell characteristics, regulation of immune responses, and resistance to radiotherapy and chemotherapy through interactions with up- or downstream targeted molecules [154, 182]. In the treatment of common digestive tract tumors, radiotherapy and chemotherapy are pivotal. However, their effectiveness is limited. Targeting m6A regulators and their downstream targets shows promise as a molecular targeted therapy for advanced gastrointestinal tract cancers.[183,184,185]. Approximately 50% of patients with gastrointestinal stromal tumors (GIST) develop resistance to imatinib treatment within two years. Recent studies have revealed increased m6A modification in imatinib-resistant GIST cells and tissues. METTL3 upregulation leads to imatinib resistance by enhancing the expression of multidrug resistance protein 1 (MRP1). This leads to a reduction in the intracellular concentration of imatinib, promoting imatinib resistance in gastrointestinal stromal tumor cells [186]. METTL3 is also an effective target for improving conventional treatment efficacy in patients with pancreatic cancer. Knocking down METTL3 expression in pancreatic cancer cell lines enhances their sensitivity to anticancer radiotherapy and chemotherapeutics such as gemcitabine, cisplatin, and 5-fluorouracil [187]. Furthermore, inhibiting the METTL3-mediated activation of frizzled10 in liver cancer cells contributes to lenvatinib resistance by regulating the downstream β-catenin/c-Jun/Mitogen-activated protein kinase kinase (MEK)/ERK axis [98]. These findings suggest the potential for METTL3-targeted therapeutic strategies and provide new directions for personalized treatment of patients with pancreatic cancer.
Besides being a well-known m6A writer, ALKBH5 also collaborates with METTL14/IGF2BPs to suppress tumor cell glycolysis by negatively regulating the JMJD8/PKM2 signaling axis, thereby slowing down colorectal cancer progression. In preclinical tumor models, an ALKBH5 mRNA delivery system restored ALKBH5 levels at the tumor site, suppressing colorectal cancer growth and offering a novel clinical target [140]. Moreover, studies have found that RNA-binding protein IGF2BP1 binds to apoptotic protease-activating factor 1 (APAF1)-binding lncRNA (ABL) and recognizes METTL3-mediated m6A modification on ABL, maintaining the stability of ABL. This interaction between IGF2BP1 and ABL leads to gastric cancer cell resistance to apoptosis induced by chemotherapy such as 5‐fluorouracil and paclitaxel [188]. ALKBH5 has emerged as a promising candidate for drug development with its significant tumor-suppressive functions and the ability to sensitize pancreatic cancer cells to chemotherapy. It mediates m6A demethylation of DNA damage-inducible transcript 4 (DDIT4) and inhibits the methylation of WIF-1. This upregulates DDIT4 antisense RNA1, activating the Wnt and the mTOR signaling pathways, impairing the chemosensitivity of pancreatic cancer cells to gemcitabine [159, 189].
Currently, several targeted drug delivery systems and clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated proteins systems have been successfully developed to regulate tumor m6A modification levels for anticancer purposes [190,191,192,193]. Engineered small extracellular vesicles were used to effectively deliver short interfering RNA targeting YTHDF1, leading to efficient depletion of YTHDF1 expression in gastric cancer tissues. This approach inhibited gastric cancer progression and metastasis by blocking the translation of frizzled7 and subsequently inactivating the Wnt/β-catenin pathway [194]. Furthermore, it also triggered a strong interferon-γ response, resulting in enhanced cytotoxic T lymphocyte response and tumor-associated macrophage-mediated phagocytosis [195]. These findings highlight the potential of targeting m6A regulators through RNA interference as a promising strategy for cancer treatment [196]. In addition, m6A modifications can also shape the immune landscape by influencing cytokine production, immune cell differentiation, and the inflammatory response, which are critical factors in cancer progression and response to therapy. Applying these insights to gastrointestinal cancers could reveal new mechanisms by which m6A modifications influence tumor immunity and help identify potential therapeutic targets for modulating the immune response to cancer [197].
However, while the current research on m6A modifications in gastrointestinal cancers has provided valuable insights, it is crucial to acknowledge the limitations and potential biases in recent research on m6A modifications in gastrointestinal tract cancers. One notable limitation is the relatively small sample sizes used in most clinical studies concerning the verification of clinical significance of targeting m6A modifications in gastrointestinal cancers. The majority of clinical studies of M6A in gastrointestinal tumors have an inadequate sample size, which may affect the reliability and generalizability of the findings. Additionally, the common use of different experimental models, such as gastrointestinal cancer cell lines versus animal models, may not accurately reflect in vivo biology and introduce variability and the difficulty of successful translation into a clinical setting for human patients. Moreover, there are differences among the methodologies used for the detection and analysis of the specific m6A modification in different studies. Overall, further studies with larger sample sizes, standardized methodologies, and comprehensive analyses are needed to fully understand the role of m6A modifications in these gastrointestinal tract cancers.
In summary, m6A modification has broad prospects in clinical diagnostics and treatment. Detecting m6A modification levels and implementing therapeutic strategies targeting m6A regulators and their downstream targets can enhance diagnostic accuracy and treatment outcomes in gastrointestinal tract cancers, ultimately improving clinical efficacy and patient outcomes.
Conclusion
Increasing evidence consistently links alterations in m6A regulatory proteins and global m6A modification patterns with the occurrence and progression of gastrointestinal tract cancers. Multiple large-scale cohorts confirm the correlations between m6A level, patient prognosis, and diagnostic reliability, presenting m6A levels as promising indicators for both prognosis and diagnosis in these cancers. Dysregulation of m6A modification impacts the pathogenesis of tumor progression by regulating biological processes such as tumor cell proliferation, invasion, and metastasis. Given the critical roles of m6A methylation in several types of gastrointestinal tract cancers, m6A modification holds promise as a potential therapeutic target. However, significant research gaps remain, particularly in understanding how m6A regulators interact with other epigenetic factors and contribute to cancer heterogeneity. The development of precise biomarkers for m6A modification and effective therapeutic strategies targeting m6A-related pathways also necessitates further investigation. Addressing these unresolved challenges is crucial for advancing our knowledge and translating it into clinical applications that significantly improve patient outcomes. Future research should focus on elucidating the intricate molecular mechanisms of m6A modifications in gastrointestinal tract cancers and developing targeted therapies that leverage this knowledge.
