Abstract
The degradation of proteasomes or lysosomes is emerging as a principal determinant of programmed death ligand 1 (PDL1) expression, which affects the efficacy of immunotherapy in various malignancies. Intracellular cholesterol plays a central role in maintaining the expression of membrane receptors; however, the specific effect of cholesterol on PDL1 expression in cancer cells remains poorly understood. Cholesterol starvation and stimulation were used to modulate the cellular cholesterol levels. Immunohistochemistry and western blotting were used to analyze the protein levels in the samples and cells. Quantitative real-time PCR, co-immunoprecipitation, and confocal co-localization assays were used for mechanistic investigation. A xenograft tumor model was constructed to verify these results in vivo. Our results showed that cholesterol suppressed the ubiquitination and degradation of PDL1 in hepatocellular carcinoma (HCC) cells. Further mechanistic studies revealed that the autocrine motility factor receptor (AMFR) is an E3 ligase that mediated the ubiquitination and degradation of PDL1, which was regulated by the cholesterol/p38 mitogenic activated protein kinase axis. Moreover, lowering cholesterol levels using statins improved the efficacy of programmed death 1 (PD1) inhibition in vivo. Our findings indicate that cholesterol serves as a signal to inhibit AMFR-mediated ubiquitination and degradation of PDL1 and suggest that lowering cholesterol by statins may be a promising combination strategy to improve the efficiency of PD1 inhibition in HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is one of the leading causes of cancer-related deaths worldwide, especially in China [1]. Recent progress in targeted therapy and immunotherapy has provided more options for precise HCC treatment [2]. Programmed death ligand 1 (PDL1) is an essential immune checkpoint protein that binds to programmed death 1 (PD1) on T cells, leading to cancer immunosuppression [3]. PDL1 expression in cancer cells is regulated by multiple signaling pathways, including nuclear factor kappa-B (NFκB), mitogenic activated protein kinase (MAPK), mammalian target of rapamycin (mTOR), signal transducer and activator of transcription (STAT), and c-Myc [4, 5]. The PDL1 protein then undergoes degradation in proteasomes or lysosomes through multiple pathways, leading to increased effectiveness of cancer immunotherapy [6,7,8,9,10,11,12]. Therefore, identifying novel proteins that modulate PDL1 degradation may offer potential therapeutic targets for combination therapy with immune checkpoint inhibitors.
Metabolic reprogramming is a hallmark of malignancy, leaving distinct vulnerabilities while promoting cancer progression [13, 14]. Cholesterol accumulation has been found in various cancers and is reported to maintain membrane receptors, such as ErbB2, MET, and epidermal growth factor receptor (EGFR) [15,16,17,18,19]. In addition, a recent study indicated that the cholesterol content in T cells upregulates immune checkpoint proteins such as PD1, 2B4, T cell immunoglobulin domain and mucin domain-3 (TIM3), and lymphocyte activation gene-3 (LAG3) [20]. However, the mechanism by which dysregulated cholesterol metabolism supports PDL1 signaling, particularly its degradation in cancer cells, remains unclear. Accordingly, elucidating the relationship between cholesterol and PDL1 and clarifying the underlying mechanisms, may help in developing potential combination therapies against HCC.
The ubiquitin proteasome system (UPS) is indispensable for both PDL1 degradation and cholesterol homeostasis in eukaryotic cell [21, 22]. The UPS involves the conjugation of ubiquitin to substrates through an E1 activating enzyme, an E2 conjugating enzyme, and an E3 ubiquitin ligase, which provides specificity towards particular target proteins [23]. Degradation generally occurs following the ubiquitination of target proteins. E3 ligases involved in cholesterol biosynthesis include AMFR, membrane associated RING-CH-type finger 6 (MARCHF6), F-box and WD repeat domain containing 7 (FBW7), RING finger protein 139 (RNF139), and synoviolin 1 (SYVN1), most of which are also involved in the ubiquitination of proteins beyond cholesterol metabolism [24,25,26,27,28,29]. Therefore, these E3 ligases may act as adapters between cholesterol and PDL1.
In the present study, we first found close correlations between the expressions of sterol regulatory element-binding protein 2 (SREBP2), a key regulator of cholesterol biosynthesis, and PDL1, explored the effects of cholesterol on AMFR-mediated PDL1 ubiquitination and degradation, and revealed the underlying mechanism of this process. Finally, we investigated the anti-tumor effects of the combination of statin, a cholesterol-lowering agent, and an anti-PD1 antibody in HCC.
Materials and methods
Cell culture
The human HCC cell line MHCC97H was obtained from the Liver Cancer Institute, Fudan University, Shanghai, China. The human HCC cell line Huh7 and human acute T lymphocyte leukemia cell line Jurkat were purchased from the Cell Resources Center, Chinese Academy of Sciences, Shanghai, China. The murine HCC cell line, Hepa 1–6, was purchased from the American Type Culture Collection, Manassas, VA, USA. All cell lines were cultured in high-glucose dulbecco’s modified eagle medium (DMEM) supplemented with 10% foetal bovine serum (FBS) in an atmosphere containing 5% CO2 at 37 °C.
Tissue microarrays and clinical specimens
HCC tissue microarrays containing 31 paired tumor and non-tumor tissue samples were purchased from Outdo Biotech. HCC specimens were collected from 36 patients who had undergone surgical resection at our Institute. None of the patients had received any preoperative cancer treatment.
Immunohistochemistry
Immunohistochemical staining was performed on paraffin sections of HCC samples using a two-step protocol with the Novolink Polymer Detection System (Leica, Germany). After antigen retrieval, the sections were incubated with primary antibody overnight at 4 °C, followed by incubation with the secondary antibody at 37 °C for 30 min. All antibodies were diluted according to the instructions. The sections were stained with DAB, counterstained with hematoxylin, dehydrated in ethanol, mounted in dimethyl benzene, and placed under a coverslip. Staining intensity and percentage of positive cells were incorporated into a semi-quantitative score describing “0” (no expression), “1.5” (moderate expression) and “3” (strong expression).
Cholesterol starvation/stimulation
The cells were rinsed once with serum-free medium and incubated in cholesterol-depleting medium, DMEM containing 0.1% methyl-beta-cyclodextrin (MCD) and 0.5% lipoprotein-depleted serum, for 1–2 h. Cells were then transferred to cholesterol-depleting medium (starvation), or to cholesterol-depleting medium supplemented with 10 µg/mL cholesterol and incubated for another 1–2 h.
Western blotting
Cells were harvested in cell lysis buffer and boiled for 15 min. The protein concentration was quantified using BCA Protein Assay Kit. All proteins were separated using 8–12% SDS-PAGE and transferred to the NC membranes. After incubated with primary antibodies overnight at 4 °C, the membranes were washed and incubated with secondary antibodies for 1 h at room temperature. All antibodies were diluted according to the instructions. β-actin or GAPDH were used as an internal control.
Quantitative real-time PCR
Total RNA was isolated using the TRIzol Reagent following the manufacturer’s protocol. Reverse transcription was performed using PrimeScript RT Master Mix. Real-time PCR was performed on a QuantStudio 6 Flex Real-Time PCR System using the PrimeSTAR HS DNA Polymerase. The primers used for the amplification of human genes are listed in Supplementary Materials (Table S1).
Protein stability assay
Briefly, cells were treated with cycloheximide (CHX; 5 µg/mL) in cholesterol-depleting medium with/without cholesterol (10 µg/mL) at indicated intervals and then harvested in cell lysis buffer for western blotting analysis.
Co-immunoprecipitation
Cells were harvested in IP lysis buffer supplemented with phosphatase and protease inhibitors, pre-cleared with Protein A/G Magnetic Beads for 1 h, and then incubated overnight with anti-Myc Magnetic Beads at 4ºC. The beads were washed in IP lysis buffer and bound proteins were eluted for further analysis.
Confocal co-localization assay
Cells grown on coverslips were stimulated with cholesterol, fixed in 4% paraformaldehyde, permeabilized, and stained with primary antibodies, followed by incubation with Alexa 488- or 594-conjugated secondary antibodies. Nuclei were counterstained with DAPI. Images were captured using a confocal microscope.
Plasmid construction
All short hairpin RNA (shRNA) constructs were obtained from Sigma-Aldrich. The AMFR overexpression plasmid was obtained by cloning the C-terminal Flag-tagged AMFR into pCDH. All plasmids were verified by sequencing. The primer sequences are shown in the Supplementary Materials (Tables S2 and S3).
Cell counting kit-8 (CCK-8) assay
Liver cancer cells were plated in 96-well plates at a density of 2000 cells/well. Following vehicle control or drug treatment, cells were incubated with the culture medium containing 10% CCK-8 reagent (Beyotime, C0037) at 37 °C for 30 min. The absorbance was measured at 450 nm.
T cell-mediated tumor cell killing assay
Cancer cells were allowed to adhere to 6-well plates overnight and then incubated with lovastatin (1 µM) in the presence or absence of pre-activated Jurkat cells. The Jurkat cells were pre-activated overnight with 50 ng/ml Phorbol-12-myristate-13-acetate (PMA). The ratio of cancer cells to activated Jurkat cells was 1:3. T cells and cell debris were removed by washing with phosphate buffer solution (PBS), and live cancer cells were quantified using a CCK-8 assay.