Data availability
Not applicable.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer statistics, 2021. CA Cancer J Clin 71:7–33. https://doi.org/10.3322/caac.21654
Bijlsma MF, Sadanandam A, Tan P, Vermeulen L (2017) Molecular subtypes in cancers of the gastrointestinal tract. Nat Rev Gastroenterol Hepatol 14:333–342. https://doi.org/10.1038/nrgastro.2017.33
Xin Z, Jiang S, Jiang P, Yan X, Fan C, Di S, Wu G, Yang Y, Reiter RJ, Ji G (2015) Melatonin as a treatment for gastrointestinal cancer: a review. J Pineal Res 58:375–387. https://doi.org/10.1111/jpi.12227
Stukalin I, Ahmed NS, Fundytus AM, Qian AS, Coward S, Kaplan GG, Hilsden RJ, Burak KW, Lee JK, Singh S, Ma C (2022) Trends and projections in national united states health care spending for gastrointestinal malignancies (1996–2030). Gastroenterology 162:1098-1110.e2. https://doi.org/10.1053/j.gastro.2021.12.244
Siegel R, DeSantis C, Virgo K, Stein K, Mariotto A, Smith T, Cooper D, Gansler T, Lerro C, Fedewa S, Lin C, Leach C, Cannady RS, Cho H, Scoppa S, Hachey M, Kirch R, Jemal A, Ward E (2012) Cancer treatment and survivorship statistics, 2012. CA Cancer J Clin 62:220–241. https://doi.org/10.3322/caac.21149
Ben-Aharon I, van Laarhoven HWM, Fontana E, Obermannova R, Nilsson M, Lordick F (2023) Early-onset cancer in the gastrointestinal tract is on the rise-evidence and implications. Cancer Discov 13:538–551. https://doi.org/10.1158/2159-8290.Cd-22-1038
Milette S, Sicklick JK, Lowy AM, Brodt P (2017) Molecular pathways: targeting the microenvironment of liver metastases. Clin Cancer Res 23:6390–6399. https://doi.org/10.1158/1078-0432.Ccr-15-1636
Calderaro J, Kather JN (2021) Artificial intelligence-based pathology for gastrointestinal and hepatobiliary cancers. Gut 70:1183–1193. https://doi.org/10.1136/gutjnl-2020-322880
Diplas BH, Ptashkin R, Chou JF, Sabwa S, Foote MB, Rousseau B, Argilés G, White JR, Stewart CM, Bolton K, Chalasani SB, Desai AM, Goldberg Z, Gu P, Li J, Shcherba M, Zervoudakis A, Cercek A, Yaeger R, Segal NH, Ilson DH, Ku GY, Zehir A, Capanu M, Janjigian YY, Diaz LA Jr, Maron SB (2023) Clinical importance of clonal hematopoiesis in metastatic gastrointestinal tract cancers. JAMA Netw Open 6:e2254221. https://doi.org/10.1001/jamanetworkopen.2022.54221
Alexandre L, Tsilegeridis-Legeris T, Lam S (2022) Clinical and endoscopic characteristics associated with post-endoscopy upper gastrointestinal cancers: a systematic review and meta-analysis. Gastroenterology 162:1123–1135. https://doi.org/10.1053/j.gastro.2021.12.270
Song WK, Wilson BC (2005) Endoscopic detection of early upper GI cancers. Best Pract Res Clin Gastroenterol 19:833–856. https://doi.org/10.1016/j.bpg.2005.04.006
Marc G, Lopes CV (2008) Endoscopic resection of superficial gastrointestinal tumors. World J Gastroenterol 14:4600–4606. https://doi.org/10.3748/wjg.14.4600
(2012) Abstracts of the 2011 World Molecular Imaging Congress. San Diego, California, USA. September 7–10, 2011. Mol Imaging Biol 14 Suppl 1:S4–996. https://doi.org/10.1007/s11307-012-0543-5
Vedeld HM, Goel A, Lind GE (2018) Epigenetic biomarkers in gastrointestinal cancers: the current state and clinical perspectives. Semin Cancer Biol 51:36–49. https://doi.org/10.1016/j.semcancer.2017.12.004
Kandimalla R, Xu J, Link A, Matsuyama T, Yamamura K, Parker MI, Uetake H, Balaguer F, Borazanci E, Tsai S, Evans D, Meltzer SJ, Baba H, Brand R, Von Hoff D, Li W, Goel A (2021) EpiPanGI Dx: a cell-free DNA methylation fingerprint for the early detection of gastrointestinal cancers. Clin Cancer Res 27:6135–6144. https://doi.org/10.1158/1078-0432.Ccr-21-1982
Kim BG, Li C, Qiao W, Mamura M, Kasprzak B, Anver M, Wolfraim L, Hong S, Mushinski E, Potter M, Kim SJ, Fu XY, Deng C, Letterio JJ (2006) Smad4 signalling in T cells is required for suppression of gastrointestinal cancer. Nature 441:1015–1019. https://doi.org/10.1038/nature04846
Grady WM, Yu M, Markowitz SD (2021) Epigenetic alterations in the gastrointestinal tract: current and emerging use for biomarkers of cancer. Gastroenterology 160:690–709. https://doi.org/10.1053/j.gastro.2020.09.058
Li XP, Qu J, Teng XQ, Zhuang HH, Dai YH, Yang Z, Qu Q (2023) The emerging role of super-enhancers as therapeutic targets in the digestive system tumors. Int J Biol Sci 19:1036–1048. https://doi.org/10.7150/ijbs.78535
Liu X, Wang S, Xia X, Chen Y, Zhou Y, Wu X, Zhang J, He S, Tan Y, Qiang F, Røe OD, Li G, Zhou J (2012) Synergistic role between p53 and JWA: prognostic and predictive biomarkers in gastric cancer. PLoS ONE 7:e52348. https://doi.org/10.1371/journal.pone.0052348
Dominissini D, Moshitch-Moshkovitz S, Salmon-Divon M, Amariglio N, Rechavi G (2013) Transcriptome-wide mapping of N(6)-methyladenosine by m(6)A-seq based on immunocapturing and massively parallel sequencing. Nat Protoc 8:176–189. https://doi.org/10.1038/nprot.2012.148
Boccaletto P, Stefaniak F, Ray A, Cappannini A, Mukherjee S, Purta E, Kurkowska M, Shirvanizadeh N, Destefanis E, Groza P, Avşar G, Romitelli A, Pir P, Dassi E, Conticello SG, Aguilo F, Bujnicki JM (2022) MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Res 50:D231-d235. https://doi.org/10.1093/nar/gkab1083
Ke S, Alemu EA, Mertens C, Gantman EC, Fak JJ, Mele A, Haripal B, Zucker-Scharff I, Moore MJ, Park CY, Vågbø CB, Kusśnierczyk A, Klungland A, Darnell JE Jr, Darnell RB (2015) A majority of m6A residues are in the last exons, allowing the potential for 3’ UTR regulation. Genes Dev 29:2037–2053. https://doi.org/10.1101/gad.269415.115
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, Sorek R, Rechavi G (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485:201–206. https://doi.org/10.1038/nature11112
Liu J, Huang T, Yao J, Zhao T, Zhang Y, Zhang R (2023) Epitranscriptomic subtyping, visualization, and denoising by global motif visualization. Nat Commun 14:5944. https://doi.org/10.1038/s41467-023-41653-4
Yang Y, Hsu PJ, Chen YS, Yang YG (2018) Dynamic transcriptomic m(6)A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res 28:616–624. https://doi.org/10.1038/s41422-018-0040-8
Meyer KD, Jaffrey SR (2014) The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat Rev Mol Cell Biol 15:313–326. https://doi.org/10.1038/nrm3785
Zhang W, Qian Y, Jia G (2021) The detection and functions of RNA modification m(6)A based on m(6)A writers and erasers. J Biol Chem 297:100973. https://doi.org/10.1016/j.jbc.2021.100973
Bi Z, Liu Y, Zhao Y, Yao Y, Wu R, Liu Q, Wang Y, Wang X (2019) A dynamic reversible RNA N(6) -methyladenosine modification: current status and perspectives. J Cell Physiol 234:7948–7956. https://doi.org/10.1002/jcp.28014