Xenograft tumor model
Notably, 4-week-old male C57BL/6 mice were obtained from Shanghai Laboratory Animal Center, Chinese Academy of Sciences. All the mice were bred in laminar flow cabinets under clean conditions. The experimental protocol was approved by the Department of Experimental Animal Science of Wenzhou Medical University. For the combination therapy experiment, Hepa 1–6 cells (1 × 107) were subcutaneously inoculated into the right flank of mice. When palpable tumors were formed, mice were randomly assigned to four groups (n = 6 for each group), which received PBS (oral administration, every alternate day), lovastatin (30 mg/kg, oral administration, every alternate day), BE0146 (100 µg, intraperitoneal injection, every three days), or combination therapy for 2 weeks, and tumor samples were then extracted for further analysis.
Statistical analysis
Data are presented as mean ± SD or mean ± SEM as indicated in figure legends. Statistical differences were determined using the Student’s t test or one-way ANOVA. p < 0.05 were considered statistically significant.
Results
Cholesterol biosynthesis correlates with PDL1 in HCC
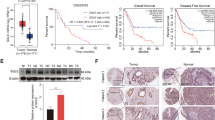
Cholesterol accumulation in cancer cells maintains membrane receptors in various cancers [18, 19]. Cellular cholesterol levels reflect the dynamic balance between biosynthesis, uptake, export, and esterification, among which cholesterol biosynthesis seems to play a central role in HCC progression [30, 31]. To reveal the potential role of cholesterol biosynthesis in PDL1 degradation in HCC, we first examined the protein levels of PDL1 and SREBP2, a master regulator of cholesterol biosynthesis, using a tissue microarray containing 31 paired tumor and non-tumor HCC tissues. Notably, HCC tumor tissues had significantly higher protein levels of both SREBP2 and PDL1, compared with that of the non-tumor tissues (Fig. 1A–C). To further explore the relationship between PDL1 and cholesterol biosynthesis, we determined the correlations between PDL1 and SREBP2 or HMG-CoA reductase (HMGCR), the principal rate-limiting enzyme of the cholesterol biosynthesis pathway, in another independent cohort of HCC specimens from our hospital. The data showed that PDL1 was significantly correlated with both SREBP2 and HMGCR in HCC tissues (Fig. 1D and E). Collectively, these findings indicate that cholesterol biosynthesis is associated with PDL1 expression in HCC.
Cholesterol biosynthesis correlates with PDL1 in HCC. A Representative IHC images of SREBP2 (upper) and PDL1 (lower) in paired tumor and non-tumor tissues of HCC tissue microarray. B and C IHC analyses of protein levels of SREBP2 (B) and PDL1 (C) in HCC tissue microarray. D and E IHC analyses and correlation of protein levels of PDL1 with SREBP2 (D) and HMGCR (E) in 36 HCC specimens. Linear regression coefficient and statistical significance were indicated. For B and C, data were presented as mean ± SD, and statistical significance was determined by student's t test. *, p < 0.05; ****, p < 0.0001. For D and E, r and p value were calculated by Pearson correlation
Cholesterol upregulates PDL1 in a post-transcriptional manner
Proteins are mainly degraded by two systems in eukaryotic cells, the UPS and the autophagic-lysosomal system, both of which are involved in PDL1 degradation [32]. To determine the role of cholesterol in regulating PDL1 expression, we determined the effects of modulating cellular cholesterol levels on PDL1 protein levels in MHCC97H and Huh7 cells. HCC cells were first fasted in cholesterol-depleting medium and then treated with cholesterol at the indicated concentrations. Data showed that cholesterol accumulation resulting from cholesterol supplement increased PDL1 protein levels in MHCC97H and Huh7 cells (Fig. 2A). Furthermore, we used lovastatin, a classic cholesterol-lowering agent, to validate the above findings. Conversely, lovastatin markedly decreased PDL1 protein levels in MHCC97H and Huh7 cells (Fig. 2B).
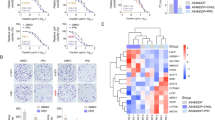
Cholesterol upregulates PDL1 post-transcriptionally. A and B Western blotting analyses of PDL1 in MHCC97H and Huh7 cells treated with cholesterol (A) or lovastatin (B). C and d qRT-PCR analyses of PDL1 in MHCC97H and Huh7 cells treated with cholesterol (C) or lovastatin (D). E Western blotting analyses of exogenous Myc-PDL1 in HEK293T cells after cholesterol or lovastatin stimulation. Myc-PDL1 were pre-transfected into HEK293T cells for at least 24 hours. For cholesterol stimulation, cells were pre-starved with cholesterol-depleting medium for 2 hours and stimulated with indicated concentrations of cholesterol for another 2 hours. For lovastatin treatment, cells were incubated with indicated concentrations of lovastatin for 48 hours. Cell lysates were then collected for Western blotting and qRT-PCR analyses. Data were presented as mean ± SD from triplicate experiments (C and D), and statistical significance was determined by one-way ANOVA. ns non-significant
Next, we explored the mechanism by which cholesterol induces an increase in PDL1 protein levels. PDL1 mRNA levels were determined in MHCC97H and Huh7 cells after cholesterol supplementation or lovastatin treatment (Fig. 2C and D). The results indicated that neither cholesterol supplementation nor lovastatin treatment affected the PDL1 mRNA levels, suggesting that the increase or decrease in PDL1 levels after cholesterol/lovastatin treatment occurred at the post-transcriptional level. Similarly, the effect of cholesterol and lovastatin on exogenous Myc-PDL1 was detected in HEK293T cells. Our results showed that exogenous Myc-PDL1 was also regulated by cellular cholesterol manipulation (Fig. 2E). Collectively, these findings demonstrate that cellular cholesterol upregulates PDL1 in a post-transcriptional manner.