Huang J, Yin P (2018) Structural insights into N(6)-methyladenosine (m(6)A) modification in the transcriptome. Genomics Proteomics Bioinform 16:85–98. https://doi.org/10.1016/j.gpb.2018.03.001
Meyer KD, Jaffrey SR (2017) Rethinking m(6)A readers, writers, and erasers. Annu Rev Cell Dev Biol 33:319–342. https://doi.org/10.1146/annurev-cellbio-100616-060758
Xue C, Chu Q, Zheng Q, Jiang S, Bao Z, Su Y, Lu J, Li L (2022) Role of main RNA modifications in cancer: N(6)-methyladenosine, 5-methylcytosine, and pseudouridine. Signal Transduct Target Ther 7:142. https://doi.org/10.1038/s41392-022-01003-0
Kisan A, Chhabra R (2023) Modulation of gene expression by YTH domain family (YTHDF) proteins in human physiology and pathology. J Cell Physiol 238:5–31. https://doi.org/10.1002/jcp.30907
Alarcón CR, Goodarzi H, Lee H, Liu X, Tavazoie S, Tavazoie SF (2015) HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell 162:1299–1308. https://doi.org/10.1016/j.cell.2015.08.011
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, Zhao BS, Mesquita A, Liu C, Yuan CL, Hu YC, Hüttelmaier S, Skibbe JR, Su R, Deng X, Dong L, Sun M, Li C, Nachtergaele S, Wang Y, Hu C, Ferchen K, Greis KD, Jiang X, Wei M, Qu L, Guan JL, He C, Yang J, Chen J (2018) Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol 20:285–295. https://doi.org/10.1038/s41556-018-0045-z
Zhu S, Wang JZ, Chen D, He YT, Meng N, Chen M, Lu RX, Chen XH, Zhang XL, Yan GR (2020) An oncopeptide regulates m(6)A recognition by the m(6)A reader IGF2BP1 and tumorigenesis. Nat Commun 11:1685. https://doi.org/10.1038/s41467-020-15403-9
Müller S, Bley N, Busch B, Glaß M, Lederer M, Misiak C, Fuchs T, Wedler A, Haase J, Bertoldo JB, Michl P, Hüttelmaier S (2020) The oncofetal RNA-binding protein IGF2BP1 is a druggable, post-transcriptional super-enhancer of E2F-driven gene expression in cancer. Nucleic Acids Res 48:8576–8590. https://doi.org/10.1093/nar/gkaa653
Bell JL, Wächter K, Mühleck B, Pazaitis N, Köhn M, Lederer M, Hüttelmaier S (2013) Insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs): post-transcriptional drivers of cancer progression? Cell Mol Life Sci 70:2657–2675. https://doi.org/10.1007/s00018-012-1186-z
Lederer M, Bley N, Schleifer C, Hüttelmaier S (2014) The role of the oncofetal IGF2 mRNA-binding protein 3 (IGF2BP3) in cancer. Semin Cancer Biol 29:3–12. https://doi.org/10.1016/j.semcancer.2014.07.006
Pereira B, Billaud M, Almeida R (2017) RNA-binding proteins in cancer: old players and new actors. Trends Cancer 3:506–528. https://doi.org/10.1016/j.trecan.2017.05.003
Li L, Miao H, Chang Y, Yao H, Zhao Y, Wu F, Song X (2021) Multidimensional crosstalk between RNA-binding proteins and noncoding RNAs in cancer biology. Semin Cancer Biol 75:84–96. https://doi.org/10.1016/j.semcancer.2021.03.007
Haussmann IU, Bodi Z, Sanchez-Moran E, Mongan NP, Archer N, Fray RG, Soller M (2016) m(6)A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature 540:301–304. https://doi.org/10.1038/nature20577
Ji X, Wang Z, Sun W, Zhang H (2023) The emerging role of m6A modification in endocrine cancer. Cancers (Basel). https://doi.org/10.3390/cancers15041033
Lokody I (2014) Gene regulation: RNA methylation regulates the circadian clock. Nat Rev Genet 15:3. https://doi.org/10.1038/nrg3638
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, Weng X, Chen K, Shi H, He C (2015) N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161:1388–1399. https://doi.org/10.1016/j.cell.2015.05.014
Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ, Liu C, He C (2017) YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res 27:315–328. https://doi.org/10.1038/cr.2017.15
Lee Y, Choe J, Park OH, Kim YK (2020) Molecular mechanisms driving mRNA degradation by m(6)A modification. Trends Genet 36:177–188. https://doi.org/10.1016/j.tig.2019.12.007
Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, Sun HY, Li A, Ping XL, Lai WY, Wang X, Ma HL, Huang CM, Yang Y, Huang N, Jiang GB, Wang HL, Zhou Q, Wang XJ, Zhao YL, Yang YG (2016) Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Mol Cell 61:507–519. https://doi.org/10.1016/j.molcel.2016.01.012
Wang X, Xie H, Ying Y, Chen D, Li J (2020) Roles of N(6) -methyladenosine (m(6) A) RNA modifications in urological cancers. J Cell Mol Med 24:10302–10310. https://doi.org/10.1111/jcmm.15750
Zhao Z, Meng J, Su R, Zhang J, Chen J, Ma X, Xia Q (2020) Epitranscriptomics in liver disease: basic concepts and therapeutic potential. J Hepatol 73:664–679. https://doi.org/10.1016/j.jhep.2020.04.009
Fustin JM, Doi M, Yamaguchi Y, Hida H, Nishimura S, Yoshida M, Isagawa T, Morioka MS, Kakeya H, Manabe I, Okamura H (2013) RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155:793–806. https://doi.org/10.1016/j.cell.2013.10.026
Tong J, Flavell RA, Li HB (2018) RNA m(6)A modification and its function in diseases. Front Med 12:481–489. https://doi.org/10.1007/s11684-018-0654-8
Wei W, Ji X, Guo X, Ji S (2017) Regulatory role of N(6) -methyladenosine (m(6) A) methylation in RNA processing and human diseases. J Cell Biochem 118:2534–2543. https://doi.org/10.1002/jcb.25967
Yang C, Hu Y, Zhou B, Bao Y, Li Z, Gong C, Yang H, Wang S, Xiao Y (2020) The role of m(6)A modification in physiology and disease. Cell Death Dis 11:960. https://doi.org/10.1038/s41419-020-03143-z
Zhu TY, Hong LL, Ling ZQ (2023) Oncofetal protein IGF2BPs in human cancer: functions, mechanisms and therapeutic potential. Biomark Res 11:62. https://doi.org/10.1186/s40364-023-00499-0
Li X, Ma B, Zhang W, Song Z, Zhang X, Liao M, Li X, Zhao X, Du M, Yu J, He S, Yan H (2023) The essential role of N6-methyladenosine RNA methylation in complex eye diseases. Genes Dis 10:505–520. https://doi.org/10.1016/j.gendis.2022.05.008
Li J, Wang F, Liu Y, Wang H, Ni B (2021) N(6)-methyladenosine (m(6)A) in pancreatic cancer: regulatory mechanisms and future direction. Int J Biol Sci 17:2323–2335. https://doi.org/10.7150/ijbs.60115
Shen C, Xuan B, Yan T, Ma Y, Xu P, Tian X, Zhang X, Cao Y, Ma D, Zhu X, Zhang Y, Fang JY, Chen H, Hong J (2020) m(6)A-dependent glycolysis enhances colorectal cancer progression. Mol Cancer 19:72. https://doi.org/10.1186/s12943-020-01190-w
Yue SW, Liu HL, Su HF, Luo C, Liang HF, Zhang BX, Zhang W (2023) m6A-regulated tumor glycolysis: new advances in epigenetics and metabolism. Mol Cancer 22:137. https://doi.org/10.1186/s12943-023-01841-8
Ma E, Li J, Shen C, Gu Y, Zhang X, Li L, Zhao J, Wang Z (2023) The m(6)A-related gene signature stratifies poor prognosis patients and characterizes immunosuppressive microenvironment in hepatocellular carcinoma. Front Immunol 14:1227593. https://doi.org/10.3389/fimmu.2023.1227593
Sun L, Chen X, Zhu S, Wang J, Diao S, Liu J, Xu J, Li X, Sun Y, Huang C, Meng X, Lv X, Li J (2024) Decoding m(6)A mRNA methylation by reader proteins in liver diseases. Genes Dis 11:711–726. https://doi.org/10.1016/j.gendis.2023.02.054
Ding SQ, Zhang XP, Pei JP, Bai X, Ma JJ, Zhang CD, Dai DQ (2023) Role of N6-methyladenosine RNA modification in gastric cancer. Cell Death Discov 9:241. https://doi.org/10.1038/s41420-023-01485-z
Xu D, Shao W, Jiang Y, Wang X, Liu Y, Liu X (2017) FTO expression is associated with the occurrence of gastric cancer and prognosis. Oncol Rep 38:2285–2292. https://doi.org/10.3892/or.2017.5904
Zhou Y, Yin Z, Hou B, Yu M, Chen R, Jin H, Jian Z (2019) Expression profiles and prognostic significance of RNA N6-methyladenosine-related genes in patients with hepatocellular carcinoma: evidence from independent datasets. Cancer Manag Res 11:3921–3931. https://doi.org/10.2147/cmar.S191565
Li Y, Zheng D, Wang F, Xu Y, Yu H, Zhang H (2019) Expression of demethylase genes, FTO and ALKBH1, is associated with prognosis of gastric cancer. Dig Dis Sci 64:1503–1513. https://doi.org/10.1007/s10620-018-5452-2
Zhao X, Chen Y, Mao Q, Jiang X, Jiang W, Chen J, Xu W, Zhong L, Sun X (2018) Overexpression of YTHDF1 is associated with poor prognosis in patients with hepatocellular carcinoma. Cancer Biomark 21:859–868. https://doi.org/10.3233/cbm-170791
Zhang B, Chen Z, Tao B, Yi C, Lin Z, Li Y, Shao W, Lin J, Chen J (2021) m(6)A target microRNAs in serum for cancer detection. Mol Cancer 20:170. https://doi.org/10.1186/s12943-021-01477-6
Hu BB, Wang XY, Gu XY, Zou C, Gao ZJ, Zhang H, Fan Y (2019) N(6)-methyladenosine (m(6)A) RNA modification in gastrointestinal tract cancers: roles, mechanisms, and applications. Mol Cancer 18:178. https://doi.org/10.1186/s12943-019-1099-7
Li J, Liang L, Yang Y, Li X, Ma Y (2021) N(6)-methyladenosine as a biological and clinical determinant in colorectal cancer: progression and future direction. Theranostics 11:2581–2593. https://doi.org/10.7150/thno.52366
Chen LJ, Liu HY, Xiao ZY, Qiu T, Zhang D, Zhang LJ, Han FY, Chen GJ, Xu XM, Zhu JH, Ding YQ, Wang SY, Ye YP, Jiao HL (2023) IGF2BP3 promotes the progression of colorectal cancer and mediates cetuximab resistance by stabilizing EGFR mRNA in an m(6)A-dependent manner. Cell Death Dis 14:581. https://doi.org/10.1038/s41419-023-06099-y
Maity A, Das B (2016) N6-methyladenosine modification in mRNA: machinery, function and implications for health and diseases. FEBS J 283:1607–1630. https://doi.org/10.1111/febs.13614
Lin S, Choe J, Du P, Triboulet R, Gregory RI (2016) The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Mol Cell 62:335–345. https://doi.org/10.1016/j.molcel.2016.03.021
(2016) The RNA methyltransferase METTL3 promotes oncogene translation. Cancer Discov 6:572. https://doi.org/10.1158/2159-8290.Cd-rw2016-083
Su R, Dong L, Li Y, Gao M, Han L, Wunderlich M, Deng X, Li H, Huang Y, Gao L, Li C, Zhao Z, Robinson S, Tan B, Qing Y, Qin X, Prince E, Xie J, Qin H, Li W, Shen C, Sun J, Kulkarni P, Weng H, Huang H, Chen Z, Zhang B, Wu X, Olsen MJ, Müschen M, Marcucci G, Salgia R, Li L, Fathi AT, Li Z, Mulloy JC, Wei M, Horne D, Chen J (2020) Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell 38:79-96.e11. https://doi.org/10.1016/j.ccell.2020.04.017
Li HB, Tong J, Zhu S, Batista PJ, Duffy EE, Zhao J, Bailis W, Cao G, Kroehling L, Chen Y, Wang G, Broughton JP, Chen YG, Kluger Y, Simon MD, Chang HY, Yin Z, Flavell RA (2017) m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature 548:338–342. https://doi.org/10.1038/nature23450
He L, Li H, Wu A, Peng Y, Shu G, Yin G (2019) Functions of N6-methyladenosine and its role in cancer. Mol Cancer 18:176. https://doi.org/10.1186/s12943-019-1109-9
Chen XY, Zhang J, Zhu JS (2019) The role of m(6)A RNA methylation in human cancer. Mol Cancer 18:103. https://doi.org/10.1186/s12943-019-1033-z
Wang T, Kong S, Tao M, Ju S (2020) The potential role of RNA N6-methyladenosine in Cancer progression. Mol Cancer 19:88. https://doi.org/10.1186/s12943-020-01204-7
Zaccara S, Jaffrey SR (2020) A unified model for the function of YTHDF proteins in regulating m(6)A-modified mRNA. Cell 181:1582-1595.e18. https://doi.org/10.1016/j.cell.2020.05.012
Chen L, Gao Y, Xu S, Yuan J, Wang M, Li T, Gong J (2023) N6-methyladenosine reader YTHDF family in biological processes: structures, roles, and mechanisms. Front Immunol 14:1162607. https://doi.org/10.3389/fimmu.2023.1162607
Yao J, Song Y, Yu X, Lin Z (2023) Interaction between N(6)-methyladenosine modification and the tumor microenvironment in colorectal cancer. Mol Med 29:129. https://doi.org/10.1186/s10020-023-00726-2
Delaunay S, Helm M, Frye M (2023) RNA modifications in physiology and disease: towards clinical applications. Nat Rev Genet. https://doi.org/10.1038/s41576-023-00645-2
Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR (2019) A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol 16:589–604. https://doi.org/10.1038/s41575-019-0186-y
Altekruse SF, McGlynn KA, Reichman ME (2009) Hepatocellular carcinoma incidence, mortality, and survival trends in the United States from 1975 to 2005. J Clin Oncol 27:1485–1491. https://doi.org/10.1200/jco.2008.20.7753
El-Serag HB (2004) Hepatocellular carcinoma: recent trends in the United States. Gastroenterology 127:S27-34. https://doi.org/10.1053/j.gastro.2004.09.013
Rich NE, Yopp AC, Singal AG, Murphy CC (2020) Hepatocellular carcinoma incidence is decreasing among younger adults in the United States. Clin Gastroenterol Hepatol 18:242-248.e5. https://doi.org/10.1016/j.cgh.2019.04.043
Altekruse SF, Henley SJ, Cucinelli JE, McGlynn KA (2014) Changing hepatocellular carcinoma incidence and liver cancer mortality rates in the United States. Am J Gastroenterol 109:542–553. https://doi.org/10.1038/ajg.2014.11
Villanueva A (2019) Hepatocellular carcinoma. N Engl J Med 380:1450–1462. https://doi.org/10.1056/NEJMra1713263
Lim H, Ramjeesingh R, Liu D, Tam VC, Knox JJ, Card PB, Meyers BM (2021) Optimizing survival and the changing landscape of targeted therapy for intermediate and advanced hepatocellular carcinoma: a systematic review. J Natl Cancer Inst 113:123–136. https://doi.org/10.1093/jnci/djaa119
Liu Z, Lin Y, Zhang J, Zhang Y, Li Y, Liu Z, Li Q, Luo M, Liang R, Ye J (2019) Molecular targeted and immune checkpoint therapy for advanced hepatocellular carcinoma. J Exp Clin Cancer Res 38:447. https://doi.org/10.1186/s13046-019-1412-8
Zhang W, Tong S, Hu B, Wan T, Tang H, Zhao F, Jiao T, Li J, Zhang Z, Cai J, Ye H, Wang Z, Chen S, Wang Y, Li X, Wang F, Cao J, Tian L, Zhao X, Chen M, Wang H, Cai S, Hu M, Bai Y, Lu S (2023) Lenvatinib plus anti-PD-1 antibodies as conversion therapy for patients with unresectable intermediate-advanced hepatocellular carcinoma: a single-arm, phase II trial. J Immunother Cancer. https://doi.org/10.1136/jitc-2023-007366