Cholesterol suppresses ubiquitination and degradation of PDL1
To further clarify the mechanism underlying cholesterol-mediated PDL1 upregulation, we determined whether cholesterol increased the stability of PDL1. CHX, a protein synthesis inhibitor, was used to delineate the degradative curves of PDL1 in MHCC97H and Huh7 cells under cholesterol starvation or stimulation. Strikingly, cholesterol stabilized PDL1 in both MHCC97H and Huh7 cells, indicating that cellular cholesterol upregulates PDL1 by inhibiting its degradation (Fig. 3A and B).
Cholesterol suppresses ubiquitination and degradation of PDL1. A and B Protein stability analyses of PDL1 in MHCC97H (A) and Huh7 (B) cells with or without cholesterol stimulation. Cells were treated with 5 μg/mL cycloheximide (CHX) at indicated intervals and were analyzed by Western blotting analyses. Representative images were shown in the left, and the quantifications of Western blotting bands were shown in the right, the relative intensity of protein bands in control group was set as 100%. C Western blotting analyses of PDL1 in MHCC97H (upper) and Huh7 (lower) cells treated with cholesterol-depleting medium containing 20 μM MG132 (a proteasome inhibitor) or HCQ (an autophagy inhibitor) for 2 hours. Methyl-beta-cyclodextrin (MCD) was used to deplete cholesterol. D Competitive CoIP assays of GFP-LC3 with Myc-PDL1 in HEK293T cells. The plasmids were pre-transfected into HEK293T cells for at least 24 hours, cells were then stimulated by 3 μg/mL cholesterol before IP analysis. E and F Exogenous ubiquitination analyses of PDL1 in HEK293T cells treated with cholesterol or lovastatin for indicated times. HA-Ubiquitin and Myc-PDL1 were pre-transfected into HEK293T cells for at least 24 hours. Data were presented as mean ± SD from triplicate experiments (A and B), and were representatives of three independent experiments. Statistical significance was determined by one-way ANOVA. **, p < 0.01; ****, p < 0.0001
Therefore, we applied MG132 (a proteasome inhibitor) and hydroxychloroquine (HCQ, an autophagy inhibitor) in cholesterol-depleted MHCC97H and Huh7 cells. The results showed that MG132 rescued PDL1 degradation, indicating the involvement of the UPS (Fig. 3C). We then determined the interaction of microtubule associated protein 1 light chain 3 (MAP1LC3 or LC3, an autophagy marker) with PDL1 in HEK293T cells treated with cholesterol and found that the LC3-PDL1 interaction was not weakened by cholesterol supplementation, which is consistent with our previous findings (Fig. 3D). Next, we performed exogenous ubiquitination analysis of PDL1 in HEK293T cells treated with cholesterol or lovastatin. Surprisingly, cholesterol decreased PDL1 ubiquitination, whereas lovastatin increased it (Fig. 3E and F). In summary, these observations support the hypothesis that cholesterol upregulates PDL1 by suppressing its ubiquitination and degradation.
AMFR induces ubiquitination and degradation of PDL1 after cholesterol starvation
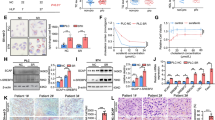
AMFR is the most extensively studied E3 ligase involved in cholesterol biosynthesis, and is regulated by sterols, including cholesterol [33, 34]. A recent study has revealed its role in targeting cholesterol-irrelevant proteins for degradation [25]. Therefore, we first explored the role of AMFR in PDL1 degradation by constructing AMFR-knockdown and -overexpressing HCC cells. The results showed that AMFR knockdown markedly upregulated PDL1 protein levels in both MHCC97H and Huh7 cells, whereas AMFR overexpression led to the opposite effect (Fig. 4A and B). Furthermore, we determined the effects of cholesterol and lovastatin on AMFR expression in MHCC97H and Huh7 cells. The results showed that AMFR expression was conversely regulated by cellular cholesterol, supporting its role in mediating starvation-induced PDL1 degradation (Fig. 4C and D).