El-Khoueiry AB, Sangro B, Yau T, Crocenzi TS, Kudo M, Hsu C, Kim TY, Choo SP, Trojan J, Welling THR, Meyer T, Kang YK, Yeo W, Chopra A, Anderson J, Dela Cruz C, Lang L, Neely J, Tang H, Dastani HB, Melero I (2017) Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet 389:2492–2502. https://doi.org/10.1016/s0140-6736(17)31046-2
Yau T, Park JW, Finn RS, Cheng AL, Mathurin P, Edeline J, Kudo M, Harding JJ, Merle P, Rosmorduc O, Wyrwicz L, Schott E, Choo SP, Kelley RK, Sieghart W, Assenat E, Zaucha R, Furuse J, Abou-Alfa GK, El-Khoueiry AB, Melero I, Begic D, Chen G, Neely J, Wisniewski T, Tschaika M, Sangro B (2022) Nivolumab versus sorafenib in advanced hepatocellular carcinoma (CheckMate 459): a randomised, multicentre, open-label, phase 3 trial. Lancet Oncol 23:77–90. https://doi.org/10.1016/s1470-2045(21)00604-5
Yau T, Hsu C, Kim TY, Choo SP, Kang YK, Hou MM, Numata K, Yeo W, Chopra A, Ikeda M, Kuromatsu R, Moriguchi M, Chao Y, Zhao H, Anderson J, Cruz CD, Kudo M (2019) Nivolumab in advanced hepatocellular carcinoma: Sorafenib-experienced Asian cohort analysis. J Hepatol 71:543–552. https://doi.org/10.1016/j.jhep.2019.05.014
Xiong C, Pan G, Wang H, Meng G, Yan L, Li R, Yan Y, Yang Y, Zhang X, Yang C, Dong Z, Li T (2023) Construction of an anoikis-related prognostic signature to predict immunotherapeutic response and prognosis in hepatocellular carcinoma. J Cancer Res Clin Oncol. https://doi.org/10.1007/s00432-023-05428-0
Llovet JM, Montal R, Sia D, Finn RS (2018) Molecular therapies and precision medicine for hepatocellular carcinoma. Nat Rev Clin Oncol 15:599–616. https://doi.org/10.1038/s41571-018-0073-4
Faivre S, Rimassa L, Finn RS (2020) Molecular therapies for HCC: looking outside the box. J Hepatol 72:342–352. https://doi.org/10.1016/j.jhep.2019.09.010
Chen M, Wong CM (2020) The emerging roles of N6-methyladenosine (m6A) deregulation in liver carcinogenesis. Mol Cancer 19:44. https://doi.org/10.1186/s12943-020-01172-y
Wang J, Yu H, Dong W, Zhang C, Hu M, Ma W, Jiang X, Li H, Yang P, Xiang D (2023) N6-methyladenosine-mediated up-regulation of FZD10 regulates liver cancer stem cells’ properties and lenvatinib resistance through WNT/β-catenin and hippo signaling pathways. Gastroenterology 164:990–1005. https://doi.org/10.1053/j.gastro.2023.01.041
Chen M, Wei L, Law CT, Tsang FH, Shen J, Cheng CL, Tsang LH, Ho DW, Chiu DK, Lee JM, Wong CC, Ng IO, Wong CM (2018) RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology 67:2254–2270. https://doi.org/10.1002/hep.29683
Chen Y, Peng C, Chen J, Chen D, Yang B, He B, Hu W, Zhang Y, Liu H, Dai L, Xie H, Zhou L, Wu J, Zheng S (2019) WTAP facilitates progression of hepatocellular carcinoma via m6A-HuR-dependent epigenetic silencing of ETS1. Mol Cancer 18:127. https://doi.org/10.1186/s12943-019-1053-8
Ma JZ, Yang F, Zhou CC, Liu F, Yuan JH, Wang F, Wang TT, Xu QG, Zhou WP, Sun SH (2017) METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6) -methyladenosine-dependent primary MicroRNA processing. Hepatology 65:529–543. https://doi.org/10.1002/hep.28885
Luo X, Cao M, Gao F, He X (2021) YTHDF1 promotes hepatocellular carcinoma progression via activating PI3K/AKT/mTOR signaling pathway and inducing epithelial-mesenchymal transition. Exp Hematol Oncol 10:35. https://doi.org/10.1186/s40164-021-00227-0
Yang Z, Li J, Feng G, Gao S, Wang Y, Zhang S, Liu Y, Ye L, Li Y, Zhang X (2017) MicroRNA-145 modulates N(6)-methyladenosine levels by targeting the 3’-untranslated mRNA region of the N(6)-methyladenosine binding YTH domain family 2 protein. J Biol Chem 292:3614–3623. https://doi.org/10.1074/jbc.M116.749689
Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F (2020) Gastric cancer. Lancet 396:635–648. https://doi.org/10.1016/s0140-6736(20)31288-5
Thrift AP, El-Serag HB (2020) Burden of gastric cancer. Clin Gastroenterol Hepatol 18:534–542. https://doi.org/10.1016/j.cgh.2019.07.045
Crew KD, Neugut AI (2006) Epidemiology of gastric cancer. World J Gastroenterol 12:354–362. https://doi.org/10.3748/wjg.v12.i3.354
Hohenberger P, Gretschel S (2003) Gastric cancer. Lancet 362:305–315. https://doi.org/10.1016/s0140-6736(03)13975-x
Tan Y, Wei Z, Liu K, Qin Y, Hui W (2023) Lifestyle habits and gastric cancer in an East Asian population: a Mendelian randomization study. Front Oncol 13:1224753. https://doi.org/10.3389/fonc.2023.1224753
Bai X, Li X, Ding S, Dai D (2023) Adherence to the Mediterranean diet and risk of gastric cancer: a systematic review and meta-analysis. Nutrients. https://doi.org/10.3390/nu15173826
Rugge M, Genta RM, Malfertheiner P, Graham DY (2023) Steps forward in understanding gastric cancer risk. Gut 72:1802–1803. https://doi.org/10.1136/gutjnl-2022-328514
Chia NY, Tan P (2016) Molecular classification of gastric cancer. Ann Oncol 27:763–769. https://doi.org/10.1093/annonc/mdw040
Ichikawa H, Nagahashi M, Shimada Y, Hanyu T, Ishikawa T, Kameyama H, Kobayashi T, Sakata J, Yabusaki H, Nakagawa S, Sato N, Hirata Y, Kitagawa Y, Tanahashi T, Yoshida K, Nakanishi R, Oki E, Vuzman D, Lyle S, Takabe K, Ling Y, Okuda S, Akazawa K, Wakai T (2017) Actionable gene-based classification toward precision medicine in gastric cancer. Genome Med 9:93. https://doi.org/10.1186/s13073-017-0484-3
Alessandrini L, Manchi M, De Re V, Dolcetti R, Canzonieri V (2018) Proposed molecular and miRNA classification of gastric cancer. Int J Mol Sci. https://doi.org/10.3390/ijms19061683
Hartgrink HH, Jansen EP, van Grieken NC, van de Velde CJ (2009) Gastric cancer. Lancet 374:477–490. https://doi.org/10.1016/s0140-6736(09)60617-6
Milewski PJ, Bancewicz J (1989) Improving the results of treating gastric cancer. BMJ 299:278–279. https://doi.org/10.1136/bmj.299.6694.278
Thrumurthy SG, Chaudry MA, Hochhauser D, Mughal M (2013) The diagnosis and management of gastric cancer. BMJ 347:f6367. https://doi.org/10.1136/bmj.f6367
Field K, Michael M, Leong T (2008) Locally advanced and metastatic gastric cancer: current management and new treatment developments. Drugs 68:299–317. https://doi.org/10.2165/00003495-200868030-00004
Goldenring JR (2017) The AGA/funderburg award in gastric cancer: twenty-five years of advances in gastric cancer research. Gastroenterology 152:1262–1266. https://doi.org/10.1053/j.gastro.2017.03.010
Holdstock G, Bruce S (1981) Endoscopy and gastric cancer. Gut 22:673–676. https://doi.org/10.1136/gut.22.8.673
Yue B, Song C, Yang L, Cui R, Cheng X, Zhang Z, Zhao G (2019) METTL3-mediated N6-methyladenosine modification is critical for epithelial-mesenchymal transition and metastasis of gastric cancer. Mol Cancer 18:142. https://doi.org/10.1186/s12943-019-1065-4
Song C, Zhou C (2021) HOXA10 mediates epithelial-mesenchymal transition to promote gastric cancer metastasis partly via modulation of TGFB2/Smad/METTL3 signaling axis. J Exp Clin Cancer Res 40:62. https://doi.org/10.1186/s13046-021-01859-0
Liu T, Yang S, Sui J, Xu SY, Cheng YP, Shen B, Zhang Y, Zhang XM, Yin LH, Pu YP, Liang GY (2020) Dysregulated N6-methyladenosine methylation writer METTL3 contributes to the proliferation and migration of gastric cancer. J Cell Physiol 235:548–562. https://doi.org/10.1002/jcp.28994
Lin S, Liu J, Jiang W, Wang P, Sun C, Wang X, Chen Y, Wang H (2019) METTL3 promotes the proliferation and mobility of gastric cancer cells. Open Med (Wars) 14:25–31. https://doi.org/10.1515/med-2019-0005
Zhang C, Zhang M, Ge S, Huang W, Lin X, Gao J, Gong J, Shen L (2019) Reduced m6A modification predicts malignant phenotypes and augmented Wnt/PI3K-Akt signaling in gastric cancer. Cancer Med 8:4766–4781. https://doi.org/10.1002/cam4.2360
Hu Y, Gong C, Li Z, Liu J, Chen Y, Huang Y, Luo Q, Wang S, Hou Y, Yang S, Xiao Y (2022) Demethylase ALKBH5 suppresses invasion of gastric cancer via PKMYT1 m6A modification. Mol Cancer 21:34. https://doi.org/10.1186/s12943-022-01522-y
Biller LH, Schrag D (2021) Diagnosis and treatment of metastatic colorectal cancer: a review. JAMA 325:669–685. https://doi.org/10.1001/jama.2021.0106
Yue Q, Zhang Y, Wang F, Cao F, Bai J, Duan X (2022) Characterization of m6A methylation modification patterns in colorectal cancer determines prognosis and tumor microenvironment infiltration. J Immunol Res 2022:8766735. https://doi.org/10.1155/2022/8766735
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Mármol I, Sánchez-de-Diego C, Pradilla Dieste A, Cerrada E, Rodriguez Yoldi MJ (2017) Colorectal carcinoma: a general overview and future perspectives in colorectal cancer. Int J Mol Sci. https://doi.org/10.3390/ijms18010197
Wong SH, Yu J (2019) Gut microbiota in colorectal cancer: mechanisms of action and clinical applications. Nat Rev Gastroenterol Hepatol 16:690–704. https://doi.org/10.1038/s41575-019-0209-8
Midgley R, Kerr D (1999) Colorectal cancer. Lancet 353:391–399. https://doi.org/10.1016/s0140-6736(98)07127-x
Saeed M, Shoaib A, Kandimalla R, Javed S, Almatroudi A, Gupta R, Aqil F (2022) Microbe-based therapies for colorectal cancer: advantages and limitations. Semin Cancer Biol 86:652–665. https://doi.org/10.1016/j.semcancer.2021.05.018
Li J, Ma X, Chakravarti D, Shalapour S, DePinho RA (2021) Genetic and biological hallmarks of colorectal cancer. Genes Dev 35:787–820. https://doi.org/10.1101/gad.348226.120
Zygulska AL, Pierzchalski P (2022) Novel diagnostic biomarkers in colorectal cancer. Int J Mol Sci. https://doi.org/10.3390/ijms23020852
Bhullar DS, Barriuso J, Mullamitha S, Saunders MP, O’Dwyer ST, Aziz O (2019) Biomarker concordance between primary colorectal cancer and its metastases. EBioMedicine 40:363–374. https://doi.org/10.1016/j.ebiom.2019.01.050
Li T, Hu PS, Zuo Z, Lin JF, Li X, Wu QN, Chen ZH, Zeng ZL, Wang F, Zheng J, Chen D, Li B, Kang TB, Xie D, Lin D, Ju HQ, Xu RH (2019) METTL3 facilitates tumor progression via an m(6)A-IGF2BP2-dependent mechanism in colorectal carcinoma. Mol Cancer 18:112. https://doi.org/10.1186/s12943-019-1038-7
Deng R, Cheng Y, Ye S, Zhang J, Huang R, Li P, Liu H, Deng Q, Wu X, Lan P, Deng Y (2019) m(6)A methyltransferase METTL3 suppresses colorectal cancer proliferation and migration through p38/ERK pathways. Onco Targets Ther 12:4391–4402. https://doi.org/10.2147/ott.S201052
Bai Y, Yang C, Wu R, Huang L, Song S, Li W, Yan P, Lin C, Li D, Zhang Y (2019) YTHDF1 regulates tumorigenicity and cancer stem cell-like activity in human colorectal carcinoma. Front Oncol 9:332. https://doi.org/10.3389/fonc.2019.00332
Ye M, Chen J, Lu F, Zhao M, Wu S, Hu C, Yu P, Kan J, Bai J, Tian Y, Tang Q (2023) Down-regulated FTO and ALKBH5 co-operatively activates FOXO signaling through m6A methylation modification in HK2 mRNA mediated by IGF2BP2 to enhance glycolysis in colorectal cancer. Cell Biosci 13:148. https://doi.org/10.1186/s13578-023-01100-9
Wu S, Yun J, Tang W, Familiari G, Relucenti M, Wu J, Li X, Chen H, Chen R (2023) Therapeutic m(6)A eraser ALKBH5 mRNA-loaded exosome-liposome hybrid nanoparticles inhibit progression of colorectal cancer in preclinical tumor models. ACS Nano 17:11838–11854. https://doi.org/10.1021/acsnano.3c03050
Fan HN, Chen ZY, Chen XY, Chen M, Yi YC, Zhu JS, Zhang J (2022) METTL14-mediated m(6)A modification of circORC5 suppresses gastric cancer progression by regulating miR-30c-2-3p/AKT1S1 axis. Mol Cancer 21:51. https://doi.org/10.1186/s12943-022-01521-z
Klein AP (2021) Pancreatic cancer epidemiology: understanding the role of lifestyle and inherited risk factors. Nat Rev Gastroenterol Hepatol 18:493–502. https://doi.org/10.1038/s41575-021-00457-x
Vincent A, Herman J, Schulick R, Hruban RH, Goggins M (2011) Pancreatic cancer. Lancet 378:607–620. https://doi.org/10.1016/s0140-6736(10)62307-0
Saluja A, Maitra A (2019) Pancreatitis and pancreatic cancer. Gastroenterology 156:1937–1940. https://doi.org/10.1053/j.gastro.2019.03.050
Sabater L, Muñoz E, Roselló S, Dorcaratto D, Garcés-Albir M, Huerta M, Roda D, Gómez-Mateo MC, Ferrández-Izquierdo A, Darder A, Cervantes A (2018) Borderline resectable pancreatic cancer. Challenges and controversies. Cancer Treat Rev 68:124–135. https://doi.org/10.1016/j.ctrv.2018.06.006
DiMagno EP (1999) Pancreatic cancer: clinical presentation, pitfalls and early clues. Ann Oncol 10(Suppl 4):140–142
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Wang Y, Yang G, You L, Yang J, Feng M, Qiu J, Zhao F, Liu Y, Cao Z, Zheng L, Zhang T, Zhao Y (2019) Role of the microbiome in occurrence, development and treatment of pancreatic cancer. Mol Cancer 18:173. https://doi.org/10.1186/s12943-019-1103-2
Tempero MA (2019) NCCN guidelines updates: pancreatic cancer. J Natl Compr Canc Netw 17:603–605. https://doi.org/10.6004/jnccn.2019.5007
Zhao Y, Tang J, Jiang K, Liu SY, Aicher A, Heeschen C (2023) Liquid biopsy in pancreatic cancer—current perspective and future outlook. Biochim Biophys Acta Rev Cancer 1878:188868. https://doi.org/10.1016/j.bbcan.2023.188868
Stoffel EM, Brand RE, Goggins M (2023) Pancreatic cancer: changing epidemiology and new approaches to risk assessment, early detection, and prevention. Gastroenterology 164:752–765. https://doi.org/10.1053/j.gastro.2023.02.012
Goldberg R (2022) Toward precision pancreatic cancer care. J Natl Compr Canc Netw 20:547–548. https://doi.org/10.6004/jnccn.2022.7019