AMFR induces ubiquitination and degradation of PDL1. A and B Western blotting analyses of PDL1 in MHCC97H and Huh7 cells after AMFR knockdown (A) or overexpression (B). C and D Western blotting analyses of AMFR in MHCC97H and Huh7 cells treated with cholesterol (C) or lovastatin (D). E and F CoIP (E) and competitive CoIP (F) assays of Flag-AMFR with Myc-PDL1 in HEK293T cells. Flag-AMFR and Myc-PDL1 were pre-transfected into HEK293T cells for at least 24 hours, cells were then incubated with DMEM containing 10% FBS (E) or stimulated by 3 μg/mL cholesterol (F) before IP analysis. G Confocal co-localization assay of AMFR (green) with PDL1 (red) in MHCC97H cells under cholesterol-depleted/stimulated conditions, scale bars represent 10 μm. Cells were starved with cholesterol-depleting medium for 2 hours before confocal microscopy. H and I Protein stability assays of PDL1 in MHCC97H cells with AMFR knockdown or overexpression under cholesterol starvation. Representative images were shown in H, and the quantifications of Western blotting bands were shown in I, the relative intensity of protein bands in control group was set as 100%. Data were presented as mean ± SD from triplicate experiments (I), and were representatives of three independent experiments. Statistical significance was determined by one-way ANOVA. ***, p < 0.001; ****, p < 0.0001
To further clarify the role of AMFR in the ubiquitination and degradation of PDL1, the interaction between AMFR and PDL1 was examined in HEK293T cells treated with or without cholesterol. Interestingly, we found that AMFR interacted with PDL1 and that cholesterol decreased this interaction (Fig. 4E and F). Next, we conducted a confocal colocalization assay of AMFR and PDL1 in MHCC97H cells under cholesterol starvation or stimulation. Our results showed that AMFR (green spots) co-localized with PDL1 (red spots) in cholesterol-starved MHCC97H cells (yellow spots), whereas no co-localization was observed in cholesterol-treated cells (Fig. 4G). Consistent with these findings, increased or decreased PDL1 protein stability was observed in AMFR-knockdown and -overexpressing MHCC97H cells, respectively (Fig. 4H and I). In conclusion, these findings revealed that AMFR is required for the ubiquitination and degradation of PDL1 in cholesterol-depleted cells.
Cholesterol inhibits AMFR-dependent degradation of PDL1 by activating p38 MAPK pathway
Next, we investigated the mechanisms underlying the inhibition of AMFR-dependent ubiquitination and degradation of PDL1 by cellular cholesterol. p38 MAPK phosphorylates AMFR and decreases AMFR protein expression [35, 36]. Therefore, we first determined the effects of cholesterol and lovastatin on the p38 MAPK pathway. Our results showed that cholesterol activated the p38 MAPK pathway and decreased AMFR protein expression, whereas lovastatin had the opposite effect (Fig. 5A and B). These phenomena indicate that the p38 MAPK pathway may be responsible for cholesterol-induced AMFR reduction. We then used SB202190, a p38 MAPK inhibitor, to investigate the direct effect of blocking the p38 MAPK pathway on AMFR and PDL1 expression in MHCC97H cells. The inhibition of p38 MAPK by SB202190 reduced PDL1 protein levels and increased AMFR protein expression, which is consistent with the above findings (Fig. 5C). Subsequently, increased co-localization of AMFR and PDL1 (yellow spots) was observed in MHCC97H cells following SB202190 treatment (Fig. 5D). Similarly, p38 MAPK inhibition enhanced the interaction between AMFR and PDL1 (Fig. 5E). Finally, we evaluated the effect of SB202190 on PDL1 stability in MHCC97H cells. The results showed that the inhibition of p38 MAPK remarkably destabilized PDL1 in MHCC97H cells (Fig. 5F and G). Taken together, these findings highlight that cholesterol inhibits the AMFR-dependent degradation of PDL1 by activating the p38 MAPK pathway.