Ye Y, Feng W, Zhang J, Zhu K, Huang X, Pan L, Su J, Zheng Y, Li R, Deng S, Bai R, Zhuang L, Wei L, Deng J, Li M, Chen R, Lin D, Zuo Z, Zheng J (2021) Genome-wide identification and characterization of circular RNA m(6)A modification in pancreatic cancer. Genome Med 13:183. https://doi.org/10.1186/s13073-021-01002-w
Guo Y, Wang R, Li J, Song Y, Min J, Zhao T, Hua L, Shi J, Zhang C, Ma P, Yang C, Zhu L, Gan D, Li S, Liu X, Su H (2021) Comprehensive analysis of m6A RNA methylation regulators and the immune microenvironment to aid immunotherapy in pancreatic cancer. Front Immunol 12:769425. https://doi.org/10.3389/fimmu.2021.769425
Wang M, Liu J, Zhao Y, He R, Xu X, Guo X, Li X, Xu S, Miao J, Guo J, Zhang H, Gong J, Zhu F, Tian R, Shi C, Peng F, Feng Y, Yu S, Xie Y, Jiang J, Li M, Wei W, He C, Qin R (2020) Upregulation of METTL14 mediates the elevation of PERP mRNA N(6) adenosine methylation promoting the growth and metastasis of pancreatic cancer. Mol Cancer 19:130. https://doi.org/10.1186/s12943-020-01249-8
Wang N, Yao F, Liu D, Jiang H, Xia X, Xiong S (2022) RNA N6-methyladenosine in nonocular and ocular disease. J Cell Physiol 237:1686–1710. https://doi.org/10.1002/jcp.30652
Chen J, Sun Y, Xu X, Wang D, He J, Zhou H, Lu Y, Zeng J, Du F, Gong A, Xu M (2017) YTH domain family 2 orchestrates epithelial-mesenchymal transition/proliferation dichotomy in pancreatic cancer cells. Cell Cycle 16:2259–2271. https://doi.org/10.1080/15384101.2017.1380125
Guo X, Li K, Jiang W, Hu Y, Xiao W, Huang Y, Feng Y, Pan Q, Wan R (2020) RNA demethylase ALKBH5 prevents pancreatic cancer progression by posttranscriptional activation of PER1 in an m6A-YTHDF2-dependent manner. Mol Cancer 19:91. https://doi.org/10.1186/s12943-020-01158-w
Tang B, Yang Y, Kang M, Wang Y, Wang Y, Bi Y, He S, Shimamoto F (2020) m(6)A demethylase ALKBH5 inhibits pancreatic cancer tumorigenesis by decreasing WIF-1 RNA methylation and mediating Wnt signaling. Mol Cancer 19:3. https://doi.org/10.1186/s12943-019-1128-6
Liu X, Feng M, Hao X, Gao Z, Wu Z, Wang Y, Du L, Wang C (2023) m6A methylation regulates hypoxia-induced pancreatic cancer glycolytic metabolism through ALKBH5-HDAC4-HIF1α positive feedback loop. Oncogene 42:2047–2060. https://doi.org/10.1038/s41388-023-02704-8
Hu Y, Tang J, Xu F, Chen J, Zeng Z, Han S, Wang F, Wang D, Huang M, Zhao Y, Huang Y, Zhuo W, Zhao G (2022) A reciprocal feedback between N6-methyladenosine reader YTHDF3 and lncRNA DICER1-AS1 promotes glycolysis of pancreatic cancer through inhibiting maturation of miR-5586-5p. J Exp Clin Cancer Res 41:69. https://doi.org/10.1186/s13046-022-02285-6
Yan F, Al-Kali A, Zhang Z, Liu J, Pang J, Zhao N, He C, Litzow MR, Liu S (2018) A dynamic N(6)-methyladenosine methylome regulates intrinsic and acquired resistance to tyrosine kinase inhibitors. Cell Res 28:1062–1076. https://doi.org/10.1038/s41422-018-0097-4
Visvanathan A, Patil V, Arora A, Hegde AS, Arivazhagan A, Santosh V, Somasundaram K (2018) Essential role of METTL3-mediated m(6)A modification in glioma stem-like cells maintenance and radioresistance. Oncogene 37:522–533. https://doi.org/10.1038/onc.2017.351
Zhou S, Bai ZL, Xia D, Zhao ZJ, Zhao R, Wang YY, Zhe H (2018) FTO regulates the chemo-radiotherapy resistance of cervical squamous cell carcinoma (CSCC) by targeting β-catenin through mRNA demethylation. Mol Carcinog 57:590–597. https://doi.org/10.1002/mc.22782
Yang S, Wei J, Cui YH, Park G, Shah P, Deng Y, Aplin AE, Lu Z, Hwang S, He C, He YY (2019) m(6)A mRNA demethylase FTO regulates melanoma tumorigenicity and response to anti-PD-1 blockade. Nat Commun 10:2782. https://doi.org/10.1038/s41467-019-10669-0
Ping Y, Huang J, Zhu J, Sun Z, Shang A, Chen C, Liu W, Li D (2023) Comprehensive analyses of molecular features, prognostic values, and regulatory functionalities of m(6)A-modified long non-coding RNAs in lung adenocarcinoma. Clin Epigenetics 15:60. https://doi.org/10.1186/s13148-023-01475-z
Zhu Y, Li J, Yang H, Yang X, Zhang Y, Yu X, Li Y, Chen G, Yang Z (2023) The potential role of m6A reader YTHDF1 as diagnostic biomarker and the signaling pathways in tumorigenesis and metastasis in pan-cancer. Cell Death Discov 9:34. https://doi.org/10.1038/s41420-023-01321-4
Huang W, Qi CB, Lv SW, Xie M, Feng YQ, Huang WH, Yuan BF (2016) Determination of DNA and RNA methylation in circulating tumor cells by mass spectrometry. Anal Chem 88:1378–1384. https://doi.org/10.1021/acs.analchem.5b03962
Ge L, Zhang N, Chen Z, Song J, Wu Y, Li Z, Chen F, Wu J, Li D, Li J, Wang C, Wang H, Wang J (2020) Level of N6-methyladenosine in peripheral blood RNA: a novel predictive biomarker for gastric cancer. Clin Chem 66:342–351. https://doi.org/10.1093/clinchem/hvz004
Oh J, Hwa C, Jang D, Shin S, Lee SJ, Kim J, Lee SE, Jung HR, Oh Y, Jang G, Kwon O, An JY, Cho SY (2022) Augmentation of the RNA m6A reader signature is associated with poor survival by enhancing cell proliferation and EMT across cancer types. Exp Mol Med 54:906–921. https://doi.org/10.1038/s12276-022-00795-z
Cho SH, Ha M, Cho YH, Ryu JH, Yang K, Lee KH, Han ME, Oh SO, Kim YH (2018) ALKBH5 gene is a novel biomarker that predicts the prognosis of pancreatic cancer: a retrospective multicohort study. Ann Hepatobiliary Pancreat Surg 22:305–309. https://doi.org/10.14701/ahbps.2018.22.4.305
Li W, Gao Y, Jin X, Wang H, Lan T, Wei M, Yan W, Wang G, Li Z, Zhao Z, Jiang X (2022) Comprehensive analysis of N6-methylandenosine regulators and m6A-related RNAs as prognosis factors in colorectal cancer. Mol Ther Nucleic Acids 27:598–610. https://doi.org/10.1016/j.omtn.2021.12.007
Qin S, Liu G, Jin H, Chen X, He J, Xiao J, Qin Y, Mao Y, Zhao L (2022) The comprehensive expression and functional analysis of m6A modification “readers” in hepatocellular carcinoma. Aging (Albany NY) 14:6269–6298. https://doi.org/10.18632/aging.204217
Yang J, Wu Z, Wu X, Chen S, Xia X, Zeng J (2022) Constructing and validating of m6a-related genes prognostic signature for stomach adenocarcinoma and immune infiltration: potential biomarkers for predicting the overall survival. Front Oncol 12:1050288. https://doi.org/10.3389/fonc.2022.1050288
Wang Q, Chen C, Ding Q, Zhao Y, Wang Z, Chen J, Jiang Z, Zhang Y, Xu G, Zhang J, Zhou J, Sun B, Zou X, Wang S (2020) METTL3-mediated m(6)A modification of HDGF mRNA promotes gastric cancer progression and has prognostic significance. Gut 69:1193–1205. https://doi.org/10.1136/gutjnl-2019-319639
Guan K, Liu X, Li J, Ding Y, Li J, Cui G, Cui X, Sun R (2020) Expression status and prognostic value of M6A-associated genes in gastric cancer. J Cancer 11:3027–3040. https://doi.org/10.7150/jca.40866