Cholesterol inhibits AMFR-dependent degradation of PDL1 by activating p38 MAPK pathway. A and B Western blotting analyses of p38 MAPK pathway in MHCC97H and Huh7 cells incubated with indicated concentrations of cholesterol (A) for 2 hours after starvation or with lovastatin (B) for 48 hours. C Western blotting analyses of PDL1 in MHCC97H and Huh7 cells incubated with indicated concentrations of SB202190 (a P38 inhibitor) for 48 hours. D Confocal co-localization assay of AMFR (green) with PDL1 (red) in MHCC97H cells after P38 inhibition, scale bars represent 10 μm. E CoIP assays of Flag-AMFR with Myc-PDL1 in HEK293T cells. The plasmids were pre-transfected into HEK293T cells for at least 24 hours, cells were then incubated with 3 μM SB202190 (a P38 inhibitor) before IP analyses. F and G Protein stability assay of PDL1 in MHCC97H cells after P38 inhibition. Representative images were shown in (F), and the quantifications of Western blotting bands were shown in (G), the relative intensity of protein bands in control group was set as 100%. Data were presented as mean ± SD from triplicate experiments (G), and were representatives of three independent experiments. Statistical significance was determined by one-way ANOVA. ***, p < 0.001; ****, p < 0.0001
Lowering cholesterol enhances the anti-tumor effect of PD1 inhibition
Finally, we investigated the effect of cholesterol lowering on the efficacy of PD1 inhibition in HCC. We first determined the effect of lovastatin on T cell cytotoxicity in MHCC97H, Huh7, and Hepa 1–6 cells. The results showed that lovastatin enhanced the killing effect of T cells on these cells (Fig. 6A–C). We then investigated combination therapy with lovastatin and PD1 inhibition in an HCC xenograft model. Briefly, Hepa 1–6 cells were injected subcutaneously into the right flank of immune-competent mice one week before treatment, and mice were sacrificed after two weeks of treatment with lovastatin, BE0146 (an anti-PD1 antibody), or a combination of both, and the tumors were extracted. The results showed that lovastatin enhanced the anti-tumor effect of PD1 inhibition in vivo, which was consistent with previous findings in vitro (Fig. 6D–F). To further clarify the underlying mechanism of the combination therapy, we performed immunohistochemistry analysis of PDL1 in Hepa 1-6-derived xenograft tumors after combination therapy. We found that lovastatin, with or without PD1 inhibition, reduced PDL1 protein levels in xenograft tumors (Fig. 6G). Moreover, cholesterol assays of xenograft tumors indicated that lovastatin reduced cholesterol levels in tumors in the presence or absence of PD1 inhibition (Fig. 6H). Taken together, our data indicated that lowering cholesterol with lovastatin enhanced the anti-tumor effect of PD1 inhibition.
The combination of lovastatin and PD1 blockade suppresses HCC growth. A–C Effects of lovastatin on in vitro cytotoxicity of T cells in MHCC97H (A), Huh7 (B), and Hepa 1-6 (C) cells. Cells were incubated with 1 μM lovastatin in the absence or presence of pre-activated Jurkat cells for 48 hours, then Jurkat cells were washed out before relative viabilities were determined by CCK-8 assays. D Photograph of the subcutaneous tumors in Hepa 1-6-derived xenograft tumor model after combination therapy of lovastatin (30 mg/kg) and BE0146 (an anti-PD1 antibody, 100 μg) for 2 weeks (n = 6). E Growth curves of the Hepa 1-6-derived xenograft tumors after indicated treatments. F Volumes of the Hepa 1-6-derived xenograft tumors at day 15. G IHC analyses of Hepa 1-6-derived xenograft tumors after combination therapy. Left, representative images of IHC staining of PDL1 in each group, scale bars represent 100 μm; right, relative quantifications of PDL1 staining. H Cholesterol assay of Hepa 1-6-derived xenograft tumors after combination therapy. Experiments were performed with at least three biological replicates and were representatives of at least three independent experiments. Statistical significance was determined by one-way ANOVA. Data were presented as mean ± SD (A-C) or mean ± SEM (E–H). *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001
Discussion
Metabolic reprogramming is a distinguishing hallmark of cancer that promotes the progression of various malignancies [13]. Cholesterol accumulation has been observed in various cancers and promotes various transmembrane signaling pathways by maintaining receptors [15,16,17,18,19]. PDL1/PD1 is perhaps the most well-known immune checkpoint pathway that induces immunosuppression in cancers. A recent study indicated that cholesterol accumulation leads to T-cell exhaustion by upregulating various immune checkpoint proteins, including PD120. However, the mechanism by which dysregulated cholesterol metabolism supports PDL1 signaling, particularly its degradation in cancer cells, remains unclear. In the present study, we found that cholesterol suppresses the ubiquitination and degradation of PDL1 in HCC cells. Further investigation revealed that AMFR, a cholesterol-related E3 ligase, mediates the ubiquitination and degradation of PDL1 in response to cholesterol accumulation through the p38 MAPK pathway and that lowering cholesterol by statins improved the efficacy of PD1 inhibition in HCC (Fig. 7).
Cholesterol suppressed the ubiquitination and degradation of PDL1. When intracellular cholesterol is sufficient, phosphorylated p38 suppresses the ubiquitination and the following degradation of PDL1 by inhibiting AMFR, which is the E3 ligase of PDL1 ubiquitination. Lowering cholesterol by statins upregulates AMFR and promotes PDL1 ubiquitination and degradation by inhibiting p38, thus enhancing the efficacy of anti-PD1 antibody
These findings are crucial supplements to the current understanding of cholesterol metabolism and immune checkpoint pathways, indicating that cholesterol in both tumors and the surrounding microenvironment can work synergistically to support immune checkpoint pathways, leading to cancer immunosuppression. Therefore, we deliberate about a possible master regulator that orchestrates cholesterol metabolic reprogramming in HCC. Since metabolic reprogramming is considered to result from genetic alterations in oncogenes, one possibility is that genetic alterations in oncogenes, especially PDL1, could drive cholesterol alterations. In fact, PDL1 signaling in cancer cells has been reported to promote cancer progression independent of PD1 binding [37, 38]. Another study revealed that direct PDL1 blockade dampens glycolysis by inhibiting mTOR activity [39]. Given the significant role of tumor mutation burden of PDL1 in predicting the response to PDL1/PD1 inhibition, the role of PDL1 in cancer cells deserves further investigation.
Cholesterol metabolism is known to be subtly controlled by the UPS [22]. The key E3 ligases involved in cholesterol metabolism include AMFR, MARCHF6, FBW7, RNF139, and SYVN1which respond to cholesterol alterations and restore cholesterol homeostasis by targeting the key players in this process. Recent studies have revealed the roles of these E3 ligases in cancer progression, which are accomplished by targeting cholesterol-irrelevant proteins for degradation [25,26,27,28,29]. Therefore, although we identified AMFR as the main adaptor for PDL1 ubiquitination and degradation, other E3 ligases may exist which act separately from the cholesterol metabolism that play roles in this process, which require further exploration.
In conclusion, our study demonstrates a cholesterol-dependent pathway of PDL1 ubiquitination and degradation, and identifies statins as a potential combination strategy to increase the efficacy of PD1 inhibition in HCC.
Data availability
No datasets were generated or analysed during the current study.
References
Maomao C et al (2022) Current cancer burden in China: epidemiology, etiology, and prevention. Cancer Biol Med 19(8):1121–1138
Yan T et al (2022) The advanced development of molecular targeted therapy for hepatocellular carcinoma. Cancer Biol Med 19(6):802–817
Topalian SL, Drake CG, Pardoll DM (2012) Targeting the PD-1/B7-H1(PD-L1) pathway to activate anti-tumor immunity. Curr Opin Immunol 24:207–212
Ritprajak P, Azuma M (2015) Intrinsic and extrinsic control of expression of the immunoregulatory molecule PD-L1 in epithelial cells and squamous cell carcinoma. Oral Oncol 51:221–228
Casey SC et al (2016) MYC regulates the antitumor immune response through CD47 and PD-L1. Science 352:227–231
Li CW et al (2016) Glycosylation and stabilization of programmed death ligand-1 suppresses T-cell activity. Nat Commun 7:12632
Mezzadra R et al (2017) Identification of CMTM6 and CMTM4 as PD-L1 protein regulators. Nature 549:106–110
Zhang J et al (2018) Cyclin D-CDK4 kinase destabilizes PD-L1 via cullin 3-SPOP to control cancer immune surveillance. Nature 553:91–95
Deng L et al (2019) Inhibition of mTOR complex 1/p70 S6 kinase signaling elevates PD-L1 levels in human cancer cells through enhancing protein stabilization accompanied with enhanced β-TrCP degradation. Oncogene 38:6270–6282
Wang H et al (2019) HIP1R targets PD-L1 to lysosomal degradation to alter T cell-mediated cytotoxicity. Nat Chem Biol 15:42–50
Zhou S et al (2019) Neddylation inhibition upregulates PD-L1 expression and enhances the efficacy of immune checkpoint blockade in glioblastoma. Int J Cancer 145:763–774
Romero Y, Wise R, Zolkiewska A (2020) Proteolytic processing of PD-L1 by ADAM proteases in breast cancer cells. Cancer Immunol Immunother 69:43–55
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674
Faubert B, Solmonson A, DeBerardinis RJ (2020) Metabolic reprogramming and cancer progression. Science 10:368
Krycer JR, Brown AJ (2013) Cholesterol accumulation in prostate cancer: a classic observation from a modern perspective. Biochim Biophys Acta 1835:219–229
Munir MT et al (2020) VD(3) and LXR agonist (T0901317) combination demonstrated greater potency in inhibiting cholesterol accumulation and inducing apoptosis via ABCA1-CHOP-BCL-2 cascade in MCF-7 breast cancer cells. Mol Biol Rep 47:7771–7782
Jun SY et al (2021) Reduction of squalene epoxidase by cholesterol accumulation accelerates colorectal cancer progression and metastasis. Gastroenterology 160:1194–1207e1128
Zhang J et al (2019) Cholesterol content in cell membrane maintains surface levels of ErbB2 and confers a therapeutic vulnerability in ErbB2-positive breast cancer. Cell Commun Signal 17:15
Shao WQ et al (2020) Liver x receptor agonism sensitizes a subset of hepatocellular carcinoma to sorafenib by dual-inhibiting MET and EGFR. Neoplasia 22:1–9
Ma XZ et al (2019) Cholesterol induces CD8(+) T cell exhaustion in the tumor microenvironment. Cell Metab 30:143–156e145
Gou Q et al (2020) PD-L1 degradation pathway and immunotherapy for cancer. Cell Death Dis 11:955
Sharpe LJ, Cook EC, Zelcer N, Brown AJ (2014) The UPS and downs of cholesterol homeostasis. Trends Biochem Sci 39:527–535