Liu J, Sun G, Pan S, Qin M, Ouyang R, Li Z, Huang J (2020) The Cancer Genome Atlas (TCGA) based m(6)A methylation-related genes predict prognosis in hepatocellular carcinoma. Bioengineered 11:759–768. https://doi.org/10.1080/21655979.2020.1787764
Wu Z, Huang X, Cai M, Huang P, Guan Z (2022) Novel necroptosis-related gene signature for predicting the prognosis of pancreatic adenocarcinoma. Aging (Albany NY) 14:869–891. https://doi.org/10.18632/aging.203846
Wang Y, Zhu GQ, Tian D, Zhou CW, Li N, Feng Y, Zeng MS (2022) Comprehensive analysis of tumor immune microenvironment and prognosis of m6A-related lncRNAs in gastric cancer. BMC Cancer 22:316. https://doi.org/10.1186/s12885-022-09377-8
Wang L, Zhang S, Li H, Xu Y, Wu Q, Shen J, Li T, Xu Y (2021) Quantification of m6A RNA methylation modulators pattern was a potential biomarker for prognosis and associated with tumor immune microenvironment of pancreatic adenocarcinoma. BMC Cancer 21:876. https://doi.org/10.1186/s12885-021-08550-9
Zhang Z, Zhang X (2021) Identification of m6A-related biomarkers associated with prognosis of colorectal cancer. Med Sci Monit 27:e932370. https://doi.org/10.12659/msm.932370
Gao Y, Wang H, Chen S, An R, Chu Y, Li G, Wang Y, Xie X, Zhang J (2022) Single-cell N(6)-methyladenosine regulator patterns guide intercellular communication of tumor microenvironment that contribute to colorectal cancer progression and immunotherapy. J Transl Med 20:197. https://doi.org/10.1186/s12967-022-03395-7
Bertero A, Brown S, Madrigal P, Osnato A, Ortmann D, Yiangou L, Kadiwala J, Hubner NC, de Los Mozos IR, Sadée C, Lenaerts AS, Nakanoh S, Grandy R, Farnell E, Ule J, Stunnenberg HG, Mendjan S, Vallier L (2018) The SMAD2/3 interactome reveals that TGFβ controls m(6)A mRNA methylation in pluripotency. Nature 555:256–259. https://doi.org/10.1038/nature25784
Xie LJ, Yang XT, Wang RL, Cheng HP, Li ZY, Liu L, Mao L, Wang M, Cheng L (2019) Identification of Flavin mononucleotide as a cell-active artificial N(6) -methyladenosine RNA demethylase. Angew Chem Int Ed Engl 58:5028–5032. https://doi.org/10.1002/anie.201900901
Zhu L, Zhu Y, Han S, Chen M, Song P, Dai D, Xu W, Jiang T, Feng L, Shin VY, Wang X, Jin H (2019) Impaired autophagic degradation of lncRNA ARHGAP5-AS1 promotes chemoresistance in gastric cancer. Cell Death Dis 10:383. https://doi.org/10.1038/s41419-019-1585-2
Xu K, Zhang Q, Chen M, Li B, Wang N, Li C, Gao Z, Zhang D, Yang L, Xu Z, Li X, Xu H (2022) N(6)-methyladenosine modification regulates imatinib resistance of gastrointestinal stromal tumor by enhancing the expression of multidrug transporter MRP1. Cancer Lett 530:85–99. https://doi.org/10.1016/j.canlet.2022.01.008
Taketo K, Konno M, Asai A, Koseki J, Toratani M, Satoh T, Doki Y, Mori M, Ishii H, Ogawa K (2018) The epitranscriptome m6A writer METTL3 promotes chemo- and radioresistance in pancreatic cancer cells. Int J Oncol 52:621–629. https://doi.org/10.3892/ijo.2017.4219
Wang Q, Chen C, Xu X, Shu C, Cao C, Wang Z, Fu Y, Xu L, Xu K, Xu J, Xia A, Wang B, Xu G, Zou X, Su R, Kang W, Xue Y, Mo R, Sun B, Wang S (2022) APAF1-binding long noncoding RNA promotes tumor growth and multidrug resistance in gastric cancer by blocking apoptosome assembly. Adv Sci (Weinh) 9:e2201889. https://doi.org/10.1002/advs.202201889
Zhang Y, Liu X, Wang Y, Lai S, Wang Z, Yang Y, Liu W, Wang H, Tang B (2022) The m(6)A demethylase ALKBH5-mediated upregulation of DDIT4-AS1 maintains pancreatic cancer stemness and suppresses chemosensitivity by activating the mTOR pathway. Mol Cancer 21:174. https://doi.org/10.1186/s12943-022-01647-0
Li J, Chen Z, Chen F, Xie G, Ling Y, Peng Y, Lin Y, Luo N, Chiang CM, Wang H (2020) Targeted mRNA demethylation using an engineered dCas13b-ALKBH5 fusion protein. Nucleic Acids Res 48:5684–5694. https://doi.org/10.1093/nar/gkaa269
Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, Gersbach CA (2015) Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol 33:510–517. https://doi.org/10.1038/nbt.3199
Dominguez AA, Lim WA, Qi LS (2016) Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation. Nat Rev Mol Cell Biol 17:5–15. https://doi.org/10.1038/nrm.2015.2
Su R, Dong L, Li C, Nachtergaele S, Wunderlich M, Qing Y, Deng X, Wang Y, Weng X, Hu C, Yu M, Skibbe J, Dai Q, Zou D, Wu T, Yu K, Weng H, Huang H, Ferchen K, Qin X, Zhang B, Qi J, Sasaki AT, Plas DR, Bradner JE, Wei M, Marcucci G, Jiang X, Mulloy JC, Jin J, He C, Chen J (2018) R-2HG exhibits anti-tumor activity by targeting FTO/m(6)A/MYC/CEBPA signaling. Cell 172:90-105.e23. https://doi.org/10.1016/j.cell.2017.11.031
Nishizawa Y, Konno M, Asai A, Koseki J, Kawamoto K, Miyoshi N, Takahashi H, Nishida N, Haraguchi N, Sakai D, Kudo T, Hata T, Matsuda C, Mizushima T, Satoh T, Doki Y, Mori M, Ishii H (2018) Oncogene c-Myc promotes epitranscriptome m(6)A reader YTHDF1 expression in colorectal cancer. Oncotarget 9:7476–7486. https://doi.org/10.18632/oncotarget.23554
You Q, Wang F, Du R, Pi J, Wang H, Huo Y, Liu J, Wang C, Yu J, Yang Y, Zhu L (2023) m(6) a reader YTHDF1-targeting engineered small extracellular vesicles for gastric cancer therapy via epigenetic and immune regulation. Adv Mater 35:e2204910. https://doi.org/10.1002/adma.202204910
Deng LJ, Deng WQ, Fan SR, Chen MF, Qi M, Lyu WY, Qi Q, Tiwari AK, Chen JX, Zhang DM, Chen ZS (2022) m6A modification: recent advances, anticancer targeted drug discovery and beyond. Mol Cancer 21:52. https://doi.org/10.1186/s12943-022-01510-2
Zhang X, Zhang S, Yan X, Shan Y, Liu L, Zhou J, Kuang Q, Li M, Long H, Lai W (2021) m6A regulator-mediated RNA methylation modification patterns are involved in immune microenvironment regulation of periodontitis. J Cell Mol Med 25:3634–3645. https://doi.org/10.1111/jcmm.16469
Acknowledgements
Not applicable.
Funding
This work was supported by the Fundamental Research Funds for the Central Universities (No. 2022ZFJH003), the National Key Research and Development Program of China (No. 2020YFE0204300, 2019YFC0840600 and 2019YFC0840609), and Zhejiang Provincial Natural Science Foundation (No. LY20H030006).
Author information
Authors and Affiliations
Contributions
D-H.Z., K-K.S., and L-L. L. designed the study and wrote the main manuscript text. X-X. O-Y. and Y-H.Z. prepared Figs. 1, 2 and Table 1. X-P.Y. and Z-H.L. prepared Fig. 3 and Table 2. S–S. A-N. and L-L.L. revised the final version. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, DH., Su, KK., Ou-Yang, XX. et al. Mechanisms and clinical landscape of N6-methyladenosine (m6A) RNA modification in gastrointestinal tract cancers. Mol Cell Biochem 479, 1553–1570 (2024). https://doi.org/10.1007/s11010-024-05040-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-024-05040-x