A H, H, H., S, E., Ciechanover A (1983) Components of ubiquitin-protein ligase system. Resolution, affinity purification, and role in protein breakdown. J Biol Chem 258:8206–8214
Jiang W, Song BL (2014) Ubiquitin ligases in cholesterol metabolism. Diabetes Metab J 38:171–180
Tsai YC et al (2007) The ubiquitin ligase gp78 promotes sarcoma metastasis by targeting KAI1 for degradation. Nat Med 13:1504–1509
Zavacki AM et al (2009) The E3 ubiquitin ligase TEB4 mediates degradation of type 2 iodothyronine deiodinase. Mol Cell Biol 29:5339–5347
Huang LY et al (2018) SCF(FBW7)-mediated degradation of Brg1 suppresses gastric cancer metastasis. Nat Commun 9:3569
Lin PH, Wm L, Chau LY (2013) TRC8 suppresses tumorigenesis through targeting heme oxygenase-1 for ubiquitination and degradation. Oncogene 32:2325–2334
Guo X et al (2021) HRD1 inhibits fatty acid oxidation and tumorigenesis by ubiquitinating CPT2 in triple-negative breast cancer. Mol Oncol 15:642–656
Luo J, Yang HY, Song BL (2019) Mechanisms and regulation of cholesterol homeostasis. Nat Rev Mol Cell Biol 21:225–245
Che L et al (2020) Cholesterol biosynthesis supports the growth of hepatocarcinoma lesions depleted of fatty acid synthase in mice and humans. Gut 69:177–186
Zhao J, Zhai B, Gygi SP, Goldberg AL (2015) mTOR inhibition activates overall protein degradation by the ubiquitin proteasome system as well as by autophagy. Proc Natl Acad Sci U S A 112:15790–15797
Song BL, Sever N, DeBose-Boyd RA (2005) Gp78, a membrane-anchored ubiquitin ligase, associates with Insig-1 and couples sterol-regulated ubiquitination to degradation of HMG CoA reductase. Mol Cell 19:829–840
Lee JN, Song B, DeBose-Boyd RA, Ye J (2006) Sterol-regulated degradation of Insig-1 mediated by the membrane-bound ubiquitin ligase gp78. J Biol Chem 281:39308–39315
Li L et al (2015) p38 MAP kinase-dependent phosphorylation of the Gp78 E3 ubiquitin ligase controls ER-mitochondria association and mitochondria motility. Mol Biol Cell 26:3828–3840
Ohtsuki Y et al (2019) Inhibition of cytochrome P450 3A protein degradation and subsequent increase in enzymatic activity through p38 MAPK activation by acetaminophen and salicylate derivatives. Biochem Biophys Res Commun 509:287–293
Gao H, Zhang J, Ren X (2019) PD-L1 regulates tumorigenesis and autophagy of ovarian cancer by activating mTORC signaling. Biosci Rep 39:BSR20191041
Escors D et al (2018) The intracellular signalosome of PD-L1 in cancer cells. Signal Transduct Target Ther 3:26
Chang CH et al (2015) Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162:1229–1241
Acknowledgements
Not applicable.
Funding
This research was supported by Zhejiang Provincial Natural Science Foundation of China under Grant. No.LY21H160009; the National Natural Science Foundation of China (grant number 82072696, 82000605, 81972737); Zhejiang Province Traditional Chinese Medicine Science and Technology Project. No. 2024ZL1008; Wenzhou City Basic Scientific Research Project. No. Y2023896; Ruian City Basic Scientific Research Project. No. MS2023054; Zhejiang Provincial Medical and Health Science and Technology Project (grant number 2024KY1632).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Wei-Qing Shao, Yi-Tong Li, Xu Zhou, Sheng-Guo Zhang, Ming-Hao Fan, Dong Zhang, Zhen-Mei Chen, Chen-He Yi, Sheng-Hao Wang, Wen-Wei Zhu, Ming Lu and Ji-Song Chen. The first draft of the manuscript was written by Wei-Qing Shao, Yi-Tong Li, Xu Zhou, Jing Lin and Yu Zhou, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. This study was approved by the Ethics Committee of Ruian People’s Hospital (Grant No. YJ2024089). All animal experiments were approved by the Tab of Animal Experimental Ethical Inspection of Laboratory Animal Centre, Wenzhou Medical University (Grant No. wydw2020-0703).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent to publish
The authors affirm that human research participants provided informed consent for publication of the images in Fig. 1.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shao, WQ., Li, YT., Zhou, X. et al. Cholesterol suppresses AMFR-mediated PDL1 ubiquitination and degradation in HCC. Mol Cell Biochem (2024). https://doi.org/10.1007/s11010-024-05106-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11010-024-05106-w